Abstract
The cellular prion protein (PrPC) is a mainly α-helical 208-residue protein located in the pre- and postsynaptic membranes. For unknown reasons, PrPC can undergo a structural transition into a toxic, β-sheet rich scrapie isoform (PrPSc) that is responsible for transmissible spongiform encephalopathies (TSEs). Metal ions seem to play an important role in the structural conversion. PrPC binds Zn(II) ions and may be involved in metal ion transport and zinc homeostasis. Here, we use multiple biophysical techniques including optical and NMR spectroscopy, molecular dynamics simulations, and small angle X-ray scattering to characterize interactions between human PrPC and Zn(II) ions. Binding of a single Zn(II) ion to the PrPC N-terminal domain via four His residues from the octarepeat region induces a structural transition in the C-terminal α-helices 2 and 3, promotes interaction between the N-terminal and C-terminal domains, reduces the folded protein size, and modifies the internal structural dynamics. As our results suggest that PrPC can bind Zn(II) under physiological conditions, these effects could be important for the physiological function of PrPC.
Similar content being viewed by others
Introduction
Misfolding and aggregation of the 208-residue prion protein (PrP) is the molecular event underlying the progressive and fatal neurodegenerative diseases collectively known as transmissible spongiform encephalopathies (TSEs)1, or prion diseases. PrP is an outer membrane glycoprotein encoded by the PRNP gene, highly conserved within mammals2, and expressed at high levels in the brain—especially in the striatum, hippocampus and cortex3. Single-nucleotide mutations or sequence expansions within the PRNP gene are the cause of genetic prion diseases such as familial Creutzfeldt-Jakob disease (fCJD), fatal familial insomnia (FFI), and Gerstmann-Sträussler-Scheinker (GSS) syndrome. Post-translational modifications of PrP produce the mature cellular form known as PrPC4,5,6, which can undergo a structural rearrangement into the aggregated, β-sheet-rich, and pathological (scrapie) isoform denoted as PrPSc. This form acts as a template for PrPC to refold into toxic conformations1,7. The infectious prion diseases arise from contact with pathogenic PrPSc via events such as organ transplantation from people with CJD (iatrogenic CJD), consumption of beef contaminated with bovine spongiform encephalopathy (variant CJD)1,7, or human cannibalistic rituals (Kuru)8. The most common prion disease in humans is however sporadic CJD, but its origins are unknown. Despite intense research, no drugs have so far been devised that can cure prion diseases9,10,11.
The human PrP protein is expressed as a 253-residue long precursor polypeptide chain. The post-translational modifications include: removal of the N-terminal 22-residue signal sequence, removal of 23 C-terminal residues, formation of one disulfide bridge (Cys179-Cys214), glycosylation of two asparagine residues (Asn181, Asn197), as well as binding of a glycosylphosphatidylinositol (GPI) anchor4,5,6 (Fig. 1). The N-terminal domain of PrPC is intrinsically disordered12, although it contains four octapeptide repeats with β-turn or polyproline II secondary structure13,14,15. The C-terminal domain, whose 3D-fold is well conserved within mammals, consists of three α-helices and two antiparallel β-sheets12,16,17,18. The first α-helix is formed by residues 144–154 and is flanked by two short β-strands, i.e. residues 128–131 and 161–164. Helices α2 and α3 consist of residues 173–194 and 200–228, and are interconnected by a disulfide bond between Cys179 and Cys21412,18,19. In vitro studies suggest that conversion of the mainly α-helical PrPC into the toxic and β-sheet-rich PrPSc isoform requires misfolding or unfolding of PrPC as an intermediate step1,7.
Schematic representation of the mature human PrPC protein. PrPC consists of an N-terminal intrinsically disordered domain and a C-terminal structured domain. The octarepeat (OR) region is marked in orange, α-helices are marked in blue, β-sheets are marked in green, hexagons represent the glycosylation sites, S–S is a disulfide bridge and GPI represents the glycosylphosphatidylinositol anchor.
The biological function of PrPC is not fully understood. The protein is involved in controlling synaptic transmission and neuronal plasticity. Studies on mouse models suggest that PrPC may be crucial for the regulation of the circadian rhythm, and the development of the central nervous system20,21. Moreover, it has been proposed that PrPC might have a significant role in the homeostasis of different metal ions21,22, as the brain distribution of metal ions correlates with the PrPC expression level23. It is still unclear if zinc dyshomeostasis, or metal imbalances in general, are part of the pathology in prion diseases, as appears to be the case in other neurodegenerative protein aggregation diseases such as Alzheimer's 24.
Human PrPC has been tested for binding of various divalent metal ions including Cu(II), Ni(II), Zn(II), and Mn(II)25,25,26,27,28,29,30. Zinc is the second-most abundant (after iron) metal in living organisms. It is a co-factor of many enzymes31, and plays an important role in cell signaling and proliferation32. Zn(II) ions are important neurotransmitters in the synaptic cleft where concentrations can reach up to 300 µM33,34. In PrPC both Cu(II) and Zn(II) mainly binds to the octarepeat region (Fig. 1, orange)25,29,35. Upon binding of Zn(II) ions, the N-terminal domain forms a tertiary contact with the C-terminal domain via the octarepeats36. This interaction is disrupted in PrPC mutants associated with TSE, suggesting that disruption of Zn(II)-mediated intramolecular interactions might cause TSE36. Even though the interaction between PrPC and Zn(II) has been investigated for the last twenty years, the dissociation constant values reported in literature for the formed complex vary in almost three orders of magnitude, i.e. from ~ 0.5 μM to ~ 200 μM, even when measured in similar environments25,37,38.
Here, we used multiple biophysical techniques, including spectroscopic, scattering, and theoretical methods, to study Zn(II) binding to the full-length human PrPC protein.
Results
CD spectroscopy reveals α-helix to β-sheet transition in PrPC upon Zn(II) binding
Circular dichroism (CD) spectroscopy was used to monitor changes in protein secondary structure induced by Zn(II) ions. The CD spectrum of the pure protein in N-ethyl morpholine (NEM) buffer corresponds to a typical α-helix, with characteristic minima at 208 and 222 nm and a maximum at 193 nm (Fig. 2A). Addition of twenty molar equivalents of Zn(II) to PrPC resulted in a general decrease in CD intensity over the whole wavelength range (Fig. 2A). The observed change corresponds to a decrease in the content of regular α-helices (helix 1, Table S1), and an increase in the content of distorted α-helices (helix 2, Table S1). Addition of Zn(II) also increased the content of antiparallel β-sheets and β-turns, and reduced the content of parallel β-sheets (Table S1). The changes in PrPC secondary structure upon addition of Zn(II) were clearly visible in the far UV region; we therefore proceeded with careful Zn(II) titrations in this region.
CD spectra of huPrPC(23–231) titrated with ZnCl2. (A) 20 μM PrPC (blue line) and 20 μM PrPC with 400 μM ZnCl2 (red line) in 10 mM NEM buffer, pH 7.4. (B) Titration of 1 μM PrPC with ZnCl2 in 10 mM NEM buffer, pH 7.4. The initial spectrum is in blue and the final one in red. (C) Titration of 0.5 μM PrPC with ZnCl2 in 10 mM sodium phosphate buffer, pH 7.4. The initial spectrum is in blue and the final spectrum is in red. (D) CD signal intensity at 217 nm derived from the spectra in Fig. 2B, plotted as a function of added ZnCl2 and fitted to Eq. (1) (Kd = 28.8 ± 1.5 μM). (E) CD signal intensity at 217 nm derived from the spectra in (C), plotted as a function of added ZnCl2 and fitted to Eq. (1) (Kd = 12.1 ± 0.7 μM).
