Abstract
Vancomycin-resistant enterococci are a global challenge currently as reported by the World Health Organization. It is also important to recognize that combating antimicrobial resistance needs to recognize the interconnections between people, animals, plants and their shared environment in creating public health, the so-called One Health approach. Although the presence of VRE has been described in many regions of the world, there is a lack of comprehensive data indicating their prevalence of in Africa. Therefore, this study aimed to aggregate the result of studies describing VRE reported across multiple regions in Africa. A literature search was conducted on PubMed, Google scholar, and Hinari with the term “Vancomycin resistance enterococcus in Africa” on August 1–3, 2019. All available articles were downloaded to “Endnote version 7.1” then to Microsoft Word 2013. Articles determined to meet our criteria for the review was extracted to Microsoft Excel 2013. Those articles that reported the prevalence of vancomycin resistance Enterococcus obtained from all sample types and published from 2010 to 2019 in the English language were included for the review. A meta-analysis was conducted with OpenMetaAnalyst version R.3.1.0 software. The effect size was determined using a binary random effect model and statically significant considered when p < 0.05. Heterogeneity determined with the inconsistency index. A leave one out analysis used to perform the sensitivity analysis. There were 151 articles identified from the database searches; of this, 36 articles included after extensive review with two independent authors. Out of 4073 samples collected, 1488 isolates identified with an overall pooled prevalence of VRE 26.8% (95% CI; 10.7–43.0%) in Africa with a one-health perspective. The analysis showed that considerable heterogeneity among the studies (I2 = 99.97%; p < 0.001). Subgroup analysis in-country, African region, laboratory method, year of publication, and sample source showed that a high prevalence was identified from South Africa (74.8%), South African regions (74.8%), PCR (959.2%), 2010–2015 years (30.3%) and environmental (52.2%), respectively. This meta-analysis indicates that there was a high-pooled prevalence of vancomycin-resistant enterococci in African. A lot should be done to prevent and control the transmission of vancomycin resistance enterococci to a human being from the environment in the continent.
Similar content being viewed by others
Introduction
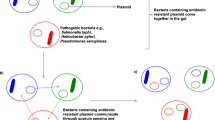
Vancomycin-resistant enterococci are defined as members of the genus, Enterococcus, that possess either intrinsic or acquired resistance to the antibiotic vancomycin used to treat serious infections caused by these bacteria1,2. Intrinsic resistance occurs in isolates of E. gallinarum and E. casseliflavus /E. flavescens, which demonstrate an innate, low-level resistance to vancomycin. These enterococci rarely cause infections in humans or animals. In contrast, high-level vancomycin resistance, most commonly seen in E. feacium and E. faecalis, may be associated with serious, life-threatening infections. High-level vancomycin resistance has also been identified in E. raffinosus, E. avium, E. durans, and several other enterococci, however, these species are rarely associated with infections. A variety of transferable genetic elements designated vanA, vanB, vanC, vanD, and vanE, may lead to resistance to vancomycin in enterococci, however, vanA and vanB are most common3,4.
VRE emerged as important nosocomial pathogens in the 1980s, and there is concern that they may be, or become, endemic in the non-hospital setting, both in human and animal reservoirs and in the general environment5. It advanced to an inoffensive colonizer of the gut of humans and animals, ranging from insects to reptiles, birds, and mammals. Whilst they are ubiquitous, they represent a minority population of the healthy human microbiome6. Presence in the environment is generally considered an indicator of human or animal faecal contamination of recreational or drinking water7.
The rise of VR Enterococcus feacium in the European Union has to lead to the sanction of avoparcin, an antibiotic that chemically related to vancomycin8. The USA never approved avoparcin for clinical use. In the years post-ban, VRE surveillance data of EU hospitals showed no obvious reduction in VRE rates. Because of very limited alternatives to vancomycin, VRE infections remain a serious clinical treatment challenge throughout the world. Surveillance studies showed zero rates of VRE in US livestock. Whole-genome sequencing data suggest that VRE might have evolved from ampicillin-resistant E. feacium from dogs9.
Some members of the genus Enterococcus are well-documented pathogens associated with serious clinical manifestations in humans, including bacteremia, infective endocarditis, intra-abdominal and pelvic infections, urinary tract infections, and, in rare cases, central nervous system infections10,11,12. Infection with VREs is associated with an increased mortality rate, illustrated by a 2.5-fold increase in mortality for patients suffering from VRE bacteremia13.
The One Health Commission defines One Health as “a collaborative, multisectoral, and trans-disciplinary approach—working at local, regional, national, and global levels—to achieve optimal health and well-being outcomes recognizing the interconnections between people, animals, plants and their shared environment.” All potentially constitute overlapping reservoirs of antimicrobial resistance14. Given the serious health threat, a common understanding of AMR, and of the need for a One Health approach to tackle it, are of fundamental importance15,16.
