Abstract
Antibiotic-resistant Escherichia coli (E. coli) are common in retail poultry products. In this study, we aimed to isolate and characterize multidrug resistant (MDR) E. coli in raw chicken meat samples collected from poultry shops in Sylhet division, Bangladesh, as well as to determine correlation between resistance phenotype and genotype. A total of 600 chicken meat swabs (divided equally between broiler and layer farms, n = 300 each) were collected and the isolates identified as E. coli (n = 381) were selected. Disc diffusion antimicrobial susceptibility assay showed resistance of these isolates to ampicillin, erythromycin, tetracycline, streptomycin, trimethoprim-sulfamethoxazole, chloramphenicol, and gentamicin. Polymerase chain reaction (PCR) identified several antibiotic resistance genes (ARGs) in our isolates. Among these ARGs, the prevalence of tetA (for tetracycline) was the highest (72.58%) in broiler chicken isolates, followed by sul1 (for sulfonamide; 44.16%), aadA1 (for streptomycin; 33.50%), ereA (for erythromycin; 27.41%), aac-3-IV (for gentamicin; 25.38%), and the two genes cmlA (24.87%) and catA1 (8.63%) for chloramphenicol. On the other hand, the respective prevalence in layer chicken isolates were 82.06%, 47.83%, 35.87%, 35.33%, 23.91%, 19.02%, and 5.43%. Furthermore, 49.23% of the isolates from broiler chicken were MDR, with the presence of multiple antibiotic resistance genes, including 3 (40.11%) and 4 (9.13%) genes. On the other hand, 51.09% of layer chicken E. coli isolates were MDR, with 3, 4 or 5 ARGs detected in 36.41%, 14.13%, and 0.54% of the isolates, respectively. We also found that 12.8% of broiler chicken E. coli isolates and 7.61% of layer chicken isolates carried genes coding for extended-spectrum SHV beta-lactamases. Lastly, we report the presence of the AmpC beta-lactamase producing gene (CITM) in 4.56% and 3.26% of broiler and layer chicken E. coli isolates, respectively. We found significant correlations between most of the antimicrobial resistant phenotypes and genotypes observed among the investigated E. coli isolates. Our findings highlight the need for the prudent use of antimicrobials in chickens to minimize the development of antibiotic-resistant bacterial strains.
Similar content being viewed by others
Introduction
Antimicrobial resistance has recently become a public health concern. In response to the problem, the World Health Organization (WHO) has recommended a global surveillance system in veterinary and human medicine. The theme of the 2011 World Health Day was “Antibiotic resistance: no action today, no cure tomorrow” and it was selected to create mass awareness among the world population.
Escherichia coli (E. coli), a member of the Enterobacteriaceae family and a major cause of foodborne infections, is a common inhabitant of gastrointestinal tract of poultry, animals, and humans1. Unhygienic slaughter practices are responsible for contamination of meat with E. coli2. It has been reported that the E. coli strains isolated from contaminated meat and meat products are resistant to commonly used antibiotics3. Excessive use of antibiotics is considered the main cause of antibiotic resistance4,5. This resistance is acquired through horizontal gene transfer or gene mutations6,7. Multidrug-resistant (MDR) bacteria usually harbour several drug resistant genes8. The rapid emergence of multidrug-resistant E. coli strains has resulted in significant morbidity and mortality in humans9.
Beta-lactamases are bacterial enzymes which confer resistance to beta-lactam antibiotics, such as penicillin and cephalosporin by hydrolysing the beta-lactam ring. In recent years, new types of beta-lactamase enzymes including extended-spectrum beta-lactamases (ESBLs) and AmpC beta-lactamases have emerged10,11,12. The most common beta-lactamases in Gram-negative bacteria are TEM, SHV, OXA, CMY, and CTX-M beta-lactamases13. ESBLs and AmpCs are mostly located on mobile genetic elements (plasmids or integrons). These mobile genetic elements get transferred to other bacterial cells through horizontal gene transfer mechanisms, including conjugation, transformation, and transduction14. Food animals as well as retail meat act as reservoir of ESBL and AmpC-producing E. coli15,16,17,18.
Bangladesh is a large poultry producer. According to the report published in 2015 by the Department of Livestock Services, there were over 115,000 farms, producing approximately 170 million broiler and layer chickens in Bangladesh. Like many other developing countries, hygienic raw food processing technology is still underdeveloped and there is lack of proper antimicrobial drug resistance surveillance in Bangladesh. Uncontrolled use of antimicrobials for the prevention and/or treatment of diseases of food animals increases the risk of emergence of resistant bacterial strains. Contamination of chicken meat with ESBL and AmpC-producing E. coli is currently becoming an emerging food safety concern in Bangladesh. However, limited information is available on the prevalence and the genotypic characteristics of antibiotic-resistant bacterial strains associated with humans’ or food animals’ ecological niches in Bangladesh.
In this study, we identified and isolated E. coli from broiler and layer chicken meat from retail poultry shops in Sylhet division of Bangladesh. In addition, the resistance of these isolates to commonly used antibiotics, such as tetracycline and others was tested. Multiplex and uniplex polymerase chain reaction (PCR) assays were used to test for several non-beta lactam antibiotic resistance genes, such as tetracycline resistance gene (tetA), as well as genes involved in beta-lactam antibiotic resistance, ESBL genes (TEM, CTX-M, CTX-M-1, CTX-M-2, SHV), and AmpC (CITM).
