Abstract
Neuromedin U (NmU) is a neuropeptide regulating diverse physiological processes. The insect homologs of vertebrate NmU are categorized as PRXamide family peptides due to their conserved C-terminal end. However, NmU homologs have been elusive in Mollusca, the second largest phylum in the animal kingdom. Here we report the first molluscan NmU/PRXamide receptor from the slug, Deroceras reticulatum. Two splicing variants of the receptor gene were functionally expressed and tested for binding with ten endogenous peptides from the slug and some insect PRXamide and vertebrate NmU peptides. Three heptapeptides (QPPLPRYa, QPPVPRYa and AVPRPRIa) triggered significant activation of the receptors, suggesting that they are true ligands for the NmU/PRXamide receptor in the slug. Synthetic peptides with structural modifications at different amino acid positions provided important insights on the core moiety of the active peptides. One receptor variant always exhibited higher binding activity than the other variant. The NmU-encoding genes were highly expressed in the slug brain, while the receptor gene was expressed at lower levels in general with relatively higher expression levels in both the brain and foot. Injection of the bioactive peptides into slugs triggered defensive behavior such as copious mucus secretion and a range of other anomalous behaviors including immobilization, suggesting their role in important physiological functions.
Similar content being viewed by others
Introduction
Neuromedin U (NmU) is a neuropeptide, identified initially from the porcine spinal cord1. Since then, NmU has been found in many vertebrates and shown to regulate diverse physiological processes including smooth-muscle contraction, blood pressure, local blood flow, ion transport in the gut, stress responses, gastric acid secretion and feeding behavior in vertebrates2,3. All NmU peptides isolated from vertebrates have a highly conserved C-terminal sequence (-PRNamide), which is closely related to the C-terminal sequence (-PRXamide, X indicates a variable amino acid) of insect PRXamide family peptides and invertebrate orthologs of NmU4.
Several mammalian NmU receptors (NmU-Rs) have been deorphanized and characterized in the gut and central nervous system (CNS) and found to be involved in feeding behavior5,6,7,8. Genetic screening of the Drosophila genome found that G protein-coupled receptors (GPCRs) (CG8784, CG8795, CG9918, and CG14575) are homologs of the vertebrate NmU-Rs9,10. These GPCRs have been identified as receptors for insect PRXamide family peptides, including pyrokinins (PKs) and CAPA peptides11,12,13,14. Upon the peptides binding to their corresponding GPCRs, G-proteins coupled are activated to open a ligand-gated calcium channel to allow the influx of extracellular Ca2+ as a second messenger, triggering a cascade of signal transduction for essential cellular processes10,15. These neuropeptides are involved in feeding behavior, muscle contraction, and fluid secretion in the digestive organs16,17,18,19. Four NmU-R homologs of the nematode Caenorhabditis elegans have also been identified as receptors for a nematode PRXamide20.
Mollusca, which includes snails and slugs, is the second largest phylum in terms of species diversity in the animal kingdom, and together with Arthropoda, Annelida, and Nematoda makes up the Protostomia clade. From an evolutionary viewpoint, therefore, the molluscan components are crucial to infer invertebrate evolution. The first Molluscan NmU/PRXamide family peptides were identified in the terrestrial snail, Helix lucorum21. However, no molluscan receptors for NmU/PRXamide family peptides have been characterized yet, although a variety of neuropeptides and physiological signals have been extensively studied in sea slugs in the genus Aplysia. The slug species, Deroceras reticulatum (Müller) (Gastropoda: Pulmonata), is a terrestrial slug native to Europe with global distribution22 that causes serious damage in a wide range of vegetables and field crops23. Unlike shelled snails, the terrestrial slugs including D. reticulatum, are highly sensitive to desiccation, sometimes showing a huddling behavior with a contracted posture when humidity is low24,25. Recently, we mined the slug transcriptome to develop a comprehensive list of neuropeptide genes including those that encode neuropeptides with a PRXamide at the C-termini26.
In this study, we identified two splicing variants of a slug receptor gene that respond to the NmU/PRXamide peptides from D. reticulatum. The receptors were heterologously expressed and tested with a variety of natural and synthetic peptide ligands, not only deorphanizing the receptors, but also characterizing the essential peptide moieties for receptor binding. This is the first report on the molluscan NmU/PRXamide receptor and its true ligands. It sheds light on the molecular evolution of the signaling system between vertebrate NmU and invertebrate PRXamide neuropeptides in animals.
Results
Identification of six genes encoding NmU-like peptides in slugs
Our previous study identified seven different genes encoding putative NmU/PRXamide peptides in the D. reticulatum transcriptome26. In the current study, their full-length genes were cloned by PCR, sequenced, and named myomodulin1, myomodulin2 and myomodulin3; pleurin1 and pleurin2; and small cardioaccelery peptides (sCAP1 and sCAP2) (GenBank Accession numbers: MW026240–MW026246). Twenty-six NmU/PRX peptides were predicted from the deduced amino acid sequences (Table 1, Fig. S1). The myomodulin1 gene produces 16 myomodulin-like peptides, including eight canonical Mmd peptides with a conserved MLRLamide(a) at their C-terminal ends (Table 1, Fig. S1). The PMSMLRLa was the most abundant with 7 out of 16 copies, and there were two copies for each of the QLSMLRLa and SLGMLRLa peptides. The myomodulin2 gene produces two myomodulin peptides, including a GQFSAARLa, a GLQMLRLa, and three NmU/PRX peptides, QPPLPRYa, FFFRPAPRGa, and AVPRPRIa. The myomodulin3 gene produces four NmU/PRXamide peptides, including two copies of QPPVPRYa, and single copies of the QPPLPRYa and SFFRPAPRGa peptides. The NmU/PRXamide peptides from the myomodulin2 and myomodulin3 genes have the same or very similar amino acid sequences.
The pleurin1 and pleurin2 genes produce six NmU/PRXamide peptides in total (Table 1, Fig. S1). They are composed of 14–15 amino acids and include four -PRIa peptides and two -PRVa peptides based on their C-terminal ends. The sCAP1 and sCAP2 genes produce four NmU/PRXamide peptides composed of nine amino acids, all with -PRMa C-terminal ends (Table 1, Figure S1). We selected eight representative peptides for the receptor-binding assays below.
Identification of the NmU receptor genes in slug
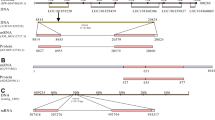
A BLAST search against the D. reticulatum transcriptome database was used to detect a 412-bp long transcript (c9058), which showed the highest homology to the vertebrate NmU receptors or the arthropod PRX receptors (E-value < 1e−59). RACE-PCR with gene-specific primers was used to amplify both ends of the gene, including 5′- and 3′-untranslated regions (UTRs). The full-length gene was then amplified from cDNA using conventional PCR and sequenced. Sequencing revealed two PCR amplicons representing two transcript variants, named DretNmU-Ra and DretNmU-Rb (GenBank Accession numbers: MW026247 and MW026248) (Fig. 1). When aligned with the slug genome sequences (unpublished data), the coding region of the receptor gene was found to contain five exons spanning about 9 Kb in the genome (Fig. 1A).