VRE is one of these multidrug resistances that need comprehensive data that indicates the pooled prevalence of VRE in Africa. Therefore, this study aimed to compile available data of VRE in Africa in a one-health perspective: a systematic review and meta-analysis.
Methods
Literature search strategy
A literature search conducted on PubMed, Google scholar, and Hinari with the term “Vancomycin resistance enterococcus in Africa” on August 1–3, 2019. Citations of all available articles were exported to “Endnote version 7.1” then to Microsoft Word 2013. All the articles that met our inclusion criteria were included for systematic review and meta-analysis. There were 151 articles obtained from the databases. Of these, 29 articles were excluded based on setting and duplications, 66 were excluded because either title or year of publication was unacceptable. A total of 56 articles underwent full-text assessment. An additional 20 articles were excluded because they failed to report the prevalence of VRE, their year of publication was before 2010, or they lacked clearly described laboratory methods or unknown sample source. Finally, 36 articles were subjected to an extensive review by two independent authors. The article selection process was conducted according to the PRISMA protocol of 201517 (Fig. 1).
Eligibility criteria
The inclusion criteria for this review were articles published in the English language that reported the prevalence of VRE and were published from 2010 through 2019. All sample sources were considered. Specifically, publications excluded from this review were any of the following: published before 2010 or after 2019, published other than the English language, has no clear laboratory methods, had unknown sample types, or failed to include the prevalence of VRE.
Data analysis
Microsoft Excel 2013 was used for data extraction and results were then exported to Microsoft Word plus 2013. Data was entered to OpenMetaAnalyst version R.3.1.018 software for each study after copying each column from Excel to the software and a meta-analysis, subgroup meta-analysis and sensitivity analysis were conducted. The result was presented as a forest plot in the figure. The pooled prevalence of VRE at 95% CI was determined with a binary random-effect model by the DerSimonian–Laird method. The statistical significance was considered when p value < 0.05.
Data quality
The quality of the study included in the review and meta-analysis evaluated with a 14 point scoring tool, an NIH quality assessment tool for observational and cross-sectional studies in which studies categorized as a good, fair, and poor quality based on the internal validity of each article19. Accordingly, nine (25%) articles were categorized as fair, eleven (30.6%) as poor, and sixteen (44.4%) articles as good quality (supplementary file).
Heterogeneity and publication bias
The heterogeneity of the publication was determined with the measure of the inconsistency index (I2) and p value. The total variations in the articles were due to heterogeneity rather than by chance with a value of < 30%, 30–60%, 61–75%, and > 75% suggestive of low, moderate, substantial, and considerable heterogeneity, respectively20. Publication bias was not checked as the study is considerably heterogeneous as recommended by Hak et al.21, if the data is heterogeneous no need of conducting publication bias.
Study features
Studies conducted in African countries that reported the prevalence of VRE and were published between 2010 and 2019 were considered. All sample types and laboratory methods employed were included for the review and meta-analysis. The following data types were extracted from each article and presented in Table 1: author name; year of publication; country of origin, source of sample (human, animal, environmental); laboratory method used (culture and polymerase chain reaction (PCR), PCR only, culture, number of different Enterococcus species isolated and the number of VRE isolated.
Country of origin for the articles
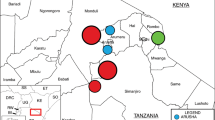
The country in which the articles originated is indicated as follows, eight articles from each country, Ethiopia22,23,24,25,26,27,28,29, and South Africa29,30,31,32,33,34,35,36,37, four articles in each country Egypt38,39,40,41, and Tunisia42,43,44,45. Another three articles from each of these countries Morocco46,47,48 and Uganda49,50,51 and two articles from each of these countries Nigeria52,53, Tanzania54,55 and Algeria56,57 were included for the study (Table 1).
Result
The pooled prevalence of vancomycin resistance Enterococci
Out of 4073 enterococci isolates described in papers meeting inclusion criteria, 1488 were identified as VRE with an overall pooled prevalence of 26.8% (95% CI; 10.7–43.0, I2 = 99.97%; p < 0.001) in Africa in a one-health perspective. The meta-analysis indicates that there was considerable heterogeneity among the articles with a consistency index (I2) = 99.97% (Fig. 2).
Sensitivity analysis
Sensitivity analysis was performed with leave one out analysis showed that there is no difference compared to pooled prevalence 26.8% (95% CI: 10.7–43.0, p < 0.001) versus 26.8% (95% CI; 10.7–43.0, p < 0.001) (Fig. 3).
