Abstract
In an attempt to evaluate the susceptibility of the mosquito Culex quinquefasciatus to bacterial agents, a population naturally infected with a Wolbachia pipientis wPipSJ native strain was tested against the action of three bacterial mosquitocides, Bacillus thuringiensis subsp. israelensis, Bacillus wiedmannii biovar thuringiensis and Lysinibacillus sphaericus. Tests were carried out on mosquito larvae with and without Wolbachia (controls). Cx. quinquefasciatus naturally infected with the native wPipSJ strain proved to be more resistant to the pathogenic action of the three mosquitocidal bacterial strains. Additionally, wPipSJ was fully characterised using metagenome-assembled genomics, PCR–RFLP (PCR-Restriction Fragment Length Polymorphism) and MLST (MultiLocus Sequence Typing) analyses. This Wolbachia strain wPipSJ belongs to haplotype I, group wPip-III and supergroup B, clustering with other mosquito wPip strains, such as wPip PEL, wPip JHB, wPip Mol, and wAlbB; showing the southernmost distribution in America. The cytoplasmic incompatibility phenotype of this strain was revealed via crosses between wildtype (Wolbachia+) and antibiotic treated mosquito populations. The results of the tests with the bacterial agents suggest that Cx. quinquefasciatus naturally infected with wPipSJ is less susceptible to the pathogenic action of mosquitocidal bacterial strains when compared with the antibiotic-treated mosquito isoline, and is more susceptible to B. thuringiensis subsp. israelensis than to the other two mosquitocidal agents.
Similar content being viewed by others
Introduction
The mosquito Culex quinquefasciatus (Diptera: Culicidae) is a known vector of pathogens of significant medical importance1 in Africa, Central and South America and Asia; successfully transmiting major parasites like filaria (Wuchereria bancrofti and Dirofilaria immitis), and well described viruses, West Nile virus (WNV) or St. Louis encephalitis (SLEV) and Venezuelan equine encephalitis2,3,4.
Like other mosquito species, adults of Cx. quinquefasciatus only fly short distances and are believed to disperse throughout the world by anthropic action. Their immature stages can be found in exposed tires that fill with water, and any type of water container on modern trucks or ships, and adults have been even detected on all kind of vehicles including airplanes or long-distance buses5. In Argentina, it occurs in different biogeographic regions6, especially urban zones7, being a competent vector of WNV and SLEV8,9.
To control mosquitoes, a number of biological methods have been applied in the past, among them the IIT (Incompatible Insect Technique) caused by Wolbachia pipientis. Despite its very first success on Culex mosquitoes in Burma, now dating over 50 years10,11, the Wolbachia IIT technique has only seen a rise in applications in the last few years. This bacterium was first detected in the common household species Culex pipiens, in 1924, by Hertig and Wolbach12. It is the most widespread bacterial endosymbiont infecting terrestrial arthropods, mainly insects, but some arachnids, freshwater crustaceans and filarial nematodes too13. Wolbachia is maternally transmitted and some of its reproductive phenotypes have been well studied, like CI or cytoplasmic incompatibility, parthenogenesis, feminization, as well as death of male offspring14; with CI being the most studied phenotype in mosquitoes and defined as an early form of embryonic lethality, since paternal chromosomes are lost during the first mitosis15.
Moreover, Wolbachia can counteract the effect of human pathogens within mosquito species16. In artificially infected Aedes aegypti with wMel Wolbachia from Drosophila melanogaster, the mosquito ability to transmit arboviruses is reduced (dengue, chikungunya and Zika)17,18. Anopheles stephensi populations infected with Wolbachia wAlbB strain, from Aedes albopictus, are resistant to Plasmodium falciparum19 (malaria parasite), but the Plasmodium interference phenotype is not the same for all vectors. For example, in an Anopheles gambiae population transiently infected with wAlbB, the number of Plasmodium berghei oocysts was higher, increasing the risk of malaria transmission20. Besides, Wolbachia-infection of Culex tarsalis enhanced the infection rate of WNV18.
It has also been demonstrated that Wolbachia infection confers Diptera species some resistance to insect pathogens21. In experimentally infected Cx. pipiens mosquitoes, it is clear an upregulation of several genes for immune effector molecules involved in antimicrobial pathways22. However, Endersby and Hoffmann while studying sensitivity of Ae. aegypti to the insecticides biphentrine, temephos and s-metoprene, as well as the bio-agent Bacillus thuringiensis, without finding significant differences comparing Wolbachia+ and uninfected mosquitos23. It has also been shown that the Wolbachia amount is higher in Cx. pipiens lines resistant to organophosphorus insecticides than in susceptible mosquitoes24; and it has been detected that this bacterial load would be responsible for fitness effects such as preimaginal mortality, and adults of smaller body size and lower fertility25. The results obtained in different mosquito trials demonstrate that Wolbachia must be carefully assessed for use as a biological control agent. The outcomes will not only depend on the mosquito species studied but also on the Wolbachia strain, and these two factors would vary in their response to other chemical or biological agents applied to the control of mosquito populations.