Materials and methods
Ethics statement
The handling of animals in the study was performed in accordance with the current Bangladesh legislation (Cruelty to Animals Act 1920, Act No. I of 1920 of the Government of the People’s Republic of Bangladesh). The specific experiments were approved by the Ethics Committee of Sylhet Agricultural University and National Institute of Biotechnology, Bangladesh.
Isolation and identification of E. coli
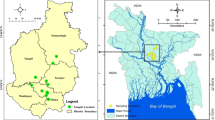
A total of 600 swabs were collected randomly from broiler (n = 300) and layer (n = 300) chicken meat samples, derived from 100 different retail poultry shops at Sylhet division of Bangladesh. Gram staining, growth characteristics on culture media (including nutrient broth, nutrient agar, MacConkey’s agar, Eosin Methylene Blue agar; all from Merck, Germany), and results of biochemical tests (including sugar fermentation, indole, methyl red (MR), Voges-Proskauer (VP), and citrate utilization tests) were used for identification and isolation of E. coli, as previously described19. Molecular confirmation of the isolates was performed using PCR targeting the 16S rRNA, using a primer set specific for E. coli, as previously described20. The isolates identified as E. coli (n = 381; 197 from broiler and 184 from layer chickens) were selected for further investigation.
Antimicrobial susceptibility testing
The susceptibilities of the 381 chicken meat-derived E. coli isolates to a panel of commonly used antibiotics were determined using the Kirby-Bauer method on Mueller–Hinton agar plates (Merck, Germany) according to the guidelines and breakpoints of the Clinical and Laboratory Standard Institute21. The antimicrobial discs used, which were all obtained from Oxoid (UK), included: trimethoprim-sulphamethoxazole (23.75 µg), chloramphenicol (30 µg), erythromycin (15 µg), gentamicin (10 µg), tetracycline (30 µg), streptomycin (10 µg), and ampicillin (10 µg). E. coli ATCC 25,922 (American Type Culture collection, Manassas, VA, USA) was used as a control strain. Test results were only validated when the diameters of the inhibition zones of the E. coli ATCC 25922 control strain were within the performance ranges. Resistant and intermediate resistant isolates were considered as non-susceptible as previously described22. E. coli was defined as multidrug resistant isolate when it was found non-susceptible to at least one agent in three or more different classes of antimicrobial agents, excluding the broad-spectrum penicillins without a β-lactamase inhibitor22.
Extraction of bacterial genomic DNA
All E. coli isolates (n = 381) were cultured overnight in nutrient broth at 37 °C and then bacterial genomic DNA was extracted using the Phenol–Chloroform Isoamyl Alcohol (PCI) method, as described previously23. The average concentration and purity of the extracted DNA were determined using the Nano Drop™ 2000c spectrophotometer (ThermoScientific, USA).
PCR amplification and detection of antibiotic resistant genes
All the tested E. coli isolates (n = 381) were PCR-screened for the presence of seven non-beta-lactam and six beta-lactam antibiotic resistant genes (ARGs) using a combination of two uniplex and two multiplex assays. The resistant genes for tetracycline (tetA) and streptomycin (aadA1) were amplified individually using set 1 and 2 primers, respectively (Table 1). Set 3 and 4 primers were used to detect some other ARGs (sul1, catA1, cmlA, ereA, aac-3-IV, blaSHV, and CITM) and four types of ESBL genes (blaTEM, blaCTX-M, blaCTX-M-1, and blaCTX-M-2), respectively (Table 1). All PCR amplifications were conducted in a thermal cycler (Gene Atlas, Japan) using the conditions listed below. The basic setup of the uniplex PCR amplification for set 1 consisted of an initial denaturation step at 95 °C for 15 min, followed by denaturation at 94 °C for 30 s, annealing at 56 °C for 30 s, and extension at 72 °C for 1 min. This cycle was repeated 30 times followed by a final extension step at 72 °C for 10 min24. The uniplex PCR amplification conditions for set 2 consisted of an initial denaturation step at 95 °C for 3 min, followed by denaturation at 94 °C for 1 min, annealing at 58 °C for 90 s, and extension at 72 °C for 1 min. This cycle was repeated 35 times followed by a final extension step at 72 °C for 10 min25. Regarding the basic setup of the multiplex PCR for primer set 3, it consisted of an initial denaturation step at 95 °C for 15 min, followed by denaturation at 94 °C for 1 min, annealing at 58 °C for 30 s, and extension at 72 °C for 1 min. This cycle was repeated 30 times followed by a final extension step at 72 °C for 10 min24. Finally, the thermal profile of the multiplex PCR with primer set 4 included an initial denaturation step at 94 °C for 3 min, followed by denaturation at 94 °C for 1 min, annealing at 55 °C for 1 min, and extension at 72 °C for 1 min. This cycle was repeated 30 times followed by a final extension step at 72 °C for 10 min26,27. After amplification, 10 µl of each PCR reaction was separated on a 1.5% (w/v) agarose gel electrophoresis using QA-AgaroseTM (MP Biomedical, USA), stained with ethidium bromide (0.5 mg/ml), and visualized using a gel documentation system (Uvitech, UK). A molecular weight marker with 100 bp increments (100 bp DNA ladder, Invitrogen™, Massachusetts, USA) was used as a size standard. Strains of E. coli O157:K88ac:H19, CAPM 5933 and E. coli O159:H20, CAPM 6006 were used as positive controls, while distilled water was used as a negative control.
