Abstract
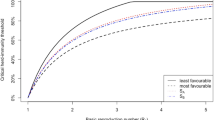
After a year of living with the COVID-19 pandemic and its associated consequences, hope looms on the horizon thanks to vaccines. The question is what percentage of the population needs to be immune to reach herd immunity, that is to avoid future outbreaks. The answer depends on the basic reproductive number, R0, a key epidemiological parameter measuring the transmission capacity of a disease. In addition to the virus itself, R0 also depends on the characteristics of the population and their environment. Additionally, the estimate of R0 depends on the methodology used, the accuracy of data and the generation time distribution. This study aims to reflect on the difficulties surrounding R0 estimation, and provides Spain with a threshold for herd immunity, for which we considered the different combinations of all the factors that affect the R0 of the Spanish population. Estimates of R0 range from 1.39 to 3.10 for the ancestral SARS-CoV-2 variant, with the largest differences produced by the method chosen to estimate R0. With these values, the herd immunity threshold (HIT) ranges from 28.1 to 67.7%, which would have made 70% a realistic upper bound for Spain. However, the imposition of the delta variant (B.1.617.2 lineage) in late summer 2021 may have expanded the range of R0 to 4.02–8.96 and pushed the upper bound of the HIT to 90%.
Similar content being viewed by others
Introduction
On 11 March 2020, the World Health Organization declared the COVID-19 pandemic, and by 11 March 2021, 2.63 million people had died because of it1. However, although these are the published figures, there were probably many more undocumented virus related deaths that were not recorded due to lack of tests2,3,4. After a year of struggling, restrictions to lessen the spread of the virus, a downturn in the economy and the cost of human lives, most people are wondering when the pandemic will end. The year 2020 ended with the hopeful approval of some vaccines5, but how many people must be vaccinated to return to pre-pandemic life? The answer is quite complicated since vaccines do not provide 100% protection against infections6,7 nor fully block the transmissibility of the virus8,9,10. However, it is theoretically interesting to study when the herd immunity threshold (HIT) will be reached, if possible, under the assumptions that immune population (recovered and vaccinated people) get permanent immunisation against the different mutations of the SARS-CoV-2 virus and will not transmit the virus any further. In Spain, there is a general opinion that the HIT will be reached when 70% of the population becomes immune, which is not equivalent to 70% of vaccinated population in real life. Note that there is no single definition of HIT11 and this can lead to misunderstandings. In this study, HIT will refer to the minimum proportion of the immune population that will produce a monotonic decrease of new infections, even if restrictions are lifted and society returns to a pre-pandemic level of social contact. The question is how realistic is a HIT of 70% for Spain.
The HIT is usually defined in terms of the effective reproduction number, Re(t), which is the average number of secondary infections produced by an infected individual at time t. Any outbreak starts with Re > 1, stabilizes with Re = 1, and declines with Re < 1. Therefore, the HIT will be reached when Re = 1 and Re < 1 afterwards. Given the number of susceptible individuals, that is, those that can get infected, Re(t) can be estimated in an unmitigated epidemic as12,13
where S(t) is the number of susceptible individuals at time t; N is the total number of the population; and R0 is the basic reproductive number, that is, the expected number of secondary infections produced by an infected individual in a population where all individuals are susceptible and there are no measures to reduce transmission12,14. The proportion of susceptible, S(t)/N, can be written as 1 − q, where q is the proportion of immune population. Then, if Re(t) = 1 (and Re(t) < 1 afterwards), HIT equals q by definition. Replacing these equalities in Eq. (1) and operating, we get15,16
Note the direct relationship: the larger the R0, the larger the HIT; and that Eq. (2) makes sense only when R0 > 1, since for values R0 < 1 the disease will disappear naturally and the concept of HIT loses its sense. In Eq. (2) it is intrinsically assumed that recovered individuals cannot become susceptible again, that is, they cannot get re-infected nor transmit the virus after recovery. R0 is used to quantify the transmissibility of the virus, which depends on the virus itself and the characteristics of the population that is being infected. Regarding other infectious diseases, typical values of R0 are 0.9–2.1 for seasonal flu and 1.4–2.8 for the 1918 flu17, ~ 3 for SARS-CoV-118 and < 0.8 for MERS19. For COVID-19 in 2020, a systematic review of 21 studies, mainly in China, found R0 ranging from 1.9 to 6.520, which leads to HIT values between 47 and 84%. However, in 62% of these studies, the R0 was between 2 and 3 (HIT between 50 and 67%). In Western Europe in 2020, an average R0 was estimated at 2.2 (95% CI = [1.9, 2.6])21, with a HIT value of 55% (95% CI = [47, 62]). Therefore, 70% is an upper bound of HIT in 2020 in most of the cited cases, but not in all.
Theoretically, R0 can only be observed at the very beginning of the pandemic, while the whole population are susceptible and no control measures are in force (e.g., social distancing, the use of masks, etc.). This is the case in the above-mentioned studies20,21. However, during the COVID-19 pandemic the virus has mutated into more transmissible variants, with a higher R0. In consequence, the HIT has been increasing during the course of the pandemic, but its estimated value cannot be directly updated because the new variants did not exist at the beginning of the pandemic when the R0 should have been observed.
