Abstract
Reporter gene imaging allows for non-invasive monitoring of molecular processes in living cells, providing insights on the mechanisms underlying pathology and therapy. A lysine-rich protein (LRP) chemical exchange saturation transfer (CEST) MRI reporter gene has previously been developed and used to image tumor cells, cardiac viral gene transfer, and oncolytic virotherapy. However, the highly repetitive nature of the LRP reporter gene sequence leads to DNA recombination events and the expression of a range of truncated LRP protein fragments, thereby greatly limiting the CEST sensitivity. Here we report the use of a redesigned LRP reporter (rdLRP), aimed to provide excellent stability and CEST sensitivity. The rdLRP contains no DNA repeats or GC rich regions and 30% less positively charged amino-acids. RT-PCR of cell lysates transfected with rdLRP demonstrated a stable reporter gene with a single distinct band corresponding to full-length DNA. A distinct increase in CEST-MRI contrast was obtained in cell lysates of rdLRP transfected cells and in in vivo LRP expressing mouse brain tumors (\(p=0.0275\), n = 10).
Similar content being viewed by others
Introduction
Magnetic resonance (MR) reporter genes can non-invasively monitor transgene expression in vivo at high resolution with unlimited tissue penetration1. These genes can be utilized for the evaluation of cell and gene-based therapy, as investigated for cancer, neural degeneration, and myocardial infarction2. Specific applications include tracking of dividing/differentiating stem cells3, imaging of dendritic cells4, monitoring of viral therapeutics5, and detection of micro-environment changes6.
The adaptability of MRI and the different contrast mechanism that can be used, fostered creativity among scientists that allowed the development of numerous genetically encoded reporters7,8. In a pioneering work in the field, Louie et al., used beta-galactosidase to catalyze an MRI specific probe9. This study set the stage for a variety of genetically encoded reporters based on T\(_{1}\) relaxation. Bartelle, Turnbull and their colleagues have taken a creative approach to express a multi-component reporter specifically in endothelial cells10. Another strategy relied on overexpression of the organic anion transporting protein (Oatp1a1). These transporters facilitate the uptake of Gd(\(^{3+}\))-DTPA11,12. A different approach was to develop genetically encoded reporters based on T\(_{2}\) relaxation. By using iron transporters13,14,15 or iron storage proteins16,17,18, scientists sought to modulate the intracellular T\(_{2}\) relaxivity. An alternative approach was to use hyperpolarized xenon as a substrate for enhancing the MRI contrast of a genetically encoded reporter19. Nevertheless, all these innovative approaches relied on administration of a substrate or a secondary compound that is not accessible to all tissues and requires repetitive injections for longitudinal imaging.
A different category of MRI reporter genes, which do not require any exogenous material co-administration, includes gas-filled protein nanostructures that affect the magnetic susceptibility20, the human water channel aquaporin 1 that affects water diffusivity21, the urea transporter UT-B that affects the transmembrane water exchange22, and chemical exchange saturation transfer (CEST) based reporters23. CEST is an alternative and particularly useful contrast mechanism for MR reporters23,24,25,26,27,28. In a typical CEST-MRI experiment, compounds containing exchangeable protons are selectively saturated using selective radio-frequency (RF) irradiation and undergo chemical exchange with the bulk water protons. A signal associated with the compound concentration and its labile proton exchange rate can then be detected using the water signal29,30,31. CEST MRI reporter genes offer a few attractive advantages: (1) The artificially engineered CEST proteins are non-metallic and bio-compatible. (2) The imaging procedure does not require the injection of a concomitant labeling agent. (3) The reporter contrast can be switched on/off by the use of a selective MR RF saturation pulse, allowing the acquisition of additional and uncontaminated anatomical/functional MRI information at the same imaging session. Of particular promise was the development of the lysine-rich protein (LRP) reporter gene23, with demonstrated applicability for imaging cells in tumors32, oncolytic virus delivery and spread5, and cardiac viral vector expression33. However, thus far, the gap between the promise of MR reporters and actual clinical use has not been bridged due to the low sensitivity and specificity of the detection. In particular, the LRP reporter gene in its current form is unstable, as the highly repetitive DNA sequence encoding the reporter leads to DNA recombination events and hence the expression of a range of truncated protein fragments with fewer lysine amide protons, resulting in decreased and limited CEST sensitivity5,32.
Here, we report the redesign of an improved, stable, and robust LRP-based CEST-MRI reporter gene (rdLRP), using an RHGP amino acid motif that consisted of arginine, histidine, glycine, and proline amino acids separated by different numbers of lysines. The reporter stability and contrast enhancement ability were evaluated in cell lysates from human embryonic kidney (HEK293T) and murine glioma (GL261N4) cell lines as well as from in vivo LRP expressing brain tumors in mice.
