Abstract
The severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) pandemic has devastated global public health systems and economies, with over 52 million people infected, millions of jobs and businesses lost, and more than 1 million deaths recorded to date. Contact with surfaces contaminated with droplets generated by infected persons through exhaling, talking, coughing and sneezing is a major driver of SARS-CoV-2 transmission, with the virus being able to survive on surfaces for extended periods of time. To interrupt these chains of transmission, there is an urgent need for devices that can be deployed to inactivate the virus on both recently and existing contaminated surfaces. Here, we describe the inactivation of SARS-CoV-2 in both wet and dry format using radiation generated by a commercially available Signify ultraviolet (UV)-C light source at 254 nm. We show that for contaminated surfaces, only seconds of exposure is required for complete inactivation, allowing for easy implementation in decontamination workflows.
Similar content being viewed by others
Introduction
Towards the end of 2019, an outbreak of life-threatening pneumonia caused by a novel betacoronavirus occurred in the Hubei Province of China1. The virus, named severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), has since spread across the world at an alarming rate to cause a debilitating and ongoing pandemic, with only a few islands not reporting any cases to date. While SARS-CoV-2 is thought to be of zoonotic origin2, intense and extensive human-to-human transmission has mainly been driven by the inhalation of respiratory droplets and virus-bearing particles spread through the air3, or by contact with surfaces contaminated with settled droplets4. Although academic institutions and pharmaceutical organizations worldwide have banded together to develop countermeasures against the virus, there are still no licensed vaccines or therapeutics available. The disruption of transmission chains is therefore crucial for managing the outbreak and preventing additional infections.
Ultraviolet (UV) irradiation is an extensively tested, widely used and effective no-contact method for inactivating viral pathogens5,6,7. There are three types of UV, including UV-A (315–400 nm), UV-B (280–315 nm) and UV-C (100–280 nm), of which UV-C is most commonly employed in germicidal applications. At a wavelength of 254 nm, viral inactivation can be attributed to direct UV-C light absorption and photochemical damage to nucleic acid, leading to the disruption of viral replication8. Despite its wide use, limited data exists on the effectiveness of UV-C on inactivating wet and dried SARS-CoV-2 on contaminated surfaces. In particular, the efficacy of UV-C for inactivating SARS-CoV-2 in fluids needs to be determined, as the UV absorbance characteristics of fluid constituents may influence the dose required to achieve complete viral inactivation.
In this paper, we describe the complete and rapid inactivation of SARS-CoV-2 in both wet and dried droplets using 254 nm UV-C irradiation. Our results suggest that UV-C is an affordable and effective tool for preventing SARS-CoV-2 contact transmission that can easily be deployed to manage the coronavirus disease outbreak.
Results and discussion
Estimation of viral decay time
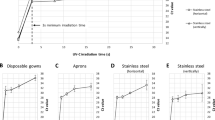
To examine the inactivation efficacy of UV-C on SARS-CoV-2, virus was applied to plastic tissue culture dishes and exposed to UV radiation as either wet or dried droplets for varying amounts of time ranging from 0.8 to 120 s. Under a UV-C irradiance of 0.849 mW/cm2, partial inactivation occurred from 0.8 s of exposure, while SARS-CoV-2 virus infectivity was reduced to below detectable levels in as few as 9 s for dried virus (Table 1; Fig. 1A) and 4 s for wet virus (Table 1; Fig. 1B).
Reduction in infectivity of SARS-CoV-2 after exposure to UV-C irradiation. The virus was exposed to UV-C as dried droplets (A) or wet droplets (B). Each set of data (dry samples and wet samples) shows a decrease of the remaining infectivity as a function of time, normalized to 1. Blue lines indicate single exponential decay functions while red lines indicate double exponential decay functions.
Virus inactivation by UV light is expected to be an exponential process9. Therefore, to estimate the decay time, we used linear regression methods with single and double exponential decay functions (Fig. 1). The single exponential decay function has the form \({\mathrm{y}}= {\mathrm{e}}^{-t/\uptau }\), while the double exponential function has the form \(\mathrm{y}= {\left(1-f\right)\mathrm{e}}^{-t/{\uptau }_{1}}+f{\mathrm{e}}^{-t/{\uptau }_{2}}\). τ, τ1 and τ2 are the decay times of the linear regressions. In case of double exponential decay, f is the fraction of the viruses that survive the first decay. For the analysis, data points were normalized so that the initial condition t = 0 corresponds to 100% infectivity with no irradiance.
