Abstract
Xanthomonas fragariae is a quarantine bacterial pathogen that causes angular leaf spot on strawberry. The aim of our study was to analyse the mechanism of interaction of this bacterium with its host plant at the transcriptome level. For this purpose, mRNAs of X. fragariae growing in Wilbrink’s medium and from infected strawberry cv. Elsanta plants were isolated and sequenced using the Illumina MiSeq platform. The expression profiles of the bacteria in Wilbrink’s medium and in planta were very diverse. Of the 3939 CDSs recorded, 1995 had significantly different expression in planta (966 and 1029 genes were down- and upregulated, respectively). Among the genes showing increased expression in planta, those with eggNOG/COG (evolutionary genealogy of genes: Non-supervised Orthologous Groups/Cluster of Orthologous Groups) categories associated with bacterial cell motility, signal transduction, transport and metabolism of inorganic ions and carbohydrates and transcription were overrepresented. Among the genes with the most increased expression in planta, genes primarily associated with flagella synthesis and chemotaxis were found. It is also interesting to note that out of the 31 genes localized on a plasmid, 16 were expressed differently in planta, which may indicate their potential role in plant–pathogen interactions. Many genes with differentiated expression that were localized on chromosome and plasmid encode proteins of unknown function.
Similar content being viewed by others
Introduction
Xanthomonas fragariae (Xf), the causal agent of angular leaf spot (ALS) of strawberries, is listed in Europe as a quarantine pathogen on the EPPO A2 list1. In the early phase of disease development, the symptoms of ALS are water-soaked angular spots visible on the abaxial leaf surface that enlarge over time, become reddish, necrotic and irregular and become visible on the adaxial leaf surface. When viewed with transmitted light, the spots are translucent. Under favourable conditions (high moisture and a temperature over 20 °C), a yellow bacterial exudate is produced on the leaf surface and calyces2,3.
The pathogen has a very narrow host range. It is found mostly on Fragaria × ananassa and incidentally on F. virginiana, F. vesca and two species of Potentilla–P. fruticosa and P. glandulosa, which show disease symptoms after artificial inoculation1,2. This limited host range in comparison to those of other plant pathogenic Xanthomonas species reflects a smaller genome (4.2 Mb vs. typical for this genus ~ 5 Mb) with a reduced repertoire of genes responsible for pathogenic abilities. On the other hand, the Xf genome reveals a high amount of DNA gained via horizontal gene transfer, which results in a set of uncommon genes possibly involved in pathogenicity4.
The pathogenic abilities of Xanthomonas bacteria are determined by several factors, and the major virulence-related gene regions are present in the Xf genome. Xf possesses a hrp gene cluster coding for structural elements of the type III secretion system (T3SS) and a distinct T3SS effector (T3E) set and an essential part of the gum cluster coding for xanthan—extracellular polysaccharide (EPS) synthesis4. It also contains a type IV secretion system (T4SS) and an xps-coded type II secretion system (T2SS) and, although in reduced form, TonB-dependent transporters, which do not play a direct role in pathogenesis but allow Gram-negative bacteria to take up scarce resources from nutrient-limiting environments by transporting molecules such as siderophores and hemes. Xf possesses the smallest repertoire of cell wall-degrading enzymes (CWDEs). In the Xf genome, there is a lack of xylan degradation and β-ketoadipate pathways commonly present in the majority of Xanthomonas pathogens. This can be related to the method and process of infection because Xf does not cause extensive tissue destruction as the necrotrophic pathogens with an abundance of CWDEs5.
Although X. fragariae is an important pathogen in the propagation of strawberry plant material6,7, there is limited knowledge of the molecular aspects of this plant pathogen. Generally, the genetic analysis of bacterial plant pathogen genes involved in pathogenic interactions with plants has been performed mostly through the production and screening of mutants, which usually allows the study of only a limited number of genes by “in vivo expression technology” (IVET)8. Other approaches, such as metabolomics, proteomics and DNA microarrays9, have several limitations. For instance, assigning an identity to the biomarkers can be problematic10, the detection limits may be high, and the possibility to simultaneously monitor bacterial activity may be limited11. By using next-generation sequencing, it is possible to characterize the whole transcriptome of pathogenic organisms, and this technique was successfully used for the study of both human and plant bacterial pathogens12,13.
The aim of our study was to determine the changes in gene expression of X. fragariae during infection of strawberry at the transcriptome level. For this purpose, we applied an RNA-seq technique to reveal the global changes in gene expression of X. fragariae. Until now, no detailed studies on the mechanism of action of this pathogen on hosts with different susceptibility levels have been carried out.
Results
Overview of RNA-seq results
For each biological replicate, a library was constructed and sequenced on a MiSeq sequencer (Illumina). For each sample, 4,770,496 to 6,554,258 reads were obtained, and 4,553,963 to 6,059,845 reads were mapped to the genome of X. fragariae IPO 3485 (Supplementary Table S1). The biological replicates showed a very high level of correlation (r ≥ 0.95). Principal component analysis (PCA) of the log2-transformed normalized expression values showed a large difference in values between the RNA expression of bacteria grown in planta and in liquid medium and small differences between replicate samples (Fig. 1).
