Abstract
Our main aim was to investigate the diversity, origin and biogeographical affiliations of freshwater gammarids inhabiting the Aegean Islands by analysing their mtDNA and nDNA polymorphism, thereby providing the first insight into the phylogeography of the Aegean freshwater gammarid fauna. The study material was collected from Samothraki, Lesbos, Skyros, Evia, Andros, Tinos and Serifos islands as well as from mainland Greece. The DNA extracted was used for amplification of two mitochondrial (COI and 16S) and two nuclear markers (28S and EF1-alpha). The multimarker time-calibrated phylogeny supports multiple origins and different diversification times for the studied taxa. Three of the sampled insular populations most probably represent new, distinct species as supported by all the delimitation methods used in our study. Our results show that the evolution of freshwater taxa is associated with the geological history of the Aegean Basin. The biogeographic affiliations of the studied insular taxa indicate its continental origin, as well as the importance of the land fragmentation and the historical land connections of the islands. Based on the findings, we highlight the importance of studying insular freshwater biota to better understand diversification mechanisms in fresh waters as well as the origin of studied Aegean freshwater taxa.
Similar content being viewed by others
Introduction
The Mediterranean islands are considered natural laboratories of evolution, exhibiting high levels of diversity and endemism, making them a vital part of one of the globally most precious biodiversity hotspots and a model system for studies of biogeography and evolution1,2,3,4. It is estimated that the Mediterranean region, including continental Europe, Northern Africa and Middle East covers approximately 2 million km2 which equals ca. 1.3% of the world’s land surface. Yet it is inhabited by ca. 6% of the world’s freshwater species with at least 43% of them considered to be local endemics5, however, the freshwater fauna of the Mediterranean islands remains largely understudied.
The Aegean Sea is one of the major sea basins within the Mediterranean, which houses around 7500 islands and islets occurring at a variety of isolation levels and exhibiting different topographic features. The Aegean region is recognized to have, globally, the highest number of islands in a single sea basin6. The largest of the Aegean islands is Crete, being also the fifth largest in the Mediterranean region. The majority of the islands were once part of a single landmass called Aegeis, which emerged probably around 23 Ma7. In the Middle Miocene, the movement of the African and Eurasian continental plates led eventually to the first fragmentation of Aegeis and to the isolation of Crete from the Peloponnese3,8. Further isolations of the land units within the Aegean basin were caused by the formation of Mid-Aegean Trench (around 12–9 Ma), which led to the separation of the central Aegean from the east Aegean islands3,9. Another crucial event in the history of the entire region was the closure of the Mediterranean basin that took place at the end of Miocene (6–5 Ma, Messinian Salinity Crisis), leading to complete desiccation and, consequently, to the mass extinction of the marine biota. During that time, many formerly isolated islands regained their connections with the mainland and/or the other islands, which resulted in the exchange of terrestrial and freshwater faunas3. Afterwards, some of the connections with the mainland were again temporarily re-established during the Pleistocene, due to eustatic sea-level changes and recurrent glaciation events, with most of the islands gaining their present shape and the current level of isolation at the end of Pleistocene10.
Gammarid amphipods are among the most abundant aquatic macroinvertebrates in fresh waters in Europe and, particularly, in the Mediterranean Region11. They are known to shape the freshwater macrozoobenthic communities, being recognised as aquatic keystone species12. Given the exclusively aquatic life cycle and high level of diversity, freshwater gammarids are considered to be particularly interesting evolutionary models13. However, the freshwater gammarid fauna is relatively poorly known with most of the studies on the Mediterranean amphipods being focused on the marine biota. Currently, about 120 freshwater gammarid species have been reported from the Mediterranean region, while only 24 species have been reported from the Mediterranean islands. They belong mainly to two genera: Gammarus Fabricius, 1775 and Echinogammarus Stebbing, 189914,15,16,17,18. Given that recently a high rate of overlooked diversity was discovered in both the above-mentioned genera [e.g.19,20,21,22,23,24,25,26], it may be reasoned that the current number of species already described from the Mediterranean islands is largely underestimated. To date, there have been five species reported from the Aegean Islands with three of them being Cretan endemics (E. kretensis Pinkster, 1993, E. platvoeti Pinkster, 1993 and the recently described G. plaitisi Hupało, Mamos, Wrzesińska & Grabowski, 2018) and with single records of G. komareki Schäferna, 1923 from Gökçeada and G. uludagi G.S. Karaman, 1975 from Lesbos and Evia14,15,18,27,28.
In this paper, we investigate the presence of freshwater populations of Gammarus on other Aegean islands. Given the high rate of endemism already observed in numerous organisms from the Aegean archipelago3, including freshwater Gammarus14,18, one can hypothesise that there will be a high level of local endemism on other Aegean islands as well. However, given that final separation of particular Aegean islands occurred at different geological times, with some being isolated since the end of Miocene (e.g. Crete) and some still bearing recurrent land connections with the mainland as late as during Pleistocene (e.g. Evia, Lesbos, Samothraki), it can also be hypothesised that the level of endemism will vary among islands. In this case, the islands that still had land connections in Pleistocene, will perhaps not have endemic lineages, due to possible faunal exchanges with the mainland. These hypotheses are tested by performing DNA polymorphism analyses on four molecular markers: two mitochondrial (COI and 16S rDNA) and two nuclear (28S rDNA and EF1-alpha) in the freshwater populations of gammarids from the Aegean islands and neighbouring, continental regions. By doing so, the aim is to reveal their biogeographic affiliations and the possible origin to provide the first insight into the phylogeography of the Aegean freshwater amphipod fauna.
Materials and methods
Sample collection and identification
The study material consisted of 50 individuals of the genus Gammarus collected between 2008 and 2018 from 13 sampling sites, including nine sites from freshwater habitats on the Aegean islands, namely Samothraki, Skyros, Evia, Lesbos, Andros, Tinos and Serifos and four sites from mainland Greece (Table 1). Gammarids were sampled using a variety of methods, including collection from gravel, rocks and aquatic vegetation with a hand net or using rectangular kick sample nets (aperture 25 × 25 cm and 0.5 mm mesh size). Samples were sorted on-site, and gammarids were immediately fixed in 96% ethanol. Specimens were examined under a Nikon SMZ 800 stereomicroscope and identified using the available Mediterranean freshwater gammarid taxonomic literature14,15,16,17.
