Abstract
Cryptococcus deneoformans is an opportunistic fungal pathogen that infects the lungs via airborne transmission and frequently causes fatal meningoencephalitis. Claudins (Cldns), a family of proteins with 27 members found in mammals, form the tight junctions within epithelial cell sheets. Cldn-4 and 18 are highly expressed in airway tissues, yet the roles of these claudins in respiratory infections have not been clarified. In the present study, we analyzed the roles of Cldn-4 and lung-specific Cldn-18 (luCldn-18) in host defense against C. deneoformans infection. luCldn-18-deficient mice exhibited increased susceptibility to pulmonary infection, while Cldn-4-deficient mice had normal fungal clearance. In luCldn-18-deficient mice, production of cytokines including IFN-γ was significantly decreased compared to wild-type mice, although infiltration of inflammatory cells including CD4+ T cells into the alveolar space was significantly increased. In addition, luCldn-18 deficiency led to high K+ ion concentrations in bronchoalveolar lavage fluids and also to alveolus acidification. The fungal replication was significantly enhanced both in acidic culture conditions and in the alveolar spaces of luCldn-18-deficient mice, compared with physiological pH conditions and those of wild-type mice, respectively. These results suggest that luCldn-18 may affect the clinical course of cryptococcal infection indirectly through dysregulation of the alveolar space microenvironment.
Similar content being viewed by others
Introduction
The two sister species, Cryptococcus neoformans (formerly C. neoformans var. grubii, serotype A) and C. deneoformans (formerly C. neoformans var. neoformans, serotype D), are yeast-type fungal pathogens with thick capsules composed of polysaccharides such as glucuronoxylomannan and galactoxylomannan1. These fungi infect the lungs via airborne transmission and cause life-threatening meningoencephalitis in patients with impaired cell-mediated immunity such as acquired immunodeficiency syndrome2. As both of these Cryptococcus spp. are intracellularly growing yeasts, they are eradicated primarily by the cellular immune mechanism, which is critically regulated by the Th1-Th2 immune balance2,3,4,5,6. The Th1-mediated immune response supports host defense by inducing production of nitric oxide (NO), which enhances macrophages’ ability to kill Cryptococcus, and by promoting the formation of granulomas at infection sites, which prevents the spread of this fungal pathogen to the surrounding lung tissues and to the CNS6,7,8,9. The Th2 immune response, in contrast, disturbs host defense by suppressing macrophages’ killing of Cryptococcus and granuloma formation6,7,10. In a recent study, we demonstrated that IL-17A regulates Th1-mediated host defense against cryptococcal infection13; other studies have reported other roles for this cytokine12,13,14,15,16.
Claudins (Cldns), cell–cell adhesion molecules with four transmembrane domains located at the tight junctions (TJs) between the cells in epithelial cell sheets, play various roles in the formation of paracellular barriers and channels17,18,19,20. Twenty-seven members of the Cldn protein family occur in mammals, and each of these subtypes is expressed in an organ-specific and developmental stage-dependent manner21,22,23. Ohta et al. have reported that Cldn-1, 3, 4, 7, and 10 are components of airway TJs, whereas Cldn-5 and 18 are expressed in alveolar TJs22. In addition, LaFemina et al. have demonstrated that Cldn-3, 4, and 18 are the predominant claudins expressed in primary rat alveolar epithelial cells23. Cldn-4 has been reported to function as a paracellular sodium barrier and to play an undefined role in protecting the lungs from acute injury, although its contribution to normal lung physiology is limited24. High expression of Cldn-18 has been described in lung and stomach epithelium, with lung-specific (Cldn-18.1 or luCldn-18) and stomach-specific (Cldn-18.2 or stCldn-18) isoforms of Cldn-18 regulated by alternative promoter usage25,26. luCldn18 has been shown to be involved in alveolarization, alveolar epithelial permeability, and alveolar fluid clearance27,28. The role of these claudins in respiratory infections has not been clarified, however. To correct this, in the present study, we investigated the role of claudins, especially Cldn-4 and luCldn-18, in host protection and immune response against infection with C. deneoformans using mice genetically lacking either Cldn-4 or luCldn-18.
Materials and methods
Ethical statement
This study was performed in strict accordance with the Fundamental Guidelines for Proper Conduct of Animal Experiments and Related Activities in Academic Research Institutions under the jurisdiction of the Ministry of Education, Culture, Sports, Science and Technology in Japan, 2006. All experimental procedures involving animals followed the Regulations for Animal Experiments and Related Activities at Tohoku University, Sendai; Osaka University, Osaka; and RIKEN, Japan and were approved by the Institutional Animal Care and Use Committee at Tohoku University, Osaka University, and RIKEN Kobe branch. All experiments were performed under anesthesia, and all efforts were made to minimize the suffering of the animals. The study was carried out in compliance with the ARRIVE guidelines (https://arriveguidelines.org/).
Establishment of Cldn-4 and lung-specific Cldn-18-deficient mice
The Cldn-4 targeting construct was made using the Pnt1.1 vector, which contains a neomycin-resistance gene (provided by M. Okabe, University of Osaka, Osaka, Japan) (Fig. S1). The 5′ arm and 3′ arm were amplified by PCR using a BacPac vector, which included the Cldn-4 gene from C57BL/6, as the template. The amplified DNA fragments were confirmed by sequencing and inserted into the upstream and downstream regions of the PGK-Neo cassette, respectively. The targeting construct was linearized and electroporated into 129-derived embryonic stem cells. After selection with G418, drug-resistant clones were picked up and screened by PCR and Southern blot analysis. Cldn-4-targeted embryonic stem clone was microinjected into blastocysts derived from C57BL/6 mice and transferred to pseudopregnant females. The resulting chimeric mice were bred with C57BL/6 mice to obtain germline transmitted heterogenous (+/−) mice. The resulting Cldn4+/− mice were intercrossed to obtain Cldn4−/− mice. Genotyping was performed by PCR (CL4-WTF2: 5′-CAGTTGCCCACCTCGTAGCAACGAC-3′, CL4-WTR2; 5′-ATCCACCAGCAATTTGGATGTAAGC-3′, CL4-neo; 5′-CCGGTGGATGTGGAATGTGTGCGAGGCC-3′). The product of the wild-type allele is ~ 300 bp, and the targeted allele yields a ~ 500-bp product. For lung-specific claudin-18 (Cldn18.1)-deficient mice, the chimeric mice were generated as described elsewhere (http://www2.clst.riken.jp/arg/methods.html) and crossed with C57BL/6 mice to obtain heterozygous lung-specific Cldn18.1 flox/wt mice (Accession No. CDB0809K: http://www2.clst.riken.jp/arg/mutant%20mice%20list.html) (Fig. S2). Crossing these mice with CAG-Cre Tg mice29 yielded Cldn18.1+/− mice which were then intercrossed to obtain Cldn18.1−/− mice. Genotyping was performed by PCR (Cldn18-F: 5′-GTGAATCCCAGCAAGTTTTGTTATAGA-3′, Cldn18-WTR; 5′-CTCAGTGTTTGCTCATACTGTTTTCCTT-3′, Cldn18-KOR; 5′-GAATGCAAAGTGCCTATGATGC-3′). The product of the wild-type allele is ~ 310 bp, and the targeted allele yields a ~ 640-bp product.
