Abstract
Digital PCR (dPCR) technology has been proven to be highly sensitive and accurate in detecting copy number variations (CNV). However, a higher-order multiplexing dPCR assay for measuring SMN1 and SMN2 copy numbers in spinal muscular atrophy (SMA) samples has not been reported. Described here is a rapid multiplex SMA dPCR genotyping assay run on a fully integrated dPCR instrument with five optical channels. The hydrolysis probe-based multiplex dPCR assay quantifies SMN1, SMN2, and the total SMN (SMN1 + SMN2) while using RPPH1 gene as an internal reference control. The quadruplex assay was evaluated with characterized control DNA samples and validated with 15 blinded clinical samples from a previously published study. SMN1 and SMN2 copy numbers were completely concordant with previous results for both the control and blinded samples. The dPCR-based SMA copy number determination was accomplished in 90 min with a walk-away workflow identical to real-time quantitative PCR (qPCR). In summary, presented here is a simple higher-order multiplexing solution on a novel digital PCR platform to meet the growing demand for SMA genotyping and prognostics.
Similar content being viewed by others
Introduction
Proximal spinal muscular atrophy (SMA) is an early-onset neurodegenerative disease characterized by the loss of α-motor neurons in the anterior horn of the spinal cord which leads to muscle weakness and atrophy1,2. SMA has an autosomal recessive inheritance pattern with an incidence of 1 in 6000–10,000 births3,4. SMA results from the loss or mutation of survival motor neuron 1 (SMN1) on the q arm (q13) of chromosome 5 and retention of the paralogous survival motor neuron 2 (SMN2)5. SMN1 and SMN2 are nearly identical save 5 key nucleotide differences at the 3′ ends of the genes. SMN2 is functionally distinguishable from SMN1 by a single nucleotide difference (c.840C>T) in exon 7 that disrupts an exonic splice enhancer6,7. Even though close to 95% of SMA cases result from the loss of SMN1 and retention of SMN2, there is a wide spectrum of clinical disease severity based on the motor development milestones that are achieved8. Numerous studies using different assays and patient cohorts (reviewed in9) have demonstrated a strong inverse correlation between SMN2 copy number and disease severity. There are currently three USA Food and Drug Administration (FDA) approved therapeutic options; nusinersen (Spinraza, Biogen), onasemnogene abeparvovec (Zolgensma, AveXis/Novartis) and risdiplam (Evrysdi, Roche)10,11,12,13,14. These agents target SMN2 expression. SMN2 copy number is used to design the therapeutic regimens for SMA patients receiving these approved therapies, with immediate treatment for patients with 2 or 3 copies of SMN215,16. A simple yet highly accurate and sensitive solution for SMN1 and SMN2 copy number detection is needed to meet the rapidly growing demand for informed SMA treatment.
Current genotyping and copy number determination methods for SMA are complex, time consuming or lack resolution with higher copy numbers. Multiplex ligation-dependent probe amplification (MLPA) and real-time quantitative PCR (qPCR) can detect SMN1 deletions17,18,19,20. MLPA is a multi-step method requiring post-PCR capillary electrophoresis and a long time-to-result (~ 20 h)18,19. Alternatively, qPCR requires normalization with standard curves. More importantly, neither MLPA nor qPCR can consistently distinguish unit differences in SMN1 or SMN2 when the copy number is greater than 320,21,22. Genomic sequencing (GS) technology has made progress on copy number detection. A recent publication reported SMA diagnosis and carrier identification using whole genome sequencing and advanced copy number analysis software23. However, there are still limitations to GS for diagnosis including the cost, sample processing time, complex data analysis and the need for orthogonal validation.
Digital PCR (dPCR) alleviates the limitations previously noted to provide absolute quantification of a target gene within a sample for molecular diagnostics and prognostics24. With dPCR, the prepared PCR sample is distributed across a large number of physically isolated micro-reaction partitions so that each partition will have single digit counts of template DNA or none at all25,26. The absolute concentration (and confidence level) of the target gene(s) can be calculated by counting the number of positive partitions (containing at least one target molecule) and the number of negative partitions (containing no target molecules) and using a Poisson-based statistical correction. dPCR can reliably and accurately measure SMN1 and SMN2 copy numbers over a wide range, between 0 and 6 copies26,27. Due to the optical configuration of first-generation dPCR platforms, most of the currently available multiplex assays are duplex or variations of a duplex assay. A recently published report on multiplex dPCR describes simultaneous identification of SMN1 and SMN2 copy numbers by using different signal amplitudes within the same optical channel28. The Combinati (Palo Alto, California) Absolute Q Digital PCR System (Fig. 1) is a single benchtop instrument with 5 optical channels and a qPCR-equivalent, walkaway workflow29. Instead of generating air and water emulsions, the Absolute Q features rigid micro-injection-molded partitioning consumable plates which ensure highly consistent partition yield and volume. In this study, the Absolute Q is used to demonstrate the first “higher order” multiplex dPCR assay for simultaneous SMA diagnostic, i.e. loss of SMN1, and prognostic, i.e. SMN2 copy number, assessment.
