Abstract
The outbreak of COVID-19 has raised interest in the kinin–kallikrein system. Viral blockade of the angiotensin-converting enzyme 2 impedes degradation of the active kinin des-Arg(9)-bradykinin, which thus increasingly activates bradykinin receptors known to promote inflammation, cough, and edema—symptoms that are commonly observed in COVID-19. However, lean and reliable investigation of the postulated alterations is currently hindered by non-specific peptide adsorption, lacking sensitivity, and cross-reactivity of applicable assays. Here, an LC–MS/MS method was established to determine the following kinins in respiratory lavage fluids: kallidin, bradykinin, des-Arg(10)-kallidin, des-Arg(9)-bradykinin, bradykinin 1-7, bradykinin 2-9 and bradykinin 1-5. This method was fully validated according to regulatory bioanalytical guidelines of the European Medicine Agency and the US Food and Drug Administration and has a broad calibration curve range (up to a factor of 103), encompassing low quantification limits of 4.4–22.8 pg/mL (depending on the individual kinin). The application of the developed LC–MS/MS method to nasal lavage fluid allowed for the rapid (~ 2 h), comprehensive and low-volume (100 µL) determination of kinins. Hence, this novel assay may support current efforts to investigate the pathophysiology of COVID-19, but can also be extended to other diseases.
Similar content being viewed by others
Introduction
In March 2020, the World Health Organization declared a pandemic of coronavirus disease 2019 (COVID-19) due to alarming levels of spread and severity. COVID-19 is caused by infection with the novel severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2). Hospitalized patients commonly present with symptoms of fever, cough, and dyspnea and can develop pulmonary edema in early disease1. It has been postulated that these symptoms are in connection with a dysregulated kinin-kallikrein-system (KKS)2,3,4,5. As yet, concentrations of peptides within the KKS have not been reported in COVID-19 patients and their role remains unclear.
Activation of the KKS, i.e. by tissue and plasma kallikrein, leads to the formation of the kinins bradykinin and kallidin (lys-bradykinin), both potent activators of bradykinin-2 receptors on endothelial cells6. The activation of these receptors promotes vasodilation, inflammation, and capillary leakage leading to edema (Fig. 1)6,7. Bradykinin and kallidin are cleaved by carboxypeptidase N and M into des-Arg(9)-bradykinin and des-Arg(10)-kallidin, respectively, which are ligands for the bradykinin-1 receptor, as demonstrated in in vitro experiments using human tissues6,8. The in vivo activation of bradykinin-1 receptors in animals was shown to mediate inflammation, bronchoconstriction, and extravasation, which causes (pulmonary) edema9,10. The receptor is furthermore upregulated during inflammation, thus providing increased receptor binding sites for des-Arg(9)-bradykinin and des-Arg(10)-kallidin6,7,11. While des-Arg(10)-kallidin is further cleaved into des-Arg(9)-bradykinin, the latter is mainly degraded by the angiotensin-converting enzyme (ACE) 212,13. ACE 2 has been identified as the binding site of SARS-CoV-2, enabling it to enter cells4,14. It is thus assumed that cleavage of active des-Arg(9)-bradykinin into the inactive bradykinin 1-7 is considerably reduced during SARS-CoV-2 infection. In this context, viral attenuation of ACE 2 activity contributed to the pathogenesis of lung inflammation that was concomitant with increased bradykinin-1 receptor expression12. Therefore, the KKS is suggested to be involved in the pathogenesis of COVID-19 via the viral blockade of ACE 2, leading to elevated active des-Arg(9)-bradykinin levels (Fig. 1)4,15. Monitoring of seven kinin peptides (bradykinin, kallidin, des-Arg(9)-bradykinin, des-Arg(10)-kallidin, bradykinin 1-7, bradykinin 1-5 and bradykinin 2-9) may provide insights into the hypothetically altered kinin metabolism during SARS-CoV-2 infection.
Because ACE 2 is highly expressed in the nasal epithelium, the nose is presumed to represent the main entry point of SARS-CoV-2 prior to further spread within the host16. Therefore, saline lavage fluids from the respiratory tract would likely be the most suitable matrix to investigate alterations in the KKS peptide levels, as these fluids originate from the main area of viral infection and clinical symptoms in mild to severe courses (e.g. cough, nasal congestion, pulmonary inflammation and edema)5,17. While sampling of epithelial lining fluid in the lower airways by bronchoalveolar lavage is limited by its invasiveness requiring anesthesia, the use of nasal lavage fluid (NLF) offers the advantage of non-invasive, cheap and easy sampling18. In the NLF of healthy volunteers, kinin levels were reported to be typically less than 100 pg/mL by immunometric detection, however, no quantitative differentiation of the kinins was possible due to the underlying analytical technique applied19,20. The susceptibility to cross-reactivity with similar structures—which is the case for kinin peptides (Fig. 1)—represents the main disadvantage of immunoassays. Nevertheless, highly sensitive immunoassay-based quantification methods that can distinguish kinin peptides have been developed using plasma or tissues, but they require extensive effort for sample purification, including (multiple) solid-phase extractions (SPEs), liquid–liquid extraction, and chromatographic separation prior to immunoassay21,22.
Liquid chromatography coupled to tandem mass spectrometry (LC–MS/MS) represents a rational choice to overcome immunoassay-related limitations. Nevertheless, to date, no LC–MS/MS method has been reported that can comprehensively determine kinin peptides in respiratory saline lavage fluids. Validated LC–MS/MS methods are available for the determination of single kinin peptides (bradykinin, des-Arg-(9)-bradykinin and bradykinin 1-5) from plasma, serum, or whole blood and lack sensitivity in the desired low pg/mL range23,24,25. Owing to the substantial dilution of epithelial lining fluid by a factor of 60–120 during lavage, a suitable assay must be highly sensitive26. The reliable analysis of low levels of endogenous peptides in diluted matrix, whereby the peptides are often characterized by non-specific peptide adsorption, requires extensive method development27,28. For that, design of experiments (DoE) has proven its usefulness as a lean tool for method development of multifactor-dependent settings and contributes to signal increase in LC–MS/MS29,30.