The accuracy of the RNA-seq data was verified by RT-qPCR (Reverse Transcribed quantitative Polymerase Chain Reaction). Fold changes in expression values under different environmental experimental conditions obtained with these two techniques were plotted on a scatter graph, with fold change values obtained from RT-qPCR on the x-axis and those obtained from RNA-seq on the y-axis (Fig. 2). The high value for the Pearson correlation coefficient (r = 0.938; p < 0.001;) indicated a positive correlation between the two techniques.
Expression of X. fragariae IPO 3485 genes in planta
Out of 3939 annotated CDSs on the genome of X. fragariae IPO 3485 strain, 1995 were differentially expressed in planta compared to bacteria in pure culture in liquid Wilbrink’s medium. Of all DEGs (Differentially Expressed Genes), 966 and 1029 were down- and upregulated in planta, respectively (Fig. 3, Supplementary Table S2).
For every transcript, the fold change between bacteria growing in liquid Wilbrink’s medium and in planta 15 days after inoculation is plotted against the p value. Statistically significant differentially expressed genes (DEGs) with a log2 fold change ≥ 1.5 or ≤ − 1.5 are depicted as red dots, insignificant as blue dots. The numbers aside the arrow pointing up represent the number of upregulated genes and the numbers aside arrow pointing down represent the number of downregulated genes in planta.
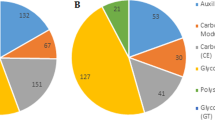
The 1995 DEGs were classified into eggNOG/COG categories. In the case of downregulated genes, 7 of 25 eggNOG/COG categories were over-represented. The over-represented categories corresponding to the lowest p values in increasing order included energy production and conversion (C), lipid transport and metabolism (I), nucleotide transport and metabolism (F), amino acid transport and metabolism (E), translation (J), intracellular trafficking and secretion (U), and cell wall/membrane/envelope biogenesis (M). Among the 1029 upregulated genes, 7 eggNOG/COG categories were over-represented. The over-represented categories with the lowest p values in increasing order included the following: cell motility (N), signal transduction mechanisms (T), function unknown (S), posttranslational modification, protein turnover, chaperones (O), inorganic ion transport and metabolism (P), transcription (K), and carbohydrate transport and metabolism (G) (Supplementary Table S3).
The introduction of X. fragariae cells to strawberry leaf tissue also influenced the metabolic pathways—KEGG (Kyoto Encyclopedia of Genes and Genomes) pathways of the bacteria. Among the downregulated pathways, energy metabolism, nucleotide metabolism and translation were mostly over-represented. On the other hand, among the upregulated genes, over-representation of genes playing roles in the pathways of cell motility and signal transduction was observed (Supplementary Table S4).
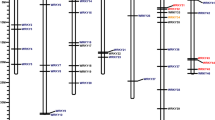
Out of all X. fragariae genes located on the chromosome and a 21 kb plasmid, in all experimental combinations, the highest abundance of transcripts was observed for the gene NBC2815_00663, which was annotated to code the OmpA family outer membrane protein, followed by the genes NBC2815_00750, coding for OmpW family protein; groL (molecular chaperone GroEL); and NBC2815_02547, coding for a hypothetical protein. Proteins OmpA and OmpW are membrane proteins.
The most upregulated X. fragariae gene in planta was NBC2815_00636 (fold change (FCh) = 105.91), coding for the synthesis of the NADPH-sulfite reductase flavoprotein subunit. Among other highly upregulated genes, genes involved in flagella synthesis and chemotaxis and genes coding for hypothetical proteins were found (Supplementary Table S5). Of the downregulated genes, the gene NBC2815_03825, coding for the synthesis of phage-related capsid scaffold protein, was the most downregulated gene in planta (FCh = − 44.97). Other highly downregulated genes included type IV secretion system genes, outer membrane adhesin protein XadA and a few genes coding for hypothetical proteins (Supplementary Table S6).
Among genes expected to be important for the pathogenicity of X. fragariae, different regulation of their expression in planta was observed4. The gumBCDE genes located in the gum cluster, encoding xanthan synthesis, were upregulated, while gumKLMN genes were downregulated. No changes in the expression of the nucleotide precursor of xanthan synthesis genes xanA and xanB (NBC2815_00824) were observed. Among the rpf genes known to be involved in the regulation of pathogenicity factors in other Xanthomonas species14, only rpfA was highly upregulated in planta, whereas rpfF and rpfG were downregulated and the rest of the rpf genes did not show any change in expression.
All genes involved in cell motility, including flagellar genes, were significantly upregulated. Out of 29 genes playing a role in chemotaxis, 26 revealed higher expression in planta than in pure culture. Among TonB-dependent transporters and receptors involved in the transport of e.g., siderophores, which are critical for successful niche settlement, different changes in expression were observed. Some of them, such as the gene fhuA or the genes NBC2815_01036 and NBC2815_02594, annotated as TonB-like proteins or receptors, were upregulated in planta (FCh = 3.99–6.97). On the other hand, the genes fyuA and iroN_2, coding for receptors, were significantly downregulated in planta (FCh < − 8.48).
All but one (hrpD6) T3SS genes were upregulated in planta. The hrpD6 gene also showed upregulation (FCh = 1.6), but the FDR (False Discovery Rate) was higher than 0.05. Among 40 genes annotated as T3SS or coding for avirulence proteins, 20 were upregulated, two (xopAF and NBC2815_00544) were downregulated, and the others showed no change in expression in planta (Supplementary Table S2).
