Abstract
The solid-state photo-chemically induced dynamic nuclear polarization (photo-CIDNP) effect generates non-Boltzmann nuclear spin magnetization, referred to as hyperpolarization, allowing for high gain of sensitivity in nuclear magnetic resonance (NMR). Well known to occur in photosynthetic reaction centers, the effect was also observed in a light-oxygen-voltage (LOV) domain of the blue-light receptor phototropin, in which the functional cysteine was removed to prevent photo-chemical reactions with the cofactor, a flavin mononucleotide (FMN). Upon illumination, the FMN abstracts an electron from a tryptophan to form a transient spin-correlated radical pair (SCRP) generating the photo-CIDNP effect. Here, we report on designed molecular spin-machines producing nuclear hyperpolarization upon illumination: a LOV domain of aureochrome1a from Phaeodactylum tricornutum, and a LOV domain named 4511 from Methylobacterium radiotolerans (Mr4511) which lacks an otherwise conserved tryptophan in its wild-type form. Insertion of the tryptophan at canonical and novel positions in Mr4511 yields photo-CIDNP effects observed by 15N and 1H liquid-state high-resolution NMR with a characteristic magnetic-field dependence indicating an involvement of anisotropic magnetic interactions and a slow-motion regime in the transient paramagnetic state. The heuristic biomimetic design opens new categories of experiments to analyze and apply the photo-CIDNP effect.
Similar content being viewed by others
Introduction
The solid-state photo-CIDNP (photo-chemically induced dynamic nuclear polarization) effect1,2,3 allows to enhance nuclear magnetic resonance (NMR) signals by a build-up of transient nuclear spin-hyperpolarization caused by light-induced transient spin-correlated radical pairs (SCRPs) in electron-transfer proteins4,5. The effect was discovered by Zysmilich and McDermott in 1994 by performing 15N magic-angle spinning (MAS) NMR studies on an isolated, frozen and quinone-blocked photosynthetic reaction center protein of the purple bacterium Rhodobacter sphaeroides6. Since its discovery, the effect has been observed in plenty of other photosynthetic reaction centers (RCs) of plants7,8,9,10, algae11,12, diatoms13,14, purple bacteria15,16,17,18, heliobacteria19,20, and green sulfur bacteria21. The spin-chemical machinery pumping nuclear polarization has been probed by field-dependent22,23, time-resolved24,25,26, and preparation-dependent27,28 experiments, and is interpreted by the occurrence of up to three mechanisms running in parallel22,24,29: In the Differential Relaxation (DR) mechanism, also called “cyclic reactions mechanism”, the symmetry between both decay branches is broken due to a difference in relaxation caused by the paramagnetic molecular triplet state30,31,32. In the electron–electron nuclear Three-Spin Mixing (TSM) mechanism, symmetry breaking is driven by the pseudosecular part of the hyperfine interaction during the evolution of the SCRP33,34. The differences in decay kinetics between the SCRP in its singlet and its triplet state cause the Differential Decay (DD) mechanism35. Recently, the coherent electron–electron–nuclear spin-dynamics has been described in terms of a unified theoretical approach, based on level crossings (LCs) and level anti-crossings (LACs)36,37,38, which allowed the establishment of CIDNP sign rules and to determine conditions optimal for polarization formation.
Besides photosynthetic RCs, there is another class of proteins acting as molecular spin-machines pumping non-Boltzmann nuclear spin-state upon illumination: biological photoreceptors that contain a flavin cofactor as chromophore with characteristic absorption maxima in the UV-A and blue light regions. Merely one family of flavoproteins, phototropins, was reported to show the photo-CIDNP effect39,40,41. Generally, phototropins are blue-light receptors that harbor two light-sensing light-oxygen-voltage (LOV) domains, each of which incorporates one flavin mononucleotide (FMN) cofactor non-covalently. Upon excitation, the FMN in a wild-type LOV domain of phototropin undergoes adduct formation with a close-by conserved cysteine42,43. However, so far, no photo-CIDNP effect was observed from a wild-type LOV domain. The first successful solid-state photo-CIDNP effect observation was demonstrated for a frozen cysteine-lacking LOV domain of phototropin from the green alga Chlamydomonas reinhardtii (Crphot-LOV1-C57S)41,44. In this case, the photo-excited FMN undergoes inter-system crossing to the triplet state, causes intra-protein electron transfer from a tryptophan to the FMN and thus gives rise to formation of a transient SCRP. Spin evolution of this SCRP allows for the build-up of the solid-state photo-CIDNP selectively on FMN and tryptophan. In Crphot-LOV1-C57S, the edge-to-edge distance (\(r_{FW}\)) between FMN (F) and tryptophan (W) is ~ 11 Å. Remarkably, Weber et al. first observed photo-CIDNP in a similar LOV protein, the LOV2 domain of phototropin 1 from oat, Avena sativa (Asphot1-LOV2-C450A), by 13C liquid-state NMR40,45,46. The crystal structure of Asphot1-LOV2-C450A is almost identical to that of Crphot-LOV1-C57S, given that they share 47% of amino acid sequences. The \(r_{FW}\) value is very similar for the two proteins. This is a very rare example that a nuclear-spin hyperpolarization effect generated in one system could be observed by both liquid-state and solid-state NMR. To explore the action of anisotropic mechanisms in the liquid sample that are expected to be sensitive to the magnetic field strength44,46, a solution-state NMR spectrometer equipped with a field-cycling device47 was employed to study 1H, 13C and 15N photo-CIDNP generated in Crphot-LOV1-C57S within a broad field range from 0.01–9.4 T39.
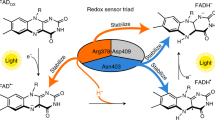
The LOV domain is the blue-light mediating motif of not only phototropin but also of some other proteins, e.g., aureochromes48. Aureochromes represent a class of LOV proteins with an unusual inversed domain arrangement, as they carry, in contrast to the majority of LOV-domain proteins, the functional/signaling domain N-terminally to the LOV domain and not C-terminally; they exhibit great potential in controlling DNA binding49 as a natural optogenetic module. Despite a different domain organization of the full-length protein, the LOV domain of aureochromes seem to show a high degree of conservation to canonical LOV domains with respect to sequence and secondary structure50. Like in other LOV domains, the optical excitation of FMN chromophore leads to formation of an adduct with a conserved cysteine residue. Furthermore, a recently discovered flavoprotein from the radiation-resistant bacterium Methylobacterium radiotolerans, Mr4511, unusually lacks the single tryptophan conserved in 75% of LOV domains51. Due to the absence of tryptophan to quench the triplet state of FMN, 3FMN, after excitation by light, the cysteine-lacking mutant of Mr4511 can serve as an efficient singlet-oxygen generator51. The variety of LOV domains raises the question whether LOV domains can be generally constructed in such a way that they show a photo-CIDNP effect.