The specific action of bacterial mosquitocidal proteins from Bacillus thuringiensis subsp. israelensis and Lysinibacillus sphaericus as biological insecticide has been widely researched, and not only the way of action of some of these proteins such as Cry, Bim and Cyt has been demonstrated, but also the synergistic interactions between them, increasing their toxic action or reversing the insects resistance26,27,28. It has been recently reported that the epithelial lesions produced by B. thuringiensis crystal proteins in Spodoptera littoralis could induce the reduction of the host immune response permitting the midgut bacteria replication resulting in a lethal septicemia due to the microbiota resident in the host midgut29. However, the interaction between the microbial flora present in mosquitoes, including symbiotic bacteria, with entomopathogenic bacteria that could modify the susceptibility to toxins produced by them or be responsible for other effects, has not yet been deeply analyzed and should be considered within the context of integrated management programs of mosquitoes.
The IIT method makes use of Wolbachia-induced CI, resulting in the massive release and spread of mosquito males that are Wolbachia-incompatible with the wild-type females30,31. IIT can also take benefit of the Wolbachia-induced CI to replace a natural population with a drive-wanted phenotype32. To optimize the combined application of control methods, towards a much wiser vector control, like the combination of IIT, that relies on Wolbachia infection, with the use of its host response to entomopathogens, more research on susceptibility of w+-mosquito lines to mosquitocidal bacteria is needed.
As a new strain of Wolbachia has been found naturally infecting Cx. quinquefasciatus in southern South America, Argentina, the aim of this study was to examine the impact of three entomopathogenic bacteria on Cx. quinquefasciatus larvae infected with this Wolbachia CI-strain. This strain is fully characterised here using metagenome-assembled genomics, PCR–RFLP (PCR-Restriction Fragment Length Polymorphism) and MLST (MultiLocus Sequence Typing) analyses; and its CI phenotype studied via crosses between wildtype and antibiotic cured Cx. quinquefasciatus.
Results
Wolbachia effect on Cx. quinquefasciatus fitness
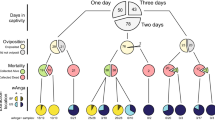
Fecundity (egg laid per female) and fertility (total eggs hatched) rates of mosquitoes infected with Wolbachia (w+) or treated with antibiotics (w−) show no significant differences for both tests (Mann–Whitney test, p > 0.99) (Fig. 1a). However, when analyzing time of larval development, statistical differences were observed between the mosquito lines (Fig. 1b); and Kaplan–Meier survival curves showed that the Wolbachia infected lines presented higher survival rates than the antibiotic-treated (Log-rank (Mantel-Cox) test: χ2 = 54.08, p < 0.01).
Susceptibility of Wolbachia (w +)- or (w −)-Cx. quinquefasciatus larvae to entomopathogenic bacteria
In all, the three entomopathogens used in this experiment, B. thuringiensis subsp. israelensis (Bti strain H14), Bacillus wiedmannii biovar thuringiensis (Bwt strain FCC 41) and L. sphaericus (Ls strain 2362) were required in greater concentrations for (w+)-larvae. LC50 values were significantly different between (w+)- and (w−)-larvae, and L. sphaericus showed the highest toxic activity difference (ratio tests, p < 0.05). Comparisons of LC50 values between lines, for each different entomopathogenic bacteria are shown in Table 1. Furthermore, mortality of w+ and w− larvae were compared when they were subjected to similar concentrations of the different entomopathogenic bacteria. Mortalities were significantly higher in the antibiotic-treated lines (Bti H14, p = 0.0117; Bwt FCC 41, p = 0.0508; Ls 2362, p = 0.0001) (Fig. 2a,c,e). Plots of mortality values for each line per concentration and per strain allow visualization of differences by concentration gradient (Fig. 2b,d,f).
Effect of Bacillus thuringiensis sp. israelensis (Bti H14), Bacillus wiedmannii biovar thuringiensis (Bwt FCC 41) and Lysinibacillus sphaericus (Ls 2362) on (w+) or (w−) larvae. (a, c, e) Average of dead larvae (w+ and w-) by entomophatogenic bacteria strain. Means with different letters are significantly different. (b, d, f) Linear regressions of mortality per concentration between w+ and w− lines with different entomopathogenic bacteria strain [Inset b: magnification of the curves at initial concentrations to visualize differences].
Molecular characterization of a native Wolbachia wPip strain
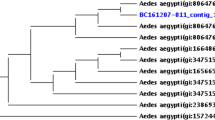
According to the resulting phylogenetic tree using five concatenated MLST genes (2079 bp), the wPip strain of the Cx. quinquefasciatus San Juan line (wPipSJ) of this study belongs to the B supergroup, and it has a 100% similarity to other wPip strains previously reported (Fig. S1 and Table S1).