In the linear regression of dried droplets, the reduced χ2 for double exponential decay (0.36) was lower than the one corresponding to single exponential decay (0.52). The R2 for double exponential was higher than the R2 of the single exponential. Hence, we used the double exponential decay to estimate the decay times, obtaining \({\uptau }_{1}=0.48\pm 0.09\mathrm{ s}\), and \({\uptau }_{2}= 1.60\pm 1.17\mathrm{ s}\).
In case of wet droplets, we observed the opposite: the χ2 for the double exponential (1.0) was higher than the one corresponding to the single exponential (0.8). The R2 for the double exponential and the single exponential was the same (0.9). We therefore used the single exponential decay as a best fit of the data to estimate the decay time, translating into an average decay time of \(\uptau =1.0\pm 0.1\mathrm{ s}\). Within one standard deviation, the decay times of wet and dried droplets are congruent. This is most likely due to the limited resolution of the measurements. In addition, this indicates that given the observation limits, UV-C absorption by media constituents did not significantly affect virus inactivation at a wavelength of 254 nm.
It should be noted that the experiments for this study were performed under specific and controlled conditions. Factors such as humidity, textured surfaces and the presence of dust and other particles may reduce the effectiveness of UV-C and influence the dose required to achieve complete viral inactivation6. It is also important to consider the composition of respiratory droplets when evaluating the effectiveness of 254 nm UV-C irradiation. Droplets are likely to be in solvent with a variety of other biological fluids such as respiratory mucus (phlegm) which may include viral glycoproteins, and UV-C absorption of these fluids and particles may result in a reduction in viral inactivation efficiencies. The results obtained in this study should therefore be interpreted as the minimum dose of radiation required to achieve viral inactivation.
A recent study looked at the combined effectiveness of UV-A/UV-C irradiation of SARS-CoV-2 in comparison to UV-A or UV-C alone10. The study found that UV-A was a poor means of inactivating SARS-CoV-2 in comparison to inactivation by 1.94 mW/cm2 UV-C. Of note, our results were generated using a lower dose of UV-C (0.849 mW/cm2) and a quicker time (4 – 9 s rather than 9 min with UV-C alone). This supports our observations showing the minimum required UV-C irradiance and time needed for complete SARS-CoV-2 inactivation. Another recent study investigated the effectiveness of light-emitting diode (LED) UV-C irradiation (280 nm)11 in inactivating SARS-CoV-2 in wet droplets. Similar to our findings, these results indicate inactivation of wet virus within 10 s at a UV dose of 3.75 mW/cm2. Using linear regression methods of analysis, our data provides a comprehensive overview of viral decay as a function of time for both wet (4 s) and dried (9 s) virus. Given the extended survival of SARS-CoV-2 on surfaces such as bank notes, glass and stainless steel14, the ability of countermeasures such as UV devices to inactivate virus on previously contaminated surfaces is crucial and was demonstrated using dried virus in our study.
Although it was beyond the scope of this study, future studies should address the effects of humidity, surface conformation and the natural matrices in which the virus may exist on the required UV-C inactivation doses. Direct exposure of the skin and eyes to 254 nm UV-C light can present a serious health hazard, such as corneal irritation and burns12,13 and as such, 254 nm UV-C light should only be used with proper training or where people are not at risk of being exposed. Forthcoming studies will explore the viral inactivation effectiveness of far UV-C wavelengths (207–222 nm) that have been proposed to be a safer alternative to 254 nm UV-C light7.
Although several techniques exist for inactivating SARS-CoV-2, the lack of proven effective tools and interventions have allowed for the unmanageable spread of the virus in the human population. Our results show that UV-C is a powerful tool that can be applied extensively in a wide range of public institutions including hospitals, nursing homes, workplaces, schools, airports and shopping centers to disinfect contaminated equipment and surfaces to prevent and reduce SARS-CoV-2 contact transmission.
Methods
UV-C device
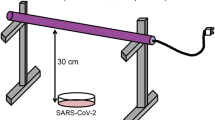
A test apparatus was designed, optimized, fabricated and calibrated to enable accurate and controlled UV-C treatment of test samples (Fig. 2). A collimated beam setup was fabricated based on a dual chamber construction. The top chamber contains the UV-C light source, the electronic driver, and a shutter system to control the exposure times of the samples while keeping the lamp output stable.