The aim of this work was to extend the range of biological systems amenable to the solid-state photo-CIDNP effect. To reach this goal, we at first produced a cysteine-lacking LOV domain of aureochrome1a from the diatom Phaeodactylum tricornutum (Ptaureo1a-LOV-C287S)52,53 and show that it generated 1H-, 13C-, and 15N photo-CIDNP effects in aqueous solution. By using a field-cycling system, the magnetic field-dependencies of the 1H, 13C, and 15N hyperpolarization effects have been obtained revealing that the magnetic field for maximal photo-CIDNP depends on the nuclear gyromagnetic ratio. Such a behavior is a characteristic feature of the solid-state photo-CIDNP effect36,39, closely resembling that of Crphot-LOV1-C57S. Based on this result, we propose that, like in phototropin, the anisotropic magnetic interactions might play a decisive role in photo-CIDNP formation in the LOV domain of aureochrome in solution.
Furthermore, in the heuristic approach, biomimetic protein design is used to control the conditions for the occurrence of the photo-CIDNP effect. We employed Mr4511 for an extended protein mutation strategy allowing us to change parameters such as the distance between the partners governing the electron transfer reactions that give rise to SCRP formation and recombination, and to tune magnetic parameters of the SCRP. By doing so, we were able to probe the spin dynamics in the SCRP by magnetic-field dependent photo-CIDNP studies. We could also elucidate the role of different photo-CIDNP mechanisms that are responsible for the formation of nuclear spin-polarization. Last but not least, we discovered a non-tryptophan-induced photo-CIDNP effect generated by the cysteine-devoid Mr4511, in which tryptophan is absent.
Results
To rationalize the key properties of molecular spin-machines that can be used to generate photo-CIDNP, we proposed a design strategy based on mutations, supported by field-dependent CIDNP studies. By using various mutations, as described below, we were able to vary the distance between the electron donor and acceptor. In this way, we affected the rate of SCRP formation and recombination, and also varied the electron–electron spin–spin interaction in the SCRP. To probe the reaction and spin dynamics in the SCRP, we used the field dependence of photo-CIDNP.
Screening LOV domains for induction of photo-CIDNP
Aiming for designed molecular spin-machines producing light-induced nuclear hyperpolarization, we have designed a series of protein mutants, which will be presented in parts (i)—(iii) (Table 1).
-
(i)
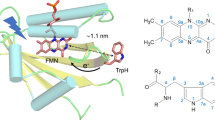
So far, the occurrence of the solid-state photo-CIDNP effect was limited to cysteine-lacking LOV domains of phototropin39,40,41; for this reason, here we explored other potential LOV-based light-induced hyperpolarization generators. Alignment and comparison of the amino-acid sequences of Asphot1-LOV2, Crphot-LOV1 and Ptaureo1a-LOV show about 50% of identity (Fig. 1A) and the crystal structures show almost identical tertiary structures (Fig. 1B). In particular, the distance between FMN and tryptophan (Fig. 1C), which determines the strength of the spin–spin coupling in the SCRP and is therefore central to generate photo-CIDNP, is nearly the same, being approximately 11 Å. Therefore, we used the mutant Ptaureo1a-LOV-C287S (\(r_{FW}\) ~ 11 Å) for the liquid state photo-CIDNP NMR experiment, in which the conserved cysteine 287 is replaced by serine.
Figure 1 (A) Alignments of the amino-acid sequence of wild-type phototropin 1-LOV2 from Avena sativa (Asphot1-LOV2), phototropin-LOV1 from Chlamydomonas reinhardtii (Crphot-LOV1), aureochrome1a-LOV from Phaeodactylum tricornutum (Ptaureo1a-LOV), and 4511 from Methylobacterium radiotolerans (Mr4511): segments highlighted in green and blue refer to α-helices, and β-sheets, respectively, indicating the secondary structure of the proteins. The conserved positions of cysteine located in Eα and tryptophan in Hβ in the LOV domains are typeset in orange. The five positions to introduce tryptophan in Mr4511 via mutagenesis are highlighted in red. (B) Alignment of the crystal structures of Crphot-LOV1 (PDB: 1N9L, green)56, Ptaureo1a-LOV (PDB: 5A8B, red)57, and the simulated structure of Mr4511 (yellow) without FMN. The simulation is performed with SWISS-MODEL based on the crystal structure of aureochrome1a-LOV from Vaucheria frigida (PDB entry: 3UE6)58. The information about structural modeling is listed in Supplementary Information Table S1. Five mutants of cysteine-lacking Mr4511 were generated, one with tryptophan placed at the canonical position, Q112W, \(r_{FW}\) ~ 11 Å, the other four at non-canonical positions F130W, Y116W, Y129W and K57W with increasing \(r_{FW}\). The figure was created by the PyMOL Molecular Graphics System, Version 1.2r3pre, Schrödinger, LLC. (C) IUPAC numbering of atomic positions in the isoalloxazine ring of FMN and the side chain of tryptophan.
-
(ii)
Formation of an SCRP by electron transfer to excited FMN can occur if a nearby tryptophan is present to act as the electron donor. It has been shown that the amino acid tryptophan is able to provide this function41,54. Therefore, in the LOV protein Mr4511, lacking the conserved tryptophan, we introduced a tryptophan at its canonical position Q112 by mutation resulting in the Mr4511-C71S-Q112W double mutant. Previously, transient absorption experiments have been used to test the function of these mutants. In Mr4511, when the cysteine residue was mutated to serine (C71S) or glycine (C71Q) and no tryptophan was present, the lifetime of 3FMN, \(\tau_{T}\), was around 240 μs51. Introduction of tryptophan to the canonical position, Mr4511-C71S-Q112W, gave rise to faster quenching of 3FMN reducing \(\tau_{T}\) to ~ 24 μs51, a value very close to ~ 27 μs observed in Crphot-LOV1-C57S (\(r_{FW}\) ~ 11 Å)43. Hence, the double mutant Mr4511-C71S-Q112W (\(r_{FW}\) ~ 11 Å) is the second candidate for the generation of the solid-state photo-CIDNP effect.
-
(iii)
Finally, we introduced the electron-donating tryptophan at non-canonical positions. The introduction of tryptophan to a new position of the protein allows to change the distance between FMN and tryptophan, their relative orientation and chemical environments and, therefore, to affect the kinetic and magnetic parameters, critical for the formation of the solid-state photo-CIDNP effect. It is difficult to fine-tune all relevant reaction and magnetic resonance parameters simultaneously, therefore, we focused on creating mutants with different \(r_{FW}\). Lacking a crystal structure of Mr4511, the design relied on a structural model created using SWISS-MODEL and the crystal structure of aureochrome1a-LOV (PDB: 3UE6) from a eukaryotic photosynthetic stramenopile as template58. The report of the modeling parameters is provided in Supplementary Information Table S1. Additionally, comparison of the amino-acid sequence (Fig. 1A) allowed us to predict the occurrence of α-helix and β-sheet secondary structures and to reconstruct the tertiary structure of Mr4511. As targets for mutation, we considered amino acids that do not interact with the FMN50 and also have a bulky side chain similar as tryptophan. Using these ideas, we have designed the following mutants with different positions of tryptophan with various \(r_{FW}\) values: Mr4511-C71S-F130W (~ 6 Å), Mr4511-C71S-Y116W (~ 9 Å), Mr4511-C71S-Y129W (~ 11 Å) and Mr4511-C71S-K57W (~ 17 Å) (Table 1).
