Abstract
The role of the gut microbiome is increasingly being recognized by health scientists and veterinarians, yet its role in wild animals remains understudied. Variations in the gut microbiome could be the result of differential diets among individuals, such as variation between sexes, across seasons, or across reproductive stages. We evaluated the hypothesis that diet alters the avian gut microbiome using stable isotope analysis (SIA) and 16S rRNA gene sequencing. We present the first description of the thick-billed murre (Uria lomvia) fecal microbiome. The murre microbiome was dominated by bacteria from the genus Catellicoccus, ubiquitous in the guts of many seabirds. Microbiome variation was explained by murre diet in terms of proportion of littoral carbon, trophic position, and sulfur isotopes, especially for the classes Actinobacteria, Bacilli, Bacteroidia, Clostridia, Alphaproteobacteria, and Gammaproteobacteria. We also observed differences in the abundance of bacterial genera such as Catellicoccus and Cetobacterium between sexes and reproductive stages. These results are in accordance with behavioural observations of changes in diet between sexes and across the reproductive season. We concluded that the observed variation in the gut microbiome may be caused by individual prey specialization and may also be reinforced by sexual and reproductive stage differences in diet.
Similar content being viewed by others
Introduction
The role of bacteria in higher trophic level predator interactions is increasingly being appreciated1,2. The gut microbiome may be crucial for the adaptation of the vertebrate host to ecological pressures, such as rapid environmental change or adverse conditions3,4,5,6. Bacterial communities play an important role in processing food, and matching microbiome with diet may be one mechanism for animals to overcome changes in their diet throughout their life cycle7,8. Variation in diet composition was linked to changes in the gut microbiome in wild sticklebacks and perches, two fish species with individual prey specialization (IPS)7,8. IPS is a widely distributed phenomenon, occurring in a large number of taxa where individuals of the same population use prey resources differentially9,10. In particular, individuality in prey selection is not explained by obvious differences, such as by sex or age9,10. These studies also show that other inter-individual variables can change the feeding behaviour (e.g. differential diets between sexes) leading to variation in the microbiome6,8.
Most studies describing diet effects on the avian gut microbiome are based on domestic or captive birds such as chickens11,12,13 or raptors14. These studies have mainly focused on the detection of specific groups or species of bacteria (mostly pathogens)15,16,17. Recently, gut microbial communities for a few wild bird species have been described18. Maul and collaborators19 characterized cloacal bacteria according to carbon substrate utilization in culture plates (EcoPlates) and clustered these bacterial communities of various passerine bird species according to bird diet. The gut microbiome of two passerine species differed between spring and fall migrations6 and between migrating and resident shorebirds20, suggesting that the environment, and possibly diet, has a strong influence on wild bird microbiomes. Dewar et al.21 reported individual variation in the gastrointestinal microbiota of four penguin species. One meta-analysis illustrated that, even among phylogenetically and behaviourally diverse species, diet has an important effect on gut microbiome composition22. Lower microbial diversity of house sparrow guts from urbanized areas compared to rural sparrows shed some light to the way in which microbiomes can adapt to reduced niches (a city, compared to the countryside), but may also lose plasticity23. Hird and collaborators24 studied the intestinal microbiome of 59 Neotropical bird species and determined that host species was the most important determinant for the intestinal microbial composition, followed by host ecology (i.e. broad dietary preferences and habitat)24. They also proposed the inclusion of gut sampling as part of the field sampling protocol for collection of museum specimens as it could provide valuable information to the “microdiversity” within wild macroorganisms24. Similar results were obtained for the gut microbiomes of 74 Equatorial Guinean bird species in terms of the importance of host species, diet, and location as determinants of the microbial composition25. The gut microbiomes of Darwin’s finches were shown to be conserved across species with the exception of the blood-feeding vampire finch, showing that extreme diet specialization can change microbial gut composition26.
Thick-billed murres (Uria lomvia; hereafter ‘murres’) are the most abundant seabirds in the Canadian Arctic. In the low Arctic, murres are generalists, with the most diverse diet of any Uria population. However, despite being population-wide generalists, there is a high degree of IPS that is maintained across time with some individuals disproportionately catching rare prey types year after year27,28. Sex-specific feeding behaviours are also present as males tend to feed on amphipods and shallow benthic prey at night, while females tend to feed on deep benthic and schooling prey only available during the day29,30. In addition, females feed at a higher trophic level when rearing chicks compared to when they are incubating their egg, but males do not29.
In this study we used 16S rRNA gene sequencing of fecal samples as a proxy to describe for the first time the gut microbiome of the thick-billed murre. We tested the hypothesis that diet alters gut microbiome. For this, we used stable isotope ratios as direct estimators of diet. As 15N content in an organism increases systematically as trophic level increases, the nitrogen isotope ratio (δ15N) is useful for describing trophic position31. The carbon isotope ratio (δ13C), also referred to as the proportional use of littoral carbon8, can help to determine diet according to the habitat animals feed on as, for example, there is a systematic enrichment of 13C in benthic feeding organisms, relative to pelagic feeding organisms32. Sulfur stable isotopes present can also be used to determine diet through changes in habitat as the most-enriched δ34S values are found in the benthos of marine systems33. We also examined variation in the murre gut microbiome between sex and reproductive stage (incubation vs. chick-rearing), that would be consistent with known variations in murre diet between sexes and reproductive stages29.