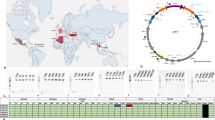
The ank2 and pk1 allelic profiling (for wPip-I to wPip-V) of the Wolbachia San Juan strain line resulted in a band pattern matching with wPip-III group (by the specific PCR–RFLP analysis). Additionally, seven polymorphic wPip genes of the Cx. quinquefasciatus line were analysed to allow assignment to a wPip haplotype. Based on this, wPipSJ belongs to haplotype I (Table S2). Up to date, this is the strain showing the southernmost distribution of haplotype I and wPip-III group in Cx. quinquefasciatus in America (Fig. S2).
The metagenome-assembled genome of the wPipSJ strain consists of 1.3 Mbps, from 121 contigs and an N50 of 20,098 bp, and an average G + C content of 34.5%. This draft genome has 1171 open reading frames, 5 rRNA and 39 tRNA genes; and its total coding percentage is 82.3%. According to the core genome analysis of the wPipSJ and additional 29 available Wolbachia genomes, the phylogenetic reconstruction placed the wPipSJ strain in Wolbachia supergroup B, in the same cluster as other mosquito wPip strains, such as wPip PEL, wPip JHB, wPip Mol, and wAlbB (Fig. 3 and Table S3).
Maximum-likelihood phylogenetic tree reconstruction performed with GET_PHYLOMARKERS and IQ-TREE (500 ultrafast boostrap replicates) according to the Wolbachia core genome from 30 available genomes. Numbers in the branches indicate ultrafast bootstrap value, and letters on the right indicate the Wolbachia supergroup. The symbol // indicates a trimmed branch.
Cytoplasmic incompatibility analysis of the wPipSJ strain
Cx. quinquefasciatus crosses were performed between the wild type w+PipSJ and the antibiotic-treated w−PipSJ populations. Crosses resulted incompatible when w+ males mated w− females, obtaining 0% of eggs hatched per female, or 100% embryo mortality (Fig. 4). Both crossing types, w+FF x w+MM and w+FF x w−MM were compared with a t-test (unpaired), and highly significant differences were observed, p < 0.001. Extra confirmation of successful mating was visualised by dissection of females, showing sperm inside the spermatheca in all females with no hatched eggs (Supplementary Fig. S3). Detection of Wolbachia in all males from the incompatible crossings was confirmed by using PCR with the wsp gene primers, further description of the protocol is described in experimental procedures.
Complete unidirectional cytoplasmic incompatibility in Culex quinquefasciatus lines. Egg hatching percentage in crosses between the wild type (w+PipSJ) and tetracycline-treated (w−PipSJ) strains of Cx. quinquefasciatus. Sixteen couples were used in each type of crossing. Each dot represents a mosquito egg raft from a unique female, carrying approximately 150 eggs per raft. The median number of hatched eggs/egg-raft are shown in horizontal bars.
Discussion
The wPipSJ strain expresses complete unidirectional CI (100% embryo mortality) in Cx. quinquefasciatus crossings, showing promising application for IIT strategies30.
The variability in the response of mosquitoes carrying the native Wolbachia strain and Wolbachia-free isoline to the different entomopathogenic bacteria, used in this work, could be explained by the differential production of putative virulence factors and extracellular degradative enzymes, such as bacteriocins, phospholipases C, enterotoxins, hemolysins, and some antibacterial proteins27,33,34; all this might cause modifications or imbalances in the intestinal microflora of insects that, in some way, could participate in the pathogenic activity (Table 1). Moreover, the mosquitocidal bacterial strains Bti H14 and Ls 2362 showed significant differences in the mortality of Cx. quinquefasciatus isoline (wild-type) naturally infected with Wolbachia, (Fig. 2), while there was no significant difference in the mortality of the mosquito isolines when exposed to the native or Argentinean mosquitocidal agent Bwt FCC41. The differences observed in the pathogenic action between bacterial pathogens could be justified with the increase in gene expression related to the mosquito immune system induced under the presence of Wolbachia like promoting the induction of reactive oxygen species or ROS which regulates the activation of immune genes to secrete proteins (mosquito innate response)35. In relation to the activation of an immune response, in a recent study on Wolbachia proteins, specifically WSP, it has been found that engineered WSP in Asaia bacteria carried by Ae. aegypti and An. stephensi mosquitoes, the mosquito (host) immune system is activated inhibiting the development of filarial parasites36. Another reason of the mosquito/host lower susceptibility could be related to improve nutrition in Wolbachia infected insects as it happens with several bloodsucking arthropods. In general, vertebrate blood as food source is deficient in some micronutrients, like B vitamins, required by hematophagous insects being supplied by the metabolism of some microorganisms associated with them as part of this microflora. For vitamins B2 and B7, complete biosynthetic pathways were found from the genome sequence analysis of Wolbachia while studies on diet revealed that vitamins derived from Wolbachia contribute to the nutrition of the host arthropod37.