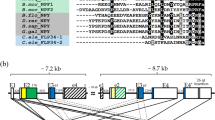
Structure of NmU receptor from Deroceras reticulatum. (A) Gene structure of the receptor composed of five exons, where two splicing variants (DretNmU-Ra and DreNmU-Rb) are derived. (B) Scheme of the two receptor variants, depicting the variable regions at the C-terminal intracellular domains. EC, extracellular (EC) domains. (C) Maximum-likelihood phylogenetic tree of the NmU receptor sequences from D. reticulatum (Dret), Aplysia californica (Ac), Drosophila melanogaster (Dm), Caenorhabditis elegans (Ce), and Homo sapiens (Hs). A human gonadotropin-releasing hormone receptor (GNRH-R) was used as an outgroup.
A comparison with the genome sequence revealed that the two transcript variants, -Ra and -Rb, seemed to be produced by alternative splicing events in exon 4. The DretNmU-Rb has 11 more nucleotides in exon 4 but 20 fewer nucleotides in exon 5 than DretNmU-Ra (Fig. 1A). A large intron gap (2250 bp) was found between exons 4 and 5 by PCR amplification using the specific primers for the two exons and genomic DNA. We confirmed the 11-bp retained in the variant -Rb and the resulting shift in the amino acid coding frame. Overall, DretNmU-Rb encodes a protein 3 aa-shorter (502 aa) than the variant -Ra (505 aa) (Fig. 1B, Supplementary data, Fig. S3). Protein sequence alignment by Clustal W algorithm and transmembrane (TM) domain prediction by TMHMM Server (v. 2.0, http://www.cbs.dtu.dk/services/TMHMM/) showed that the variable portion of these proteins is in the intracellular C-terminal domain, which is beyond the predicted seventh TM domain (Figs. 1B, S2). The phylogenetic tree of the NmU and PRX receptors indicated that the two slug receptors are close to Drosophila PK receptors and human NmU receptors (Fig. 1C).
Functional testing of receptors
Screening with peptide ligands
NmU/PRXamide ligands from slug (ten peptides), insect (eight peptides), and mammals (two peptides) were tested against the two slug receptors stably expressed in Sf9 cells. To determine the binding strength of the peptides to the receptors, fluorescence intensity induced by the calcium influx into the cell was measured (Fig. 2). In the initial tests using 500 nM concentration of each ligand, two slug peptides, QPPLPRYa and QPPVPRYa, evoked distinct binding to both variants, as did AVPRPRIa. However, the other PRXamide peptides from the slug failed to trigger a significant response to either receptor (Fig. 2). The insect PRX peptides and the mammalian NmU and FMRF peptides tested in this study failed to show any significant binding activities compared to the control (Fig. 2). Between the two receptor variants, variant Rb was always more responsive to the peptide ligands than variant A. The empty Sf9 control cells had no activity (Fig. 2).
Ligand screening of the binding activity of two receptor variants identified from Deroceras reticulatum. Peptides derived from various sources including the slug were assayed at 500 nM concentration against the Sf9 cell lines stably expressing either receptor variant Ra or Rb. Wild type Sf9 cells were also tested as a negative control. The mean response from four wells receiving the same treatment in the same plate was regarded as one replicate, and three to four replicates from independent assays on different cell plates were analyzed (mean + SEM). Peptide and species abbreviations: MM, myomodulin; NmU, neuromedinU; PK, pyrokinin; ETH, ecdysis triggering hormone; PBAN, pheromone biosynthesis activating neuropeptide; DH, diapause hormone; Dm, Drosophila melanogaster; Hz, Helicoverpa zea; Hh, Halyomorpha halys.
Responses of the natural peptides
The three most active peptides (QPPLPRYa, QPPVPRYa, and AVPRPRIa) were serially diluted and evaluated for dosage dependent binding to DretNmU-Ra and DretNmU-Rb (Fig. 3). They are all heptapeptides and share the -PRXamide. DretNmU-Rb was more active to the peptide ligands than DretNmU-Ra. The maximum binding activity of receptor variant Rb was twice that of receptor variant Ra for the peptide ligands tested, although the EC50 values of the receptor variant Ra is lower than those of the variant Rb.
Normalized concentration-responses and EC50 values (nM) of the two NmU receptor variants, DretNmU-Ra (A) and DretNmU-Rb (B), from Deroceras reticulatum to the three bioactive peptides that elicited high responses in the screening experiment. Data represent the mean ± SEM response of cells from four or five independent experiments. The mean response from four wells receiving the same treatment in the same plate was regarded as one replicate, and four to five replicates from independent assays on different cell plates were analyzed (mean ± SEM).
Responses of the modified peptides
To determine the core amino acids in the peptide ligand, the most active peptide (QPPLPRYa) was modified by replacing different amino acids or by removing one amino acid from the N-terminus. The various peptides were synthesized and tested at 500 nM concentration for binding to DretNmU-Ra and -Rb (Fig. 4). The glutamine (Q) at the N-terminus of the peptide was replaced with asparagine (N), which has the same polar and uncharged R group, NPPLPRYa. The binding strength to both receptors was similar to the natural ligand. The tyrosine (Y) at the C-terminus of the peptide was replaced with phenylalanine (F) or tryptophan (W), which have a similar aromatic amino acid group, QPPLPRFa or QPPLPRWa. The QPPLPRFa peptide showed a similar binding response to the receptors as the original ligand, but the QPPLPRWa peptide was dramatically reduced in its binding activity (Fig. 4). Each of seven amino acids (Q1P2P3L4P5R6Y7) of the natural ligand was replaced in series with alanine (A). When the P5, R6, or Y7 were replaced with alanine (A), these modified peptides almost lost their ability to bind the receptors. Interestingly, when the other amino acids, Q1, P2, P3 or L4, were replaced with an A in the natural ligand, they were still able to bind the receptors. Finally, with serial deletions of amino acids from the N-terminus of the natural ligand a gradual decrease of the binding ability was observed until the pentapeptide, PLPRYa, but then all activity was lost going from the tetra-peptide (LPRYa) to the di-peptide (RYa) (Fig. 4).
Structure–activity test for two receptor variants identified from Deroceras reticulatum. Synthetic peptides modified at various positions were assayed at 500 nM concentration against the Sf9 cell lines stably expressing either the receptor variant Ra or Rb. Amino acid substitutions compared to the true peptide ligand sequence (QPPLPRYa) are highlighted in bold. N, asparagine; F, phenylalanine; W, tryptophan; A, alanine. The mean response from four wells receiving the same treatment in the same plate was regarded as one replicate, and three to four replicates from independent assays on different cell plates were analyzed (mean + SEM).
