Abstract
This study aims to develop an artificial intelligence (AI)-based model to assist radiologists in pneumoconiosis screening and staging using chest radiographs. The model, based on chest radiographs, was developed using a training cohort and validated using an independent test cohort. Every image in the training and test datasets were labeled by experienced radiologists in a double-blinded fashion. The computational model started by segmenting the lung field into six subregions. Then, convolutional neural network classification model was used to predict the opacity level for each subregion respectively. Finally, the diagnosis for each subject (normal, stage I, II, or III pneumoconiosis) was determined by summarizing the subregion-based prediction results. For the independent test cohort, pneumoconiosis screening accuracy was 0.973, with both sensitivity and specificity greater than 0.97. The accuracy for pneumoconiosis staging was 0.927, better than that achieved by two groups of radiologists (0.87 and 0.84, respectively). This study develops a deep learning-based model for screening and staging of pneumoconiosis using man-annotated chest radiographs. The model outperformed two groups of radiologists in the accuracy of pneumoconiosis staging. This pioneer work demonstrates the feasibility and efficiency of AI-assisted radiography screening and diagnosis in occupational lung diseases.
Similar content being viewed by others
Introduction
Pneumoconiosis is a major occupational lung disease with increasing prevalence and severity worldwide. The term “pneumoconiosis” summarizes all lung diseases caused by excessive exposure to dust (e.g., silica, asbestos, coal, and mixed dust), which often occurs in the workplace. It is an irreversible disease with no cure1. In 2013 alone, it caused approximately 260,000 deaths globally2. In the United States, the prevalence of coal worker’s pneumoconiosis (CWP; a major type of pneumoconiosis) has been increasing over the last decade and now exceeds 10% among long-tenured miners nationwide and 20% in central Appalachia1. Recently, there has been a surprising resurgence of progressive massive fibrosis, a disabling and fatal form of CWP, in central Appalachia3,4. Lung transplantation is also increasingly common among CWP patients5. According to the Centers for Disease Control and Prevention (CDC), during 1999–2016, the mean years of potential life lost attributed to CWP increased from 8.1 to 12.6 years6. In addition, more than one million workers in the United States are exposed to crystalline silica, which potentially leads to silicosis7, an irreversible, often disabling type of pneumoconiosis8. Furthermore, in recent years, more reports have indicated that dental technicians also suffer from the disease caused by inhalation of various airborne particles9. Because of the lengthy and unnoticeable progression of pneumoconiosis, and the seriousness of its outcomes, regular screening of the population at potential risk is the key to the early intervention and prevention of pneumoconiosis.
Current clinical diagnosis of pneumoconiosis is mainly based on the examination of chest radiographs (i.e. X-ray images). In 1980, the International Labor Organization (ILO) established a standardized system to classify radiographic abnormalities of pneumoconiosis according to the profusion level of small opacities observed in the lung10. This system has greatly facilitated pneumoconiosis screening and staging by providing a commonly accepted standard and guidelines. However, radiograph-based diagnosis of pneumoconiosis still requires a well-trained and experienced radiologist to visually identify subtle graphic patterns and features described in the ILO guidelines. This process is laborious and subject to considerable inter- and intra-observer variations. For example, the concordance in pneumoconiosis diagnosis is between 85 and 90% among expert radiologists11, and around 80% in general medical staff12 in the United States. The consistency is likely to be even worse for the screening programs conducted at community sites, especially in developing countries. Therefore, early detection and intervention of pneumoconiosis still demand a solution to timely, accurate, and efficient screening at a relatively low economic cost.
In order to improve the diagnostic efficiency and accuracy among radiologists, a variety of methods have been developed to analyze the chest radiographs. Since 1975, researchers have been studying the feasibility of computer-based classification of profusion in chest radiographs. For example, Hall et al.13 used pattern recognition techniques with spatial moments for the analysis of radiology images. Ledley et al.14 developed a texture analysis method to classify chest chest radiographs. Savol et al.15 investigated an adaptive object-growing algorithm based on image intensity in order to recognize small rounded opacities. Since 2010, with the rise of computer-aided detection (CAD), several image analysis methods for pneumoconiosis diagnosis based on texture features in chest radiographs have been proposed16,17,18,19. Several image analysis methods have been developed to assist radiologists by reducing their workload or improving the performance in various tasks in disease diagnosis20,21,22. However, all of these methods rely on certain level of “handcrafted” feature definition, which is technically challenging and time-consuming, especially for complex tasks23 like pneumoconiosis diagnosis and staging.