Results
CEST-MRI of various polypeptides
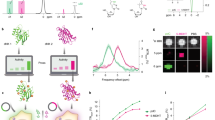
A number of different lysine polypeptides with different amino acid motifs separating lysine residues were screened to find a peptide that disrupts the repetitive lysine sequences and has optimal CEST contrast. Of these, the CEST contrast of a lysine polypeptide containing an arginine, histidine, glycine, and proline (RHGP) amino acid motif stood out as this displayed virtually the same CEST contrast as poly-L-lysine (PLL) for the amide exchangeable protons at 3.6 ppm chemical shift (Fig. 1, MTR\(_{asym}\) of 6.0% and 6.3% for RHGP and PLL, respectively, while the spectral noise at a blank PBS region was 0.5%). In addition, only for the RHGP was a strong CEST contrast observed at the chemical shift of \(\sim \) 2 ppm attributed to the guanidyl amines of the arginine residues (Fig. 1b). Based on the contrast advantages described above, the rdLRP was engineered using RHGP units with different lysine spacers and a total of 100 amino acids (Fig. 2a). It shall be noted, that If we would produce a protein with only multiple copies of the RHGP polypeptide, the stability of the DNA/RNA and the protein could be reduced. Therefore, we also incorporated scrambled partial motifs from all other examined peptides, as they all had a considerable contribution to the 3.6 ppm contrast.
Screening various polypeptides in search of an optimal LRP-based reporter gene backbone. (a) Z-spectra for a variety of polypeptides with different amino acid motifs. (b) The corresponding CEST contrast observed as a function of saturation frequency offset (MTR\(_{asym}\)). (c) Polypeptide sequences and their respective CEST contrast at the amide exchangeable proton chemical shift of 3.6 ppm. The amide proton CEST contrast for the RHGP polypeptide (containing the arginine, histidine, glycine and proline amino acid motif) is similar in magnitude to that of PLL. A large CEST contrast is also observed at 2 ppm for the guanidyl amine exchangeable proton of the RHGP polypeptide. A: Alanine, PLL: poly-L-lysine.
RNA characterization of the rdLRP reporter. (a) Amino acid sequence of the rdLRP consisting of RHGP units with different lysine spacers with a total of 100 amino acid residues. (b,c) RT-PCR of total mRNA from HEK293T cell lysates transfected with rdLRP, evaluating the BGH reverse tag (b) and the v5 epitope tag (c) sites. A single well-defined band is observed from cell lysates transfected with rdLRP, but not from control cell lysates. The base-pair lengths are consistent with expression of full-length rdLRP mRNA (b, 557 bp; c, 491 bp). Raw gel images are shown in Supplementary Information Fig. S2.
Stability analysis of the rdLRP reporter
The rdLRP contains no DNA repeats or GC rich regions resulting in a significantly more stable protein with 30% less positively charged amino acids compared to the original LRP (see Supplementary Information Fig. S1). The improved stability is evident in RT-PCR of total mRNA from cell lysates transfected with rdLRP, where only a single well-defined DNA band is observed at the base pair length expected for full length rdLRP (Fig. 2b,c, Supplementary Information Fig. S2). In contrast, a ladder profile of bands of different base-pair lengths was observed previously in RT-PCR of cell lysates transfected with the original LRP reporter gene (see Gilad et al.23 and Farrar et al.5).
In vitro CEST-MRI of rdLRP transfected cell lysates
A distinct increase in the CEST magnetization transfer ratio asymmetry (MTR\(_{asym}\)) was observed for both human embryonic kidney (HEK293T) cells and murine glioma (GL261N4)34 cells transfected with the rdLRP relative to control, non transfected cells (Fig. 3). The two main amplitude peaks in the MTR\(_{asym}\) of the cell lysates were at the guanidyl amine (\(\sim \) 2 ppm) and amide (\(\sim \) 3.6 ppm) exchangeable proton frequencies, in agreement with the RHGP peptide CEST analysis (Fig. 1).
In vitro CEST-MRI contrast evaluation of the rdLRP reporter. (a,b) Z-spectra of control (blue trace) and rdLRP (red trace) in transfected HEK293T (a) and GL261N4 (b) cell lysates acquired at 4.7T. (c,d) The corresponding MTR\(_{asym}\) spectra. Increased CEST contrast is observed at \(\sim \) 2 ppm corresponding to the guanidyl amine exchangeable protons of arginine amino acid residues and at \(\sim \) 3.6 ppm corresponding to the amide exchangeable protons of the peptide bond generated by coupling two amino acids.