Subgroup analyses
The subgroup analysis performed based on country indicates that the highest prevalence of VRE was in South Africa 74.8% (95% CI; 51–99%; I2 = 99.9%; p < 0.001) observed followed by, Egypt 37.2% (95% CI; − 17–92%; I2 = 99.7%; p < 0.001), Uganda 9.8% (95% CI; − 0.027–0.223%; I2 = 90.2%; p < 0.001), Morocco 8.2% (95% CI = − 3.0–20.0%; I2 = 88.7%; p < 0.001), Ethiopia 7.9% (95% CI; 5.0–11.0% ; I2 = 60.7%; p = 0.01), Tunisia 6.5% (95% CI; 4.0–9.0%, I2 = 0; p = 0.95), Tanzania 6.1% (95% CI; 3.4–8.8%; I2 = 9.27%; p < 0.294), Nigeria 2.8% (95% CI; − 3.0–9.0%; I2 = 79.1%; p = 0.03) and Algeria 2.8% (95% CI; 1.0–5.0%; I2 = 0; p = 0.71).
Our study conducted a subgroup analysis of VRE based on the laboratory method employed by each study. Accordingly, the laboratory methods grouped into culture and polymerase chain reaction (PCR), PCR only and culture only. It indicates that the highest rates of VRE are from article conducted with PCR 59.2% (95% CI: − 6.8–125.3%; I2 = 99.3%; p < 0.001), followed by culture and PCR 38.9% (95% CI: 16.1–61.6%; I2 = 99.9% p < 0.001) and culture only 7.3% (95% CI: 4.8–9.8%; I2 = 72.2%; p < 0.00).
Our study tried to perform a subgroup analysis of the prevalence of VRE dividing the study publication year into two categories as 2010–2015 and 2016–2019. Accordingly, the prevalence of VRE was higher in the range of 2010–2015 as compared to 2016–2019 (30.3% vs. 25.1%) which is statically significant with p < 0.000.
Studies included for our review were from four African regions as defined by African Union commission: South, North, West, and East Africa. No studies were found from countries in the Central Africa Region. The greatest numbers of studies (75%) were from the North an East Africa Regions. Hence, our subgroup analysis indicates that a higher prevalence of VRE was from South African regions 74.8% (95% CI: 51.1–98.5%, I2 = 99.9%, p < 0.001) followed by, East 17.9% (95% CI: 1.1–34.8%, I2 = 99.5%, p < 0.001), North 15.9% (95% CI: − 0.6–32.4%, I2 = 99.3%, p < 0.001) and West 2.8% (95% CI: − 3.3–9.0%, I2 = 79.1%, p = 0.02). The difference is statically significant with p < 0.000.
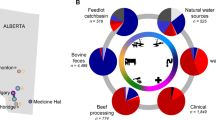
Subgroup analysis performed based on the source of sample categorizing as non- human and human source. It indicates that a higher prevalence of VRE detected from environmental sample sources 52.2% (95% CI: 22.5–82.0%, I2 = 99.6, p < 0.001) followed by animal 30.5% (95% CI: 8.4–52.5%, I2 = 99.9%, p < 0.001), human 10.2% (95% CI: 6.8–13.7%, I2 = 84.5%, p < 0.001) and animal and human 3.7% (95% CI: − 1.2–8.6%, I2 = 85.2%, p < 0.001) (Table 2).
Discussions
Vancomycin is one of a limited number of antibiotics that can be used to treat infections in humans resulting from Gram-positive multidrug-resistant organisms (MDRO) including Enterococci58. In the late 1980s, the emergence of VRE in European hospitals followed by isolation from Danish raw minced pork and frozen poultry generated global concern59. One Health is the concept that the optimum health for people, animals, and the environment should all be considered through the ongoing cooperative efforts of scientists and practitioners in a variety of disciplines60.
Our study based on the data available from studies in Africa on VRE in which animal, human, and environmental sources of samples had been specified were analyzed to determine the pooled prevalence of VRE. The overall prevalence of VRE was (26.8%) in Africa from different sample sources. This prevalence is higher than reported in the studies conducted in Iraq (14%)61, Europe (2.7%)1, (13%)62, Thailand (10.3%)63, South America (6%)64. However, it is comparable to a study from Latin America (30%)65. These differences may be due to the source of the sample we used for the analysis is from different sources and may be due to the enterococcus population structure selected overtime which is highly resistant for environmental conditions and different antibiotics2,66.
The subgroup analysis at the country level showed that there is a pronounced difference of VRE in different countries, which ranged from (74.8%) in South Africa to (2.8%) in Algeria and Nigeria which is statically significant with p < 0.000. This variation might be due to sampling source difference, sample size, laboratory method used, year of publication, and the number of studies included for the meta-analysis.