Results
The composition of the murre fecal microbiome is dominated by the genus Catellicoccus
After denoising and filtering, a total of 7,232,617 16S rRNA gene reads with an average of 76,942 ± 4960 (mean ± SE; n = 94) reads per sample were obtained. We observed a large degree of inter-individual variation in the composition of the gut microbiome within the studied individuals (Fig. 1). Firmicutes was the predominant phylum and ranged from over 11.9% to over 99.8% of the microbiome of the sampled murres with Catellicoccus being the dominant genus detected and comprising 1.6% to 98.8% of an individual’s gut microbiota (Supplementary Figs. 1 and 2). Other major phyla in the murre gut microbiome include Fusobacteria, Proteobacteria, and Actinobacteria. There is a strong influence of some amplicon sequence variants (ASVs) on the composition of the murre gut microbiome (Fig. 2). The most important features determining the weighted and unweighted UniFrac distance matrix showed the influence of ASVs from the genera Catellicoccus, Breznakia, Cetobacterium, Escherichia/Shigella (these two genera cannot not be easily distinguished solely based on 16S rRNA gene sequences), and Campylobacter on the overall community composition of the murre fecal microbiome (Fig. 2 and Supplementary Fig. 3).
Principal Coordinates Analysis (PCoA) plot for the gut community composition using the unweighted UniFrac metric with samples identified by sex and reproductive stage. Biplots show the taxonomy of the ASVs with the top 5 effects on the community composition with position of the arrow indicating the direction of the effect.
Diet variation measured between sexes and reproductive stages is associated with individual ASV relative abundances
Females fed at a higher trophic position and had larger δ34S values compared to males (Table 1). Individuals of both sexes fed at a higher trophic position during the chick-rearing stage than during the incubation stage (Table 1). Quasibinomial generalized linear models (GLMs) showed that 20.8% of the relative abundances of the most common (> 0.01% relative abundance) ASVs are correlated with a combination of the proportional use of littoral carbon and the quadratic effect of δ34S (Table 2). A χ2 test showed that this proportion statistically differs from chance with the expected 5% false detection rate (FDR) expected from performing multiple comparisons. For this group of models, ASV abundance tended to be mostly positively associated with the proportional use of littoral carbon and negatively associated with the quadratic effect of δ34S (Fig. 3). After performing an FDR correction, 19.6% of the models showed a correlation. Trophic position and δ34S also correlated with ASV relative abundance at a larger proportion than the 5% FDR (Fig. 3; Table 2). After the FDR correction, 1.2% of the ASVs were correlated with the proportional use of littoral carbon, 1.4% were correlated with trophic position, and 0.5% with δ34S.
Quasibinomial effects of various diet metrics on the abundance of the 424 most abundant ASVs of the murre fecal microbiome. Columns contain the models of a diet metric for ASVs from a bacterial class in each row. Vertical bars represent an ASV within the given class that accounts for > 0.01% mean relative abundance and that has a positive or negative association with a particular metric. For ASVs with a red bar, its relative abundance increases with the metric. For ASVs with a blue bar, relative abundance decreased with the metric. Numbers under the different diet metrics indicate the number of ASVs for which an effect of the metric was observed. Numbers with an asterisk (*) indicate that the proportion of ASVs with an effect of the metric surpasses the expected 5% false positive rate. Metrics with a double dagger (‡) form part of the same model which combined the linear effect of the proportion of littoral carbon and the quadratic effect of δ34S.
Microbial community metrics differ between reproductive stages and sexes
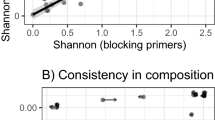
No differences in Faith’s phylogenetic diversity (Faith’s PD) were found between sexes (Fig. 4a; Table 3). Incubating birds had a more phylogenetically diverse microbiome than chick rearing birds (Fig. 4b; Table 3). No differences were observed between sexes or reproductive stages in terms of Shannon’s diversity index (Fig. 4c,d; Table 3). Murres appeared to have different microbial communities between reproductive stages in terms of qualitative community composition (Fig. 2a; Supplementary Fig. 4a): unweighted UniFrac distances (PERMANOVA: pseudo-F = 1.99, R2 = 0.042, P = 0.007 ). No differences for unweighted UniFrac distances were observed between sexes. It must be noted that the multivariate dispersions are not homogeneous between incubating and chick-rearing birds (PERMDISP: F = 9.89, P = 0.015) with greater dispersion among incubating birds than among chick-rearing birds. There were no statistical differences between the dispersions of males and females. No differences were observed between sexes or reproductive stages for the weighted UniFrac distances (Supplementary Figs. 3 and 4b). We also observed different abundances of individual ASVs between sexes as 34 ASVs were more abundant in males than in females (Fig. 5a; Supplementary Table 1). There were 13 ASVs which were more abundant for birds during the chick-rearing stage and 24 ASVs that were more abundant for birds during the incubation stage (Fig. 5b; Supplementary Table 1).
Differences in Faith’s phylogenetic diversity for the gut microbiome with no differences between sexes (a) but evidencing differences between reproductive stages (b) and showing no differences in terms of Shannon’s Diversity Index (c) between sexes or (d) between reproductive stages. Letters above boxplots represent differences between groups with p < 0.05.
Discussion
As expected, diet—as measured by stable isotope analysis (SIA)—was associated with variation in gut microbiome. The association was linear for trophic position and carbon source but was non-linear for sulfur source. We observed large amounts of individual-level variation that could be caused by diet specialization observed previously for this species27,28,29. Moreover, microbiome changed with sex and reproductive stage, which is also consistent with known dietary differences.
The gut microbiome of the thick-billed murre is consistent with other avian microbiomes, which are mainly comprised of the phyla Firmicutes, Actinobacteria, Bacteroidetes, and Proteobacteria18. Even though ethanol preservation can lead to changes in the relative abundance of some bacterial groups, these changes are smaller or comparable to observed differences among technical replicates34. Because of this, we do not consider that the patterns in our study showing the dominance of Firmicutes to be an artifact of the preservation method. However, we did observe that there was only a small proportion of Bacteroidetes in our samples compared to other studies18 (Fig. 1). Bacteroidetes have been associated with degradation of complex biopolymers, such as cellulose, in mammal guts and it has been suggested that this association may also exist for birds35. Given that plant material is not a regular part of the murre diet, this could explain the low abundance of Bacteroidetes in our study.