ZnCl2 was titrated to PrPC, both in 10 mM NEM, pH 7.4 (Fig. 2B), and in 10 mM sodium phosphate buffer, pH 7.4 (Fig. 2C). The measurements in NEM buffer displayed an isodichroic point at ~ 242 nm (Fig. 2B). In phosphate buffer no isodichroic point was visible (Fig. 2C). The presence of this isodichroic point shows that the Zn(II)-induced general loss of CD signal intensity is caused by a two-phase structural transition, and not by a lowered protein concentration related to e.g. possible Zn-related aggregation and precipitation of the protein. Figure 2D,E show plots of the CD intensity at 217 nm vs Zn(II) concentration, derived from the spectra in Fig. 2B,C. Fitting binding Eq. (1) to the Zn(II) titration data in NEM buffer (Fig. 2B) produced a dissociation constant (Kd) of 28.8 ± 1.5 μM (Fig. 2D), while the titration data in sodium phosphate buffer (Fig. 2C) suggested stronger binding of Zn(II) to PrPC. Fitting to Eq. (1) yielded the dissociation constant Kd = 12.1 ± 0.7 μM (Fig. 2E). Clearly, the binding behavior of the Zn(II) ions is to some extent influenced by the environmental conditions, such as buffer composition. Because possible binding of Zn(II) ions to the buffer is not taken into account, the calculated Kd values should be considered to be apparent.
Fluorescence spectroscopy shows that tryptophan residues are in close proximity to the PrPC-Zn(II) binding site
The PrPC protein has seven tryptophan residues located in the N-terminal unstructured domain. Four of these residues, i.e. Trp65, Trp73, Trp81 and Trp89, are located in the octarepeat region where they appear to be indirectly involved in copper binding35. Addition of Cu(II) ions quench the tryptophan fluorescence signal25. Here, a similar effect is observed upon addition of Zn(II) ions to 0.5 μM PrPC protein in 10 mM sodium phosphate buffer, pH 7.4 (Fig. 3A). The fluorescence spectrum of apo-PrPC showed a single maximum at ~ 347 nm, indicating full exposure of the tryptophan residues to the solvent39.
(A) Fluorescence spectra (excitation wavelength = 280 nm) of 0.5 μM huPrPC protein titrated with 0 to 70 µM ZnCl2, see (B), in 10 mM sodium phosphate buffer, pH 7.4 at 25 °C. The initial spectrum is in blue and the final in red. (B) Changes in the relative fluorescence intensity at 347 nm, fitted to Eq. (2) (Kd = 16.8 ± 0.9 μM).
The titration with ZnCl2 solution resulted in a reduced fluorescence intensity, a slight blue shift of the maximum to 343 nm, and revealed the presence of two additional peaks at 333 nm and 313 nm (Fig. 3A). The blue shift of tryptophan fluorescence to ~ 342 nm corresponds to a change in tryptophan environment and exposure to the bound water molecules, while the two new peaks at 333 nm and 313 nm may be attributed to buried and structured tryptophan residues, respectively39. In the last fluorescence spectrum of the titration, corresponding to a Zn(II) concentration of 70 µM, the maximum at 343 nm is still visible. Plotting the fluorescence intensity at 347 nm versus ZnCl2 concentration and fitting the data to Eq. (2) suggested binding of Zn(II) ions to the PrPC protein (Fig. 3B) with an apparent dissociation constant Kd = 16.8 ± 0.9 μM.
NMR spectroscopy shows Zn(II) binding promotes interactions between the PrPC N- and C-terminal domains
To investigate the structural alteration of PrPC under Zn(II) saturation, high-resolution 2D heteronuclear 1H-15 N HSQC NMR solution spectra of the protein were acquired in 50 mM deuterated HEPES buffer, 50 mM NaCl, pH 7.0, before and after addition of two molar equivalents of ZnCl2 (Fig. 4A). Comparison of the spectra collected without and with Zn(II) ions revealed no differences in peak positions. Reduced cross-peak intensities for certain amino acids, however, indicate residue-specific binding interactions with Zn(II) ions (Fig. 4A). As Zn(II) ions are not paramagnetic, this loss of signal intensity is likely caused by chemical exchange on an intermediate NMR time scale. Substantial intensity changes were observed for cross-peaks corresponding to amino acids in the folded domain of the PrPC 3D structure—in particular in the C-terminal region with α-helices 2 and 3. In α-helix 2, decreased cross-peak amplitudes were detected for residues in the last two turns of the helix, e.g. Thr183, His187, and Thr188 (Fig. 4A).
(A) 2D NMR 1H-15 N-HSQC spectrum of 300 µM huPrPC protein in 50 mM HEPES, 50 mM NaCl, pH 7.0, under saturation with Zn(II) ions. The 1H traces shown as insets reveal decreased peak amplitudes under Zn(II) saturation (red) compared to the spectra for the apo form (blue). (B) Relative amplitudes of the resonance peaks versus gradient strength, used to calculate translational diffusion coefficients for the apo (blue) and Zn(II)-saturated (red) forms of the PrPC protein. (C) Relative ratios between 15N R2 relaxation rates obtained for the apo (R2(apo)) and Zn-bound (R2(Zn)) states of PrPC residues. Data are presented only for residues in the C-terminal domain. (D) A ribbon presentation of the C-terminal fragment of the PrPC protein. Residues exhibiting decreased R2 relaxation rates under saturation with Zn(II) ions are shown in dark.
A more compact structure of the PrPC protein in presence of Zn(II) ions was demonstrated by measurements of the translational diffusion coefficient (Dtr) by PGSE-NMR experiments. Addition of ZnCl2 resulted in higher translational mobility of the protein compared to the apo form, with Dtr values increasing from 1.09 ± 0.02 10–10 m2/s up to 1.23 ± 0.03 10–10 m2/s (Fig. 4B).
To further explore molecular dynamic processes in the PrPC sample, the 15N spin–spin relaxation rates (R2) were measured. The data acquired for the Zn-bound state were compared with previously collected R2 values obtained for the apo form of PrPC at the same conditions. For the 71 amide nitrogens assigned in the C-terminal domain, this comparison revealed decreased 15N R2 relaxation rates for the majority of the 15N backbone resonances under Zn(II) saturation (Fig. 4C). Strong effects of Zn(II)-binding were seen in a number of C-terminal residues. For example, significantly decreased R2 values for Gly126 and Tyr128 suggest changes towards more mobility for the short β–sheet fragment 128YMLG131. In addition, the significantly decreased R2 values for the Cys214, Thr216, Ser222, and Gln223 residues in the third α-helix suggest increased local dynamics also in this region upon Zn(II) binding (Fig. 4C).
The changes in molecular dynamic processes, observed under saturation by Zn(II), are highlighted in the 3D structure of the C-terminal domain of the protein (Fig. 4D). Together with an overall lower structural stability deduced from the 15N R2 data for Zn(II)-bound PrPC, we conclude that there are pronounced changes in the dynamics of the C-terminal region consisting of α–helices 2 and 3 joined with a β-strand motif (Fig. 4D). Our data suggest that even though local dynamics increase in the C-terminal domain by Zn(II) binding, the resulting effect is to promote interactions between the N-terminal and C-terminal domains, resulting in an overall more compact protein structure. Such an effect is in agreement with previous studies of Zn(II) binding to PrPC36,38.
Molecular dynamics simulations show the Zn(II) binding coordination, and consequent structural and dynamic changes in the PrPC C-terminal domain
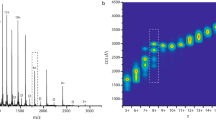
As reported previously, the Zn(II) ion can be coordinated in PrPC(23–231) by four imidazole ring nitrogens from histidine His61, His69, His77, and His85 in the octarepeat fragment36. Our NMR data showed that addition of Zn(II) to PrPC at pH 7.0 shifted only 13Cε1-1Hε1 resonances (Fig. S1). Despite lacking residue-specific assignments for the 13C and 1H histidine side-chain resonances, there are clear chemical shifts in four out of nine histidine 13Cε1 signals (Fig. 5A). We speculate that these shifts are caused by Zn(II)-binding to the four histidine residues in the octarepeat region. Thus, molecular dynamic simulations were used to create the 3D structure of the Zn(II)-binding motif shown in Fig. 5. Due to the spherical geometry of Zn(II) coordination, it is difficult to predict the exact geometry of the Zn(II)-binding motif. Nevertheless, in the second coordination sphere we note the existence of two oxygens from the carbonyl groups of Gly71 and Trp89, which may compensate for the lack of negative charges in the histidine imidazole rings40.