This study encompasses a detailed analysis of the HIT of the ancestral variant, that was the dominant variant at the beginning of the pandemic, from different approaches and quantifies the influence of three key factors: (1) source/quality of data; (2) infectiousness evolution over time; and (3) methodology to estimate R0. Finally, we indirectly estimate the R0 of the current dominant variants using Eq. (1) and comparisons between Re values of several variants. The HIT values derived from these new R0 estimates are discussed in the last section.
Data
Three COVID-19 daily infection datasets for Spain were used, from 1 January to 29 November 2020: (1) official infections published by the Instituto de Salud Carlos III (ISCIII,22); and Infections estimated with the REMEDID algorithm23 from (2) official COVID-19 deaths22, and (3) excess of all-causes deaths (ED) from European Mortality Monitoring surveillance system (MoMo,24). The REMEDID-derived infection data are more realistic than official infection data since they assimilate seroprevalence studies data25 and known dynamics of COVID-19 (see23, for further discussion). As the last national longitudinal seroprevalence study in Spain finished on 29 November 2021, our REMEDID time series has been estimated up to that date. This is not a limitation for this study since only data up to March 2020 will be used (see next section).
Intrinsic growth rate
At the beginning of an outbreak the infections, I(t), increase exponentially12,16 and can be fitted to the model
where \(\varepsilon \left( t \right)\) accounts for errors in the fitting; t is time; a is a positive number determining the point where the function crosses the ordinate axis, and then depends on where the origin of time has been set; and r is a positive number called intrinsic growth rate or Malthusian number, that defines the increasing rate of the exponential growth. r is usually the first property that epidemiologists estimate in an outbreak. The higher the r, the higher the speed in the increase of cases. When comparing diseases, r is an indicator of contagiousness, as is R0. In fact, with enough information about the latent and infectious periods, r (t−1 units) can be used to estimate R0 (dimensionless), although the relationship is not simple26. In the latent period (exposed in a Susceptible-Exposed-Infected-Recovered (SEIR) model), an infected individual cannot produce a secondary infection, unlike in the infectious period, where secondary infections may be produced.
When estimating r, it must be kept in mind that I(n) (Fig. 1a), where n denotes time discretized in days, increases exponentially during a short period of time. Consequently, the first problem is to figure out the latest day, n0, before I(n) will abandon the strictly exponential growth because of the diminishing of the number of susceptible individuals. To estimate n0, we use the property that during the exponential growth I(n) is not only rising, but is accelerating with an increasing acceleration. Then, n0 is the day where the first maximum of I″(n), the second (discrete) derivative of I(n), is reached. For REMEDID I(n), from both official and MoMo data, n0 is 23 February 2020 (Fig. 1c). Figure 2 shows the least-squares best fit of Eq. (3) to REMEDID I(n) truncated at n0, whose parameters are:
-
(1)
a = 11.86 (95% CI = [11.01, 12.70]) and r = 0.1592 (95% CI = [0.1576, 0.1609]), when MoMo ED are used;
-
(2)
a = 10.11 (95% CI = [9.25, 10.96]) and r = 0.1591 (95% CI = [0.1571, 0.1610]), when official deaths are used.
(a) Daily new infections: black thin line reflects official data, and thick black line is its 7-days moving average; red and blue lines are infections inferred from REMEDID methodology applied to MoMo excess of dead and to official COVID-19 deaths, respectively. (b) and (c) are the first and second discrete derivative of time series shown in (a). Official I′(n) (I″(n)) is estimated from the 7-days running mean of official I(n) (I′(n)). Panels (b) and (c) show the smoothed versions of I′(n) and I″(n), respectively.
Daily new infections from official data (black dots) for a period of time of 65 days (1 January to 5 March 2020) and inferred from REMEDID applied to MoMo excess of dead (red dots) and official COVID-19 deaths (blue dots) for a period of time of 46 days (January 9 to February 23). Solid lines are the exponential fitting (Eq. 1) to them.
Considering the Bonferroni correction, the difference between the two estimates of r has a CI = [− 0.0034, 0.0038], which has at least a 90% of confidence level. Since the CI includes the value 0, there is no evidence that these two parameters are different. Besides, a linearization of the model allows to perform a contrast of hypothesis on r, that confirms that there is no significant discrepancy between the two estimates of r. Then, REMEDID I(n) will be estimated from MoMo ED hereafter. Applying the same hypothesis for contrast, it can be observed that the a parameters are significatively different. However, since a value is not relevant to determine the growth rate, which is our aim here, we will not discuss its estimated values. If the same analysis were carried out with official I(n), which were not reliable at the beginning of the pandemic, we would get r = 0.2322 (95% CI = [0.2266, 0.2377]) and the end of the exponential growth on 5 March 2020. This value is significantly different, at least at 90% confidence level after Bonferroni correction, from the r estimated from any REMEDID I(n) since the CI of their differences do not include the 0. A contrast of hypothesis confirms this discrepancy. Note that despite the larger value of r from official I(n) the fitted exponential is smaller than those estimated from REMEDID I(n) (Fig. 2) because of the horizontal shift due to differences in the a parameter. The end of the exponential growth has been estimated from 7-days running averaged versions of I(n), I′(n), and I″(n) (Fig. 1a–c respectively). It has to be said that at the beginning of the outbreak, the official data underestimated the number of infections due to the low sampling capability.