The genes of the xps cluster constituting a T2SS showed no change in expression or were downregulated in planta. The downregulated genes were xpsD, which codes for secretin (an outer membrane protein that, as a pentadecamer, provides a pore through the membrane), and the three genes xpsIJK, coding minor pseudopilins. Generally, T2SSs secrete toxins and degradative enzymes (such as xylanases, cellulases or proteases) that are transported across the inner bacterial membrane and often contribute to the virulence of plant pathogenic bacteria, e.g., by facilitating bacterial infection by degrading plant cell walls15. Although the role of T2SSs has not yet been studied in Xf, in the genome of Xf, several genes responsible for the synthesis of degradative enzymes, such as cellulases, proteases, polygalacturonases, lyases, amylases and lipases, were found. Many of them, such as all genes coding for amylases, polygalacturonases, and xylanase and the majority of genes coding for proteases, were upregulated in planta (Supplementary Table S7).
In the X. fragariae genome, genes coding for the type VI secretion system (T6SS) were found4. Of the T6SS genes coding for machinery proteins, increased expression of three genes, impJ, tssJ (NBC2815_01934) and evpG (NBC2815_01941), was observed. In the genome of X. fragariae, several copies of genes for T6SS effectors were found: 17 for VgrG, 15 for the Rhs protein family and one PAAR protein (proline–alanine–alanine–arginine repeat protein). Of them, 15 VgrG and 8 Rhs protein-coding genes were upregulated, while PAAR protein and 2 Rhs genes were downregulated.
Xf also possesses a whole cluster of a T4SS, which is responsible for intercellular DNA transfer, and almost all the genes were differentially expressed. Out of three virD4 copies, only a traG copy located on the plasmid was upregulated, while two other copies on the chromosome were downregulated. Highly downregulated genes were involved in pilin synthesis: pilW, NBC2815_01578 and pilE (FCh > 12). A majority of virB genes were slightly upregulated. Type IV secretion systems have been shown to contribute to virulence, for instance, in Xanthomonas campestris pv. campestris16; however, its role in Xf pathogenicity is not known.
Only a limited change in the expression of multidrug efflux system genes was observed. Only three genes, NBC2815_00396, NBC2815_01613 and NBC2815_01676, were upregulated, and four, norM, NBC2815_00639, NBC2815_01397 and NBC2815_02129, were downregulated. Increased expression in strawberry plants was observed for three genes (NBC2815_01235, NBC2815_01669, NBC2815_02506) related to heat shock, which are primarily involved in defence against stress. Simultaneously, decreased expression was observed for two genes related to heat shock (NBC2815_01461, NBC2815_02622) and two related to cold shock (scoF, NBC2815_01420).
Differences in the expression of some genes coding for membrane proteins were detected between bacteria grown on medium and in planta. Membrane proteins create a selective barrier and protect bacteria from the environment by preventing entry of many toxic molecules into the cell; additionally, they are members of transport systems. Out of 126 genes annotated as coding for membrane proteins, differences in expression were observed for 58: 23 were upregulated, and 35 were downregulated in strawberry tissue (Supplementary Table S2). Among membrane protein genes with increased expression in planta, tsr6 (coding for a protein chemotaxis sensory transducer) and phuR_2 (coding for a hemin receptor) were the most upregulated, while NBC2815_03739 (coding for the adhesion protein XadA) and genes of proteins involved in LPS biosynthesis, WxcD, WxcE and WxcO, were the most downregulated in planta. The downregulation of LPS synthesis can be involved in the pathogen strategy to avoid inducing the immune response of plants because LPSs are recognized as pathogen-associated molecular patterns (PAMPs)17.
In addition to chromosomal DNA, strain IPO 3485 (NBC2815) possesses one 21.045 kb plasmid (pNBC2815-21). A total of 31 annotated genes are localized on this plasmid, and almost half of them have unknown functions. Of all plasmid genes, 17 were differentially expressed in planta: three were downregulated, and 14 were upregulated. The upregulated genes included traG (a member of the T4SS) and genes coding for recombinases and partitioning proteins. On the plasmid, genes potentially involved in pathogenicity, such as xopT (T3E) and NBC2815_04017, encode avirulent proteins, but their expression was not changed in planta.
Cyclic-di-GMP (c-di-GMP), a universal bacterial secondary messenger signalling compound, regulates different aspects of cell life, including processes involved in effective pathogenicity and responses to a variety of environmental stimuli, including stress. The effects of c-di-GMP are dose dependent, and their level is controlled by the opposing actions of diguanylate cyclases (DGC), including GGDEF domain proteins, and phosphodiesterases (PDEs), possessing EAL or HD-GYP domains. These two groups of enzymes control the synthesis and degradation of c-di-GMP, respectively18. In the genome of Xf IPO 3485, 10 genes possessing these characteristic domains were annotated, and eight of them (NBC2815_01280, NBC2815_01823, NBC2815_02702, NBC2815_03946, NBC2815_01824, NBC2815_01207, NBC2815_00405, NBC2815_00406) were upregulated in planta.