(A) Region of NMR cross-peaks for aromatic hydrogens, from 2D 1H-13C HSQC spectra acquired for 13C,15N-labeled huPrPC(23–231) protein in apo (blue) and Zn-saturated (red) form; (B) 3D model of the huPrPC Zn(II)-binding motif, obtained from MD simulations. The histidine nitrogen atoms coordinating the Zn(II) ion are shown in blue. The two additional carbonyl oxygens (Gly71 and Trp89) present in the second coordination shell are shown in red.
Molecular dynamics simulations were performed also to investigate changes in the C-terminal α-helices upon Zn(II) binding. In the first step, a 195 ns trajectory for apo-PrPC with an extended N-terminal domain was performed. After ~ 25 ns of simulation the RMSD values converged (Fig. S2), and the N-terminal domain formed a compact structure with three antiparallel β-sheets (Pro51-Gly53, Gln67-His69, Gly72-Gly74) located around the C-terminal region of α-helix 3, which became partially unfolded (Fig. 6A, model marked in blue).
(A) Overlaid models of apo-PrPC (blue) and Zn(II)-bound PrPC (red), obtained at the end of the molecular dynamics simulations. The bound Zn(II) ion (gray) is coordinated by four histidine side chains from the PrPC octarepeat region. (B) The changes in α-helicity of the PrPC C-terminal domain during the whole time of the simulations for apo-PrPC (blue) and Zn(II)-bound PrPC (red), compared to the 1QLX13 NMR model of PrPC (dashed black line). (C,D) Torsion angles χ2 versus χ1, characterizing the side chain conformations of tryptophan residues from the N-terminal domain of PrPC in apo form (C) and with a bound Zn(II) ion (D). Trp65 is blue, Trp73 is green, Trp81 is violet, and Trp89 is yellow.
In the second step we generated a 100 ns trajectory for holo-PrPC bound to a Zn(II) ion. Because our NMR experiments suggest an involvement of histidine Nε2 atoms in the Zn(II) coordination (Fig. S1), our initial model involved Zn(II) coordinated by the four deprotonated (i.e., neutral) Nε2 atoms from the His61, His69, His77, and His85 residues, located in the octarepeat region. The distances between the histidine ε2 nitrogen atoms and the Zn(II) ion were stable over the simulation time, with average values around ~ 2.15 Å (Fig. S3) and fluctuations in the range from 1.96 to 2.49 Å.
The simulation for holo-PrPC also converged after ~ 25 ns (Fig. S2). During the simulation time the N-terminal domain moved closer to the C-terminal domain, in a similar way as for the apo form, although the resulting structure was less compact (Fig. 6A, model marked in red). The Zn(II)-bound octarepeat region moved to the C-terminus of α-helix 2 and unfolded this region (Fig. 6B).
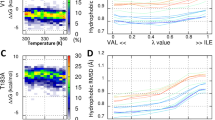
Surprisingly, the main changes in PrPC after Zn(II) binding involved stabilization of α-helix 3 (Fig. 6B). We therefore used MD simulations to investigate possible mechanisms for this stabilization. In apo form the C-terminal region of α-helix 3, involving Ser222-Arg228, becomes unfolded during a 195 ns simulation (Fig. 6B). This process may be mediated by Met166 from the β2-α2 loop. In the initial steps of the apo-PrPC simulation, the C-terminal fragment of α-helix 3 forms hydrogen bonds with the β2-α2 loop, stabilizing α-helix 3 (Fig. 7A, Fig. S4). As the N-terminal domain can move freely, the Nδ1 atom of His111 forms a hydrogen bond with the Oε1 atom of Glu168, and disrupts the hydrogen bond between Oε2 of Glu168 and OH of Tyr226 (Fig. S5). The newly formed hydrogen bond allows Met166 to rotate (Fig. S4, Fig. S5), and then form a hydrogen bond between Sδ of Met166 and NH2 of Arg228 (Fig. S5). In the final step, Arg228 forms a hydrogen bond involving OH of Tyr163 (Fig. 7B, S6), resulting in unfolding of the C-terminal part of α-helix 3. In difference from the apo-PrPC simulation, such unfolding of α-helix 3 was not observed in the simulation of the Zn(II)-PrPC complex. In the holo form, α-helix 3 is initially stabilized by the β2-α2 loop in a similar manner as initially in the apo form (Fig. 7C, Fig. S7). In the next step, however, the three hydrophobic residues Ala120, Val121, and Val122 intercalate between the β2-α2 loop and α-helix 3, thereby separating Met166 and Arg228 (Fig. 7D, Fig. S8). Finally, α-helix 3 becomes stabilized by the N-terminal domain, where the interaction involves four possible hydrogen bonds (Fig. 7D, Fig. S9).
Different behavior of α-helix 3 in apo (blue) and holo (red) PrPC. The C-terminal region of α-helix 3 is stabilized by the β2-α2 loop (A,C). In the apo form formation of a His111-Glu168 hydrogen bond results in rotation of Met166, which then affects the hydrogen bond formation between Arg228 and Tyr163, resulting in partial α-helix 3 unfolding (B). In the holo form the hydrophobic amino acids Ala120, Val121, Val122 separate the β2-α2 loop from α-helix 3, and the N-terminal domain stabilizes α-helix 3 (D).
Our fluorescence experiments showed a blue shift for the PrPC fluorescence maximum upon Zn(II) binding (Fig. 3), which suggests less exposure to the solvent and/or partial structure induction around the tryptophan residues. To investigate if such a phenomenon correlates with our MD simulations, we analyzed the χ1 and χ2 torsion angles for all tryptophan residues. The χ2- χ1 plots for the tryptophan residues located in the octarepeat region are presented in Fig. 6C,D, while χ2- χ1 plots for other tryptophan residues are presented in Figure S10. From the seven analyzed tryptophan residues two residues from the octarepeat region (i.e., Trp65 and Trp89) showed a decrease in mobility, defined as a narrower range of preferred χ2-χ1 angles, for Zn(II)-bound PrPC. Interestingly, two tryptophan residues not located in the octarepeat region, i.e. Trp57 and Trp99, also showed a decrease in mobility upon Zn(II) binding. This suggests that Zn(II) binding slows down PrPC dynamics not only in the octarepeat region itself, but also in the regions before and after it. In addition, χ1 for Trp81 and Trp89 from the octarepeat region more often adopted an unstable gauche- conformation when Zn(II) was bound to PrPC. Gauche- conformations were also prominent for Trp31 and Trp57, which are located outside the octarepeat region, when Zn(II) was present. This observation suggests that Zn(II) binding can act as a driving force for PrPC to access a different conformational pool, that rarely is available for apo-PrPC.
Characterization of the PrPC-Zn(II) complex using small angle X-ray scattering
Earlier results of Spevacek et al.36 suggest that Zn(II) binding to the PrPC protein induces a tertiary fold between the N-terminal and C-terminal domains. To investigate this hypothesis we performed small angle X-ray scattering (SAXS) experiments. In the initial SAXS analysis, we investigated scattering parameters that can be easily and directly obtained from the SAXS curve without any external structural models, namely the radius of gyration (Rg) and the pair distance distribution function P(r). The latter is a probability function for finding two particles at a given distance, and it allows us to easily calculate the maximum diameter of a protein. The Guinier plot (Fig. 8A, inset) for apo-PrPC and Zn(II)-bound PrPC was linear (s·Rg < 1.3), indicating a monodisperse sample. This observation allowed us to calculate the radius of gyration (Rg) from the Guinier approximation. Our calculated Rg values for the apo-PrPC (2.51 ± 0.11 nm) and Zn(II)-bound PrPC (2.66 ± 0.07 nm) were the same within experimental error. Despite no clear differences in the Rg values, we proceeded with calculation of the P(r) function. This function, and also the determined maximum particle diameter (Dmax), differed significantly between apo- and Zn(II)-bound PrPC. The Dmax values were ~ 11.6 nm for apo-PrPC and ~ 10.2 nm for Zn(II)-bound PrPC, suggesting a reduction in the maximum PrPC diameter of ~ 1.4 nm (Fig. 8B).