Statistical analysis
The antibiotic resistance data are expressed as percentages or frequency of the E. coli isolates. A two-way analysis of variance (ANOVA) without replication was used to determine the significant differences in the levels of resistance prevalence among the selected antibiotics, between broiler and layer chickens, as well as among the four districts under study. A P value of < 0.05 was considered to be statistically significant. These statistical analyses were carried out using the GraphPad Prism (version 6; GraphPad Software Inc.; USA).
Results
Prevalence of E. coli in broiler and layer chicken meat swabs
We collected a total of 600 chicken meat swab samples (75 from broiler and 75 from layer chickens from each of the four districts of Sylhet division; Sylhet, Moulavibazar, Sunamganj, and Habiganj). Out of the 600 samples, 381 E. coli isolates (63.5%) (197 from broiler and 184 from layer chicken) were identified using staining, cultural, and biochemical tests (Table 2). There was no statistically significant difference in the prevalence of E. coli between samples from broiler and layer chickens (P = 0.06), nor among the four districts (P = 0.37).
Prevalence of antimicrobial resistance
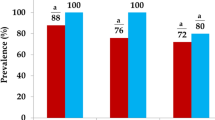
All E. coli isolates (n = 381) were tested for resistance to seven different antimicrobial agents by disc diffusion method. As shown in Table 3, 286 (75.06%) of the isolates were MDR. Resistance to ampicillin, erythromycin, and tetracycline were the most prevalent in the isolates (98.95%, 89.5%, and 85.3%, respectively). No significant difference in resistance patterns was observed in isolates from broiler and layer chicken meat (P = 0.18).
Prevalence of non-beta-lactam ARGs
The overall prevalence of non-beta-lactam ARGs among the investigated E. coli isolates in relation to type of hens (broiler or layer) in Sylhet division are given in Table 4. The most prevalent gene was that of tetA (for tetracycline resistance; harboured by 77.17% of the isolates), which was followed by sul1 (for sulphonamide resistance; 45.94%), aadA1 (for streptomycin resistance; 34.65%), ereA (for erythromycin resistance; 31.23%), aac-3-IV (for gentamicin resistance; 24.67%), and the two genes cmlA (22.05%) and catA1 (7.09%) for chloramphenicol resistance. Additionally, there was a significant difference in the prevalence of the seven non-beta-lactam ARGs among the E. coli isolates (P = 0.0001) but there was no significant difference in the prevalence of each antibiotic resistance gene in broiler versus layer chickens (P = 0.42).
As mentioned previously, the investigated E. coli isolates were from broiler and layer chickens that have been collected from four districts within Sylhet division. The prevalence of E. coli isolates from broiler or layer chicken harbouring non-beta-lactam resistance genes in relation to these districts is shown in Table 5. There was no significant difference in the prevalence of sul1, catA1, ereA, aac-3-IV, tetA and aadA1genes between broiler and layer chickens, nor between the four districts (P > 0.05). On the other hand, the prevalence of cmlA gene was significantly higher in broiler chicken than in layer chicken. Moreover, the prevalence of cmlA gene was significantly different among the four districts under study (P < 0.05).
Prevalence of MDR genes
MDR analysis was carried out according to the definition proposed previously22. The analysis was performed against five antimicrobial categories (representative antimicrobials tested in this analysis and the respective genes involved are shown in brackets): aminoglycosides (gentamicin-aac-3-IV, streptomycin-aadA1), tetracyclines (tetracycline-tetA), phenicols (choloramphenicol-cmlA and catA1), macrolides (erythromycin-ereA) and folate pathway inhibitors (sulfonamide/trimethoprim-sul1). The analysis showed 26 resistance profiles (Fig. 1), the most frequent among which (n = 16) correlated with isolates from layer chickens and harboured the resistance genes for sulfonamide, erythromycin and tetracycline.
MDR profiles of E. coli isolates from broiler (n = 97) and layer (n = 94) chickens. SUL sulfonamide (a representative of folate pathway inhibitors), CML chloramphenicol (a representative of phenicols), ERE erythromycin (a representative of macrolides), TET tetracycline (a representative of tetracyclines), GEN gentamicin, STP streptomycin (representatives of aminoglycosides).
A total of 191 (50.13%) out of 381 E. coli isolates from broiler and layer chickens of Sylhet division carried more than 3 ARGs in their genomes. The majority of the E. coli isolates harboured resistance genes to three classes of antibiotics (146 isolates; 38.32%), while 44 isolates (11.55%) and 1 isolate (0.26%) possessed four and five antibiotic resistance genes, respectively (Table 6). Within the triple-antibiotic resistant E. coli isolates, 79 isolates were from broiler chickens, while 67 isolates were from layer chickens. Out of the 44 isolates that carried four antibiotic resistance genes, 18 (9.13%) were from broiler chickens and 26 (14.13%) were from layer chickens. However, in contrast to the one E. coli layer chicken isolate (0.54%), none of the broiler chicken isolates carried five antibiotic resistance genes. There was a significant difference in the prevalence of MDR genes (P = 0.01); however, there was no significant difference in their prevalence between broiler and layer chickens (P = 0.83).