Discussion
X. fragariae is a rather weak pathogen with a relatively slow growth rate, a limited possibility of systematically colonizing plants, and a restricted capacity to survive outside plant tissue19. Moreover, the pathogen survives poorly on leaf surfaces, and a significant decrease in the population density in the first 3 days after inoculation was found6. This drop in the population can indicate that this pathogen uses a strategy of avoidance20 to escape environmental stresses and enters the plant leaves to survive. The population gradually rebuilds during later phases of infection, reaching the preparatory stage of the bacteria before the exudation phase at ca. 14 days post inoculation6. During disease development, symptoms start from single small spots of water-soaked lesions bounded by leaf vines. With time, the number of angular spots increases, and as observed by electron microscopy techniques3, bacterial cells suspended in copious amounts of EPS colonize mesophyll air spaces. In the next steps, plasmolysis of mesophyll cells and disruption of organelles occur, and bacteria ooze from stomates. In our study, we analysed Xf mRNA in planta with RNA-seq 15 days after inoculation, when well-developed water soaking symptoms were observed but strawberry tissue was not necrotized yet. We could assume that the bacterial cell stages and gene expression of the cells that were located in the water-soaked leaf parts and on the border of healthy and already diseased plant tissue were different. The bacterial population may be localized in the intercellular spaces and partially move into the vascular tissue; however, a large part is in degraded tissue. The obtained results of transcriptome analysis are thus an average expression of cells in different steps of interaction with plant tissue.
In all experimental combinations, the highest abundance of transcripts was observed for the genes coding membrane proteins OmpA and OmpW, groL (encoding molecular chaperone GroEL), and one gene of a hypothetical protein. The high number of transcripts of the gene coding OmpA can be caused by the fact that it is the most abundant outer membrane (OM) protein in many bacterial species, e.g., OmpA in Escherichia coli is present at 100,000 copies per cell21. Although this protein is also found to play a role in the pathogenesis of, e.g., Xanthomonas albilineans22, its expression was lower in planta than in Wilbrink’s medium, similar to the groL gene behaviour. Chaperone GroEL is involved in protein folding and cell proliferation/survival23. OmpW is involved in the transport of small hydrophilic molecules across the bacterial outer membrane24. Studies in Salmonella typhimurium suggest that this porin may have a role in the protection of bacteria against various forms of environmental stress by operating as an efflux channel for toxic compounds25, and in the case of Xanthomonas axonopodis pv. citri, OmpW is suspected to be involved in protecting biofilms26.
The T6SS system is widespread in Gram-negative bacteria and can deliver toxic effector proteins into eukaryotic cells or competing bacteria; it can also participate in uptake of metal ions, such as iron, manganese, and zinc. T6SS effector translocation apparatuses are very similar to an inverted bacteriophage-puncturing device composed of at least 13 proteins identified as core components27,28. In our study, three genes coding for the elements of this translocation apparatus and many coding for T6SS effectors were upregulated in planta, which suggests their importance in relation to host plants; however, the potential role of this system in the pathogenic relation of Xf is unknown.
Among the upregulated genes in planta, seven out of 25 eggNOG/COG categories were over-represented. Although the in planta Xf gene expression was analysed 15 days after strawberry leaf inoculation, the groups of upregulated Xf genes are congruent with those of Xanthomonas oryzae pv. oryzae (Xoo) in an in vitro system with rice leaf extract during the first 60 min after contact with the medium29. However, taking into account the low virulence of Xf and problems with maintaining a high population in early contact with plants, it is difficult to compare what time since inoculation can be equivalent to the first minutes of contact in the case of Xoo. In Xoo, the most over-represented categories among the differentially expressed genes were inorganic ion transport and metabolism (P) and cell motility (N). Uptake of inorganic ions, such as iron, seems to be crucial for plant pathogenic bacteria that need to compete with their host to obtain nutrients. The key regulator HrpG is a repressor of chemotaxis and flagella biosynthesis-related genes and consequent bacterial motility30, which was confirmed in a study on Xoo29. In the case of our study, motility, chemotaxis and flagella synthesis genes were all significantly upregulated together with high upregulation of hrpG and other hrp genes, which again can represent the average gene expression of Xf cells in the leaves.
Pili are involved in twitching motility and may be involved in the adhesion of Xanthomonas species31. The genes coding for the synthesis of pili are downregulated in Xf present in water-soaked lesions, possibly because the process of attachment to the host had already passed. Conversely, the genes for flagella synthesis were upregulated. For X. axonopodis pv. citri, it has been demonstrated that flagella and flagella-dependent motility are important not only for adherence to biotic and abiotic surfaces but also for maturing of the biofilm, symptom development and extension and dispersion of the biofilms32, indicating that maturation and extension of the biofilm in the lesions still takes place. The upregulation of chemotaxis and flagella genes may also indicate a preparation for a release and dispersal of the pathogen via moisture from the lesions33.
Looking at the expression of genes known as important for pathogenicity, out of 12 genes of xanthan synthesis, only four gumBCDE were upregulated, the next four genes in the operon showed no change in expression, and the last four genes were downregulated. In xanthan biosynthesis, the upregulated gumBCE genes are responsible for polymerization and/or export of the polymer, while the others are mostly responsible for the synthesis34, so it seems that at this stage of plant infection, synthesis of xanthan was not relatively intensive. Out of the rpf gene cluster, which is known to be involved in the regulation of pathogenicity factors, an increase in expression was observed only for the rpfA gene. Mutation of this gene resulted in reduced levels of extracellular enzymes and EPS in Xanthomonas campestris pv. campestris (Xcc). Additionally, a possible connection between rpfA and iron metabolism was suggested35. Simultaneously, no change or decrease in expression was recorded for two other genes, rpfB and rpfF, which are essential for the synthesis of a small diffusible signal molecule that is involved in the regulation of virulence factor synthesis in response to physiological or environmental changes36.