Mice
Male or female mice at 6 to 8 weeks of age and 16 to 24 g of weight were used in the analyses. Mice were randomly allocated to the various experimental groups. All mice were kept under specific pathogen-free conditions at the Institute for Animal Experimentation, Tohoku University Graduate School of Medicine; the Osaka University Laboratory of Biological Science, Graduate School of Frontier Biosciences and Medicine; or RIKEN Kobe branch. The conditions of the breeding room were as previously described11,30,31,32,33,34,35.
Inoculation with Cryptococcus deneoformans
A strain of C. deneoformans, designated as B3501 (a kind gift from Dr. Kwong Chung, National Institute of Health, Bethesda, MD, USA) was used. The yeast cells were cultured on potato dextrose agar (Eiken, Tokyo, Japan) plates for 2–3 d before use. Mice were anaesthetized by an intramuscular injection of 0.3 mg/kg of midazolam (Fuji Phama, Tokyo, Japan) and 0.02 mg/kg of Medetomidine hydrochloride (Nippon Zenyaku Kogyo, Fukushima, Japan) and an intraperitoneal injection of 15 mg/kg of pentobarbital (Abbott Laboratory, North Chicago, IL, USA). Live C. deneoformans (1 × 106 cells, 50 μl) was inoculated into the trachea of each mouse using a 24-gauge catheter (Terumo, Tokyo, Japan).
Preparation of bronchoalveolar lavage fluids and lung interstitial homogenates
Mice were sacrificed at various timepoints after infection. Bronchoalveolar lavage fluids (BALFs) were prepared as previously described34,35. After the BALFs were collected, lung homogenates were prepared as previously described11,30,31,32,33.
Enumeration of viable C. deneoformans
Evaluation of the fungal burdens in the lungs and brains were carried out as previously described11,30,31,32,33. Briefly, the lungs and brains were homogenized separately in 5 ml (lungs) or 1 ml (brains) of distilled water through a stainless-steel mesh at room temperature. In some experiments, BALFs and lung homogenates after BAL were analyzed separately in 3 ml and 5 ml, respectively, of distilled water. These samples were diluted, inoculated on PDA plates, and cultured before the resulting colonies were counted.
Histological examination
Histopathological specimens of the lungs were obtained as previously described11,30,31,32,33,34,35. Lung specimens were stained with hematoxylin–eosin (H-E) or periodic acid-Schiff (PAS) at the Biomedical Research Core, Animal Pathology Platform of Tohoku University Graduate School of Medicine. The granulomatous area was calculated for the H&E-stained sections as the proportion of granulomatous area relative to the total lung area using ImageJ software (https://imagej.nih.gov/ij/; provided in the public domain by the National Institutes of Health).
Measurement of BALF protein
Protein concentration in BALFs was measured using a bicinchoninic acid (BCA) protein assay reagent kit (Thermo Fisher Scientific K.K., Yokohama, Japan). The detection limit was 125 μg/mL.
Flow cytometry
The BALF cells were washed three times in PBS containing 1% fetal calf serum (FCS) and 0.1% sodium azide and then stained with APC-anti-CD3ε mAb (clone 145-2C11; BioLegend), FITC-anti-CD4 mAb (clone GK1.5; BioLegend), APC/Cy7-anti-CD8 mAb (clone 53–6.7; BioLegend), and PE/Cy7-anti-NK1.1 mAb (clone PK136; BioLegend), or FITC-anti-CD11c mAb (clone N418; BioLegend), PE-anti-CD11b mAb (clone M1/70; BioLegend), PE/Cy7-anti-Gr-1 mAb (clone RB6-8C5; BioLegend), and Pacific Blue-anti-F4/80 mAb (clone BM8; BioLegend). Isotype-matched IgG was used for control staining. The stained cells were analyzed using a BD FACS Canto II flow cytometer (BD Bioscience) as previously described11,31,32,33.
Cytokine assay
BALFs were collected in 1 ml PBS. After BAL, the lungs were excised and then homogenized separately in 5 ml PBS through a stainless-steel mesh. After centrifugation, the supernatants were collected and stored at − 70 °C before use. Concentrations of each cytokine in BALF and lung homogenates were measured using an ELISA kit as previously described11,30,31,32,33,34,35.
Measurement of ion concentration and pH in BALFs
BALFs were collected in 1 ml saline from 8-week-old mice under deep anesthesia. Na+ and K+ ion concentrations in BALFs were directly measured using Na+ and K+ compact ion meters (Horiba, Kyoto, Japan) according to the manufacturer’s instructions. Cl− ion concentrations in BALFs diluted with MQ water were measured using the QuantiChrom Chloride Assay Kit (BioAssay Systems, Hayward, CA, USA). The electrical potentials were measured using the LAQAtwin compact pH meter (Horiba, Kyoto, Japan) according to the manufacturer’s instructions. A standard curve was prepared from the electrical potential of saline adjusted to pH 4, 7, and 9 with HCl or NaOH. The pH values of the BALF samples were calculated from this standard curve after the measured values were corrected for the electrical potential of saline.