Another aspect relevant for rational design of a biomimetic light-driven spin-machine for production of photo-CIDNP is the possibility to introduce isotope labels. In particular, for the measurement of the 15N photo-CIDNP we employed 15NH4Cl as the sole nitrogen source in the bacterial growth medium during protein expression and produced uniformly 15N-labelled protein and cofactor (see “Methods”). For the 13C photo-CIDNP experiment, different labelling strategies were previously applied, either by incorporating the 13C-labelled FMN into a natural abundant protein moiety of the phototropin-LOV domain40,46 or by selective 13C-labelling of the single tryptophan of the phototropin-LOV domain39. This enables unambiguous assignment of hyperpolarized 13C signals and analysis of the photo-CIDNP effect generated by electron donor and acceptor separately. A complete picture of the effect, however, involving both electron donor and acceptor is still missing. Therefore, in the present work we produced a uniformly 13C-labelled Crphot-LOV1-C57S (\(r_{FW}\) ~ 11 Å), aiming to compare the photo-CIDNP effect of FMN and tryptophan under the same conditions. The hyperpolarization effect in combination with isotope labelling paves the way to field-dependent NMR measurements, providing knowledge of the relationship between enhancement factor and magnetic field, which might provide the key data for future theoretical analysis of the exact photo-CIDNP mechanism.
Comparison of the photo-CIDNP effect between phototropin and aureochrome
Figure 2A shows the 1H photo-CIDNP effect and its field dependence observed in Crphot-LOV1-C57S (\(r_{FW}\) ~ 11 Å). The protein produced for the experiment initially contained all the nuclei in their natural abundance. Then the protonated buffer of the sample was exchanged to a deuterated buffer (see “Methods” section). The final protein solution may contain ~ 0.4% residual 1H. From this sample, the effect has not been observed directly on the protons of FMN and tryptophan, however, the light-minus-dark spectra show a negative enhancement (emissive signal, i.e., opposite to the thermal polarization) of the HDO signal at 4.7 ppm, which agrees with the previous publication39. A closer look at the light-minus-dark spectra shows that the entire range (– 2 to 10 ppm) in the proton NMR spectra exhibits hyperpolarization, in particular the aliphatic region (0 to 2.5 ppm) as well as the HDO signal. Integrating either the HDO signal or the range of 0 to 2.5 ppm or the range of – 2 to 10 ppm, we obtain a field dependence with a maximum at 0.6 T, as shown in Fig. 3A. Since the position of the maximum is the same for all protons, it is plausible that the 1H hyperpolarization has originated from the same SCRP and has been distributed over the whole protein as well as to the residual protons in the deuterated solvent. We assume that the spread of hyperpolarization under liquid-state conditions is due to cross relaxation or due to spin diffusion mediated by non-averaged proton-proton dipolar couplings59,60.
(A) 1H, (B) 13C and (C) 15N photo-CIDNP spectra of Crphot-LOV1-C57S (~ 11 Å) detected at 289 K at different magnetic field strengths by using a 9.4 T liquid-state NMR spectrometer equipped with a field-cycling device. Hereafter, all photo-CIDNP spectra are light-minus-dark NMR spectra. The upper trace includes signal assignments (blue) with 13C chemical shifts of hyperpolarized signals mentioned in Supplementary Information Table S2 and 15N chemical shifts in Table 2. The lower trace shows stacked spectra of the CIDNP effects of the corresponding nuclei measured at five different magnetic fields.
Intensity of selected hyperpolarized signals of Crphot-LOV1-C57S (\(r_{FW}\) ~ 11 Å), integrated and plotted against the magnetic field strength for (A) 1H, (B) 13C and (C) 15N. For convenient comparison, the magnitude of the maximum polarization of each nucleus is arbitrarily set to − 1 and the signal enhancement is normalized to this value. The plus and minus signs at the y-axis represent the enhanced absorptive (positive) and emissive polarization (negative) relative to the thermal polarization. Uniformly 13C- and uniformly 15N-enriched Crphot-LOV1-C57S (~ 11 Å) samples were employed for the 13C and 15N photo-CIDNP NMR experiments, respectively. Regarding 1H photo-CIDNP, the signal of HDO, the aliphatic region (0 to 3 ppm), and the broader region (− 2 to 10 ppm) of the 1H photo-CIDNP spectrum were integrated for comparison.
13C photo-CIDNP of a uniformly 13C-labelled Crphot-LOV1-C57S (\(r_{FW}\) ~ 11 Å) induced at various field strengths ranging from 0.5 to 9.4 T and detected always at 9.4 T is shown in Fig. 2B. The hyperpolarized 13C signals are tentatively assigned according to previous studies39,46,59,61, and are summarized in Table S2 in Supplementary Information. Figure 3B presents the integrated areas of some selected hyperpolarized carbon signals of FMN and tryptophan against the magnetic field strength. Overall, the carbons on the tryptophan indole ring seem to be stronger polarized than those on the isoalloxazine ring of FMN. The hyperpolarized carbon nuclei of tryptophan show emissive polarization at all fields and maximal polarization at around 3 T in agreement with previous results39. The selected 13C atoms of the isoalloxazine ring of FMN show different enhancement patterns. As the magnetic field increases, the signal of, e.g., FMN C-8 stays negative, while FMN C-2 changes the sign of polarization from emission to enhanced absorption. Due to the difference of labelling strategies, the comparison of the 13C photo-CIDNP spectrum of Crphot-LOV1-C57S (\(r_{FW}\) ~ 11 Å) with a previously published 13C photo-CIDNP spectrum of Asphot1-LOV2-C450A (\(r_{FW}\) ~ 11 Å)40,46 is not straightforward. Despite that, we can compare the signal of Trp C-3 carbon in both spectra. In Asphot1-LOV2-C450A (\(r_{FW}\) ~ 11 Å) where the tryptophan nucleus is at natural abundance, strong hyperpolarization at Trp C-3 occurs. The chemical shift of C-3 of Crphot-LOV1-C57S (109.0 ppm) slightly differs from δ (C-3) = 113.5 ppm for Asphot1-LOV2-C450A which might result from the different protein environment of the tryptophan residues. In both cases, the sign of the signal of Trp C-3 is always emissive at all fields studied. Nevertheless, the field at which the maximum polarization at Trp C-3 occurs, \(B_{max}\) = 3 T for Crphot-LOV1-C57S and \(B_{max}\) = 7 T for Asphot1-LOV2-C450A, is well distinguished. We assume that the difference in the label positions causes the difference in field dependence although details are not yet understood. Furthermore, in a previous work by Kothe et al.46 the photo-CIDNP effect has been measured at four different magnetic fields (5.9, 7.1, 9.4, and 11.8 T), limiting comparability of data points and localization of the maximum \(B_{max}\), while in the present work, we overcame this problem by using a shuttle system (10 nT < Bo < 9.4 T).
Figures 2C and 3C depict the field dependence of the 15N photo-CIDNP obtained in uniformly 15N-labelled Crphot-LOV1-C57S (\(r_{FW}\) ~ 11 Å) at 11 magnetic field strengths ranging from 0.1 to 9.4 T. In Fig. 2C, the three hyperpolarized signals and their field dependencies are shown (for assignment, see Table 2). The Trp N-1 CIDNP is a low absorptive polarization at low fields, turning into strong emissive polarization at about 1 T field and having a maximum at around 5 T. In contrast, the photo-CIDNP of FMN N-5 is always absorptive with a maximum at 3 T. At increasing field, the hyperpolarization of FMN N-10 turns from enhanced absorption to emission at around 3 T, similar to the photo-CIDNP of Trp N-1 and in clear contrast to the photo-CIDNP of FMN N-5. A common feature for these three nuclei is that the high-field maximum occurs at around 4 T.