Therefore, this study aimed to develop and validate a novel LC–MS/MS method characterized by a broad calibration curve range to comprehensively and sensitively determine KKS peptides (bradykinin, kallidin, des-Arg(9)-bradykinin, des-Arg(10)-kallidin, bradykinin 1-7, bradykinin 1-5 and bradykinin 2-9) to enable reliable insights into their alterations in COVID-19—or other disease states (e.g. allergies or lung cancer)—in comparison to controls. Information regarding these alterations will contribute to the understanding of the pathophysiology related to these peptides and may help to identify new therapeutic targets.
Materials and methods
Chemicals and reagents
Kallidin trifluoroacetic acid (TFA) salt (96.9%, HPLC; Tocris, Bristol, UK), bradykinin acetate (99.0%, HPLC; Sigma-Aldrich, St. Louis, MO, USA), and their metabolites des-Arg(9)-bradykinin acetate (98.7%, HPLC; Santa Cruz Biotechnology, Dallas, TX, USA), bradykinin 1-7 TFA salt (≥ 95.0%, HPLC; GenScript, Piscataway Township, NJ, USA), bradykinin 1-5 TFA salt (≥ 95.0%, HPLC; GenScript), bradykinin 2-9 TFA salt (≥ 95.0%, HPLC; GenScript), and des-Arg(10)-kallidin TFA salt (95.9%, HPLC) were used in this study. [Phe8Ψ(CH-NH)-Arg9]-bradykinin TFA salt (97.5%, HPLC) was applied as the internal standard. Formic acid (FA, ≥ 98%) and TFA (100.3%) were supplied by Sigma Aldrich. HPLC-grade methanol, water, and dimethyl sulfoxide (DMSO, ≥ 99.9%), and MS-grade methanol and ammonium acetate (99.5%) were obtained from Fisher Scientific (Loughborough, UK). Furthermore, HPLC-grade acetonitrile (ACN, Applichem, Darmstadt, Germany), MS-grade water (Honeywell Fluka, Seelze, Germany), and ammonia (30.9%) (VWR Chemicals, Radnor, PA, USA) were utilized. Isotonic saline solution 0.9% was provided by B. Braun (Melsungen, Germany).
Sampling of NLF in healthy volunteers was performed in compliance with the ethical principles of the Declaration of Helsinki and was approved by the ethics committee of the medical faculty at the Heinrich Heine University of Duesseldorf in October 2017 (study number: 6112). Written informed consent was obtained from all participants before enrolment. Bioanalysis was conducted in accordance with Good Clinical Laboratory Practice.
Preparation of stock and working solutions
Lyophilized kinin peptides were dissolved and diluted separately in 0.3% TFA in 25/75 ACN/water (v/v/v) prior to the preparation of a combined working solution containing 500 ng/mL of each peptide salt. [Phe8Ψ(CH-NH)-Arg9]-bradykinin as an internal standard was dissolved in 0.1% FA in water (v/v) and subsequently diluted to achieve a working solution of 500 ng/mL in 0.3% TFA in 25/75 ACN/water (v/v/v). All peptide solutions were prepared using low protein-binding tubes (Sarstedt, Nümbrecht, Germany).
Sample preparation
A 0.9% isotonic saline solution was used as blank surrogate matrix for the respiratory saline lavage fluids. Owing to the endogenous presence of kinins and the long half-life of bradykinin 1-5, no reliable kinin-free human blank matrix could be generated. Optimized inhibitors were applied to effectively prevent the generation and degradation of the kinin peptides, based on previously published suitable inhibitor cocktails31. SPE was performed by applying 96-well Oasis® weak cation exchange (WCX) µ-elution plates (Waters, Milford, MA, USA). All cartridges were conditioned with 200 µL of methanol, followed by 200 µL of 5% aqueous ammonium hydroxide (v/v). Subsequently, the wells were prefilled with 150 µL of 3 ng/mL internal standard in 5% aqueous ammonium hydroxide (v/v), and 100 µL of sample was then loaded. Washing was performed with 300 µL of 5% aqueous ammonium hydroxide (v/v) and 300 µL of 10% methanol in water (v/v). Elution was conducted three times with 50 µL of 1% TFA in 75/25 ACN/water (v/v/v). The resulting eluates were evaporated to dryness under a gentle stream of nitrogen at 60 °C while shaking at 300 rpm. The residues were dissolved in 75 µL of 10/10/80 FA/methanol/water (v/v/v).
LC–MS/MS
Chromatography was performed on an Agilent 1200 SL series system (Agilent Technologies, Ratingen, Germany) consisting of a degasser (G1379B), a binary pump SL (G1379B) and a column oven TCC SL (G1316B). A Phenomenex Synergi™ 2.5 µm Hydro-RP 100 Å column (100 × 2.0 mm; Torrance, CA, USA) was used for the chromatographic separation. The mobile phases were composed of water and methanol (B) both containing 3.2% DMSO and 0.1% FA (v/v). A 7.5 min binary gradient was applied, maintaining the amount of mobile phase B at 5% for 1.5 min, increasing it to 20% until 2.2 min, to 27% until 2.7 min, to 35% until 3.1 min, and finally to 95% after 6.2 min. Mobile phase B was kept constant at 95% until 6.7 min before decreasing it to 5% and column re-equilibration for 3 min. The flow rate was set to 400 µL/min, and the injection volume of 50 µL was applied with an HTC PAL autosampler (CTC Analytics AG, Zwingen, Switzerland). After every injection, the autosampler syringe was rinsed twice with 0.2% FA in 80/20 ACN/water (v/v/v). Samples were stored at 18 °C in the autosampler.