Xf has a distinct T3E repertoire comprising multiple rare effectors and several putative new effectors4. Of genes annotated T3E or coding for avirulence proteins, only some were upregulated in planta. Among the five genes annotated xopP, only one xopP_1 had higher expression in planta than in pure culture, while four other copies showed no change in expression. XopP was found to be a crucial T3E in X. campestris37, conserved among different pathovars of this species and other Xanthomonas species38. Its function was recently described as blocking peptidoglycan- and chitin-triggered immunity in rice by inhibiting the U-box ubiquitin ligase OsPUB4439. All five copies are located in one region of the genome next to each other. Interestingly, several putative new effectors found by Vandroemme et al.4 in the Xf genome were differentially expressed in strawberry plants. Effectors of foreign origin (such as XopC1) with the highest homology to proteins in Ralstonia (e.g., XopAD_1, which shows the highest similarity to an unidentified protein in Mesorhizobium) were upregulated in planta, similar to the majority of genes coding putative XopD and showing distant homology to a T3E of Pseudomonas. These results suggest that the foreign origin genes coding for T3E can be functional in Xf during the pathogenesis process.
The role of T2SS, used for secretion of degradative enzymes, has not yet been studied in X. fragariae. In other Xanthomonas species, such as X. campestris pv. vesicatoria40 and X. axonopodis pv. citri41, the xps cluster constituting the T2SS has been identified. It is likely that this secretion system also plays a role in Xf, as several genes responsible for the synthesis of degradative enzymes, such as cellulases, proteases, polygalacturonases, lyases, amylases and lipases, were upregulated in planta.
During infection of plants, bacteria are exposed to a variety of antimicrobial compounds produced by the host. One of the mechanisms underlying plant resistance to diseases can be related to increased concentrations of toxic pathogen compounds, as is found in apple cultivars resistant to Erwinia amylovora13. Multidrug efflux pumps and permeases recognize and efficiently expel a wide range of structurally diverse compounds from the bacterial cell and play a very important role in the success of pathogens42. This observation could suggest that the compounds produced by strawberry leaf tissue are not toxic for Xf although they have antimicrobial properties43, and therefore, the expression of genes coding for proteins involved in the detoxification of bacterial cells is not increased in planta.
Plasmids are the extrachromosomal elements widely spread in plant pathogenic bacteria. The range of plasmid-coded features, including virulence, toxin and hormone production and resistance to bactericides, has been reported in many phytopathogenic bacteria. In the case of Xanthomonas, plasmid biology is still not well understood44; however, there are some reports on organisms such as X. campestris pv. malvacearum showing that in cotton, a number of plasmid-borne avr genes together encode the ability to cause watersoaking in the host plant45. The role of plasmids in X. fragariae has not been studied at all; however, the presence of a gene coding for T3E on the plasmid and the change in expression of almost half of the plasmid genes in planta suggest its potential importance in the interaction with plants.
Conclusions
The presented RNA-seq analysis describes the transcriptional response of X. fragariae in strawberry cv. Elsanta leaves 15 days after inoculation. The majority of genes known to be important for pathogenicity had increased expression in planta. Among genes with the greatest fold change in expression between experimental conditions or with the highest transcript abundance, there are many genes not assigned functions that have never been tested for their role in pathogenicity. Thus, although their role is unknown, their function in interacting with the host plant may be important. This study provides the first transcriptional profile by RNA-seq of X. fragariae during infection of a host plant.
Methods
Sample collection and RNA isolation
Strain X. fragariae IPO (Instituut voor Plantenziektenkundig Onderzoek) 3485, isolated from an infected strawberry plant in 2011 in the Netherlands, was used in this study. For RNA isolation, young leaves of strawberry cv. Elsanta grown in a quarantine greenhouse (25–30 °C; relative humidity level of 70–90%, natural light conditions) were inoculated with a water bacterial suspension of 106 cfu mL−1 grown for 20 h in liquid Wilbrink’s medium46 by high-pressure spraying of the abaxial and adaxial surfaces of leaves as previously described47.
Fifteen days after inoculation, samples were processed, and total RNA was isolated using a Total RNA Purification Kit (Norgen Biotek) after the Genomic Mini AX SOIL Spin kit (A&A Biotechnology, Gdynia, Poland) as described by Kałużna et al.47. At this time point, RNA was isolated separately from three samples of 1.5 to 2.5 g of water-soaked parts of strawberry leaves. Before isolation, the surface of the leaves was sterilized6. Additionally, RNA was isolated from a pure culture of X. fragariae IPO 3485 grown at 26 °C with 150 rpm shaking for 20 h in liquid Wilbrink’s medium. DNA was removed from samples by DNase treatment (Deoxyribonuclease I, ThermoScientific, Lithuania). The efficiency of DNA removal was tested by PCR with the primers 245A and 245B48, which were designed for the detection and identification of X. fragariae. Determination of the quality and concentration of obtained DNA-free RNA was tested on an Agilent 2100 Bioanalyzer using the Agilent RNA 6000 Nano Kit according to the manufacturer’s instructions. Two samples of pure bacterial culture and three samples of RNA isolated from infected strawberry leaves of the best quality (RIN) were subjected to rRNA depletion using a Ribo-Zero Magnetic Kit (Gram-Negative Bacteria, Illumina); they constituted biological replicates for each experimental combination.