Gene expression analysis
NmU/PRXamide peptide-encoding gene expression
The peptide-encoding genes, including myomodulins, pleurins and sCAPs, tested in this study were highly expressed exclusively in the CNS, compared to the other tissues in the slug (Fig. 5). The myomodulin1 and myomodulin2 genes were expressed at relatively higher levels than the other peptide-encoding genes and at about 100-fold greater than myomodulin3. The expression level of pleurin1 was almost eightfold greater than pleurin2. The expression level of sCAP1 was twice that of sCAP2 in the CNS. Interestingly, the sCAP1 gene also had high expression levels in the buccal mass of the slug.
Expression analysis of the NmU/PRXamide peptide-encoding genes and the two receptor variants-encoding genes in different tissues of Deroceras reticulatum. Nine different tissues were dissected from adult slugs. Total RNAs were extracted from the tissues, RNA concentration was adjusted to 100 ng/μl, and cDNAs were synthesized from 1 μg of total RNAs. Quantitative real-time PCR (qRT-PCR) was conducted with cDNA templates from the different tissues using the SYBR Green method, and specificity was confirmed by melt curve analysis. Primer efficiencies were calculated from a cDNA template dilution series. The 26S proteasome regulatory subunit 8 (rpt6) was used as a reference gene. Three biological replicates per tissue and two technical replicates per reaction were analyzed. CNS, central nervous system.
Receptor gene expression
The receptor genes were also highly expressed in the CNS, followed by expression levels in the foot, whereas the expression levels were relatively lower in other tissues (Fig. 5). The receptor DretNmU-Rb was expressed at levels two to threefold greater than the -Ra in all tissues tested in general. Interestingly, the DretNmU-Rb receptor was highly expressed in the foot compared to the other tissues except for the CNS.
Bioassay of the peptides in the slug
Based on the functional tests and response to the receptors above, we selected the two most active peptide ligands, QPPLPRYa and QPPVPRYa, and investigated their in vivo effects on D. reticulatum. When the peptides were injected, the test slugs compared to the control slugs showed a series of atypical behaviors within 30 min: the slugs displayed very rapid crawling, flaring of the mantle, rapid retraction and extension of the ocular tentacles, hyperextension of the ocular tentacles, abnormal body shapes, and immobilization. In addition, after 3 days some slugs died (QPPLPRYa: dry conditions—100% mortality; wet conditions—50%; QPPVPRYa: dry conditions—50%; wet conditions—0%). Furthermore, the slugs injected with the peptides secreted large amounts of milky mucus, causing weight loss. None of the control slugs displayed these behaviors, other than secreting a small volume of milky mucus after injection. The treated slugs that were kept in a wet environment ultimately showed mild weight loss, while those kept under dry conditions, showed substantial weight loss (Fig. 6 and Supplementary_Data_2.mp4). The relative weight losses of the injected slugs over 3 days were greater than twice that of the control slugs treated with water and kept under wet or dry conditions.
Injection bioassays of the NmU/PRXamide peptides for Deroceras reticulatum. (A) Effects of two NmU/PRXamide peptides on the weight loss (mean + SEM) of the slug (n = 4). Injection of 10 nmol of the peptide dissoved in 5 μl water using a Hamilton syringe and resulted in a loss of weight in both wet and dry conditions, although the effect was more clearly demonstrated under dry conditions. Different letters denote significant differences (P < 0.05) between treatments and control determined by One-way ANOVA followed by Dunnett’s multiple comparisons test using GraphPad Prism version 8.0. (B) Injection of the peptide (either QPPLPRYa or QPPVPRYa) triggered a series of distinct, atypical behaviors such as initial rapid crawling, flaring of the mantle, repeated retraction and extension of the ocular tentacles, hyperextension of the ocular tentacles, abnormal body shapes, and secretion of copious milky mucus (see the video in Supplementary_Data_2.mp4). (a) water injection and (b) peptide injection.
Discussion
Neuromedin U (NmU) is a bioactive peptide originally isolated from the porcine spinal cord1 and is highly conserved among vertebrates, including humans3. Since the Drosophila genome has been released27, insect PRXamide family peptides and their receptors have been characterized as vertebrate NmU orthologs and NmU receptors (NmU-Rs) due to their sequence similarity4,9,10,11,28,29,30. In evolutionary lower animal groups such as nematodes, the NmU/PRXamide orthologs and receptors have also been identified from Caenorhabditis elegans20.
The first molluscan NmU and PRXamide orthologs have been reported in a terrestrial snail21. The gene structure and expression of the NmU and PRXamide orthologs in the terrestrial snail are similar to the slug pleurin1 and pleurin2 genes expressed in the central nervous system in our studies26 (Figs. 5 and S3). These peptides have also been considered as homologs of the peptides produced by the hugin, capa, and eth genes in Drosophila31. However, little is known concerning the molluscan receptors for the NmU and PRXamide family peptides. In this study, we utilized the slug transcriptome to identify a complete list of genes encoding NmU and PRXamide peptides and determined their ability to activate NmU-Rs identified in D. reticulatum. Then, we deorphanized the receptors with three natural ligands, QPPLPRYa, QPPVPRYa, and AVPRPRIa, encoded from two myomodulin genes. This is the first report on the molluscan NmU-R and its agonistic peptide ligands.
The slug NmU-Rs contain seven transmembrane (TM) domains, indicating that the receptors belong to the typical G protein-coupled receptor (GPCR) family. The conserved ERY motif found next to TM3 is modified to DRY in this receptor (Fig. S2). Two transcript variants (DretNmU-Ra and DretNmU-Rb) of the receptor gene are derived by alternative splicing events that occur at the interface between exon 4 and exon 5. Due to a frame shift within the 11-bp intron region in variant Rb, the two variants contain a different intracellular segment (about 40 aa) at their C-termini. As shown in the protein sequences, the two GPCRs are mostly conserved, except for their C-terminal intracellular end. The remaining protein sequence, including the seven TM domains and extracellular domains, are derived from exons 1 through 4, indicating the 3-dimensional structures and binding sites should be similar between two GPCR variants. The extracellular domains of GPCRs are critical for maintaining the binding pockets to receive their specific ligands. The GPCR structures of the insect PRXamide peptides have been characterized through molecular site-directed mutations, 3D-modeling, and exchanging extracellular domains for the binding interactions and signaling32,33,34,35,36,37.