Results
Multiplex SMA genotyping assay
The hydrolysis probe-based multiplex assay is designed to use FAM, VIC, TYE665, and TAMRA to identify SMN1, SMN2, total SMN (SMN1 + 2), and RPPH1, respectively (Fig. 2). The SMN1 probe (FAM) hybridizes to nucleotide c.840C while the SMN2 probe (VIC) hybridizes to nucleotide c.840T. The total SMN (SMN1 + 2, TYE665) targets intron 1—a region that is identical between SMN1 and SMN2. The internal reference control is designed to target the highly conserved RPPH1 gene (TAMRA), which is consistently present as 2 copies30.
Schematic representation of the 4-color multiplex assay design showing the positions of the 4 probes. Arrows represent forward and reverse primers of each assay. The bars connecting two circles represent probes while the color-filled circles represent different fluorophores and the black circles represent dark quenchers. The dashed lines (//) indicate discontinuous sequence.
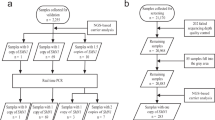
The workflow with the Absolute Q dPCR System involved fewer steps when compared to the SMN1 and SMN2 copy number assays previously developed on the QuantStudio 3D dPCR System (Fig. 3)27. As the components of the dPCR assay are integrated into a single unit, the Combinate Absolute Q dPCR assay required less hands-on time for the operator. Furthermore, the multiplex SMA genotyping assay was accomplished in 90 min on the Absolute Q dPCR System. The SMN1 and SMN2 copy number analysis was completed in 5.5 h on the QuantStudio 3D dPCR System27.
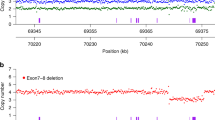
The optimized assay was first tested on control DNA samples containing SMN1 only (NA17117), SMN2 only (NA23255), or both SMN1 and SMN2 (NA03815). Representative images of the dPCR results from the analysis software (Fig. 4A) shows strong resolution of the partitions. All SMN probes (SMN1 exon 7, SMN2 exon 7 and total SMN intron 1) were normalized against the reference gene (RPPH1) that contains 2 copies per genome. The scatter plots for SMN1 exon 7 (Fig. 4B), SMN2 exon 7 (Fig. 4C), and total SMN intron 1 (Fig. 4D) shows robust separation between the target and reference probes. The multiplex SMA genotyping assay run on the Absolute Q shows high specificity and sensitivity in classifying the control samples with SMN1:SMN2 copy numbers of 0:3 (NA23255), 3:0 (NA17117), and 1:1 (NA03815). For NA03815, the Absolute Q dPCR data showed copy numbers for SMN1 exon 7, SMN2 exon 7, and total SMN intron 1 as 1.1, 1.1 and 2.2, respectively.
Representative partition image (A) and scatter plots (B–D) of the SMA 4-color multiplexing dPCR assay using NA03815 genomic DNA as the sample. The scatter plots are shown for SMN1 exon 7 (B; FAM), SMN2 exon 7 (C; VIC) and total SMN intron 1 (D; TYE665), all relative to the reference gene RPPH1 (TAMRA). The analysis software subtracted the pre-PCR image intensities from the post-PCR image intensities so as to remove fluorescent signals which did not exhibit amplification behavior, thereby eliminating false positive signals. If the pre-PCR partitions were brighter than the post-PCR partitions, then the analysis software would record negative fluorescent units. These negative units, however, did not impact the copy number quantification results.
The Absolute Q dPCR assays require less template genomic DNA (2.5 ng) than the QuantStudio 3D dPCR assays which uses 30–60 ng of EcoRI-digested genomic DNA. Genomic DNA up to 25 ng was also tested on the Absolute Q. There are no apparent differences in SMN1 and SMN2 copy number quantification completed with 2.5 ng template or with 25 ng genomic DNA (Supplementary Table S1).