Library preparation and sequencing
The rRNA-depleted sample concentration was measured using a 2100 Bioanalyzer (Agilent) and an RNA 6000 Pico Kit (Agilent, 5067-1513). The maximum allowable volume (6 µl) of mRNA was used for library construction using the NEBNext Ultra Directional RNA Library Preparation Kit for Illumina (New England Biolabs, E7420S). The libraries were sequenced on a MiSeq (Illumina) using the MiSeq Reagent Kit v2 (500 cycles) (Illumina, MS-102-2003) in the PE250 read mode. The resulting reads were additionally trimmed with Cutadapt v. 1.12 at default parameters49 These sequence data have been submitted to the ArrayExpress (EMBL) databases under accession number E-MTAB-8189.
Bioinformatic analysis
Low-quality sequence ends (ambiguous base limit: 2, quality limit: 0.05) were trimmed using the CLC Genomics Workbench v. 8.1 (Qiagen) Trim Sequences tool. High-quality sequences were aligned to the X. fragariae strain IPO 3485 (= NBC 2815) genome (LT853880, LT853881)50 using the CLC RNA-seq reference mapping algorithm with settings appropriate for prokaryotic genomes (mapping to gene regions only). A quality control to check whether the overall variability of the samples reflected their grouping and the reproducibility between repetitions was performed with PCA. For DEGs analysis, expression values were normalized using the Trimmed Mean of M values (TMM)51, and DEGs were analysed using the empirical analysis of DGE tool based on Exact Test incorporated in the EdgeR Bioconductor package and implemented in CLC Genomics Workbench. A gene was considered to be differentially regulated between two conditions when the gene showed a total read number larger than 5, a > 1.5-fold absolute fold-change ratio and an FDR-adjusted p value < 0.05.
To summarize the pathway information protein sequence, fasta files were submitted to KAAS (KEGG automatic annotation server)52,53, and KEGG orthology assignments were obtained (Supplementary Table S8). The eggNOG 4.5 database53 was used to annotate genes with common denominators or functional categories (i.e., derived from the original COG categories). Enrichment of COG and KEGG terms was evaluated by a hypergeometric distribution at FDR < 0.05.
RT-qPCR validation
The transcription expression reported in the present study was validated through RT-qPCR using 18 candidate genes selected from highly and lowly expressed bacterial genes with newly designed primers (Supplementary Table S2). Herein, three biological replicates were used to evaluate the transcriptional expression of the X. fragariae strain IPO 3485 in each biological replicate. For gene amplification, total RNA was isolated, reverse transcribed and amplified with real-time PCR, as described by Kałużna et al.47. The qPCR runs were performed on a Bio-Rad CFX96 thermocycler with SsoAdvanced SYBR Green Supermix (Bio-Rad, Hercules, CA) under the conditions described47 using the comparative 2 − ΔΔCT method.
The genes with the most stable expression based on previous analysis47 were used for normalization of RT-qPCR expression analysis of tested genes: gyrB (DNA gyrase B), ffh (signal recognition particle protein), and pykA (pyruvate kinase). For assessment of the association between RNA-seq and RT-qPCR, Pearson's correlation method was used.
Ethics approval and consent to participate
Plant material was bought from the commercial nursery. No field permissions were necessary to collect the plant samples. No specimens have been deposited as vouchers. Research Institute of Horticulture possess a permission of Main Inspectorate of Plant Health and Seed Inspection, Poland for work with quarantine pathogen Xanthomonas fragariae (Decision No. WF-411d-6/14).
References
Anonymous. PM 7-65 (1) Xanthomonas fragariae. Bull. OEPP/EPPO Bull. 36, 135–144 (2006).
Kennedy, B. W. & King, T. H. Angular leaf spot of strawberry caused by Xanthomonas fragariae sp. nov.. Phytopathology 52, 873–875 (1962).
Allan-Wojtas, P. et al. Low temperature and anhydrous electron microscopy techniques to observe the infection process of the bacterial pathogen Xanthomonas fragariae on strawberry leaves. J. Microsc. 239, 249–258 (2010).
Vandroemme, J., Cottyn, B., Baeyen, S., De Vos, P. & Maes, M. Draft genome sequence of Xanthomonas fragariae reveals reductive evolution and distinct virulence-related gene content. BMC Genom. 14, 829 (2013).
Cantu, D., Vicente, A. R., Labavitch, J. M., Bennett, A. B. & Powell, A. L. T. Strangers in the matrix: plant cell walls and pathogen susceptibility. Trends Plant Sci. 13, 610–617 (2008).
Kastelein, P. et al. Development of Xanthomonas fragariae populations and disease progression in strawberry plants after spray-inoculation of leaves. Plant Pathol. 63, 255–263 (2014).
Kastelein, P., Evenhuis, A., van der Zouwen, P. S., Krijger, M. & van der Wolf, J. M. Spread of Xanthomonas fragariae in strawberry fields by machinery. EPPO Bull. 48, 569–577 (2018).