In-vitro culture of C. deneoformans
C. deneoformans was cultured on PDA plates for two to three days before use. The yeast cells were suspended in RPMI 1640 medium with 20 mM HEPES and without NaHCO3 (Sigma-Aldrich, St. Louis, MO, USA), prepared at 1 × 104 cells/ml, and cultured at 37 °C for 24 h at different pH values (6.8 and 7.3). The number of organisms was counted using a hemocytometer. The yeast cells were concentrated onto microscope slides using a cytospin, stained with PAS (Muto Pure Chemicals, Tokyo, Japan), and observed under a microscope. The budding ratio of each PAS-stained specimen was estimated as the number of budding yeast cells out of 200 cells.
Cell division assay
C. deneoformans was cultured on PDA plates for two to three days before use. The yeast cells were stained with 50 μM carboxyfluorescein succinimidyl ester (CFSE) for 30 min in 10 ml PBS at 30 °C, and washed three times in PBS to remove excess CFSE. CFSE-labeled C. deneoformans was used for in vitro culture and in vivo infection by the method described above. The mean fluorescent intensity (MFI) of CFSE in the cultured yeast cells or CD45− cells in BALFs after infection was analyzed using a BD FACS Canto II flow cytometer.
Statistical analysis
Data were analyzed using JMP Pro 11.2.0 software (SAS Institute Japan, Tokyo, Japan). Data are expressed as mean ± SD. Differences between groups were examined for statistical significance using Welch’s t-test. A p value of less than 0.05 was considered significant.
Results
Effect of claudin deficiencies on host protection and inflammatory response against cryptococcal infection
To explore the effects of Cldn-4 and luCldn-18 deficiency on host defense against cryptococcal infection, Cldn-4-deficient mice, luCldn-18-deficient mice, and control mice for each claudin-deficient genotype were infected with C. deneoformans, and the growth of this fungal pathogen in the lungs and brains was examined. After cryptococcal infection, mouse body weights were not lower in Cldn-4−/− mice compared with Cldn-4+/+ mice (Fig. 1a). In addition, the fungal burdens in the lungs were not significantly different between Cldn-4+/+ and Cldn-4−/− mice on day 14 post-infection (Fig. 1b). In contrast, the lung and brain burdens of yeast cells were higher in luCldn-18−/− mice than in luCldn-18+/+ mice on days 14 and 28 post-infection (Fig. 1c-e). We also conducted a histological analysis to determine how luCldn-18 deficiency affected the inflammatory response in the lungs after cryptococcal infection. Defective alveolar formation and increased alveolar macrophage counts were observed in uninfected luCldn-18−/− mice compared to luCldn-18+/+ mice (Fig. 2a). In addition, massive multiplication of yeast cells with poor granulomatous responses was observed in luCldn-18−/− mice, whereas luCldn-18+/+ mice showed markedly fewer yeast cells with granulomatous responses (Fig. 2b, c).
Effect of claudin deficiencies on C. deneoformans infection. (a, b) Cldn-4+/+ (n = 4) and Cldn-4−/− mice (n = 5) were infected intratracheally with C. deneoformans. (a) Mouse weight is presented as weight relative to the initial weight of each mouse. (b) The numbers of live colonies in the lungs were counted on day 14 post-infection. Each symbol represents a separate mouse, and bars indicate the mean ± standard deviation (SD). (c-e) luCldn-18+/+ (day 14, n = 7; day 28, n = 7) and luCldn-18−/− mice (day 14, n = 5; day 28, n = 6) were infected intratracheally with C. deneoformans. (c) Mouse weight is presented as weight relative to the initial weight of each mouse. The numbers of live colonies in the lungs on days 14 and 28 (d) and in the brains on day 28 post-infection (e) were counted. Each symbol represents a separate mouse, and bars indicate the mean ± SD. The dashed line indicates the detection limit. *, p < 0.05. NS, not significant. There were no mice excluded from this analysis.
Effects of luCldn-18 deficiency on pulmonary histological findings after infection with C. deneoformans. (a) Lung sections of uninfected luCldn-18+/+ (n = 3) and luCldn-18−/− mice (n = 3) were stained with H-E and observed under a light microscope at ×40 (upper panels) and ×200 (under panels). Red arrows show alveolar macrophages. (b) luCldn-18+/+ and luCldn-18−/− mice were infected intratracheally with C. deneoformans. Sections of the lungs on day 14 post-infection were stained with H-E or PAS and observed under a light microscope. Original magnifications: upper panels, the upper lobe of the left lungs; middle panels, ×40; lower panels, ×200. Representative pictures are shown. Black arrows show granulomas. (c) The proportions of granulomatous areas were calculated. Each column represents the mean ± SD of 3 mice. *, p < 0.05.
To examine the localization of the fungus in the lungs, we compared the numbers of live C. deneoformans colonies in BALFs (alveolar space) and lung homogenate after BAL (interstitial space) between luCldn-18+/+ and luCldn-18−/− mice. The fungal burdens in both BALFs and lung tissue after BAL were greater in luCldn-18−/− mice than in luCldn-18+/+ mice on days 1, 3, 7, and 14 post-infection (Fig. 3a, b). In particular, the increase in the fungal burden in the alveolar space of luCldn-18−/− mice was remarkable on days 3, 7, and 14 post-infection (Fig. 3c).
Effect of luCldn-18 deficiency on the localization of C. deneoformans. luCldn-18+/+ (day 1, n = 4; day 3, n = 4; day 7, n = 5; day 14, n = 5) and luCldn-18−/− mice (day 1, n = 4; day 3, n = 4; day 7, n = 5; day 14, n = 4) were infected intratracheally with C. deneoformans. The numbers of live colonies in BALFs (a, alveolar space) and lung interstitial homogenates after BAL (b, interstitial space) on days 1, 3, 7, and 14 post-infection were counted. (c) The ratio of the number of live colonies in BALFs to the number in the lungs was calculated for each mouse. Each symbol represents a separate mouse, and bars indicate the mean ± SD. *, p < 0.05. There were no mice excluded from this analysis.