Quantification of control gDNA SMN1 and SMN2 copy numbers using absolute Q dPCR
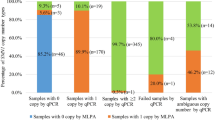
Copy numbers of SMN1, SMN2 and total SMN in 10 genomic DNA test samples from Coriell Cell Repositories were quantified using the SMA multiplex 4-color assay. All 10 Coriell DNA samples were run in triplicate on the Absolute Q dPCR System for assay verification and repeatability. The intra-assay variability—measured by %CV—between the SMN1 (Fig. 5A), SMN2 (Fig. 5B) and total SMN (Fig. 5C) is low, demonstrating strong repeatability. The intraclass correlation coefficients (ICCs) were 0.993 (95% confidence interval (CI) 0.979–0.998) for the SMN1 assay, 0.995 (95% CI 0.985–0.999) for the SMN2 assay and 0.987 (95% CI 0.963–0.996) for the total SMN assay. The SMN1 and SMN2 copy numbers measured with this multiplex assay are concordant with those provided by Coriell Cell Repositories, including one sample with 5 copies of SMN2. Additionally, the 6 copies of total SMN for NA03814 correctly matched the corresponding sums of SMN1 and SMN2 copy numbers. The total SMN copy numbers in all of the samples except for NA11254 were equal to their corresponding sums of SMN1 and SMN2 copy numbers.
Repeatability of the Absolute Q dPCR assays for SMA genotyping. The control DNA samples (n = 10) from Coriell Cell Repositories were assayed for SMN1 (A), SMN2 (B) and total SMN (C) copy numbers. Each sample was assayed in triplicate. The intra-assay precision, measured by %CV, for each assay was listed below each sample on the plots. The intra-assay precision measurements were not calculated (N/C) for those samples with a copy number less than 1.00.
SMN1 and SMN2 copy number validation of blinded clinical samples
Absolute Q and QuantStudio 3D array dPCR SMA assays were compared on a set of blinded samples derived from patient lines (n = 15). This set of samples contained gDNAs from 13 SMA patients and 2 non-SMA controls. Using the SMN1 copy number assay, the Absolute Q dPCR correctly identified all SMA samples with homozygous deletions of SMN1 (12 out of 15) within the cohort. The Absolute Q dPCR identified one of the patient-derived sample (MND10) as containing 1 copy of SMN1 and one copy of SMN2. This sample was from an individual harboring a SMN1(p.A2G) missense mutation27. The Absolute Q dPCR SMN1 (Fig. 6A), SMN2 (Fig. 6B) and total SMN (Fig. 6C) copy number measurements of the blinded clinical samples were concordant with those results obtained using QuantStudio 3D array dPCR27,31, including a sample (MND12) with 5 copies of total SMN. Bland Altman analysis of the results from SMN1 (Fig. 6D), SMN2 (Fig. 6E) and total SMN (Fig. 6F and Table 1) probes demonstrated strong agreement between Absolute Q and QuantStudio 3D dPCR assays. For each assay, 93% of the test samples fell within the limits of agreement. Additionally, the total SMN copy numbers from 14 blinded clinical samples agree with the sum of the copy numbers from SMN1 and SMN2 assays. In a single sample (MND03), the total SMN copy number at exon 7 (4 copies) was greater than the sum of the SMN1 exon 7 (0 copies) and SMN2 exon 7 (2 copies) copy numbers. Based on this result, this sample was hypothesized to have 2 copies of either SMN1 or SMN2 containing a partial deletion of exon 7. The partial deletion of exon 7 in this sample was confirmed orthogonally by junctional PCR (data not shown) and by GS23.
Agreement between the QuantStudio 3D and the Absolute Q dPCR assays. (A–C) Scatterplots for SMN1 (A), SMN2 (B) and total SMN (C) copy numbers measured with the QuantStudio 3D and the Absolute Q dPCR assays. The line of equality for each set of assays is shown as a dashed line in each plot. (D–F) Bland Altman difference plots for SMN1 (D), SMN2 (E) and total SMN (F) assays. For each assay, the bias is shown as a solid blue line that is labeled as the mean and the limits of agreement defined and labeled as the mean ± 1.96SD (standard deviation).
Discussion
Accurate quantification of SMN1 and SMN2 copy numbers is essential for the diagnosis of SMA as well as for the development of therapeutic strategies to treat the disease. This study describes the development of a multiplex dPCR assay that provides accurate and rapid SMN1 and SMN2 copy number quantification. The Absolute Q dPCR System provides reliable measurements of SMN1 and SMN2 copy numbers with a workflow that is similar to qPCR, reducing hands-on time and time-to-results when compared to other currently available dPCR platforms.