In vivo imaging of rdLRP expressing mice brain tumors
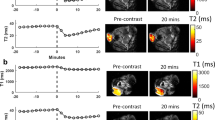
To verify that the contrast enhancement ability of the rdLRP reporter is retained for in vivo settings and to demonstrate the reporter effectiveness for a molecular-genetic imaging scenario, a preclinical tumor monitoring experiment was conducted. The rdLRP was expressed in a murine glioma GL261N4 cell line and transplanted into mouse brains. In vivo CEST-MR images were acquired at 7T and the amide-proton based contrast was analyzed (Fig. 4). As the molecular environment of the living brain/tumor has a more diverse multi-compound content compared to in vitro cells, the extraction of the CEST amide proton amplitude was performed using a Lorentzian fitting model (see the “Methods” section), aimed to isolate the signal of interest from the confounding contribution stemming from lipid/semi-solid macro-molecules and aliphatic protons (rNOE). A significant increase in the tumor amide-CEST contrast was obtained for LRP expressing tumors (\(5.88 \pm 1.44\)%, mean ± STD) compared to control (\(4.34 \pm 1.28\)%, mean ± STD), as shown in Fig. 4h (two-tailed t-test, \(p=0.0275\), n = 10 imaging data points per group, t = 2.399, df = 18, difference between means: \(1.542 \pm 0.643\)% (mean ± SEM)). These findings demonstrate that the rdLRP can be expressed in vivo in a cell-selective manner and can be used for molecular MR studies. It shall be noted, that due to evident edema (Fig. 4c–e), the protein content in the tumor region was diluted, resulting in a general decrease in the CEST signal compared to the contralateral region (Fig. 4a,b, Supplementary Information Fig. S3), as previously reported in other CEST tumor studies35,36,37. The edema has similarly affected the control and the rdLRP group mice, as reflected by the similar increase in T\(_{2}\) value for both groups (from 48.18 ± 0.75 ms to 72.86 ± 12.05 ms for the contralateral and tumor ROIs in the control mice group, respectively, and from 48.33 ± 0.79 ms to 73.08 ± 7.92 ms in the rdLRP mice group contralateral and tumor ROIs, respectively). The difference between the control and rdLRP mice groups T\(_{2}\) tumor values was not statistically significant (two-tailed t-test, \(p = 0.961\), t = 0.0491, df = 18, n = 10 imaging data points per group). Hence, although sharing similar conditions for the endogenous amide content and edema extent, the rdLRP mice group has presented a distinct amide CEST contrast increase, resulting from the presence of the rdLRP. The variability in the amide CEST signal over a single slice and between different subjects of the same group may come from B\(_{1}\) inhomogeneity, reference signal drift over time, and in-vivo variability of the T\(_{1}\) and T\(_{2}\) of water38. Nevertheless, the amide CEST signal increase effect was sufficiently strong to overcome these sources of uncertainty, as manifested in the statistically significant effect shown in Fig. 4h.
In vivo imaging of the rdLRP. (a,b) Amide proton CEST amplitude maps of representative mice implanted with either GL261N4 glioma cells (a) or with rdLRP transfected GL261N4 cells (b). (c) T\(_{2}\) relaxation time analysis for the two imaged groups, demonstrating the evident edema at the tumor region. Notably, no statistically significant differences were calculated at the tumor region between the rdLRP and control group mice (two-tailed t-test, \(p=0.961\), t = 0.0491, df = 18, n = 10 imaging data points per group). (d,e) T\(_{2}\)-weighted images for the mice imaged at (a,b), respectively. (f,g) The amide proton CEST contrast, obtained using a frequency selective saturation pulse (a,b) is overlaid atop the tumor region in the T\(_{2}\)-weighted images presented in (d) and (e), respectively. (h) Quantitative group comparison for the amide proton CEST amplitude. Statistically significant CEST signal increase was obtained in the rdLRP group (two-tailed t-test, \(p=0.0275\), n = 10 imaging data points per group, t = 2.399, df = 18, difference between means: \(1.542 \pm 0.643\)% (mean ± SEM)). The box plots represent the median, interquartile range, Tukey style whiskers, outliers (yellow) and individual data points.
Discussion
Stability and reproducibility are two essential factors required for the successful application of a reporter gene39. Furthermore, to allow the correct quantification of the biological/therapeutic process of interest (e.g., the extent of viral replication and spread during oncolytic virotherapy5), it is essential to reliably know the exact composition of the contrast-generating substance in the reporter (e.g., the number of amide bonds of the backbone, as affected by the environment induced by the presence of lysine amino acids) and avoid contaminating effects, such as random protein truncation. However, engineering highly predictable and reliable reporters is technically challenging, as demonstrated in the gene expression imperfections of previously developed CEST-MRI reporter genes5,23. Here, the LRP reporter was redesigned for overcoming its previous limitations and enhance its performance. The results have demonstrated a marked improvement in stability, as evident in RT-PCR (Fig. 2), where only full-length cDNA was observed.
The rationale of this study was to increase the LRP cellular expression and make it a more stable reporter. This will allow to use a smaller protein, half the size of the original LRP, and still ensure sufficient contrast. This is a major consideration when designing a viral gene delivery system with limited DNA payload. To that end we modified the LRP at the translational, transcriptional and protein level. For transcriptional efficacy we minimized the GC content and eliminated CpG dinucleotides content, cryptic splicing sites, negative CpG islands and TATA boxes. For improving translation efficiency, we optimized codon usage for mammalian cells. This approach was proven very efficient in the expression of another MRI reporter gene, the human PRM1, where codon optimization turned the protein from practically unexpressed to highly expressed in E. Coli24. The new rdLRP sequence (Supplementary Information Fig. S1) also encodes to less than 60% lysines, unlike previous generation that encodes close to 100% lysines and was double in length. In other words, the rdLRP uses only 35% of the number of Lysines residues per protein. Assuming a strong gene promoter is used, which is often the case, the original sequence may deplete the cellular lysine reservoirs and slow down the translation by overusing the lysine-tRNALys. Therefore, reducing the lysine content can significantly reduce the burden on the cellular protein production machinery. In addition, we eliminated mRNA secondary structures, RNA instability motifs, removed ribosomal binding sites and minimized protein refolding. We incorporated protein motifs that were suggested to increase peptide resistance to degradation and improve peptides stability40. Together, these manipulations, lead to a more stable LRP that is smaller but can be overexpressed in higher concentrations in cells and consequently increase the reporter detectability.