The LC system was coupled to an API 4000 (AB Sciex, Darmstadt, Germany) mass spectrometer equipped with a Turbo V source for detection. The electrospray interface was operated in positive mode with multiple reaction monitoring mode. The curtain gas was maintained at 31 psi, the collision gas at 8 psi, the nebulizer gas at 45 psi, the heater gas at 65 psi, the ion spray voltage at 5500 V, and the source temperature at 350 °C. Peptide-specific parameters are displayed in Table 1.
Data acquisition was conducted using Analyst® 1.6.2 software (AB Sciex, Darmstadt, Germany), and raw data evaluation was performed using Multiquant™ 3.0.2 (AB Sciex, Darmstadt, Germany).
Method development
Adaption of optimized injection solvent and sample collection material
A previously conducted DoE approach to optimize the injection solvent conjointly with the sample collection material to reduce non-specific adsorption of bradykinin and thus increase sensitivity30, had to be adapted to avoid peak broadening or breakthrough of the more hydrophilic kinin peptides. By using the D-optimal optimization model, an amount of 5–20% organic fraction in the injection solvent was investigated. This range correlated with the binary gradient, as no breakthrough of the kinins was expected. Furthermore, the calculations included a maximum intensity loss of 15% of the predicted intensity of the optimized injection solvent, based on current bioanalytical guidelines for the accuracy limits32. Six injection solvent compositions were calculated with distinct organic fractions and analyzed in triplicates by LC–MS/MS measurement, and responses and peak shapes were compared to the original injection solvent for bradykinin only (8.7% FA in 5.3/36.6/49.4 methanol/DMSO/water (v/v/v/v)).
Improvement of SPE recovery
The method development focused on maximizing recovery to reduce peptide loss during washing steps and to enable the detection of endogenous concentrations in the low pg/mL range. A previously developed SPE protocol for bradykinin only30 had to be adapted, since the kinin peptides differ in their amounts of hydrophobic and positively charged amino acids (Fig. 1). Mixed-mode strong cation exchange (MCX) and WCX µelution SPE were considered to evaluate which material would fit best to all analytes. All experiments investigating the washing and elution solvents were conducted using neat solution in duplicate.
Validation
Bioanalytical method validation was conducted considering the regulatory bioanalytical guidelines of the US Food and Drug administration32. Linearity, accuracy, precision, sensitivity, recovery, matrix effect, carry-over, and stability were addressed during the validation process.
Linearity
Linearity was determined in six runs using nine to eleven distinct calibrator levels (depending on the lower limit of quantification (LLOQ) of the individual peptide), which were analyzed in single determinations. In compliance with bio-analytical guidelines, the actual concentration of 75% of all calibration curve standards had to deviate less than ± 15% (± 20% at the LLOQ) from their nominal concentration (relative error (RE))32.
Accuracy, precision and sensitivity
Accuracy and precision were assessed using up to seven quality control (QC) levels covering the whole calibration range on three distinct days. The number of QCs depended on the magnitude of the calibration range per peptide. Five replicates per QC level were analyzed each day. Accuracies were determined as the deviation of the actual concentration from the nominal concentration (RE) for within-run and for between-run accuracy. Using one-way ANOVA, within-run precision was calculated as repeatability and between-run precision as day-different intermediate precision (coefficient of variation (CV)). In line with regulatory guidelines, accuracy and precision were not allowed to exceed ± 15% (± 20% at the LLOQ)32. The signal-to-noise ratio (S/N) had to be higher than 5:1 at the LLOQ. The limit of detection (LoD) was further calculated as follows using the results from six calibration curves Eq. (1)33:
Calculation of limit of detection. σ: standard deviation of the y-intercept, S: mean slope of the calibration curves.
Carry-over
Carry-over was evaluated by alternatingly injecting blank samples and upper limit of quantification (ULOQ) standards six times. According to regulatory bioanalytical guidelines, carry-over in the blank sample following the ULOQ was not allowed to exceed 20% of the LLOQ signal and 5% of the internal standard signal32.
Recovery and absolute matrix effect
Recovery was determined at four distinct QC levels covering the calibration curve range (high, middle, low and around the LLOQ) in triplicate. Pre-spiked extracted samples were compared to processed blank samples spiked with the same concentrations after µelution SPE. Matrix effects of the peptides were analyzed at the same four distinct QC levels by comparison of post-spiked samples to neat solutions (n = 3).
Stability
Stability studies were conducted at four QC levels (high, middle, low and around the LLOQ) under different storage conditions. Benchtop stability was investigated by placing the prepared QC samples at room temperature for one and three hours (n = 5). Freeze–thaw stability (room temperature to − 80 °C) was evaluated by analyzing the QC levels after one and three cycles (n = 5). Between each cycle, samples were frozen for at least 12 h. The autosampler stability was assessed by keeping QC samples in the autosampler at 18 °C for 18 h and then repeating the QC sample measurement. Finally, short-term stability of processed and evaporated samples was analyzed after storage for 24 h at + 4 °C (n = 5). Stability at the specific conditions was proven if the mean concentration at each level did not exceed ± 15%. To evaluate the stability of the analyte working solution, peak areas of the kinin peptides on 15 distinct days after 15 freeze–thaw cycles, measured routinely during method performance qualification (1 ng/mL, neat solution), were analyzed for their CV (acceptance criterion ≤ 15%).