Recently, artificial intelligence (AI) has been making remarkable success in medical image analysis owing to the rapid progress of “deep learning” 24,25,26,27. For example, studies have demonstrated the feasibility of using AI for diagnosis of lung abnormalities such as lung nodules, pulmonary tuberculosis, cystic fibrosis, and pneumoconiosis28. AI algorithms have achieved performance comparable to radiology experts in interpreting radiographs29. However, there is a lack of AI-based diagnostic tools for pneumoconiosis because of the complexity of the disease and the limited availability of well-annotated pneumoconiosis chest radiographs for training AI models.
Results
Staging consistency and accuracy for experienced radiologists
For the 405 pneumoconiosis patients in the training cohort, radiologist groups A and B diagnosed the stage independently through first-round reading (with discussion only within group and not between groups). The two groups reached the same staging results for 304 patients, while differed for the remaining 101 patients. Using the final stage as the ground truth (Fig. 1B), the staging accuracy was 0.84 and 0.87 for groups A and B, respectively, which underlines the challenge of accurate diagnosis even with a group of experienced radiologists.
Model development
The subregion-based model training process stopped at the 1200th epoch after validation accuracy failed to improve after 600 epochs. The model with the highest validation accuracy was selected as the final model.
Model evaluation and validation
Model evaluation results using the independent test dataset are summarized in Table 1. At subregion level, the top-right, top-left, and middle-right subregions had outstanding performance in screening accuracy (> 0.9), while the other three subregions also had acceptable performance (> 0.8). The screening sensitivity of the two middle subregions were over 0.9, while that of the two bottom subregions were slightly over 0.5. For the screening specificity, all subregions had good performance (0.88–0.94).
At subject level, the screening accuracy was 0.973 for the independent test cohort (n = 411), with both sensitivity and specificity above 0.97 (Table 1). The staging accuracy was 0.927 (Table 1), better than that achieved by human experts (0.84 and 0.87 for radiologist group A and B, respectively). Table 2 lists the confusion matrix of the final staging results on the test dataset.
Discussion
According to the ILO guidelines, diagnosis and staging of pneumoconiosis are mainly based on the profusion level of small opacities and the presence of large opacities/assembly of small opacities in chest radiographs. Accurate diagnosis and staging of pneumoconiosis requires a well-trained and experienced radiologist to distinguish subtle graphic patterns in the chest radiographs. This process is time-consuming, subjective, and often results in inconsistent and unreliable outcomes, especially for pneumoconiosis screening programs implemented in underdeveloped regions. In this study, we leveraged powerful deep learning algorithms and well-annotated chest radiographs to develop and validate a deep learning-based model for screening and staging of pneumoconiosis. In order to have a set of reliable labels as the “ground truth” to train the deep learning model, we assembled a team of eight experienced radiologists and performed a two-round reading in a double-blinded fashion. Half of these radiologists have participated in developing the latest diagnosis standard of pneumoconiosis in China, which ensured the reliability of the labels.
The major differences between the proposed deep learning-based model and the other models for screening of pneumoconiosis are: (1) The deep learning model developed in our study has a higher screening accuracy, compared to the reported values in other papers. The screening accuracies in other reported studies are 0.79 (Cai et al., 2012), 0.92 (Zhu et al., 2014) and 0.94 ~ 0.95 (Yu et al., 2011), while the screening accuracy for our model is 0.97. (2) Our model not only provides a screening result (disease or not), but also the stage of the disease, which can greatly facilitate the diagnosis and treatment planning. (3) Our model for screening and staging pneumoconiosis is based on the profusion level of small opacities and the presence of large opacities in the six subregions of a lung field, which provides more enriched radiographic information for early diagnosis and intervention. (4) The data augmentation methods are widely adopted in the computer vision and deep learning field. These are the most common techniques to increase sample size and avoid overfitting in training a deep learning model30,31. With proper augmentation, our deep learning model gained better generalization power and achieved the same level of accuracy and stability on unseen (testing) dataset compared with the training dataset.