Estimates of R 0
Generation time
During the infectious period, an infected individual may produce a secondary infection. However, the individual’s infectiousness is not constant during the infectious period, but it can be approximated by the probability distribution of the generation time (GT), which accounts for the time between the infection of a primary case and the infection of a secondary case. Unfortunately, such distribution is not as easy to estimate as that of the serial interval, which accounts for the time between the onset of symptoms in a primary case to the onset of symptoms of a secondary case. This is because the time of infection is more difficult to detect than the time of symptoms onset. Ganyani et al.27 developed a methodology to estimate the distribution of the GT from the distributions of the incubation period and the serial interval. Assuming an incubation period following a gamma distribution with a mean of 5.2 days and a standard deviation (SD) of 2.8 days, they estimated the serial interval from 91 and 135 pairs of documented infector-infectee in Singapore and Tianjin (China). Then, they found that the GT followed a gamma distribution with mean = 5.20 (95% CI = [3.78, 6.78]) days and SD = 1.72 (95% CI = [0.91, 3.93]) for Singapore (hereafter GT1), and with mean = 3.95 (95% CI = [3.01, 4.91]) days and SD = 1.51 (95% CI = [0.74, 2.97]) for Tianjin (hereafter GT2). Ng et al.28 applied the same methodology to 209 pairs of infector-infectee in Singapore and determined a gamma distribution with mean = 3.44 (95% CI = [2.79, 4.11]) days and SD 2.39 (95% CI = [1.27, 3.45]; hereafter GT3). Figure 3 shows the probability density functions (PDF) of such distributions, fGT. The differences between them are remarkable. For example, the 54.5%, 81.0%, and 80.7% of the contagions are produced in a pre-symptomatic stage (in the first 5.2 days after primary infection) assuming GT1, GT2, and GT3, respectively.
Probability density function of the generation time distribution, fGT(t), of GT1 (blue line; Singapore27), GT2 (yellow line; Tianjin27), GT3 (red line; Singapore28), and GTth (black line; theoretical distribution). Bars are the discretized version, \(\widetilde{{f_{GT} }}\left( n \right)\), of the PDF of GTth.
Theoretically, assuming that the incubation periods of two individuals are independent and identically distributed, which is quite plausible, the expected/mean values of the GT and the serial interval should be equal29,30. The mean of the serial interval is easier to estimate than that of the GT. For that reason, we assume a mean serial interval as estimated from a meta-analysis of 13 studies involving a total of 964 pairs of infector-infectee, which is 4.99 days (95% CI = [4.17, 5.82])31, is more reliable than the aforementioned means of the GT. This value is within the error estimates of the means of GT1 and GT2, but not for GT3. Then, we construct a theoretical distribution for the GT that follows a gamma distribution (hereafter GTth) with mean = 4.99 days and SD = 1.88 days. This theoretical distribution can be seen in Fig. 3 and approximates the average PDF of three gamma distributions with mean = 4.99 and the SD of GT1, GT2, and GT3. We assume a conservative CI = [1.51, 2.39] for the theoretical SD, defined with the minimum and maximum SD values of GT1, GT2, and GT3. GTth shows 63.1% of pre-symptomatic contagions.
R 0 from r
In theory, the basic reproduction number R0 can be estimated as far as the intrinsic growth rate r, and the distributions of both the latent and infectious periods are known26,32,33,34. The latent period accounts for the period during which an infected individual cannot infect other individuals. It is observed in diseases for which the infectious period starts around the end of the incubation period, as happened with influenza35 and SARS36. However, from Fig. 3 it is inferred that COVID-19 is transmissible from the moment of infection, and we will assume a null latent period. Then, if the GT follows a gamma distribution, R0 can be estimated from the formulation of Anderson and Watson32, which was adapted to null latent periods by Yan26 as
where meanGT is the mean GT and shapeGT is one of the two parameters defining the gamma distribution, which can be estimated as
For GTth, we get R0 = 1.50 (CI = [1.41, 1.61]) for REMEDID I(n) and R0 = 1.76 (CI = [1.60, 1.94]) for official I(n). For the other three GT distributions, R0 ranges from 1.39 (CI = [1.27, 1.58]) to 1.51 (CI = [1.34, 1.80]) for REMEDID I(n) and from 1.59 (CI = [1.40, 1.88]) to 1.78 (CI = [1.51, 2.23]) for official I(n) (Table 1). In all cases, R0 from GTth are within those from the three known GT distributions and indistinguishable from them within the error estimates. The lower (upper) bound of the CI is estimated as the minimum (maximum) R0 obtained from all the possible combinations of 100 evenly spaced values covering the CI of r, meanGT and SDGT. Then, following the Bonferroni correction, the reported CI present at least a 85% of confidence level for GT1, GT2, and GT3, but it cannot be assured for GTth since the CI of its SD is unknown. In general, all these R0 estimates are lower than those summarised by Park et al.20.