The murre gut microbiome was dominated by bacteria belonging to the genus Catellicoccus (phylum Firmicutes). This genus was first described with isolates from harbour porpoise (Phocoena phocoena) and grey seal (Halichoerius grypus) carcasses36 and, since then, have been found to be ubiquitous in the gut microbiome of various avian species. Catellicoccus marimammalium is used as a marker for detection of gull fecal contamination37,38,39,40. Bacteria from the genus Catellicoccus have also been found in the gastrointestinal tract of zebra finches (Taeniopygia guttata)41; barn swallows (Hirundo rustica)42; red knots (Calidris canutus) and ruddy turnstones (Arenaria interpres)43; black-tailed godwits (Limosa limosa), black-winged stilts (Himantopus himantopus) and common redshanks (Tringa totanus)44; black-headed gulls (Chroicocephalus ridibundus)45 and waterfowl46. The genome of Catellicoccus marimammalium revealed that this bacterium encodes various functions such as nutrient transport and bile acid hydrolysis suggesting a symbiotic lifestyle of this species46. Possible beneficial effects for the host such as immune modulation and gut maturation have also been proposed for bacteria of this genus inhabiting the avian gut41. Additionally, members of the Firmicutes are widely involved in the production of short-chain fatty acids which can be absorbed as an energy source by the host gut wall35. Firmicutes have also been associated with weight gain, increase of nutrient uptake and metabolic efficiency in chickens35. It is then possible to hypothesize that Catellicoccus detected in the murre gut are aiding these birds to optimize their nutrition as the murres face the harsh conditions of the Arctic and the limited feeding events during the reproductive season.
High abundances of bacteria from the phylum Fusobacteria have also been observed in the guts of other seabirds such as gentoo and king penguins (Pygoscelis papua and Aptenodytes patagonicus, respectively)21,47, common diving petrels (Pelecanoides urinatrix)48, gulls40, vultures and carnivorous mammals18,49, and humans50. Although bacteria from this phylum are known to be pathogenic, recently it has been observed that they could aid their host to metabolize nutrients18,49. Fusobacteria are known butyrate producers and can ferment amino acids and glucose47,48,50 and, in chickens, they boost the host immune system and adiposity48. The most influential genus observed for this phylum that shaped the overall community composition (Fig. 2) was Cetobacterium. Cetobacterium someare isolated from the intestinal tract of freshwater fish produce vitamin B12 and acetic acid which suggest the beneficial effects of this bacterium for its host51.
The biplot analysis for the UniFrac PCoA plots (Fig. 2 and Supplementary Fig. 3) showed that one of the most influential ASVs shaping the microbial community structure belonged to the genus Breznakia. This genus is relatively novel and not much is known about these bacteria, but this genus and others in the family Erysipelotrichaceae have been frequently isolated from the guts of mammals and insects, making this family mostly comprised of inhabitants of animal intestinal tracts52. Many bacterial genera associated with opportunistic pathogens, such as Campylobacter, Helicobacter, Escherichia/Shigella, Corynebacterium, Mycobacterium, Neisseria and Ornithobacterium, among others, were found in the murre gut. However, no risk of disease is believed to exist solely based of the presence of these genera as many of them contain representatives that have also been found in other healthy birds21,43,44,47,53. The presence of these taxa should continue to be monitored given that reverse zoonosis of various Campylobacter strains associated with humans occurs in Antarctic seabirds54.
The gut microbiome of males and females are different in terms of qualitative community composition (Fig. 2) and abundances of individual ASVs (Fig. 5a). The SIA also showed that females are feeding at a higher trophic position than males, suggesting that trophic position influences the fecal microbiome of murres. These results are consistent with the social behaviour of murre mating pairs at Coats Island in which males stay in the colony during the day and leave their nest to feed at night while females feed during the day and go back to the colony during the night29. This leads to different types of prey being caught by each sex, with males preferring amphipods and shallow diving fish and females preferring deep water benthic prey29. These observations are also consistent with the differences we detected in terms of δ34S as females had a larger average value for this isotopic signature, which suggests that they are feeding at higher depths than males. Additionally, risk-prone diet for females (i.e. schooling fish)29 could provide less consistent food sources which helps to explain why there was a smaller phylogenetic diversity for females and that some key genera, such as Cetobacterium, were more abundant in males.
We detected differences in phylogenetic diversity (Fig. 4b) and in qualitative composition of bacterial communities between reproductive stages (Fig. 2). We also observed individual ASVs that changed in abundance between the chick-rearing and incubating birds (Fig. 5b). This may occur because, for example, chick-rearing females feed at higher trophic levels than incubating females, as chick-rearing females must locate fish to bring back and feed their offspring29. SIA confirmed that chick-rearing murres were in fact feeding at a higher tropic level than those sampled during the incubation stage. Diet variation caused by changes in prey composition throughout the breeding season could potentially cause the differences in the gut microbiome that we observe. As prey from higher trophic levels, such as fish, tend to be encountered less frequently29, dietary intake can become less consistent which could in turn modify the gut microbiome. Despite the fact that we detected heterogeneity of multivariate dispersions between reproductive stages for the unweighted UniFrac distances, PERMANOVA results have been observed to become too conservative for unbalanced designs when there is a greater dispersion for the largest group, as is the present case55. This suggests that the observed differences might in fact be larger than those detected by the PERMANOVA analysis between reproductive stages. Additionally, the lack of observed differences for the weighted UniFrac distances between sexes and reproductive stages suggest that the differences that we observe are caused due to differences in microbial richness, rather than in abundance in terms of the overall community composition.