Some dPCR copy number variation assays rely on digestion of the template genomic DNA with a restriction endonuclease23. This is because sequence repeats of two or more copies can be in close proximity to each other, increasing the probability that they will be partitioned in the same microchamber and result in a miscalculation of the copy number. Restriction endonuclease digestion of the template DNA helps to reduce the probability of proximal gene copies partitioning in the same microchamber. In this study, SMN1 and SMN2 copy numbers were accurately measured without restriction endonuclease digestion of the sample genomic DNA. Each target-specific assay was designed with distinct fluorophores that can reliably differentiate between SMN1 and SMN2; even though these genes share highly homologous DNA sequences. Additionally, the high partition numbers of the microfluidic device (20,000 partitions) and the low DNA input amount (~ 2 to 3 ng per reaction) facilitate single molecule partitioning. Finally, SMN1 and SMN2 genes, each approximately 28 kb in length, are both located in the same region of chromosome 5 but they are separated from each other by about 875 kb of genomic sequence. Mechanical sheering of the genomic DNA during DNA sample isolation may further fragment this region thereby reducing the likelihood of both genes being in the same partition. Eliminating the restriction digestion step in the copy number assay, therefore, simplifies the workflow of the assay. Future studies would assess the effects of specimen source and DNA isolation procedure on the need for either a shearing or digestion step prior to dPCR analysis.
In addition to eliminating the need for a pre-PCR digestion step, the dPCR workflow is simplified by integration of multiple systems into a single instrument. The first-generation of dPCR systems have separate instruments for microfluidic partitioning of the PCR reactions and template DNA, PCR amplification and raw data acquisition. This required significant hands-on-time and training for each unique workflow, which, in turn, increased the probability of operator error or process variability. The Absolute Q dPCR System combines sample partitioning, thermocycling, and raw data acquisition into a single benchtop instrument. As a result, the Absolute Q dPCR System has an average run-time that is 73% faster than first-generation dPCR systems27,31. The shortened run-time along with minimal hands-on operation could facilitate platform adoption into diagnostic and prognostic settings.
In addition to determining SMN1 and SMN2 copy numbers, dPCR can also identify partial deletions in SMN1/SMN2 and gene conversion events that generate hybrid SMN genes. By comparing total SMN copy number to the sum of SMN1 exon 7 and SMN2 exon 7 copy numbers, partial deletions of SMN1 or SMN2 can be identified. One of the patient-derived cell lines tested in this study (MND003) contained a partial deletion of exon 7, which was confirmed orthogonally (data not shown). Additionally, one of the control samples (NA11254) possibly contains a partial deletion as the total SMN copy number was found to be 3 at intron 1 while the sum of SMN1 and SMN2 copy numbers at exon 7 was 2. The discrepant results with this sample are not a consequence of a lack of specificity in the intron 1 primers or probe since the in silico specificity scores for these sequences were high, i.e. all had a BLAT score of 22, which exceeds the minimal threshold of 2032. These results need to be confirmed orthogonally by junctional PCR or GS in order to verify that the partial deletion is indeed the reason for the discrepancy. Partial SMN deletions, particularly resulting from the loss of exons 7 and/or 8 (SMN1/2Δ78), have been previously reported in the literature using several approaches including MLPA and GS23,33. Gene conversion events can create hybrid SMN genes where part of the gene is SMN1 and another part is SMN234. The most commonly identified hybrid gene is SMN1 with c.840T (i.e. SMN2), however there are other SMN1/SMN2 hybrid gene variations. Other SMN1/SMN2 hybrid genes have different exon 7 inclusion efficiencies compared to SMN1 or SMN235,36. It is important to correctly identify hybrid SMN genes as they can potentially lead to unexpected responses to current therapeutic options. Future work involving the development of a panel of assays targeting the single nucleotide difference between SMN1 and SMN2 at exon 8, as well as the intronic single nucleotide differences, would be useful for the identification of partial deletions in SMN genes as well as hybrid SMN genes.
A limitation of the copy number assay described in this study is its inability to detect silent SMA carriers (2 + 0) where both copies of SMN1 are present in cis on the same chromosome. A single nucleotide polymorphism within SMN1 (g.27134T>G; rs143838139) is associated with the cis configuration of SMN137. In addition, this dPCR assay cannot detect intragenic point mutations in SMN1, which account for 3–5% of SMA cases8. Future studies are needed to develop dPCR assays that are able to detect these single nucleotide changes. These assays could potentially be integrated into an expanded multiplexing assay panel alongside the current SMN1/SMN2 differentiating dPCR assays described here.
Newborn screening of SMA is becoming more available and routine in the United States as well as many other countries38,39. Dried blood spots (DBS) are a common source material for SMA newborn screening. A 3-mm diameter DBS typically yields ~ 140 ng gDNA (in a volume of 50 μL) using column-based DNA extraction40. This study demonstrates that SMN1 and SMN2 copy numbers can be accurately measured with as little as 2–3 ng gDNA, so the Absolute Q and MAP plates could reliably be used for orthogonal validation of SMA carrier status or SMN2 copy number. As the technology is further developed, quadraplex dPCR could potentially become a primary SMA screening tool.