DNA extraction, PCR amplification, sequencing
The DNA was extracted using the standard phenol/chloroform method29, implementing the protocol described previously14. The extracted DNA was stored at 4 °C until amplification and finally long-term stored at − 20 °C. In the first step, a fragment of the cytochrome c oxidase subunit I gene (COI) was amplified using three different primer pairs, depending on the amplification success. In the second step, at least one individual per the delimited Molecular Operational Taxonomic Unit (MOTU) (see below) was amplified for additional markers that were used in phylogeny reconstruction—mitochondrial 16S rRNA and nuclear markers being 28S rRNA and the EF1-alpha gene. All the primer sequences, PCR conditions and original references for all the molecular markers used in this study are provided in Table S1. Afterwards, all PCR products (5 µl) were cleaned up using exonuclease I (ThermoFisher Scientific) and alkaline phosphatase FastAP (ThermoFisher Scientific) according to the manufacturer’s guidelines. Direct sequencing was performed using the same forward primers as for amplification, using the BigDye terminator technology in Macrogen sequencing company. Since the nuclear 28S rRNA marker is over 1100 bp long, it was sequenced both ways, which allowed for obtaining the full coverage, additionally providing better resolution and identification of polymorphic sites. Given that other nuclear marker used in this study, EF1-alpha, has a shorter length and did not exhibit a significant level of polymorphic sites, it was sequenced using only the forward primer.
Sequence data authentication, editing, alignment, deposition and reference material
All obtained sequences were confirmed as belonging to Gammarus via BLASTn searches in GenBank30. Subsequently, they were assembled, aligned and trimmed to 625 (COI), 391 (16S), 1107 (28S) and 602 (EF1-alpha) base pairs respectively, using the Geneious 10.0.9 software package31. The sequences of two gene-coding markers (COI and EF1-alpha) were translated to check for stop codons. Alignments were performed using the MAFFT plugin with G-INS-i algorithm in Geneious. In case of double peaks and low-quality regions detected in some of the sequences of 28S, the two strands were compared and the dominant signal was chosen at each problematic site. No ambiguous sites were detected in the analysed 28S sequences.
All the sequences were deposited in GenBank (accession numbers to be provided upon acceptance). Additionally, the sequences of all markers used in this study were compiled in the dataset and deposited in the public repository of the Barcode of Life Data Systems (BOLD)32, where all the relevant metadata information and sequence trace files are available (https://doi.org/10.5883/DS-GAEG).
The dataset was also supplemented by multimarker dataset of 56 reference sequences (30—COI, 10—16S, 11- 28S, 5—EF1-alpha) from the public repositories, representing the related Gammarus species from the Aegean islands, mainland Greece and adjacent regions were added to the dataset, along with a single sequence of isopod Asellus aquaticus used for rooting the tree (Table S2).
MOTU delimitation and interspecific relationships
The Molecular Operational Taxonomic Units (MOTUs) were delimited using COI sequence dataset. Firstly, according to the distance-based Automatic Barcode Gap Discovery (ABGD) methodology33. The results of the genetic distance-based, ABGD MOTU delimitation were cross-validated with the phylogenetic tree based delimitation methods, namely: Generalized Mixed Yule Coalescent (GMYC) single and multiple models34,35 using GMYC Web Server (https://species.h-its.org/gmyc/) and multi-rate Poisson tree processes (mPTP)36 using MCMC chain of 50 million iterations with a burn-in of 1 million. Additionally, we have measured patristic distances using Patristic 1.037, according to a patristic distance threshold of 16% proposed for crustaceans at the COI locus38, based on the prior observations of morphological differences across multiple crustacean species. We have also measured K2p distances in MEGA7 software to compare them with obtained patristic distances39. For tree-based MOTU delimitation methods, we have obtained a consensus tree using BEAST 2.4.7 after performing three MCMC runs of 10 M iterations, sampled every 1000 iterations, using Tamura-Nei model, selected as a best-fit model of evolution with bModel test40 and Birth–Death tree model set as a prior, chosen as best-fit tree model using path-sampling. MCMC runs were examined using Tracer v1.7.1 and all the sampled parameters for each studied MOTU achieved sufficient effective sample sizes (ESS > 200). None of the reference sequences mentioned in Table S2 were included in the MOTU delimitation processing.
For final visualisation, the neighbour-joining tree of all COI sequences, using the Kimura 2-parameter (K2p) model of evolution with 1000 bootstrap replicates, was created in MEGA7 software39.
Phylogeny reconstruction, time calibration and history of the diversification
The dataset used for reconstruction of time-calibrated phylogeny consisted of a single representative individual per each delimited MOTU, according to ABGD and patristic distance delimitation methods along with 11 reference Gammarus sequences as well as individuals, which represented respective calibration points described below. The substitution saturation was tested in DAMBE 7.0.2841, using the index proposed42, to assess the potential loss of phylogenetic signal. No significant saturation was detected (Iss value lower than Iss.c value; p < 0.01; Table S3) for any molecular marker. Based on the best partitioning scheme selected by the PartitionFinder43, we divided the molecular data into seven partitions: 16S, 28S with three codon positions of COI and two partitions of EF1-alpha, one comprising codon positions 1. and 2. and another one with 3. codon position. The time-calibrated phylogeny was reconstructed in BEAST 2.4.7 package44, performing three MCMC chains of 100 million iterations, sampled every 2000 iterations, using the best-fit substitution models (all listed in Table S4) determined by bModel test40. The optimal molecular clock as well as tree model were chosen via path sampling/stepping-stone procedures using three runs per clock model and tree model, analysing the marginal likelihoods and Bayes factors (BF)45, using BEAST 2.4.7. Provided the strict clock was rejected for each partition (BF > 50), we used the uncorrelated log-normal relaxed clock46. Given that Yule and Birth–Death tree models were equally supported by path-sampling, we have chosen the Yule model as a tree prior, being a simpler model. Additionally, two more MCMC chains of 100 million iterations, with sampling every 2000 iterations with same substitution models, but with joint COI codon positions and TN93 with G and I set as a consensus best-fit substitution model were performed to determine the estimated COI substitution rate.
For molecular clock calibration, we used five primary calibration points known from literature and related to geological events, as well as one secondary calibration point, which helped to validate the clock calibration based upon the primary calibration points (Table S5). The most recent calibration point is based on the radiation of the endemic Gammarus species flock in Lake Ohrid, coinciding with the emergence of the lake itself47,48, which most probably took place ca. 2 Ma. The second calibration point is based on the split between the Black Sea and the Caspian Sea populations of Pontogammarus maeoticus, estimated at about 4 Ma, connected with the shifts of continental plates, causing the of former Pontian Lake into Black and Caspian Sea, respectively49,50. The third calibration point reflects the estimated time of diversification of the Gammarus fossarum species complex that took place in the Carpathians, being caused by the Middle Miocene subsidence event between ca. 15 and ca. 17 Ma19. The fourth one marks the origin of the Acanthogammaridae family, endemic to Lake Baikal, estimated at 28–30 Ma51,52,53. The oldest calibration point reflects the connection between the Eocene regression of the Paratethys Sea at ca. 37 Ma and the divergence between Sarothrogammarus and Rhipidogammarus genera23,27. The constraints of the calibration points were applied by imposing priors on the respective tree nodes, using lognormal distribution of the MRCA priors, which allowed the incorporation of the possible uncertainty of the data. For cross-validating the ages of the nodes obtained using primary calibration points listed above, we have used four additional calibration points based on the fossil record, using three fossil amphipods and one basal eumalacostracan as an outgroup (Table S6). A detailed description of the fossil calibration points is provided in54. Besides using the sequences available for the molecular clock calibration, the reference sequences of Gammarus available from the literature were used in the reconstruction of phylogeny to provide further insights into the phylogenetic and biogeographical affinities within the Aegean gammarids (all individuals listed in Table S2).