Our study also performed a subgroup meta-analysis based on the laboratory method used for isolation and identification of VRE. It showed that the higher the technique engaged by the studies for identification of VRE, the more sensitive and specific for detection of VRE in which studies conducted with PCR primers for isolation of higher prevalence of VRE, whereas those conducted with conventional culture were less likely to detect VRE. Some studies reported in a comparison of PCR and culture supports that the former technique is more sensitive and specific than later one for the identification of VRE67,68,69.
Our study revealed that a reduction of VRE from (2010–2015) to (2016–2019) (30.3%) versus (25.1%). In contrast to our finding a study from Europe bared increment in VRE from 2012 to 2018 which was (8.1%) to (19.1%)62, in Brazil from 2006 to 2009 from (2.5%) to (15.5%)70. The disagreement might be due to study period variance, the area covered, sample type used, the ability of detection of laboratories dissimilarity.
Analysis of VRE in African regions showed that there was a high prevalence in the South African region (74.8%) almost twenty-six times of West Africa and four times than of North and East African regions. The difference can be explained it might be due to the laboratory method used for detection and identification of VRE67,69, the sample difference71 and overall antibiotics usage in human72,73 and animal74,75,76.
The sample source in which we categorized in human, animal and environmental sources for the sake of subgroup meta-analysis showed that a higher prevalence of VRE was isolated from environmental, followed by the animal source as compared to a human source. This may be due to most of the articles included based on our inclusion criteria is from non-human sources as different wild and domestic animal wastes or by-products, poultry, birds, and the environmental sample was compiled for analysis. The other reason is probably due to the intensive conditions in which these animals maintained for different antibiotics as a kind of growth promoter77,78. This part of the study strained the one health approach, which is an important way of combating antibiotics resistance that worsens the world; now a day’s77,79.
Strength and limitation of the study
The strength of our review and meta-analysis is, it presented an all-inclusive data about VRE in Africa. It offered a subgroup analysis of data based on country, laboratory method used, African regions, year of publication, and source of the sample. Even if we included the most common databases for searching, our data has a limitation of addressing all search engines. It also did not identify which species of enterococci with resistances are commonly reported.
Conclusion
Our meta-analysis finding demonstrated that there is a high prevalence of VRE circulating in Africa. The subgroup analysis indicates that a high prevalence of VRE isolated from South African region. Similarly, studies conducted with PCR laboratory method isolated the highest VRE. Additionally, our study showed that the prevalence decreasing over time. Environmental sample source is with a higher VRE as compared to the human and animal sample source. Thus, a means of prevention and control targeting humans, animals, and environments based on regional, and country perspectives should be practised in the continent to alleviate the infection with VRE.
Data availability
All the data supporting the findings can be obtained from the corresponding author.
References
Schouten, M., Hoogkamp-Korstanje, J., Meis, J., Voss, A. & Group, E. V. S. Prevalence of vancomycin-resistant enterococci in Europe. Eur. J. Clin. Microbiol. Infect. Dis. 19, 816–822 (2000).
Ramadhan, A. & Hegedus, E. Survivability of vancomycin-resistant enterococci and fitness cost of vancomycin resistance acquisition. J. Clin. Pathol. 58, 744–746 (2005).
Werner, G. et al. Emergence and spread of vancomycin resistance among enterococci in Europe. Eurosurveillance 13, 19046. https://doi.org/10.2807/ese.13.47.19046-en (2008).
Widmer, A.F.-X. Vancomycin-resistant enterococci: an ongoing challenge for infection control. Swiss Med. Wkly. 142, w13554. https://doi.org/10.4414/smw.2012.13554 (2012).
Leong, K. W. et al. Emergence of vancomycin-resistant Enterococcus faecium at an Australian hospital: a whole-genome sequencing analysis. Sci. Rep. 8, 6274 (2018).
Lebreton, F., Willems, R. J. & Gilmore, M. S. in Enterococci: from commensals to leading causes of drug-resistant infection [Internet] (Massachusetts Eye and Ear Infirmary, 2014).
Gouliouris, T. The Relative Importance of Human and Animal Sources of Vancomycin-Resistant Enterococcus faecium in Immunocompromised Patients in the Hospital in Immunocompromised Patients in the Hospital (University of Cambridge, Cambridge, 2019).
Tchente Nguefack, C. et al. Clinical presentation, risk factors and pathogens involved in bacteriuria of pregnant women attending antenatal clinic of 3 hospitals in a developing country: a cross-sectional analytic study. BMC Pregnancy Childbirth 19, 143. https://doi.org/10.1186/s12884-019-2290-y (2019).
McEwen, S. A. & Collignon, P. J. Antimicrobial resistance: a one health perspective. Antimicrob. Resist. Bacteria Livestock Compan. Anim. 521–547 (2018).