The acquisition cost for the Absolute Q dPCR System is comparable to high-end qPCR instrumentation, and somewhat lower than other currently available dPCR platforms. In this study, the consumable (SMA reagents and MAP plates) costs were $2.36 per test. This is less than typical per-sample costs for available droplet dPCR systems ($3.80 per sample41) and other technologies such as mass spectrometry ($3.00 per sample42). These per-sample costs are exclusive of the cost associated with labor, which is reduced with the lower hands-on time of the Absolute Q dPCR System.
The Absolute Q dPCR quadruplex assay approach could be extended to identification of copy number variations associated with other diseases, in addition to SMA. dPCR has been used to identify rare gene variants as well as differences in copy number of multiple genes associated with pediatric-onset disorders (reviewed in24). We have demonstrated that this system can identify a rare single nucleotide variant (EGFR(T790M)) associated with non-small cell lung carcinoma as well as fusion genes (BCR-ABL1 and CCDC88C-FLT3) that are associated with different types of cancer29.
Accurate quantification of SMN1 and SMN2 copy numbers is essential for the proper diagnosis of SMA and the identification of carriers. Similarly, accurate measurement of SMN2 copy number is essential to determine therapeutic regimens for treating children with SMA15,16. Digital PCR—including the Absolute Q dPCR System presented here—can accurately, reliably and rapidly quantify SMN1 and SMN2 copy numbers in genomic DNA samples. Future work will assess the feasibility of this dPCR platform for diagnostic as well as prognostic purposes using well-established assay validation approaches and different types of DNA samples43.
Materials and methods
DNA samples
Control samples of human genomic DNA (n = 10) were obtained from Coriell Cell Repositories (Camden, NJ; Table 2). Most of these samples (n = 8) had well characterized SMN1 and SMN2 copy numbers provided by Coriell Cell Repositories. Among the 10 samples, 5 were derived from SMA patients with 0 copies of SMN1. The copy numbers for NA11254 and NA17117 were measured using high-resolution melting analysis and array digital PCR (personal communications). NA03814 and NA03815 samples were from known SMA carriers, i.e. 1 copy of SMN1.
Patient-derived samples (n = 15) were procured from the Motor Neuron Diseases Research Laboratory (Nemours Alfred I. duPont Hospital for Children). Genomic DNA was isolated from patient-derived fibroblast and lymphoblastoid cell lines and the SMN1 and SMN2 copy numbers were measured using the QuantStudio 3D Digital PCR System (Life Technologies, Waltham, MA) as described previously27,29. Approval for cell line generation was provided by the Nemours Institutional Review Board and this study is registered on ClinicalTrials.gov (NCT01754441 and NCT02532244). This study was approved by the Nemours Institutional Review Board (#764456). Written informed consent or assent was obtained for each patient-derived cell line. Each cell line was de-identified. For all samples tested, the tester was blinded to the previously measured SMN1 and SMN2 copy numbers. All procedures performed in this study involving human participants were in accordance with the ethical standards of the institutional and/or research committees and with the 1964 Helsinki declaration and its later amendments of comparable ethical standards.
Absolute Q digital PCR system and microfluidic array partitioning (MAP) plates
The Combinati Absolute Q Digital PCR System consists of Microfluidic Array Partitioning (MAP) consumable plates and a fully integrated instrument that automates partitioning of reagents in the plate, PCR thermocycling and 5-color fluorescence image acquisition. The MAP plate has a standard microplate footprint capable of running up to 16 samples per dPCR run29. Each unit is loaded with 10 µL of sample mixture and more than 90% of the loaded sample is partitioned and analyzed in ~ 20,000 pico-scale partitions. The partition volume in MAP plates is defined by the physical dimensions of the microarray chambers and not by a stochastic process such as fluid emulsions, which is important because consistent partition volume is a critical component of the dPCR statistical correction model. The physical array ensures that all samples across all plates yield ~ 20,000 analyzed partitions and minimal sample is lost to dead volume or compromised partitions.
The Absolute Q digital PCR System has a walk-away workflow identical to traditional qPCR. The MAP plate is loaded via pipette with 10 μL of PCR mix and an overlay of 10 μL Isolation Buffer in each well. The wells are then capped with specialized gaskets. The plate is then placed into the Absolute Q tray and retracted into the system. The Absolute Q uses positive pressure from an on-board compressor to partition the sample within the MAP plate without the need for microfluidic valves, sealing films or other moving parts. The MAP slides are constructed of a cyclo olefin polymer (COP; 80 μm) film that seals the microfluidic features that are molded into a separate piece of thicker material. Four identical slides are bonded to a rigid, microtiter format plate frame that includes the loading wells to complete the plate assembly. The thin film becomes gas permeable when positive pressure is applied to a well containing reagents. As the reagent enters the microfluidic features, air is passed out of the partition through the film. This allows reagent to completely fill dead-ended partitions and prevents any bubbles from forming inside of the microfluidic features. The Isolation Buffer overlay follows the reagent and physically separates the reagent reaction volumes to complete the partitioning. Positive pressure applied to the consumable during the PCR thermocycling prevents any evaporation and ensures that bubbles will not form and disrupt the isolated micro-reactions.