Endersby and Hoffmann studied a number of Ae. aegypti w+ lines for their susceptibility to Bti H14 and, in general, only found slight differences in the response between w+ lines, but, it is not possible to know from this study how many backcrossings they received and if their genetic makeup was that of a lab naïve population23.
In order to demonstrate that the Wolbachia strain from Cx. quinquefasciatus in Argentina is the B supergroup described by Baldo et al.38, several methods were used, such as characterization of the wsp gene by PCR, application of MLST markers, plus, the analysis of the core genome, by comparison with other available sequences in public databases. Identification of Wolbachia haplotypes from different geographic regions allows to track their distribution and diversity, providing information on the evolution and dynamics of Wolbachia in its mosquito hosts39. Wolbachia strains vary, but the addition of a new element, a just discovered mobile putative plasmid (pWCP), made of 9.23 kbp, including 14 genes, adds even more diversity to mosquito’s Wolbachia, and, consequently more challenges to the dynamics of host phenotypes40.
For the first time, the southernmost distribution of Wolbachia haplotype I in Cx. quinquefasciatus in America is reported. This strain, wPipSJ, belongs to the wPip-III group as it was previously described in Cx. pipiens in North America41. Cx. quinquefasciatus naturally infected with wPipSJ is less susceptible to the pathogenic action of mosquitocidal bacterial strains when compared with the antibiotic-treated mosquito isoline. The lack of fitness costs, normally associated with strong cytoplasmic incompatibility, suggests that the newly characterized wPipSJ from Argentina could be used in mosquito population suppression strategies based on IIT. These results encourage more research on native Wolbachia strains infecting mosquitoes in different parts of the world analysing the effect of them on the mosquito response to entomopathogens that could be of further help in future integrated mosquito vector management programs.
Methods
Permit for research with animals
Protocols for blood feeding the mosquitos on mice were reviewed and approved by the Animal Experimental Committee at the Faculty of Exact and Natural Sciences, Mar del Plata University (Institutional Committee on Care and Use of Experimental Animals (CICUAL) No. 2555-04-14). Mice were handled in strict accordance with National Health Service and Food Quality (SENASA) guidelines (Argentina) following the 2011 revised form of The Guide for the Care and Use of Laboratory Animals published by the U.S. National Institutes of Health.
Mosquito rearing
Mosquito larvae were collected in natural breeding sites from San Juan province, Argentina. Insects were identified based on the larval fourth stage or adult females’ morphological characters and male genitalia (those reared to adult) according to Rossi et al.42 and Harbach43 respectively. Mosquito laboratory lines were established from a naturally infected population and were maintained at the Insect Biological Control Laboratory of the INBIOTEC-CONICET, FIBA (Argentina) following lab conditions such as 24 °C and 80 ± 5% RH and a 12 h light: 12 h dark photoperiodicity. Larvae were fed using fish food (Shulet Carassius). Adult stages were provided with a 10% solution of sucrose was given to adults and females were allowed to feed blood from mice.
Wolbachia detection
Detection of Wolbachia followed PCR using the wsp-specific primers wsp-81F and wsp-691R44 from DNA extracted from single female mosquitos (twenty by colony) using PureLink Genomic DNA Mini Kit (Invitrogen, Grand Island, New York, USA) according to the manufacturer’s instructions. To verify the quality of the DNA, the amplification of the cytochrome c oxidase (COI) gene was used according to Folmer et al.45.
Generation of antibiotic-treated Culex quinquefasciatus line
In order to obtain a Wolbachia free line, a Cx. quinquefasciatus wild type colony was treated with tetracycline as previously described31,46. A tetracycline solution (0.1 mg/mL) was used for exposure of first larvae during development. Pupae were transferred to clean water without antibiotic and reared to adulthood, and adults allowed to feed ad libitum with sucrose 10% containing 0.05 mg/mL of tetracycline hydrochloride (final concentration, Sigma, St Louis, MO; Cat. No. T33 83) for three consecutive generations. To confirm Wolbachia in each mosquito generation, ten individual females and ten larvae were screened using the wsp-specific amplification as described above. The line wild type naturally infected with Wolbachia (w+) and Wolbachia free treated with tetracycline (w−) were kept under same environmental conditions previously described. The antibiotic was removed in the fourth generation and the experimental work was initiated after at least three generations permitting capture of environmental microbiota, and as well as recovery from the side effects of the treatment with antibiotic. The absence of Wolbachia in mosquito adults cured with tetracycline was confirmed by PCR amplification of the wsp gene.
