Abstract
Chronic kidney disease (CKD) of unknown etiology (CKDu) mostly affects agricultural communities in Central America, South Asia, Africa, but likely also in North America and Australia. One such area with increased CKDu prevalence is the Medawachchiya District Secretariat Division of the Anuradhapura District in the North Central Province of Sri Lanka. Recent research has focused on the presence of various microbial pathogens in drinking water as potential causal or contributing factors to CKDu, yet no study to date has performed a more comprehensive microbial and water chemistry assessment of household wells used for domestic water supply in areas of high CKDu prevalence. In this study, we describe the chemical composition and total microbial content in 30 domestic household wells in the Medawachchiya District Secretariat Division. While the chemical composition in the tested wells mostly lies within standard drinking water limits, except for high levels of fluoride (F), magnesium (Mg), sodium (Na), chloride (Cl) and calcium (Ca) in some samples, we find a frequent presence of cyanotoxin-producing Microcystis, confirming earlier studies in Sri Lanka. Since the total microbial content of drinking water also directly influences the composition of the human gut microbiome, it can be considered an important determinant of health. Several bacterial phyla were previously reported in the gut microbiome of patients with CKD. Using these bacteria phyla to define operational taxonomic units, we found that these bacteria also occur in the microbiome of the sampled well water. Based on available environmental data, our study demonstrates associations between the abundances of these bacteria with geographical distribution, well water temperature and likely fertilizer use in the local surface water catchment area of the individual household wells. Our results reinforce the recommendation that household wells with stagnant or infrequently used water should be purged prior to use for drinking water, bathing and irrigation. The latter is suggested because of the reported potential accumulation of bacterial toxins by agricultural crops. The observation that bacteria previously found in chronic kidney disease patients are also present in household wells requires a more detailed systematic study of both the human gut and drinking water microbiomes in CKDu patients, in relation to disease prevalence and progression.
Similar content being viewed by others
Introduction
The 2016 Annual Health Statistics of Sri Lanka lists chronic kidney disease (CKD) as the leading cause of hospital deaths in Anuradhapura and Polonnaruwa, the two districts of North Central Province1. The above average prevalence of kidney disease is attributed to a chronic kidney disease of unknown or undetermined origin (CKDu), that appears to have emerged over the past 20 years. Though there is neither full consensus nor a comprehensive case definition, by general agreement the disease is not associated with known risk factors for CKD, such as diabetes or hypertension, and occurs mostly in poorer young and middle-aged individuals in agricultural communities. Progressive and asymptomatic until the late stages, its characteristic chronic tubulointerstitial tissue pathology with secondary glomerulosclerosis2 has recently resulted in a change of terminology from CKDu to CINAC, chronic interstitial nephritis in agricultural communities (CINAC)3.
In Sri Lanka’s North Central region, notably the Medawachchiya District Secretariat Division of the Anuradhapura District which has some of the highest prevalence of CKDu4, 80% of the mostly farming population relies on groundwater for their daily water needs5, including drinking water. Previous reports have investigated potential environmental6,7, genetic8, occupational and social3,9,10 risk factors for CKDu without generating conclusive evidence for specific causes. Recent studies have pointed towards an association between CKDu, groundwater chemistry and the water quality in household wells11,12,13,14, suggesting that the disease could be caused by hydrogeochemical factors such as high fluoride and water hardness, heavy metals and microbial contamination or combinations thereof. CKDu is presently viewed as a complex disease mostly found in certain rural populations and provisionally defined by the absence of consistently observed causes commonly associated with chronic kidney disease. Recent research supports the hypothesis that CKDu is likely caused by a combination of intrinsic and external factors such as malnutrition, dehydration, and the consumption of poor quality drinking water including those high in fluoride (F) concentrations14,15, or pathogens and bacterial toxins. However, the actual hierarchy of importance of the various contributing disease causes or, importantly, any risk-enhancing or mitigating interactions between these remain to be established15,16,17,18. Heavy metal exposure from groundwater has largely been ruled out as the major causal factor for CKDu in Sri Lanka7,19, however, contamination by pathogens and bacterial toxins in water used for irrigation, domestic and potable water supply remains a valid hypothesis17,18,20,21,22,23.