Zhao, Y., Blumer, S. E. & Sundin, G. W. Identification of Erwinia amylovora genes induced during infection of immature pear tissue. J. Bacteriol. 187, 8088–8103 (2005).
Pandey, S. S., Patnana, P. K., Lomada, S. K., Tomar, A. & Chatterjee, S. Co-regulation of iron metabolism and virulence associated functions by iron and XibR, a novel iron binding transcription factor, in the plant pathogen Xanthomonas. PLoS Pathog. 12(11), e1006019 (2016).
Johnson, C. H. & Gonzalez, F. J. Challenges and opportunities of metabolomics. J. Cell. Physiol. 227, 2975–2981 (2012).
Bumgarner, R. Overview of dna microarrays: types, applications, and their future. Curr. Protoc. Mol. Biol. 101, 22–1 (2013).
Brankatschk, K., Kamber, T., Pothier, J. F., Duffy, B. & Smits, T. H. M. Transcriptional profile of Salmonella enterica subsp. enterica serovar Weltevreden during alfalfa sprout colonization. Microb. Biotechnol. 7, 528–544 (2014).
Puławska, J., Kałuzna, M., Warabieda, W. & Mikiciński, A. Comparative transcriptome analysis of a lowly virulent strain of Erwinia amylovora in shoots of two apple cultivars—susceptible and resistant to fire blight. BMC Genom. 18, 868 (2017).
Lu, H. et al. Acquisition and evolution of plant pathogenesis-associated gene clusters and candidate determinants of tissue-specificity in Xanthomonas. PLoS ONE 3(11), e3828 (2008).
Cianciotto, N. P. & White, R. C. Expanding role of type II secretion in bacterial pathogenesis and beyond. Infect. Immun. 85, e00014-17 (2017).
Qian, W. et al. Comparative and functional genomic analyses of the pathogenicity of phytopathogen Xanthomonas campestris pv. campestris. Genome Res. 15(6), 757–767 (2005).
Li, Y. et al. Mechanism of plant-microbe interaction and its utilization in disease-resistance breeding for modern agriculture. Physiol. Mol. Plant Pathol. 83, 51–58 (2013).
Schirmer, T. & Jenal, U. Structural and mechanistic determinants of c-di-GMP signalling. Nat. Rev. Microbiol. 7, 724–735 (2009).
Maas, J. L. Compendium of Strawberry Diseases. 2nd ed. (APS Press, 1998).
Beattie, G. A. & Lindow, S. E. The secret life of foliar bacterial pathogens on leaves. Annu. Rev. Phytopathol. 33, 145–172 (1995).
Koebnik, R., Locher, K. P. & Van Gelder, P. Structure and function of bacterial outer membrane proteins: Barrels in a nutshell. Mol. Microbiol. 37(2), 239–253 (2000).
Fleites, L. A. et al. Xanthomonas albilineans OmpA1 appears to be functionally modular and both the OMC and C-like domains are necessary for leaf scald disease of sugarcane. Mol. Plant-Microbe Interact. 26(10), 1200–1210 (2013).
Gottesman, S., Wickner, S. & Maurizi, M. R. Protein quality control: triage by chaperones and proteases. Genes Dev. 11, 815–823 (1997).
Hong, H., Patel, D. R., Tamm, L. K. & Van Den Berg, B. The outer membrane protein OmpW forms an eight-stranded β-barrel with a hydrophobic channel. J. Biol. Chem. 281(11), 7568–7577 (2006).
Gil, F. et al. SoxS regulates the expression of the Salmonella enterica serovar Typhimurium ompW gene. Microbiology 155(Pt 8), 2490–2497 (2009).
Zimaro, T. et al. Insights into Xanthomonas axonopodis pv. citri biofilm through proteomics. BMC Microbiol. 13, 186 (2013).
Boyer, F., Fichant, G., Berthod, J., Vandenbrouck, Y. & Attree, I. Dissecting the bacterial type VI secretion system by a genome wide in silico analysis: What can be learned from available microbial genomic resources?. BMC Genom. 10, 104 (2009).
Silverman, J. M., Brunet, Y. R., Cascales, E. & Mougous, J. D. Structure and regulation of the type VI secretion system. Annu. Rev. Microbiol. 66, 453–472 (2012).
Kim, S. et al. Time-resolved pathogenic gene expression analysis of the plant pathogen Xanthomonas oryzae pv. oryzae. BMC Genom. 17, 345 (2016).
Guo, Y., Figueiredo, F., Jones, J. & Wang, N. HrpG and HrpX play global roles in coordinating different virulence traits of Xanthomonas axonopodis pv. citri. Mol. Plant-Microbe Interact. 6, 649–661 (2011).
Mhedbi-Hajri, N., Jacques, M. A. & Koebnik, R. Adhesion mechanisms of plant-pathogenic Xanthomonadaceae. Adv. Exp. Med. Biol. 715, 71–89 (2011).
Malamud, F. et al. The Xanthomonas axonopodis pv. citri flagellum is required for mature biofilm and canker development. Microbiology 157, 819–829 (2011).
Kamoun, S. & Kado, C. I. Phenotypic switching affecting chemotaxis, xanthan production, and virulence in Xanthomonas campestris. Appl. Environ. Microbiol. 56, 3855–3860 (1990).