However, we did observe some differences in individual ASVs’ abundance that cannot be explained by trophic position, sex, or reproductive stage but that can still be attributed to diet. The abundances of a large proportion of the most abundant ASVs were associated with the proportional use of littoral carbon and the quadratic effect of sulfate availability (δ34S). A differential use of littoral carbon suggests that an individual’s feeding preference on benthic or littoral food sources changes the abundance of certain groups of bacteria. δ34S is an isotope signature that has been used to differentiate between coastal and marine feeding birds56 and between sources of sulfate originated from the surface and the sediments33. Given that we observed both positive and negative quadratic effect of this isotope ratio, certain microbes appear to benefit from a more generalized diet and that some others benefit more from more specialized diets. Given the larger proportion of GLMs including the source of the sulfate and the use of littoral carbon compared to the GLMs using trophic position, the source of the food in terms of depth is likely a more important factor in shaping the microbiome than the trophic position of the prey that the birds feed on. These results support the behavioural observations of IPS at the murre colony in Coats Island28.
We conclude that the changing diet of the thick-billed murre between sexes and through the reproductive stage is linked to variation of the gut microbial communities. However, the correlational associations observed in the present study cannot be used to establish a causal effect of dietary preferences shaping the gut microbiome. There is an additional need for studies of wild animal microbiomes given that the behaviours and environmental conditions that might be encountered in nature could result in changes in the gut microbiome that might not be observable under laboratory conditions. For example, our study design was biased towards males (because they are more easily sampled at the colony), and future studies could focus on species that can be sexed morphologically allowing direct incorporation of sex into study design and capture protocol. We also corroborate the need to characterize diet for particular individuals instead of generalizing diets of populations as a whole9,10. Future studies could also focus on linking the trends in the composition of the microbiome that we describe with the fitness of individuals to determine how certain groups of bacteria are positively or negatively affecting their host birds.
Methods
Fecal sample collection and preparation
Methodologies were approved by the Macdonald Campus Facility Animal Care Committee under Animal Use Protocol number 2015–7599 and adhered to all regulations and guidelines. Thick-billed murres (N = 48; 9 females, 31 males, and 8 with undetermined sex) were captured with a noose pole and sampled between July and August 2017 at a breeding colony at Coats Island (62°98′N, 82°00′W) located in northern Hudson Bay, Nunavut, Canada. Fecal samples were aseptically taken by inserting sterile swabs into the cloaca. The swab was swirled inside the cloaca to stimulate the release of feces. The recovered fecal matter was collected in sterile polypropylene tubes which were kept on ice until they were taken back to the base camp (less than an hour from the moment of sampling). Samples were then mixed with absolute ethanol (4:1 ethanol to feces ratio) and stored at − 20 °C until processed (between 2 and 3 months after they were frozen). This process was done at two different time points with 2–7 days in between each sampling event (Supplementary Table 2)57. Ethanol was removed from the samples by centrifugation: polypropylene tubes containing the samples were centrifuged at 4500 RPM and 4 °C for 5 min and the supernatant was discarded. The pellet was resuspended in 2 mL of sterile deionized water, centrifuged at 4500 RPM and 4 °C for 5 min and the supernatant was discarded once more to assure any remaining ethanol was removed from the collected feces.
DNA extraction and sequencing
Due to the high content of contaminants present in avian feces (e.g. uric acid) which might inhibit DNA extractions58, we modified the protocols of commercially used DNA extraction and purification kits to improve the quality of the obtained nucleic acids. DNA was extracted from the resulting pellets of the fecal samples using the DNeasy PowerLyzer PowerSoil kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions with minor modifications: samples were heated at 65 °C for 10 min before bead beating, and, for the final elution step, nuclease-free water was heated to 70 °C and used to elute the DNA. Extracted DNA was purified using Monarch PCR & DNA Cleanup Kit (New England Biolabs, Ipswitch, MA) according to the manufacturer’s instructions with minor modifications: 20 µL of nuclease-free water heated to 70 °C were used to elute the DNA.
The 16S rRNA gene was amplified using primers 515F-Y (5′-GTGYCAGCMGCCGCGGTAA) and 926R (5′-CCGYCAATTYMTTTRAGTTT)59 containing Illumina overhang adapter sequences. These primers were selected as they have been shown to be more accurate and produce longer amplicons than the 515F-C/806R primer pair, helping to differentiate previously unresolvable taxa59. 25 µL PCR reactions containing 1X HotStarTaq Master Mix (Qiagen, Hilden, Germany), 0.6 µM of each primer, 0.4 mg mL−1 BSA (Sigma-Aldrich, St. Louis, MO), and 1 µL of DNA were performed under the following conditions: initial denaturation at 95 °C for 15 min, followed by 25 to 35 cycles (depending on the DNA concentration of the sample) of 94 °C for 1 min, 50 °C for 45 s, 72 °C for 1 min, and a final extension step at 72 °C for 10 min. Reactions were purified using Agencourt AMPure XP magnetic beads (0.8 bead-to-PCR volume ratio; Beckman Coulter, USA). Indexing was performed using the Nextera XT index kit (Illumina, San Diego, CA) following manufacturer’s instructions with a minor modification of 15 min at 95 °C in the initial denaturation used to activate the polymerase (Qiagen HotStarTaq Master Mix). Indexed samples were purified with AMPure XP beads (1.12 bead-to-PCR volume ratio) and quantified using the Qubit fluorometer (Invitrogen, Thermo Fisher Scientific, USA). Samples (including negative controls of PCR reactions) were pooled in equimolar ratios of 4 nM and sequenced with a 2 × 250 bp run with v2 chemistry on a MiSeq platform (Illumina, San Diego, CA). Adapters and indices were removed by the Illumina FASTQ file generation pipeline.