In particular, the toxin-producing cyanobacteria genera, including Microcystis and Cylindrospermopsis, as well as Leptospira interrogans can lead to kidney disease14,18,24,25 and have, therefore, been put forward as potential causes of CKDu21,26,27. Significant increases of cyanobacteria in freshwater resources in Sri Lanka have occurred over the past century as a result of eutrophication and rising temperatures21,28, demonstrating that altered water chemistry is likely to promote the presence of pathogenic bacteria in this region.
There has not yet been a study that specifically probes into the association of the water chemistry in individual household wells with the composition of their complete microbiome.
We suggest that well water chemistry, including temperature, pH, dissolved oxygen (DO) and nutrients (phosphorus (P),potassium (K) and nitrate (NO3)) will favor the emergence of typical well water microbiomes, that are both determined by and determining the chemical characteristics of the well water.
The sampling for this project was undertaken, in an area of Sri Lanka which is highly affected by CKDu, as a pilot study to determine possible links between altered water chemistry and the microbiome of household well water used as a regular or occasional drinking, washing or cooking water source. The rationale for the total microbiome analysis of household well water beyond those bacteria already known to produce nephrotoxins is based on the most recent observation that the source and microbial content of drinking water (in addition to food) itself significantly determines the microbial composition of the human gut microbiome29,30,31. In turn, it is increasingly recognized that an individual’s human microbiome composition has a close, disease modifying, association across a wide range of health and disease conditions, including CKD32.
Methods
Sampling
Thirty domestic well water samples, comprised of shallow groundwater (between 1 and 9 m depth below ground surface), were collected from the Medawachchiya region between the 15th of April 2018 and 23rd of April 2018. Sample locations are shown in Fig. 1.
Sri Lanka map showing dry (< 1750 mm year−1), intermediate (1750–2500 mm year−1) and wet (> 2500 mm year−1) zones. Inset shows the study region in Medawachchiya with groundwater sampling locations shown as blue crosses. CKDu prevalence represents numbers of CKDu patients in each Grama Niladhari (GN) Division, showing high prevalence within the dry zone. Figure prepared in Surfer v.11.0.642 (www.goldensoftware.com/products/surfer).
Sampled water was obtained from a combination of lined and unlined wells and springs. During sampling, lining materials were observed to be either brick, concrete, or a combination of both. Rock types in the wells were predominantly weathered quartz-feldspathic gneiss overlain with between 0–9 m of soil, surrounded by coconut trees, rice paddies, grass, bushland and home gardens.
To sample, a peristaltic pump was placed 1 m (m) below the standing water level where possible, or between the base of the well and the surface where standing water level was < 1 m. Electrical conductivity (EC), water temperature, dissolved oxygen (DO) and pH were measured using a flow cell connected to the pump and a YSI meter. During pumping wells simultaneously refilled with water, therefore water was pumped until field parameters stabilized prior to sampling. A high flow Waterra 0.45 µm filter was then connected to the pump outlet. Each sample bottle was rinsed three times with the filtered water sample and then filled. Cation samples were collected in 60 ml high-density polyethylene (HDPE) containers respectively, acidified using 0.4 ml of ultrapure nitric acid (HNO3) and refrigerated. Anion samples were collected in 60 ml HDPE bottles. Stable water isotope samples were collected in 2 × 30 ml HDPE bottles. Samples collected for the measurement of 13C of dissolved inorganic carbon (13CDIC) were collected in 2 × 12 ml glass vials ensuring no headspace. The samples were inverted to ensure no bubbles were present and refrigerated after collection. The filter unit was retained for microbiome analyses and frozen. The particulate organic carbon (13CPOC) samples were collected by adding a Millipore POC filter with 0.7 µm filter paper to the peristaltic pump outlet and ~ 2 L of water were passed through the filter. 13CPOC samples were then frozen. Tape was placed around the lid of all water samples to limit atmospheric exchange.