Applicability
Nasal lavage with 10 mL of 0.9% isotonic saline (5 mL per nostril) was performed in nine healthy volunteers (6 female/3 male). Volunteers were healthy adults above the age of 18 years without any signs of a respiratory infection or allergy. The volunteers were asked to tip their head backwards, hold the breath, and refrain from swallowing. The obtained fluid was collected directly into the inhibitor cocktail and was immediately vortexed after completing the sampling. The recovered volume was documented to enable the non-invasive normalization of kinin levels. At least 30% of the instilled volume had to be recovered during the lavage in line with the American Thoracic Society guideline for bronchoalveolar lavage34. The samples were centrifuged at 4 °C for 15 min at 500×g to remove cells, mucus, and debris. A 100 µL aliquot of the supernatant was analyzed.
Results
Method development
Improvement of sensitivity
Analyzing the contour plots of the D-optimal optimization model for the original injection solvent composition and the contour plots with restricted organic amount, showed clearly that high acidic amounts were necessary for the injection solvent, as well as the highest organic amount that would be compatible with peak shapes of the hydrophilic peptides (Fig. 2). Of the six investigated injection solvent compositions, 10/10/80 FA/methanol/water (v/v/v) gave best peak shapes for bradykinin 1-5, by maintaining good signal intensity for bradykinin with 98.8 ± 2.3% of the optimized injection solvent composition (predicted: 91.4% with a log(D) of − 0.49 and probability of failure of 0.12%).
Contour plots of the D-optimal optimization for the response of bradykinin in distinct injection solvent compositions. In comparison to the contour plot including the original injection solvent for bradykinin with a fixed fraction of 0.366 dimethyl sulfoxide (a), restriction of the organic amount in context with the comprehensive determination of seven kinin peptides (b and c) lead to a shift to highest compatible organic and acidic fractions with regard to the hydrophilic peptides. White areas indicate regions, that were not investigated or were excluded due to peak distortion for hydrophilic peptides. cps: counts per second.
Improvement of recovery
Using MCX SPE, the more hydrophobic or charged peptides (bradykinin, des-Arg(10)-kallidin, kallidin, and the internal standard) could not be satisfactorily eluted, when applying two 100 µL elutions of 5% of ammonium hydroxide in ACN, as indicated by low recoveries of 0–3%. Increasing the elution volume or adding ammonium acetate to the elution solvent to produce a salting-out effect did not substantially improve recoveries. Therefore, a customized protocol using WCX with modified washing steps was developed. In particular, the more hydrophilic kinins and those containing fewer amino functional groups (bradykinin 1-5 and bradykinin 1-7) were not robust regarding the recovery, as they were washed out when using an organic amount exceeding 10% methanol, subsequently affecting sensitivity and precision (Fig. 3). An increased amount of washing solvent (300 µL of 10% methanol) did not affect peptide recoveries but resulted in more precise values and was subsequently applied as described in the sample preparation section.
Recoveries during evaluation of washing steps using neat solutions and applying weak cation exchange µelution solid-phase extraction (n = 2). Peptide characteristics were calculated using ProtParam43. ACN: acetonitrile, GRAVY: grand average of hydropathicity index, MeOH: methanol, NH3: ammonium.
Method validation
The results for linearity measurements gave best fits using quadratic regression except for bradykinin 1-5, where linear regression was applied (mean r ≥ 0.9960 for all analytes). The broad dynamic calibration curve ranges were (depending on the analyte) between 4.4 and 22.8 pg/mL for the LLOQ and between 4505.9 and 8255.5 pg/mL for the ULOQ. More detailed results for the linearity and the respective calibration curve ranges are presented in Table 2 and an example in supplementary Figure S1.
The within-run and between-run precision as well as accuracy results for all investigated QC levels are presented in Table 3. Guideline-compliant results were obtained for all kinin peptides32. In line with the maximally allowed deviation of 20% for accuracy and precision at the LLOQ, the LLOQ was set to 4.4 pg/mL for kallidin (S/N: 94), to 6.7 pg/mL for bradykinin (S/N: 96), to 10.6 pg/mL for des-Arg(10)-kallidin (S/N: 199), to 7.3 pg/mL for (des-Arg(9)-bradykinin (S/N: 403), to 6.5 pg/mL for bradykinin 1-7 (S/N: 155), to 8.1 pg/mL for bradykinin 2-9 (S/N: 65) and to 22.8 pg/mL for bradykinin 1-5 (S/N: 39). Representative chromatograms for the blank, the respective LLOQ, and the high QC are displayed in Fig. 4. The LoD was calculated as 3.5 pg/mL for kallidin, 2.5 pg/mL for bradykinin, 2.5 pg/mL for des-Arg(10)-kallidin, 3.0 pg/mL for des-Arg(9)-bradykinin, 4.4 pg/mL for bradykinin 1-7, 5.6 pg/mL for bradykinin 2-9 and 13.6 pg/mL for bradykinin 1-5.
The carry-over following the injection of an ULOQ sample was below 20% of the signal of the LLOQ for all analytes (kallidin: 19.0%, bradykinin: 17.0%, des-Arg(10)-kallidin: 16.8%, des-Arg(9)-kallidin: 19.1%, bradykinin 1-7: 11.9%, bradykinin 2-9: 18.9%, bradykinin 1-5: 4.4%). No carry-over was observed for the internal standard.
At the distinct QC levels, recoveries did not vary substantially, as indicated by the low CVs of ≤ 5% between the different levels. The more hydrophilic analytes bradykinin 1-5 (mean 34.1%) and bradykinin 1-7 (mean 45.8%) presented lower recoveries compared to the peptides with more lipophilic or additional amine functional groups (mean 74.1% to 88.4%) (Fig. 5). Mean ion suppression of the four levels ranged from − 16.8% (bradykinin 1-5) to − 4.3% (des-Arg(10)-kallidin) (Fig. 5).