Human protamine-1 (hPRM1)24,41 is another CEST reporter gene described recently that has demonstrated efficient expression and stability in cells24 (Supplementary Information Table S1). Findings and conclusions from this reporter were integrated into the design of the rdLRP. Briefly, hPRM1 is a relatively short protein, which encodes to 51 amino acids. Moreover, it has high content (47%) of arginine residues, thus, cells have limited capacity of synthesizing high quantities of it. Taken together these two factors are limiting the production of the hPRM1 protein and preventing the cloning of multiple copies of the hPRM1 in tandem, resulting in limited contrast. Those limitations were one of the motivations of exploring the rdLRP as the next generation of CEST reporter gene. Importantly, the CEST-MRI contrast enhancement ability was fully retained compared to the original LRP reporter gene. The in vivo CEST contrast improvement calculated in this work (\(1.542 \pm 0.643\)%, mean ± SEM) is similar to that reported in previous LRP studies, where an in vivo increased CEST contrast of 1.1%5 or less32 was observed. As mentioned earlier, the smaller rdLRP reporter gene contains only 35% of the total lysine content theoretically available in the original LRP reporter23. However, this change in the reporter gene composition is compensated by its superior stability, allowing to de facto take advantage of its full contrast enhancement potential, without any unexpected decreases in sensitivity due to random DNA/protein truncation. With a stable reporter gene on hand, the size of the reporter gene could be extended for a further gain in CEST sensitivity.
The rdLRP imaging studies were performed on a variety of magnetic field strengths (4.7, 7, and 11.7T). Generally, as the B\(_{0}\) field increases, the signal from fast exchanging protons (e.g., from the guanidyl amine protons) is better visualized, the T\(_{1}\) of water is increased (allowing a longer chemical exchange and hence a stronger CEST signal), and the different spectral components are better resolved38. Although a direct quantitative comparison between the contrast improvement at different fields and conditions is challenging due to the variety of contributing factors (endogenous amide/amine signal in-vivo, the effect of saturation pulse power, field-dependent water T\(_{1}\) and T\(_{2}\), etc.), in all cases, a detectable rdLRP CEST contrast enhancement effect was obtained, demonstrating the potential and usefulness of the suggested reporter.
Another beneficial contrast feature of the rdLRP was demonstrated in its CEST MTR\(_{asym}\) (Fig. 3), where an additional and strong CEST contrast peak was observed at the chemical shift of 2 ppm, attributed to the guanidyl amines of the arginine residue. This may provide a further boost in the CEST sensitivity as well as provide a means for performing guanidyl amine/amide ratiometric CEST imaging for obtaining quantitative pH measurements independent of the reporter concentration (as performed for iodine based CEST contrast agents42,43). In addition, the very rapid exchange rate (\(\ge 1\) kHz) of the guanidyl amine exchangeable protons may provide a means for distinguishing the CEST reporter from endogenous proteins with slow proton chemical exchange by exploiting exchange rate specific CEST methods such as Frequency Labeled EXchange transfer (FLEX)44. However, the resonance frequency of guanidyl amine protons is close to water. Therefore, to avoid direct water saturation, the saturation pulse power must be limited, preventing sufficient amplification of the fast exchanging amine proton signal. Given these two challenging amine-related considerations (a very fast exchange rate together with a resonance frequency close to water), we have chosen to focus our in-vivo investigations on the amide-based signal. Nevertheless, a potential future solution, allowing to exploit the contrast enhancement stemming from both amide and guanidyl proton pools might come from implementing a multi-saturation power acquisition scheme, as performed in CEST MR-fingerprinting45,46,47, with the potential addition of CEST neural networks for separating out the contribution of each contrast-generating compound37.
In summary, a second generation of the LRP CEST reporter gene was engineered, characterized, and thoroughly evaluated using RT-PCR, cell lysates from two cell lines, and in vivo LRP-expressing tumors in mice. Significant improvement in stability and contrast enhancement ability were demonstrated. Future work is underway to optimize the protein sequence even more with emphasis on reporters that generate CEST contrast at 5 ppm where the endogenous CEST background is lower.
Conclusion
The redesigned LRP reporter demonstrates excellent stability with distinct CEST contrast observed for both amide and guanidyl amine exchangeable protons of the redesigned reporter. The redesigned reporter gene was detectable in vivo in a mouse tumor model engineered to express the rdLRP.