Samples were treated in the bottom chamber using deep UV-C light generated with a classical Mercury type TUV PLL 35 W light source, generating a peak wavelength at 254 nm. Multiple sensor-based safety measures were applied to protect the user against incidental exposure to the UV-C radiation. The irradiance level for three different lamps inside the treatment chamber was measured using a calibrated UV-C sensor system (Spectroradiometer GL Optic Spectis 5.0 Touch with detector GL Opti Probe 5.1.50), which provided irradiance patterns and levels shown in Fig. 3 from which optimal treatment locations could be deduced.
Virus inactivation procedures
All experiments were performed in the biosafety level 4 laboratory of the National Emerging Infectious Diseases Laboratories of Boston University. A volume of 100 µl SARS-CoV-2 (7.33 × 103 PFU/ml) (USA/WA1-2020)15 was plated onto the surface of 60 mm plastic tissue culture dishes (TPP) in 5 µl aliquots. The virus was allowed to dry for approximately 2 h on a subset of the dishes while the rest were processed immediately in the prototype UV-C device. Briefly, a pair of dishes (one to be treated and one control wrapped tightly in aluminum foil) were placed in the center of the device at an irradiance level of 0.849 mW/cm2, and towards the side of the UV-C device, respectively. The dishes were UV-C-treated for either 0.8, 2, 3, 4, 5, 6, 9, 15, 30 or 120 s, with each treatment time tested in triplicate. Dishes containing dried virus were treated in the same manner. Following treatment, the wet and dried virus were resuspended in 1.9 ml or 2 ml, respectively, of high glucose Dulbecco's Modified Eagle Medium (DMEM)(Gibco) containing 0.04 mM phenol red, 1 × antibiotic–antimycotic (Gibco), 1 × non-essential amino acids (Gibco), 1 × GlutaMAX-I (Gibco), 1 mM sodium pyruvate (Gibco) and 2% fetal bovine serum (FBS)(Gibco). The resuspended virus was then serially diluted from 1 × 100 to 1 × 10–2.5 using half-logarithmic dilutions. A back-titration of the virus was included for each experiment.
Confirmation of virus inactivation by plaque assay
Vero E6 cells maintained in high glucose DMEM (Gibco) supplemented with 1 × GlutaMAX-I, 1 mM sodium pyruvate, 10% FBS (Gibco) and 1 × non-essential amino acids (Gibco) were seeded into 6-well CellBIND plates (Corning) at a density of 8.0 × 105 cells per well. The cells were incubated at 37 °C and 5% CO2 overnight. The media was removed from each well and 200 µl of each dilution prepared from resuspended virus was added to the respective wells of a 6-well plate. One well containing only DMEM with 2% FBS was included as a control on each plate. A back-titer of the virus used to prepare the 60 mm dishes was performed in triplicate by inoculating each well of a 6-well plate with 1 × 10–2 to 1 × 10–6 dilutions of the virus, respectively. Plates were incubated at 37 °C and 5% CO2 for 1 h with intermittent rocking. Cells were then overlaid with 2 ml of a 1:1 solution of 2.5% Avicel RC-591 (DuPont Nutrition and Health) and 2 × Temin's Modified Eagle Medium (Gibco) without phenol red, supplemented with 10% FBS (Gibco), 2 × antibiotic–antimycotic (Gibco) and 2 × GlutaMAX-I (Gibco). The cells were incubated at 37 °C and 5% CO2 for 2 days. Plates were fixed in 10% neutral buffered formalin (ThermoFisher Scientific), followed by staining with 0.2% Gentian Violet (Ricca Chemical) in 10% neutral buffered formalin. The number of plaques per virus dilution were determined by eye and used to calculate the titer of the virus using the following formula:
Statistical analysis
The statistical package Microcal Origin was used to analyze the data. A detailed explanation of the statistical methods used is provided with the results.
Data availability
Additional data supporting the findings of this study are available from the corresponding author upon request.
References
Zhu, N. et al. A novel coronavirus from patients with pneumonia in China, 2019. N Engl J Med 382, 727–733. https://doi.org/10.1056/NEJMoa2001017 (2020).
Zhou, P. et al. A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 579, 270–273. https://doi.org/10.1038/s41586-020-2012-7 (2020).
Zhang, R. et al. Identifying airborne transmission as the dominant route for the spread of COVID-19. Proc Natl Acad Sci 117, 14857–14863. https://doi.org/10.1073/pnas.2009637117 (2020).