Alternatively, R0 can be estimated by applying the Euler–Lotka equation29,33,
In this case, we get values closer to previous estimates20. In particular, for GTth, we get R0 = 2.12 (CI = [1.81, 2.48]) for REMEDID I(n) and R0 = 2.92 (CI = [2.28, 3.75]) for official I(n). For the other three GT distributions, R0 ranges from 1.63 (CI = [1.43, 1.90]) to 2.21 (CI = [1.59, 2.95]) for REMEDID I(n) and from 1.97 (CI = [1.59, 2.54]) to 3.11 (CI = [1.84, 4.90]) for official I(n) (Table 1). The CI are estimated as in Eq. (4).
R0 from a dynamical model
We designed a dynamic model with Susceptible-Infected-Recovered (SIR) as stocks that accounts for the infectiousness of the infectors. Such a model is a generalisation of the Susceptible-Exposed-Infected-Recovered (SEIR) model37. Births, deaths, immigration and emigration are ignored, which seems reasonable since the timescale of the outbreak is too short to produce significant demographic changes. For the sake of simplicity, the recovered stock includes recoveries and fatalities, and it is denoted as R(t). A random mixing population is assumed, that is a population where contacts between any two people are equally probable. Time is discretized in days, so the real time variable t is replaced by the integer variable n. As a consequence, the derivatives in the differential equations defining the dynamic model explained below are discrete derivatives.
The size of the population is fixed at N = 100,000, and then, for any day n we get
where \(\tilde{S}\left( n \right)\), \(\tilde{I}\left( n \right)\), and \(\tilde{R}\left( n \right)\) are the discretized versions of S(t), I(t), and R(t) and \(\tilde{I}\) is assumed to be null for negative integers. The summation is a consequence of the infectiousness, which is approximated according to the GT, whose PDF is discretized as
from n = 1 to 20. Figure 3 shows \(\widetilde{{f_{GT} }}\left( n \right)\) for GTth. Truncating at n = 20 accounts for 99.99% of the area below the PDF of all the GT. Then, an infected individual at day n0 is expected to produce on average
infections n days later, where \(\widetilde{{R_{e} }}\left( n \right)\) is the discretized version of Re(t). From this expression, it is obvious that values of \(\widetilde{{R_{e} }}\left( n \right) < 1\) will produce a decline of infections. Conversely, infections at day n0 are produced by all individuals infected during the previous 20 days as
whose continuous version has been reported in previous studies29,38. The expression in brackets is called total infectiousness of infected individuals at day n039. According to Eq. (1), Eq. (10) can be expressed in terms of R0 as
As we want a dynamic model capable of providing \(\tilde{I}\left( {n_{0} } \right)\) from the stocks at time step n0 − 1, we replaced \(\tilde{S}\left( {n_{0} } \right)\) by \(\tilde{S}\left( {n_{0} - 1} \right)\) in Eq. (11). This assumption makes sense in a discrete domain since the infections at time n0 take place in the susceptible population at time n0 − 1. Then, assuming that all stocks are set to zero for negative integers, our dynamic model can be expressed in terms of Eq. (7) and the following differential equations:
where \(\delta \tilde{I}\), \(\delta \tilde{S}\), and \(\delta \tilde{R}\) are the (discrete) derivatives of \(\tilde{I}\), \(\tilde{S}\), and \(\tilde{R}\), respectively. Applying the initial conditions \(\tilde{S}\left( 0 \right) = N - 1\), \(\tilde{I}\left( 0 \right) = 1\), and \(\tilde{R}\left( 0 \right) = 0\), it is assumed that the outbreak was produced by only one infector. The latter is not true in Spain, since several independent introductions of SARS-CoV-2 were detected40. However, for modelling purposes it is equivalent to introducing a single infection at day 0 or M infections produced by the single infection n days later. Then, the date of the initial time n = 0 is accounted as a parameter date0, which is optimised, as well as R0, to minimise the root-mean square of the residual between the model simulated \(\tilde{I}\left( n \right)\) and the REMEDID and official I(n) for the period from date0 to n0.
The model was implemented in Stella Architect software v2.1.1 (www.iseesystems.com) and exported to R software v4.1.1 with the help of deSolve (v1.28) and stats (v4.1.1) packages, and the Brent optimisation algorithm was implemented. For REMEDID I(n) and GTth, we obtained date0 = 13 December 2019 and R0 = 2.71 (CI = [2.33, 3.15]). Optimal solutions combine lower/higher R0 and earlier/later date0 (Fig. 4), which highlights the importance of providing an accurate first infection date to estimate R0. When the other three GT distributions were considered, we obtained similar date0, ranging from 12 to 17 December 2019, and R0 values ranging from 2.08 (CI = [1.86, 2.42]) to 2.85 (CI = [2.05, 3.25]; see Table 1). For official infections, date0 was set to 1 January 2020 for all cases, and R0 ranged from 1.81 (CI = [1.64, 2.07]) to 2.41 (CI = [1.80, 2.91]). The CI are estimated as in Eq. (4).
Root-mean square (RMS) of the residuals between infections from the model, which depends on date0 (x-axis) and R0 (y-axis), and REMEDID (from MoMo ED) and official infections. Parameters optimizing the model are highlighted in purple. RMS larger than 1275 (left panel) and 103 (right panel) are saturated in white.