For comparison of the photo-CIDNP field dependencies for the three different NMR active nuclear isotopes, we have chosen the most enhanced signal or spectral region for each nucleus, 1H, 13C and 15N: Trp C-3, Trp N-1 and the 1H spectrum intensity integrated from – 2 to 10 ppm of Crphot-LOV1-C57S (\(r_{FW}\) ~ 11 Å) (Fig. 4A). The field at which the 1H, 13C and 15N photo-CIDNP production reached its maximum follows the relationship: 0.6 T (1H) < 3 T (13C) < 5 T (15N). Under similar conditions, the field maxima for photo-CIDNP enhancement of the corresponding nuclei in Ptaureo1a-LOV-C287S (\(r_{FW}\) ~ 11 Å) (tryptophan is selectively isotope-labelled) were obtained. The positions of the maxima are in the same order (Fig. 4B); the corresponding 1D spectra are shown in Figure S1. Hence, a high similarity in the field dependence is observed for the two proteins. All three types of nuclei in both samples exhibit emissive NMR signals at the maxima; the positions of maxima depend on the nuclear gyromagnetic ratios (the maximum is positioned at a lower field for a nucleus with a higher gyromagnetic ratio). In both cases, the 1H hyperpolarization reaches its emissive maximum at a field around 0.6 T, while the maxima of 13C and 15N photo-CIDNP appear to be at higher fields. Although the primary sequence of the two proteins does not have a high identity, their tertiary structures (Fig. 1B) and, as shown here, their functional mechanisms are almost identical. Therefore, the production of the photo-CIDNP does not depend crucially on the primary sequence of individual amino acids, whereas the distance between the two redox partners forming the SCRP and their mutual orientation plays of a decisive role.
Comparison of the field dependencies of the 1H and 13C and 15N photo-CIDNP effects generated by (A) Crphot-LOV1-C57S (\(r_{FW}\) ~ 11 Å) and (B) Ptaureo1a-LOV-C287S (\(r_{FW}\) ~ 11 Å). Detailed experimental parameters are provided in the “Methods” section. The hyperpolarized 1H signal has been obtained by integrating the spectral region of − 2 to 10 ppm (black square); the 13C signal from C-3 of tryptophan (red triangle) and the 15N signal from indole of tryptophan (blue circle) are also shown. The dashed lines are added to guide the eye. The maximum emissive polarization of all nuclei is scaled to − 1. The minus sign of the y-axis implies that the hyperpolarization is emissive. The 1D spectra of the photo-CIDNP effect observed in Ptaureo1a-LOV-C287S are shown in Supporting Information Figure S1.
Tryptophan and non-tryptophan derived photo-CIDNP effect generated in Mr4511 mutants
To generate a photo-CIDNP effect in Mr4511, the conserved cysteine was first replaced with the inactive serine resulting in the mutant Mr4511-C71S. Furthermore, tryptophan was introduced to different locations of the Mr4511-C71S protein, generating five additional mutants (Table 1). The magnetic field dependence of the photo-CIDNP effect under liquid-state conditions obtained by 15N and 1H NMR is summarized in Fig. 5, plotted in the same way as in Fig. 3. The corresponding 15N NMR spectra are shown in Figure S2 and the 15N chemical shifts are provided in Table 2.
Magnetic-field dependencies of the 1H (left column) and 15N (right column) photo-CIDNP effect generated in six Mr4511-C71S mutants having different \(r_{FW}\): (A) Mr4511-C71S-F130W (\(r_{FW}\) ~ 6 Å), (B) Mr4511-C71S-Y116W (\(r_{FW}\) ~ 9 Å), (C) Mr4511-C71S-Q112W (\(r_{FW}\) ~ 11 Å), (D) Mr4511-C71S-Y129W (\(r_{FW}\) ~ 11 Å), (E) Mr4511-C71S-K57W (\(r_{FW}\) ~ 17 Å) and (F) Mr4511-C71S. Regarding the 1H photo-CIDNP, the region of chemical shifts ranging from − 2 to 10 ppm of the spectrum is integrated and plotted against the magnetic field (A1–F1). The three 15N photo-CIDNP hyperpolarized signals correspond to FMN N-5 (green), FMN N-10 (yellow) and Trp N-1 (blue); the signals are plotted against the magnetic field strength (A2–F2) in the same range and scale as for the 1H signal. For straightforward comparison, the signal strength at the maximal hyperpolarization magnitude of each graph is normalized to unity. The positive and negative signs of the y-axis indicate absorptive and emissive hyperpolarization, respectively. Since the mutant F130W shows no detectable photo-CIDNP effect, as presented in the Supplementary Information, Figures S3 and S4, the data points shown in (A1) and (A2) were measured the over the same spectra regions with the same noise level as in the corresponding spectra for Y129W shown in (D1) and (D2).
The mutant Mr4511-C71S-F130W (\(r_{FW}\) ~ 6 Å), which has the shortest distance between the redox partners of the SCRP, shows neither a 1H nor a 15N photo-CIDNP in the magnetic field range from 0.1 to 9.4 T (Fig. 5A1,A2; Supplementary Information Figure S3 and S4). Time-resolved optical absorption analysis on F130W suggests that the triplet state 3FMN is not formed to an observable extent in this protein, most likely, because of the ultra-fast electron transfer between FMN and tryptophan (Aba Losi, personal communication). Accordingly, for the construction of light-induced artificial spin-machines pumping nuclear hyperpolarization, the information of a minimum distance of the redox partners of the SCRP is highly relevant.
Except for Mr4511-C71S-F130W (\(r_{FW}\) ~ 6 Å), all other Mr4511-C71S mutants generated both, the 15N and 1H photo-CIDNP effect under liquid-state conditions. As shown in Fig. 5B2, Mr4511-C71S-Y116W (\(r_{FW}\) ~ 9 Å) shows hyperpolarization for the nitrogen on FMN and tryptophan, therefore it is referred to as “tryptophan-derived photo-CIDNP”. This protein shows a maximal 1H hyperpolarization at 2.4 T (Fig. 5B1), which is a higher field than observed for any other mutant (see below).
The \(r_{FW}\) value of ~ 11 Å in the phototropin and aureochrome LOV domains was known to generate a photo-CIDNP effect40,41. Here we compare the two cases when the tryptophan is at the canonical position, Mr4511-C71S-Q112W (\(r_{FW}\) ~ 11 Å) (Fig. 5C1,C2) and the non-canonical position but with nearly the same \(r_{FW}\) distance, Mr4511-C71S-Y129W (\(r_{FW}\) ~ 11 Å) (Fig. 5D1, D2). Although Mr4511-C71S-Q112W and Mr4511-C71S-Y129W exhibit a similar field maximum for the 1H photo-CIDNP effect, their field-dependent 15N photo-CIDNP effects are significantly different. This means that the efficiency of photo-CIDNP formation does not only depend on the spatial distance. Different orientations and different local mobility of the residues might be considered as the origin of this difference. However, the phase of hyperpolarized Trp N-1 signal is allways negative for mutants Y116W, Y129W and Q112W.