The variable sequences of the intracellular C-terminal regions should not affect the GPCR 3D-structure. Therefore, the C-terminal intracellular part (40 aa) presumably contributes to the discrepancy between the ligand-receptor binding strength of the two GPCR variants. The intracellular domain at the C-terminal end is expected to activate G proteins, opening specific calcium channels necessary for the influx of Ca2+ as a secondary messenger, triggering a signal transduction cascade10,15. The responses of DretNmU-Rb to the ligands tested in this study were always more active (~ twofold) than those of the variant Ra. This suggests that the C-terminus intracellular domain may contribute to the conformational transition of the GPCR to activate G proteins and increase the Ca2+ influx, once the ligand binds to the receptor. In insect PBAN GPCRs, the C-terminus intracellular sequences are functionally significant for ligand-induced receptor internalization to regulate the strength and duration of receptor-mediated cell signaling38,39,40,41. Further studies on the intracellular components linked to GPCR signal transduction are needed to better understand these processes in slugs.
In arthropods, multiple PRXamide peptides are often encoded from the same gene, but generally members from different families of peptides are not encoded within a gene4,42,43. However, the slug myomodulin2 gene produces peptides from two different peptide families, Myomodulin (MM) (-MLRLa) and NmU/PRXamide (Table 1 and Fig. S1). This is similar to what has been observed in Aplysia where a single gene was found to encode not only the canonical MM peptides but also three NmU/PRXamide peptides (called MMG2-derived peptides; MMG2-DPs) including QPPLPRYa (MMG2-DPb), AVALPRIa (MMG2-DPd), and AVPRPRIa (MMG2-DPf)44. Interestingly, both QPPLPRYa and AVPRPRIa are considered natural ligands of the NmU receptor in D. reticulatum (Table 1, Fig. 3), suggesting these peptides are highly conserved at least in Gastropoda.
Although the myomodulin3 gene encodes only NmU/PRXamide peptides, its DNA sequence is similar to the myomodulin1 and myomodulin2 genes, although the myomodulin3 encodes all three NmU/PRXamide peptides in this study. The canonical MM peptides were originally isolated from a feeding muscle in Aplysia45 and the gene encoding the MM peptides was cloned later46,47. More myomodulin genes and peptides have been reported from molluscan species either from transcriptomes/genomes or from proteomics26,48,49.
Pleurin genes and sCAP genes produce three and two NmU/PRXamide peptides, respectively. Due to their high sequence similarity to insect PRXamide peptides, they have been characterized as orthologs of insect pyrokinin or CAPA peptides6,31,50,51. However, no functional studies have been reported yet. If additional receptors are identified from the slug, they could be deorphanized by these peptides.
The natural ligand’s structural modification stuides indicate that the C-terminal three amino acids (PRXamide) comprise a core motif for the ligand-receptor binding. When any of the three amino acids were replaced by the simplest non-polar and aliphatic amino acid alanine (A), as in QPPLPRAa, QPPLPAYa, and QPPLARYa, the receptor response to the ligands almost completely disappeared. When the C-terminal tyrosine (Y) was replaced by aromatic and non-polar phenylalanine (F) or tryptophan (W), QPPLPRFa retained normal activity to the receptors, but QPPLPRWa activity was eliminated. Interestingly, all three amino acids, Y, F, and W, have the aromatic functional group, and tryptophan (W) only contains an indole side chain that could interfere with the ligand–receptor binding. Additional structural–activity relationships developed through ligand modification or mimics will be helpful for finding the core residues in the ligands and developing potential targets for pest control52,53.
In contrast, the N-terminal glutamine (Q) in the peptide did not have an effect on ligand–receptor binding. For example, when Q was replaced with asparagine (N) or alanine (A), or just eliminated from the ligand, there was no significant decrease in the ligand-receptor binding activity. However, the second and fourth amino acids from the N-terminus seem to be more critical, because when they were replaced with alanine (A), binding activity decreased about 42–53% for the variant Rb and about 22–36% for the variant Ra, whereas the third position proline (P) between these two essential amino acids was not critical for binding activity.
A minimum peptide length appeared to be critical for ligand–receptor binding; at least five amino acids with the core motif (PRXamide) were found to be essential to activate the receptors. From the overall structural analyses, it seems that the core C-terminal three amino acids (PRXamide) and a minimum five amino acids are critical to activate the slug receptors and trigger signal transduction. It is also consistent in different animal groups in general4. More studies on peptide 3D modelling and additional structural relationships would help to elucidate the agonistic and/or antagonistic effect on the target receptor. It is noteworthy that the families of insect PRXamide peptides and the mammalian NmU peptide tested were not able to activate the slug receptors (Fig. 3). It would be very interesting to examine the activity of slug peptides with insect PRX-Rs or vertebrate NmU-Rs.
In vertebrates, the NmU peptides function in the activation of ion transport and contractile activity in intestinal and arterial musculature via NmU-R3. In insects, many PRXamide peptides have various functions, including heartbeat modulation, visceral muscle contraction, (anti)diuresis, pheromone biosynthesis, cuticle melanization, etc54,55,56,57,58. In Aplysia, immunocytochemistry studies on two MM peptides (original names, MMs and myomodulin gene2-derived peptides) indicate that the two myomodulin genes are differentially expressed in the buccal ganglion, the buccal musculature, and in the pedal ganglion. These organs are involved in multiple actions including feeding and muscle contractions, suggesting that these peptides play a broad role in feeding behavioral plasticity44. The gene expression studies of various organs in D. reticulatum showed that the active peptide-encoding genes (myomodulins and pleurins) are highly expressed in the CNS, whereas the receptor gene (DretNmU-Rb) was highly expressed in both CNS and foot. The results are similar to the studies of Aplysia.
In the injection bioassays, the NmU/PRXamide peptide triggered atypical behaviors in the slug, including rapid movement, abnormal body shapes, flaring of the mantle, and rapid retraction and extension of the ocular tentacles, which might be derived from excessive muscle contractions. This likely immoderate muscle contraction in the slug induced by the peptide is a similar physiological change observed in the insects above. It is noteworthy that treated specimens secreted copious amounts of milky mucus, which is a well-known defense response for D. reticulatum usually against physical attack59,60,61,62. This suggests that the pedal mucus gland could be an additional location for receptors, which active peptides could directly or indirectly target. It will be interesting to perform more detailed investigations to gain a better understanding of how slugs respond to the peptides and the underlying mechanisms involved, which will increase our knowledge of the physiological function of the peptides and aid in the development of novel pesticides against molluscan pests.
Methods
Slug
Deroceras reticulatum species were collected in the Willamette Valley, Oregon, USA, and maintained in a humid container with carrot and lettuce in a controlled incubator (18 °C, 90% RH, 12:12 = light:dark, dim light).