Effect of luCldn-18 deficiency on the immune response after infection with C. deneoformans
During infection with various microorganisms, an inflammatory response is induced, increasing total protein concentrations and inflammatory cell counts36. The next series of experiments examined the effect of luCldn-18 deficiency on the immune response to C. deneoformans. Total protein concentrations and cell counts in BALFs were significantly greater in luCldn-18−/− mice than in luCldn-18+/+ mice on days 0, 1, 3, 7, and 14 post-infection (Fig. 4a, b).
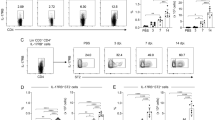
Effects of luCldn-18 deficiency on inflammatory response in BALFs after infection with C. deneoformans. luCldn-18+/+ (day 0, n = 4; day 1, n = 5; day 3, n = 5; day 7, n = 6; day 14, n = 6) and luCldn-18−/− mice (day 0, n = 4 ;day 1, n = 4; day 3, n = 6; day 7, n = 6; day 14, n = 7) were infected intratracheally with C. deneoformans. Protein concentration (a) and cell counts (b) in BALFs were measured on days 0 (uninfected), 1, 3, 7, and 14 post-infection. Each column represents the mean ± SD. (c) BALF cells were prepared on day 3 post-infection. The cell population was analyzed using flow cytometry, and the number in each subset was calculated. Each column represents the mean ± SD. Cytokine production in BALF (d) and lung interstitial homogenates after BAL (e) at the indicated time intervals after C. deneoformans infection. Each column represents the mean ± SD. +/+, luCldn-18+/+ mice. −/−, luCldn-18−/− mice. ND, not detected. *, p < 0.05. There were no mice excluded from this analysis.
The distributions of various cell types in the BALF were analyzed using flow cytometry on day 3 post-infection (when cell counts peaked). The numbers of neutrophils, alveolar macrophages, inflammatory monocytes, NK, CD4+T, CD8+T, and NKT cells were significantly greater in luCldn-18−/− mice than in luCldn-18+/+ mice (Fig. 4c). Host defense against cryptococcal infection is largely regulated by the balance between the Th1 and Th2 immune responses2,3,4,5,6,7. Our previous study showed that IL-17A is involved in the negative regulation of local host defenses against C. deneoformans infection through suppression of the Th1 response11. Here, therefore, we examined the effect of luCldn-18 deficiency on cytokine production in response to C. deneoformans infection by measuring cytokine concentrations in BALFs and lung homogenate after BAL on days 3, 7, and 14 post-infection. Production of IFN-γ in BALF on days 3 and 7 and in lung homogenate after BAL on day 14 were significantly lower in luCldn-18−/− mice than in luCldn-18+/+ mice (Fig. 4d, e). Similarly, production of IL-4 and IL-13 in BALF on day 14 and in the lungs on days 3 and 14 were significantly lower in luCldn-18−/− mice than in luCldn-18+/+ mice (Fig. 4d, e). In addition, production of IL-17A in BALF on day 7 and in the lungs on day 14 were significantly lower in luCldn-18−/− mice than in luCldn-18+/+ mice (Fig. 4d, e).
Effect of luCldn-18 deficiency on homeostasis of the lung microenvironment
luCldn-18 is involved in alveolar fluid homeostasis by regulating solute and ion permeability between alveolar epithelial cells27. The next series of experiments examined the effect of luCldn-18 deficiency on the homeostasis of the lung microenvironment. In a steady state, K+ ion concentration in BALFs was significantly higher in luCldn-18−/− mice than in luCldn-18+/+ mice, while pH was significantly lower (Fig. 5a). Airway surface liquid pH can be altered by disease and lung inflammation, in part because airway inflammation promotes acidification37. Alveolus acidification was observed after cryptococcal infection in both genotypes (Fig. 5b). In addition, alveolar pH remained significantly lower in luCldn-18−/− mice than in luCldn-18+/+ mice until at least seven days after infection (Fig. 5b).
Effects of luCldn-18 deficiency on ion concentration and pH in BALFs. (a) The concentrations of several ions (Na+, Cl−, K+) and pH were measured in BALFs of uninfected luCldn-18+/+ (n = 3) and luCldn-18−/− mice (n = 3). Each column represents the mean ± SD. (b) luCldn-18+/+ (n = 4) and luCldn-18−/− mice (n = 4) were infected intratracheally with C. deneoformans. pH in BALFs was measured on day 3 and 7 post-infection. Each column represents the mean ± SD. *, p < 0.05. NS, not significant. There were no mice excluded from this analysis.
Effect of luCldn-18 deficiency on fungal growth
Cryptococcal replication has been reported to be enhanced at acidic pH as opposed to physiological pH38,39. In this study, similarly, the numbers of yeast cells and the budding rate after in vitro culture were significantly greater at BALF’s pH condition of luCldn-18−/− mice (pH 6.8) than in those of luCldn-18+/+ mice (pH 7.3) (Fig. 6a, b). In addition, in the cell division assay using CFSE, CFSE labeling of yeast cells after in vitro culture, which is correlated with cell propagation, was significantly lower at pH 6.8 than at pH 7.3 (Fig. 6c, d). In our in vivo infection experiment, likewise, CFSE expression on CD45− cells in BALFs after cryptococcal infection was significantly lower in luCldn-18−/− mice than in luCldn-18+/+ mice (Fig. 6e, f), suggesting that cryptococcal replication was increased at the acidic pH that occurs in luCldn-18−/− mice as opposed to the physiological pH that occurs in luCldn-18+/+ mice.
Effect of luCldn-18 deficiency on fungal growth. C. deneoformans was cultured in RPMI 1640 medium containing 20 mM HEPES at 37 °C for 24 h at different pH values (6.8 and 7.3, n = 3 cultures for each condition). (a) The number of organisms was counted using a hemocytometer. Each symbol represents a separate culture, and bars indicate the mean ± SD. (b) The budding ratio was calculated in the PAS-stained specimens. Each column represents the mean ± SD. CFSE-labeled C. deneoformans was cultured in RPMI 1640 medium containing 20 mM HEPES at 37 °C for 24 h at different pH values (6.8 and 7.3) and cell division was analyzed using flow cytometry (c, d). Representative histograms (c) and the MFI of CFSE (d) are shown. CFSE-labeled C. deneoformans cultured in PDA plates was used as a positive control. Each column represents the mean ± SD. luCldn-18+/+ (n = 3) and luCldn-18−/− mice (n = 3) were infected intratracheally with CFSE-labeled C. deneoformans (e, f). BALF cells were prepared at 24 h post-infection. CD45−CFSE+ cells in the alveolar space were defined as fungal cells and analyzed for CFSE fluorescence intensity using flow cytometry. (e) Representative histograms are shown. CFSE-labeled C. deneoformans cultured in PDA plates was used as a positive control. (f) The MFI of CFSE was calculated. Each column represents the mean ± SD. *, p < 0.05. There were no mice excluded from this analysis.