Sequencing data processing
Unless stated otherwise, all analyses were performed in R 3.6.1. Chimeras were removed from the raw sequencing data and a preliminary taxonomic assignment was undertaken with the Silva database60 version 132 release and using DADA261. Sequencing data was then analyzed using Qiime 262 (release 2020.8). Based on the preliminary classification, ASVs predominantly present in the negative controls (50% prevalence threshold) and also present in the fecal sample data were removed (n = 114), samples with fewer than 1000 sequences (n = 1), singletons (n = 70), and the negative controls (Supplementary Figure 7) were removed from further analysis. A Taxonomic classification was performed with the q2-feature-classifier multinomial naive Bayes classifier63 using the SSU Ref NR 99% Silva database60 version 132 release. ASVs that were not assigned a taxonomic classification at the domain level (n = 33), and mitochondrial and chloroplastic ASVs (n = 34), were removed. A phylogenetic tree was inferred using the approximately maximum-likelihood method implemented by FastTree 264.
Stable isotope analysis (SIA)
Blood was taken from the brachial vein using heparinized syringes. Red blood cells (RBCs) were separated from plasma by centrifugation and they were frozen and transported in gaseous nitrogen to the processing facility where they were stored at − 80 °C until processed. We followed the methodology previously used to perform SIA for murre blood samples from the same colony2. We freeze-dried and powdered the RBCs and lipids were removed from the powder using a 2:1 chloroform:methanol soak and rinse65. Stable isotope analyses for nitrogen (trophic position), carbon (proportional use of littoral carbon), and sulfur (feeding habitat depth) stable isotopes were performed by the G.G Hatch Stable Isotope Laboratory (Ottawa, ON) for 43 of the sampled birds that where also part of the Northern Contaminants Project of Aboriginal Affairs and Northern Development Canada. 1 mg subsamples were used for stable nitrogen and carbon isotope assays. All measurements are reported in parts per thousand (‰) relative to the AIR international standard. For every 10 samples, replicate measurements of internal laboratory standards calibrated to international standards were also run indicating a measurement error of ± 0.2‰. 10 mg subsamples were used to measure stable sulfur isotope ratios. All measurements are reported in parts per thousand (‰) relative to the VDCT international standard. Data was normalized with a precision of ± 0.4‰ using calibrated internal standards. Sample duplicates for every 10 samples were used for quality control to assure that the relative percent difference is less than 5%.
We used the δ15N and δ13C values obtained from the murre blood samples along with those of pelagic and littoral prey items commonly captured by murres2 in order to determine proportional use of littoral carbon and trophic position8 in R 3.6.1.
Statistical analyses
Alpha level for all tests was 0.05. Unless stated otherwise, all analyses were performed in R 3.6.1. ASV abundances were converted into relative abundances for sequencing data normalization66. Repeatability was evaluated for birds where two samples were obtained using the q2-longitudinal plugin67 to compare differences in terms of alpha diversity with the Wilcoxon signed-rank test68. We also evaluated repeatability for the community composition between the two time points with a PERMANOVA test69 for unweighted and weighted UniFrac distances70. These repeatability analyses showed that samples from the first time point were not statistically different from samples for the second time point (Supplementary Figs. 5 and 6). Given these results, only the sequencing data from the first sampling point will be analyzed to account for the fact that the two sample points are not statistically independent.
Diversity analyses were performed with the q2-diversity plugin. Faith’s PD71 and Shannon’s diversity index72 were calculated for each sex and reproductive stage (incubating vs. chick-rearing). Kruskal–Wallis rank tests73 were used to observe statistical differences in Faith’s PD and Shannon’s diversity index between these groups. Unweighted and weighted UniFrac distances70 were calculated to observe the community structure of the murre gut microbiome. After testing for the homogeneity of multivariate dispersions (PERMDISP)74, differences in community structure for the two distance measurements between sexes and between reproductive stages were calculated using the permutational multivariate analysis of variance (PERMANOVA)69. Principal Coordinates Analysis (PCoA) plots and biplots75 were created with Emperor76,77 to visualize the differences obtained with PERMANOVA. Additionally, NMDS plots were created with phyloseq78 to provide a different visualization of the community composition.
We used DESeq279,80 to observe differences in ASV abundance between sexes and reproductive stages. Given that this approach already corrects for differences in sample counts80, the differential expression analysis using a negative binomial distribution was applied on pre-normalisation by proportions data set as implemented by the DESeq function of the DESeq2 package79.
We tested differences of trophic position between sexes using Student’s t-test and we tested differences of isotopic signatures between sexes and reproductive stages with a Mann–Whitney test81, given the non-parametric nature of the isotope data for these groups. We also observed the effect that isotopic signatures had on the relative abundance the ASVs in our samples following a similar strategy as developed by Bolnick and collaborators7,8. We selected the most abundant ASVs that had a mean relative abundance of > 0.01% obtaining a total of 926 ASVs. We then used GLMs with a quasibinomial error created with the MuMIn package82 to evaluate the dependency of the abundance of individual ASVs on proportional use of littoral carbon, trophic position, and δ34S. Additionally, we evaluated if there were non-linear effects on ASV abundance using quadratic terms for the three isotopic signatures. To account for false positives, we used a chi-squared test to identify whether the number of GLMs with p < 0.05 exceeded the 5% null expectation. We then applied an FDR analysis to obtain the number of models for which q < 0.05.
Data availability
The sequencing data was deposited in Sequence Read Archive of NCBI under accession number PRJNA594033. Isotope data and sex data is included as the Supplementary Table 2 and it also available in the Mendeley Data repository, http://dx.doi.org/10.17632/r69xp7xsky.357. Detailed parameters for the bioinformatic and statistical analyses used for the study is available in the GitHub repository, https://github.com/estebangongora/murre-microbiome.
References
Kuwae, T. et al. Biofilm grazing in a higher vertebrate: The Western Sandpiper, Calidris mauri. Ecology 89, 599–606 (2008).