Sievert, D. et al. National Healthcare Safety Network (NHSN) Team and Participating NHSN Facilities. Antimicrobial-resistant pathogens associated with healthcare-associated infections: summary of data reported to the National Healthcare Safety Network at the Centers for Disease Control and Prevention, 2009–2010. Infect. Control Hosp. Epidemiol. 34, 1–14 (2013).
Hill, E. E. et al. Infective endocarditis: changing epidemiology and predictors of 6-month mortality: a prospective cohort study. Eur. Heart J. 28, 196–203 (2006).
Wang, J. S., Muzevich, K., Edmond, M. B., Bearman, G. & Stevens, M. P. Central nervous system infections due to vancomycin-resistant enterococci: case series and review of the literature. Int. J. Infect. Dis. 25, 26–31 (2014).
DiazGranados, C. A., Zimmer, S. M., Mitchel, K. & Jernigan, J. A. Comparison of mortality associated with vancomycin-resistant and vancomycin-susceptible enterococcal bloodstream infections: a meta-analysis. Clin. Infect. Dis. 41, 327–333 (2005).
Anteneh, Z. A., Andargie, K. & Tarekegn, M. Prevalence and determinants of acute diarrhea among children younger than five years old in Jabithennan District, Northwest Ethiopia, 2014. BMC Public Health 17, 1–8 (2017).
Hakanen, A., Jalava, J. & Kaartinen, L. The National Action Plan on Antimicrobial Resistance 2017–2021. https://julkaisut.valtioneuvosto.fi/handle/10024/161022 (2017).
Commission, O. H. Definitions of One Health. https://www.onehealthcommission.org/en/why_one_health/what_is_one_health/ (2019).
Moher, D. et al. Preferred reporting items for systematic review and meta-analysis protocols (PRISMA-P) 2015 statement. Syst. Rev. 4, 1. https://doi.org/10.1186/2046-4053-4-1 (2015).
Wallace, B. C., Issa J. Dahabreh, Thomas A. Trikalinos, Joseph Lau, Paul Trow, and Christopher H. Schmid. . OpenMetaAnalyst: "Closing the gap between methodologists and end-users: R as a computational back-end." J. Stat. Softw. 49 (2012).
National, N. Quality Assessment Tool for observational cohort and cross-sectional studies. https://www.nhlbi.nih.gov/health-topics/study-quality-assessment-tools (2017).
Singh, S. How to conduct and interpret systematic reviews and meta-analyses. Clin. Transl. Gastroenterol. 8, e93 (2017).
Hak, T., Van Rhee, H. J., & Suurmond, R. How to interpret the results of a meta-analysis. (Version 1.3) Rotterdam, The Netherlands: Erasmus Rotterdam Institute of Management, www.erim.eur.nl/researchsupport/meta-essentials/downloads (2016).
Abamecha, A., Wondafrash, B. & Abdissa, A. Antimicrobial resistance profile of Enterococcus species isolated from intestinal tracts of hospitalized patients in Jimma Ethiopia. BMC Res. Notes 8, 213. https://doi.org/10.1186/s13104-015-1200-2 (2015).
Abebe, W., Endris, M., Tiruneh, M. & Moges, F. Prevalence of vancomycin-resistant Enterococci and associated risk factors among clients with and without HIV in Northwest Ethiopia: a cross-sectional study. BMC Public Health 14, 185. https://doi.org/10.1186/1471-2458-14-185 (2014).
Agegne, M., Abera, B., Derbie, A., Yismaw, G. & Shiferaw, M. B. Magnitude of vancomycin-resistant enterococci (VRE) colonization among HIV-infected patients attending ART clinic in West Amhara Government Hospitals. Int. J. Microbiol. https://doi.org/10.1155/2018/7510157 (2018).
Ali, S., Alemayehu, M., Dagnew, M. & Gebrecherkos, T. Vancomycin-resistant enterococci and its associated risk factors among HIV-positive and -negative clients attending dessie referral hospital, Northeast Ethiopia. Int. J. Microbiol. 4753460, 1–9. https://doi.org/10.1155/2018/4753460 (2018).
Ferede, Z. T., Tullu, K. D., Derese, S. G. & Yeshanew, A. G. Prevalence and antimicrobial susceptibility pattern of Enterococcus species isolated from different clinical samples at Black Lion Specialized Teaching Hospital, Addis Ababa Ethiopia. . BMC Res. Notes 11, 793. https://doi.org/10.1186/s13104-018-3898-0 (2018).
Solomon, F. B. et al. Antibiotic-resistant airborne bacteria and their multidrug resistance pattern at University teaching referral Hospital in South Ethiopia. Ann. Clin. Microbiol. Antimicrob. 16, 29. https://doi.org/10.1186/s12941-017-0204 (2017).