Next, we aimed to determine the prevalence of E. coli isolates (from broiler or layer chicken) harbouring MDR genes (non-beta lactam antibiotics) in relation to the four districts of Sylhet division under study (Table 7). Within the three categories of MDR isolates (those having 3, 4, or 5 MDR genes), all the prevalence differences between the four districts were found to be non-significant (P > 0.05). Similarly, there were no significant differences in the prevalence of MDR isolates between broiler and layer chickens, with the exception of isolates with 4 MDR genes. For this category, we observed a significant difference (P = 0.01) in the prevalence of MDR isolates between the two types of hens.
Detection of beta-lactamase coding genes (ESBL and AmpC)
Out of 381 E. coli isolates, 53 (13.91%) harboured beta-lactam ARGs in their genomes. Additionally, 38 isolates (10%) were positive for the SHV ESBL gene, whereas 15 (3.93%) were positive for AmpC (CITM) gene (Table 8). Out of 197 resistant E. coli isolates from broiler chicken, 24 (12.18%) and 9 (4.56%) were positive for SHV and CITM genes, respectively. Similarly, out of 184 E. coli isolates from layer chicken, 14 (7.61%) and 6 (3.26%) were positive for SHV and CITM genes, respectively. None of the investigated E. coli isolates (whether from broiler or layer chickens) contained TEM, CTX-M, CTX-M-1, or CTX-M-2 genes. Therefore, in total, 16.75% (n = 33) of broiler chicken E. coli isolates carried either ESBL or AmpC genes, compared to 10.87% (n = 20) of layer chicken E. coli isolates. Statistically significant differences between the prevalence of both types of beta-lactamase coding genes (ESBL and AmpC) were observed (P = 0.001) but there were no significant differences between broiler and layer chickens (P = 0.24).
The data regarding the prevalence of beta-lactam (ESBL and AmpC) ARGs among the investigated isolates (from broiler or layer chickens) in relation to the four districts of Sylhet division are shown in Table 9. There was a statistically significant higher prevalence of isolates possessing the SHV (ESBL) gene in broiler chicken than in layer chicken (P = 0.01), while there were no significant differences among the four districts (P = 0.07). In the case of the CITM gene (AmpC), there was no significant differences in the prevalence of isolates possessing this gene between broiler and layer chickens, nor between the four districts (P > 0.05).
Correlation between antimicrobial resistant phenotypes and genotypes
In this study, significant correlations (r2 > 0 and P < 0.05) were found between most of the antimicrobial resistant (AMR) phenotypes and genotypes observed among the investigated E. coli (n = 381) isolates (Table 10). Comparatively stronger correlation was found between Gentamycin and aac-3-IV among the E. coli strains isolated from layer chicken (r2 = 0.791 and P < 0.001). On the other hand, no significant correlation was observed between chloramphenicol AMR phenotype and catA1 gene among isolates from broiler chicken (r2 = 0.018 and P = 0.067). Similarly, in the case of layer chicken, no significant correlation was observed between erythromycin AMR phenotype and ereA gene among the E. coli isolates (r2 = 0.001 and P = 0.672).
Discussion
Chicken meat is a potential source of multi-drug resistant ESBL-producing E. coli strains, which are responsible for serious human health concerns worldwide32. In this molecular study, we isolated E. coli from chicken meat and examined the existence of ARGs. The high prevalence of antibiotic resistant E. coli isolates in our findings indicates that the raw chicken meat from retail poultry shops could be contaminated with antimicrobial-resistant E. coli. This is alarming for developing countries like Bangladesh, where retail poultry shops hardly maintain proper hygienic condition during processing of chicken meat.
In the current study, the overall prevalence of E. coli in chicken meat was 63.5% whereas, 65.67% of broiler and 61.33% of layer meat swabs tested positive for E. coli. Other research groups detected high frequency of E. coli in poultry meat33,34,35. In contrast, Ranjbar et al.36, Moawad et al.37 and Younis et al.38 showed lower prevalence of E. coli in raw chicken meat. Another study in India found that 78% of broiler chicken meat specimens from retail shops were contaminated with E. coli39. Jakaria et al.40 reported that in Bangladesh, the prevalence rates of E. coli in layer, broiler, and indigenous chicken were 78.67%, 82% and 70%, respectively.
In this study, the disc diffusion method showed relatively higher frequency of antibiotic resistance and multidrug resistance among the investigated E. coli isolates than the genotypic analysis. This may be due to the possible protective role of the tested genes against multiple (often related) antimicrobial drugs that are not structurally or mechanistically related41. We found that more than 80% of the tested E. coli isolates were resistant to the common medically used antibiotics, such as ampicillin, erythromycin, and tetracycline. These findings were more or less similar to the findings of other researchers42,43. A similar study in Ethiopia showed that E. coli isolates from broiler chicken were resistant to tetracycline (90%), streptomycin (78%), ampicillin (60%) and highly sensitive to gentamicin (77%)44.
In this study, among the seven tested non-beta-lactam antibiotic resistance genes, the prevalence of tetracycline (tetA), sulphonamides (sul1), and streptomycin (aadA1) resistance genes were the highest (72.58%, 44.67% and 33.50%, respectively), followed by erythromycin (ereA) (31.23%) and the two chloramphenicol resistant genes cmlA (22.05%) and catA1(7.09%). Our findings indicate that chicken meat E. coli could be a reservoir of resistance genes, which may later become transferred to other common pathogens.