Parameters of all three runs were examined in Tracer 1.7.1 and reached the ESS values above 200. The runs were combined and resampled with LogCombiner 2.4.7 with 30% burn-in, with the maximum clade credibility chronograms being annotated using TreeAnnotator 2.4.4 and visualised using FigTree 1.4.444.
The history of diversification was inferred using the lineage through time (LTT) plot generated in Tracer 1.7.1 from the 1000 trees, subsampled in Logcombiner, obtained from the Bayesian time-calibrated phylogeny reconstruction performed in BEAST 2.4.7 software.
Results
Taxonomic assignment and geographical distribution
All the collected Gammarus individuals were morphologically assigned to the so-called Gammarus pulex group [sensu15] reported from Greece and adjacent areas. Specimens from Andros, Evia and Skyros, were morphologically identified as the pan-European morphospecies Gammarus pulex (Linnaeus, 1758). Species from Tinos and Serifos were identified as Gammarus plaitisi, previously considered as a Cretan endemic. Specimens from Lesbos and Samothraki were ascribed to Gammarus uludagi and Gammarus arduus G.S. Karaman, 1975, respectively. The latter has been previously reported from mainland Greece and from the Balkan Peninsula, but never from any of the Aegean islands. Gammarus arduus was present in two samples from mainland Greece, namely Arisvi (Thraki) and Amfitriti (Thraki). Material from mainland Greece included samples from the Pelion Peninsula and Sofades (Karditsa, Thessaly) containing other members of Gammarus pulex group [sensu15] reported from adjacent regions; Gammarus crenulatus, described from Greece with its locus typicus, being the same as sampled in this study. Since some of the individuals could not be assigned with certainty to any known morphospecies, we have decided to use Open Nomenclature (ON) qualifiers55, widely accepted in taxonomic nomenclature. All of those individuals were identified as members of genus Gammarus. The individuals from Evia, Skyros and Andros were identified as Gammarus aff. pulex, fitting some of the diagnostic features of the species with a noticeable variation in setation patterns. Given those uncertainties, they were classified as Gammarus sp.1, Gammarus sp.2 and Gammarus sp.3. The individuals from Pelion were identified as G. aff. birsteini. However, given that G. birsteini is so far known only from limited areas in eastern Turkey and Kazakhstan and that not all of the morphological features fitted the studied specimens, we consider it as a different, yet undiscovered, species, thus assigning it to Gammarus sp.4.
MOTU delimitation, diversity, affiliations and distribution
The ABGD, along with MOTU delimitation based on the patristic distance, supported the existence of eight distinct lineages of Gammarus present in our material collected from the islands and from mainland Greece (Fig. 1). For tree-based delimitation methods, the single species hypothesis was rejected for both single and multiple approaches (result of Likelihood Ratio tests < 0.0001). The delimitation results slightly differed among the methods used, namely GMYC single threshold model, mPTP and GMYC multiple threshold. The first two methods indicated the presence of ten MOTUs, with two extra lineages within G. plaitisi and G. arduus, respectively. Moreover, GMYC multiple threshold model supported twelve MOTUs, splitting G. plaitisi into three units, as well as two extra lineages within Gammarus from Skyros and Andros. We have chosen both ABGD, cross-validated with patristic distance method as the main delimitation methods due to its most conservative approaches (Fig. 1). Moreover, the patristic distances seemed to be also congruent with the K2p distances, which further supported the delimited MOTUs (Table S8). All of the insular MOTUs grouped together with other Gammarus pulex sequences, accompanied by other members of the Gammarus pulex species group including G. uludagi, G. lacustris or G. alpinus (sensu Karaman & Pinkster, 1977a). However, the samples from mainland Greece with G. crenulatus and G. sp.4 grouped, with high bootstrap value, together with a reference sequence of G. roeselii. The latter belongs to the so-called Gammarus roeselii group [sensu16 (Fig. 1).
Neighbor-joining COI tree from data obtained in the study (tip labels marked in bold) and mined from NCBI GenBank with the addition of the outgroups. The numbers by respective nodes indicate bootstrap values ≥ 0.75. The scale bar corresponds to the K2p distance. The bars represent different delimitation methods used; colours used for ABGD MOTUs correspond to those presented in other figures.
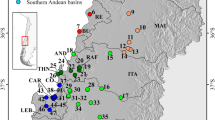
Each of the studied islands is presumably inhabited by a single MOTU only, with the exception of Evia, where both MOTUs of Gammarus sp.1 from this study and G. uludagi are present (Fig. 2). In most cases, a MOTU found on one island is present neither on any other island nor on mainland Greece. The only exception is Gammarus plaitisi present on Tinos and Serifos as well as on Crete14 and Gammarus arduus present on both Samothraki and on the mainland (Fig. 2).
Map of the sampling sites on Aegean islands. The colours and symbols correspond to those presented in other figures. The map was generated using QGIS 2.18.381 (https://www.qgis.org/en/site/).
Phylogeny reconstruction, molecular dating and history of the diversification
The estimated COI substitution rate reached the value of 0.0118 ± 0.00677 substitutions/site/My, which is congruent with the substitution rates reported in other studies dating divergences in other crustaceans and arthropods, including freshwater amphipods [0.007–0.0177 substitutions/site/My]; [e.g.25,54,56,57,58].
The ages of the nodes obtained using primary calibration points were generally congruent with the dating of the nodes through fossil calibration, however the 95% HPD values were generally wider than those obtained through primary calibration scheme (Fig. 3, Table S7). It was expected as the calibration points used in fossil calibration scheme derive from outgroups more distant to the target taxa used in this study and the dates have considerable uncertainty. Since the topology of the obtained consensus tree is congruent (Fig. S1), and all the well-supported nodes were retrieved as well as their dates see
Maximum clade credibility, time-calibrated Bayesian reconstruction of phylogeny of Aegean Gammarus MOTUs. Phylogeny was inferred from sequences of the mitochondrial: COI, 16S rRNA markers and nuclear: 28S rRNA,EF1-α markers. The numbers by respective nodes indicate Bayesian posterior probability values ≥ 0.8. The coloured bars represent ABGD delimitation method, with colours, and symbols and shapes corresponding to those presented in other figures. Dark grey node bars represent 95% HPD obtained from the primary calibration points, whereas the light grey node bars represent 95% HPD obtained from the fossil calibration points. EOC—Eocene, OLI—Oligocene, MIO—Miocene, PLI—Pliocene, PLE—Pleistocene.
m to be supported by both calibration schemes, we have decided to present and discuss the nodes obtained via primary calibration scheme as reliable proxy for species’ divergence.