Chia, P. Y. et al. Detection of air and surface contamination by SARS-CoV-2 in hospital rooms of infected patients. Nat Commun 11, 2800. https://doi.org/10.1038/s41467-020-16670-2 (2020).
Darnell, M. E. R. et al. Inactivation of the coronavirus that induces severe acute respiratory syndrome SARS-CoV. J Virol Methods 121, 85–91. https://doi.org/10.1016/j.jviromet.2004.06.00 (2004).
McDevitt, J. J. et al. Aerosol susceptibility of influenza virus to UV-C light. Appl Environ Microbiol 78, 1666–1669. https://doi.org/10.1128/AEM.06960-11 (2012).
Buonanno, M. et al. Far-UVC light (222 nm) efficiently and safely inactivates airborne human coronaviruses. Sci Rep 10, 10285. https://doi.org/10.1038/s41598-020-67211-2 (2020).
Ariza-Mateos, A. et al. RNA self-cleavage activated by ultraviolet light-induced oxidation. Nucleic Acid Res 40, 1748–1766. https://doi.org/10.1093/nar/gkr822 (2012).
Kowalski, W. J. et al. Mathematical modeling of ultraviolet germicidal irradiation for air disinfection. Quant Microbiol 2, 249–270. https://doi.org/10.1023/A:1013951313398 (2002).
Heilingloh, C. S. et al. Susceptibility of SARS-CoV-2 to UV irradiation. Am J Infect Control 48, 1273–1275. https://doi.org/10.1016/j.ajic.2020.07.031 (2020).
Inagaki, I. et al. Rapid inactivation of SARS-CoV-2 with deep-UV LED irradiation. Emerg Microbes Infect 9, 1744–1747. https://doi.org/10.1080/22221751.2020.1796529 (2020).
Zaffina, S. et al. Accidental exposure to UV radiation produced by germicidal lamp: case report and risk assessment. Photochem Photobiol 88, 1001–1004. https://doi.org/10.1111/j.1751-1097.2012.01151.x (2012).
Trevisan, A. et al. Unusual high exposure to ultraviolet-C radiation. Photochem Photobiol 82, 1077–1079. https://doi.org/10.1562/2005-10-27-RA-728 (2008).
Riddell, S. et al. The effect of temperature on persistence of SARS-CoV-2 on common surfaces. Virol J 17, 145. https://doi.org/10.1186/s12985-020-01418-7 (2020).
Harcourt, J. et al. Severe acute respiratory syndrome coronavirus 2 from patient with coronavirus disease, United States. Emerg Infect Dis 26, 1266–1273. https://doi.org/10.3201/eid2606.200516 (2020).
Acknowledgements
The authors acknowledge Lauren E. Malsick for technical assistance. This work was supported by Signify Research.
Author information
Authors and Affiliations
Contributions
A.G., S.N.D., N.S. and L.G.A.M. conceptualized the study design. S.N.D., N.S., L.G.A.M. and R.I.J. performed the experiments. N.S., S.N.D., L.G.A.M., G.C. and A.G. analyzed the data and wrote the first draft of the paper with input from D.B., M.d.S. and W.W. All authors read, revised and approved the final paper.
Corresponding author
Ethics declarations
Competing interests
AG, NS, LGAM, SND, and RIJ have no competing interests to declare. MdS, GC, DB, and WW are employees of Signify Research.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Storm, N., McKay, L.G.A., Downs, S.N. et al. Rapid and complete inactivation of SARS-CoV-2 by ultraviolet-C irradiation. Sci Rep 10, 22421 (2020). https://doi.org/10.1038/s41598-020-79600-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-79600-8
This article is cited by
-
Photoinactivation of the bacteriophage PhiX174 by UVA radiation and visible light in SM buffer and DMEM-F12
BMC Research Notes (2024)
-
Estimation of the UV susceptibility of aerosolized SARS-CoV-2 to 254 nm irradiation using CFD-based room disinfection simulations
Scientific Reports (2024)
-
Germicidal efficacy of continuous and pulsed ultraviolet-C radiation on pathogen models and SARS-CoV-2
Photochemical & Photobiological Sciences (2024)
-
Wavelength dependence of ultraviolet light inactivation for SARS-CoV-2 omicron variants
Scientific Reports (2023)
-
Ultrafast inactivation of SARS-CoV-2 by 254-nm UV-C irradiation on porous and non-porous media of medical interest using an omnidirectional chamber
Scientific Reports (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.