Nucleic acids isolation
Mucus was removed by scraping the surface of live slugs using sterilized forceps before homogenization. A mature slug was homogenized in ATL buffer provided in the QIAamp DNA Mini Kit (Qiagen, USA) using a PYREX glass pestle grinder (Corning, USA). The slimy homogenate was diluted with additional ATL buffer, further homogenized and the homogenate was pooled in a microcentrifuge tube. The genomic DNA was isolated using QIAamp DNA Mini Kit according to the manufacturer’s instructions, and stored at − 20 °C until use. Three mature slugs with mucus-removed were individually homogenized in 1 ml lysis buffer provided in the PureLink RNA Mini Kit (Invitrogen, USA) using a PYREX glass pestle as above. The slimy homogenate from the three samples was diluted with additional lysis buffer, further homogenized and pooled into a 15 ml-Falcon tube. Total RNA was extracted from the combined sample using the PureLink RNA Mini Kit following the manufacturer’s instructions, quantified on a NanoDrop2000 spectrophotometer (Thermo Fisher Scientific), and stored at − 80 °C.
cDNA synthesis
For the random amplification of the cDNA ends (RACE-PCR), 5′- and 3′-RACE-ready cDNAs were synthesized from 1 µg of total RNA (see above) using the SMARTer RACE 5′/3′ Kit (Clontech, USA) according to the manufacturer’s instructions. For conventional PCR, first-strand cDNAs were also synthesized from 1 µg of total RNA using the SuperScript IV First-Strand Synthesis System (Invitrogen) according to the manufacturer’s instructions. Both cDNAs were stored at − 20 °C until use.
BLAST search
We have previously reported seven PRXamide-encoding genes including myomodulin1, myomodulin2, myomodulin3 (partial), pleurin1, pleurin2, sCAP1, and sCAP2 in the D. reticulatum transcriptome26. For the NmU/PRXamide receptor genes, BLAST searches against the slug transcriptome found a partial transcript (c90580) that was most similar to the mammalian NmU-Rs and insect PK/CAPA receptors. The full-length sequences of these genes were amplified using a combination of RACE-PCR and conventional PCR, cloned, and sequenced. (see Supplementary Data 1 for detailed methods).
Expression vector construction
The full-length sequence of the receptor gene was PCR amplified from the previously cloned-pJET1.1 vector constructs using Phusion DNA Polymerase (Thermo Fisher Scientific) with a sense primer, including the Kozak sequence (GCCACCATGG; underlined ATG = start codon), and an anti-sense primer including the stop codon. PCR was performed under the following conditions: 98 °C for 30 s; 35 cycles of 98 °C for 10 s, 55 °C for 20 s, and 72 °C for 1 min; then 72 °C for 10 min. Primer sequences used in this study are listed in Table S1. PCR products were purified using the GeneJET Gel Extraction Kit (Thermo Fisher Scientific), and ligated into the pIB/V5-His TOPO-TA expression vector (Invitrogen). The vectors containing the receptor genes were sequenced and the orientation of the sequences was confirmed.
Expressing receptors in Sf9 cells
Sf9 cells were cultured in a 50 ml-Erlenmeyer flask while shaking at 145 rpm at 28 °C using Insectagro Sf9 Medium (Corning) supplemented with 5% heat-inactivated fetal bovine serum (HyClone, USA) and antibiotics (penicillin and streptomycin; Gibco). Three µg of either pIB/DretNmU-Ra or pIB/DretNmU-Rb plasmid was added to 100 μl of serum-free medium, and 15% CellFectin II (Invitrogen) in 100 μl serum-free medium was prepared, and both were placed at room temperature for 5 min. Samples were combined by pipetting and placed at room temperature for another 20 min. The plasmid-CellFectin II mixture was then added to a T-25 flask containing a layer of Sf9 cells covering about 70% of the surface, and incubated overnight. The next morning, the medium was replaced with fresh medium containing 50 μg/ml blasticidin. The medium was replaced daily with 50 μg/ml blasticidin containing medium for 1 week, and then the blasticidin concentration was gradually reduced to 30 μg/ml for the next 2 weeks. When culture was confluent, the recombinant cells were lifted and transferred to suspension cultures and grown until the exponential growth phase, and then stock cultures were frozen in liquid nitrogen. These frozen stock cell lines were thawed, cultivated for several passages in media containing 20 μg/ml blasticidin, and used for the ligand binding assays.
Peptide ligands
Analysis of the D. reticulatum transcriptome for neuropeptides26 revealed seven genes encoding putative PRXamide peptides (Table 1). For confirmation, the full-length sequences were PCR amplified, cloned, and Sanger sequenced (details above). In particular, the partial transcript of myomodulin2 gene was extended by RACE-PCR to identify 5′- and 3′-untranslated regions (see details above). With experimentally confirmed full-length sequences, the putative peptide sequences were predicted according to the mono- and dibasic cleavage site rules51. Among twenty-six different peptides amidated in their C-terminal ends, eight representative slug peptides were selected and synthesized by Peptide 2.0 (Chantily, USA). Several insect PRXamide peptides and vertebrate peptides (FMRF amide and Neuromedin U) were also synthesized for comparison.
Functional testing of receptors
Two days before the binding assay, ~ 50,000 Sf9 cells expressing the DretNmU-Ra or -Rb from suspension cultures were dispensed into each well of a black 96-well plate (Corning C3603) and incubated at 28 °C overnight. At 48 h, cells were rinsed once with 100 μl of fresh medium without FBS. After removing the media, cells were incubated with 95 μl of 1 × FLIPR Calcium 6 reagent (Molecular Devices, USA) containing 2.5 mM probenecid at room temperature in the dark for 1 h. The Calcium 6 reagent-loaded cells were transferred to the Flexstation 3 multi-mode microplate reader (Molecular Devices) to measure fluorescence change (∆fluorescence) before and after loading 5 μl of peptide ligands (20×), detecting emission at 535 nm with excitation at 485 nm. Fluorescence measurements from each well on the column were taken every 5 s for 4 min. The peptide ligand was added by automatic pipettor once at 30 s from the beginning, and 5 μl of 5 μM ionomycin was added at 3 min to confirm calcium activity. Baseline fluorescence was determined by averaging 5 time points from each well prior to treatment with ligand and the resulting response was expressed as a percent increase in fluorescence relative to the baseline value.
For the initial screening and structure–activity tests, each cell line was exposed to a single concentration of 500 nM of each peptide or control. For the dosage-response assay, threefold serial dilutions of selected ligands with water were used, giving final ligand concentrations between 8 pM and 1.5 μM. The mean response from four wells receiving the same treatment in the same plate was regarded as one replicate, and at least three replicates from independent assays on different cell plates were analyzed. Data were transformed into a log scale and fit into the non-linear regression curve. EC50 values were obtained by statistical analysis using the sum of squares F-test function using GraphPad Prism version 8.0 (San Diego, CA, USA).