Before and after PCR thermocycling, entire arrays are imaged with up to five optical channels configured for the most commonly used dyes, including a ubiquitous quality control dye (ROX) used to verify proper partition filling and finding. The images taken before PCR are subtracted from the after-PCR images to remove background noise. Combinati Analysis software automatically applies optical crosstalk compensation and classifies the partitions using a convoluted neural network algorithm to eliminate false positives/ negatives and ensure robust quantification results. The full dPCR process occurs within the single benchtop instrument without operator’s interaction after setting up the protocol parameters (Fig. 1). The streamlined workflow reduces the potential for contamination, minimizes human handling errors, and reduces the time to result. No fluids ever contact instrument components, so minimal system maintenance is required.
Digital PCR primer and probe design
The primer and probe oligos were designed using PrimerQuest Tool (Integrated DNA Technologies, Coralville, IA). All primers were checked for target sequence specificity using NCBI Primer-BLAST44 and UCSC In-Silico PCR (https://genome.ucsc.edu/, last accessed May 27, 2020) using default settings. For Primer-BLAST, each primer was required to have at least 2 total mismatches to unintended targets, including at least 2 mismatches with the last 5 base pairs at the 3′ end. For In-Silico PCR, the minimum match size was 15 and a minimum BLAT32 score of 20, using the GRCh38/hg38 human genome assembly. Primers and probes were also evaluated for primer dimers and cross primer interactions using Multiple Primer Analyzer (Thermo Fisher Scientific, Waltham, MA) with the optimal sensitivity value (3) for dimer detection. All of the oligos were synthesized by Integrated DNA Technologies and Thermo Fisher Scientific. The assay includes a total of 3 pairs of primers and 4 probes with the SMN1 and SMN2 targets sharing a common pair of primers. The FAM-labeled SMN1 and VIC-labeled SMN2 probes were designed utilizing the signature c.840 single nucleotide difference between exon 7 of SMN1 (c.840C) and SMN2 (c.840T). The oligos for total SMN copy number were designed upstream in intron 1 targeting both SMN1 and SMN2 genes, and the probe was labeled with TYE665. The TAMRA-labeled probe for RNase P (RPPH1; OMIM #608513) targeted the single exon region of RPPH1 gene (Fig. 2). All probes have either QSY or IBRQ dark quenchers on their 3′ ends (Table 3).
Digital PCR
The control DNA samples were diluted to 10 ng/µL working stock. The blinded DNA samples were measured using Qubit 4 fluorometer (Invitrogen) and diluted to 10 ng/µL. The PCR reaction mix contained 1X Combinati MasterMix, 2.5–25 ng human gDNA, 1800 nmol/L SMN1/SMN2 primers, 900 nmol/L RPPH1 primers, 900 nmol/L total SMN primers, and 250 nmol/L of each probe. Each MAP plate well was loaded by hand via pipette with 10 µL of PCR reaction mix then overlaid with 10 µL Isolation Buffer. Gasket caps were placed and the plate was put into the Absolute Q and run with the following conditions: 3 min activation at 95 °C, 40 cycles of 5 s at 95 °C and 30 s at 62 °C. The fully automated Absolute Q Control Software (v1.0.20, https://www.combinati.com/) controls the sample digitization into the MAP partitions, thermal cycling, and imaging.
Data analysis
The copy numbers were determined using the Absolute Q Analysis Software (v10.5.4). The software automatically calculates the optimal positive/ negative threshold and in this case the positive signals were five times higher than the negative signals. For each unit, the software displays the total viable partition count and the positive partition count. The software has a setting to identify the sample types as a copy number variation assay and applies the appropriate calculations to automatically display the copy numbers for the samples. The copy numbers of SMN1, SMN2 and total SMN are normalized by the reference control RPPH1 using the following equations:
where N = total number of viable partitions, N1 = number of SMN1 positive partitions, N2 = number of SMN2 positive partitions, N3 = number of total SMN positive partitions and N4 = number of RPPH1 positive partitions.
Statistical analysis
The coefficient of variability (%CV), defined as the standard deviation divided by the mean value for each set of replicates, was used to assess repeatability, or intra-assay precision. Reliability was also measured by the intraclass correlation coefficient (ICC) using SPSS v.25 (IBM, Armonk, NY). A Bland Altman agreement analysis45 using SigmaPlot v.12.0 (Systat Software, Inc., San Jose, CA) was used to measure the agreement between dPCR methods.
Data availability
The datasets generated or analyzed during the current study are available from the corresponding author on reasonable request.