Samples were removed from the refrigerator or freezer when field work was complete and the samples were ready to be posted. Frozen samples were packed with other frozen samples in a foam cool box to ensure they would stay as cold as possible during transport. Once samples arrived at Australian Nuclear Science and Technology Organisation (ANSTO), samples for freezing or refrigeration were immediately flash frozen and refrigerated respectively and gamma-irradiated at 50 kGray prior to transfer to the respective labs for analysis at ANSTO.
Analysis
All water samples were analysed at Australian Nuclear Science and Technology Organisation (ANSTO). Major element compositions and trace element compositions were determined by inductively coupled plasma atomic emission spectroscopy (ICP-AES) using a Varian 820MS and inductively coupled plasma mass spectrometry (ICP-MS) using a Thermo Fisher iCAP7600 Duo respectively. Anions were measured by ion chromatography using a Dionex ICS-2100. The number of decimal places in the lab reports are determined for each sample based on the relative standard deviation (RSD) of the lab standard for the element at the relevant sample concentration. For ICPAES, ICPMS and IC, RSD are between 1 and 5% for data that are 10 times the limit of reporting (LOR). For samples with data that are < 10 times the LOR, the RSD are > 5%, and up to 50% if the sample is at the LOR. Stable water isotopes and stable dissolved inorganic carbon (DIC) isotopes were measured using Gas Bench II coupled to a continuous flow delta V advantage isotope ratio mass spectrometer. A two-point calibration was used to normalize the data which consisted of two standards that bracket the samples being analyzed. The results were reported relative to IAEA secondary standards which are certified relative to VPDB using a two point calibration, with final sample results accurate to ± 0.3 ‰ for δ13CDIC. The POC isotopes were measured using an elemental analyzer-isotopic ratio mass spectrometer.
To perform the microbiome analysis, DNA was extracted from the filters using the PowerSoil DNA isolation kit (MO BIO Laboratories) following the manufacturer’s instructions. The variable V3 and V4 regions of the 16S rRNA gene was then amplified via PCR (16S Forward primer: 5′-TCGTCGGCAGCGTCAGATGTGTATAAGAGACAGCCTACGGGNGGCWGCAG and 16S Reverse primer: 5′-GTCTCGTGGGCTCGGAGATGTGTATAAGAGACAGGACTACHVGGGTATCTAATCC). Each reaction consisted of 1 × KAPA HiFi HotSart ReadyMix (KAPA Biosystems), 0.2 µM of each primer and 12.5 ng of sample DNA in a final volume of 25 µl. The thermocycler conditions were 95 °C for 3 min, then 25 cycles at 98 °C for 30 s, 55 °C for 30 s and 72 °C for 30 s, then a final extension at 72 °C for 5 min. PCR products were then purified using AMPure XP beads (Beckman Coulter) following the Illumina 16S Metagenomic Sequencing Library preparation guide. At this point, samples were transferred to the Ramaciotti Centre for Genomics (UNSW Sydney, Australia) for further processing. Briefly, dual indices and Illumina sequencing adaptors were attached using the Nextera XT Index Kit. Samples were cleaned again using AMPure XP beads and then quantified, normalized and pooled before being sequenced on the Illumina MiSeq platform using paired 300 bp reads and MiSeq v3 reagents.
QIIME (Quantitative Insights Into Microbial Ecology) version 1.9.1 was used to process the sequence data33. The mapping file was validated using validate_mapping_file.py. Forward and reverse reads were then joined using the script multiple_join_paired_ends.py and then multiple_extract_barcodes.py was used to remove the primer sequences and process the data for subsequent steps. Further processing and quality filtering was performed using multiple_split_libraries_fastq.py, chimeras were identified with identify_chimeric_seqs.py and the usearch tool, then the chimeras removed with filter_fasta.py. Finally, operational taxonomic units (OTUs) were picked using pick_open_reference_otus.py, usearch and the SILVA rRNA database release 132. OTUs with total counts across all samples of less than 10 were removed.
Statistical analyses and figures were generated in R (v3.5.1). Species accumulation curves were generated using the vegan package. Stacked barplots, heatmaps and microbial networks were generated using the physeq package. Redundancy analysis (RDA) and principle component analysis (PCA) were performed in R and used to compare microbes to water chemistry parameters, and group co-occurrent CKD-associated microbe groups respectively. Count data have been used as scaling problems were encountered due to some microbes being present in very low abundances. Correlations between microbes and water chemistry parameters were also performed in R. Quantile–Quantile plots and constant variance were checked for linear correlations to ensure normality and constant variance of residuals. Where data was heavily skewed, the response variable was log or square root transformed to normalize. Non-parametric Spearman correlations were used where data remained heavily skewed after transformation.