The time-calibrated phylogeny revealed that divergence within the Gammarus pulex group from the studied region started around 12 Ma (95% HPD: 18.9—8.8 Ma) (Fig. 3). All of the deeper divergence events between the insular taxa seem to take place between 12 and 8 million years ago. At that time, G. platisi seemed to diverge from the G. pulex lineage from mainland Greece. Gammarus uludagi from Lesbos diverged from its conspecific from Evia as well as G. pulex from Evia and Peloponnese and G. pulex from Skyros and Andros diverged from other lineages (Fig. 3). During the same timescale, about 11.5 Ma (95% HPD: 20.8–5.5 Ma), G. crenulatus diverged from G. sp.4. These two species, along with G. roeselii, could have separated from Gammarus pulex group members already in the Eocene, around 40 Ma. More recently, in the Pliocene and Pleistocene, diversification events probably took place only in the insular populations of G. plaitisi, about 2.5 Ma (95% HPD: 4.1–0.9 Ma) as well as in G. arduus, where the insular population from Samothraki diverged from the mainland conspecifics, most probably about 2 Ma (95% HPD: 2.8–0.7 Ma). The youngest divergence seemed to take place in late Pleistocene, about 0.2 Ma (95% HPD: 0.45–0.03 Ma), when the population of G. plaitisi from Serifos apparently diverged from the one from Crete (Fig. 3).
The lineages-through-time plot (Fig. S3) shows that the accumulation of lineages remained rather constant over time, with no significant increase in lineage accumulation.
Discussion
Diversity and distribution of Aegean insular freshwater fauna
In this study, we provide the first evidence of the presence of freshwater populations of Gammarus on five Aegean islands, namely Samothraki, Skyros, Andros, Tinos and Serifos, with three of these populations most probably representing the new, distinct species supported by all delimitation methods used in this study. The same holds true for another, possibly new species, inhabiting the limited area in mainland Greece. The ABGD approach used in this study as the main MOTU delimitation method is considered to be more conservative compared to tree-based methods like GMYC and closer to the species distinction provided by taxonomists [e.g.33,59]. In some cases, these methods over-split putative entities, depending on the overall genetic distances differential or sampling bias60,61. However, cross-validating those methods with the conservative approach of patristic distance raises up the probability of the congruence of delimited taxa38. On the other hand, it still poses the question of unanimity of the phylogenetically delimited species with the biological species concept62. Recently, based on the experimental observations done on freshwater gammarids, it was argued that lineages separated by a genetic distance exceeding 4% are less likely to form precopulatory pairs and thus, might be reproductively isolated63. Even though amphipods diverged by ca. 16% still formed precopulatory pairs under laboratory conditions, this was never observed in the field. However, no further evidence supporting the cross-lineage fertility and presence of hybrid offspring produced in the laboratory conditions was provided in referenced study. It was then argued by the authors that lineages separated by ca. 16% living in sympatry seem to exhibit prezygotic barriers, preventing them from mating with divergent counterparts. In this case, the majority of the insular Aegean species were isolated by more than 16% genetic distance from the closest related lineage, both when using commonly applied for calculating genetic distances Kimura-2-parameter model (K2p) as well as the patristic distance (comparison in Table S8). According to the 4.3% K2p genetic distance threshold proposed for gammarids by64 and even more conservative, 16% patristic distance threshold proposed by38, one may suppose that they are likely to represent separate and reproductively isolated lineages.
Up to now, in the Aegean basin, members of the genus Gammarus were reported only from Lesbos, Gökçeada (Imbros), Thasos, Evia and Crete15,27,28,65. Interestingly, none of these records indicated any endemic insular species. Our results reveal that almost every lineage is endemic to one island only (Fig. 2). Apart from the recent study by14, describing the new species G. plaitisi and G. uludagi from Evia provided by27, there were no other molecular studies conducted on any Aegean gammarids. Given that molecular studies on freshwater insular gammarids are scarce, one could expect a high number of overlooked diversity on the islands. Considering the fact that Aegean archipelago is characterized by an exceptionally high level of endemism confirmed in numerous biota3, it is also probable that some of the overlooked Aegean lineages may, in fact, represent undiscovered endemic taxa. That is a particularly valid assumption for the distinct lineages from Evia, Skyros and Andros, as no individuals belonging to these MOTUs have been reported from elsewhere nor detected in our study.
Miocene diversification of the Aegean freshwater insular gammarids
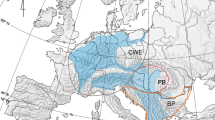
Our results suggest that diversification of freshwater gammarids on the Aegean islands started in the Middle Miocene, around 12 Ma (Fig. 3). Up to that point, the Aegean region remained a single landmass (Aegeis), comprising not only all the present islands but also the Balkan Peninsula and Asia Minor7. Around 12 Ma, the fragmentation of the Aegeis landmass began, due to collision of the African tectonic plate with the Eurasian plate in the Middle Miocene66. In fact, the movements of landmasses during that time supposedly induced divergence events in numerous freshwater crustaceans including amphipods, both epigean20,21,25 and subterranean67, isopods68 and crabs69. Moreover, at that time, the formation of the Mid-Aegean Trench started and was fully accomplished by 10–9 Ma, resulting in separation of the western part of the Aegean region from the eastern one (Fig. 4)9. These events have led to numerous isolation and diversification episodes in the Aegean fauna, which are reflected in modern distribution patterns3. Interestingly, in the Aegean, Middle Miocene events are, in general, known to affect the divergence mainly of terrestrial taxa, such as snails, beetles, isopods and scorpions3. Seemingly, all of the Aegean freshwater biota diverging after the end of Miocene or more recently, in Pliocene and/or Pleistocene including species of crabs, crayfish and snails, used temporarily existing land connections70,71,72. Our results suggest, however, that the divergence of the Aegean insular freshwater gammarids was most probably affected by the Middle Miocene land movements and fragmentation, which also might have had a significant impact on their current distribution. For example, the divergence of G. uludagi from Lesbos, belonging to the eastern Aegean islands, from its conspecific from Evia, coincides with the approximate time of the formation of the Mid-Aegean Trench. The same is true for the divergence of G. pulex inhabiting the same island from the lineage inhabiting Peloponnese, being on the western side of Aegean basin. This supports the possible connectivity of the divergence with the geological history. It is also suggested that the formation of the Mid-Aegean Trench might have played a role in the evolution and dispersal of freshwater Pseudorientalia snails73. The authors suggest that limited distribution of Pseudorientalia, currently inhabiting only Aegean islands east of the Mid-Aegean Trench, might be associated with the formation of this biogeographical barrier and subsequent regional land fragmentation in the Middle Miocene. Similarly, the land fragmentation in the Middle Miocene most likely affected other divergence events shown in our data, such as the divergence of G. plaitisi from Crete, Tinos and Serifos from G. pulex inhabiting mainland Greece, including the Peloponnese, which coincides with the isolation of Crete and the separation on the islands from Peloponnese (Figs. 3, 3). It also supports the results of the previous study on the divergence of Cretan G. plaitisi (Hupało et al.14). The land separation at that time most probably resulted also in the divergence between the Gammarus from Andros and the one from Skyros (Figs. 3, 4). However, given the low posterior probability of that node, still more data is needed to determine the actual relationships between these taxa.