Discussion
TJs function as the primary structural component controlling paracellular permeability and as barriers in the spaces between the alveolar epithelial cells40, yet their contribution to respiratory infection has not been thoroughly analyzed. Cldn-4 and luCldn-18 are among the components of airway TJs22,23. In the present study, luCldn-18 deficiency increased susceptibility to pulmonary C. deneoformans infection, while Cldn-4 deficiency did not affect fungal clearance. TJs assemble through homotypic and heterotypic cis- and trans-interactions between claudins in paracellular localization41,42. In lung epithelial cells, Kage et al. found that Cldn-4 deficiency did not alter the expression of other Cldns (Cldn-3, 5, 7, and 18) or ion permselectivity at steady state24. In alveolar epithelial cells, in contrast, Li et al. demonstrated that luCldn-18 deficiency increased expression of Cldn-3 and Cldn-4 in whole lungs and ion permeability at steady state27. In addition, impaired alveolarization was observed in luCldn-18-deficient mice28. In the current study, likewise, luCldn-18 deficiency led to altered expression of other Cldns (Cldn-3 and 5) in the lungs, with increased K+ ion concentration, decreased pH in BALF, and impaired alveolar formation. These data suggest that luCldn-18 may have an important role in maintaining homeostasis of the microenvironment within alveolar spaces and that its impairment may increase susceptibility to cryptococcal infection.
luCldn18 deficiency increased fungal burden not only in the lungs but also in the brain. The blood–brain barrier (BBB) protects the central nervous system (CNS) by restricting the passage of molecules and microorganisms43. It has been suggested that C. deneoformans may enter the blood or brain from the lung through a Trojan horse mechanism, crossing the BBB as a passenger inside host phagocytes44. The increased fungal dissemination into the brain in luCldn-18-deficient mice may be related to pressure changes due to the increased fungal burden in the lungs. Berndt et al. have reported that Cldn-5, 11, 12, and 25 are expressed in murine brain capillaries45, and some pathogens are known to infiltrate the CNS by regulating the expression of these Cldns46: human immunodeficiency virus type 1 (HIV-1), for example, has been shown to disrupt BBB integrity via modification of Cldn-5, thereby allowing HIV-1 to enter the brain47. Group B streptococcus disrupts the paracellular pathway of the BBB by downregulating claudin-5 via upregulation of Snail1 and enhances bacterial invasion into the CNS48. Yet the effect on fungal passage of the expression of these Cldns within the brain has not been analyzed. Since the present study did not analyze whether the expression of Cldns in the brain was altered in luCldn-18-deficient mice, further investigations are required to address the possibility that the passage of fungus, including phagocytosed fungus, might be enhanced in luCldn-18-deficient mice due to altered Cldn expression patterns in their brains.
In luCldn-18-deficient mice, the increased population of C. deneoformans was more remarkable in the alveolar space than in the lung interstitium. In addition, acidification in the alveolar spaces of luCldn-18-deficient mice was observed during infection with this fungal pathogen. In previous studies, stCldn-18-deficient mice have been found to have lower pH in the stomach due to gastric acid leakage, leading to gastritis and gastric tumorigenesis18,49,50. Alveolar macrophages, which constitute a sizable fraction of alveolar subphase fluids, are a substantial source of metabolic H+ production under physiological conditions37. In the current study, the number of alveolar macrophages was significantly greater in luCldn-18−/− mice than in luCldn-18+/+ mice after cryptococcal infection, suggesting that acidification in the alveolar spaces of luCldn-18-deficient mice may be caused by increased production of metabolic H+ by alveolar macrophages. In addition, the enhanced cell destruction of increased numbers of macrophages and airway epithelial cells may have induced high K+ ion concentrations in alveolar subphase fluids, although our limited analysis of this matter prevents us from drawing firm conclusions.
A Th1-mediated immune response contributes toward the eradication of C. deneoformans through inducing macrophage activation and accelerating the formation of granuloma at the infection sites6,7,8,9, whereas the Th2-mediated immune response counteracts these host protective responses, worsening infection6,7,10. In the present study, the production of Th1-related cytokines such as IL-12p40 and IFN-γ was significantly decreased in luCldn-18−/− mice compared to luCldn-18+/+ mice after cryptococcal infection. Unexpectedly, however, Th2 cytokines such as IL-4 and IL-13, which suppress host defense against cryptococcal infection by inhibiting macrophage activation and enhancing mucin production6,7,10,30, were not increased but rather also decreased in luCldn-18−/− mice compared to luCldn-18+/+ mice. In addition, the synthesis of IL-17A, which is involved in the negative regulation of local host defense against C. deneoformans infection11, was significantly lower in luCldn-18−/− mice than in luCldn-18+/+ mice. Thus, the changes in production of host defense-related cytokines were not consistent with the impaired fungal elimination observed in luCldn-18−/− mice. Similar inconsistent results were observed in the infiltration of inflammatory cells including CD4+ T cells into the alveolar spaces, which was drastically increased in luCldn-18−/− mice compared to luCldn-18+/+ mice. There are no reports of a single luCldn-18 deficiency leading to increasing paracellular cell transport. In the current study, luCldn-18 deficiency induced Cldn expression changes, but the expression of many TJ proteins, including occludin, was maintained. Therefore, the possibility of increased mechanical paracellular transport through alveolar epithelial cells is thought to be low. Previously, Leblebicioglu and co-workers reported that chemotactic responses of human peripheral blood polynuclear leukocytes were significantly increased under decreased pH conditions in in vitro cultures51. Lower pH induces activation of monocytes/macrophages and dendritic cells but suppresses T cell immune responses36,52,53. Higher levels of extracellular K+ ions are reported to suppress the activation of T cell-mediated immune responses54,55, which is also observed under the luCldn-18-deficient condition in the current study. These findings suggest that acidification in the alveolar spaces of luCldn-18−/− mice may promote the infiltration of inflammatory cells into the alveolar space while also suppressing the T cell immune response to C. deneoformans through acidification and increased levels of K+ ions.