In a similar study, the prevalence rates of tetA, aadA1 and sul1ARGs in E. coli isolated from Vietnam were found to be 81%, 81%, and 27.1%, respectively24, while their respective rates in Portugal were 41.1%, 70.6% and 23.5%45. Additionally, Moawad et al.37 reported that E. coli isolates from poultry meat were resistant to tetracycline (80.9%), streptomycin (61.9%) and trimethoprim/sulphamethoxazole (61.9%), which is higher than the prevalence rates reported in our present study. In Bangladesh, 37–100% of the poultry-derived E. coli strains were found resistant to chloramphenicol, tetracycline, streptomycin, erythromycin and penicillin, as reported by Rahman et al. and Islam et al.46,47. The lowest prevalence rate was observed with aac-3-IV (gentamicin resistance gene; 24.67%) which may be attributed to the very low absorption rate of gentamycin in poultry48.
We showed that 75.06% of our E. coli isolates were resistant to at least three antibiotics. In a study conducted in Iran, a high prevalence rate (64.91%) of MDR strains among E. coli isolates from commercial chicken meat has been reported49. Studies have demonstrated even higher prevalence rates of MDR E. coli in broiler (94%) and layer (60%) chicken in India50 and in Nepal (80.0%)51. Such high prevalence of MDR isolates may be due to misuse of antibiotics, which may ultimately replace the drug sensitive microorganisms in an antibiotic saturated environment52.
In our study, a total of 53 (13.91%) beta-lactam antibiotic resistant E. coli isolates were identified from broiler (n = 33; 16.75%) and layer (n = 20; 10.87%) chicken meat. Among the 5 types of bla genes (TEM, CTX-M, CTX-M-1, CTX-M-2 and SHV) tested by multiplex PCR in the current study, only the blaSHV gene was detected in our E. coli isolates. In a study conducted in Netherlands, blaCTX-M-1(58.1%) has been found to be the most common gene in chicken meat, followed by blaTEM-52 (14%) and blaSHV-12 (14%)53. In another study conducted in Egypt, TEM, CTX-M and SHV genes have been detected in 57.55%, 46.23% and 23.58% of the isolates, respectively54. Highly prevalent ESBL-producing E. coli strains in meat samples from broiler (87%) and layer (42%) chickens have been previously reported in India50, Nepal (36.9%)51 and Vietnam (37%)55. In a previous study, the AmpC gene has been found to be mostly plasmid-associated whereas the chromosomal AmpC was found in a small percentage of E. coli56. In Vietnam and Italy, 84.2%24 and 11.2%57, respectively, of the chicken meat isolates carried plasmid-associated AmpC genes, which is higher than the prevalence among the isolates of the current study. Over expression of ESBL and AmpC beta-lactamases in gram-negative bacteria may reduce therapeutic options for treatment of their infections by providing resistance to most beta-lactam antibiotics58.
The strong correlation between the phenotypes and genotypes of AMR in bacteria may indicate that the resistance to these antibiotics is mainly attributed to the presence of certain AMR genes in their genome. Based on this, a number of strong correlations between the phenotypes and genotypes of AMR in E. coli were found in our study, such as between gentamicin and aac-3-IV, tetracycline and tetA, and streptomycin and aadA1, indicating that the resistance to some antimicrobials may be mediated, at least partially, by a single gene. This finding is more or less similar to the results of previous studies59,60.
On the other hand, interestingly, we found that some strains possessed resistance phenotypes but did not have the corresponding ARGs and vice versa. This finding is similar to the results reported by Rosengren et al.60. A possible explanation is that resistance phenotypes can be expressed upon the stimulation of many different genetic factors, and that each factor may present a unique epidemiological character61,62. Also, this may be due to the co-selection pressure of one antimicrobial class on another. It is known that the use of a particular antimicrobial agent can select for resistance not only to its own, but also act as potential co-selection marker for other antimicrobials agents. It means the use of single antimicrobial agent can lead to the selection and co-selection of multiple resistance phenotypes and ARGs63. It is reported that the high prevalence of MDR isolates as well as their persistence is governed by co-selection processes, even in the absence of antibiotic selection pressure64.
In the same context, in the current study we found comparatively large number of phenotypic erythromycin resistance E. coli strains than those carrying the ereA gene. This might be the result of the carriage of erythromycin resistance genes other than that included in our study. Thus, further detailed investigation is necessary to unveil the exact mechanism of AMR in E. coli isolates of broiler and layer chickens produced in Bangladesh.
There is very limited data on antibiotic use in chicken production in Bangladesh. In addition to the AMR genes that could be detected given the available resources, there may be other AMR genes that can be revealed in future studies. Further studies on the roles of TEM, SHV, and CTX genes will be important.