The selection of paleogeographical maps of the Aegean Region, including its islands (after Popov et al.82). The dotted line represents the approximate location of the Mid-Aegean Trench. The maps were generated using QGIS 2.18.381 (https://www.qgis.org/en/site/).
Most recent Plio–Pleistocene diversification events
Although major divergence events in Aegean freshwater gammarids took place in the Middle Miocene, some of them occurred more recently, in the Plio-Pleistocene (Fig. 3). The Pliocene water level fluctuations, as well as Pleistocene glaciation events with associated sea regression and climate aridification, strongly affected the river systems and promoted the diversification of numerous biota in the Mediterranean Region3,21,74. Several studies have confirmed that major European peninsulas, including the Balkan Peninsula, served as refugia and diversification hotspots for numerous taxa, eventually becoming starting points for their dispersal21,25,75. In the Aegean, this was also the time of intense diversification events for numerous vertebrate taxa, mostly involving ‘herptiles’ and mammals3 and references therein. Fossil evidence indicates the presence of two pygmy elephant species in the Aegean region, which diversified in the Pleistocene and Holocene3. Our data suggest that the separation between G. arduus from Samothraki and the mainland conspecifics took place during the same time, from the late Pliocene to the beginning of the Pleistocene (Fig. 3). This makes Samothraki, based on the up-to-date molecular evidence, the only known Aegean island not inhabited by gammarid endemics (Fig. 1). Favourably, this divergence event could be associated with the recurrent Pleistocene land connections between the island and the continent3,10. One could argue then for the plausibility of a similar scenario for the other confirmed records of gammarid presence in the Aegean, given that islands like Lesbos, Gökçeada and Thasos—all sharing the temporal land connections with the continent at a similar time as Samothraki did10. However, up to now, no molecular evidence is available to confirm or reject such a hypothesis. On the other hand, although Evia was connected with the mainland in Pleistocene, the lineage most likely diverged earlier, in Miocene (Fig. 3). However, this might also be due to undersampling from the mainland and neighbouring islands.
Even more recent is the diversification within G. plaitisi from the three Aegean islands: Crete, Tinos and Serifos (Figs. 2, 3). By confirming the presence of G. plaitisi on Tinos and Serifos, we are rejecting the earlier-proposed alleged endemism of this species on Crete14. This finding, along with the very low intraspecific haplotypic diversity on Crete, confirms Plio/Pleistocene dispersal of this species suggested by the authors14. Still, it is unclear how the species dispersed from Serifos to Crete, as there were no known temporal land connections between these two islands in Pleistocene (Fig. 4)3. It is equally puzzling to the evolutionary history and dispersal of the freshwater Potamon crabs, that diverged and dispersed to Crete, as well as, to Cyprus during Pleistocene, where no land connections were known to exist between the islands and the mainland70. It is suggested by the authors that early humans might have aided in the dispersal of certain taxa, including crabs, which would then mean that the arrival of these freshwater biotas to Crete may be very recent. Another plausible scenario involves the passive dispersal by birds [e.g.76], already suggested for freshwater Daphniola snails inhabiting distant eastern Aegean islands77. It is argued that the snail lineages probably diverged recently, in Pleistocene, where no land connection was presumably present between those islands, and thus, the dispersal was possible either due to the mediating factor or through the successive dispersal through neighbouring islands77. It might be the case also for G. plaitisi with possible intermediate, yet still undiscovered, populations, e.g. on Milos or Kythera and Antikythera islands.
Taxonomic affiliations of the Aegean Gammarus
The results of our study provide new evidence for rejecting the monophyly of Gammarus pulex with three potentially new distinct taxa within this morphospecies (Fig. 3). These findings confirm the recently observed high cryptic diversity and lack of monophyly in numerous widespread European freshwater gammarids [e.g.19,20,21,22,26,78,79]. Interestingly, our data also support the polyphyly of Gammarus uludagi. This finding questions the reliability of the specimen recorded from Evia, as the species was originally described and reported from the western part of Asia Minor and the island of Lesbos15. Nevertheless, the incongruences in the species’ integrity raise further questions about the taxonomic congruence of other formerly described species (Fig. 3). This seems to be especially valid for specimens assigned to Gammarus pulex. The species exhibits a significant level of intraspecific morphological variation in several characters such as e.g. the number of segments in flagella, the shape of epimeral plates, the number of spines and the setation pattern on various body parts15. In our study, we have observed mostly the differences in setation on epimers, pereiopods and uropod 3. Although we did not recognize any stable patterns which could reliably distinguish the specimens from particular lineages, we believe that a detailed morphometric study could help to resolve the taxa boundaries. Moreover, the taxonomic affiliation of G. crenulatus shown in our results (Figs. 1, 3) also suggests the incongruence of morphogroups, the so-called Gammarus pulex, Gammarus roeselii and Gammarus balcanicus groups, formerly described by15,16,17. Even though G. crenulatus was originally assigned to the Gammarus pulex group, our results suggest that it is, in fact, more closely related to G. roeselii than to G. pulex (Fig. 3). Most of the recent thorough phylogenetic studies on gammarids23,27 did not include any molecular data on G. crenulatus, so more studies are needed to fully resolve this matter. The divergence between G. roeselii from mainland Greece, G. crenulatus and G. sp.4 took place around 20 Ma, well before the fragmentation of Aegeis. It suggests that other processes might have played a role in their divergence, however again more data is needed to reveal the detailed evolutionary history of this group. These data, along with other recent findings21 support the need for a comprehensive revision of Gammarus pulex and further studies on Gammarus roeselii morphospecies, incorporating an integrative approach using the detailed morphological information, ideally, combined with extensive molecular data. Using such a perspective would provide a major step towards fully resolving the species’ phylogenetic relationships.
Conclusions
The results of our study suggest a high level of local lineage endemism for gammarids in the Aegean islands, which is in agreement with previous studies showing that several taxonomic groups exhibit a high level of local endemism in this part of the Mediterranean. . The presence of G. arduus on Samothraki supports our hypothesis that the level of endemism will vary between the islands, with the absence of endemic lineages on islands that were still connected with the mainland during the Pleistocene. On the other hand, the presence of distinct, separate MOTUs on Evia and Lesbos leads to reject this hypothesis; however, more data are needed both from the regions adjacent to Evia as well as from inland waters of Turkey, where G. uludagi has been reported. Moreover, the results of our study support the presence of at least four yet undescribed gammarid species, three of which are endemic to respective Aegean islands they inhabit, namely Evia, Skyros and Andros.