Herd immunity threshold and discussion
HIT of the ancestral variant was estimated from R0 via Eq. (2) and values are shown in Table 1, which range between 28.1 (CI = [21.3, 36.7]) and 64.9% (CI = [51.2, 69.2]) for REMEDID I(n) (hereafter HITR), and between 37.0 (CI = [28.4, 46.7]) and 67.8% (CI = [45.7, 79.6]) for official I(n) (Hereafter HITO). The differences between the estimations are determined by three key factors: (1) source/quality of data; (2) GT distribution; and (3) methodology to estimate R0.
In general, official infection data are of poor quality, but if death records and seroprevalence studies were available, the REMEDID algorithm would provide more reliable infections time series23. The maximum difference between HITR and HITO is 13.1 percentage points, corresponding to the Eq. (6) estimate, although such difference is not significant within the errors estimates. Moreover, official data vary depending on the date of publication. For example, the maximum HITO is 67.7%. from data available in February 2021, and 80.1% from data available a year before, in March 2020. The latter is similar to the 80.7% published by Kwok et al.41 in March 2020, which was obviously based on data available at that time. The February 2021 version of the data is more realistic than the March 2020 one, and the REMEDID-derived infections are more realistic than both of them23. In consequence, results based on REMEDID data should be more reliable.
The most influential factor for estimating the HIT is the methodology to estimate R0, which may produce differences of ~ 30 percentage points for HITR and ~ 20 points for HITO for the same dataset and GT distribution. Such differences are significant within the error estimates for all GT in HITR and only for GTth in HITO. For each GT, the lowest HIT values were obtained from Eq. (4), but the largest HITR and HITO are obtained from the dynamic model and Eq. (6), respectively. The CI from Eq. (6) and the dynamic model are longer than those from Eq. (4), meaning that the former are more sensitive to errors in the involved parameters. Moreover, the largest errors are obtained from Eq. (6) for both HITR and HITO, although they are larger for HITO. It means that Eq. (6) is the methodology most sensitive to parameters and data quality. In general, results from Eq. (6) are reconcilable with the other two within the error estimates, but Eq. (4) and the dynamic model are only reconcilable for official data (Table 1).
The selection of a GT produces HIT differences up to 6 percentage points when R0 is estimated from Eq. (4); 18.7 from Eq. (6); and 13.7 from the dynamic model, although in no case are significantly different within the error estimates. It is more difficult to estimate the GT than the serial interval. For that reason, many studies approximate the GT by a serial interval (e.g.39,41). However, though GT and serial interval have the same mean, serial interval presents a larger variance30, which will underestimate R0 when using Eq. (6) 29. HIT values from Eq. (4) for any GT are included in the CI obtained for the other GT. On the contrary, although all the CI estimated from Eq. (6) overlap among them, only some HIT values are included in the CI estimated for other GT. This is also the situation for the HIT estimated from the dynamic model.
The influential factors should be kept in mind when interpreting R0 estimates. For example, Locatelli et al.21 estimated an average R0 of 2.2 (CI = [1.9, 2.6]) for Western Europe by using official data available in September 2020, a theoretical approximation of GT, and Eq. (6). For any GT in Table 1 it can be observed that: (1) official data produces the highest R0 values for Eq. (6) with respect to Eq. (4), and the dynamic model; and that (2) the more realistic REMEDID data also produces lower R0 values when Eq. (6) is used. Then, it could be conjectured that the R0 reported by Locatelli et al.21 is in the upper bound of all the possible R0 estimates for Western Europe.
In summary, accurately estimating HIT is quite complicated. In any case, assuming that REMEDID-derived infection data are more accurate than official data, 70% seems to be a good upper bound of HIT for the ancestral variant. However, the upper bound increases to 80% (accounting for the CI) if we rely on official data. Besides, the most important impediment to determine the value of the HIT is that it is variable in time. The more transmissible new SARS-CoV-2 variants present higher Re, and in consequence a higher (theoretical) associated R0 and higher HIT values. For example, the B.1.1.7 lineage (also known as alpha variant), which was first detected in England in September 202042, and thereafter rapidly spread around the world. In Spain, at the beginning of January 2021, the alpha variant was ~ 30% of the circulating SARS-CoV-2 variants, but it was over 80% from March to May 202143. On the other hand, the B.1.617.2 lineage (also known as delta variant), first detected in India in December 202044, has represented over 95% of the SARS-CoV-2 variants in Spain from late July to at least up to October 202143. Both alpha and delta displaced the previous variants because of their higher transmissibly. Although the R0 cannot be directly estimated for these variants since they appeared in the middle of the pandemic, the Re can. It has been estimated that the Re of the alpha variant is ~ 70% higher than in previous existing variants45. On the other hand, the Re of the delta variant is also ~ 70% higher than in alpha variant46. Following Eq. (1) and assuming that the variations of Re from one variant to another are not produced by changes in the control measures, it can be inferred that the R0 of the alpha and delta variants are 70% and 189% higher than the ancestral variant, respectively. Therefore, if we take the highest estimate of R0 in Table 1 (R0 = 3.11, CI = [1.84, 4.90]; for GT1, Eq. (6), official data) as an upper bound of the R0 of the ancestral variant, we get that 8.99 (CI = [5.32, 14.16]) is an upper bound estimate of the delta variant R0. In that case, we can conclude that an upper bound of the HIT at present in Spain is 88.9% (CI = [81.2, 92.9]). For a more realistic upper bound, we could alternatively take the maximum R0 for REMEDID data in Table 1 (R0 = 2.85, CI = [2.05, 3.25]; for GT1, dynamic model) as an upper bound, which would produce R0 = 8.24 (CI = [5.92, 9.39]) and HIT = 87.9% (CI = [83.1, 89.4%]) as upper bound for the delta variant, in agreement with previous estimates46. Then, a HIT of 90% seems to be realistic for Spain with a predominant delta variant as in October 2021.