Fitness parameters analysis
In order to analyze developmental differences Cx. quinquefasciatus lines were compared. Following Baton et al.31 3 variables were considered to study mosquito fitness (w+ and w−): survival, fecundity and fertility. For that, egg laid, hatching rate, developmental stages and individual survivor number per line were daily monitored. Four rafts of eggs from each line were placed in individual containers with dechlorinated water until neonates hatched. Larvae were provided with commercial fish food (Shulet Carassius) to pupal stage. Eggs, larvae, pupae, and adults were quantified under a binocular microscope (Nikon SMZ800) until adults emerged. The full experiment was repeated 4 times considering four different generations of mosquito lines from the laboratory.
Larvicidal activity of bacterial entomopathogens
Entomopathogenic effectivity of the B. thuringiensis subsp. israelensis (H14), B. wiedmannii biovar thuringiensis (FCC 41) and L. sphaericus (2362) strains was determined on larvae of Cx. quinquefasciatus (w+) and (w−) laboratory rearing lines, as previously described47. Twenty L2 instar larvae were placed in 20 mL of dechlorinated water. Six concentrations (0.01 to 1 µg/mL) of the spore–crystal complexes were used, and fish food was included as negative control. The bioassay was repeated three times. Examination of larvae was performed after 24 h, and these specimens were incubated at 28 °C. Using Probit analysis48 mean lethal concentration or LC50 was estimated.
Wolbachia strain molecular characterization
The wPip infection was characterized through the analysis of 13 different Wolbachia markers, which were amplified by PCR and Sanger sequenced, according to methodologies previously described. The markers were: (i) the Wolbachia surface protein gene (wsp)44; (ii) the analysis of five house-keeping gene sequences for the Wolbachia MLST methodology according to Baldo et al.38. For that, multiple alignment is performed with the concatenated sequences of the genes (gatB, coxA, hcpA, ftsZ, and fbpA, downloaded from https://pubmlst.org/wolbachia/ and detailed in Table S2). The phylogeny was estimated by Mr Bayes program, using members of all Wolbachia supergroups; (iii) the analysis of two ANK Wolbachia markers (ank2 and pk1) by a specific PCR–RFLP assay described by Dumas et al.41 that allows to distinguish the 5 known wPip groups (wPip-I to wPip-V); and (iv) the identification of wPip haplotypes based on the polymorphism of seven genes: the former two ANK Wolbachia markers plus five additional genes, the DNA mismatch repair protein gene MutL, the ANK gene pk2, the methylase gene GP12, the putative secreted protein gene GP15 and the regulatory protein gene RepA49. In this case, the haplotypes of each gene are identified based on the comparison of the sequence of the gene with all of the sequences of the described haplotypes methodology, according to Atyame et al.50.
DNA extraction, library building construction, metagenome assembly, and analysis
Total DNA of 10 fourth stage Cx. quinquefasciatus larvae was extracted using the DNeasy Blood & Tissue kit (QIAGEN) and the metagenomic library was prepared using the TruSeq DNA Nano kit. Paired-end sequencing was conducted in a Novaseq platform with a 150 bp read length, generating 32 Gbp in raw reads. Quality check of reads was performed with FastQC (http://www.bioinformatics.babraham.ac.uk/projects/fastqc/). Filtering of Wolbachia reads was computed with Bowtie251 using publicly available Wolbachia from Cx. quinquefasciatus genomes. With the extracted Wolbachia reads the genome was assembled using SPAdes52. The completeness of the genome was checked with BUSCO53 and an alignment with a public available wPip genome was performed using Mauve software54 to check for genomic rearrangements and differences. The phylogenetic reconstruction was performed with GET_PHYLOMARKERS55 using 29 available Wolbachia genomes listed in NCBI genome database (Table S3). This pipeline identifies marker genes and estimates genome phylogenies from the orthologous clusters computed by GET_HOMOLOGUES56. The phylogeny was computed with IQ-TREE57 and the tree was edited with iTOL58.
Crossing experiments and cytoplasmic incompatibility analysis
In order to determine the CI level, experimental crosses between FF (females) w−PipSJ × MM (males) w+PipSJ and FF w−PipSJ × MM w−PipSJ were performed. Pupae of each line were separated in individual glass tubes to avoid mating once adults emerged. Two days after emergence mass crosses were performed between 25 virgin adults of each sex during two days in a single cage (30 cm × 30 cm). After three days, females were blood-fed for 4 h and twenty-four hours later were removed and individually placed into smaller cages (15 × 15 cm) for oviposition. The eggs raft from each female was placed in containers with 10 mL of dechlorinated water and eggs hatched were quantified under a binocular microscope. Crossing experiments were repeated up to achieve 16 eggs raft per crossing, using five generations of laboratory mosquito lines. The spermatheca of all females that laid eggs was checked to determine the occurrence of spermatozoa and to confirm mating. Infection was detected by using the diagnostic PCR assay.