Methods
CEST-MRI of various polypeptides
Five synthetic peptides (Genscript, Piscataway, NJ) were dissolved in PBS (Fig. 1) and their CEST Z-spectra were obtained on a 11.7T vertical-bore MRI scanner (Bruker Biospin, Ettlingen, Germany) with imaging parameters as described previously23,32. Briefly, 5 mg/ml peptides were placed in 5 mm NMR tubes and the collection placed within the scanner at 37 \(^{\circ }\)C. A modified rapid acquisition with relaxation enhancement (RARE) sequence was implemented, with repetition time/echo time (TR/TE) = 5000/20 ms, RARE factor = 16, slice thickness = 1mm, field of view (FOV) = \(1.7 \times 1.7\) cm, matrix size = \(128 \times 64\), resolution = \(0.17 \times 0.34\) mm, and a saturation pulse (amplitude = 3.6 \(\upmu \)T, duration = 3 s), repeated over a frequency offset range of − 5 ppm to 5 ppm with 0.1 ppm steps. The water saturation shift reference (WASSR) method48 was used for B\(_{0}\) corrections, using a similar protocol with TR = 15 s, saturation pulse amplitude, duration, and range of 0.5 \(\upmu \)T, 0.5 s, and − 1 ppm to 1 ppm with 0.1 ppm steps, respectively.
Redesigned LRP (rdLRP) reporter gene synthesis
The rdLRP was designed using the peptides in Fig. 1 as a starting point for the assembly of the gene with emphasis on amino acid motif that consisted of arginine, histidine, glycine, and proline amino acids separated by different numbers of lysines. Next, the gene was optimized by considering transcriptional efficacy (i.e., GC content, CpG dinucleotides content, cryptic splicing sites, negative CpG islands and TATA boxes) as well as translation efficiency (including: codon usage bias, GC content, mRNA secondary structure, RNA instability motif, stable free energy of mRNA and ribosomal binding sites) and protein refolding (e.g., codon usage bias, interaction of codon and anti-codon and RNA secondary structures). For that end we use dedicated algorithms from GenScript (Piscataway, NJ). A full length DNA sequence of the gene was custom synthesized by GenScript (Piscataway, NJ) and was further subcloned into expression vector.
Stability characterization of the rdLRP reporter
The Taq polymerase-amplified PCR product of the rdLRP was inserted into the TOPO Cloning site of a pcDNA3.1/V5-His-TOPO vector (Thermo Fisher Scientific, MA). The vector was transfected to \(6 \times 10^5\) cells of HEK293T cells using Opti-MEM (Thermo Fisher Scientific, MA) and a Lipofectamine reagent (Thermo Fisher Scientific, MA) according to the vendor’s protocol. RNA was extracted from the transfected cell pellets using TRIzol reagent (Thermo Fisher Scientific, MA). Purified RNA template was reverse-transcribed to cDNA by using iScript cDNA Synthesis Kit (Bio-Rad, CA) and polymerase chain reaction (PCR) was performed on T-100 Thermal Cycler (Bio-Rad, CA). Two kinds of combinations of primers (for T7 priming site/BGH Reverse priming site, and for T7 priming site/V5 epitope) were used. The sizes of the PCR products were checked by electrophoresis on 1% agarose gel.
In vitro CEST-MRI of cell lysates
The rdLRP plasmid, under control of the cytomegalovirus (CMV) promoter, was transfected into HEK293T and GL261N4 cells using lipofectamine. After 24 h incubation, the cells were lysed in pH 7.5 RIPA buffer and underwent 3 freeze/thaw/ sonication cycles. Protein content of the supernatant was measured and protein concentrations of control and LRP transfected cell lysates were adjusted to be equivalent (typically 10 \(\upmu \)g/\(\upmu \)l protein). Control and rdLRP cell lysate samples were loaded into 5 mm od sample tubes and placed into 50 ml Falcon tubes containing water. CEST Z-spectra were acquired on a 4.7T MRI scanner (Bruker Biospin, Ettlingen, Germany) using a CEST-EPI pulse sequence, with flip-angle = 90\(^{\circ }\) and TE/TR = 20/8000 ms. The saturation pulse power, duration, and frequency offsets were 3.6 \(\upmu \)T, 5000 ms, and 7 to − 7 ppm with 0.25 ppm increments, respectively.
Animal model
All experimental protocols were approved by the Institutional Animal Care and Use Committees (IACUC) of the Brigham and Women’s Hospital. All animal experiments were carried out in accordance with approved IACUC guidelines and regulations. Six-to-8-week-old female C57/BL6 mice were purchased from Envigo (n = 10). 100,000 cells of GL261N4/rdLRP-GL261N4 were implanted stereotactically with a Kopf stereotactic frame (David Kopf Instruments) in the right frontal lobe of the mice (ventral 3.0 mm, rostral 0.5 mm, and right lateral 2.0 mm from bregma).