In the current study, fungal replication of C. deneoformans was significantly enhanced both in acidic culture conditions in vitro and in the alveolar spaces of luCldn-18-deficient mice in vivo compared with physiological pH and control mice, respectively. Several investigators have demonstrated that acidic conditions enhance cryptococcal growth while alkaline conditions inhibit it37,38,56,57. Islam et al. have reported that the replication of Cryptococcus including the B3501 strain was increased at acidic pH compared to physiological pH conditions56. Levitz et al. have reported that C. neoformans is able to grow at pH values ranging from 5 to 8, with optimal growth at pH 557. Meanwhile, Vidotto et al. have demonstrated that K+ ion concentration does not affect C. neoformans phenoloxidase activity, which has long been known as a virulence factor58. Collectively, these results suggest that acidification in the alveolar spaces of luCldn-18−/− mice may not only suppress T cell activation but also enhance the growth of the fungal pathogen C. deneoformans.
Diseases in which the expression of luCldn-18 changes are not well known, such as diabetes mellitus, hematological diseases and collagen diseases, are known to be risk factors for cryptococcosis in non-HIV patients59. In addition, luCldn-18 may be impaired by inflammatory responses and tissue damage induced after cryptococcal infection, but the present study did not analyze whether the expression of luCldn-18 in the lungs was decreased after infection. Analysis of luCldn-18 gene mutations and expression in high-risk patients and luCldn-18 injury mechanisms involved in cryptococcal infection may be useful in finding therapies for infection with this pathogen.
In conclusion, the present study demonstrates that luCldn-18 deficiency probably suppresses T cell activation during cryptococcal infection through acidification and increased extracellular K+ ion concentration in the alveolar spaces. As C. deneoformans has greater replication efficiency in lower pH conditions, the acidification of the alveolar spaces may increase the lung burden of this fungal pathogen in luCldn-18−/− mice. Thus, luCldn-18 may be indirectly involved in limiting the progression of cryptococcal infection in the lungs through maintaining the homeostatic condition in the alveolar space microenvironment, as shown in Fig. 7. The current findings offer a novel approach to and a possibly better understanding of the pathogenic mechanism of cryptococcal infection at the mucosal surfaces of the bronchoalveolar spaces.
Summary of the role of luCldn-18 in host defense against infection with C. deneoformans. luCldn-18 deficiency induces acidification and high K+ ion concentration in the alveolar space. Alveolar acidification and high K+ ion concentrations inhibit the activation of T cells in response to cryptococcal infection and thereby suppress granuloma formation. In addition, the increased replication efficiency of this fungus at low pH values increases the fungal burden in luCldn-18-deficient mice. Consequently, luCldn-18 indirectly helps prevent cryptococcal infection by maintaining homeostasis of the alveolar space microenvironment. AVSE, alveolar subphase fluid. TJ, tight junction. AMφ, alveolar macrophages. AT1, alveolar type I cells. AT2, alveolar type II cells. This image was drawn by Ko Sato.
Abbreviations
- Cldn:
-
Claudin
- luCldn-18:
-
Lung-specific Cldn-18
- stCldn-18:
-
Stomach-specific Cldn-18
- TJ:
-
Tight junction
- PDA:
-
Potato dextrose agar
- PBS:
-
Phosphate buffered saline
- BALF:
-
Bronchoalveolar lavage fluid
- H-E:
-
Hematoxylin–eosin
- PAS:
-
Periodic acid-Schiff
- BCA:
-
Bicinchoninic acid
- FCS:
-
Fetal calf serum
- CFSE:
-
Carboxyfluorescein succinimidyl ester
- BBB:
-
Blood–brain barrier
- CNS:
-
Central nervous system
- HIV-1:
-
Human immunodeficiency virus type 1
- MFI:
-
Mean fluorescent intensity
- AVSF:
-
Alveolar subphase fluid
- SD:
-
Standard deviation
- ND:
-
Not detected
- NS:
-
Not significant
- AMφ:
-
Alveolar macrophages
- AT1:
-
Alveolar type I cells
- AT2:
-
Alveolar type II cells
References
Hagen, F. et al. Recognition of seven species in the Cryptococcus gattii/Cryptococcus neoformans species complex. Fungal Genet. Biol. 78, 16–48 (2015).
Perfect, J. R. & Casadevall, A. Cryptococcosis. Infect. Dis. Clin. North Am. 16, 837–874, v–vi (2002).
Feldmesser, M., Tucker, S. & Casadevall, A. Intracellular parasitism of macrophages by Cryptococcus neoformans. Trends Microbiol. 9, 273–278 (2001).
Gilbert, A. S., Wheeler, R. T. & May, R. C. Fungal pathogens: Survival and replication within macrophages. Cold Spring Harb. Perspect. Med. 5, a019661 (2015).
Lim, T. S. & Murphy, J. W. Transfer of immunity to cryptococcosis by T-enriched splenic lymphocytes from Cryptococcus neoformans-sensitized mice. Infect. Immun. 30, 5–11 (1980).
Koguchi, Y. & Kawakami, K. Cryptococcal infection and Th1-Th2 cytokine balance. Int. Rev. Immunol. 21, 423–438 (2002).
Arora, S. et al. Effect of cytokine interplay on macrophage polarization during chronic pulmonary infection with Cryptococcus neoformans. Infect. Immun. 79, 1915–1926 (2011).
Tohyama, M., Kawakami, K., Futenma, M. & Saito, A. Enhancing effect of oxygen radical scavengers on murine macrophage anticryptococcal activity through production of nitric oxide. Clin. Exp. Immunol. 103, 436–441 (1996).
Hardison, S. E. et al. Pulmonary infection with an interferon-gamma-producing Cryptococcus neoformans strain results in classical macrophage activation and protection. Am. J. Pathol. 176, 774–785 (2010).