Statistical analyses
Normality and homogeneity of variance were analyzed using the Shapiro-Wilk and Levene tests respectively59. The differences between % of hatched eggs per female were analyzed with Mann–Whitney U tests (p = 0.05). The Kaplan–Meier survival estimator was used for generating survival functions. The differences between average of dead larvae per concentration per entomopathogenic bacteria strain were analyzed with Wilcoxon matched-pairs singed rank tests. Linear regressions were plotted to observe the differences in mortality per concentration between w+PipSJ and w−PipSJ lines. For hatching rate data, an unpaired t-test comparison was used. Statistical analyses of the survival data were performed using GraphPad Prism version 4.01 for Windows (GraphPad Software, La Jolla California USA, www.graphpad.com). To establish the 24 h LC50 Log-Probit was applied, adding the 95% confidence intervals (CI) from R, specifically the package ‘ecotox’, v1.3.3, (CRAN60). Significant differences between the LC50 of the different lines were evaluated with ratio tests: if the 95% confidence interval is including 1, the difference of LC50 values is not considered significant61.
References
Turell, M. J. Members of the Culex pipiens complex as vectors of viruses. J. Am. Mosq. Control Assoc. 28, 123–126 (2012).
Simonsen, P. E. & Mwakitalu, M. E. Urban lymphatic filariasis. Parasitol. Res. 112, 35–44 (2013).
Go, Y. Y., Balasuriya, U. B. R. & Lee, C. Zoonotic encephalitides caused by arboviruses: transmission and epidemiology of alphaviruses and flaviviruses. Clin. Exp. Vaccine Res. 3, 58–77 (2014).
Hadfield, J. et al. Twenty years of West Nile virus spread and evolution in the Americas visualized by Nextstrain. PLoS Pathog. 15, e1008042 (2019).
Lounibos, L. P. Invasions by insect vectors of human disease. Annu. Rev. Entomol. 47, 233–266 (2002).
Rossi, G. C. Annotated checklist, distribution, and taxonomic bibliography of the mosquitoes (Insecta: Diptera: Culicidae) of Argentina. Check List. 11, 1712 (2015).
Cardo, M. V., Rubio, A., Vezzani, D. & Carbajo, A. E. Assessment of Culex pipiens bioforms in the world’s southernmost distribution limit. Mem. Inst. Oswaldo Cruz. 115, e190390 (2020).
Artsob, H. et al. West Nile Virus in the New World: trends in the spread and proliferation of West Nile Virus in the Western Hemisphere. Zoonoses Public Health. 56, 357–369 (2009).
Diaz, L. A. et al. Transmission of endemic St Louis encephalitis virus strains by local Culex quinquefasciatus populations in Cordoba, Argentina. Trans. R. Soc. Trop. Med. Hyg. 107, 332–334 (2013).
Zhang, D., Lees, R. S., Xi, Z., Bourtzis, K. & Gilles, J. R. L. Combining the sterile insect technique with the incompatible insect technique: III-robust mating competitiveness of irradiated triple Wolbachia-infected Aedes albopictus males under semi-field conditions. PLoS ONE 11, e0151864 (2016).
Laven, H. Eradication of Culex pipiens fatigans through cytoplasmic incompatibility. Nature 216, 383–384 (1967).
Hertig, M. & Wolbach, B. Studies on Rickettsia-like micro-organisms in insects. J. Med. Res. 44, 329–374 (1924).
Gerth, M., Gansauge, M. T., Weigert, A. & Bleidorn, C. Phylogenomic analyses uncover origin and spread of the Wolbachia pandemic. Nat. Commun. 5, 5117. https://doi.org/10.1038/ncomms6117 (2014).
McGraw, E. A. & O’Neill, S. L. Beyond insecticides: new thinking on an ancient problem. Nat. Rev. Microbiol. 11, 181–193 (2013).
Tram, U. Role of elayed nuclear envelope breakdown and mitosis in Wolbachia-induced cytoplasmic incompatibility. Science 296, 1124–1126 (2002).
Yin, C., Sun, P., Yu, X., Wang, P. & Cheng, G. Roles of symbiotic microorganisms in arboviral infection of arthropod vectors. Trends Parasitol. 36, 607–615 (2020).
Moreira, L. A. et al. A Wolbachia symbiont in Aedes aegypti limits infection with dengue, chikungunya, and Plasmodium. Cell 139, 1268–1278 (2009).
Lee, W.-S., Webster, J. A., Madzokere, E. T., Stephenson, E. B. & Herrero, L. J. Mosquito antiviral defense mechanisms: a delicate balance between innate immunity and persistent viral infection. Parasit. Vectors. 12, 165 (2019).
Bian, G. et al. Wolbachia invades Anopheles stephensi populations and induces refractoriness to Plasmodium infection. Science 340, 748–751 (2013).