The results of the time-calibrated phylogeny indicate multiple origins and different timescales of differentiation for the Aegean insular freshwater gammarids. The biogeographic affiliations of the studied insular taxa indicate their continental origin as well as the importance of the land fragmentation and the historical land connections of the islands that most probably influenced the evolutionary history of the Aegean biota. Moreover, the deep divergences inferred from the reconstructed phylogenies indicate that most gammarid species present in the Aegean islands diverged well before the final isolation of some islands, indicating that they may be considered the only known freshwater taxa belonging to ‘the old colonizers’ (sensu3). These results not only provide important evidence supporting the survival of at least some freshwater taxa of the Messinian Salinity Crisis, but also may be highly useful as calibration points e.g. formation of Mid-Aegean Trench or recent isolation of Samothraki from the mainland, for future studies of Aegean taxa.
Given the scarcity of available data, one cannot exclude further discoveries of freshwater gammarids from other Aegean islands. More molecular data and thorough sampling of the area are essential to provide a detailed picture of the evolutionary history of Aegean freshwater insular gammarids. This is especially important given that Mediterranean islands are among the most anthropogenically affected regions in the Mediterranean region and are also one of the least studied in terms of their freshwater diversity80. Considering that the vast majority of freshwater ecosystems of the Aegean islands are under significant threat due to increasing water and habitat demands for tourism development, agriculture and pastoral development, there is an urgency for more studies revealing the actual biodiversity of insular freshwater biota that may aid in planning a reasonable and sustainable strategy for their conservation80. The results presented here provide not only additional evidence indicating strong connectivity between the evolution of the freshwater biota and the geological history of the Mediterranean, but also broaden the still scarce knowledge on the evolutionary mechanisms of the diversification of the insular freshwater fauna.
Data availability
All DNA sequences and original chromatograms are stored in the BOLD database (https://doi.org/10.5883/DS-GAEG), and the processed DNA sequences are also available in GenBank under following accession numbers (to be provided). The location data, respective alignments and additional metadata are also available from BOLD.
References
Myers, N., Mittermeier, R. A., Mittermeier, C. G., Da Fonseca, G. A. & Kent, J. Biodiversity hotspots for conservation priorities. Nature 403, 853 (2000).
Whittaker, R. J. & Fernández-Palacios, J. M. Island Biogeography: Ecology, Evolution, and Conservation (Oxford University Press, Oxford, 2007).
Poulakakis, N. et al. A review of phylogeographic analyses of animal taxa from the Aegean and surrounding regions. J. Zool. Syst. Evol. Res. 53, 18–32 (2015).
4Woodward, J. The Physical Geography of the Mediterranean. Vol. 8 (Oxford University Press on Demand, 2009).
De Figueroa, J. M. T., López-Rodríguez, M. J., Fenoglio, S., Sánchez-Castillo, P. & Fochetti, R. Freshwater biodiversity in the rivers of the Mediterranean Basin. Hydrobiologia 719, 137–186 (2013).
Triantis, K. A. & Mylonas, M. Greek islands biology. In Encyclopedia of Islands (eds Gillespie, R. & Glague, D. A.) 388–392 (University of California Press, Berkeley, CA, 2009).
Meulenkamp, J. The Neogene in the southern Aegean area. Opera Bot. 30, 5–12 (1971).
van der Geer, A., Dermitzakis, M. & de Vos, J. Crete before the Cretans: The reign of dwarfs. Pharos 13, 121–132 (2006).
9Dermitzakis, M. & Papanikolaou, D. In Annales geologiques des pays helleniques. 245–289.
Perissoratis, C. & Conispoliatis, N. The impacts of sea-level changes during latest Pleistocene and Holocene times on the morphology of the Ionian and Aegean seas (SE Alpine Europe). Mar. Geol. 196, 145–156 (2003).
MacNeil, C., Dick, J. T. & Elwood, R. W. The trophic ecology of freshwater Gammarus spp. (Crustacea: Amphipoda): Problems and perspectives concerning the functional feeding group concept. Biol. Rev. 72, 349–364 (1997).
Kelly, D. W., Dick, J. T. & Montgomery, W. I. The functional role of Gammarus (Crustacea, Amphipoda): Shredders, predators, or both?. Hydrobiologia 485, 199–203 (2002).
Bilton, D. T., Freeland, J. R. & Okamura, B. Dispersal in freshwater invertebrates. Annu. Rev. Ecol. Syst. 32, 159–181 (2001).
Hupało, K., Mamos, T., Wrzesińska, W. & Grabowski, M. First endemic freshwater Gammarus from Crete and its evolutionary history—An integrative taxonomy approach. PeerJ 6, e4457 (2018).
Karaman, G. S. & Pinkster, S. Freshwater Gammarus species from Europe, North Africa and Adjacent Regions of Asia (Crustacea-Amphipoda).: Part I. Gammarus pulex-Group and Related Species. Bijdragen tot de Dierkunde 47, 1–97 (1977).
Karaman, G. S. & Pinkster, S. Freshwater Gammarus species from Europe, North Africa and adjacent regions of Asia (Crustacea-Amphipoda): Part II. Gammarus roeseli-group and related species. Bijdragen tot de Dierkunde 47, 165–196 (1977).
Karaman, G. S. & Pinkster, S. Freshwater Gammarus Species from Europe, North Africa and Adjacent Regions of Asia (Crustacea-Amphipoda).: Part III. Gammarus balcanicus-Group and Related Species. Bijdragen tot de Dierkunde 57, 207–260 (1987).
Pinkster, S. A revision of the genus Echinogammarus Stebbing, 1899 with some notes on related genera (Crustacea, Amphipoda). Memorie del Museo Civico di Storia Naturale (IIa serie) Sezione Scienze della Vita (A. Biologia) 10, 1–183 (1993).
Copilaş-Ciocianu, D. & Petrusek, A. The southwestern Carpathians as an ancient centre of diversity of freshwater gammarid amphipods: Insights from the Gammarus fossarum species complex. Mol. Ecol. 24, 3980–3992 (2015).
Copilaş-Ciocianu, D. & Petrusek, A. Phylogeography of a freshwater crustacean species complex reflects a long-gone archipelago. J. Biogeogr. 44, 421–432 (2017).
Grabowski, M., Mamos, T., Bącela-Spychalska, K., Rewicz, T. & Wattier, R. A. Neogene paleogeography provides context for understanding the origin and spatial distribution of cryptic diversity in a widespread Balkan freshwater amphipod. PeerJ 5, e3016 (2017).
Grabowski, M., Wysocka, A. & Mamos, T. Molecular species delimitation methods provide new insight into taxonomy of the endemic gammarid species flock from the ancient Lake Ohrid. Zool. J. Linn. Soc. 181, 272–285 (2017).
Hou, Z., Sket, B. & Li, S. Phylogenetic analyses of Gammaridae crustacean reveal different diversification patterns among sister lineages in the Tethyan region. Cladistics 30, 352–365 (2014).
Katouzian, A.-R. et al. Drastic underestimation of amphipod biodiversity in the endangered Irano-Anatolian and Caucasus biodiversity hotspots. Sci. Rep. 6, 22507 (2016).