Quantitative real-time PCR
Quantitative real-time PCR (qRT-PCR) was conducted using the SYBR Green method in a StepOnePlus Real-Time PCR System (Applied Biosystems). Nine different slug tissues, such as central nervous system (CNS), crop (without content), intestine (with digestive gland), buccal mass, ovotestis, bursa copulatrix (with albumen gland), renal duct, mantle and foot, were dissected, homogenized and extracted for total RNAs (see above) with three biological replicates. cDNA templates were synthesized from 1 μg of total RNA using Verso cDNA Synthesis Kit with oligo dT/random hexamer primers (ThermoFisher Scientific) according to the manufacturer’s instructions. The qRT-PCR reaction conditions were performed at 95 °C for 10 min; 40 cycles of 95 °C for 15 s and 60 °C for 1 min; followed by a melting curve analysis over the range of 60–95 °C with 0.3 °C/min increments, with the specific primers (Table S1). Six different concentrations of cDNA pool were used to construct a standard curve for each primer set to determine primer efficiency. Six candidate reference genes (β-cop, rpl3, rpl40, rpt6, vha26, and vps16) were evaluated for their primer efficiency and variability through pilot tests, then the 26S proteasome regulatory subunit 8 (rpt6) gene was selected as a reference gene (Table S2) (see Supplementary Data 1 for detailed methods).
Bioassay of the peptides in the slug
Two peptides, QPPLPRYa and QPPVPRYa, were dissolved in purified water to obtain a 2 mM concentration and individually injected into the adult slugs (270–520 mg in weight). A Hamilton syringe was used to inject a dose of 10 nmol peptide in 5 μl water or water only for control into the posterior part of the foot of the slug (n = 4). The foot was chosen as the injection site in order to avoid possible damage to internal organs, which are primarily located underneath the mantle. The injected slug was observed for an initial 30 min and notes were taken on its atypical behavior compared to the control, e.g. rapid movement, abnormal body shapes, flaring of the mantle, rapid retraction and extension of the ocular tentacles, and excessive milky mucus secretion. Since the treated slugs secreted a large amount of the defensive mucus, weight loss was assessed by weighing the slugs on a VWR B2-Series Analytical Balance immediately before the treatment and again 3 days later. Post treatment weights were obtained after slugs were placed under wet conditions (slugs placed in a ventilated 8 oz plastic container lined with damp tissue paper) or under dry conditions (same container type without damp tissue paper). Slugs were placed under these conditions approximately 30 min after injection. These plastic containers were maintained in a growth chamber (Thermo Scientific Precision Model 818) at 18 °C and 12 h photoperiod. Statistical analysis was performed by One-way ANOVA followed by Dunnett’s multiple comparisons test using GraphPad Prism version 8.0 (GraphPad Software, CA, USA).
References
Minamino, N., Kangawa, K. & Matsuo, H. Neuromedin U-8 and U-25: Novel uterus stimulating and hypertensive peptides identified in porcine spinal cord. Biochem. Biophys. Res. Commun. 130, 1078–1085. https://doi.org/10.1016/0006-291x(85)91726-7 (1985).
Ivanov, T. R., Lawrence, C. B., Stanley, P. J. & Luckman, S. M. Evaluation of neuromedin U actions in energy homeostasis and pituitary function. Endocrinology 143, 3813–3821. https://doi.org/10.1210/en.2002-220121 (2002).
Brighton, P. J., Szekeres, P. G. & Willars, G. B. Neuromedin U and its receptors: Structure, function, and physiological roles. Pharmacol. Rev. 56, 231–248. https://doi.org/10.1124/pr.56.2.3 (2004).
Jurenka, R. The PRXamide neuropeptide signalling system: conserved in animals. Adv. Insect Physiol. 49, 123–170. https://doi.org/10.1016/bs.aiip.2015.07.001 (2015).
Howard, A. D. et al. Identification of receptors for neuromedin U and its role in feeding. Nature 406, 70–74. https://doi.org/10.1038/35017610 (2000).
Fujii, R. et al. Identification of neuromedin U as the cognate ligand of the orphan G protein-coupled receptor FM-3. J. Biol. Chem. 275, 21068–21074. https://doi.org/10.1074/jbc.M001546200 (2000).
Kojima, M. et al. Purification and identification of neuromedin U as an endogenous ligand for an orphan receptor GPR66 (FM3). Biochem. Biophys. Res. Commun. 276, 435–438. https://doi.org/10.1006/bbrc.2000.3502 (2000).
Szekeres, P. G. et al. Neuromedin U is a potent agonist at the orphan G protein-coupled receptor FM3. J. Biol. Chem. 275, 20247–20250. https://doi.org/10.1074/jbc.C000244200 (2000).
Hewes, R. S. & Taghert, P. H. Neuropeptides and neuropeptide receptors in the Drosophila melanogaster genome. Genome Res. 11, 1126–1142. https://doi.org/10.1101/gr.169901 (2001).
Broeck, J. V. Insect G protein-coupled receptors and signal transduction. Arch. Insect Biochem. Physiol. 48, 1–12. https://doi.org/10.1002/arch.1054 (2001).
Park, Y., Kim, Y. J. & Adams, M. E. Identification of G protein-coupled receptors for Drosophila PRXamide peptides, CCAP, corazonin, and AKH supports a theory of ligand-receptor coevolution. Proc. Natl. Acad. Sci. USA 99, 11423–11428. https://doi.org/10.1073/pnas.162276199 (2002).
Rosenkilde, C. et al. Molecular cloning, functional expression, and gene silencing of two Drosophila receptors for the Drosophila neuropeptide pyrokinin-2. Biochem. Biophys. Res. Commun. 309, 485–494. https://doi.org/10.1016/j.bbrc.2003.08.022 (2003).
Cazzamali, G., Torp, M., Hauser, F., Williamson, M. & Grimmelikhuijzen, C. J. The Drosophila gene CG9918 codes for a pyrokinin-1 receptor. Biochem. Biophys. Res. Commun. 335, 14–19. https://doi.org/10.1016/j.bbrc.2005.07.038 (2005).
Iversen, A., Cazzamali, G., Williamson, M., Hauser, F. & Grimmelikhuijzen, C. J. Molecular cloning and functional expression of a Drosophila receptor for the neuropeptides capa-1 and -2. Biochem. Biophys. Res. Commun. 299, 628–633. https://doi.org/10.1016/S0006-291X(02)02709-2 (2002).
Iismaa, T. P., Biden, T. J. & Shine, J. G Protein-Coupled Receptors (R.G. Landes Co., 1995).
Melcher, C. & Pankratz, M. J. Candidate gustatory interneurons modulating feeding behavior in the Drosophila brain. PLoS Biol. 3, e305. https://doi.org/10.1371/journal.pbio.0030305 (2005).
Terhzaz, S. et al. Mechanism and function of Drosophila capa GPCR: a desiccation stress-responsive receptor with functional homology to human neuromedinU receptor. PLoS ONE 7, e29897. https://doi.org/10.1371/journal.pone.0029897 (2012).