Hughes, G. L., Rivero, A. & Rasgon, J. L. Wolbachia can enhance Plasmodium infection in mosquitoes: implications for malaria control?. PLoS Pathog. 10, e1004182 (2014).
Panteleev, D. I. et al. The endosymbiotic bacterium Wolbachia enhances the nonspecific resistance to insect pathogens and alters behavior of Drosophila melanogaster. Genetika 43, 1277–1280 (2007).
Duron, O. et al. Variability and expression of ankyrin domain genes in Wolbachia variants infecting the mosquito Culex pipiens. J. Bacteriol. 189, 4442–4448 (2007).
Endersby, N. M. & Hoffmann, A. A. Effect of Wolbachia on insecticide susceptibility in lines of Aedes aegypti. Bull. Entomol. Res. 103, 269–277 (2013).
Berticat, C., Rousset, F., Raymond, M., Berthomieu, A. & Weill, M. High Wolbachia density in insecticide–resistant mosquitoes. Proc. R. Soc. Lond. B Biol. Sci. 269, 1413–1416 (2002).
Duron, O. et al. High Wolbachia density correlates with cost of infection for insecticide resistant Culex pipiens mosquitoes. Evolution 60, 303 (2006).
Ben-Dov, E. Bacillus thuringiensis subsp. israelensis and its dipteran-specific toxins. Toxins 6, 1222–1243 (2014).
Berry, C. The bacterium, Lysinibacillus sphaericus, as an insect pathogen. J. Invertebr. Pathol. 109, 1–10 (2012).
Federici, B. A. et al. Developing recombinant bacteria for control of mosquito larvae. J. Am. Mosq. Control Assoc. 23, 164–175 (2007).
Caccia, S. et al. Midgut microbiota and host immunocompetence underlie Bacillus thuringiensis killing mechanism. Proc. Natl. Acad. Sci. 113, 9486–9491 (2016).
Sinkins, S. P. Wolbachia and cytoplasmic incompatibility in mosquitoes. Insect Biochem. Mol. Biol. 34, 723–729 (2004).
Baton, L. A., Pacidônio, E. C., Gonçalves, D. S. & Moreira, L. A. wFlu: characterization and evaluation of a native Wolbachia from the mosquito Aedes fluviatilisas a potential vector control agent. PLoS ONE 8, e59619 (2013).
Brelsfoard, C. L. & Dobson, S. L. Wolbachia-based strategies to control insect pests and disease vectors. Asia-Pac. J. Mol. Biol. Biotechnol. 17, 10 (2009).
Salamitou, S. et al. The plcR regulon is involved in the opportunistic properties of Bacillus thuringiensis and Bacillus cereus in mice and insects. Microbiology 146, 2825–2832 (2000).
Lazarte, J. N., Lopez, R. P., Ghiringhelli, P. D. & Berón, C. M. Bacillus wiedmannii biovar thuringiensis: a specialized mosquitocidal pathogen with plasmids from diverse origins. Genome Biol. Evol. https://doi.org/10.1093/gbe/evy211 (2018).
Brennan, L. J., Keddie, B. A., Braig, H. R. & Harris, H. L. The endosymbiont Wolbachia pipientis induces the expression of host antioxidant proteins in an Aedes albopictus cell line. PLoS ONE 3, e2083 (2008).
Epis, S. et al. Chimeric symbionts expressing a Wolbachia protein stimulate mosquito immunity and inhibit filarial parasite development. Commun. Biol. 3, 105 (2020).
Douglas, A. E. The B vitamin nutrition of insects: the contributions of diet, microbiome and horizontally acquired genes. Curr. Opin. Insect Sci. 23, 65–69 (2017).
Baldo, L. et al. Multilocus sequence typing system for the endosymbiont Wolbachia pipientis. Appl. Environ. Microbiol. 72, 7098–7110 (2006).
Bourtzis, K. et al. Harnessing mosquito–Wolbachia symbiosis for vector and disease control. Acta Trop. 132, S150–S163 (2014).
Reveillaud, J. et al. The Wolbachia mobilome in Culex pipiens includes a putative plasmid. Nat. Commun. 10, 1051 (2019).
Dumas, E. et al. Population structure of Wolbachia and cytoplasmic introgression in a complex of mosquito species. BMC Evol. Biol. 13, 181 (2013).
Rossi, G. C., Mariluis, J. C., Schnack, J. A. & Spinelli, G. R. Dípteros vectores (Culicidae y Calliphoridae) de la provincia de Buenos Aires. ProBiota Cobiobo. 3, 4 (2002).
Harbach, R. E. Culex pipiens: species versus species complex taxonomic history and perspective. J. Am. Mosq. Control Assoc. 28, 10–23 (2012).