Conclusion
Our study aimed primarily to characterize the antibiotic resistant E. coli isolates from commercial broiler and layer chicken meat samples in greater Sylhet division of Bangladesh. The correlations between the phenotypic and the genotypic susceptibility were also explored. Overall, the prevalence of E. coli was 63.5% (381/600) in these samples, from which 75.06% (286/381) of the isolates were MDR and 50.13% (191/381) contained 3 to 5 MDR genes. The isolates from raw chicken meat were highly resistant to ampicillin, erythromycin and tetracycline and 13.91% (53/381) of the isolates contained beta-lactamase (ESBL and AmpC) producing genes. This situation is alarming for Bangladesh, where facilities for health care, surveillance for antibiotics medication, and facilities to detect MDR and ESBL genes are underdeveloped. Our results highlight the need to develop novel antibiotic with potent activity against MDR and ESBL-producing bacteria. At the same time, promoting the rational use of antibiotics in livestock, as well as adopting safe food handling and proper cooking practices are crucial to reduce or eliminate the risk from pathogenic antibiotic resistance bacteria originating from raw foods.
References
Levine, M. M. Escherichia coli that cause diarrhea: Enterotoxigenic, enteropathogenic, enteroinvasive, enterohemorrhagic, and enteroadherent. J. Infect. Dis. 155(3), 377–389 (1987).
Laury, A., Echeverry, A. & Brashears, M. Fate of Escherichia coli O157: H7 in Meat. In Safety of Meat and Processed Meat, 31–53. (Springer, 2009).
Molbak, K. Spread of resistant bacteria and resistance genes from animals to humans—The public health consequences. J. Vet. Med. B Infect. Dis. Vet. Public Health 51(8–9), 364–369 (2004).
Moreno, A. et al. Extended-spectrum beta-lactamases belonging to CTX-M group produced by Escherichia coli strains isolated from companion animals treated with enrofloxacin. Vet. Microbiol. 129(1–2), 203–208 (2008).
Okeke, I. N., Lamikanra, A. & Edelman, R. Socioeconomic and behavioral factors leading to acquired bacterial resistance to antibiotics in developing countries. Emerg. Infect. Dis. 5(1), 18–27 (1999).
Laxminarayan, R. & Brown, G. M. Economics of antibiotic resistance: A theory of optimal use. J. Environ. Econ. Manag. 42(2), 183–206 (2001).
Hughes, D. & Andersson, D. I. Evolutionary consequences of drug resistance: Shared principles across diverse targets and organisms. Nat. Rev. Genet. 16(8), 459–471 (2015).
Nikaido, H. Multidrug resistance in bacteria. Annu. Rev. Biochem. 78, 119–146 (2009).
de Been, M. et al. Dissemination of cephalosporin resistance genes between Escherichia coli strains from farm animals and humans by specific plasmid lineages. PLoS Genet. 10(12), e1004776 (2014).
Babic, M., Hujer, A. M. & Bonomo, R. A. What’s new in antibiotic resistance? Focus on beta-lactamases. Drug Resist. Updat. 9(3), 142–156 (2006).
Bradford, P. A. Extended-spectrum beta-lactamases in the 21st century: Characterization, epidemiology, and detection of this important resistance threat. Clin. Microbiol. Rev. 14(4), 933–951 (2001) (table of contents).
Paterson, D. L. Resistance in gram-negative bacteria: Enterobacteriaceae. Am. J. Med. 119(6 Suppl 1), S20–S28 (2006) (discussion S62–S70).
Livermore, D. M. & Woodford, N. The beta-lactamase threat in Enterobacteriaceae, Pseudomonas and Acinetobacter. Trends Microbiol. 14(9), 413–420 (2006).
Perilli, M. et al. Identification and characterization of a new metallo-β-lactamase, IND-5, from a clinical isolate of Chryseobacterium indologenes. Antimicrob. Agents Chemother. 51(8), 2988–2990 (2007).
Belmar Campos, C. et al. Prevalence and genotypes of extended spectrum beta-lactamases in Enterobacteriaceae isolated from human stool and chicken meat in Hamburg, Germany. Int. J. Med. Microbiol. 304(5–6), 678–684 (2014).
Leverstein-van Hall, M. A. et al. Dutch patients, retail chicken meat and poultry share the same ESBL genes, plasmids and strains. Clin. Microbiol. Infect. 17(6), 873–880 (2011).
Ding, H. et al. The prevalence of plasmid-mediated AmpC beta-lactamases among clinical isolates of Escherichia coli and Klebsiella pneumoniae from five children’s hospitals in China. Eur. J. Clin. Microbiol. Infect. Dis. 27(10), 915–921 (2008).
Maleki, A. et al. High Prevalence of AmpC beta-lactamases in clinical isolates of Escherichia coli in Ilam, Iran. Osong. Public Health Res. Perspect. 6(3), 201–204 (2015).
Cheesbrough, M. Medical Laboratory Manual for Tropical Countries, Vol. 2: Microbiology (Tropical Health Technology, 1984).
Sabat, G. et al. Selective and sensitive method for PCR amplification of Escherichia coli 16S rRNA genes in soil. Appl. Environ. Microbiol. 66(2), 844–849 (2000).
CLSI, Performance Standards for Antimicrobial Susceptibility Testing; Fifteenth Informational Supplement. NLSI document M100-S15. (Clinical and Laboratory Standards Institute, Wayne, 2005).
Magiorakos, A. P. et al. Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: An international expert proposal for interim standard definitions for acquired resistance. Clin. Microbiol. Infect. 18(3), 268–281 (2012).
Sambrook, J., Fritsch, E. F. & Maniatis, T. Molecular Cloning: A Laboratory Manual (Cold Spring Harbor Laboratory Press, 1989).