Góngora, E., Braune, B. M. & Elliott, K. H. Nitrogen and sulfur isotopes predict variation in mercury levels in Arctic seabird prey. Mar. Pollut. Bull. 135, 907–914 (2018).
Ben-Yosef, M., Aharon, Y., Jurkevitch, E. & Yuval, B. Give us the tools and we will do the job: Symbiotic bacteria affect olive fly fitness in a diet-dependent fashion. Proc. R. Soc. B Biol. Sci. 277, 1545–1552 (2010).
Alberdi, A., Aizpurua, O., Bohmann, K., Zepeda-Mendoza, M. L. & Gilbert, M. T. P. Do vertebrate gut metagenomes confer rapid ecological adaptation?. Trends Ecol. Evol. 31, 689–699 (2016).
Lapanje, A., Zrimec, A., Drobne, D. & Rupnik, M. Long-term Hg pollution-induced structural shifts of bacterial community in the terrestrial isopod (Porcellio scaber) gut. Environ. Pollut. 158, 3186–3193 (2010).
Lewis, W. B., Moore, F. R. & Wang, S. Characterization of the gut microbiota of migratory passerines during stopover along the northern coast of the Gulf of Mexico. J. Avian Biol. 47, 659–668 (2016).
Bolnick, D. I. et al. Individuals’ diet diversity influences gut microbial diversity in two freshwater fish (threespine stickleback and Eurasian perch). Ecol. Lett. 17, 979–987 (2014).
Bolnick, D. I. et al. Individual diet has sex-dependent effects on vertebrate gut microbiota. Nat. Commun. 5, 4500 (2014).
Bolnick, D. I., Yang, L. H., Fordyce, J. A., Davis, J. M. & Svanbäck, R. Measuring individual-level resource specialization. Ecology 83, 2936–2941 (2002).
Bolnick, D. I. et al. The ecology of individuals: Incidence and implications of individual specialization. Am. Nat. 161, 1–28 (2003).
Apajalahti, J. H. A., Kettunen, A., Bedford, M. R. & Holben, W. E. Percent G + C profiling accurately reveals diet-related differences in the gastrointestinal microbial community of broiler chickens. Appl. Environ. Microbiol. 67, 5656–5667 (2001).
Apajalahti, J. & Kettunen, A. Microbes of the chicken gastrointestinal tract. In Avian Gut Function in Health and Disease (ed. Perry, G. C.) 124–137 (CAB International, Wallingford, 2006).
Oakley, B. B. et al. The chicken gastrointestinal microbiome. FEMS Microbiol. Lett. 360, 100–112 (2014).
Bangert, R. L., Ward, A. C. S., Stauber, E. H., Cho, B. R. & Widders, P. R. A survey of the aerobic bacteria in the feces of captive raptors. Avian Dis. 32, 53–62 (1988).
Soucek, Z. & Mushin, R. Gastrointestinal bacteria of certain Antarctic birds and mammals. Appl. Microbiol. 20, 561–566 (1970).
Mead, G. C., Griffiths, N. M., Impey, C. S. & Coplestone, J. C. Influence of diet on the intestinal microflora and meat flavour of intensively-reared broiler chickens. Br. Poult. Sci. 24, 261–272 (1983).
Waldenström, J. et al. Prevalence of Campylobacter jejuni, Campylobacter lari, and Campylobacter coli in different ecological guilds and taxa of migrating birds. Appl. Environ. Microbiol. 68, 5911–5917 (2002).
Waite, D. W. & Taylor, M. W. Exploring the avian gut microbiota: Current trends and future directions. Front. Microbiol. 6, 1–12 (2015).
Maul, J. D., Gandhi, J. P. & Farris, J. L. Community-level physiological profiles of cloacal microbes in songbirds (order: Passeriformes): Variation due to host species, host diet, and habitat. Microb. Ecol. 50, 19–28 (2005).
Risely, A., Waite, D. W., Ujvari, B., Hoye, B. J. & Klaassen, M. Active migration is associated with specific and consistent changes to gut microbiota in Calidris shorebirds. J. Anim. Ecol. 87, 428–437 (2018).
Dewar, M. L. et al. Interspecific variations in the gastrointestinal microbiota in penguins. Microbiologyopen 2, 195–204 (2013).
Waite, D. W. & Taylor, M. W. Characterizing the avian gut microbiota: Membership, driving influences, and potential function. Front. Microbiol. 5, 1–12 (2014).
Teyssier, A. et al. Inside the guts of the city: Urban-induced alterations of the gut microbiota in a wild passerine. Sci. Total Environ. 612, 1276–1286 (2018).
Hird, S. M., Sánchez, C., Carstens, B. C. & Brumfield, R. T. Comparative gut microbiota of 59 neotropical bird species. Front. Microbiol. 6, 1403 (2015).
Capunitan, D. C., Johnson, O., Terrill, R. S. & Hird, S. M. Evolutionary signal in the gut microbiomes of 74 bird species from Equatorial Guinea. Mol. Ecol. 29, 829–847 (2020).
Michel, A. J. et al. The gut of the finch: Uniqueness of the gut microbiome of the Galápagos vampire finch. Microbiome 6, 167 (2018).
Elliott, K. H., Woo, K. J. & Gaston, A. J. Specialization in murres: The story of eight specialists. Waterbirds 32, 491–506 (2009).
Woo, K. J., Elliott, K. H., Davidson, M., Gaston, A. J. & Davoren, G. K. Individual specialization in diet by a generalist marine predator reflects specialization in foraging behaviour. J. Anim. Ecol. 77, 1082–1091 (2008).
Elliott, K. H., Gaston, A. J. & Crump, D. Sex-specific behavior by a monomorphic seabird represents risk partitioning. Behav. Ecol. 21, 1024–1032 (2010).
Paredes, R., Jones, I. & Boness, D. Parental roles of male and female thick-billed murres and razorbills at the Gannet Islands, Labrador. Behaviour 143, 451–481 (2006).