Bader, R. et al. Genetic dissection of neural circuit anatomy underlying feeding behavior in Drosophila: Distinct classes of hugin-expressing neurons. J. Comp. Neurol. 502, 848–856. https://doi.org/10.1002/cne.21342 (2007).
Audsley, N. & Weaver, R. J. Neuropeptides associated with the regulation of feeding in insects. Gen. Comp. Endocrinol. 162, 93–104. https://doi.org/10.1016/j.ygcen.2008.08.003 (2009).
Lindemans, M. et al. A neuromedin-pyrokinin-like neuropeptide signaling system in Caenorhabditis elegans. Biochem. Biophys. Res. Commun. 379, 760–764. https://doi.org/10.1016/j.bbrc.2008.12.121 (2009).
Bogdanov, Y. D., Balaban, P. M., Poteryaev, D. A., Zakharov, I. S. & Belyavsky, A. V. Putative neuropeptides and an EF-hand motif region are encoded by a novel gene expressed in the four giant interneurons of the terrestrial snail. Neuroscience 85, 637–647. https://doi.org/10.1016/s0306-4522(97)00561-7 (1998).
Tulli, M. C. et al. Predation on the slug Deroceras reticulatum (pulmonata: Stylommatophora) by Scarites anthracinus (Coleoptera: Carabidae). Ecol. Austral. 19, 55–61 (2009).
South, A. Terrestrial Slugs: Biology, Ecology and Control (Springer, Netherlands, 1992).
Waite, T. A. Huddling and postural adjustments in response to desiccating conditions in Deroceras reticulatum (Müller). J. Molluscan Stud. 54, 249–250. https://doi.org/10.1093/mollus/54.2.249 (1988).
Arias, R. O. The biology of the grey garden slug Deroceras reticulatum (Müller). Master thesis (Oregon State University, 1960).
Ahn, S. J., Martin, R., Rao, S. & Choi, M. Y. Neuropeptides predicted from the transcriptome analysis of the gray garden slug Deroceras reticulatum. Peptides 93, 51–65. https://doi.org/10.1016/j.peptides.2017.05.005 (2017).
Adams, M. D. et al. The genome sequence of Drosophila melanogaster. Science 287, 2185–2195. https://doi.org/10.1126/science.287.5461.2185 (2000).
Melcher, C., Bader, R., Walther, S., Simakov, O. & Pankratz, M. J. Neuromedin U and its putative Drosophila homolog hugin. PLoS Biol. 4, e68. https://doi.org/10.1371/journal.pbio.0040068 (2006).
Choi, M. Y., Fuerst, E. J., Rafaeli, A. & Jurenka, R. Identification of a G protein-coupled receptor for pheromone biosynthesis activating neuropeptide from pheromone glands of the moth Helicoverpa zea. Proc. Natl. Acad. Sci. USA 100, 9721–9726. https://doi.org/10.1073/pnas.1632485100 (2003).
Park, Y., Kim, Y. J., Dupriez, V. & Adams, M. E. Two subtypes of ecdysis-triggering hormone receptor in Drosophila melanogaster. J. Biol. Chem. 278, 17710–17715. https://doi.org/10.1074/jbc.M301119200 (2003).
Ierusalimsky, V. N. & Balaban, P. M. Neuropeptides of Drosophila related to molluscan neuropeptides: Dependence of the immunoreactivity pattern on the ontogenetic stage and functional state. Brain Res. 1152, 32–41. https://doi.org/10.1016/j.brainres.2007.03.038 (2007).
Stern, P. S. et al. Molecular modeling of the binding of pheromone biosynthesis activating neuropeptide to its receptor. J. Insect Physiol. 53, 803–818. https://doi.org/10.1016/j.jinsphys.2007.03.012 (2007).
Choi, M. Y. & Jurenka, R. A. Site-directed mutagenesis and PBAN activation of the Helicoverpa zea PBAN-receptor. FEBS Lett. 584, 1212–1216. https://doi.org/10.1016/j.febslet.2010.02.033 (2010).
Choi, M. Y., Fuerst, E. J., Rafaeli, A. & Jurenka, R. Role of extracellular domains in PBAN/pyrokinin GPCRs from insects using chimera receptors. Insect Biochem. Mol. Biol. 37, 296–306. https://doi.org/10.1016/j.ibmb.2006.12.004 (2007).
Jurenka, R. & Nusawardani, T. The pyrokinin/ pheromone biosynthesis-activating neuropeptide (PBAN) family of peptides and their receptors in Insecta: Evolutionary trace indicates potential receptor ligand-binding domains. Insect Mol. Biol. 20, 323–334. https://doi.org/10.1111/j.1365-2583.2010.01065.x (2011).
Hariton-Shalev, A., Shalev, M., Adir, N., Belausov, E. & Altstein, M. Structural and functional differences between pheromonotropic and melanotropic PK/PBAN receptors. Biochim. Biophys. Acta 1830, 5036–5048. https://doi.org/10.1016/j.bbagen.2013.06.041 (2013).
Jiang, L. et al. Isolation and functional characterization of the pheromone biosynthesis activating neuropeptide receptor of Chinese oak silkworm, Antheraea pernyi. Int. J. Biol. Macromol. 117, 42–50. https://doi.org/10.1016/j.ijbiomac.2018.05.145 (2018).
Lee, J. M. et al. Re-Evaluation of the PBAN receptor molecule: characterization of PBANR variants expressed in the pheromone glands of moths. Front. Endocrinol. 3, 6. https://doi.org/10.3389/fendo.2012.00006 (2012).
Lee, J. M. et al. Establishment of Sf9 transformants constitutively expressing PBAN receptor variants: Application to functional evaluation. Front. Endocrinol. 3, 56. https://doi.org/10.3389/fendo.2012.00056 (2012).
Hull, J. J., Ohnishi, A. & Matsumoto, S. Regulatory mechanisms underlying pheromone biosynthesis activating neuropeptide (PBAN)-induced internalization of the Bombyx mori PBAN receptor. Biochem. Biophys. Res. Commun. 334, 69–78. https://doi.org/10.1016/j.bbrc.2005.06.050 (2005).
Hull, J. J., Lee, J. M. & Matsumoto, S. Identification of specific sites in the third intracellular loop and carboxyl terminus of the Bombyx mori pheromone biosynthesis activating neuropeptide receptor crucial for ligand-induced internalization. Insect Mol. Biol. 20, 801–811. https://doi.org/10.1111/j.1365-2583.2011.01110.x (2011).
Choi, M. Y., Sanscrainte, N. D., Estep, A. S., Vander Meer, R. K. & Becnel, J. J. Identification and expression of a new member of the pyrokinin/pban gene family in the sand fly Phlebotomus papatasi. J. Insect Physiol. 79, 55–62. https://doi.org/10.1016/j.jinsphys.2015.06.001 (2015).