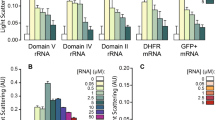
SAXS studies of PrPC in apo- (blue) and Zn(II)-bound (red) forms. (A) Experimental SAXS data fitted with GAJOE (black line χ2 = 1.03 and 0.95, respectively). SAXS curves were displaced along the vertical axis for clarity. Left bottom inlet shows linear fits (black lines) in the Guinier regime. (B) P(r) functions for the two protein forms. (C) Rg distribution for the two protein forms. (D) Dmax distribution for the two protein forms. (E,F) Pool of conformers representing apo-PrPC (E) and Zn(II)-bound PrPC (F).
At the next stage, the PrPC conformation was studied by the ensemble optimization method (EOM)41, which is commonly used for conformational analysis of intrinsically disordered proteins or proteins with intrinsically disordered domains42. In this approach, the sample is treated as a polydisperse mixture of different conformers, and the experimental SAXS curve is fitted as a sum of weighted calculated scattering intensities from all conformers. As initial conformers, we used the apo and Zn(II)-bound PrPC models generated in our molecular dynamics simulations. Thus, the SAXS curve can be represented as a population of molecules with different Rg or Dmax values. In the Rg distribution, apo-PrPC occupied three major and distinct conformations (Fig. 8C, blue line): compact (Rg ~ 2.0 nm, ~ 71%), intermediate (Rg ~ 4.0 nm, ~ 14%), and extended (Rg ~ 5.2 nm, ~ 14%). Addition of Zn(II) (Fig. 8C, red line) resulted in the appearance of a predominant conformation with Rg ~ 2.3 nm (~ 62%), reduction of the extended conformations to Rg ~ 4.2 nm (cumulatively 24%) and a reduction of the intermediate conformations to Rg ~ 3.4 (~ 12%). Similar features were observed in the Dmax distribution: apo-PrPC coexisted in three main conformations (Fig. 8D, blue line, Fig. 8E) characterized by different Dmax values: compact (Dmax ~ 7.5 nm, ~ 71%), intermediate (Dmax ~ 12 nm, ~ 14%), and extended (Dmax ~ 16 nm, ~ 14%). For PrPC bound to a Zn(II) ion, Dmax of the extended PrPC conformers was shifted to ~ 12 nm (~ 24%) and ~ 11 nm (12%), with a reduction in the amount of the compact conformations to ~ 62% and an increase in Dmax to ~ 8.1 nm (Fig. 8D, red line, Fig. 8F). Overall, the SAXS results clearly show that binding of Zn(II) ions induces a more compact fold of the PrPC protein.
Discussion
Earlier studies performed on murine PrPC have indicated that the N-terminal domain becomes more ordered and interacts with the C-terminal domain upon addition of Zn(II) ions36. To investigate such possible effects in human PrPC, and if they might correspond to changes in the secondary structure of the protein, circular dichroism spectroscopy experiments were conducted. Estimation of the secondary structure from the CD spectrum of the apo-PrPC by the BeStSel software43 showed an α-helical content similar to that reported for the PrPC NMR structure (PDB ID: 1QLX12) (Table S1). The observed small discrepancy between the amounts of other structural motifs in apo-PrPC, observed by CD spectroscopy and reported in the 1QLX NMR model, are probably caused by multiple structural states of the flexible N-terminal domain13,14,15. Addition of twenty molar equivalents of Zn(II) to PrPC immediately changed the CD spectrum (Fig. 2). The observed isodichroic point at ~ 242 nm suggests a decrease in the amount of regular α-helices and increase in the amount of parallel β-sheets43. This phenomenon probably corresponds to structural transitions within the N-terminal domain and α-helices 2 and 3, as α-helix 1 is flanked by two β-sheets (Fig. 1) and it is also more conformationally stable12. The proposed structural transitions would involve distortion of α-helices 2 and 3 together with structure induction in the octarepeat region with β-like motifs forming around the Zn(II) ion, rather than formation of parallel β-sheets. Such structural transitions suggest a tertiary contact between the Zn(II)-occupied octarepeat region and the C-terminal domain, in line with previous observations for mouse PrPC(23–230) done by Spevacek et. al36. The increase in the amount of distorted α-helices probably originates from contacts between the Zn(II)-saturated octarepeat region and helices 2 and 336, leading to local secondary structure instabilities. According to our CD titrations the Kd for PrPC-Zn(II) in phosphate buffer is 12.1 µM. This value is close to that reported in a recently published ITC study, i.e. 16.9 µM38. The Kd values for the PrPC-Zn(II) complex that we here derive with different techniques (CD and fluorescence spectroscopy) and in different buffers (NEM and phosphate buffer) are slightly different, as is to be expected, but they all are in the range of 10–30 µM (Figs. 2 and 3). Thus, we conclude that the dissociation constant for the Zn(II)-PrPC complex is in the low micromolar range.
Fluorescence spectroscopy also suggests structure induction in the N-terminal domain of PrPC in the presence of Zn(II) ions. The fluorescence peak with a maximum at 313 nm most likely originates from tyrosine residues, as PrPC does not contain any structured tryptophan residues12,39,44. Most tyrosine residues (nine out of twelve) are located in the C-terminal domain. Thus, minor changes in the tyrosine fluorescence signal likely correspond to changes in the local environment of C-terminal tyrosine residues. If such changes are induced by Zn(II) binding to the N-terminal region, they would arguably constitute evidence for interactions between the N-terminal and C-terminal domains.
During the titrations with ZnCl2 the initial fluorescence peak at 347 nm, which corresponds to multiple exposed tryptophan residues, split into two well-resolved peaks of lower intensity and with maxima at 333 nm and 343 nm. These two peaks likely correspond to tryptophan residues buried and exposed to bound water molecules, respectively39. This observation, which is connected also to fluorescence quenching of some tryptophan residues, suggests structure induction in the PrPC octarepeat region upon Zn(II) binding. As a comparison, crystallographic studies of Cu(II) ions bound to the HGGGW fragment of the octarepeat region showed that tryptophan residues participate in Cu(II) coordination by forming a hydrogen bond with a water molecule axially bound to Cu(II)35. The octarepeat region appears to bind Cu(II) and Zn(II) ions via the same histidine residues, but in different binding conformations25,29,35. The Kd values calculated from our fluorescence experiments in phosphate buffer are in good agreement with our CD results, i.e. in the 10–30 µM range, which agrees with the results of previously published ICT experiments38. This suggests that tryptophan residues might be indirectly involved in Zn(II) binding to PrPC.
Our NMR data suggest that binding of Zn(II) ions to PrPC produces relatively small alterations in the 3D structure of the folded C-terminal domain, as the position of the backbone amide resonances did not show significant changes. Nevertheless, Zn(II) binding leads to increased linewidths for several backbone resonances, indicating changes in spin–spin relaxation. This was confirmed by measuring 15N R2 values. A comparison can be made with the Amyloid-β (Aβ) peptide related to Alzheimer´s disease, which is well known to bind metal ions24,45. In analogy with the interpretations proposed in a previous NMR study of Zn(II) binding to Aβ, binding of Zn(II) to the prion protein may give rise to folding of the peptide chain around the bound metal ion46, which could explain the observed signal intensity changes in the HSQC spectrum. The NMR diffusion data furthermore suggest a more compact 3D structure of PrPC in the presence of Zn(II), characterized by stronger interactions and an increased number of contacts between the N- and C-terminal domains, which is in agreement with the previously reported data for mouse PrPC36.