Due to the large size of digital chest radiographs and the often limited computing resources at the actual screening sites, it is impractical to develop and deploy a deep learning-based tool that directly works on whole chest radiographs. Existing solutions either down-sample the image (e.g., Gaussian pyramid) or reduce the model capacity (e.g., shallow learning, small CNN, etc.) to a feasible scale. However, pneumoconiosis diagnosis highly relies on identifying subtle image features, rendering not much room for downgrading the image resolution or compromising feature richness by model simplification. Alternatively, in this study, we designed a two-stage deep learning model following a divide-and-conquer strategy, which simplifies the problem into classifying the relatively small subregions in the lung instead of the whole image. This modeling strategy also mimics the clinical diagnostic procedure, which allowed us to well incorporate human knowledge (Table 3) into the AI model. It has two major advantages: (1) it largely reduces the computational load and thus the requirement of computing resources; (2) by incorporating human knowledge on pneumoconiosis staging, it reduces the sample size needed in training data.
In practice, pneumoconiosis screening is usually conducted at community sites, where resources, image quality and expertise level may vary. In order to improve the robustness and generalizability of the developed model with a limited amount of training data, we performed extensive image augmentation, which is one of the most effective pre-processing strategies to enrich data variety and avoid overfitting. In this study, the original images were augmented with rotating, flipping, shifting, zooming and varying the image intensity, resolution and quality. These augmentation processes largely reduced the influence of sampling condition, image contrast and lung size, and reinforced the model to focus on image features associated with pneumoconiosis.
Low accuracies were observed in the three subregions: middle-left (0.881), bottom-right (0.856), and bottom-left (0.818). This is because the middle-left region has a relatively small area due to its overlaps with the heart in a projection plane. Each of the right and left lungs has a hilum that lies roughly midway down the lung. When moving from the hilum to the periphery on the bottom, there will also be a gradual diminution of the lung markings. All these anatomical features weaken the accuracy in the two bottom subregions. In contrast, the right bronchus is relatively shorter and flatter than those in the left. As a result, inhaled dust tends to be deposited there. Thus, the top-right and middle-right subregions have more easily identifiable radiographic abnormalities in pneumoconiosis, resulting in higher accuracies.
There are several limitations in our study. Firstly, the sample size in the test dataset is relatively small. More subjects are needed, especially those with stage II and III pneumoconiosis, for a more comprehensive evaluation of the model. Secondly, the current datasets only contain chest radiographs. Although chest radiographs the standard modality for pneumoconiosis diagnosis, computed tomography (CT) typically provides more details of the opacities regions in the lung and should be tested in future studies. Lastly, the current deep learning-based model is mainly trained by those well-labeled chest radiographs from pneumoconiosis patients and healthy people. It cannot differentiate pneumoconiosis from other lung diseases that share a similar pathology. The predictive model could be further improved by (1) integrating other useful key factors to pneumoconiosis, such as working history, respiratory assessment, lungs' main function, and blood pressure, and (2) feeding more comprehensively labeled chest radiographs from patients with different lung diseases.
Methods
Ethics approval and consent to participate
The University of Texas Southwestern Institutional Review Board granted approval for this research. Data were collected under information consent for study participation. All methods were performed in accordance with the relevant guidelines and regulations.
Training and test datasets
The training dataset was obtained from a cohort of 805 subjects (400 healthy controls, 163 stage I, 125 stage II, and 117 stage III pneumoconiosis patients). The test dataset was obtained from an independent cohort of 411 subjects (304 healthy controls, 73 stage I, 21 stage II and 13 stage III pneumoconiosis patients), which was used to evaluate the model developed from the training dataset. The subjects were enrolled from two different sites to ensure generalizability of the resulting model. All the enrolled chest radiographs (one image per subject) had been carefully checked for quality and graded either “Good” or “Acceptable” by radiologists, indicating no technical defects likely to impair the diagnosis and staging of pneumoconiosis.
In this study, we used the “final stage” determined by radiologists (see below) as the ground truth. To improve the diagnosis accuracy, we recruited eight experienced radiologists and divided them into three groups (Fig. 1A): three in group A and three in group B, with group C consisting of six radiologists (two from group A, two from group B, and two additional). The six radiologists in group C were selected from six different national institutions for occupational disease research in China, all of whom had more than 10 years’ experience in pneumoconiosis. Moreover, four of these radiologists participated in the development of the latest diagnostic standard of pneumoconiosis in China: “Diagnosis of occupational pneumoconiosis GBZ70-2015.”32.