Predel, R. & Wegener, C. Biology of the CAPA peptides in insects. Cell Mol. Life Sci. 63, 2477–2490. https://doi.org/10.1007/s00018-006-6187-3 (2006).
Proekt, A. et al. Identification of a new neuropeptide precursor reveals a novel source of extrinsic modulation in the feeding system of Aplysia. J. Neurosci. 25, 9637–9648. https://doi.org/10.1523/JNEUROSCI.2932-05.2005 (2005).
Cropper, E. C., Tenenbaum, R., Kolks, M. A., Kupfermann, I. & Weiss, K. R. Myomodulin: a bioactive neuropeptide present in an identified cholinergic buccal motor neuron of Aplysia. Proc. Natl. Acad. Sci. USA 84, 5483–5486. https://doi.org/10.1073/pnas.84.15.5483 (1987).
Lopez, V., Wickham, L. & Desgroseillers, L. Molecular cloning of myomodulin cDNA, a neuropeptide precursor gene expressed in neuron L10 of Aplysia californica. DNA Cell Biol. 12, 53–61. https://doi.org/10.1089/dna.1993.12.53 (1993).
Miller, M. W. et al. The myomodulin-related neuropeptides: characterization of a gene encoding a family of peptide cotransmitters in Aplysia. J. Neurosci. 13, 3358–3367. https://doi.org/10.1523/JNEUROSCI.13-08-03358.1993 (1993).
Greenberg, M. J. et al. Characterization of myomodulin-related peptides from the pulmonate snail Helix aspersa. Peptides 18, 1099–1106. https://doi.org/10.1016/s0196-9781(97)00149-6 (1997).
Kellett, E. et al. Myomodulin gene of Lymnaea: Structure, expression, and analysis of neuropeptides. J. Neurosci. 16, 4949–4957. https://doi.org/10.1523/JNEUROSCI.16-16-04949.1996 (1996).
Stewart, M. J. et al. Neuropeptides encoded by the genomes of the Akoya pearl oyster Pinctata fucata and Pacific oyster Crassostrea gigas: a bioinformatic and peptidomic survey. BMC Genomics 15, 840. https://doi.org/10.1186/1471-2164-15-840 (2014).
Veenstra, J. A. Neurohormones and neuropeptides encoded by the genome of Lottia gigantea, with reference to other mollusks and insects. Gen. Comp. Endocrinol. 167, 86–103. https://doi.org/10.1016/j.ygcen.2010.02.010 (2010).
Kim, Y. J., Nachman, R. J., Aimanova, K., Gill, S. & Adams, M. E. The pheromone biosynthesis activating neuropeptide (PBAN) receptor of Heliothis virescens: Identification, functional expression, and structure-activity relationships of ligand analogs. Peptides 29, 268–275. https://doi.org/10.1016/j.peptides.2007.12.001 (2008).
Jiang, H., Wei, Z., Nachman, R. J. & Park, Y. Molecular cloning and functional characterization of the diapause hormone receptor in the corn earworm Helicoverpa zea. Peptides 53, 243–249. https://doi.org/10.1016/j.peptides.2013.11.005 (2014).
Paluzzi, J. P., Park, Y., Nachman, R. J. & Orchard, I. Isolation, expression analysis, and functional characterization of the first antidiuretic hormone receptor in insects. Proc. Natl. Acad. Sci. USA 107, 10290–10295. https://doi.org/10.1073/pnas.1003666107 (2010).
Matsumoto, S. et al. Functional diversity of a neurohormone produced by the suboesophageal ganglion: Molecular identity of melanization and reddish colouration hormone and pheromone biosynthesis activating neuropeptide. J. Insect Physiol. 36, 427–432. https://doi.org/10.1016/0022-1910(90)90060-S (1990).
Raina, A. K. et al. Identification of a neuropeptide hormone that regulates sex pheromone production in female moths. Science 244, 796–798. https://doi.org/10.1126/science.244.4906.796 (1989).
Schoofs, L., Holman, G. M., Hayes, T. K., Nachman, R. J. & De Loof, A. Isolation, identification and synthesis of locustamyoinhibiting peptide (LOM-MIP), a novel biologically active neuropeptide from Locusta migratoria. Regul. Pept 36, 111–119. https://doi.org/10.1016/0167-0115(91)90199-Q (1991).
Predel, R., Kellner, R., Kaufmann, R., Penzlin, H. & Gade, G. Isolation and structural elucidation of two pyrokinins from the retrocerebral complex of the American cockroach. Peptides 18, 473–478. https://doi.org/10.1016/S0196-9781(97)00067-3 (1997).
Werneke, S. W., Swann, C., Farquharson, L. A., Hamilton, K. S. & Smith, A. M. The role of metals in molluscan adhesive gels. J. Exp. Biol. 210, 2137–2145. https://doi.org/10.1242/jeb.006098 (2007).
Mc Donnell, R. J., Paine, T. D. & Gormally, M. J. A Guide to the Invasive and Native Fauna of California (University of California Agricultural and Natural Resources Publications, 2009).
Thomas, A., Mc Donnell, R. J., Paine, T. D. & Harwood, J. D. A Field Guide to the Slugs of Kentucky (University of Kentucky Press, Lexington, 2010).
Gould, J., Valdez, J. W. & Upton, R. Adhesive defence mucus secretions in the red triangle slug (Triboniophorus graeffei) can incapacitate adult frogs. Int. J. Behav. Biol. 125, 587–591. https://doi.org/10.1111/eth.12875 (2019).
Acknowledgements
We thank Tara Neill for supplying slugs, Andrew Colton for assisting with the slug bioassay, and Kelly Donahue and Christina Flinn for technical assistance. This work was supported in part by Oregon Seed Council, Oregon Association of Nurseries, Agricultural Research Foundation, and base funding from USDA ARS CRIS 2072-22000-040-00D for M.Y.C; and by CRIS project # MIS-311360 for S.J.A.
Author information
Authors and Affiliations
Contributions
S.J.A., J.A.C. and M.Y.C. conducted molecular cloning, heterologous expression and binding assays. S.J.A., R.M.D. and M.Y.C. performed the peptide injection bioassays. S.J.A., R.M.D., R.C.M. and M.Y.C analyzed data and wrote the manuscript. All the authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Supplementary Information 2.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ahn, SJ., Mc Donnell, R.J., Corcoran, J.A. et al. Identification and functional characterization of the first molluscan neuromedin U receptor in the slug, Deroceras reticulatum. Sci Rep 10, 22308 (2020). https://doi.org/10.1038/s41598-020-79047-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-79047-x
This article is cited by
-
Improved cryopreservation media formulation reduces costs of maintenance while preserving function of genetically modified insect cells
In Vitro Cellular & Developmental Biology - Animal (2022)
-
Cell lines derived from the small hive beetle, Aethina tumida, express insecticide targets
In Vitro Cellular & Developmental Biology - Animal (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.