All kinin peptides were stable during autosampler storage for 18 h at 18 °C and throughout the short-term stability test for 24 h at 4 °C (supplementary table S2). With the exception of bradykinin 1-5, all other peptides were further stable for three freeze–thaw cycles and on the benchtop for 3 h. Bradykinin 1-5 was only stable for one hour on the benchtop, whereas after 3 h a mean decrease of − 31.0% was observed. Additionally, bradykinin 1-5 showed a tendency to degrade during freeze–thaw cycles, with increased degradation after three cycles compared to one (mean decrease: − 24.3% (1 h) vs. − 31.3% (3 h)). However, bradykinin 1-5, as well as the other kinin peptides, were stable for 15 freeze–thaw cycles measured over the course of 1 month (CV: 13.8%) in the analyte working solution. Since enzyme-free matrix was applied, the degradation was not related to insufficient inhibition of enzyme activity. Therefore, degradation might be caused by ionic interactions known to potentially affect instability. Thus, patient samples should be exposed to as few as feasible freeze–thaw cycles and be prepared freshly whenever possible.
Applicability
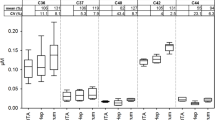
Endogenous levels of the kinin peptides in nine healthy volunteers were comprehensively determined in NLF and confirmed the method applicability. The recovery of instilled lavage volumes ranged from 33 to 89% in the healthy volunteers. Normalized mean levels for all kinin peptides were in the low pg/mL range; namely, 8.2 pg/mL for kallidin, 2.9 pg/mL for des-Arg(10)-kallidin, 22.5 pg/mL for bradykinin, 40.3 pg/mL for des-Arg(9)-bradykinin, 36.4 pg/mL for bradykinin 1-7 and 27.5 pg/mL for bradykinin 1-5 were measured. Bradykinin 2-9 was below the LoD (< 5.6 pg/mL) in all volunteer samples. Box plots of the level data are presented in Fig. 6 and a representative chromatogram is shown in Fig. 4.
Box plots of the kinin peptide concentrations of healthy volunteers (n = 9) in nasal lavage fluid. Data were normalized to the recovered volume of nasal instillation with 10 mL saline. Filled black square: interquartile range, T: 1.5 interquartile range, –: median, open square: mean, filled black rhombus: outlier.
Discussion
The presented novel LC–MS/MS assay enabled the comprehensive and accurate determination of bradykinin, kallidin, des-Arg(9)-bradykinin, des-Arg(10)-kallidin, bradykinin 2-9, bradykinin 1-7, and bradykinin 1-5 in NLF. Characterized by a high sensitivity (4.4–22.8 pg/mL depending on the kinin) despite the use of low volumes, the applicability of this method was successfully proven by determining low-abundance kinin peptides in NLF. Full validation according to regulatory bioanalytical guidelines was achieved32.
To the best of our knowledge, this study is the first report of the comprehensive determination of kinin peptides in respiratory saline lavage fluid. Previous determinations of kinins in NLF by immunometric approaches did not quantitatively differentiate between the kinins and were limited to bradykinin and kallidin19,20. Immunoassays are prone to cross-reactivity with structurally similar peptides, which impacts the accuracy and reliability of the results. Further, these methods do not allow the simultaneous investigation and differentiation of kinin peptides from one sample aliquot. Thus, disease-related alterations in the KKS cascade by inhibition or inducement of enzyme activities that affect the generation or degradation of kinin peptides cannot be comprehensively assessed. Furthermore, data obtained from available LC–MS/MS methods is limited and provides a narrow scope of information. The restricted determination of not more than two kinin peptides simultaneously and the generally inadequate sensitivity to detect endogenous peptides in the low pg/mL range by LC–MS/MS has not yet allowed for a comprehensive assessment of the KKS.
Therefore, in advance of this method validation, extensive and systematic investigation was conducted to improve the sensitivity by optimizing the mobile phase and reducing nonspecific peptide adsorption of bradykinin30. By means of the DoE approach, substantial signal intensity increases for bradykinin—by a factor of 7.7 for the mobile phase optimization and by a factor of 26.6 for the injection solvent optimization—were achieved30. Following this approach, the intensity of the other peptides could now be improved through DoE and formed the basis to facilitate the low detection limits of 6.7 pg/mL for bradykinin and the range of 4.4 to 22.8 pg/mL for the other six kinin peptides using saline matrix. As suitable assays in saline solution are lacking, the performance of the developed assay can only be discussed in relation to other human matrices. For bradykinin 1-5, Seip et al. 2015 obtained a similar LLOQ of 20.3 pg/mL (vs. 22.8 pg/mL in the here presented study), but applied larger sample volumes (1 mL blood)35. The measurement of des-Arg(9)-bradykinin by LC–MS/MS was marked by a quantification limit of 2 ng/mL25. The LC–MS/MS assay of Lindström et al., established a LLOQ of 106.2 pg/mL for bradykinin using 500 µL of plasma23, which was already a factor of 100 below previously published LC–MS/MS methods with detection limits of 10 ng/mL24,25. However, this sensitivity was not sufficient to determine endogenous levels of bradykinin throughout all their plasma samples23.