The prion protein structures deposited in PDB do not have fully defined N-terminal domain structures, as this domain is in a dynamic equilibrium between random coil, PPII helix, and β-turn secondary structures13,14,15. Two deposited structures of human PrPC proteins, one for the G127V mutation and the other for wild-type PrP (5YJ4, 5YJ5), obtained from NMR and MD studies, contain in addition to the C-terminal domain also a collection of proposed models of the N-terminal domain47. However, such models do not answer what happens to PrPC upon metal ion binding. Our MD simulations indicate that the N-terminal domain, upon Zn(II) binding, interacts with the C-terminal domain and destabilizes α-helix 2. Similar tertiary folds were previously observed for murine PrPC interacting with Cu(II), Zn(II), and Cd(II) ions36,38,48,49, and may therefore have similar functions in metal ion recognition.
The MD simulations suggest also that PrPC upon Zn(II) binding might follow a different folding path, with altered interactions between the β2-α2 loop and α-helix 3. In our proposed model for Zn(II)-bound PrPC, an interaction between His111 and Glu168 caused unfolding of the α-helix 3 C-terminal part. Such an interaction between His111 and the β2-α2 loop might be a first step in PrPC unfolding and fibrillization. Introduction of different amino acids, in the form of glutamines in the β2-α2 loop, has previously been shown to shorten the lag phase for mouse PrPC fibrillization50. As histidine residues at pH 7.4 can act both as donors and acceptors for hydrogen bonding, it is possible that His111 can form hydrogen bonds with glutamines from the β2-α2 loop, disturbing the α-helix 3 and resulting in faster PrPC fibrillization. On the other hand, Zn(II) binding to PrPC could disrupt such interactions by burying His111 in a different protein region. Previous in vitro studies on different forms of PrPC seem to support this idea, as Zn(II) inhibits PrP aggregation51,52.
The radius of gyration for apo-PrPC and Zn(II)-bound PrPC, determined by Guinier approximation, remained unchanged within experimental error, and were furthermore similar to those reported by Thakur et al. for apo and Cu(II)-bound PrPC48. The decrease in Dmax after addition of Zn(II) ions indicates that the N-terminal PrPC domain adopts a more compact shape in the presence of Zn(II) ions. The distance distribution function moreover suggests a reduction in the maximum particle dimension by ~ 1 nm in the presence of Zn(II), similar to what was earlier observed for the Cu(II)-PrPC complex48. Ensemble analysis performed by a genetic algorithm produced Rg and Dmax distributions similar to those for the Cu(II)-PrPC complex48. Our results suggest that Zn(II) binding to the PrPC shows similar features to those observed for Cu(II) binding, resulting in decreased N-terminal conformational freedom and therefore partial folding of the octarepeat region around the bound metal ion.
Using crosslinking, mass spectrometry, and NMR, McDonald et al. suggested that the conformational states of Cu(II)-bound PrPC are available for apo PrPC and vice versa, with Cu(II) acting as a switch that shifts this equilibrium53. Our CD, NMR and MD results suggest that PrPC upon Zn(II) binding adopts a distinct conformation, that might be unavailable for apo PrPC. This distinct conformation is a result of the metal binding by the histidine residues that during coordination of a metal ion cannot form contacts with other parts of the protein. On the other hand our SAXS analysis shows the overlap between the Rg and Dmax for apo and Zn(II)-bound PrPC. This might suggest that even though major conformations of apo and Zn(II)-bound PrPC differ, some conformations are similar, which is in line with previous studies53.
While Zn(II) can promote dimerization of the lipid-anchored octarepeat region54, we have recently shown that upon interaction with Zn(II) ions octarepeat peptides (consisting of PrP(58–93)) form fibrillar structures with features characteristic for amyloids: they form the characteristic cross-β structure and bind the thioflavin T and Congo Red dyes55. We proposed that the reason for fibril formation could be a lack of the previously reported tertiary contact between the metal ion-saturated octarepeat region and the C-terminal domain36,49. Indeed, the octarepeat region seems to have an important role in PrPSc formation. Studies of antibodies targeting the C-terminal domain suggest that the octarepeat region is required to maintain high PrPC toxicity, while ligands targeting octarepeat region seem to reduce PrPC toxicity56.
Despite many years of research, the PrPC function in terms of metal ion binding is still elusive, and it is still unclear if metal imbalance is part of the pathology in prion diseases. Out of three proposed functions for PrPC, i.e. zinc sensor, transporter, or sequester57,58,59, the first two seem to be the most tempting. With internalization of PrPC into the Golgi apparatus and endosomes upon interaction with Zn(II)60,61, or enhancement of zinc uptake by PrPC in neurons62 at Zn(II) concentrations around or below 100 µM, our calculated Kd values for the PrPC-Zn(II) complex in the 10–30 µM range appear reasonable and physiologically important.
In summary, our results suggest that binding of Zn(II) ions to the PrPC N-terminal domain via four His residues from the octarepeat region induces a structural transition in the C-terminal α-helices 2 and 3, promotes interaction between the N-terminal and C-terminal domains, reduces the folded protein size, and modifies the internal structural dynamics. The stabilization of α-helix 3 by Zn(II) binding to the N-terminal domain may explain why Zn(II) appears to inhibit PrPC fibrillization.
Materials and methods
Materials
The human recombinant protein PrPC(23–231) was expressed using previously published protocols63,64. The pRSETB vector (Invitrogen, USA) was used to clone plasmid containing a fusion of human PrPC protein with a thrombin cleavage site and an N-terminal HisTag. The construct was expressed in E. coli (BL21-DE3) together with 100 µg/mL ampicillin antibiotic and induced by isopropyl β-D-galactopyranoside (IPTG) at OD600 = 0.8. A buffer containing 100 mM Tris at pH 8, 10 mM K2HPO4, 10 mM glutathione (GSH), 6 M GuHCl, and 0.5 mM phenylmethane sulfonyl fluoride (PMSF) was used during sonication of the lysates. Next, the supernatant was loaded to an Ni–NTA column (GE Healthcare) and eluted with buffer E (100 mM Tris at pH 5.8, 10 mM K2HPO4, and 500 mM imidazole). The imidazole was removed with two-step dialysis. After thrombin cleavage the huPrPC(23–231) was concentrated using an Amicon Ultra filter (cutoff: 3 kDa).
The 15N-labeled and 13C,15N-double labeled forms form of huPrPC(23–231) were prepared by the same protocol, except that the M9 media was supplemented with 1 g of 15NH4Cl and 2 g of the 13C-glucose (both CIL Inc, Cambridge, UK) for one liter of E. coli culture. Mass spectrometry was used for quality control of the samples. Protein concentrations were determined by spectrophotometry, using the extinction coefficient ε280 = 57,995 M-1 cm-165.
Circular dichroism (CD) spectroscopy
The initial potassium acetate buffer was exchanged to phosphate or N-ethylmorpholine (NEM) buffer using a Amicon® Ultra-0.5 centrifugal filter device (Merck) with an NMWL cutoff of 3 kDa. After the first round of concentration, 10 mM phosphate, pH 7.4 or 10 NEM, pH 7.4 buffer was added to increase the sample volume up to 500 μl. The sample was then centrifuged again, and the whole procedure was repeated three times. After buffer exchange the huPrPC sample was filtered using an Ultrafree®-MC centrifugal filter with 0.22 μm pore size.
Circular dichroism spectra were collected using a Jasco J-815 spectropolarimeter (Jasco, Tokyo, Japan). The data were collected in a step scan mode with 0.5 nm resolution, 2 nm bandwidth and digital integration time of 4 s. The spectra in a 190–260 nm and 200–260 nm range were recorded in 0.1 mm and 10 mm cells respectively. All experiments were performed in triplicate with buffer baseline correction at 25 °C. Experimental data were fitted to tight binding equation66 (1):
where [θ]0 and [θ]∞ are the initial and saturated CD intensities, [Zn] is the Zn(II) concentration, [PrPC] is the PrPC concentration and Kdapp is the dissociation constant of Zn(II)-PrPC complex. Because possible binding of Zn(II) ions to the buffer is not taken into account, the calculated dissociation constants should be considered to be apparent.
Estimation of the secondary structure from the CD spectra was performed by the BeStSel software43.