Becker, A., Katzen, F., Pühler, A. & Ielpi, L. Xanthan gum biosynthesis and application: a biochemical/genetic perspective. Appl. Microbiol. Biotechnol. 50, 145–152 (1998).
Wilson, T. J. et al. The rpfA gene of Xanthomonas campestris pathovar campestris, which is involved in the regulation of pathogenicity factor production, encodes an aconitase. Mol. Microbiol. 28, 961–970 (1998).
Barber, C. E. et al. A novel regulatory system required for pathogenicity of Xanthomonas campestris is mediated by a small diffusible signal molecule. Mol. Microbiol. 24, 555–566 (1997).
Roux, B. et al. Genomics and transcriptomics of Xanthomonas campestris species challenge the concept of core type III effectome. BMC Genom. 16, 975 (2015).
Potnis, N. et al. Comparative genomics reveals diversity among xanthomonads infecting tomato and pepper. BMC Genom. 12, 146 (2011).
Ishikawa, K. et al. Bacterial effector modulation of host E3 ligase activity suppresses PAMP-triggered immunity in rice. Nat. Commun. 5, 5430 (2014).
Szczesny, R. et al. Functional characterization of the Xcs and Xps type II secretion systems from the plant pathogenic bacterium Xanthomonas campestris pv vesicatoria. New Phytol. 187, 983–1002 (2010).
Baptista, J. C. et al. Mutation in the xpsD gene of Xanthomonas axonopodis pv. citri affects cellulose degradation and virulence. Genet. Mol. Biol. 33, 146–153 (2010).
Martinez, J. L. et al. Functional role of bacterial multidrug efflux pumps in microbial natural ecosystems. FEMS Microbiol. Rev. 33, 430–449 (2009).
Puupponen-Pimiä, R. et al. Antimicrobial properties of phenolic compounds from berries. J. Appl. Microbiol. 90, 494–507 (2001).
Vivian, A., Murillo, J. & Jackson, R. W. The roles of plasmids in phytopathogenic bacteria: Mobile arsenals?. Microbiology 147(Pt 4), 763–780 (2001).
Yang, Y. Watersoaking function(s) of XcmH1005 are redundantly encoded by members of the Xanthomonas avr/pth gene family. Mol. Plant–Microbe Interact. 9(2), 105–113 (1996).
Koike, H. The aluminum-cap method for testing sugarcane varieties against leaf scald disease. Phytopathology 55, 317–319 (1965).
Kałużna, M., Kuras, A. & Puławska, J. mRNA extraction of Xanthomonas fragariae in strawberry and validation of reference genes for the RT-qPCR for study of bacterial gene expression. Mol. Biol. Rep. 46, 5723–5733 (2019).
Pooler, M. R., Ritchie, D. F. & Hartung, J. S. Genetic relationships among strains of Xanthomonas fragariae based on random amplified polymorphic DNA PCR repetitive extragenic palindromic. Appl. Environ. Microbiol. 62, 3121–3127 (1996).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnetjournal 17, 10–12 (2011).
Gétaz, M., van der Wolf, J. M., Blom, J. & Pothier, J. F. Complete genome sequences of three isolates of Xanthomonas fragariae, the bacterium responsible for angular leaf spots on strawberry plants. Genome Announc. 5(32), e00642-e717 (2017).
Robinson, M. D. & Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 11(3), R25 (2010).
Moriya, Y., Itoh, M., Okuda, S., Yoshizawa, A. C. & Kanehisa, M. KAAS: an automatic genome annotation and pathway reconstruction server. Nucleic Acids Res. 35, W182–W185 (2007).
Huerta-Cepas, J. et al. EGGNOG 4.5: a hierarchical orthology framework with improved functional annotations for eukaryotic, prokaryotic and viral sequences. Nucleic Acids Res. 44(D1), D286–D293 (2016).
Acknowledgements
This study has received funding from the European Union Seventh Programme (FP7/2007-2013) under Grant Agreement Number 613678 (DROPSA). The research work was co-financed from financial resources for science MNiSW in 2014–2018 granted for the implementation of the international co-funded Project 3195/7. PR/2014/2. This article is also based upon work from COST Action CA16107 EuroXanth, supported by COST (European Cooperation in Science and Technology).
Author information
Authors and Affiliations
Contributions
J.P. is author of the conception of the study, its design, bioinformatics analysis of RNA-seq data and their interpretation. M.K. designed and performed RNA isolation and RNA-seq validation by RT-qPCR. W.W. analysed data statistically. J.F.P. and M.G. sequenced the genome of Xf strain. The manuscript was written by J.P. and J.Mvd.W. All authors contributed to manuscript revision, read and approved the submitted version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Puławska, J., Kałużna, M., Warabieda, W. et al. Transcriptome analysis of Xanthomonas fragariae in strawberry leaves. Sci Rep 10, 20582 (2020). https://doi.org/10.1038/s41598-020-77612-y
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-77612-y
This article is cited by
-
Dynamic character displacement among a pair of bacterial phyllosphere commensals in situ
Nature Communications (2022)
-
Comparative transcriptional analyzes of Xanthomonas citri subsp. citri reveal mechanisms of adaptation and bacterial virulence in the early stage of citrus canker disease
European Journal of Plant Pathology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.