References
Crawford, T. O. & Pardo, C. A. The neurobiology of childhood spinal muscular atrophy. Neurobiol. Dis. 3, 97–110 (1996).
Tisdale, S. & Pellizzoni, L. Disease mechanisms and therapeutic approaches in spinal muscular atrophy. J. Neurosci. 35, 8691–8700 (2015).
Pearn, J. Incidence, prevalence and gene frequency studies of chronic childhood spinal muscular atrophy. J. Med. Genet. 15, 409–413 (1978).
Cuscó, I. et al. Prenatal diagnosis for risk of spinal muscular atrophy. Br. J. Obstet. Gynaecol. 109, 1244–1249 (2002).
Lefebvre, S. et al. Identification and characterization of a spinal muscular atrophy-determining gene. Cell 80, 155–165 (1995).
Lorson, C. L., Hahnen, E., Androphy, E. J. & Wirth, B. A single nucleotide in the SMN gene regulates splicing and is responsible for spinal muscular atrophy. Proc. Natl. Acad. Sci. USA 96, 6307–6311 (1999).
Monani, U. R. et al. A single nucleotide difference that alters splicing patterns distinguishes the SMA gene SMN1 from the copy gene SMN2. Hum. Mol. Genet. 8, 1177–1183 (1999).
Kolb, S. J. & Kissel, J. T. Spinal muscular atrophy. Neurol. Clin. 33, 831–846 (2015).
Butchbach, M. E. R. Copy number variations in the survival motor neuron genes: implications for spinal muscular atrophy and other neurodegenerative diseases. Front. Mol. Biosci. 3, 7 (2016).
Chiriboga, C. A. et al. Results from a phase I study of nusinersen (ISIS-SMNrx) in children with spinal muscular atrophy. Neurology 86, 890–897 (2016).
Finkel, R. S. et al. Treatment of infantile-onset spinal muscular atrophy with nusinersen: a phase 2, open-label, dose-escalation study. Lancet 388, 3017–3026 (2016).
Haché, M. et al. Intrathecal injections in children with spinal muscular atrophy: nusinersen clinical trial experience. J. Child Neurol. 31, 899–906 (2016).
Mendell, J. R. et al. Single-dose gene-replacement therapy for spinal muscular atrophy. N. Engl. J. Med. 377, 1713–1722 (2017).
Sturm, S. et al. A phase 1 healthy male volunteer single escalating dose study of the pharmacokinetics and pharmacodynamics of risdiplam (RG7916, RO7034067), a SMN2 splicing modifier. Br. J. Clin. Pharmacol. 85, 181–193 (2019).
Glascock, J. et al. Treatment algorithm for infants diagnosed with spinal muscular atrophy through newborn screening. J. Neuromuscul. Dis. 5, 145–158 (2018).
Glascock, J. et al. Revised recommendations for the treatment of infants diagnosed with spinal muscular atrophy via newborn screening who have 4 copies of SMN2. J. Neuromuscul. Dis. 7, 97–100 (2020).
Feldkötter, M., Schwarzer, V., Wirth, R., Wienker, T. F. & Wirth, B. Quantitative analyses of SMN1 and SMN2 based on real-time LightCycler PCR: fast and highly reliable carrier testing and prediction of severity of spinal muscular atrophy. Am. J. Hum. Genet. 70, 358–368 (2002).
Arkblad, E. L. et al. Multiplex ligation-dependent probe amplification improves diagnostics in spinal muscular atrophy. Neuromuscul. Disord. 16, 830–838 (2006).
Huang, C. H. et al. Copy number analysis of survival motor neuron genes by multiplex ligation-dependent probe amplification. Genet. Med. 9, 241–248 (2007).
Gómez-Curet, I. et al. Robust quantification of the SMN gene copy number by real-time Taqman PCR. Neurogenetics 8, 271–278 (2007).
Alías, L. et al. Accuracy of marker analysis, quantitative real-time polymerase chain reaction and multiple ligation-dependent probe amplification to determine SMN2 copy number in patients with spinal muscular atrophy. Genet. Test. Mol. Biomarkers 15, 587–594 (2011).
Prior, T. W., Nagan, N., Sugarman, E. A., Batish, S. D. & Braastad, C. Technical standards and guidelines for spinal muscular atrophy testing. Genet. Med. 13, 686–694 (2011).
Chen, X. et al. Spinal muscular atrophy diagnosis and carrier screening from genome sequencing data. Genet. Med. 22, 945–953 (2020).
Butchbach, M. E. R. Applicability of digital PCR to the investigation of pediatric-onset genetic disorders. Biomol. Detect. Quant. 10, 9–14 (2016).
Sykes, P. J. et al. Quantitation of targets for PCR by use of limiting dilution. Biotechniques 13, 444–449 (1992).