In vivo CEST MRI
The rdLRP was expressed in a murine glioma cell line and transplanted into mice brains as described above (5 mice transplanted with rdLRP-GL261N4 and 5 mice implanted with GL261N4). Imaging was performed at 11 and 18 days post implantation using a 7T preclinical MRI (Bruker Biospin, Ettlingen, Germany). The mice were anesthetized using 0.5 to 2% Isoflurane during the imaging on a dedicated heated cradle, and the respiration rate was continuously monitored using a mechanical sensor. T\(_{2}\)-weighted images were acquired using a multi-slice multi-echo (MSME) protocol, with TR/TE = 60/2000 ms. T\(_{2}\) maps were acquired using the multi-echo spin-echo protocol, TR = 2000 ms, 25 TE values between 8 and 200 ms. Z-spectra were acquired using the same CEST-EPI sequence used for the cell lysate in vitro study, with a saturation pulse power of 0.7 \(\upmu \)T. Static magnetic field B\(_{0}\) maps were acquired using the WASSR method48 , employing a saturation pulse power of 0.3 \(\upmu \)T, TR/TE = 8000/20 ms, flip-angle = 90 \(^{\circ }\), saturation duration = 3000 ms, and a saturation pulse frequency varying between − 1 to 1 ppm in 0.1 ppm increments. The field of view (19 mm \(\times \) 19 mm \(\times \) 1 mm) and image resolution (297 \(\times \) 297 \(\times \) 1000 \(\upmu \)m\(^{3}\)) were identical for all scans.
Statistical analysis
Group analysis was carried out using a two-tailed student t-test, calculated using the open source SciPy scientific computing library for Python49. Differences were considered significant at \(p<0.05\). Box plots (Fig. 4c,h) represent the median, interquartile range, Tukey style whiskers, outliers (yellow circles), and individual data points.
Image analysis
Data analysis was performed using MATLAB R2018a and Python 3.6 custom written scripts, based on previously published routines, as described below. T\(_{2}\) maps reconstruction was performed using exponential fitting. CEST images were corrected for B\(_{0}\) inhomogeneity using the WASSR method48,50, followed by cubic spline smoothing51. The magnetization transfer ratio asymmetry curve was calculated using: MTR\(_{asym}\) = (S\(^{-\Delta \omega } - \) S\(^{+\Delta \omega }\))/S\(_0\), where S\(^{\pm \Delta \omega }\) is the signal measured with saturation offset of \(\pm \Delta \omega \) around the water frequency and S\(_0\) is the unsaturated signal. Semi-quantitative mapping of the CEST molecular compound amplitudes was performed using a 5-pool (water, semi-solid macro-molecules, amide, rNOE, and guanidyl amine) Lorentzian fitting model, with the starting point and boundaries described in52. For amide proton amplitude image visualization, an image down-sampling expedited adaptive least-squares (IDEAL) fitting approach was implemented, as described in53. Mouse tumor regions of interest (ROIs) were manually delineated based on the T\(_{2}\) weighted images. Finally, the amide Lorentzian peak amplitude pixel values were overlaid atop the T\(_{2}\) weighted images at the tumor ROIs (Fig. 4).
Data availability
The datasets generated and analysed during the current study are available from the corresponding author upon request.
Code availability
Source code is available from the corresponding author upon request.
References
Bar-Shir, A., Liu, G., Greenberg, M. M., Bulte, J. W. & Gilad, A. A. Synthesis of a probe for monitoring HSV1-tk reporter gene expression using chemical exchange saturation transfer MRI. Nat. Protoc. 8, 2380 (2013).
Vandsburger, M. H., Radoul, M., Cohen, B. & Neeman, M. MRI reporter genes: applications for imaging of cell survival, proliferation, migration and differentiation. NMR Biomed. 26, 872–884 (2013).
Campan, M. et al. Ferritin as a reporter gene for in vivo tracking of stem cells by 1.5-T cardiac MRI in a rat model of myocardial infarction. Am. J. Physiol. 300, H2238–H2250 (2011).
Kim, H. S. et al. In vivo tracking of dendritic cell using MRI reporter gene, ferritin. PLoS ONE 10, e0125291 (2015).
Farrar, C. T. et al. Establishing the lysine-rich protein CEST reporter gene as a CEST MR imaging detector for oncolytic virotherapy. Radiology 275, 746–754 (2015).
Gilad, A. A., Winnard Jr, P. T., van Zijl, P. C. & Bulte, J. W. Developing MR reporter genes: promises and pitfalls. NMR Biomed. 20, 275–290 (2007).
Airan, R. D., Li, N., Gilad, A. A. & Pelled, G. Genetic tools to manipulate MRI contrast. NMR Biomed. 26, 803–809 (2013).
Gilad, A. A. & Shapiro, M. G. Molecular imaging in synthetic biology, and synthetic biology in molecular imaging. Mol. Imaging Biol. 19, 373–378 (2017).
Louie, A. Y. et al. In vivo visualization of gene expression using magnetic resonance imaging. Nat. Biotechnol. 18, 321–325 (2000).
Bartelle, B. B. et al. Novel genetic approach for in vivo vascular imaging in mice. Circ. Res. 110, 938–947 (2012).
Patrick, P. S. et al. Dual-modality gene reporter for in vivo imaging. Proc. Natl. Acad. Sci. USA 111, 415–420 (2014).