Mamos, T., Wattier, R., Burzyński, A. & Grabowski, M. The legacy of a vanished sea: A high level of diversification within a European freshwater amphipod species complex driven by 15 My of Paratethys regression. Mol. Ecol. 25, 795–810 (2016).
Mamos, T., Wattier, R., Majda, A., Sket, B. & Grabowski, M. Morphological vs molecular delineation of taxa across montane regions in Europe: The case study of Gammarus balcanicus Schäferna, (Crustacea: Amphipoda). J. Zool. Syst. Evol. Res. 52, 237–248 (2014).
Hou, Z., Sket, B., Fišer, C. & Li, S. Eocene habitat shift from saline to freshwater promoted Tethyan amphipod diversification. Proc. Natl. Acad. Sci. 108, 14533–14538 (2011).
Özbek, M. & Özkan, N. Gökçeada içsularının Amphipoda (Crustacea: Malacostraca) faunası Amphipoda (Crustacea: Malacostraca) fauna of the inland-waters of Gökçeada Island. Ege J. Fish. Aquat. Sci. 34, 63–67 (2017).
Hillis, D. M., Moritz, C., Mable, B. K. & Olmstead, R. G. Molecular systematics Vol. 23 (Sinauer Associates, Sunderland, 1996).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990).
Kearse, M. et al. Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28, 1647–1649 (2012).
Ratnasingham, S. & Hebert, P. D. BOLD: The Barcode of Life Data System (https://www.barcodinglife.org). Molecular ecology notes 7, 355–364 (2007).
Puillandre, N., Lambert, A., Brouillet, S. & Achaz, G. ABGD, automatic barcode gap discovery for primary species delimitation. Mol. Ecol. 21, 1864–1877 (2012).
Monaghan, M. T. et al. Accelerated species inventory on Madagascar using coalescent-based models of species delineation. Syst. Biol. 58, 298–311 (2009).
Pons, J. et al. Sequence-based species delimitation for the DNA taxonomy of undescribed insects. Syst. Biol. 55, 595–609 (2006).
Kapli, P. et al. Multi-rate Poisson tree processes for single-locus species delimitation under maximum likelihood and Markov chain Monte Carlo. Bioinformatics 33, 1630–1638 (2017).
Fourment, M. & Gibbs, M. J. PATRISTIC: A program for calculating patristic distances and graphically comparing the components of genetic change. BMC Evol. Biol. 6, 1 (2006).
Lefébure, T., Douady, C., Gouy, M. & Gibert, J. Relationship between morphological taxonomy and molecular divergence within Crustacea: Proposal of a molecular threshold to help species delimitation. Mol. Phylogenet. Evol. 40, 435–447 (2006).
Kumar, S., Stecher, G. & Tamura, K. MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 33, 1870–1874 (2016).
Bouckaert, R. R. & Drummond, A. J. bModelTest: Bayesian phylogenetic site model averaging and model comparison. BMC Evol. Biol. 17, 42 (2017).
Xia, X. DAMBE7: New and improved tools for data analysis in molecular biology and evolution. Mol. Biol. Evol. 35, 1550–1552 (2018).
Xia, X., Xie, Z., Salemi, M., Chen, L. & Wang, Y. An index of substitution saturation and its application. Mol. Phylogenet. Evol. 26, 1–7 (2003).
Lanfear, R., Calcott, B., Ho, S. Y. & Guindon, S. PartitionFinder: Combined selection of partitioning schemes and substitution models for phylogenetic analyses. Mol. Biol. Evol. 29, 1695–1701 (2012).
Bouckaert, R. et al. BEAST 2: A software platform for Bayesian evolutionary analysis. PLoS Comput. Biol. 10, e10003537 (2014).
Baele, G., Li, W. L. S., Drummond, A. J., Suchard, M. A. & Lemey, P. Accurate model selection of relaxed molecular clocks in Bayesian phylogenetics. Mol. Biol. Evol. 30, 239–243 (2012).
Drummond, A. J., Ho, S. Y., Phillips, M. J. & Rambaut, A. Relaxed phylogenetics and dating with confidence. PLoS Biol. 4, e88 (2006).
Wysocka, A. et al. A tale of time and depth: Intralacustrine radiation in endemic Gammarus species flock from the ancient Lake Ohrid. Zool. J. Linn. Soc. 167, 345–359 (2013).
Wysocka, A. et al. Origin of the Lake Ohrid gammarid species flock: Ancient local phylogenetic lineage diversification. J. Biogeogr. 41, 1758–1768 (2014).
Cristescu, M. E., Hebert, P. D. & Onciu, T. M. Phylogeography of Ponto-Caspian crustaceans: A benthic–planktonic comparison. Mol. Ecol. 12, 985–996 (2003).
Nahavandi, N., Ketmaier, V., Plath, M. & Tiedemann, R. Diversification of Ponto-Caspian aquatic fauna: Morphology and molecules retrieve congruent evolutionary relationships in Pontogammarus maeoticus (Amphipoda: Pontogammaridae). Mol. Phylogenet. Evol. 69, 1063–1076 (2013).
Macdonald Iii, K. S., Yampolsky, L. & Duffy, J. E. Molecular and morphological evolution of the amphipod radiation of Lake Baikal. Mol. Phylogenet. Evol. 35, 323–343 (2005).
Mats, V., Shcherbakov, D. Y. & Efimova, I. Late Cretaceous-Cenozoic history of the Lake Baikal depression and formation of its unique biodiversity. Stratigr. Geol. Correl. 19, 404 (2011).
Sherbakov, D. Y. On the phylogeny of Lake Baikal amphipods in the light of mitochondrial and nuclear DNA sequence data. Crustaceana 72, 911–919 (1999).
Copilaş-Ciocianu, D., Sidorov, D. & Gontcharov, A. Adrift across tectonic plates: Molecular phylogenetics supports the ancient Laurasian origin of old limnic crangonyctid amphipods. Organ. Divers. Evol. 19, 191–207 (2019).
Sigovini, M., Keppel, E. & Tagliapietra, D. Open Nomenclature in the biodiversity era. Methods Ecol. Evol. 7(10), 1217–1225 (2016).
Ketmaier, V., Argano, R. & Caccone, A. Phylogeography and molecular rates of subterranean aquatic stenasellid isopods with a peri-Tyrrhenian distribution. Mol. Ecol. 12, 547–555 (2003).
Knowlton, N. & Weigt, L. A. New dates and new rates for divergence across the Isthmus of Panama. Proc. R. Soc. Lond. Ser. B Biol. Sci. 265, 2257–2263 (1998).
Papadopoulou, A., Anastasiou, I. & Vogler, A. P. Revisiting the insect mitochondrial molecular clock: The mid-Aegean trench calibration. Mol. Biol. Evol. 27, 1659–1672 (2010).