Toru, M. et al. Prevalence and phenotypic characterization of Enterococcus species isolated from clinical samples of pediatric patients in Jimma University Specialized Hospital, south-west Ethiopia. BMC Res. Notes 11, 281. https://doi.org/10.1186/s13104-018-3382-x (2018).
Yilema, A. et al. Isolation of enterococci, their antimicrobial susceptibility patterns and associated factors among patients attending at the University of Gondar Teaching Hospital. BMC Infect. Dis. 17, 276–276. https://doi.org/10.1186/s12879-017-2363-3 (2017).
Ateba, C. N., Lekoma, K. P. & Kawadza, D. T. Detection of vanA and vanB genes in vancomycin-resistant enterococci (VRE) from groundwater using multiplex PCR analysis. J. Water Health 11, 684–691. https://doi.org/10.2166/wh.2013.037 (2013).
Iweriebor, B. C., Gaqavu, S., Obi, L. C., Nwodo, U. U. & Okoh, A. I. Antibiotic susceptibilities of Enterococcus species isolated from hospital and domestic wastewater effluents in Alice, eastern cape province of South Africa. Int. J. Environ. Res. Public Health 12, 4231–4246. https://doi.org/10.3390/ijerph120404231 (2015).
Iweriebor, B. C., Obi, L. C. & Okoh, A. I. Virulence and antimicrobial resistance factors of Enterococcusspp. isolated from faecal samples from piggery farms in Eastern Cape, South Africa. BMC Microbiol. 15, 136. https://doi.org/10.1186/s12866-015-0468-7 (2015).
Iweriebor, B. C., Obi, L. C. & Okoh, A. I. Macrolide, glycopeptide resistance and virulence genes in Enterococcus species isolates from dairy cattle. J. Med. Microbiol. 65, 641–648. https://doi.org/10.1099/jmm.0.000275 (2016).
Matlou, D. P. et al. Virulence profiles of vancomycin-resistant enterococci isolated from surface and groundwater utilized by humans in the North West Province South Africa: a public health perspective . Environ. Sci. Pollut. Res. 26, 15105–15114. https://doi.org/10.1007/s11356-019-04836-5 (2019).
Molale, L. G. & Bezuidenhout, C. C. Antibiotic resistance, efflux pump genes and virulence determinants in Enterococcus spp. from surface water systems. Environ. Sci. Pollut. Res. Int. 23, 21501–21510. https://doi.org/10.1007/s11356-016-7369-7 (2016).
Molechan, C. et al. Molecular epidemiology of antibiotic-resistant Enterococcus spp. from the farm-to-fork continuum in intensive poultry production in KwaZulu-Natal, South Africa. Sci. Total Environ. 692, 868–878. https://doi.org/10.1016/j.scitotenv.2019.07.324 (2019).
Tatsing Foka, F. E. & Ateba, C. N. Detection of virulence genes in multidrug-resistant Enterococci isolated from feedlots dairy and beef cattle: implications for human health and food safety. BioMed Res. Int. https://doi.org/10.1155/2019/5921840 (2019).
Hammad, A. M., Hassan, H. A. & Shimamoto, T. Prevalence, antibiotic resistance and virulence of Enterococcus spp. in Egyptian fresh raw milk cheese. Food Control 50, 815–820. https://doi.org/10.1016/j.foodcont.2014.10.020 (2015).
Hassan, R. M., Ghaith, D. M., Ismail, D. K. & Zafer, M. M. Reduced susceptibility of Enterococcus spp. isolates from Cairo University Hospital to tigecycline: Highlight on the influence of proton pump inhibitors. J. Glob. Antimicrob. Resist. 12, 68–72. https://doi.org/10.1016/j.jgar.2017.12.005 (2018).
Moemen, D., Tawfeek, D. & Badawy, W. Healthcare-associated vancomycin-resistant Enterococcus faecium infections in the Mansoura University Hospitals intensive care units Egypt. Braz. J. Microbiol. [Publ. Braz. Soc. Microbiol. 46, 777–783. https://doi.org/10.1590/s1517-838246320140403 (2015).
Osman, K. M. et al. Poultry as a vector for emerging multidrug-resistant Enterococcus spp.: the first report of vancomycin (van) and the chloramphenicol-florfenicol (cat-fex-cfr) resistance genes from pigeon and duck faeces. Microb. Pathog. 128, 195–205. https://doi.org/10.1016/j.micpath.2019.01.006 (2019).
Ben Said, L. et al. Prevalence, antimicrobial resistance and genetic lineages of Enterococcus spp. from vegetable food, soil and irrigation water in farm environments in Tunisia. J. Sci. Food Agric. 96, 1627–1633. https://doi.org/10.1002/jsfa.7264 (2016).