Nyström, N. N. et al. Longitudinal visualization of viable cancer cell intratumoral distribution in mouse models using Oatp1a1-enhanced magnetic resonance imaging. Invest. Radiol. 54, 302–311 (2019).
Weissleder, R. et al. In vivo magnetic resonance imaging of transgene expression. Nat. Med. 6, 351–354 (2000).
Deans, A. E. et al. Cellular MRI contrast via coexpression of transferrin receptor and ferritin. Magn. Reson. Med. 56, 51–59 (2006).
Koretsky, A., Lin, Y., Schorle, H. & Jaenisch, R. Genetic control of MRI contrast by expression of the transferrin receptor. In Proc. Int. Soc. Magn. Reson. Med., Vol. 4, 23 (ISMRM Berkeley, CA, 1996).
Cohen, B., Dafni, H., Meir, G. & Neeman, M. Ferritin as novel MR-reporter for molecular imaging of gene expression. In Proc. Int. Soc. Magn. Reson. Med., Vol. 11, 1707 (2004).
Genove, G., DeMarco, U., Xu, H., Goins, W. F. & Ahrens, E. T. A new transgene reporter for in vivo magnetic resonance imaging. Nat. Med. 11, 450–454 (2005).
Cohen, B. et al. MRI detection of transcriptional regulation of gene expression in transgenic mice. Nat. Med. 13, 498–503 (2007).
Shapiro, M. G. et al. Genetically encoded reporters for hyperpolarized xenon magnetic resonance imaging. Nat. Chem. 6, 629 (2014).
Lu, G. J. et al. Acoustically modulated magnetic resonance imaging of gas-filled protein nanostructures. Nat. Mater. 17, 456–463 (2018).
Mukherjee, A., Wu, D., Davis, H. C. & Shapiro, M. G. Non-invasive imaging using reporter genes altering cellular water permeability. Nat. Commun. 7, 1–9 (2016).
Schilling, F. et al. MRI measurements of reporter-mediated increases in transmembrane water exchange enable detection of a gene reporter. Nat. Biotechnol. 35, 75–80 (2017).
Gilad, A. A. et al. Artificial reporter gene providing MRI contrast based on proton exchange. Nat. Biotechnol. 25, 217–219 (2007).
Bar-Shir, A. et al. Human protamine-1 as an MRI reporter gene based on chemical exchange. ACS Chem. Biol. 9, 134–138 (2014).
Bar-Shir, A. et al. Transforming thymidine into a magnetic resonance imaging probe for monitoring gene expression. J. Am. Chem. Soc. 135, 1617–1624 (2013).
Airan, R. D. et al. MRI biosensor for protein kinase a encoded by a single synthetic gene. Magn. Reson. Med. 68, 1919–1923 (2012).
Liu, G., Bulte, J. W. & Gilad, A. A. CEST MRI reporter genes. In Magnetic Resonance Neuroimaging (eds Modo, M. & Bulte, J. W.) 271–280 (Springer, Berlin, 2011).
Bar-Shir, A., Bulte, J. W. & Gilad, A. A. Molecular engineering of nonmetallic biosensors for CEST MRI. ACS Chem. Biol. 10, 1160–1170 (2015).
Van Zijl, P. C. & Yadav, N. N. Chemical exchange saturation transfer (CEST): what is in a name and what isn’t? Magn. Reson. Med. 65, 927–948 (2011).
Zaiss, M. & Bachert, P. Chemical exchange saturation transfer (CEST) and MR Z-spectroscopy in vivo: a review of theoretical approaches and methods. Phys. Med. Biol. 58, R221 (2013).
Van Zijl, P. C., Jones, C. K., Ren, J., Malloy, C. R. & Sherry, A. D. MRI detection of glycogen in vivo by using chemical exchange saturation transfer imaging (glycoCEST). Proc. Natl. Acad. Sci. USA 104, 4359–4364 (2007).
Minn, I. et al. Tumor-specific expression and detection of a CEST reporter gene. Magn. Reson. Med. 74, 544–549 (2015).
Meier, S. et al. Non-invasive detection of adeno-associated viral gene transfer using a genetically encoded CEST-MRI reporter gene in the murine heart. Sci. Rep. 8, 1–10 (2018).
Nakashima, H. et al. Toxicity and efficacy of a novel GADD34-expressing oncolytic HSV-1 for the treatment of experimental glioblastoma. Clin. Cancer Res. 24, 2574–2584 (2018).
Mehrabian, H., Desmond, K. L., Soliman, H., Sahgal, A. & Stanisz, G. J. Differentiation between radiation necrosis and tumor progression using chemical exchange saturation transfer. Clin. Cancer Res. 23, 3667–3675 (2017).
Xu, J. et al. On the origins of chemical exchange saturation transfer (CEST) contrast in tumors at 9.4 t. NMR Biomed. 27, 406–416 (2014).
Perlman, O. et al. AI boosted molecular MRI for apoptosis detection in oncolytic virotherapy. bioRxiv (2020).
van Zijl, P. C., Lam, W. W., Xu, J., Knutsson, L. & Stanisz, G. J. Magnetization transfer contrast and chemical exchange saturation transfer MRI. features and analysis of the field-dependent saturation spectrum. Neuroimage 168, 222–241 (2018).