For each subject, the radiologists in group A and B independently performed the first-round reading according to the national standard, GBZ70-2015. The lung field in each chest radiographs had been divided into six mutually exclusive subregions (see “Lung field segmentation” section for details): top-right, top-left, middle-right, middle-left, bottom-right, and bottom-left (Fig. 2). Radiologist groups A and B independently annotated for each subregion the profusion level of small opacities (level 0, 1, 2, or 3) and the presence of big opacities (see Table 4 for an example record form). Then, the stage of pneumoconiosis was determined following GBZ70-2015 (Table 3). For each subject, if the staging by groups A and B agreed, it was used as the final stage. Otherwise, the radiologist group C would read the image again and determine the final stage. This procedure was applied to the entire training and test dataset (Fig. 1B).
Model strategy
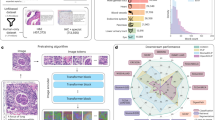
According to the routine diagnostic procedure for pneumoconiosis, we developed a two-stage deep learning model based on a divide-and-conquer strategy, which segmented a lung field into six subregions, trained subregion-specific models individually, and combined subregion-based predictions for staging (Fig. 2).
Lung field segmentation
To identify the lung field in each chest radiographs, we developed a segmentation model by adapting a U-Net semantic segmentation model together with a ResNet backbone. We first downsized the images to 256 × 256 and fed them into the model, and then resized the resulting mask image back to the original size. This resizing step allowed the model to process the images more efficiently. Then, we refined the mask with post-processing steps and extracted the lung field from the raw image (Fig. 2). To improve the accuracy and robustness of the model, we further performed extensive data augmentation, including rotating, flipping, shifting, zooming, and varying the image intensity, resolution and quality.
After segmentation, the lung field was divided into six subregions (by even distribution along the vertical direction; Fig. 2). Each subregion was only less than 1/8 of the whole image (less than 1024 × 1024), which makes the developed model feasible for the computational resources when deployed at real-world screening sites.
CNN for subregion classification
Next, we developed a subregion-based CNN classification model. We adapted a ResNet model to classify the profusion level of small opacities as well as the presence of large opacities for each of the six subregions in the lung field. Each subregion was classified as level 0, 1, 2, or 3 based on its opacity level. At the model development stage, the training cohort (805 subjects) was randomly separated into training (72 healthy controls, 72 stage I, 96 stage II, and 96 stage III), validation (24 healthy controls, 18 stage I, 8 stage II, and 8 stage III) and test sets (304 healthy controls, 73 stage I, 21 stage II, and 13 stage III). The developed model was finally evaluated in the independent test cohort of 411 subjects. Last, each subject was assigned into one of the four stages: normal, stage I, II, or III, by integrating the subregion-based classifications following GBZ70-2015 (Table 3).
Statistical analyses
To evaluate the screening performance, we used three common metrics for binary classification systems: (1) accuracy (ACCscr, the percentage of radiographs being correctly classified as normal or disease), sensitivity (SEN, the percentage of pneumoconiosis radiographs being correctly classified as abnormal), and specificity (SPE, percentage of normal radiographs being correctly classified as normal). To evaluate the staging performance, we only used accuracy (ACCsta , the percentage of radiographs being correctly classified as normal, stage I, II, or III). All statistical analyses were carried out using R version 3.6.1.
Computing environment
In this study, computational works were conducted in a computing environment using the interface of Python 3.0 based on a TensorFlow deep learning framework, which was installed and executed on a server with Linux version 3.16.0-69-generic and Ubuntu 4.8.2-19 in 64 bits. This server also includes two Intel Xeon CPU E5-2680 v3 processors of 2.50 GHz and a 30 Mb Cache, where each processor has 12 cores and the total number of logical CPU cores is 48. The server has 132 Gb RAM and an NVIDIA Tesla K40 m GPU with 2880 stream cores, 12 Gb maximum memory, 288 Gb/s maximum memory bandwidth, and 6 GHz memory clock speed.
References
Blackley, D. J., Halldin, C. N. & Laney, A. S. Continued increase in prevalence of coal workers’ pneumoconiosis in the United States, 1970–2017. Am. J. Public Health 108, 1220–1222. https://doi.org/10.2105/ajph.2018.304517 (2018).