Fluorescence spectroscopy
Intrinsic tryptophan fluorescence was measured on a FP-8300 spectrofluorimeter (Jasco, Tokyo, Japan). Samples were excited with a wavelength of 280 nm and spectra were collected from 300 to 450 nm with 5 nm excitation and emission bandwidth. All titrations were acquired in triplicate at 25 °C. The fluorescence data were fitted with a modified Hill equation67 (2):
where f0 and f∞ are the initial and saturated fluorescence intensities, [Zn] is the Zn(II) concentration, Kdapp is the apparent dissociation constant, nH is the Hill coefficient. During the fitting procedure nH was 1.58 ± 0.13.
Nuclear magnetic resonance (NMR) spectroscopy
NMR experiments were conducted at 298 K on an Agilent DDR2 800 MHz spectrometer operated at a magnetic field of 18.8 T (1H resonance frequency 799.838 MHz). 2D 1H-15N-HSQC and diffusion experiments were recorded for a sample containing 300 µM uniformly 15N-labeled human PrPC(23–231) protein dissolved in 50 mM HEPES-d18 (CIL, Cambridge, UK) at pH 7.0 with 50 mM NaCl added.
Due to limited amount of the 13C, 15N-double labeled protein the aromatic 1H-13C HSQC was collected for the sample prepared in a 3 mm sample tube at a concentration of 100 µM also in 50 mM HEPES-d18, 50 mM NaCl, pH 7.0. NMR spectra were recorded before and after addition of ZnCl2 (in two steps of respectively 50 μM and 120 μM). The recorded spectra were referenced indirectly to DSS (sodium 2,2-dimethyl-2-silapentane-5-sulfonate) using a Ξ = 0.251449530 and 0.101329118 ratio for 13C and 15N resonances, respectively68. All NMR data were processed with NMRPipe69 and analyzed with the Sparky70 software.
The 15N spin–spin relaxation rates (R2) were determined at 18.8 T using a pulse sequence based on previously published experiments71 and present in the BioPack library (Agilent Inc., PaloAlto, CA, USA). Due to fast relaxation of amide groups in human PrPC(23–231), the 15N R2 values were calculated only with six delays—10, 30, 50, 70, 90, and 110 ms. The experimental errors were estimated as standard deviation from 500 Monte Carlo simulations with the Relax (version 4.0.3) software72.
The diffusion data were collected using the DPFGDSTE (Double Polar Field Gradient Double Stimulated Echo) pulse sequence73. 28 and 15 data points were acquired to extract information about the translational diffusion coefficients (Dtr), respectively for the apo and the Zn(II)-bound forms of the huPrPC(23–231) protein. The DOSY data was processed using either VnmrJ v4.3 (Agilent Technologies Inc., USA) or MnovaNMR (Mestrelab Research SL. Santiago de Compostela, Spain) software. The Dtr experimental values were calculated according to the Stejskal-Tanner equation74 (3):
where γH is the 1H gyromagnetic ratio, δ is gradient duration (2 ms), Δ is diffusion time (150 ms), and G is the gradient strength.
Molecular dynamics simulations
The initial model of full length human PrPC was constructed by adding the missing N-terminal domain and three C-terminal residues to the 1QLX NMR structure 12. The Zn(II) ion was placed near four histidine residues from the octarepeat domain using a sculpting tool implemented in pymol. Molecular dynamics simulations were performed in GROMACS 2019.275 using the GROMOS 53A676 force field, which contains nonbonded parameters for Zn(II). Both apo and Zn(II) bound PrPC models were placed in a rectangular box with periodic boundary conditions (PBC) 5.0 nm from the box wall and solvated with a single point charge (SPC)77 water model restrained by the SETTLE algorithm78 using Van der Waals radii79. All systems were neutralized with Cl- ions and the energy was minimized with steepest descent minimization up to 5,000 steps. Temperature and pressure were equilibrated over 100 ps with a 1 fs time step using a modified Berendsen thermostat80 and the Parrinello-Rahman barostat81, respectively using the particle mesh Ewald (PME) method82. For both the apo and the Zn(II)-bound PrPC molecule, the final trajectories were generated at 300 K over 195 ns with 2 fs time step or 100 ns with 1 fs time step, respectively. All covalent bonds were constrained using the LINCS83 algorithm. The final trajectories were analyzed in the VMD software84.
Small angle X-ray scattering
The small angle X-ray scattering (SAXS) data for human PrPC(23–231) protein in solution were collected at the P12 beamline, operated by EMBL Hamburg at the PETRA III storage ring (DESY, Hamburg, Germany)85 using synchrotron radiation with a wavelength of 1.24 nm. The range of the scattering vector was from 0.105 to 3.793 nm-1.
For the SAXS experiments, the initial buffer was exchanged to MOPS buffer using an Amicon Ultra-0.5 centrifugal filter device (Merck) with an NMWL cutoff of 3 kDa, as described above. During the experiments the concentration of the huPrPC protein was 2 mg/ml. ZnCl2 was added from a 50 mM stock solution to a final concentration of 88 μM, corresponding to a 1:1 Zn(II):huPrPC(23–231) molar ratio.
The SAXS data were processed and analyzed using the PRIMUS software86 from the ATSAS 2.8 package87. The radius of gyration was obtained via the Guinier approximation for s ⋅Rg < 1.3. The pair distribution function, P(r), and the maximum intramolecular distance, Dmax, were calculated using the GNOM software88. Because the N-terminal domain of apo-PrPC is unstructured and forms multiple conformations, the SAXS data were fitted with 3D conformations from our molecular dynamics simulations using 100 cycles of GAJOE41.
References
Prusiner, S. B. Nobel lecture: Prions. Proc. Natl. Acad. Sci. 95, 13363–13383 (1998).
Harrison, P. M., Khachane, A. & Kumar, M. Genomic assessment of the evolution of the prion protein gene family in vertebrates. Genomics 95, 268–277 (2010).
Salès, N. et al. Cellular prion protein localization in rodent and primate brain: PrP c brain localization. Eur. J. Neurosci. 10, 2464–2471 (1998).
Stahl, N. Scrapie prion protein contains a phosphatidylinositol glycolipid. Cell 51, 229–240 (1987).
Haraguchi, T. et al. Asparagine-linked glycosylation of the scrapie and cellular prion proteins. Arch. Biochem. Biophys. 274, 1–13 (1989).
Heller, U., Winklhofer, K. F., Heske, J., Reintjes, A. & Tatzelt, J. Post-translational import of the prion protein into the endoplasmic reticulum interferes with cell viability: A critical role for the putative transmembrane domain. J. Biol. Chem. 278, 36139–36147 (2003).
Colby, D. W. & Prusiner, S. B. Prions. Cold Spring Harb. Perspect. Biol. 3, a006833–a006833 (2011).
Gajdusek, D. Unconventional viruses and the origin and disappearance of kuru. Science 197, 943–960 (1977).
Aguzzi, A., Lakkaraju, A. K. K. & Frontzek, K. Toward therapy of human prion diseases. Annu. Rev. Pharmacol. Toxicol. 58, 331–351 (2018).
Söderberg, K. L., Guterstam, P., Langel, Ü. & Gräslund, A. Targeting prion propagation using peptide constructs with signal sequence motifs. Arch. Biochem. Biophys. 564, 254–261 (2014).
Gielnik, M. et al. The Engineered Peptide Construct NCAM1-Aβ Inhibits Aggregation of the Human Prion Protein (PrP). https://doi.org/10.1101/2021.01.04.425177 (2021).
Zahn, R. et al. NMR solution structure of the human prion protein. Proc. Natl. Acad. Sci. 97, 145–150 (2000).
Gill, A. C. Post-translational hydroxylation at the N-terminus of the prion protein reveals presence of PPII structure in vivo. EMBO J. 19, 5324–5331 (2000).
Zahn, R. The octapeptide repeats in mammalian prion protein constitute a pH-dependent folding and aggregation site. J. Mol. Biol. 334, 477–488 (2003).