Ben Yahia, H. et al. Antimicrobial resistance and genetic lineages of faecal enterococci of wild birds: Emergence of vanA and vanB2 harbouring Enterococcus faecalis. Int. J. Antimicrob. Agents 52, 936–941. https://doi.org/10.1016/j.ijantimicag.2018.05.005 (2018).
Dziri, R. et al. Multidrug-resistant enterococci in the hospital environment: detection of novel vancomycin-resistant E. faecium clone ST910. J. Infect. Dev. Countries 10, 799–806. https://doi.org/10.3855/jidc.8014 (2016).
Naouel, K. et al. Diversity of species and antibiotic resistance among faecal enterococci from wild birds in Tunisia. Detection of vanA-containing Enterococcus faecium isolates. Eur. J. Wildl. Res. https://doi.org/10.1007/s10344-014-0884-2 (2014).
Bouamamaa, L. et al. Antibiotic resistance patterns of bacterial strains isolated from Periplaneta americana and Musca domestica in Tangier, Morocco. J. Infect. Dev. Countries 4, 194–201 (2010).
Bouymajane, A. et al. Occurrence, molecular and antimicrobial resistance of Enterococcus spp. isolated from raw cow’s milk trade by street trading in Meknes city, Morocco. Germs 8, 77–84. https://doi.org/10.18683/germs.2018.1134 (2018).
Hannaoui, I. et al. Intestinal carriage of vancomycin-resistant enterococci in a community setting in Casablanca Morocco. J. Glob. Antimicrob. Resist. 6, 84–87. https://doi.org/10.1016/j.jgar.2016.03.008 (2016).
Kateete, D. P. et al. Species, antibiotic susceptibility profiles and van gene frequencies among enterococci isolated from patients at Mulago National Referral Hospital in Kampala Uganda. BMC Infect. Dis. 19, 486. https://doi.org/10.1186/s12879-019-4136-7 (2019).
Kateete, D. P. et al. Prevalence and antimicrobial susceptibility patterns of bacteria from milkmen and cows with clinical mastitis in and around Kampala Uganda. PLoS ONE 8, e63413. https://doi.org/10.1371/journal.pone.0063413 (2013).
Ngonzi, J. et al. Risk factors for vaginal colonization and relationship between bacterial vaginal colonization and in-hospital outcomes in women with obstructed labor in a Ugandan Regional Referral Hospital. Int. J. Microbiol. https://doi.org/10.1155/2018/6579139 (2018).
Anyanwu, M. Prevalence and antibiogram of generic enterococci in ready-to-slaughter beef cattle. Notulae Sci. Biol. 7, 390–399. https://doi.org/10.15835/nsb749681 (2015).
Ngbede, E. O., Raji, M. A., Kwanashie, C. N. & Kwaga, J. K. P. Antimicrobial resistance and virulence profile of enterococci isolated from poultry and cattle sources in Nigeria. Trop. Anim. Health Prod. 49, 451–458. https://doi.org/10.1007/s11250-016-1212-5 (2017).
Katakweba, A. A. S. et al. Antimicrobial resistance in faecal samples from buffalo, wildebeest and zebra grazing together with and without cattle in Tanzania. J. Appl. Microbiol. 118, 966–975 (2015).
Katakweba, A. A. S. et al. First report on a randomized investigation of antimicrobial resistance in faecal indicator bacteria from Livestock, Poultry, and humans in Tanzania. Microb. Drug Resist. 24, 260–268 (2018).
Djahmi, N. et al. Molecular epidemiology of Enterococcus sp. isolated in a university hospital in Algeria. Scand. J. Infect. Dis. 44, 656–662. https://doi.org/10.3109/00365548.2012.673232 (2012).
Bourafa, N. et al. Identification of vancomycin-susceptible major clones of clinical Enterococcus from Algeria. J. Glob. Antimicrob. Resist. 6, 78–83. https://doi.org/10.1016/j.jgar.2016.03.009 (2016).
Faron, M. L., Ledeboer, N. A. & Buchan, B. W. Resistance mechanisms, epidemiology, and approaches to screening for vancomycin-resistant enterococcus in the health care setting. J. Clin. Microbiol. 54, 2436–2447 (2016).
McEwen, S. A. & Collignon, P. J. Antimicrobial resistance: a one health perspective. Microbiol. Spectr. https://doi.org/10.1128/microbiolspec.ARBA-0009-2017 (2018).
Kahn, L. H. Antimicrobial resistance: a one health perspective. Trans. R. Soc. Trop. Med. Hyg. 111, 255–260. https://doi.org/10.1093/trstmh/trx050 (2017).
Moghimbeigi, A. et al. Prevalence of vancomycin resistance among isolates of enterococci in Iran: a systematic review and meta-analysis. Adolesc. Health Med. Ther. 9, 177 (2018).