Van, T. T. et al. Safety of raw meat and shellfish in Vietnam: an analysis of Escherichia coli isolations for antibiotic resistance and virulence genes. Int. J. Food Microbiol. 124(3), 217–223 (2008).
Szczepanowski, R. et al. Detection of 140 clinically relevant antibiotic-resistance genes in the plasmid metagenome of wastewater treatment plant bacteria showing reduced susceptibility to selected antibiotics. Microbiology 155(Pt 7), 2306–2319 (2009).
Dierikx, C. et al. Increased detection of extended spectrum beta-lactamase producing Salmonella enterica and Escherichia coli isolates from poultry. Vet. Microbiol. 145(3–4), 273–278 (2010).
Hasman, H. et al. beta-Lactamases among extended-spectrum beta-lactamase (ESBL)-resistant Salmonella from poultry, poultry products and human patients in The Netherlands. J. Antimicrob. Chemother. 56(1), 115–121 (2005).
Randall, L. P. et al. Antibiotic resistance genes, integrons and multiple antibiotic resistance in thirty-five serotypes of Salmonella enterica isolated from humans and animals in the UK. J. Antimicrob. Chemother. 53(2), 208–216 (2004).
Olesen, I., Hasman, H. & Aarestrup, F. M. Prevalence of beta-lactamases among ampicillin-resistant Escherichia coli and Salmonella isolated from food animals in Denmark. Microb. Drug Resist. 10(4), 334–340 (2004).
Mulvey, M. R. et al. Characterization of the first extended-spectrum beta-lactamase-producing salmonella isolate identified in Canada. J. Clin. Microbiol. 41(1), 460–462 (2003).
Eckert, C. et al. Dissemination of CTX-M-type beta-lactamases among clinical isolates of Enterobacteriaceae in Paris, France. Antimicrob. Agents Chemother. 48(4), 1249–1255 (2004).
Trkov, M. et al. Molecular Characterization of Escherichia coli strains isolated from different food sources. Food Technol. Biotechnol. 52(2), 255–262 (2014).
Rashid, M. et al. Prevalence, genetic profile of virulence determinants and multidrug resistance of Escherichia coli isolates from foods of animal origin. Vet. World 6(3), 139–142 (2013).
Adeyanju, G. T. & Ishola, O. Salmonella and Escherichia coli contamination of poultry meat from a processing plant and retail markets in Ibadan, Oyo State, Nigeria. SpringerPlus 3(1), 139 (2014).
Park, H. J. et al. Antibiotic resistance and virulence potentials of Shiga toxin-producing Escherichia coli Isolates from raw meats of slaughterhouses and retail markets in Korea. J. Microbiol. Biotechnol. 25(9), 1460–1466 (2015).
Ranjbar, R. et al. Shiga (Vero)-toxin producing Escherichia coli isolated from the hospital foods; virulence factors, o-serogroups and antimicrobial resistance properties. Antimicrob. Resist. Infect. Control 6, 4 (2017).
Moawad, A. A. et al. Occurrence of Salmonella enterica and Escherichia coli in raw chicken and beef meat in northern Egypt and dissemination of their antibiotic resistance markers. Gut. Pathog. 9(1), 57 (2017).
Younis, G. A. et al. Virulence and extended-spectrum beta-lactamase encoding genes in Escherichia coli recovered from chicken meat intended for hospitalized human consumption. Vet. World 10(10), 1281–1285 (2017).
Hussain, A. et al. Risk of transmission of antimicrobial resistant Escherichia coli from commercial broiler and free-range retail chicken in India. Front. Microbiol. 8, 2120 (2017).
Jakaria, A., Islam, M. A. & Khatun, M. M. Prevalence, characteristics and antibiogram profiles of Escherichia coli isolated from apparently healthy chickens in Mymensingh, Bangladesh. Microbes Health 1(1), 27–29 (2012).
Gomez, J.E. et al. Ribosomal mutations promote the evolution of antibiotic resistance in a multidrug environment. Elife. 6 (2017).
Lee, C. W. et al. Characterization of highly pathogenic H5N1 avian influenza A viruses isolated from South Korea. J. Virol. 79(6), 3692–3702 (2005).
Akond, M. A. et al. Antibiotic resistance of Escherichia coli isolated from poultry and poultry environment of Bangladesh. Am. J. Environ. Sci. 5(1), 47–52 (2009).
Tesfaheywet, Z. & Berhanu. Antimicrobial resistant pattern of fecal Escherichia coli in selected broiler farms of eastern Haarge Zone, Ethiopia. Int. J. Appl. Biol. Pharm. Technol. 4(4) (2013).
Costa, D. et al. Prevalence of extended-spectrum beta-lactamase-producing Escherichia coli isolates in faecal samples of broilers. Vet. Microbiol. 138(3–4), 339–344 (2009).
Rahman, M., Rahman, B. M. & Rahman, B. Antibiogram and plasmid profile analysis of isolated Escherichia coli from broiler and layer. Res. J. Microbiol 3(2), 82–90 (2008).
Islam, M. J. et al. Isolation of plasmid mediated multidrug resistant E. coli from poultry. Int. J. Sustain. Crop Prod. 3(5), 46–50 (2008).