Atwell, L., Hobson, K. A. & Welch, H. E. Biomagnification and bioaccumulation of mercury in an arctic marine food web: Insights from stable nitrogen isotope analysis. Can. J. Fish. Aquat. Sci. 55, 1114–1121 (1998).
Carr, M. K. et al. Stable sulfur isotopes identify habitat-specific foraging and mercury exposure in a highly mobile fish community. Sci. Total Environ. 586, 338–346 (2017).
Peterson, B. J. & Fry, B. Stable isotopes in ecosystem studies. Annu. Rev. Ecol. Syst. 18, 293–320 (1987).
Song, S. J. et al. Preservation methods differ in fecal microbiome stability, affecting suitability for field studies. mSystems 1, e00021-e116 (2016).
Grond, K., Sandercock, B. K., Jumpponen, A. & Zeglin, L. H. The avian gut microbiota: Community, physiology and function in wild birds. J. Avian Biol. 49, e01788 (2018).
Lawson, P. A., Collins, M. D., Falsen, E. & Foster, G. Catellicoccus marimammalium gen. nov., sp. nov., a novel Gram-positive, catalase-negative, coccus-shaped bacterium from porpoise and grey seal. Int. J. Syst. Evol. Microbiol. 56, 429–432 (2006).
Sinigalliano, C. D. et al. Multi-laboratory evaluations of the performance of Catellicoccus marimammalium PCR assays developed to target gull fecal sources. Water Res. 47, 6883–6896 (2013).
Ryu, H. et al. Comparison of gull feces-specific assays targeting the 16S rRNA genes of Catellicoccus marimammalium and Streptococcus spp. Appl. Environ. Microbiol. 78, 1909–1916 (2012).
Koskey, A. M., Fisher, J. C., Traudt, M. F., Newton, R. J. & McLellan, S. L. Analysis of the gull fecal microbial community reveals the dominance of Catellicoccus marimammalium in relation to culturable enterococci. Appl. Environ. Microbiol. 80, 757–765 (2014).
Lu, J., Santo Domingo, J. W., Lamendella, R., Edge, T. & Hill, S. Phylogenetic diversity and molecular detection of bacteria in gull feces. Appl. Environ. Microbiol. 74, 3969–3976 (2008).
Benskin, C. M. H., Rhodes, G., Pickup, R. W., Wilson, K. & Hartley, I. R. Diversity and temporal stability of bacterial communities in a model passerine bird, the zebra finch. Mol. Ecol. 19, 5531–5544 (2010).
Kreisinger, J. et al. Temporal stability and the effect of transgenerational transfer on fecal microbiota structure in a long distance migratory bird. Front. Microbiol. 8, 1–19 (2017).
Grond, K., Ryu, H., Baker, A. J., Santo Domingo, J. W. & Buehler, D. M. Gastro-intestinal microbiota of two migratory shorebird species during spring migration staging in Delaware Bay, USA. J. Ornithol. 155, 969–977 (2014).
Santos, S. S. et al. Diversity of cloacal microbial community in migratory shorebirds that use the Tagus estuary as stopover habitat and their potential to harbor and disperse pathogenic microorganisms. FEMS Microbiol. Ecol. 82, 63–74 (2012).
Laviad-Shitrit, S., Izhaki, I., Lalzar, M. & Halpern, M. Comparative analysis of intestine microbiota of four wild waterbird species. Front. Microbiol. 10, 1–13 (2019).
Weigand, M. R., Ryu, H., Bozcek, L., Konstantinidis, K. T. & Santo Domingo, J. W. Draft genome sequence of Catellicoccus marimammalium, a novel species commonly found in gull feces. Genome Announc. 1, 12–13 (2013).
Dewar, M. L. et al. Influence of fasting during moult on the faecal microbiota of penguins. PLoS ONE 9, e99996 (2014).
Dewar, M. L., Arnould, J. P. Y., Krause, L., Dann, P. & Smith, S. C. Interspecific variations in the faecal microbiota of Procellariiform seabirds. FEMS Microbiol. Ecol. 89, 47–55 (2014).
Roggenbuck, M. et al. The microbiome of New World vultures. Nat. Commun. 5, 5498 (2014).
Potrykus, J., White, R. L. & Bearne, S. L. Proteomic investigation of amino acid catabolism in the indigenous gut anaerobe Fusobacterium varium. Proteomics 8, 2691–2703 (2008).
Tsuchiya, C., Sakata, T. & Sugita, H. Novel ecological niche of Cetobacterium somerae, an anaerobic bacterium in the intestinal tracts of freshwater fish. Lett. Appl. Microbiol. 46, 071018031740002–000 (2007).
Tegtmeier, D., Riese, C., Geissinger, O., Radek, R. & Brune, A. Breznakia blatticola gen. nov. sp. nov. and Breznakia pachnodae sp. nov., two fermenting bacteria isolated from insect guts, and emended description of the family Erysipelotrichaceae. Syst. Appl. Microbiol. 39, 319–329 (2016).
Vandamme, P. et al. Ornithobacterium rhinotracheale gen. nov., sp. nov. isolated from the avian respiratory tract. Int. J. Syst. Bacteriol. 44, 24–37 (1994).
Cerdà-Cuéllar, M. et al. Do humans spread zoonotic enteric bacteria in Antarctica?. Sci. Total Environ. 654, 190–196 (2019).
Anderson, M. J. & Walsh, D. C. I. PERMANOVA, ANOSIM, and the Mantel test in the face of heterogeneous dispersions: What null hypothesis are you testing?. Ecol. Monogr. 83, 557–574 (2013).
Lott, C. A., Meehan, T. D. & Heath, J. A. Estimating the latitudinal origins of migratory birds using hydrogen and sulfur stable isotopes in feathers: Influence of marine prey base. Oecologia 134, 505–510 (2003).