Results
Geochemistry
Piper plots (Supplementary Figure 1) show that the sampled water types include calcium chloride and mixed water types34. Table 1 summarizes the concentration of various inorganic water quality parameters measured in this study compared to the World Health Organization (WHO) and Sri Lanka Standards Institution (SLSI) guideline values for drinking water35,36. Our data show high levels of fluoride (F) in the Medawachchiya region with 30% of wells sampled exceeding the WHO guideline value of 1.5 mg L−1 for F, and 57% exceeding the Sri Lanka specification of 1.0 mg L−1 for F in potable water. All other parameters including chromium (Cr), arsenic (As), barium (Ba), uranium (U), copper (Cu), nitrate (NO3) and nickel (Ni) fall below the WHO guideline values for drinking water for all samples (Table 1).
Microbial diversity
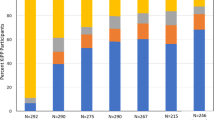
A total of 58 unique phyla categories were detected (Fig. 2). The microbial composition is dominated by 16 phyla which have abundances of ≥ 1%. These include Acidobacteria, Actinobacteria, Bacteroidetes, Chlamydiae, Chloroflexi, Cyanobacteria, Dependentiae, Epsilonbacteraeota, Firmicutes, Unassigned bacteria (Other), Patescibacteria, Planctomycetes, Proteobacteria, Spirochaetes, Verrucomicrobia and WPS-2 (Fig. 2).
Stacked bar chart showing abundance (counts) of microbial phyla in each sample. Data normalized to the median sequencing depth. NB: microbe data for samples CKDU-2-15 and CKDU-2-18 not available. Phylum shown with an asterisk symbol represent groups containing microbes identified in higher concentrations in CKD patients compared to healthy patients. These include Verrucomicrobia, Gammaproteobacteria, Enterobacteriacea, Methylobacterium and Desulfovibrio (of the Proteobacteria phylum), Leptospira (of the Spirochaetes phylum), and Holdemania, Turicibacter and Clostridium sensu stricto (of the Firmicutes phylum).
A heatmap of microbial counts in each sample and co-occurrence networks for these more abundant phyla show a negative association between Proteobacteria and Cyanobacteria for most samples (Fig. 3). Sample CKDU-2-04 however shows high Proteobacteria and Cyanobacteria counts, whilst CKDU-2-08, CKDU-2-12 and CKDU-2-30 show low Proteobacteria and low Cyanobacteria. Figure 3B indicates an association between Cyanobacteria, Patescibacteria and Verrucomicrobia.
(A) Heatmap of microbial phyla (> 1% abundance), with cluster analyses for sample grouping (upper cluster diagram) and microbial groupings (left cluster diagram). Data are scaled by rows to have a mean of zero and a standard deviation of one. (B) Network map based on Bray–Curtis distance showing similarities between microbial genera counts for phyla with abundance > 1%. Thicker lines represent greater similarity between phyla counts, thinner lines represent less similarity between phyla counts. Data normalized to the median sequencing depth.
Species accumulation curves (Supplementary Figure 2) indicate that there is a relatively low proportion of diversity in the samples, with 80% of phyla identified in the total dataset reached within approximately 4 of the 28 samples, and 95% reached within approximately 15 of the 28 samples. This suggests that sampling an additional 13 sites would only result in the identification of approximately 3 additional phyla which may not have been present at the other 15 sampling sites. 80% of the genera are reached within approximately 7 of the samples, and 95% reached within approximately 19 of the 28 samples, suggesting that the sampling of an additional 9 sites resulted in the identification of approximately 83 additional microbial genera.