Chaturvedi, P., Zhao, B., Zimmerman, D. L. & Belmont, A. S. Stable and reproducible transgene expression independent of proliferative or differentiated state using BAC TG-EMBED. Gene Ther. 25, 376–391 (2018).
Walker, J., Altman, R., Warren, J. & Altman, E. Using protein-based motifs to stabilize peptides. J. Pept. Res. 62, 214–226 (2003).
Oskolkov, N. et al. Biophysical characterization of human protamine-1 as a responsive CEST MR contrast agent. ACS Macro Lett. 4, 34–38 (2015).
Longo, D. L. et al. Iopamidol as a responsive MRI-chemical exchange saturation transfer contrast agent for pH mapping of kidneys: in vivo studies in mice at 7 T. Magn. Reson. Med. 65, 202–211 (2011).
Chen, L. Q. et al. Evaluations of extracellular pH within in vivo tumors using acidoCEST MRI. Magn. Reson. Med. 72, 1408–1417 (2014).
Yadav, N. N. et al. Detection of rapidly exchanging compounds using on-resonance frequency-labeled exchange (FLEX) transfer. Magn. Reson. Med. 68, 1048–1055 (2012).
Cohen, O., Huang, S., McMahon, M. T., Rosen, M. S. & Farrar, C. T. Rapid and quantitative chemical exchange saturation transfer (CEST) imaging with magnetic resonance fingerprinting (MRF). Magn. Reson. Med. 80, 2449–2463 (2018).
Zhou, Z. et al. Chemical exchange saturation transfer fingerprinting for exchange rate quantification. Magn. Reson. Med. 80, 1352–1363 (2018).
Heo, H.-Y. et al. Quantifying amide proton exchange rate and concentration in chemical exchange saturation transfer imaging of the human brain. Neuroimage 189, 202–213 (2019).
Kim, M., Gillen, J., Landman, B. A., Zhou, J. & van Zijl, P. C. Water saturation shift referencing (WASSR) for chemical exchange saturation transfer (CEST) experiments. Magn. Reson. Med. 61, 1441–1450 (2009).
Virtanen, P. et al. SciPy 1.0: fundamental algorithms for scientific computing in Python. Nat. Methods 17, 1–12 (2020).
Liu, G., Gilad, A. A., Bulte, J. W., van Zijl, P. C. & McMahon, M. T. High-throughput screening of chemical exchange saturation transfer MR contrast agents. Contrast Media Mol. Imaging 5, 162–170 (2010).
Stancanello, J. et al. Development and validation of a smoothing-splines-based correction method for improving the analysis of CEST-MR images. Contrast Media Mol. Imaging 3, 136–149 (2008).
Windschuh, J. et al. Correction of B1-inhomogeneities for relaxation-compensated CEST imaging at 7 T. NMR Biomed. 28, 529–537 (2015).
Zhou, I. Y. et al. Quantitative chemical exchange saturation transfer (CEST) MRI of glioma using image downsampling expedited adaptive least-squares (IDEAL) fitting. Sci. Rep. 7, 1–10 (2017).
Acknowledgements
The work was supported by the US National Institutes of Health Grants R01-CA203873, P41-RR14075, as well as by 1R01NS110942-01A1 and P01 CA163205 (EAC and HN) and NIH R01 NS098231; R01 NS104306 and P41 EB024495 (AAG). The Brigham and Women’s Small Animal Imaging Laboratory (SAIL) was funded by a G20 Grant (1G20RR031051-01) as part of the American Recovery and Reinvestment Act as a part of the construction of the Brigham and Women’s MRI Research Center (BWMRC). The 7T Bruker Small Bore Animal Magnet was partially funded by an S10 Grant (1S10OD010705-01) through the National Institutes of Health. This project has received funding from the European Union’s Horizon 2020 research and innovation programme under the Marie Skłodowska-Curie Grant Agreement No 836752 (OncoViroMRI). This paper reflects only the author’s view and the Research Executive Agency of the European Commission is not responsible for any use that may be made of the information it contains.
Author information
Authors and Affiliations
Contributions
A.A.G., M.T.M., and C.T.F. conceptualized the problem. O.P., H.I., A.A.G., M.T.M., H.N., E.A.C., and C.T.F. contributed to experimental design. A.A.G. designed the reporter gene. M.T.M. performed the peptide imaging study. H.I. and C.T.F. performed and analyzed the cell lysates study. H.I. performed the mice tumor implantation. O.P., H.I., and C.T.F. performed the in vivo imaging. O.P. and C.T.F. analyzed the in vivo data. O.P., H.I., A.A.G., M.T.M, H.N., E.A.C., and C.T.F. wrote and/or substantially revised the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Perlman, O., Ito, H., Gilad, A.A. et al. Redesigned reporter gene for improved proton exchange-based molecular MRI contrast. Sci Rep 10, 20664 (2020). https://doi.org/10.1038/s41598-020-77576-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-77576-z
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.