Góngora, E., Elliott, K. & Whyte, L. Dataset from Gut microbiome is affected by inter-sexual and inter-seasonal variation in diet for thick-billed murres (Uria lomvia). Mendeley Data v4, (2020).
Eriksson, P., Mourkas, E., González-Acuna, D., Olsen, B. & Ellström, P. Evaluation and optimization of microbial DNA extraction from fecal samples of wild Antarctic bird species. Infect. Ecol. Epidemiol. 7, 1386536 (2017).
Parada, A. E., Needham, D. M. & Fuhrman, J. A. Every base matters: Assessing small subunit rRNA primers for marine microbiomes with mock communities, time series and global field samples. Environ. Microbiol. 18, 1403–1414 (2016).
Quast, C. et al. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 41, D590–D596 (2012).
Callahan, B. J. et al. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 13, 581–583 (2016).
Bolyen, E. et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 37, 852–857 (2019).
Bokulich, N. A. et al. Optimizing taxonomic classification of marker-gene amplicon sequences with QIIME 2’s q2-feature-classifier plugin. Microbiome 6, 90 (2018).
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree 2—Approximately maximum-likelihood trees for large alignments. PLoS ONE 5, e9490 (2010).
Braune, B. M., Gaston, A. J., Hobson, K. A., Gilchrist, H. G. & Mallory, M. L. Changes in food web structure alter trends of mercury uptake at two seabird colonies in the Canadian arctic. Environ. Sci. Technol. 48, 13246–13252 (2014).
Callahan, B. J., Sankaran, K., Fukuyama, J. A., McMurdie, P. J. & Holmes, S. P. Bioconductor workflow for microbiome data analysis: From raw reads to community analyses. F1000Research 5, 1492 (2016).
Bokulich, N. A. et al. q2-longitudinal: Longitudinal and paired-sample analyses of microbiome data. mSystems 3, 1–9 (2018).
Wilcoxon, F. Individual comparisons by Ranking methods. Biometrics Bull. 1, 80 (1945).
Anderson, M. J. A new method for non-parametric multivariate analysis of variance. Austral. Ecol. 26, 32–46 (2001).
Lozupone, C. & Knight, R. UniFrac: A new phylogenetic method for comparing microbial communities. Appl. Environ. Microbiol. 71, 8228–8235 (2005).
Faith, D. P. Conservation evaluation and phylogenetic diversity. Biol. Conserv. 61, 1–10 (1992).
Shannon, C. E. & Weaver, W. The Mathematical Theory of Communication (University of Illinois Press, Champaign, 1949).
Kruskal, W. H. & Wallis, W. A. Use of ranks in one-criterion variance analysis. J. Am. Stat. Assoc. 47, 583–621 (1952).
Anderson, M. J. Distance-based tests for homogeneity of multivariate dispersions. Biometrics 62, 245–253 (2006).
Legendre, P. & Legendre, L. Ordination in reduced space. In Numerical Ecology Vol. 24 (eds Legendre, P. & Legendre, L.) 425–520 (Elsevier, Amsterdam, 2012).
Vázquez-Baeza, Y., Pirrung, M., Gonzalez, A. & Knight, R. EMPeror: A tool for visualizing high-throughput microbial community data. Gigascience 2, 16 (2013).
Vázquez-Baeza, Y. et al. Bringing the dynamic microbiome to life with animations. Cell Host Microbe 21, 7–10 (2017).
McMurdie, P. J. & Holmes, S. phyloseq: An R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 8, e61217 (2013).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
McMurdie, P. J. & Holmes, S. Waste not, want not: Why rarefying microbiome data is inadmissible. PLoS Comput. Biol. 10, e1003531 (2014).
Mann, H. B. & Whitney, D. R. On a test of whether one of two random variables is stochastically larger than the other. Ann. Math. Stat. 18, 50–60 (1947).
Bartoń, K. MuMIn: Multi-Model Inference. (2019).
Acknowledgements
This work was supported by Environment and Climate Change Canada; the Canada Research Chairs in Arctic Ecology and Polar Microbiology; the Natural Sciences and Engineering Research Council Discovery Grant Program [241061 and 74221]; and the Northern Contaminants Program of Indigenous and Northern Affairs Canada. The authors would like to thank Mira Okshevsky for critically revising the manuscript. We thank Allison Patterson, Émile Brisson-Curadeau, Sarah Poole, Jean-Hugues Martin, François St-Aubin, Scott Flemming, and Josiah Nakoolak for help collecting the field samples. Thanks also to the Polar Continental Shelf Program of Natural Resources Canada and Rick Armstrong at the Nunavut Research Institute for helping with logistics.
Author information
Authors and Affiliations
Contributions
E.G., K.H.E, and L.W. designed the study. E.G. led the sample collection, conducted the experimental work, data analysis, and wrote the paper. K.H.E. and L.W. reviewed and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Góngora, E., Elliott, K.H. & Whyte, L. Gut microbiome is affected by inter-sexual and inter-seasonal variation in diet for thick-billed murres (Uria lomvia). Sci Rep 11, 1200 (2021). https://doi.org/10.1038/s41598-020-80557-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-80557-x
This article is cited by
-
Gut fungi of black-necked cranes (Grus nigricollis) respond to dietary changes during wintering
BMC Microbiology (2024)
-
From islands to infectomes: host-specific viral diversity among birds across remote islands
BMC Ecology and Evolution (2024)
-
No evidence for associations between brood size, gut microbiome diversity and survival in great tit (Parus major) nestlings
Animal Microbiome (2023)
-
Cloacal microbiota are biogeographically structured in larks from desert, tropical and temperate areas
BMC Microbiology (2023)
-
Parental care contributes to vertical transmission of microbes in a skin-feeding and direct-developing caecilian
Animal Microbiome (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.