The presented results are valid for a randomly mixing population with a spread dynamic similar to Spain as a whole. However, even Spanish regions show different dynamics between themselves23, which may lead to specific HIT values for each region. It should be kept in mind that none of the three vaccines administered in Spain are able to completely prevent the transmission of the virus. Then, even with a 90% of the population vaccinated, the HIT will probably not be reached. However, it is true that the risk of infection is significatively reduced for vaccinated susceptible individuals6,7, which directly reduces the R0. Besides, in case of infection, the transmission of the virus is also reduced8,9,10, which modifies the associated GT, and reduces the R0 and the HIT of a vaccinated population. So, even if transmission is not completely prevented by vaccines, the greater the proportion of the vaccinated population, the lower the HIT. Therefore, it is expected that the HIT of a highly vaccinated population will be below the estimated 90% upper bound. However, all this may change with the emergence and spread of new variants with re-infection capacity47. In any case, even if the HIT is reached, it will not be the panacea. First, if HIT is reached in most places in a country but there are some specific regions or population subgroups in a region with a percentage of immune individuals below HIT, local outbreaks will be possible for those regions or subgroups. Second, the final size of an epidemic in a randomly mixing population with HIT = 70% and 90% is reached at 95.9% and 99.9% of infections, respectively15,37. This means that if the ancestral variant would have not been replaced, the decreasing rate of infections after reaching a HIT of 70% may still produce a non-negligible 25.9% of infections, that is 12.2 million infections in Spain. Third, interpretation of HIT values must be done carefully and overoptimistic messages should be avoided as has been learnt from Manaus in the Brazilian state of Amazonas. In October 2020, it was thought that Manaus had reached the HIT with 76% of infected population48, which led to a relaxation of the control measures. However, either because the percentage of infected population was not accurately estimated or because the new SARS-CoV-2 P.1 variant was capable of re-infecting, Manaus had a second wave in January 2021 with a higher mortality rate than in the first one49. Therefore, health authorities should strictly ensure an adaptive and proactive management of the new situation after theoretical herd immunity is reached.
References
Our world in data, https://ourworldindata.org/coronavirus-data-explorer). Accessed 1 Apr 2021.
Defunciones según la Causa de Muerte—Avance enero-mayo de 2019 y de 2020. Notas de prensa del Instituto Nacional de Estadística (2020). https://www.ine.es/dyngs/INEbase/es/operacion.htm?c=Estadistica_C&cid=1254736176780&menu=ultiDatos&idp=1254735573175. Accessed 16 Feb 2021.
Kung, S. et al. Underestimation of COVID-19 mortality during the pandemic. ERJ Open Res. 7(1), 00766. https://doi.org/10.1183/23120541.00766-2020 (2021).
Modi, C., Böhm, V., Ferraro, S., Stein, G. & Seljak, U. Estimating COVID-19 mortality in Italy early in the COVID-19 pandemic. Nat. Commun. 12, 2729. https://doi.org/10.1038/s41467-021-22944-0 (2021).
European Medicines Agency, https://www.ema.europa.eu/en/human-regulatory/overview/public-health-threats/coronavirus-disease-covid-19/treatments-vaccines/covid-19-vaccines. Accessed 1 Apr 2021.
Hall, V. J. et al. COVID-19 vaccine coverage in health-care workers in England and effectiveness of BNT162b2 mRNA vaccine against infection (SIREN): A prospective, multicentre, cohort study. The Lancet. 397(10286), 1725–1735. https://doi.org/10.1016/S0140-6736(21)00790-X (2021).
Thompson, M. G. et al. Interim estimates of vaccine effectiveness of BNT162b2 and mRNA-1273 COVID-19 vaccines in preventing SARS-CoV-2 infection among health care personnel, first responders, and other essential and frontline workers—Eight US Locations, December 2020–March 2021. MMWR Morb. Mortal. Wkly. Rep. 70, 495–500. https://doi.org/10.15585/mmwr.mm7013e3 (2021).
Levine-Tiefenbrun, M. et al. Initial report of decreased SARS-CoV-2 viral load after inoculation with the BNT162b2 vaccine. Nat. Med. 27, 790–792. https://doi.org/10.1038/s41591-021-01316-7 (2021).