Ayobami, O., Willrich, N., Reuss, A., Eckmanns, T. & Markwart, R. The ongoing challenge of vancomycin-resistant Enterococcus faecium and Enterococcus faecalis in Europe: an epidemiological analysis of bloodstream infections. Emerg. Microbes Infect. 9, 1180–1193 (2020).
Daniel, D. S., Lee, S. M., Dykes, G. A. & Rahman, S. Public health risks of multiple-drug-resistant Enterococcus spp Southeast Asia. Appl. Environ. Microbiol. 81, 6090–6097. https://doi.org/10.1128/AEM.01741-15 (2015).
Panesso, D. et al. Molecular epidemiology of vancomycin-resistant Enterococcus faecium: a prospective, multicenter study in South American hospitals. J. Clin. Microbiol. 48, 1562–1569 (2010).
Centres for Disease Control and Prevention (US). Antibiotic resistance threats in the United States. Centres for Disease Control and Prevention, US Department of Health and Human Services. https://www.cdc.gov/drugresistance/threat-report-2013/pdf/ar-threats-2013-508.pdf (2013).
Eliopoulos, G. M. & Gold, H. Vancomycin-resistant enterococci: mechanisms and clinical observations. Clin. Infect. Dis. 33, 210–219 (2001).
Deschaght, P. et al. Comparison of the sensitivity of culture, PCR and quantitative real-time PCR for the detection of Pseudomonas aeruginosain sputum of cystic fibrosis patients. BMC Microbiol. 9, 244 (2009).
Sloan, L. et al. Comparison of the Roche LightCycler vanA/vanB detection assay and culture for detection of vancomycin-resistant enterococci from perianal swabs. J. Clin. Microbiol. 42, 2636–2643 (2004).
d’Azevedo, P. A. et al. Rapid detection of vancomycin-resistant enterococci (VRE) in rectal samples from patients admitted to intensive care units. Braz. J. Infect. Dis. 13, 289–293 (2009).
Conceição, N., Oliveira, C. D. C. H. B. D., Silva, P. R. D., Ávila, B. G. M. & Oliveira, A. G. D. Trends in antimicrobial resistance among clinical isolates of enterococci in a Brazilian tertiary hospital: a 4-year study. Rev. Soc. Bras. Med. Trop. 44, 177–181 (2011).
Ubeda, C. et al. Vancomycin-resistant Enterococcus domination of intestinal microbiota is enabled by antibiotic treatment in mice and precedes bloodstream invasion in humans. J. Clin. Investig. 120, 4332–4341 (2010).
de Bruin, M. A. & Riley, L. W. Does vancomycin prescribing intervention affect vancomycin-resistant enterococcus infection and colonization in hospitals? A systematic review. BMC Infect. Dis. 7, 24 (2007).
Edmond, M. B. et al. Vancomycin-resistant Enterococcus faecium bacteremia: risk factors for infection. Clin. Infect. Dis. 20, 1126–1133 (1995).
DeLisle, S. & Perl, T. M. Vancomycin-resistant enterococci: a road map on how to prevent the emergence and transmission of antimicrobial resistance. Chest 123, 504S-518S (2003).
Klare, I. et al. Decreased incidence of VanA-type vancomycin-resistant enterococci isolated from poultry meat and faecal samples of humans in the community after discontinuation of avoparcin usage in animal husbandry. Microb. Drug Resist. 5, 45–52 (1999).
Levy, S. Reduced antibiotic use in livestock: how Denmark tackled resistance. Environ. Health Perspect. 122, A160–A165. https://doi.org/10.1289/ehp.122-A160 (2014).
Butaye, P., Devriese, L. A. & Haesebrouck, F. Antimicrobial growth promoters used in animal feed: effects of less well-known antibiotics on gram-positive bacteria. Clin. Microbiol. Rev. 16, 175–188. https://doi.org/10.1128/cmr.16.2.175-188.2003 (2003).
Hughes, P. & Heritage, J. Antibiotic growth-promoters in food animals. FAO Animal Production and Health Paper, 129–152 (2004).
Nadimpalli, M. et al. Combating global antibiotic resistance: emerging one health concerns in lower-and middle-income countries. Clin. Infect. Dis. 66, 963–969 (2018).
Acknowledgements
We want to acknowledge Dr. Donald R. Callihan (Ph.D.) and his wife Sally Callihan for helping as for Scientific as well as English language editing.
Author information
Authors and Affiliations
Contributions
T.A. and M.H. equally conceived the idea, conducted literature search, extracted data: T.A. performed the analysis and prepared the manuscript and M.H. revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Alemayehu, T., Hailemariam, M. Prevalence of vancomycin-resistant enterococcus in Africa in one health approach: a systematic review and meta-analysis. Sci Rep 10, 20542 (2020). https://doi.org/10.1038/s41598-020-77696-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-77696-6
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.