Braig, H. R., Zhou, W., Dobson, S. L. & O’Neill, S. L. Cloning and characterization of a gene encoding the major surface protein of the bacterial endosymbiont Wolbachia pipientis. J. Bacteriol. 180, 2373–2378 (1998).
Folmer, O., Black, M., Hoeh, W., Lutz, R. & Vrijenhoek, R. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol. Mar. Biol. Biotechnol. 3, 294–299 (1994).
Dobson, S. L. & Rattanadechakul, W. A novel technique for removing Wolbachia infections from Aedes albopictus (Diptera: Culicidae). J. Med. Entomol. 38, 844–849 (2001).
Berón, C. M. & Salerno, G. L. Cloning and characterization of a novel crystal protein from a native Bacillus thuringiensis isolate highly active against Aedes aegypti. Curr. Microbiol. 54, 271–276 (2007).
Robertson, J. L., Russell, R. M. & Savin, N. E. POLO: a user’s guide to Probit Or LOgit analysis. (Vol. 38). Department of Agriculture, Forest Service, Pacific Southwest Forest and Range Experiment Station (1980).
Atyame, C. M., Delsuc, F., Pasteur, N., Weill, M. & Duron, O. Diversification of Wolbachia endosymbiont in the Culex pipiens mosquito. Mol. Biol. Evol. 28, 2761–2772 (2011).
Atyame, C. M. et al. Multiple Wolbachia determinants control the evolution of cytoplasmic incompatibilities in Culex pipiens mosquito populations: Wolbachia CI evolution in mosquitoes. Mol. Ecol. 20, 286–298 (2011).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods. 9, 357–359 (2012).
Bankevich, A. et al. SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. J. Comput. Biol. 19, 455–477 (2012).
Simão, F. A., Waterhouse, R. M., Ioannidis, P., Kriventseva, E. V. & Zdobnov, E. M. BUSCO: assessing genome assembly and annotation completeness with single-copy orthologs. Bioinformatics 31, 3210–3212 (2015).
Darling, A. C. E. Mauve: Multiple alignment of conserved genomic sequence with rearrangements. Genome Res. 14, 1394–1403 (2004).
Vinuesa, P., Ochoa-Sánchez, L. E. & Contreras-Moreira, B. GET_PHYLOMARKERS, a software package to select optimal orthologous clusters for phylogenomics and inferring pan-genome phylogenies, used for a critical geno-taxonomic revision of the genus stenotrophomonas. Front. Microbiol. 9, 771 (2018).
Contreras-Moreira, B. & Vinuesa, P. GET_HOMOLOGUES, a versatile software package for scalable and robust microbial pangenome analysis. Appl. Environ. Microbiol. 79, 7696–7701 (2013).
Nguyen, L.-T., Schmidt, H. A., von Haeseler, A. & Minh, B. Q. IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol. Biol. Evol. 32, 268–274 (2015).
Letunic, I. & Bork, P. Interactive Tree Of Life (iTOL) v4: recent updates and new developments. Nucl. Acids Res. 47, W256–W259 (2019).
Rohlf, J. F. & Sokal, R. R. Biometry: The Principles and Practice of Statistics in Biological Research (WH Freeman Co., New York, 1980).
Hlina, B. L. Ecotox: Analysis of Ecotoxicology in R (2017).
Wheeler, M. W., Park, R. M. & Bailer, A. J. Comparing median lethal concentration values using confidence interval overlap or ratio tests. Environ. Toxicol. Chem. 25, 1441–1444 (2006).
Acknowledgements
We especially thank Prof. Ana Tassi (former Professor of English Grammar, Universidad Nacional de Mar del Plata) for proof-editing of the draft manuscript. This study was supported by Grants of the Agencia Nacional de Promoción Científica y Tecnológica (ANPCyT PICT-2015-0575), CONICET PUE 2017-0101 and Universidad Nacional de Mar del Plata (15/E883 EXA925/19).
Author information
Authors and Affiliations
Contributions
Conception and designed the experiments: L.M.D.N., M.F.G., J.N.L., M.A.P. and C.M.B., performed the experiments: L.M.D.N., M.F.G., J.N.L., and C.M.B., acquisition of data: L.M.D.N., M.F.G., J.N.L. and C.M.B., interpretation of data: L.M.D.N., M.F.G., M.A.P. and C.M.B., write and revise the manuscript: L.M.D.N., M.F.G., M.A.P. and C.M.B.. Contributed reagents, materials, analysis tools: L.M.D.N. and C.M.B. All authors read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Díaz-Nieto, L.M., Gil, M.F., Lazarte, J.N. et al. Culex quinquefasciatus carrying Wolbachia is less susceptible to entomopathogenic bacteria. Sci Rep 11, 1094 (2021). https://doi.org/10.1038/s41598-020-80034-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-80034-5
This article is cited by
-
Wolbachia dominance influences the Culex quinquefasciatus microbiota
Scientific Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.