Hamilton, C. A., Hendrixson, B. E., Brewer, M. S. & Bond, J. E. An evaluation of sampling effects on multiple DNA barcoding methods leads to an integrative approach for delimiting species: A case study of the North American tarantula genus Aphonopelma (Araneae, Mygalomorphae, Theraphosidae). Mol. Phylogenet. Evol. 71, 79–93 (2014).
Fontaneto, D., Flot, J.-F. & Tang, C. Q. Guidelines for DNA taxonomy, with a focus on the meiofauna. Mar. Biodivers.s 45, 433–451 (2015).
Yu, G., Rao, D., Matsui, M. & Yang, J. Coalescent-based delimitation outperforms distance-based methods for delineating less divergent species: The case of Kurixalus odontotarsus species group. Sci. Rep. 7, 1–13 (2017).
De Queiroz, K. Species concepts and species delimitation. Syst. Biol. 56, 879–886 (2007).
Lagrue, C. et al. Confrontation of cryptic diversity and mate discrimination within G ammarus pulex and G ammarus fossarum species complexes. Freshw. Biol. 59, 2555–2570 (2014).
Costa, F., Henzler, C., Lunt, D., Whiteley, N. & Rock, J. Probing marine Gammarus (Amphipoda) taxonomy with DNA barcodes. Syst. Biodivers. 7, 365–379 (2009).
Karaman, G. New data on some gammaridean amphipods (Amphipoda, Gammaridea) from Palearctic. Glasnik Sect. Natl. Sci. Monten Acad. Sci. Arts 15, 20–37 (2003).
Steininger, F. F. & Rögl, F. Paleogeography and palinspastic reconstruction of the Neogene of the Mediterranean and Paratethys. Geol. Soc. Lond. Spec. Publ. 17, 659–668 (1984).
Lefébure, T. et al. Phylogeography of a subterranean amphipod reveals cryptic diversity and dynamic evolution in extreme environments. Mol. Ecol. 15, 1797–1806 (2006).
Sworobowicz, L. et al. Revisiting the phylogeography of Asellus aquaticus in Europe: Insights into cryptic diversity and spatiotemporal diversification. Freshw. Biol. 60, 1824–1840 (2015).
Shih, H. T., Yeo, D. C. & Ng, P. K. The collision of the Indian plate with Asia: Molecular evidence for its impact on the phylogeny of freshwater crabs (Brachyura: Potamidae). J. Biogeogr. 36, 703–719 (2009).
Jesse, R., Grudinski, M., Klaus, S., Streit, B. & Pfenninger, M. Evolution of freshwater crab diversity in the Aegean region (Crustacea: Brachyura: Potamidae). Mol. Phylogenet. Evol. 59, 23–33 (2011).
Szarowska, M., Osikowski, A., Hofman, S. & Falniowski, A. Do diversity patterns of the spring-inhabiting snail Bythinella (Gastropoda, Bythinellidae) on the Aegean Islands reflect geological history?. Hydrobiologia 765, 225–243 (2016).
Trontelj, P., Machino, Y. & Sket, B. Phylogenetic and phylogeographic relationships in the crayfish genus Austropotamobius inferred from mitochondrial COI gene sequences. Mol. Phylogenet. Evol. 34, 212–226 (2005).
Szarowska, M., Hofman, S., Osikowski, A. & Falniowski, A. Divergence preceding island formation among Aegean insular populations of the freshwater snail genus Pseudorientalia (Caenogastropoda: Truncatelloidea). Zoolog. Sci. 31, 680–686 (2014).
Previšić, A., Walton, C., Kučinić, M., Mitrikeski, P. T. & Kerovec, M. Pleistocene divergence of Dinaric Drusus endemics (Trichoptera, Limnephilidae) in multiple microrefugia within the Balkan Peninsula. Mol. Ecol. 18, 634–647 (2009).
Gonçalves, H. et al. Multilocus phylogeography of the common midwife toad, Alytes obstetricans (Anura, Alytidae): Contrasting patterns of lineage diversification and genetic structure in the Iberian refugium. Mol. Phylogenet. Evol. 93, 363–379 (2015).
Rachalewski, M., Banha, F., Grabowski, M. & Anastácio, P. M. Ectozoochory as a possible vector enhancing the spread of an alien amphipod Crangonyx pseudogracilis. Hydrobiologia 717, 109–117 (2013).
Szarowska, M., Hofman, S., Osikowski, A. & Falniowski, A. Daphniola Radoman, 1973 (Caenogastropoda: Truncatelloidea) at east Aegean islands. Folia Malacologica 22 (2014).
Hupało, K. et al. Persistence of phylogeographic footprints helps to understand cryptic diversity detected in two marine amphipods widespread in the Mediterranean basin. Mol. Phylogenet. Evol. 132, 53–66 (2019).
Weiss, M., Macher, J. N., Seefeldt, M. A. & Leese, F. Molecular evidence for further overlooked species within the Gammarus fossarum complex (Crustacea: Amphipoda). Hydrobiologia 721, 165–184 (2014).
Hopkins, L. IUCN and the Mediterranean Islands: Opportunities for biodiversity conservation and sustainable use (Gland, Switzerland, International Union for Conservation of Nature, 2002).
QGIS Development Team. QGIS Geographic Information System. Open source geospatial foundation project. (2016).
Popov, S. V. et al. Lithological-paleogeographic maps of paratethys: 10 maps late eocene to pliocene. Cour. Forsch Senckenberg 250, 1–46 (2004).
Acknowledgements
We would like to thank the colleagues involved in sampling expeditions: Karolina Bącela-Spychalska, Piotr Gadawski, Aleksandra Bańkowska, Anna Wysocka, Grzegorz Michoński, Magdalena Kłosowska, Andrzej Zawal, Agnieszka Szlauer-Łukaszewska. The first author would also like to thank Till-Hendrik Macher for valuable help with data analysis. This work was supported by the Polish National Science Center (projects no. 2014/15/B/NZ8/00266, 2015/17/N/NZ8/01628 and 2018/28/T/NZ8/00022) and partially by the statutory funds of the Department of Invertebrate Zoology and Hydrobiology of the University of Lodz. Kamil Hupało was supported by the German Research Foundation (DFG) (project LE 2323/9-1) and Tomasz Mamos was supported by the Scholarship of the Polish National Agency for Academic Exchange (The Bekker Programme: PPN/BEK/2018/1/00225).
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
M.G. and K.H. conceived and designed the study. I.K., K.H., T.M., M.G. did the field sampling. K.H. and M.G. did the morphological analyses. K.H. performed the laboratory part of the study and generated the dataset. K.H., T.M. and M.G. analysed and interpreted the data. K.H. wrote the manuscript with a substantial contribution from M.G., T.M., I.K.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hupało, K., Karaouzas, I., Mamos, T. et al. Molecular data suggest multiple origins and diversification times of freshwater gammarids on the Aegean archipelago. Sci Rep 10, 19813 (2020). https://doi.org/10.1038/s41598-020-75802-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-75802-2
This article is cited by
-
Multiple genetic species in a halophilic non-marine ostracod (Crustacea)
Hydrobiologia (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.