Müller, U. et al. IL-13 induces disease-promoting type 2 cytokines, alternatively activated macrophages and allergic inflammation during pulmonary infection of mice with Cryptococcus neoformans. J. Immunol. 179, 5367–5377 (2007).
Sato, K. et al. Production of IL-17A at innate immune phase leads to decreased Th1 immune response and attenuated host defense against infection with Cryptococcus deneoformans. J. Immunol. 205, 686–698 (2020).
Zhang, Y. et al. Robust Th1 and Th17 immunity supports pulmonary clearance but cannot prevent systemic dissemination of highly virulent Cryptococcus neoformans H99. Am. J. Pathol. 175, 2489–2500 (2009).
Murdock, B. J., Huffnagle, G. B., Olszewski, M. A. & Osterholzer, J. J. Interleukin-17A enhances host defense against cryptococcal lung infection through effects mediated by leukocyte recruitment, activation, and gamma interferon production. Infect. Immun. 82, 937–948 (2014).
Szymczak, W. A., Sellers, R. S. & Pirofski, L. IL-23 dampens the allergic response to Cryptococcus neoformans through IL-17-independent and -dependent mechanisms. Am. J. Pathol. 180, 1547–1559 (2012).
Hardison, S. E., Wozniak, K. L., Kolls, J. K. & Wormley, F. L. Interleukin-17 is not required for classical macrophage activation in a pulmonary mouse model of Cryptococcus neoformans infection. Infect. Immun. 78, 5341–5351 (2010).
Wozniak, K. L., Hole, C. R., Yano, J., Fidel, P. L. & Wormley, F. L. Characterization of IL-22 and antimicrobial peptide production in mice protected against pulmonary Cryptococcus neoformans infection. Microbiology 160, 1440–1452 (2014).
Tsukita, S., Furuse, M. & Itoh, M. Multifunctional strands in tight junctions. Nat. Rev. Mol. Cell Biol. 2, 285–293 (2001).
Tsukita, S., Tanaka, H. & Tamura, A. The claudins: From tight junctions to biological systems. Trends Biochem. Sci. 44, 141–152 (2019).
Rüffer, C. & Gerke, V. The C-terminal cytoplasmic tail of claudins 1 and 5 but not its PDZ-binding motif is required for apical localization at epithelial and endothelial tight junctions. Eur. J. Cell Biol. 83, 135–144 (2004).
Furuse, M., Fujita, K., Hiiragi, T., Fujimoto, K. & Tsukita, S. Claudin-1 and -2: Novel integral membrane proteins localizing at tight junctions with no sequence similarity to ccluding. J. Cell Biol. 141, 1539–1550 (1998).
Tamura, A. & Tsukita, S. Paracellular barrier and channel functions of TJ claudins in organizing biological systems: Advances in the field of barriology revealed in knockout mice. Semin. Cell Dev. Biol. 36, 177–185 (2014).
Ohta, H., Chiba, S., Ebina, M., Furuse, M. & Nukiwa, T. Altered expression of tight junction molecules in alveolar septa in lung injury and fibrosis. Am. J. Physiol. Lung Cell Mol. Physiol. 302, L193–L205 (2012).
LaFemina, M. J. et al. Keratinocyte growth factor enhances barrier function without altering claudin expression in primary alveolar epithelial cells. Am. J. Physiol. Lung Cell Mol. Physiol. 299, L724–L734 (2010).
Kage, H. et al. Claudin 4 knockout mice: Normal physiological phenotype with increased susceptibility to lung injury. Am. J. Physiol. Lung Cell Mol. Physiol. 307, L524–L536 (2014).
Niimi, T. et al. Claudin-18, a novel downstream target gene for the T/EBP/NKX2.1 homeodomain transcription factor, encodes lung- and stomach-specific isoforms through alternative splicing. Mol. Cell Biol. 21, 7380–7390 (2001).
Türeci, O. et al. Claudin-18 gene structure, regulation, and expression is evolutionary conserved in mammals. Gene 481, 83–92 (2011).
Li, G. et al. Knockout mice reveal key roles for claudin 18 in alveolar barrier properties and fluid homeostasis. Am. J. Respir. Cell Mol. Biol. 51, 210–222 (2014).
LaFemina, M. J. et al. Claudin-18 deficiency results in alveolar barrier dysfunction and impaired alveologenesis in mice. Am. J. Respir. Cell Mol. Biol. 51, 550–558 (2014).
Matsumura, H., Hasuwa, H., Inoue, N., Ikawa, M. & Okabe, M. Lineage-specific cell disruption in living mice by Cre-mediated expression of diphtheria toxin A chain. Biochem. Biophys. Res. Commun. 321, 275–279 (2004).
Sato, K. et al. Cryptococcus neoformans infection in mice lacking type I interferon signaling leads to increased fungal clearance and IL-4-dependent mucin production in the lungs. PLoS ONE 10, e0138291 (2015).
Sato, Y. et al. Limited role of mincle in the host defense against infection with Cryptococcus deneoformans. Infect Immun. 88, e00400-e420 (2020).
Nakamura, Y. et al. Dectin-2 deficiency promotes Th2 response and mucin production in the lungs after pulmonary infection with Cryptococcus neoformans. Infect Immun. 83, 671–681 (2015).
Yamamoto, H. et al. Defect of CARD9 leads to impaired accumulation of gamma interferon-producing memory phenotype T cells in lungs and increased susceptibility to pulmonary infection with Cryptococcus neoformans. Infect Immun. 82, 1606–1615 (2014).
Ishizuka, S. et al. Effect of CARD9 deficiency on neutrophil-mediated host defense against pulmonary infection with Streptococcus pneumoniae. Infect Immun. 89, e00305-e320 (2020).
Toyama, M. et al. Attenuated accumulation of regulatory T cells and reduced production of interleukin 10 lead to the exacerbation of tissue injury in a mouse model of acute respiratory distress syndrome. Microbiol Immunol. 62, 111–123 (2018).
Chen, L. et al. Inflammatory responses and inflammation-associated diseases in organs. Oncotarget 9, 7204–7218 (2017).