Vogelstein, B. & Kinzler, K. W. Digital PCR. Proc. Natl. Acad. Sci. USA 96, 9236–9241 (1999).
Stabley, D. L. et al. SMN1 And SMN2 copy numbers in cell lines derived from patients with spinal muscular atrophy as measured by array digital PCR. Mol. Genet. Genom. Med. 3, 248–257 (2015).
Vidal-Folch, N. et al. Multiplex droplet digital PCR method applicable to newborn screening, carrier status and assessment of spinal muscular atrophy. Clin. Chem. 64, 1753–1761 (2018).
Dueck, M. E. et al. Precision cancer monitoring using a novel, fully integrated, microfluidic array partitioning digital PCR platform. Sci. Rep. 9, 19606 (2019).
Baer, M., Nilsen, T. W., Costigan, C. & Altman, S. Structure and transcription of a human gene for H1 RNA, the RNA component of human RNase P. Nucleic Acids Res. 18, 97 (1990).
Stabley, D. L. et al. Establishing a reference dataset for the authentication of spinal muscular atrophy cell lines using STR profiling and digital PCR. Neuromuscul. Disord. 27, 439–446 (2017).
Kent, W. J. BLAT: the BLAST-like alignment tool. Genome Res. 12, 656–664 (2002).
Vijzelaar, R. et al. The frequency of SMN gene variants lacking exon 7 and 8 is highly population dependent. PLoS ONE 14, E0220211 (2019).
Burghes, A. H. M. When is a deletion not a deletion? When it is converted. Am. J. Hum. Genet. 61, 9–15 (1997).
Kashima, T., Rao, N. & Manley, J. L. An intronic element contributes to splicing repression in spinal muscular atrophy. Proc. Natl. Acad. Sci. USA 104, 3426–3431 (2007).
Wu, X. et al. A-44G transition in SMN2 intron 6 protects patients with spinal muscular atrophy. Hum. Mol. Genet. 26, 2768–2780 (2017).
Luo, M. et al. An ashkenazi jewish SMN1 haplotype specific to duplication alleles improves pan-ethnic carrier screening for spinal muscular atrophy. Genet. Med. 16, 149–156 (2014).
Chien, Y. H. et al. Presymptomatic diagnosis of spinal muscular atrophy through newborn screening. J. Pediatr. 190, 124–129 (2017).
Kraszewski, J. N. et al. Pilot study of population-based newborn screening for spinal muscular atrophy in New York state. Genet. Med. 20, 608–613 (2018).
Durie, D. et al. Quantification of DNA in neonatal dried blood spots by adenine tandem mass spectrometry. Anal. Chem. 90, 801–806 (2018).
Lin, Y. et al. Newborn screening for spinal muscular atrophy in China using DNA mass spectrometry. Front. Genet. 10, 1255 (2019).
Vidal-Folch, N. et al. A droplet digital PCR method for severe combined immunodeficiency newborn screening. J. Mol. Diagn. 19, 755–765 (2017).
Milosevic, D. et al. Applying standard clinical chemistry assay validation to droplet digital PCR quantitative liquid biopsy testing. Clin. Chem. 64, 1732–1742 (2018).
Ye, J. et al. Primer-BLAST: a tool to design target-specific primers for polymerase chain reaction. BMC Bioinform. 13, 134 (2012).
Bland, J. M. & Altman, D. G. Statistical methods for assessing agreement between two methods of clinical measurement. Lancet 327, 307–310 (1986).
Acknowledgements
We would like to thank those SMA patients and their families who provided samples for this study. This work was supported by Combinati, the Nemours Foundation (M.E.R.B.) and the National Institutes of Health (R44OD023028 to P.H. and P20GM103464 to M.E.R.B.).
Author information
Authors and Affiliations
Contributions
L.J. and M.E.R.B. conceived and designed the experiments. L.J., R.L., S.G., A.Z., M.E.R.B. and P.H. performed the experiments and analyzed the data. L.J., R.L., A.Z., M.E.R.B. and P.H. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jiang, L., Lin, R., Gallagher, S. et al. Development and validation of a 4-color multiplexing spinal muscular atrophy (SMA) genotyping assay on a novel integrated digital PCR instrument. Sci Rep 10, 19892 (2020). https://doi.org/10.1038/s41598-020-76893-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-76893-7
This article is cited by
-
Increased AID results in mutations at the CRLF2 locus implicated in Latin American ALL health disparities
Nature Communications (2024)
-
Next generation sequencing is a highly reliable method to analyze exon 7 deletion of survival motor neuron 1 (SMN1) gene
Scientific Reports (2022)
-
Detection of SMN1 to SMN2 gene conversion events and partial SMN1 gene deletions using array digital PCR
neurogenetics (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.