In the current study, for the first time in NLF, concentrations of endogenous levels of specific kinin peptides were detectable in saline matrix and allowed for their comprehensive determination. Proud et al. 1983, measured kinin peptides in eight controls by immunometric detection, seven had levels below the LLOQ (< 20 pg/mL) and one had a level of 100 pg/mL. As mentioned above, a quantitative breakdown of the total kinin concentration compared to the respective peptides could not be made19. Turner et al., determined kinin levels of 68 (43–183) pg/mL (combined bradykinin and kallidin without distinction) (median (80% central range), n = 8)20. This is in line with the measured levels of healthy volunteers in NLF, where distinguished mean levels of 22.5 pg/mL for bradykinin and 8.2 pg/mL for kallidin were obtained. Levels for bradykinin 1-7, des-Arg(9)-bradykinin, des-Arg(10)-kallidin and bradykinin 1-5 were also in the low pg/mL range, as expected. Whereas bradykinin, bradykinin 1-7 and des-Arg(9)-bradykinin were detectable in all samples, presence of the other kinins in NLF varied individually and bradykinin 2-9 was below the quantification limit in all samples. Because levels of the cleaved peptides are not published elsewhere, a reliable classification of the concentrations is only possible to a limited extent, and it is rather necessary to ascertain these endogenous levels in larger healthy and diseased cohorts in future studies. Levels in patients are expected to exceed those in healthy volunteers if the hypothesis of a dysregulated KKS in COVID-19 can be confirmed. Therefore, the broad calibration curve range of the developed assay, covering a span of a factor of 250–1000 depending on the analyte, is expected to be suitable because it allows measuring endogenous levels in healthy controls, as well as detecting possible elevations. Further, application of the assay can easily be extended to other diseases in which alterations within the KKS are to be expected and provides the advantage of non-invasive and easy-to-handle sampling. Thus, it allows to investigate e.g. lung cancer, respiratory allergic reactions, and bradykinin-mediated side effects of ACE inhibitors36,37,38.
The developed LC–MS/MS assay further outmatched previously published immunoassay methods separating kinin peptides (bradykinin 1-7, des-Arg(9)-bradykinin, and bradykinin) in the low-abundant endogenous range regarding sample preparation effort, as it makes the final results available within 2 h of sampling. Campbell et al. 1993 applied a combination of C18 SPE followed by liquid–liquid extraction and chromatographic separation prior to immunoradiometric detection21. Duncan et al. purified their samples through five rounds of SPE followed by chromatographic separation before the fractions were analyzed by immunoassay22. Lower limits of quantification (0.3–0.4 pg/mL) were reached using these approaches; however, also 1 mL of blood was applied, which subsequently shows a sensitivity nearly equal to the presented LC–MS assay (100 µL sample volume). Advantages of the LC–MS assay are that falsification due to cross-reactivity can be excluded, and the obtained values are attributed to single peptides. Further, the significantly reduced sample preparation effort achieved in combination with a fast analysis time (~ 2 h vs. ~ 1 day), provides the opportunity to reduce the time working with potentially infectious patient samples.
Isotonic saline was chosen as the surrogate matrix for preparation of calibration curve and quality control samples to obtain blank matrix and avoid any interference with endogenous kinin peptides in respiratory lavage fluids. This was presumed to be an adequate approach, as NLF is mostly made up of saline owing to instillation with saline during lavage, with a reported dilution of factor 60–12026. Further, mucus, debris, and cells are separated by centrifugation. An important issue in quantitative lavage analysis is that the volume infused during saline lavage is not always equal to the volume sampled39. Therefore, when comparing data sets of determined peptide levels, e.g. at different time points or in different patients, normalizing against an endogenous dilution marker, such as albumin, total protein abundance, or urea has been proposed40. However, results might be influenced by capillary leakage in many respiratory disorders and an additional blood sampling is required to calculate the dilution40,41. Currently, no standardization regarding normalization of lavage fluids is available as stated in a consensus statement of the International Society for Heart and Lung Transplantation in 202042. To maintain the non-invasive manner of the assay, in this study the recovered volume was used for normalization, which is the most commonly applied normalization strategy42. Because saline is used as surrogate matrix, the assay is not limited to the investigation of respiratory saline lavage fluids in humans, but can also be used in animal models. Thereby, the innovative assay allows to investigate the pathophysiology of COVID-19 but might also support the identification of possible new therapeutic targets if the hypothesis of an altered KKS can be confirmed in COVID-19.
In conclusion, the novel LC–MS/MS assay facilitates the comprehensive determination of kallidin, bradykinin, des-Arg(10)-kallidin, des-Arg(9)-bradykinin, bradykinin 1-7, bradykinin 2-9 and bradykinin 1-5 for the first time in saline. The method is well-suited for research purposes considering its high sensitivity and broad calibration curve range in combination with low applied volumes. The successfully validated method will contribute to elucidate the pathophysiology of SARS-CoV-2 by facilitating the investigation of the postulated connection between a dysregulated KKS and other clinical syndromes (e.g. COVID-19).
Data availability
The (raw) datasets in the context of the here presented validation are made available on a public repository (https://doi.org/10.5281/zenodo.4431745).
References
Wang, D. et al. Clinical characteristics of 138 hospitalized patients with 2019 novel coronavirus-infected pneumonia in Wuhan, China. JAMA https://doi.org/10.1001/jama.2020.1585 (2020).
de Maat, S., de Mast, Q., Danser, A. J., van de Veerdonk, F. L. & Maas, C. Impaired breakdown of bradykinin and its metabolites as a possible cause for pulmonary edema in COVID-19 infection. Semin. Thromb. Hemost. https://doi.org/10.1055/s-0040-1712960 (2020).
Roche, J. A. & Roche, R. A hypothesized role for dysregulated bradykinin signaling in COVID-19 respiratory complications. FASEB J. 34, 7265–7269. https://doi.org/10.1096/fj.202000967 (2020).
van de Veerdonk, F. L. et al. Kallikrein-kinin blockade in patients with COVID-19 to prevent acute respiratory distress syndrome. Elife https://doi.org/10.7554/eLife.57555 (2020).