Mortality, G. B. D. & Causes of Death, C. Global, regional, and national age-sex specific all-cause and cause-specific mortality for 240 causes of death, 1990–2013: a systematic analysis for the Global Burden of Disease Study 2013. Lancet 385, 117–171. https://doi.org/10.1016/S0140-6736(14)61682-2 (2015).
Blackley, D. J., Halldin, C. N. & Laney, A. S. Resurgence of a debilitating and entirely preventable respiratory disease among working coal miners. Am. J. Respir. Crit. Care Med. 190, 708–709. https://doi.org/10.1164/rccm.201407-1286LE (2014).
Blackley, D. J. et al. Progressive massive fibrosis in coal miners from 3 clinics in VirginiaProgressive massive fibrosis in coal miners from 3 clinics in VirginiaLetters. JAMA 319, 500–501. https://doi.org/10.1001/jama.2017.18444 (2018).
Blackley, D. J., Halldin, C. N., Cummings, K. J. & Laney, A. S. Lung transplantation is increasingly common among patients with coal workers’ pneumoconiosis. Am. J. Ind. Med. 59, 175–177. https://doi.org/10.1002/ajim.22551 (2016).
Mazurek, J. M., Wood, J., Blackley, D. J. & Weissman, D. N. Coal workers’ pneumoconiosis-attributable years of potential life lost to life expectancy and potential life lost before age 65 years—United States, 1999–2016. Morb. Mortal. Wkly Rep. 67, 819 (2018).
Steenland, K. & Ward, E. Silica: a lung carcinogen. CA Cancer J. Clin. 64, 63–69 (2014).
Raymond, L. W. & Wintermeyer, S. Medical surveillance of workers exposed to crystalline silica. J. Occup. Environ. Med. 48, 95–101 (2006).
Leggat, P. A., Kedjarune, U. & Smith, D. R. Occupational health problems in modern dentistry: a review. Ind. Health 45, 611–621 (2007).
Welch, L. S. et al. Variability in the classification of radiographs using the 1980 International Labor Organization Classification for Pneumoconioses. Chest 114, 1740–1748. https://doi.org/10.1378/chest.114.6.1740 (1998).
Binay, S. et al. Does periodic lung screening of films meets standards?. Pak J. Med. Sci. 32, 1506–1511. https://doi.org/10.12669/pjms.326.11267 (2016).
Morgan, R. H. et al. Decision processes and observer error in the diagnosis of pneumoconiosis by chest roentgenography. Am. J. Roentgenol. 117, 757–764. https://doi.org/10.2214/ajr.117.4.757 (1973).
Hall, E. L., Crawford, W. O. & Roberts, F. E. Computer classification of pneumoconiosis from radiographs of coal workers. IEEE Trans. Biomed. Eng. 6, 518–527 (1975).
Ledley, R. S., Huang, H. & Rotolo, L. S. A texture analysis method in classification of coal workers’ pneumoconiosis. Comput. Biol. Med. 5, 53–67 (1975).
Savol, A., Li, C. & Hoy, R. Computer-aided recognition of small rounded pneumoconiosis opacities in chest X-rays. IEEE Trans. Pattern Anal. Mach. Intell. 6, 479–482 (1980).
Cai, C. X., Zhu, B. Y. & Chen, H. Computer-aided diagnosis for pneumoconiosis based on texture analysis on digital chest radiographs. Appl. Mech. Mater. 241–244, 244–247. https://doi.org/10.4028/www.scientific.net/AMM.241-244.244 (2012).
Okumura, E., Kawashita, I. & Ishida, T. Computerized analysis of pneumoconiosis in digital chest radiography: effect of artificial neural network trained with power spectra. J. Digit. Imaging 24, 1126–1132. https://doi.org/10.1007/s10278-010-9357-7 (2011).
Okumura, E., Kawashita, I. & Ishida, T. Development of CAD based on ANN analysis of power spectra for pneumoconiosis in chest radiographs: effect of three new enhancement methods. Radiol. Phys. Technol. 7, 217–227. https://doi.org/10.1007/s12194-013-0255-9 (2014).
Zhu, B. et al. The development and evaluation of a computerized diagnosis scheme for pneumoconiosis on digital chest radiographs. Biomed. Eng. Online 13, 141 (2014).