Ng, A. W., Bidani, A. & Heming, T. A. Innate host defense of the lung: Effects of lung-lining fluid pH. Lung 182, 297–317 (2004).
Howard, D. H. Some factors which affect the initiation of growth of Cryptococcus neoformans. J. Bacteriol. 82, 430–435 (1961).
DeLeon-Rodriguez, C. M. & Casadevall, A. Cryptococcus neoformans: Tripping on acid in the phagolysosome. Front. Microbiol. 7, 164 (2016).
Overgaard, C. E., Mitchell, L. A. & Koval, M. Roles for claudins in alveolar epithelial barrier function. Ann. N. Y. Acad. Sci. 1257, 167–174 (2012).
Suzuki, H., Tani, K., Tamura, A., Tsukita, S. & Fujiyoshi, Y. Model for the architecture of claudin-based paracellular ion channels through tight junctions. J. Mol. Biol. 427, 291–297 (2015).
Suzuki, H. et al. Crystal structure of a claudin provides insight into the architecture of tight junctions. Science 344, 304–307 (2014).
Serlin, Y., Shelef, I., Knyazer, B. & Friedman, A. Anatomy and physiology of the blood–brain barrier. Semin. Cell Dev. Biol. 38, 2–6 (2015).
Santiago-Tirado, F. H., Onken, M. D., Cooper, J. A., Klein, R. S. & Doering, T. L. Trojan horse transit contributes to blood-brain barrier crossing of a eukaryotic pathogen. MBio 8, 02183–16 (2017).
Berndt, P. et al. Tight junction proteins at the blood–brain barrier: Far more than claudin-5. Cell Mol. Life Sci. 76, 1987–2002 (2019).
Greene, C., Hanley, N. & Campbell, M. Claudin-5: Gatekeeper of neurological function. Fluids Barriers CNS. 16, 3 (2019).
András, I. E. & Toborek, M. HIV-1-induced alterations of claudin-5 expression at the blood–brain barrier level. Methods Mol. Biol. 762, 355–3570 (2011).
Kim, B. J. et al. Bacterial induction of Snail1 contributes to blood-brain barrier disruption. J. Clin. Invest. 125, 2473–2483 (2015).
Hayashi, D. et al. Deficiency of claudin-18 causes paracellular H+ leakage, up-regulation of interleukin-1β, and atrophic gastritis in mice. Gastroenterology 142, 292–304 (2012).
Suzuki, K. et al. Deficiency of stomach-type claudin-18 in mice induces gastric tumor formation independent of H pylori infection. Cell. Mol. Gastroenterol. Hepatol. 8, 119–142 (2019).
Leblebicioglu, B., Lim, J. S., Cario, A. C., Beck, F. M. & Walters, J. D. pH changes observed in the inflamed gingival crevice modulate human polymorphonuclear leukocyte activation in vitro. J. Periodontol. 67, 472–477 (1996).
Erra Díaz, F., Dantas, E. & Geffner, J. Unravelling the interplay between extracellular acidosis and immune cells. Mediators Inflamm. 2018, 1218297 (2018).
Bosticardo, M. et al. Biased activation of human T lymphocytes due to low extracellular pH is antagonized by B7/CD28 costimulation. Eur. J. Immunol. 31, 2829–2838 (2001).
Eil, R. et al. Ionic immune suppression within the tumour microenvironment limits T cell effector function. Nature 537, 539–543 (2016).
Ong, S. T. et al. Extracellular K+ dampens T cell functions: Implications for immune suppression in the tumor microenvironment. Bioelectricity. 1, 169–179 (2019).
Islam, A. et al. An acidic microenvironment increases NK cell killing of Cryptococcus neoformans and Cryptococcus gattii by enhancing perforin degranulation. PLoS Pathog. 9, e1003439 (2013).
Levitz, S. M., Harrison, T. S., Tabuni, A. & Liu, X. Chloroquine induces human mononuclear phagocytes to inhibit and kill Cryptococcus neoformans by a mechanism independent of iron deprivation. J. Clin. Invest. 100, 1640–1646 (1997).
Vidotto, V. et al. Effect of different K+ concentrations on Cryptococcus neoformans phenoloxidase activity. Mycopathologia 156, 171–176 (2003).
Kohno, S. et al. Clinical features of pulmonary cryptococcosis in non-HIV patients in Japan. J Infect Chemother. 21, 23–30 (2015).
Acknowledgements
We thank the Biomedical Research Unit of Tohoku University Hospital and the Biomedical Research Core, Animal Pathology Platform of Tohoku University Graduate School of Medicine for providing technical support. This work was supported in part by a Grant-in-Aid for Scientific Research (B) (18H02851) and Early-Career Scientists (19K17920 and 21K16314) from the Ministry of Education, Culture, Sports, Science and Technology of Japan; by the Research Program on Emerging and Reemerging Infectious Diseases from the Japan Agency for Medical Research and Development, AMED (JP19fk0108094 and JP20fk0108094); by the Strategic International Collaborative Research Program (SICORP), AMED (JP19jm0210073 and JP20jm0210073); by the MSD Life Science Foundation, Public Interest Incorporated Foundation (ID-014); and by the Joint Usage/Research Program of the Medical Mycology Research Center, Chiba University (20-02, 21-04).
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: Ko S, IM, JK, HY, and Kazuyoshi K. Performed the experiments: Ko S, IM, Koya S, AM, TZ, TK, AO, Kotone K, YK, and AU. Analyzed the data: Ko S, IM, Koya S, MT, DT, EK, HT, KI, and Kazuyoshi K. Contributed reagents/materials/analysis tools: AT, AS and HK. Contributed to the writing of the manuscript: Ko S, IM, AT, ST, and Kazuyoshi K. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sato, K., Matsumoto, I., Suzuki, K. et al. Deficiency of lung-specific claudin-18 leads to aggravated infection with Cryptococcus deneoformans through dysregulation of the microenvironment in lungs. Sci Rep 11, 21110 (2021). https://doi.org/10.1038/s41598-021-00708-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-00708-6
This article is cited by
-
Tetrahydrocurcumin (THC) enhanced the clearance of Cryptococcus deneoformans during infection in vivo
Antonie van Leeuwenhoek (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.