Nicolau, L. A. D., Magalhães, P. J. C. & Vale, M. L. What would Sérgio Ferreira say to your physician in this war against COVID-19: How about kallikrein/kinin system?. Med. Hypotheses 143, 109886. https://doi.org/10.1016/j.mehy.2020.109886 (2020).
Ellis, K. M. & Fozard, J. R. Species differences in bradykinin receptor-mediated responses of the airways. Autonom. Auta Pharm. 22, 3–16. https://doi.org/10.1046/j.1474-8673.2002.00230.x (2002).
Magerl, M. et al. Bradykinin in health and disease. In: Proceedings of the Bradykinin Symposium 2012, Berlin 23–24 August 2012. Inflammation research : official journal of the European Histamine Research Society … [et al.] Vol. 63, 173–178. https://doi.org/10.1007/s00011-013-0693-1 (2014).
Blais, C. et al. Contribution of angiotensin-converting enzyme to the cardiac metabolism of bradykinin: An interspecies study. Am. J. Physiol. Heart Circ. Physiol. 273, 2263–2271 (1997).
McLean, P. G., Perretti, M. & Ahluwalia, A. Kinin B 1 receptors as novel anti-inflammatory targets. Emerg. Ther. Targets 4, 127–141. https://doi.org/10.1517/14728222.4.2.127 (2005).
Gobeil, F. et al. Pharmacological profiles of the human and rabbit B1 receptors. Can. J. Physiol. Pharmacol. 75, 591–595 (1997).
Broadley, K. J., Blair, A. E., Kidd, E. J., Bugert, J. J. & Ford, W. R. Bradykinin-induced lung inflammation and bronchoconstriction. Role in parainfluenze-3 virus-induced inflammation and airway hyperreactivity. J. Pharmacol. Exp. Ther. 335, 681–692. https://doi.org/10.1124/jpet.110.171876 (2010).
Sodhi, C. P. et al. Attenuation of pulmonary ACE2 activity impairs inactivation of des-Arg9 bradykinin/BKB1R axis and facilitates LPS-induced neutrophil infiltration. Am. J. Physiol. Lung Cell. Mol. Physiol. 314, L17–L31. https://doi.org/10.1152/ajplung.00498.2016 (2018).
Vickers, C. et al. Hydrolysis of biological peptides by human angiotensin-converting enzyme-related carboxypeptidase. J. Biol. Chem. 277, 14838–14843. https://doi.org/10.1074/jbc.M200581200 (2002).
Hoffmann, M. et al. SARS-CoV-2 cell entry depends on ACE2 and TMPRSS2 and is blocked by a clinically proven protease inhibitor. Cell 181, 271-280.e8. https://doi.org/10.1016/j.cell.2020.02.052 (2020).
Tolouian, R., Vahed, S. Z., Ghiyasvand, S., Tolouian, A. & Ardalan, M. COVID-19 interactions with angiotensin-converting enzyme 2 (ACE2) and the kinin system; looking at a potential treatment. J. Renal Injury Prev. 9, e19. https://doi.org/10.34172/jrip.2020.19 (2020).
Sungnak, W. et al. SARS-CoV-2 entry factors are highly expressed in nasal epithelial cells together with innate immune genes. Nat. Med. https://doi.org/10.1038/s41591-020-0868-6 (2020).
Cascella, M. et al. Features, Evaluation, and Treatment of Coronavirus [Updated 2020 Oct 4]. StatPearls [Internet]. Treasure Island (FL): StatPearls Publishing, (2020) https://www.ncbi.nlm.nih.gov/books/NBK554776/.
Pitrez, P. M. C., Brennan, S., Turner, S. & Sly, P. D. Nasal wash as an alternative to bronchoalveolar lavage in detecting early pulmonary inflammation in children with cystic fibrosis. Respirology 10, 177–182 (2005).
Proud, D. et al. Kinins are generated in vivo following nasal airway challenge of allergic individuals with allergen. J. Clin. Investig. 72, 1678–1683 (1983).
Turner, P. J., Dear, J. W. & Foreman, J. C. Involvement of kinins in hyperresponsiveness induced by platelet activating factor in the human nasal airway. Br. J. Clin. Pharmacol. 129, 525–532. https://doi.org/10.1038/sj.bjp.0703095 (2000).
Campbell, D. J., Kladis, A. & Duncan, A. M. Bradykinin peptides in kidney, blood, and other tissues of the rat. Hypertension 21, 155–165. https://doi.org/10.1161/01.HYP.21.2.155 (1993).
Duncan, A. M. et al. Kinins in humans. Am. J. Physiol. Regul. Integr. Comp. Physiol. 278, R897–R904. https://doi.org/10.1152/ajpregu.2000.278.4.R897 (2000).
Lindström, M. et al. Plasma bradykinin concentrations during septic shock determined by a novel LC-MS/MS assay. Clin. Chim. Acta 493, 20–24. https://doi.org/10.1016/j.cca.2019.02.023 (2019).
Baralla, E. et al. Quantitative assay for bradykinin in rat plasma by liquid chromatography coupled to tandem mass spectrometry. J. Pharm. Biomed. Anal. 54, 557–561. https://doi.org/10.1016/j.jpba.2010.09.041 (2011).
van den Broek, I., Sparidans, R. W., Schellens, J. H. M. & Beijnen, J. H. Quantitative assay for six potential breast cancer biomarker peptides in human serum by liquid chromatography coupled to tandem mass spectrometry. J. Chromatogr. B Anal. Technol. Biomed. Life. Sci. 878, 590–602. https://doi.org/10.1016/j.jchromb.2010.01.011 (2010).