Kouda, T. & Kondo, H. Computer-aided diagnosis for pneumoconiosis using neural network. Int. J. Biomed. Soft Comput. Hum. Sci. 7, 13–18 (2001).
Xu, H. et al. in Proceedings of the Third International Workshop on Pulmonary Image Analysis, September 20. 129–138.
Yu, P. et al. An automatic computer-aided detection scheme for pneumoconiosis on digital chest radiographs. J. Digit. Imaging 24, 382–393. https://doi.org/10.1007/s10278-010-9276-7 (2011).
Wang, S. et al. Artificial intelligence in lung cancer pathology image analysis. Cancers (Basel). https://doi.org/10.3390/cancers11111673 (2019).
LeCun, Y., Bengio, Y. & Hinton, G. Deep learning. Nature 521, 436–444. https://doi.org/10.1038/nature14539 (2015).
Krizhevsky, A., Sutskever, I. & Hinton, G. E. ImageNet classification with deep convolutional neural networks. Commun. ACM 60, 84–90. https://doi.org/10.1145/3065386 (2017).
Goodfellow, I., Bengio, Y., Courville, A. & Bengio, Y. Deep Learning Vol. 1 (MIT Press, Cambridge, 2016).
Zhang, M. et al. GeNeCK: a web server for gene network construction and visualization. BMC Bioinform. 20, 12. https://doi.org/10.1186/s12859-018-2560-0 (2019).
Singh, R. et al. Deep learning in chest radiography: detection of findings and presence of change. PLoS ONE 13, e0204155. https://doi.org/10.1371/journal.pone.0204155 (2018).
Rajpurkar, P. et al. Deep learning for chest radiograph diagnosis: a retrospective comparison of the CheXNeXt algorithm to practicing radiologists. PLoS Med. 15, e1002686. https://doi.org/10.1371/journal.pmed.1002686 (2018).
Perez, L. & Wang, J. The effectiveness of data augmentation in image classification using deep learning. ArXiv abs/1712.04621 (2017).
Shorten, C. & Khoshgoftaar, T. M. A survey on image data augmentation for deep learning. J. Big Data 6, 60. https://doi.org/10.1186/s40537-019-0197-0 (2019).
Cai, Z. C. Comprehension of GBZ 70–2015 “Diagnosis of Occupational Pneumoconiosis”. Zhonghua Lao Dong Wei Sheng Zhi Ye Bing Za Zhi 34, 866–867. https://doi.org/10.3760/cma.j.issn.1001-9391.2016.11.025 (2016).
Acknowledgements
This work was partially supported by the Medical Scientific Research Foundation of Guangdong, China [A2020279] and the Key Technologies R&D Program of Shenzhen, China [20160291]. We would like to acknowledge the assistance of all participants in this study. We also would like to acknowledge the contributions of the physicians, nurses, and technicians.
Author information
Authors and Affiliations
Contributions
R.R., Y.X., Z.L. and G.X. provided conception and design. R.R., Q.L., D.M.Y., Y.X., and G.X. developed the model. R.R., B.Y. and D.L. developed software. L.Z., R.R., Q.L. D.M.Y, Z.L. and G.X. provided data analysis and interpretation. L.Z., X.Z., XF.Z, J.L., Y.L. X.Y. X.J.., Y.S., Z.D.L. and Z.M.L. collected and assembled data. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhang, L., Rong, R., Li, Q. et al. A deep learning-based model for screening and staging pneumoconiosis. Sci Rep 11, 2201 (2021). https://doi.org/10.1038/s41598-020-77924-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-77924-z
This article is cited by
-
An Analysis of Deep Transfer Learning-Based Approaches for Prediction and Prognosis of Multiple Respiratory Diseases Using Pulmonary Images
Archives of Computational Methods in Engineering (2024)
-
Artificial Intelligence Techniques to Predict the Airway Disorders Illness: A Systematic Review
Archives of Computational Methods in Engineering (2023)
-
The significance of serum S100 calcium-binding protein A4 in silicosis
BMC Pulmonary Medicine (2022)
-
Use data augmentation for a deep learning classification model with chest X-ray clinical imaging featuring coal workers' pneumoconiosis
BMC Pulmonary Medicine (2022)
-
Pneumoconiosis computer aided diagnosis system based on X-rays and deep learning
BMC Medical Imaging (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.