The tryptophan residue in Mr4511-C71S-K57W (\(r_{FW}\) ~ 17 Å) is the most remote electron donor from FMN in the studied set of mutants and, thus, the reaction rate constant of electron transfer in this protein mutant is expected to be the smallest in this series62. The protein shows a 15N hyperpolarization solely on the FMN N-5, while the FMN N-10 and Trp N-1 signals do not exhibit any enhancement in a wide range of magnetic fields (Fig. 5E). The same photo-CIDNP experiments were also performed on Mr4511-C71S in which no tryptophan was present leading to very similar results (Fig. 5F) with somewhat weaker single emissive FMN N5 signal (Figure S2 graphs D and E). This surprising result clearly indicates that the 15N photo-CIDNP effect reported here is not derived solely from the involvement of the added tryptophan residue. It is well-known that tyrosine can also act as an electron donor in biological systems63,64. According to the amino-acid sequence and our structural model of Mr4511, there are four tyrosine residues located in proximity to FMN in the range of 9 to 12 Å. In line with this speculation is the fact that the 15N photo-CIDNP effect is not observable for tyrosine, since tyrosine does not contain 15N in the side chain. Therefore, the light-driven molecular spin-machines can probably also rely on SCRPs containing a tyrosine radical.
Discussion
Here, we show that a photo-CIDNP effect originating from the SCRP of FMN and tryptophan can be produced in artificially designed flavoproteins. We employ a systematic mutation strategy to vary reaction and magnetic parameters of the paramagnetic centers generated by light. It appears that the \(r_{FW}\) distance of ~ 6 Å between the FMN and tryptophan is too short to provide conditions suitable for photo-CIDNP formation, whereas in the range of ~ 9 Å to ~ 11 Å, the effect has been observed (Fig. 5; Table 2). The data on the field dependence of the photo-CIDNP effect generated by the designed LOV domains show complex dependencies, which are not expected for the liquid-state photo-CIDNP effect. In particular, the differences in field dependence obtained for LOV domains having the same distance of donor and acceptor suggest that anisotropic spin interactions come into play as they are expected for solids. In addition to the field dependence, a distance dependence has been documented. Apparently, further parameters are involved, presumably the relative orientation of donor and acceptor as well as their local dynamics. Both, anisotropy and relaxation effects require further studies. Furthermore, the effect of different label patterns requires a future study.
The presence of solid-state mechanisms in LOV domains in liquids implies that the transient SCRP occurs in a slow-motion regime, during which the anisotropic electron–nuclear interactions are conserved for the build-up of hyperpolarization. In contrast, on NMR time scale, all the anisotropic nuclear interactions, i.e., nuclear dipolar coupling and chemical shift anisotropy as present in solids are averaged out and thus the hyperpolarized signals in the NMR spectra exhibit no obvious anisotropic features. Such phenomenon was previously observed for a photosynthetic RC protein-membrane complex corresponding to ~ 1 MDa, measured by 13C liquid-state NMR28. For LOV proteins, having the molecular weight of less than 20 kDa, the occurrence of anisotropic mechanisms in liquids likely relies on the formation of dimers53,57,65 or higher multimers in solution. The slow tumbling rate may lead to the presence of residual proton-proton couplings which allow for the 1H hyperpolarization transfer from the center of the photo-reaction into the environment (Fig. 2A).
So far, flavoproteins and photosynthetic RCs are the only reported electron transfer protein systems that show solid-state photo-CIDNP effect. Even despite the different cofactor arrangements and spin-dynamics, they might share the same mechanisms. Consequently, similar features of CIDNP are expected regarding the sign change of nuclear spin hyperpolarization and the similar field at which the maximum polarization occurs22,23. LCs and LACs analysis suggested that a solid-state photo-CIDNP effect is not only field-dependent, but also strongly orientation-dependent because of the anisotropic interactions governing in spin dynamics of the SCRP in solids36,37. To the present experiments conducted under liquid-state conditions, the same theory will be applied to understand the sign change that occurred in the LOV proteins as shown in Fig. 3B,C as well as Fig. 5B2–D2.
Summarizing these considerations, we can propose the following interpretation of the experimental findings.
By increasing \(r_{FW}\), we decrease two parameters: The SCRP recombination rate and the electron–electron spin–spin coupling, \(J_{SCRP}\), within the SCRP. When the \(r_{FW}\) distance is too short, photo-CIDNP formation is suppressed, most probably, due to two reasons: The first reason is that \(J_{SCRP}\) is too large, introducing an energy gap between the singlet and triplet SCRP spin states. This gap cannot be overcome by the relatively small hyperfine couplings, and singlet–triplet interconversion in the SCRP is thus suppressed. The second reason is that the spin-evolution of the SCRP requires sufficient time for photo-CIDNP formation: fast SCRP recombination interrupts this process and thus no photo-CIDNP is formed.
As \(r_{FW}\) increases further, we enter the regime in which the SCRP lifetime is sufficiently long and \(J_{SCRP}\) is sizeable, but not too large to suppress singlet–triplet mixing, giving rise to photo-CIDNP formation. In this situation, the TSM scenario comes into play and the maximum position, \(B_{max}\), in the photo-CIDNP field dependence is given37,39 by the matching condition \(\left| {\gamma_{N} } \right|B_{max} \approx J_{SCRP}\), with \(\gamma_{N}\) being the nuclear gyromagnetic ratio; the sign of polarization of the three different kinds of nuclei (1H, 13C and 15N) is also consistent with previous theoretical considerations37,39. Hence, the \(B_{max}\) field is different for different nuclei, which is consistent with the experimental data.
When \(r_{FW}\) increases further, \(J_{SCRP}\) is decreased and other photo-CIDNP mechanisms37 come into play. In this situation, polarization formation is no longer sensitive to the \(\gamma_{N}\) value, i.e., to the nuclear spin isotopes, so that different kinds of nuclei exhibit a similar behavior. The sign changes of photo-CIDNP can be rationalized in terms of changing dominance of enhancement mechanisms, as it happens in RCs37.
The design strategy also leads to the discovery of a new-type of photo-CIDNP effect generated by Mr4511-C71S in which no tryptophan is present. The same effect (Fig. 5E,F) also occurs in Mr4511-C71S-K57W (\(r_{FW}\) ~ 17 Å). Based on the present results, we are unable to unravel the origin of the new-type photo-CIDNP effect. Recent research on a designed cysteine-lacking Asphot1-LOV2 domain indicated that, without presence of the tryptophan, the FMN was reduced, however to less extend compared to the case when the tryptophan was present. Kinetic data suggested that one of the tyrosines in the LOV domain acts as counter radical66. Therefore, we proposed that tyrosine might act as electron donor in the absence of tryptophan also in our case.
With this, the present work significantly extends the class of light-driven molecular spin machines, which pump nuclear spin-hyperpolarization. The LOV systems are particularly suitable for such biomimetic design, while photosynthetic RCs due to their structural complexity allow for limited manipulations only. The biomimetic design that affects the parameters of the photo-CIDNP effect provides new categories of experiments to analyze the conditions for its occurrence.