Ginns, C. A. et al. Antimicrobial resistance and epidemiology of Escherichia coli in broiler breeder chickens. Avian Pathol. 25(3), 591–605 (1996).
Momtaz, H., Rahimi, E. & Moshkelani, S. Molecular detection of antimicrobial resistance genes in E. coli isolated from slaughtered commercial chickens in Iran. Vet. Med. 57(4), 193–197 (2012).
Brower, C. H. et al. The prevalence of extended-spectrum beta-lactamase-producing multidrug-resistant Escherichia coli in poultry chickens and variation according to farming practices in Punjab, India. Environ. Health Perspect. 125(7), 077015 (2017).
Shrestha, A. et al. Multi-drug resistance and extended spectrum beta lactamase producing Gram negative bacteria from chicken meat in Bharatpur Metropolitan, Nepal. BMC Res. Notes 10(1), 574 (2017).
Van de Boogard, A. E. & Stobberingh, E. E. Epidemiology of resistance to antibiotics links between animals and humans. Int. J. Antimicrob. Agents 14, 327–335 (2000).
Overdevest, I. et al. Extended-spectrum beta-lactamase genes of Escherichia coli in chicken meat and humans, The Netherlands. Emerg. Infect. Dis. 17(7), 1216–1222 (2011).
Abdallah, H. M. et al. Extended-spectrum beta-lactamases and/or carbapenemases-producing Enterobacteriaceae isolated from retail chicken meat in Zagazig, Egypt. PLoS ONE 10(8), e0136052 (2015).
Nguyen, V. T. et al. Prevalence and risk factors for carriage of antimicrobial-resistant Escherichia coli on household and small-scale chicken farms in the Mekong Delta of Vietnam. J. Antimicrob. Chemother. 70(7), 2144–2152 (2015).
Philippon, A., Arlet, G. & Jacoby, G. A. Plasmid-determined AmpC-type beta-lactamases. Antimicrob. Agents Chemother. 46(1), 1–11 (2002).
Ghodousi, A. et al. Extended-spectrum ss-lactamase, AmpC-producing, and fluoroquinolone-resistant Escherichia coli in retail broiler chicken meat, Italy. Foodborne Pathog. Dis. 12(7), 619–625 (2015).
Perez-Perez, F. J. & Hanson, N. D. Detection of plasmid-mediated AmpC beta-lactamase genes in clinical isolates by using multiplex PCR. J. Clin. Microbiol. 40(6), 2153–2162 (2002).
Gow, S. P. et al. Associations between antimicrobial resistance genes in fecal generic Escherichia coli isolates from cow-calf herds in western Canada. Appl. Environ. Microbiol. 74(12), 3658–3666 (2008).
Rosengren, L. B., Waldner, C. L. & Reid-Smith, R. J. Associations between antimicrobial resistance phenotypes, antimicrobial resistance genes, and virulence genes of fecal Escherichia coli isolates from healthy grow-finish pigs. Appl. Environ. Microbiol. 75(5), 1373–1380 (2009).
Boerlin, P. et al. Antimicrobial resistance and virulence genes of Escherichia coli isolates from swine in Ontario. Appl. Environ. Microbiol. 71(11), 6753–6761 (2005).
Lanz, R., Kuhnert, P. & Boerlin, P. Antimicrobial resistance and resistance gene determinants in clinical Escherichia coli from different animal species in Switzerland. Vet. Microbiol. 91(1), 73–84 (2003).
O’Connor, A., Poppe, C. & McEwen, S. Changes in the prevalence of resistant Escherichia coli in cattle receiving subcutaneously injectable oxytetracycline in addition to in-feed chlortetracycline compared with cattle receiving only in-feed chlortetracycline. Can. J. Vet. Res. 66(3), 145 (2002).
Sundqvist, M. et al. Little evidence for reversibility of trimethoprim resistance after a drastic reduction in trimethoprim use. J. Antimicrob. Chemother. 65(2), 350–360 (2010).
Acknowledgements
We would like to thank the University Grants Commission (UGC) of Bangladesh and the Sylhet Agricultural University Research System (SAURES) for facilitating part of the study.
Author information
Authors and Affiliations
Contributions
M.M.R. and H.M.A. contributed to the conception, design, supervision, and administration of the study. All authors contributed to the data acquisition and/or analysis, writing the original version, and revising the manuscript. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Rahman, M.M., Husna, A., Elshabrawy, H.A. et al. Isolation and molecular characterization of multidrug-resistant Escherichia coli from chicken meat. Sci Rep 10, 21999 (2020). https://doi.org/10.1038/s41598-020-78367-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-78367-2
This article is cited by
-
Genome mining of Escherichia coli WG5D from drinking water source: unraveling antibiotic resistance genes, virulence factors, and pathogenicity
BMC Genomics (2024)
-
Determination of antibiotic resistance patterns and genotypes of Escherichia coli isolated from wild birds
Microbiome (2024)
-
Multi-drug-resistant Escherichia coli in adult male patients with enlarged prostate attending general hospitals in Benue state
Brazilian Journal of Microbiology (2024)
-
Evaluation of in-feed supplementation of formic acid and thymol as non-antibiotic growth promoters and assessing their effect on antimicrobial resistant E.coli isolated in Turkey
Veterinary Research Communications (2024)
-
Isolation and molecular characterization of multidrug‑resistant Escherichia coli from chicken meat
3 Biotech (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.