Franciosi, L., Govorukhina, N., ten Hacken, N., Postma, D. & Bischoff, R. Protemics of epithelial lining fluid obtained by bronchscopic microprobe sampling. Nanoproteomics Methods Mol. Biol. 790, 17–28. https://doi.org/10.1007/978-1-61779-319-6_2 (2011).
John, H., Walden, M., Schäfer, S., Genz, S. & Forssmann, W.-G. Analytical procedures for quantification of peptides in pharmaceutical research by liquid chromatography-mass spectrometry. Anal. Bioanal. Chem. 378, 883–897 (2004).
Goebel-Stengel, M., Stengel, A., Taché, Y. & Reeve, J. R. The importance of using the optimal plasticware and glassware in studies involving peptides. Anal. Biochem. 414, 38–46. https://doi.org/10.1016/j.ab.2011.02.009 (2011).
Feickert, M. & Burckhardt, B. B. A design of experiments concept for the minimization of nonspecific peptide adsorption in the mass spectrometric determination of substance P and related hemokinin-1. J. Sep. Sci. 43, 818–828. https://doi.org/10.1002/jssc.201901038 (2019).
Gangnus, T. & Burckhardt, B. B. Improving sensitivity for the targeted LC-MS/MS analysis of the peptide bradykinin using a design of experiments approach. Talanta 218, 121134. https://doi.org/10.1016/j.talanta.2020.121134 (2020).
Nussberger, J. et al. Plasma bradykinin in angio-oedema. Lancet 351, 1693–1697. https://doi.org/10.1016/S0140-6736(97)09137-X (1998).
U.S. Food and Drug Administration. Bioanalytical Method Validation. Guidance for Industry. https://www.fda.gov/files/drugs/published/Bioanalytical-Method-Validation-Guidance-for-Industry.pdf (2018).
Committee for Medicinal Products for Human Use/International HMP/International Counsil for Harmonisation of Technical Requirements for Pharmaceuticals for Human Use. ICH Topic Q 2 (R1) Validation of Analytical Procedures: Text and Methodology (1995).
Meyer, K. C. et al. An Official American Thoracic Society clinical practice guideline: The clinical utility of bronchoalveolar lavage cellular analysis in interstitial lung disease. Am. J. Respir. Crit. Care Med. 185, 1004–1014 (2012).
Seip, K. F. et al. Bradykinin analysis revived–a validated method for determination of its stable metabolite in whole blood by LC-MS/MS. J. Chromatogr. B Anal. Technol. Biomed. Life Sci. 947–948, 139–144. https://doi.org/10.1016/j.jchromb.2013.12.033 (2014).
Hicks, B. M. et al. Angiotensin converting enzyme inhibitors and risk of lung cancer. Population based cohort study. BMJ 4, k4209. https://doi.org/10.1136/bmj.k4209 (2018).
Borghi, C. & Veronesi, M. Cough and ACE inhibitors. The truth beyond placebo. Clin. Pharmacol. Ther. 105, 550–552. https://doi.org/10.1002/cpt.1040 (2018).
Golias, C. H., Charalabopoulos, A., Stagikas, D., Charalabopoulos, K. & Batistatou, A. The kinin system—bradykinin: Biological effects and clinical implications. Multiple role of the kinin system—bradykinin. Hippokratia 3, 124–128 (2007).
Bowen, C. L. & Licea-Perez, H. Development of a sensitive and selective LC–MS/MS method for the determination of urea in human epithelial lining fluid. J. Chromatogr. B 917–918, 24–29. https://doi.org/10.1016/j.jchromb.2012.11.035 (2013).
Rennard, S. I. et al. Estimation of volume of epithelial lining fluid recovered by lavage using urea as marker of dilution. J. Appl. Physiol. 60, 532–538. https://doi.org/10.1152/jappl.1986.60.2.532 (1986).
de Blic, J. et al. Bronchoalveolar lavage in children. ERS Task Force on bronchoalveolar lavage in children. European Respiratory Society. Eur. Respir. J. 15, 217–231 (2000).
Martinu, T. et al. International Society for Heart and Lung Transplantation consensus statement for the standardization of bronchoalveolar lavage in lung transplantation. J. Heart Lung Transpl. 39, 1171–1190. https://doi.org/10.1016/j.healun.2020.07.006 (2020).
Gasteiger, E. et al. The Proteomics Protocols Handbook. Protein Identification and Analysis Tools on the ExPASy Server 571–607 (Humana Press, Boca Raton, 2005).
Funding
Open Access funding enabled and organized by Projekt DEAL. This work was supported by Life Sciences Research Foundation (formerly called D Collen Research Foundation). The funder was not involved in the conceptualization, design, data collection, analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
T.G. conceptualization, methodology, validation, writing—original draft, review and editing, visualization. B.B.B. conceptualization, methodology, writing—review and editing, supervision.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gangnus, T., Burckhardt, B.B. Sensitive mass spectrometric determination of kinin-kallikrein system peptides in light of COVID-19. Sci Rep 11, 3061 (2021). https://doi.org/10.1038/s41598-021-82191-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-82191-7
This article is cited by
-
Mass spectrometric study of variation in kinin peptide profiles in nasal fluids and plasma of adult healthy individuals
Journal of Translational Medicine (2022)
-
Analysis on the characteristics of spatio-temporal evolution and aggregation trend of early COVID-19 in mainland China
Scientific Reports (2022)
-
Targeted LC-MS/MS platform for the comprehensive determination of peptides in the kallikrein-kinin system
Analytical and Bioanalytical Chemistry (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.