Methods
Protein preparation
The plasmid (i) encoding the LOV-C287S mutant of aureochrome1a from P. tricornutum comprising the flanking Jα and A’α helices (238–378) was received from Peter Kroth (University of Konstanz)53. The plasmid (ii) that encodes Mr4511 from M. radiotolerans (1–164) was generated in our own group by genome cloning51. On that basis, we first constructed the cysteine-lacking Mr4511-C71S mutant. Subsequently, additional five mutants encoding tryptophan situated at different positions were created, Mr4511-C71S-F130W (~ 6 Å), Mr4511-C71S-Y116W (~ 9 Å), Mr4511-C71S-Q112W (~ 11 Å), Mr4511-C71S-Y129W (~ 11 Å) and Mr4511-C71S-K57W (~ 17 Å) with primers shown in the Supporting Information Table S3. All genetic manipulations were according to standard protocols. Plasmid (iii) encodes the LOV1-C57S mutant of phototropin from C. reinhardtii (16–133) carrying a 15 × His-tag at the N-terminus67. Further information about all the mutants employed in this work is summarized in Table 1. The protocol of heterologous overexpression and isotope-labelling of all the mutants in Escherichia coli has been reported elsewhere39. Isotopically enriched material, 15NH4Cl, [15N] indole, [u-13C6] glucose, and [u-13C8] indole employed in this research were purchased from Cambridge Isotope Laboratories, Inc. (Andover, MA, USA). Use of 15NH4Cl or [u-13C6] glucose as the sole source in the growth medium yields a uniformly 15N or 13C labelled protein, while supplementation of 13C or 15N isotope-enriched indole as precursor to the normal medium results in a selective labelling of the tryptophan side chain. The 15N and 13C labelled proteins were used for corresponding 15N and 13C NMR measurements. For 1H NMR measurement, the employed proteins are in their natural abundance, and they were washed with deuterated phosphate buffer (300 mM NaCl, 50 mM KsPO4 in D2O, pH 8.0) to a final solution containing approximately 0.4% residual protons. The final concentration of the flavoproteins were controlled at about 16 μM (\({\upvarepsilon }_{{450{\text{nm}}}}\) = 12,500 M−1 cm−1) before photo-CIDNP measurement.
Photo-CIDNP solution-state NMR
The field-dependent 15N-, 13C-, and 1H photo-CIDNP experiments of the LOV proteins were carried out on an NMR spectrometer operating at a magnetic field of 9.4 T (1H frequency of 400 MHz) (Bruker Avance III HD) equipped with a home-built field-cycling device47. It transfers the sample to variable magnetic fields within the range 10 nT < Bo < 9.4 T at which the sample is illuminated and returns it for the NMR detection at 9.4 T. For the 13C and 15N photo-CIDNP experiments, pulse-acquire with WALTZ-16 proton decoupling was employed. The pulse sequence of the 1H photo-CIDNP experiments starts with a pre-saturation composite pulse sequence68 at 9.4 T, followed by the sample shuttle cycle that includes the sample transfer to the chosen magnetic field for illumination by LED (called “light”) or the same cycle without illumination (called “dark”) during the fixed time and the return to 9.4 T, and it ends with the detection sequence. For all photo-CIDNP experiments, the samples were measured in dark and light, respectively, with the same number of scans. The illumination source was a 400-nm 2-W LED (Chanzon, China) and the illumination time was optimized to 0.5 s. A fresh aliquot of the sample stock was taken for a measurement at each magnetic field to compensate the effect of photo-bleaching. The temperature was 289 K for all samples with the exception that Ptaureo1a-LOV-C287S was measured at 277 K. The line-broadening for 15N and 13C NMR spectra were set to 30 Hz and for 1H spectra was set to 1 Hz. The 15N and 13C NMR spectra were phased to the external standard, a mixture of 0.1 M 15N labelled urea and 0.1 M 13C labelled methanol in DMSO. The chemical shifts of 15N NMR spectra were relative to liquid ammonia and referenced externally to urea 15N at 76.3 ppm69. To present the field-dependence of the photo-CIDNP effect, the selected hyperpolarized signal was integrated and plotted against the field at which the sample was illuminated. For straightforward comparison, the signal strength at the hyperpolarization maximum is set to unity and the other signals are normalized to this value. The positive and negative signs of the y-axis indicate absorptive and emissive hyperpolarization, respectively. The error bars of the 1H photo-CIDNP data represent the standard deviation of the mean value obtained from three measurements; the uncertainty of the 13C and 15N photo-CIDNP data represents the noise level relative to the corresponding hyperpolarized signal area. The spectra shown in Fig. 2 and Supplementary Information Figure S1, S2, and S4 were created with OriginPro Version 2017.
References
Daviso, E. Photochemically induced dynamic nuclear polarization (photo-CIDNP) magic-angle spinning NMR. In Biophysical Techniques in Photosynthesis (eds Aartsma, T. & Matysik, J.) 385–399 (Springer, Berlin, 2008).
Matysik, J., Diller, A., Roy, E. & Alia, A. The solid-state photo-CIDNP effect. Photosynth. Reson. 102, 427–435 (2009).
Bode, B. E., Thamarath, S. & Matysik, J. The solid-state photo-CIDNP effect and its analytical application photo-CIDNP MAS NMR to study radical pairs. In Hyperpolarization methods in NMR spectroscopy Vol. 338 (ed. Kuhn, L. T.) 105–122 (Springer, Berlin, 2013).
Closs, G. L., Forbes, M. D. E. & Norris, J. R. Spin-polarized electron paramagnetic resonance spectra of radical pairs in micelles. Observation of electron spin–spin interactions. J. Phys. Chem. 91, 3592–3599 (1987).
Hore, P. J., Hunter, D. A., McKie, C. D. & Hoff, A. J. Electron paramagnetic resonance of spin-correlated radical pairs in photosynthetic reactions. Chem. Phys. Lett. 137, 495–500 (1987).
Zysmilich, M. G. & McDermott, A. Photochemically induced dynamic nuclear polarization in the solid-state I5N spectra of reaction centers from photosynthetic bacteria Rhodobacter sphaeroides R-26. J. Am. Chem. Soc. 116, 8362–8363 (1994).
Alia, et al. Photochemically induced dynamic nuclear polarization in photosystem I of plants observed by 13C magic-angle spinning NMR. J. Am. Chem. Soc. 126, 12819–12826 (2004).
Diller, A. et al. Photo-CIDNP solid-state NMR on photosystems I and II: what makes P680 special?. Photosynth. Res. 84, 303–308 (2005).
Diller, A. et al. Signals in solid-state photochemically induced dynamic nuclear polarization recover faster than signals obtained with the longitudinal relaxation time. J. Phys. Chem. B 111, 10606–10614 (2007).
Janssen, G. J. et al. Photochemically induced dynamic nuclear polarization NMR on photosystem II: donor cofactor observed in entire plant. Sci. Rep. 8, 17853 (2018).
Janssen, G. J. et al. Observation of the solid-state photo-CIDNP effect in entire cells of cyanobacteria Synechocystis. Photosynth. Res. 104, 275–282 (2010).
Janssen, G. J., Roy, E., Matysik, J. & Alia, A. 15N photo-CIDNP MAS NMR to reveal functional heterogeneity in electron donor of different plant organisms. Appl. Magn. Reson. 42, 57–67 (2012).