Emary, K. R. W. et al. Efficacy of ChAdOx1 nCoV-19 (AZD1222) vaccine against SARS-CoV-2 variant of concern 202012/01 (B.1.1.7): An exploratory analysis of a randomised controlled trial. The Lancet. 397(10282), 1351–1362. https://doi.org/10.1016/S0140-6736(21)00628-0 (2021).
Shah, A. S. V. et al. Effect of vaccination on transmission of SARS-CoV-2. N. Engl. J. Med. https://doi.org/10.1056/NEJMc2106757 (2021).
Fine, P., Eames, K. & Heymann, D. L. “Herd immunity”: A rough guide. Clin. Infect. Dis. 52(7), 911–916. https://doi.org/10.1093/cid/cir007 (2011).
Anderson, R. M. & May, R. M. Infectious Diseases of Humans: Dynamics and Control (Oxford University Press, 1991).
Hannon, B. & Ruth, M. Dynamic Modeling of Diseases and Pests (Springer, 2009).
Heesterbeek, J. A. P. A brief history of R0 and a recipe for its calculation. Acta. Biotheor. 50, 189–204 (2002).
Kermack, W. O. & McKendrick, A. G. A contribution to the mathematical theory of epidemics. Proc. R. Soc. Lond. A. 115, 700–721 (1927).
Keeling, M. & Rohani, P. Modeling Infectious Diseases in Humans and Animals (Princeton University Press, 2008). https://doi.org/10.2307/j.ctvcm4gk0.
Coburn, B. J., Wagner, B. G. & Blower, S. Modeling influenza epidemics and pandemics: Insights into the future of swine flu (H1N1). BMC Med. 7, 30. https://doi.org/10.1186/1741-7015-7-30 (2009).
Bauch, C. T., Lloyd-Smith, J. O., Coffee, M. P. & Galvani, A. P. Dynamically modeling SARS and other newly emerging respiratory illnesses: Past, present, and future. Epidemiology 16, 791–801. https://doi.org/10.1097/01.ede.0000181633.80269.4c (2005).
Breban, R., Riou, J. & Fontanet, A. Interhuman transmissibility of Middle East respiratory syndrome coronavirus: Estimation of pandemic risk. Lancet https://doi.org/10.1016/S0140-6736(13)61492-0 (2013).
Park, M., Cook, A. R., Lim, J. T., Sun, Y. & Dickens, B. L. A systematic review of COVID-19 epidemiology based on current evidence. J. Clin. Med. 9, 967. https://doi.org/10.3390/jcm9040967 (2020).
Locatelli, I., Trächsel, B. & Rousson, V. Estimating the basic reproduction number for COVID-19 in Western Europe. PLoS ONE 16(3), e0248731. https://doi.org/10.1371/journal.pone.0248731 (2021).
Instituto de Salud Carlos III, https://cnecovid.isciii.es/covid19/#documentaci%C3%B3n-y-datos. Accessed 12 Feb 2021.
García-García, D. et al. Retrospective methodology to estimate daily infections from deaths (REMEDID) in COVID-19: The Spain case study. Sci. Rep. 11, 11274. https://doi.org/10.1038/s41598-021-90051-7 (2021).
European Mortality Monitoring surveillance system, http://www.euromomo.eu. Accessed 16 Feb 2021.
Pollán, M. et al. Prevalence of SARS-CoV-2 in Spain (ENE-COVID): A nationwide, population-based seroepidemiological study. The Lancet. 396(10250), 535–544. https://doi.org/10.1016/S0140-6736(20)31483-5 (2020).
Yan, P. Separate roles of the latent and infectious periods in shaping the relation between the basic reproduction number and the intrinsic growth rate of infectious disease outbreaks. J. Theor. Biol. 251, 238–252. https://doi.org/10.1016/j.jtbi.2007.11.027 (2008).
Ganyani, T. et al. Estimating the generation interval for coronavirus disease (COVID-19) based on symptom onset data. Euro Surveill. 25, 2000257. https://doi.org/10.2807/1560-7917.ES.2020.25.17.2000257 (2020).
Ng, S. et al. Estimating transmission parameters for COVID-19 clusters by using symptom onset data, Singapore, January–April 2020. Emerg. Infect. Dis. 27(2), 582–585. https://doi.org/10.3201/eid2702.203018 (2021).
Britton, T. & Tomba, G. S. Estimation in emerging epidemics: Biases and remedies. J. R. Soc. Interface 16, 20180670. https://doi.org/10.1098/rsif.2018.0670 (2019).
Lehtinen, S., Ashcroft, P. & Bonhoeffer, S. On the relationship between serial interval, infectiousness profile and generation time. J. R. Soc. Interface. 18, 20200756. https://doi.org/10.1098/rsif.2020.0756 (2021).
Fonfría, E. S. et al. COVID-19 epidemiological parameters for clinical and mathematical modeling: Mini-review and meta-analysis from Asian studies during early phase of pandemic. Front. Med. https://doi.org/10.1101/2020.06.17.20133587v1 (2021).
Anderson, D. & Watson, R. On the spread of a disease with gamma distributed latent and infectious periods. Biometrika 67(1), 191–198. https://doi.org/10.1093/biomet/67.1.191 (1980).