Presence and geospatial mapping of previously identified CKD-associated bacteria
We identify bacteria which have been positively associated with CKD in previous studies37,38,39. These include Verrucomicrobia, Gammaproteobacteria, Enterobacteraceae, Microcystis, Leptospira, Clostridium sensu stricto, Desulfovibrio, Holdemania, Turicibacter and Methylobacterium (Table 2), which have been identified in higher abundances in CKD fecal samples compared to healthy patients37,38,39. The spatial distributions of these microbes are shown in Supplementary Figure 3.
However, we do not identify any Clostridium IV, Enterococcus or Paraprevotella or only identify very small counts of Alloprevotella (up to 2 counts in 4 wells, which may be a result of sequencing error) all of which have also been shown to be enriched in CKD patient stool37. We did not identify any cyanotoxin-producing Cylindrospermopsis, but do observe the cyanotoxin-producing genus Microcystis, both of which can cause kidney damage14,18,25. We identify only a very small number of Akkermansia (11 counts), found only in one well (CKDU-2-04). Akkermansia is a probiotic which has been shown to be significantly higher in abundance in healthy individuals compared to those with CKD37.
Water chemistry impacts on CKD-related bacteria counts
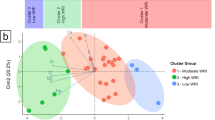
The redundancy analysis in Fig. 4 shows the associations between CKD-related microbes present in this study which have been previously identified in higher concentrations in fecal samples from CKD patients, and water chemistry parameters. The four clusters represent microbes most closely associated with each other (Supplementary Figure 4).
RDA results (Fig. 4 and Supplementary Table 1) show that microbes associated with Cluster 1, 2 and 4 correspond with RDA2 which is most highly represented by phosphorus (P), potassium (K), pH and 15N of particulate organic matter (15NPOM). Microbes associated with Cluster 3 correspond with RDA1 (Microcystis) and a combination of RDA1 and RDA2 (Leptospira and Verrucomicrobia). RDA1 is most highly associated with chemical parameters Mn and 13CDIC. The chemistry parameters used in the RDA explain approximately 47.8% of the data variability on RDA1 (x-axis) and RDA2 (y-axis) combined, suggesting that the variability of microbial abundances presented may also be highly influenced by other factors such as the abundances or presence of other OTUs not included in the analysis.
Linear regression analysis shows that Microcystis, Turicibacter and Leptospira are significantly positively correlated with pH (p = 0.03, p = 0.009 and p = 0.045 respectively). Cyanobacteria (including Microcystis) and Leptospira are positively correlated with DO (p = 0.018 and p = 0.009 respectively) and 13CDIC (p = 0.023 and p = 0.017 respectively), and negatively correlated with 13CPOC (p = 0.025 and p = 0.003 respectively). We also observe a positive correlation between pH and 13CDIC (p < 0.001) with increasing pH associated with increased 13CDIC enrichment. Methylobacterium appear to be temperature dependent with increasing temperatures resulting in increasing Methylobacterium counts (p = 0.004). Nutrients K and P appear to be linked to Actinobacteria and Gammaproteobacteria (Fig. 4) however linear regression for P or K and Actinobacteria shows that the relationships are not significant (p > 0.05). In contrast, Gammaproteobacteria is significantly positively linked to increasing K and P (p = 0.007 and p < 0.001 respectively). It is noted that P concentrations are low for all samples (> 0.01 mg L−1–0.24 mg L−1).