Zill, J. C. et al. Photo-CIDNP in the reaction center of the diatom Cyclotella meneghiniana observed by 13C MAS NMR. Z. Phys. Chem. 231, 347–367 (2017).
Zill, J. C. et al. 15N photo-CIDNP MAS NMR on both photosystems and magnetic field-dependent 13C photo-CIDNP MAS NMR in photosystem II of the diatom Phaeodactylum tricornutum. Photosynth. Res. 140, 151–171 (2019).
Matysik, J. et al. Photo-CIDNP 13C magic angle spinning NMR on bacterial reaction centres: exploring the electronic structure of the special pair and its surroundings. Biol. Chem. 382, 1271–1276 (2001).
Prakash, S., De Groot, H. J. M., Matysik, J., Gast, P. & Jeschke, G. 13C chemical shift map of the active cofactors in photosynthetic reaction centers of Rhodobacter sphaeroides revealed by photo-CIDNP MAS NMR. Biochemistry 46, 8953–8960 (2007).
Daviso, E. et al. The electronic structure of the primary electron donor of reaction centers of purple bacteria at atomic resolution as observed by photo-CIDNP 13C NMR. Proc. Natl. Acad. Sci. U.S.A. 106, 22281–22286 (2009).
Paul, S. et al. 15N photo-CIDNP MAS NMR analysis of a bacterial photosynthetic reaction center of Rhodobacter sphaeroides wildtype. J. Chem. Phys. 151, 195101 (2019).
Roy, E. et al. Characterization of the primary radical pair in reaction centers of Heliobacillus mobilis by 13C photo-CIDNP MAS NMR. Biochemistry 47, 4629–4635 (2008).
Thamarath, S. S. et al. Whole cell nuclear magnetic resonance characterization of two photochemically active states of the photosynthetic reaction center in heliobacteria. Biochemistry 51, 5763–5773 (2012).
Roy, E. et al. 13C photo-CIDNP MAS NMR on the reaction center of the green sulphur bacterium at two different magnetic fields. In Energy from the sun (eds Allen, J. et al.) 173–176 (Springer, Dordrecht, 2008).
Thamarath, S. S. et al. Electron spin density distribution in the special pair triplet of Rhodobacter sphaeroides R26 revealed by magnetic field dependence of the solid-state photo-CIDNP effect. J. Am. Chem. Soc. 134, 5921–5930 (2012).
Gräsing, D. et al. Field-cycling NMR with high-resolution detection under magic-angle spinning: determination of field-window for nuclear hyperpolarization in a photosynthetic reaction center. Sci. Rep. 7, 12111 (2017).
Daviso, E. et al. Electron–nuclear spin dynamics in a bacterial photosynthetic reaction center. J. Phys. Chem. C 113, 10269–10278 (2009).
Daviso, E. et al. Nanosecond-flash 15N photo-CIDNP MAS NMR on reaction centers of Rhodobacter sphaeroides R26. Appl. Magn. Reson. 37, 49–63 (2010).
Sai Sankar Gupta, K. B. et al. Spectral editing through laser-flash excitation in two-dimensional photo-CIDNP MAS NMR experiments. J. Magn. Reson. 246, 9–17 (2014).
Matysik, J., Gast, P., van Gorkom, H. J., Hoff, A. J. & de Groot, H. Photochemically induced nuclear spin polarization in reaction centers of photosystem II observed by 13C-solid-state NMR reveals a strongly asymmetric electronic structure of the P680+ primary donor chlorophyll. Proc. Natl. Acad. Sci. 97, 9865–9870 (2000).
Daviso, E. et al. A 10 000-fold nuclear hyperpolarization of a membrane protein in the liquid phase via a solid-state mechanism. J. Am. Chem. Soc. 133, 16754–16757 (2011).
Jeschke, G. & Matysik, J. A reassessment of the origin of photochemically induced dynamic nuclear polarization effects in solids. Chem. Phys. 294, 239–255 (2003).
Closs, G. L. On the Overhauser mechanism of chemically induced nuclear polarization as suggested by Adrian. Chem. Phys. Lett. 32, 277–278 (1975).
Goldstein, R. A. & Boxer, S. G. Effects of nuclear spin polarization on reaction dynamics in photosynthetic bacterial reaction centers. Biophys. J. 51, 937–946 (1987).
McDermott, A., Zysmilich, M. G. & Polenova, T. Solid state NMR studies of photoinduced polarization in photosynthetic reaction centers: mechanism and simulations. Solid State Nucl. Magn. Reson. 11, 21–47 (1998).
Jeschke, G. A new mechanism for chemically induced dynamic nuclear polarization in the solid state. J. Am. Chem. Soc. 120, 4425–4429 (1998).
Jeschke, G. Electron–electron–nuclear three-spin mixing in spin-correlated radical pairs. J. Chem. Phys. 106, 10072–10086 (1997).
Polenova, T. & Mcdermott, A. E. A coherent mixing mechanism explains the photoinduced nuclear polarization in photosynthetic reaction centers. J. Phys. Chem. Burn. Phys. Chem. B 103, 535–548 (1999).
Sosnovsky, D. V., Jeschke, G., Matysik, J., Vieth, H. M. & Ivanov, K. L. Level crossing analysis of chemically induced dynamic nuclear polarization: towards a common description of liquid-state and solid-state cases. J. Chem. Phys. 144, 144202 (2016).
Sosnovsky, D. V. et al. Magnetic field and orientation dependence of solid-state CIDNP. J. Chem. Phys. 150, 094105 (2019).
Morozova, O. B., Yurkovskaya, A. V., Vieth, H. M., Sosnovsky, D. V. & Ivanov, K. L. Light-induced spin hyperpolarisation in condensed phase. Mol. Phys. 115, 2907–2943 (2017).
Ding, Y. et al. Nuclear spin-hyperpolarization generated in a flavoprotein under illumination: experimental field-dependence and theoretical level crossing analysis. Sci. Rep. 9, 18436 (2019).
Richter, G. et al. Photochemically induced dynamic nuclear polarization in a C450A mutant of the LOV2 domain of the Avena sativa blue-light receptor phototropin. J. Am. Chem. Soc. 127, 17245–17252 (2005).
Thamarath, S. S., Heberle, J., Hore, P. J., Kottke, T. & Road, S. P. Solid-state photo-CIDNP effect observed in phototropin LOV1-C57S by magic-angle spinning nmr spectroscopy. J. Am. Chem. Soc. 132, 15542–15543 (2010).
Salomon, M., Christie, J. M., Knieb, E., Lempert, U. & Briggs, W. R. Photochemical and mutational analysis of the FMN-binding domains of the plant blue light receptor, phototropin. Biochemistry 39, 9401–9410 (2000).
Kottke, T., Heberle, J., Hehn, D., Dick, B. & Hegemann, P. Phot-LOV1: photocycle of a blue-light receptor domain from the green alga Chlamydomonasreinhardtii. Biophys. J. 84, 1192–1201 (2003).
Wang, X. J., Thamarath, S. S., Alia, A., Bode, B. E. & Matysik, J. The solid-state photo-CIDNP effect. Wuli Huaxue Xuebao Acta Phys. Chim. Sin. 32, 399–404 (2016).
Eisenreich, W. et al. Tryptophan 13C nuclear-spin polarization generated by intraprotein electron transfer in a LOV2 domain of the blue-light receptor phototropin. Biochem. Soc. Trans. 37, 382–386 (2009).