Smith, C. J. et al. Conformational properties of the prion octa-repeat and hydrophobic sequences. FEBS Lett. 405, 378–384 (1997).
Riek, R., Hornemann, S., Wider, G., Glockshuber, R. & Wüthrich, K. NMR characterization of the full-length recombinant murine prion protein, m PrP(23–231). FEBS Lett. 413, 282–288 (1997).
Pérez, D. R., Damberger, F. F. & Wüthrich, K. Horse prion protein NMR structure and comparisons with related variants of the mouse prion protein. J. Mol. Biol. 400, 121–128 (2010).
Biljan, I. et al. Structural basis for the protective effect of the human prion protein carrying the dominant-negative E219K polymorphism. Biochem. J. 446, 243–251 (2012).
Ciric, D. & Rezaei, H. Biochemical insight into the prion protein family. Front. Cell Dev. Biol. 3, 15 (2015).
Wulf, M.-A., Senatore, A. & Aguzzi, A. The biological function of the cellular prion protein: An update. BMC Biol. 15, 34 (2017).
Castle, A. R. & Gill, A. C. Physiological functions of the cellular prion protein. Front. Mol. Biosci. 4, 633 (2017).
Kawahara, M., Kato-Negishi, M. & Tanaka, K. Neurometals in the pathogenesis of prion diseases. Int. J. Mol. Sci. 22, 1267 (2021).
Pushie, M. J. et al. Prion protein expression level alters regional copper, iron and zinc content in the mouse brain. Metallomics 3, 206 (2011).
Wärmländer, S. K. T. S. et al. Metal binding to the amyloid-β peptides in the presence of biomembranes: Potential mechanisms of cell toxicity. JBIC J. Biol. Inorg. Chem. 24, 1189–1196 (2019).
Jackson, G. S. et al. Location and properties of metal-binding sites on the human prion protein. Proc. Natl. Acad. Sci. 98, 8531–8535 (2001).
Giese, A., Levin, J., Bertsch, U. & Kretzschmar, H. Effect of metal ions on de novo aggregation of full-length prion protein. Biochem. Biophys. Res. Commun. 320, 1240–1246 (2004).
Jószai, V. et al. Mixed metal copper(II)-nickel(II) and copper(II)-zinc(II) complexes of multihistidine peptide fragments of human prion protein. J. Inorg. Biochem. 112, 17–24 (2012).
Treiber, C., Thompsett, A. R., Pipkorn, R., Brown, D. R. & Multhaup, G. Real-time kinetics of discontinuous and highly conformational metal-ion binding sites of prion protein. JBIC J. Biol. Inorg. Chem. 12, 711–720 (2007).
Evans, E. G. B. & Millhauser, G. L. Copper- and zinc-promoted interdomain structure in the prion protein: A mechanism for autoinhibition of the neurotoxic N-terminus. in Progress in Molecular Biology and Translational Science, Vol. 150. 35–56 (Elsevier, 2017).
Millhauser, G. L. Chapter 12. Copper and prion protein function: A brief review of emerging theories of neuroprotection. in Drug Discovery (eds. Milardi, D. & Rizzarelli, E.). 249–258. https://doi.org/10.1039/9781849733014-00249 (Royal Society of Chemistry, 2011).
Vallee, B. L. & Auld, D. S. Zinc coordination, function, and structure of zinc enzymes and other proteins. Biochemistry 29, 5647–5659 (1990).
MacDonald, R. S. The role of zinc in growth and cell proliferation. J. Nutr. 130, 1500S-1508S (2000).
Weiss, J. H., Sensi, S. L. & Koh, J. Y. Zn2+: a novel ionic mediator of neural injury in brain disease. Trends Pharmacol. Sci. 21, 395–401 (2000).
Assaf, S. Y. & Chung, S.-H. Release of endogenous Zn2+ from brain tissue during activity. Nature 308, 734–736 (1984).
Burns, C. S. et al. Molecular features of the copper binding sites in the octarepeat domain of the prion protein †. Biochemistry 41, 3991–4001 (2002).
Spevacek, A. R. et al. Zinc drives a tertiary fold in the prion protein with familial disease mutation sites at the interface. Structure 21, 236–246 (2013).
Walter, E. D., Stevens, D. J., Visconte, M. P. & Millhauser, G. L. The prion protein is a combined zinc and copper binding protein: Zn 2+ alters the distribution of Cu 2+ coordination modes. J. Am. Chem. Soc. 129, 15440–15441 (2007).
Markham, K. A., Roseman, G. P., Linsley, R. B., Lee, H.-W. & Millhauser, G. L. Molecular features of the Zn2+ binding site in the prion protein probed by 113Cd NMR. Biophys. J. 116, 610–620 (2019).
Reshetnyak, Y. K. & Burstein, E. A. Decomposition of protein tryptophan fluorescence spectra into log-normal components. II. The statistical proof of discreteness of tryptophan classes in proteins. Biophys. J. 81, 1710–1734 (2001).
Le Brun, N. et al. Charge compensated binding of divalent metals to bacterioferritin: H+ release associated with cobalt(II) and zinc(II) binding at dinuclear metal sites. FEBS Lett. 397, 159–163 (1996).
Tria, G., Mertens, H. D. T., Kachala, M. & Svergun, D. I. Advanced ensemble modelling of flexible macromolecules using X-ray solution scattering. IUCrJ 2, 207–217 (2015).
Kikhney, A. G. & Svergun, D. I. A practical guide to small angle X-ray scattering (SAXS) of flexible and intrinsically disordered proteins. FEBS Lett. 589, 2570–2577 (2015).
Micsonai, A. et al. Accurate secondary structure prediction and fold recognition for circular dichroism spectroscopy. Proc. Natl. Acad. Sci. 112, E3095–E3103 (2015).
Lakowicz, J. R. Principles of Fluorescence Spectroscopy. (Springer, 2010).
Wärmländer, S. et al. Biophysical studies of the amyloid β-peptide: Interactions with metal ions and small molecules. ChemBioChem 14, 1692–1704 (2013).
Abelein, A., Gräslund, A. & Danielsson, J. Zinc as chaperone-mimicking agent for retardation of amyloid β peptide fibril formation. Proc. Natl. Acad. Sci. 112, 5407–5412 (2015).
Zheng, Z. et al. Structural basis for the complete resistance of the human prion protein mutant G127V to prion disease. Sci. Rep. 8, 13211 (2018).
Thakur, A. K., Srivastava, A. K., Srinivas, V., Chary, K. V. R. & Rao, C. M. Copper alters aggregation behavior of prion protein and induces novel interactions between Its N- and C-terminal regions. J. Biol. Chem. 286, 38533–38545 (2011).
Evans, E. G. B., Pushie, M. J., Markham, K. A., Lee, H.-W. & Millhauser, G. L. Interaction between prion protein’s copper-bound octarepeat domain and a charged C-terminal pocket suggests a mechanism for N-terminal regulation. Structure 24, 1057–1067 (2016).
Avbelj, M., Hafner-Bratkovič, I. & Jerala, R. Introduction of glutamines into the B2–H2 loop promotes prion protein conversion. Biochem. Biophys. Res. Commun. 413, 521–526 (2011).
Bocharova, O. V., Breydo, L., Salnikov, V. V. & Baskakov, I. V. Copper(II) inhibits in vitro conversion of prion protein into amyloid fibrils †. Biochemistry 44, 6776–6787 (2005).
Pan, K., Yi, C.-W., Chen, J. & Liang, Y. Zinc significantly changes the aggregation pathway and the conformation of aggregates of human prion protein. Biochim. Biophys. Acta BBA Proteins Proteom. 1854, 907–918 (2015).
McDonald, A. J. et al. Altered domain structure of the prion protein caused by Cu2+ binding and functionally relevant mutations: Analysis by cross-linking, MS/MS, and NMR. Structure 27, 907-922.e5 (2019).
Kenward, A. G., Bartolotti, L. J. & Burns, C. S. Copper and zinc promote interactions between membrane-anchored peptides of the metal binding domain of the prion protein †. Biochemistry 46, 4261–4271 (2007).