Wallinga, J. & Lipsitch, M. How generation intervals shape the relationship between growth rates and reproductive number. Proc. R. Soc. B. 274, 599–604 (2007).
Roberts, M. G. & Heesterbeek, J. A. P. Model-consistent estimation of the basic reproduction number from the incidence of an emerging infection. J. Math. Biol. 55, 803–816 (2007).
Lau, L. L. et al. Viral shedding and clinical illness in naturally acquired influenza virus infections. J Infect Dis. 201(10), 1509–1516. https://doi.org/10.1086/652241 (2010).
Peiris, J. S. et al. Clinical progression and viral load in a community outbreak of coronavirus-associated SARS pneumonia: A prospective study. Lancet 361(9371), 1767–1772 (2003).
Ma, J. & Earn, D. J. D. Generality of the final size formula for an epidemic of a newly invading infectious disease. Bull. Math. Biol. 68, 679–702. https://doi.org/10.1007/s11538-005-9047-7 (2006).
Park, S. W. et al. Forward-looking serial intervals correctly link epidemic growth to reproduction numbers. PNAS 118(2), e2011548118. https://doi.org/10.1073/pnas.2011548118 (2021).
Cori, A., Ferguson, N. M., Fraser, C. & Cauchemez, S. A new framework and software to estimate time-varying reproduction numbers during epidemics. Am. J. Epidemiol. 178(9), 1505–1512. https://doi.org/10.1093/aje/kwt133 (2013).
Gómez-Carballa, A. et al. Phylogeography of SARS-CoV-2 pandemic in Spain: A story of multiple introductions, micro-geographic stratification, founder effects, and super-spreaders. Zool. Res. 41(6), 605–620. https://doi.org/10.24272/j.issn.2095-8137.2020.217 (2020).
Kwok, K. O., Lai, F., Wei, W. I., Wong, S. Y. S. & Tang, J. W. T. Herd immunity—Estimating the level required to halt the COVID-19 epidemics in affected countries. J. Infect. 80(6), e32–e33. https://doi.org/10.1016/j.jinf.2020.03.027 (2020).
Public Health England, “Investigation of novel SARS-COV-2 variant: Variant of Concern 202012/01” (2020); www.gov.uk/government/publications/investigation-of-novel-sars-cov-2-variant-variant-of-concern-20201201.
Actualización de la situación epidemiológica de las variantes de SARS-CoV-2 de preocupación (VOC) e interés (VOI) en salud pública en España, 18 de octubre de 2021, https://www.mscbs.gob.es/profesionales/saludPublica/ccayes/alertasActual/nCov/documentos/COVID19_Actualizacion_variantes_20211018.pdf. Accessed 22 Oct 2021.
Cherian S, Potdar V, Jadhav S, Yadav P, Gupta N et al. Convergent evolution of SARS-CoV-2 spike mutations, L452R, E484Q and P681R, in the second wave of COVID-19 in Maharashtra, India. BioRxiv. 2021. Preprint at https://doi.org/10.1101/2021.04.22.440932.
Davies, N. G. et al. Estimated transmissibility and impact of SARS-CoV-2 lineage B.1.1.7 in England. Science 372, 6538. https://doi.org/10.1126/science.abg3055 (2021).
Liu, Y. & Rocklöv, J. The reproductive number of the Delta variant of SARS-CoV-2 is far higher compared to the ancestral SARS-CoV-2 virus. J. Travel Med. 28(7), taab124. https://doi.org/10.1093/jtm/taab124 (2021).
Uriu K, Kimura I, Shirakawa K, Takaori-Kondo A, Nakada T, Kaneda A, The genotype to phenotype Japan (G2P-Japan) consortium, So Nakagawa, Kei Sato. Ineffective neutralization of the SARS-CoV-2 Mu variant by convalescent and vaccine sera. bioRxiv; 2021. https://doi.org/10.1101/2021.09.06.459005
Buss, L. F. et al. Three-quarters attack rate of SARS-CoV-2 in the Brazilian Amazon during a largely unmitigated epidemic. Science 371, 288–292. https://doi.org/10.1126/science.abe9728 (2021).
Taylor, L. Covid-19: Is Manaus the final nail in the coffin for natural herd immunity?. BMJ 372, n394. https://doi.org/10.1136/bmj.n394 (2021).
Funding
This work was supported by the University of Alicante [COVID-19 2020-41.30.6P.0016 to CB] and the Montgó-Dénia Research Station (Agreement Ajuntament de Dénia-O.A. Parques Nacionales, Ministry of the Environment—Generalitat Valenciana -Conselleria de Agricultura, Desarrollo Rural, Emergencia Climática y Transición Ecológica, Spain, Spain) [2020-41.30.6O.00.01 to CB].
Author information
Authors and Affiliations
Contributions
D.G-G. and C.B. designed the study, and D.G-G. wrote the first version of the manuscript. D.G-G., I.V. and E.M. performed the mathematical analysis. All authors reviewed the last version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
García-García, D., Morales, E., Fonfría, E.S. et al. Caveats on COVID-19 herd immunity threshold: the Spain case. Sci Rep 12, 598 (2022). https://doi.org/10.1038/s41598-021-04440-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-04440-z
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.