Kothe, G. et al. Detecting a new source for photochemically induced dynamic nuclear polarization in the LOV2 domain of phototropin by magnetic-field dependent 13C NMR spectroscopy. J. Phys. Chem. B 118, 11622–11632 (2014).
Zhukov, I. V. et al. Field-cycling NMR experiments in an ultra-wide magnetic field range: relaxation and coherent polarization transfer. Phys. Chem. Chem. Phys. 20, 12396–12405 (2018).
Takahashi, F. et al. Aureochrome, a photoreceptor required for photomorphogenesis in stramenopiles. Proc. Natl. Acad. Sci. U.S.A. 104, 19625–19630 (2007).
Nakatani, Y. & Hisatomi, O. Molecular Mechanism of photozipper, a light-regulated dimerizing module consisting of the bZIP and LOV domains of aureochrome-1. Biochemistry 54, 3302–3313 (2015).
Suetsugu, N. & Wada, M. Evolution of three LOV blue light receptor families in green plants and photosynthetic stramenopiles: phototropin, ZTL/FKF1/LKP2 and aureochrome. Plant Cell Physiol. 54, 8–23 (2013).
Consiglieri, E. et al. Single mutation in a novel bacterial LOV protein yields a singlet oxygen generator. Photochem. Photobiol. Sci. 18, 2657–2660 (2019).
Schellenberger Costa, B. et al. Aureochrome 1a is involved in the photoacclimation of the diatom Phaeodactylum tricornutum. PLoS ONE 8, e74451 (2013).
Herman, E., Sachse, M., Kroth, P. G. & Kottke, T. Blue-light-induced unfolding of the Jα helix allows for the dimerization of aureochrome-LOV from the diatom Phaeodactylum tricornutum. Biochemistry 52, 3094–3101 (2013).
Eisenreich, W., Joshi, M., Weber, S., Bacher, A. & Fischer, M. Natural abundance solution 13C NMR studies of a phototropin with photoinduced polarization. J. Am. Chem. Soc. 130, 13544–13545 (2008).
Guex, N., Peitsch, M. C. & Schwede, T. Automated comparative protein structure modeling with SWISS-MODEL and Swiss-PdbViewer: a historical perspective. Electrophoresis 30, 162–173 (2009).
Fedorov, R. et al. Crystal structures and molecular mechanism of a light-induced signaling switch: the Phot-LOV1 domain from Chlamydomonas reinhardtii. Biophys. J. 84, 2474–2482 (2003).
Banerjee, A., Herman, E., Kottke, T. & Essen, L. O. Structure of a native-like Aureochrome 1a LOV domain dimer from Phaeodactylum tricornutum. Structure 24, 171–178 (2016).
Mitra, D., Yang, X. & Moffat, K. Crystal structures of aureochrome1 LOV suggest new design strategies for optogenetics. Structure 20, 698–706 (2012).
Eisenreich, W. et al. Strategy for Enhancement of 13C-photo-CIDNP NMR spectra by exploiting fractional 13C-labeling of tryptophan. J. Phys. Chem. B 119, 13934–13943 (2015).
Hore, P. J., Egmond, M. R., Edzes, H. T. & Kaptein, R. Cross-relaxation effects in the photo-CIDNP spectra of amino acids and proteins. J. Magn. Reson. 49, 122–150 (1982).
Kiryutin, A. S., Morozova, O. B., Kuh, L. T., Yurkovskaya, A. V. & Hore, P. J. 1H and 13C hyperfine coupling constants of the tryptophanyl cation radical in aqueous solution from microsecond time-resolved CIDNP. J. Phys. Chem. B 111, 11221–11227 (2007).
Moser, C. C., Keske, M. J., Warncke, K., Farid, S. R. & Dutton, P. L. Nature of biological electron transfer. Nature 355, 796–802 (1992).
Barry, B. A. & Babcock, G. T. Tyrosine radicals are involved in the photosynthetic oxygen-evolving system. Proc. Natl. Acad. Sci. U.S.A. 84, 7099–7103 (1987).
Nagai, H., Fukushima, Y., Okajima, K., Ikeuchi, M. & Mino, H. Formation of interacting spins on flavosemiquinone and tyrosine radical in photoreaction of a blue light sensor BLUF protein TePixD. Biochemistry 47, 12574–12582 (2008).
Nakasone, Y., Ohshima, M., Okajima, K., Tokutomi, S. & Terazima, M. Photoreaction dynamics of LOV1 and LOV2 of phototropin from Chlamydomonas reinhardtii. J. Phys. Chem. B 122, 1801–1815 (2018).
Kopka, B. et al. Electron transfer pathways in a light, oxygen, voltage (LOV) protein devoid of the photoactive cysteine. Sci. Rep. 7, 13346 (2017).
Guo, H., Kottke, T., Hegemann, P. & Dick, B. The phot LOV2 domain and its interaction with LOV1. Biophys. J. 89, 402–412 (2005).
Shaka, A. J., Keeler, J. & Freeman, R. Evaluation of a new broadband decoupling sequence: WALTZ-16. J. Magn. Reson. 53, 313–340 (1983).
Wishart, D. S. et al. 1H, 13C and 15N chemical shift referencing in biomolecular NMR. J. Biomol. NMR 6, 135–140 (1995).
Acknowledgements
Y. D. and J. M. thank the Deutsche Forschungsgemeinschaft (MA 4972/11-1). Y. D. also acknowledges the Russian Foundation of Basic Research (Grant No. 19-33-50094) and the Deutscher Akademischer Austauschdienst (Referat ST22) for supporting his travel and work in Novosibirsk. A. S. K., A. V. Y., R. Z. S. and K. L. I. acknowledge the Russian Foundation of Basic Research (Grant No. 19-29-10028) and the Russian Ministry for Science and Higher Education (Project No. AAAA-A16-116121510087-5). T. K. acknowledges support by a Heisenberg fellowship of the Deutsche Forschungsgemeinschaft (KO 3580/4-2). We thank Wolfgang Gärtner (Univ. Leipzig), Tanja Gulder (Univ. Leipzig), Aba Losi (Univ. Parma) and Igor Schapiro (Hebrew University) for helpful discussions.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Contributions
R.Z.S., K.L.I., A.V.Y. and J.M. designed the research. The sample preparation was mainly performed by Y.D., Z.Z., Q.X., P.K., S.B. and T.K. Most of the field-dependent photo-CIDNP experiments are done by Y.D. and A.S.K. Y.D., A.S.K., K.L.I., A.V.Y. and J.M. interpreted field-dependent CIDNP data. The figures are created by Y.D. The manuscript was written through contributions of all authors. All authors have given approval to the final version of the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ding, Y., Kiryutin, A.S., Zhao, Z. et al. Tailored flavoproteins acting as light-driven spin machines pump nuclear hyperpolarization. Sci Rep 10, 18658 (2020). https://doi.org/10.1038/s41598-020-75627-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-75627-z
This article is cited by
-
Detection of sub-nmol amounts of the antiviral drug favipiravir in 19F MRI using photo-chemically induced dynamic nuclear polarization
Scientific Reports (2024)
-
Photo-CIDNP in Solid State
Applied Magnetic Resonance (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.