Gielnik, M. et al. PrP (58–93) peptide from unstructured N-terminal domain of human prion protein forms amyloid-like fibrillar structures in the presence of Zn 2+ ions. RSC Adv. 9, 22211–22219 (2019).
Sonati, T. et al. The toxicity of antiprion antibodies is mediated by the flexible tail of the prion protein. Nature 501, 102–106 (2013).
Watt, N. T. & Hooper, N. M. The prion protein and neuronal zinc homeostasis. Trends Biochem. Sci. 28, 406–410 (2003).
Lehmann, S. Metal ions and prion diseases. Curr. Opin. Chem. Biol. 6, 187–192 (2002).
Watt, N. T., Griffiths, H. H. & Hooper, N. M. Neuronal zinc regulation and the prion protein. Prion 7, 203–208 (2013).
Pauly, P. C. & Harris, D. A. Copper stimulates endocytosis of the prion protein. J. Biol. Chem. 273, 33107–33110 (1998).
Brown, L. R. & Harris, D. A. Copper and zinc cause delivery of the prion protein from the plasma membrane to a subset of early endosomes and the Golgi: Copper and zinc effect on prion protein localization. J. Neurochem. 87, 353–363 (2003).
Watt, N. T. et al. Prion protein facilitates uptake of zinc into neuronal cells. Nat. Commun. 3, 1134 (2012).
Morillas, M., Swietnicki, W., Gambetti, P. & Surewicz, W. K. Membrane environment alters the conformational structure of the recombinant human prion protein. J. Biol. Chem. 274, 36859–36865 (1999).
Zahn, R., von Schroetter, C. & Wüthrich, K. Human prion proteins expressed in Escherichia coli and purified by high-affinity column refolding. FEBS Lett. 417, 400–404 (1997).
Wilkins, M. R. et al. Protein identification and analysis tools in the ExPASy server. in 2-D Proteome Analysis Protocols, Vol. 112. 531–552 (Humana Press Inc., 1999).
Williams, M. A. Protein–ligand interactions: Fundamentals. in Protein-Ligand Interactions (eds. Williams, M. A. & Daviter, T.) Vol. 1008. 3–34 (Humana Press, 2013).
Bindslev, N. Hill in hell. in Drug-Acceptor Interactions. 257–282. https://doi.org/10.3402/bindslev.2008.14 (Co-Action Publishing, 2008).
Wishart, D. S. et al. 1H, 13C and 15N chemical shift referencing in biomolecular NMR. J. Biomol. NMR 6, 135–140 (1995).
Delaglio, F. et al. NMRPipe: A multidimensional spectral processing system based on UNIX pipes. J. Biomol. NMR 6, 313 (1995).
Lee, W., Tonelli, M. & Markley, J. L. NMRFAM-SPARKY: Enhanced software for biomolecular NMR spectroscopy. Bioinformatics 31, 1325–1327 (2015).
Farrow, N. A. et al. Backbone dynamics of a free and a phosphopeptide-complexed Src homology 2 domain studied by 15N NMR relaxation. Biochemistry 33, 5984–6003 (1994).
d’Auvergne, E. J. & Gooley, P. R. Optimisation of NMR dynamic models I. Minimisation algorithms and their performance within the model-free and Brownian rotational diffusion spaces. J. Biomol. NMR 40, 107–119 (2008).
Wu, D. H., Chen, A. D. & Johnson, C. S. An improved diffusion-ordered spectroscopy experiment incorporating bipolar-gradient pulses. J. Magn. Reson. A 115, 260–264 (1995).
Stejskal, E. O. & Tanner, J. E. Spin diffusion measurements: Spin echoes in the presence of a time-dependent field gradient. J. Chem. Phys. 42, 288–292 (1965).
Abraham, M. J. et al. GROMACS: High performance molecular simulations through multi-level parallelism from laptops to supercomputers. SoftwareX 1–2, 19–25 (2015).
Oostenbrink, C., Villa, A., Mark, A. E. & Van Gunsteren, W. F. A biomolecular force field based on the free enthalpy of hydration and solvation: The GROMOS force-field parameter sets 53A5 and 53A6. J. Comput. Chem. 25, 1656–1676 (2004).
Berendsen, H. J. C., Postma, J. P. M., van Gunsteren, W. F. & Hermans, J. Interaction models for water in relation to protein hydration. in Intermolecular Forces (ed. Pullman, B.), Vol. 14. 331–342 (Springer Netherlands, 1981).
Miyamoto, S. & Kollman, P. A. Settle: An analytical version of the SHAKE and RATTLE algorithm for rigid water models. J. Comput. Chem. 13, 952–962 (1992).
van der Bondi, A. Waals volumes and radii. J. Phys. Chem. 68, 441–451 (1964).
Bussi, G., Donadio, D. & Parrinello, M. Canonical sampling through velocity rescaling. J. Chem. Phys. 126, 014101 (2007).
Parrinello, M. & Rahman, A. Polymorphic transitions in single crystals: A new molecular dynamics method. J. Appl. Phys. 52, 7182–7190 (1981).
Essmann, U. et al. A smooth particle mesh Ewald method. J. Chem. Phys. 103, 8577–8593 (1995).
Hess, B., Bekker, H., Berendsen, H. J. C. & Fraaije, J. G. E. M. LINCS: A linear constraint solver for molecular simulations. J. Comput. Chem. 18, 1463–1472 (1997).
Humphrey, W., Dalke, A. & Schulten, K. V. M. D. Visual molecular dynamics. J. Mol. Graph. 14, 33–38 (1996).
Blanchet, C. E. et al. Versatile sample environments and automation for biological solution X-ray scattering experiments at the P12 beamline (PETRA III, DESY). J. Appl. Crystallogr. 48, 431–443 (2015).
Konarev, P. V., Volkov, V. V., Sokolova, A. V., Koch, M. H. J. & Svergun, D. I. PRIMUS : A Windows PC-based system for small-angle scattering data analysis. J. Appl. Crystallogr. 36, 1277–1282 (2003).
Franke, D. et al. ATSAS 2.8: A comprehensive data analysis suite for small-angle scattering from macromolecular solutions. J. Appl. Crystallogr. 50, 1212–1225 (2017).
Svergun, D. I. Determination of the regularization parameter in indirect-transform methods using perceptual criteria. J. Appl. Crystallogr. 25, 495–503 (1992).
Acknowledgements
This research was supported by a research grant (2014/15/B/ST4/04839) from National Science Centre (Poland). The synchrotron SAXS data were collected at beamline P12 operated by EMBL Hamburg at the PETRA III storage ring (DESY, Hamburg, Germany). We would like to thank the beamline staff for kind assistance. S.K.T.S.W. was supported by a grant from the Magnus Bergvall foundation, and A.G. by grants from the Swedish Research Council and the Brain Foundation in Sweden. High-performance computing at University of Rijeka is supported by European Fund for Regional Development (ERDF) and by the Ministry of Science, Education and Sports of the Republic of Croatia under the project number RC.2.2.06-0001.
Author information
Authors and Affiliations
Contributions
L.Z. expressed the PrPC protein. M.G. performed CD, fluorescence, MD and SAXS experiments. I.Z. performed NMR experiments. M.G., S.K.T.S.W., Ž.S., A.G. and M.K. analysed the CD and fluorescence data. I.Z., S.K.T.S.W. and A.G. analysed the NMR data. M.G., Ž.S. and I.Z. analysed the MD data. M.G., M.T. and M.K. analysed the SAXS data. M.K., W.M.K. and I.Z. conceptualized the study. M.G., I.Z., Ž.S. and M.K. designed the study. M.G. prepared the initial draft. M.G., S.K.T.S.W., I.Z., A.G. and M.K. reviewed and edited the manuscript. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Gielnik, M., Taube, M., Zhukova, L. et al. Zn(II) binding causes interdomain changes in the structure and flexibility of the human prion protein. Sci Rep 11, 21703 (2021). https://doi.org/10.1038/s41598-021-00495-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-00495-0
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.