Discussion
Our water chemistry data confirm previous reports of increased levels of F in areas of high CKDu prevalence16, with 30% of household wells in the Medawachchiya region exceeding WHO recommendations, and more than half exceeding the Sri Lankan standard for potable water. In our association analysis of water chemistry and microbial abundances we find indication for a complex interaction between the microbial composition and water chemistry. The significant positive relationships observed between Microcystis and Cyanobacteria with DO and 13CDIC, as well as the negative correlation with 13CPOC suggests that these photosynthetic bacteria effect a significant increase in DO levels and play a role in carbon turnover due to their preferential fixation of 12C in CO2 via the Calvin cycle and subsequent release of oxygen. The utilization of the 12C-enriched carbon from the DIC pool results in the enrichment of 13C in the remaining DIC (i.e. more positive δ13CDIC values). At the same time this causes an enrichment in of 12C in the POC pool (i.e. more negative δ13CPOC values) when algae utilize this DIC40. For δ13CPOC, we observe values as low as − 43.1‰, whilst δ13CPOC values observed from most aquatic environments (oceans, rivers, lakes) typically range from approximately − 30 to − 19‰41,42,43,44. Some plankton sources, however, have shown δ13CPOC values as low as − 42‰45. Fractionation of stable carbon isotopes of CO2 by Cyanobacteria has previously been observed in a lake system46. The enrichment of 13CDIC caused by these photosynthetic bacteria is associated with a significant increase in pH, likely due to the reduction of carbonic acid resulting from the removal of dissolved CO2. The increase in Cyanobacteria and pH alters microbial composition resulting in an increase in Turicibacter, Microcystis and Leptospira. Significant correlations between DO, δ13CDIC and δ13CPOC with Letospira, a non-photosynthetic bacterium is also observed, which appears to be due to its co-occurrence with Cyanobacteria (p = 0.022). The presence of Microcystis is relevant for domestic well waters being used for drinking water and where these waters are being used on crops. Cyanotoxins have been shown to bioaccumulate in plants which may ultimately impact on human health when consumed47. We suggest that further research focus on identifying the presence and concentrations of cyanotoxins in household wells. Major cyanotoxin groups to consider include cylindrospermopsin, nodularin, microcystins, anatoxins and saitoxins48. Recent research identified a negative association between the use of boiled water and CKD probability49, however short-term boiling does not statistically decrease the concentrations of cyanotoxins50. Further, behavioural differences within the same household particularly under economic constraints would merit closer scrutiny. We note that whilst low DO concentrations may indicate decreased Cyanobacterial abundances, cyanotoxins may build up and remain in high concentrations in stagnant waters. Furthermore, the presence of Leptospira in these waters indicates that well waters in the region may not only be harmful to drink, but also when used for bathing as this can allow Leptospira to enter the body through cuts or abrasions on the skin51.
Predicated on the notion that the microbiome of the entire alimentary system reflects host diet52,53,54 environment and lifestyle factors55 and is also strongly influenced by the quality and microbial content of drinking water30,31,56,57,58,59,60, we undertook a speculative interrogation of the microbial composition and abundance in the water contained in household wells based on bacterial phyla and genera that have previously found the gut microbiome of patients with chronic kidney disease. Bacterial abundance studies of the stool from patients with CKD show significantly reduced abundances of the phyla Actinobacteria, the abundances of Verrucomicrobia and Enterobacteriaceae family are higher. On the genus level, stool from CKD patients contains fewer Akkermansia, Alloprevotella and Parasutterella, and significantly higher Enterococcus, Clostridium IV, Paraprevotella, Clostridium sensu stricto and Desulfovibrio37,38,39. Higher abundances of potentially harmful Gammaproteobacteria, and lower abundances of beneficial genera including Rumincoccacea, Bifidobacteria and Lactobacilli have also been identified in CKD patients38,39.
While we found that most bacteria linked to the CKD OTUs of interest showed no clear relationship with P or K concentration in water, we observed a significant positive correlation between Gammaproteobacteria with increased K and P. One possible source of increased P could be the increased use of P fertilizer over the past few decades due to the intensification of crop production61. Research has demonstrated that soils treated with P fertilizers appear to have higher levels of Gammaproteobacteria62 which may leach into surface waters and shallow groundwaters.
Methylobacterium are opportunistic pathogens which can result in infection in immunocompromised patients including those with acute and chronic renal failure63. Methylobacterium infection in patients with kidney disease can manifest as peritonitis, pneumonia, abdominal pain and cloudy dialysate64. The optimum growth of these bacteria occurs between 25 and 30 °C with limited growth beyond 40°C65. We observe significant increases in this genus with increased temperature in our dataset (range = 27–32 °C). This may suggest that future climate change has the potential to alter spatial or temporal distributions of these pathogens. For example, presence of Methylobacterium in surface waters may decline in the warmer months with increasing ambient temperatures reaching closer to 40 °C, and increase in cooler months, or in cooler regions such as the south-central intermediate rainfall zone where average temperatures may be within the optimum growth range for Methylobacterium. The variable presence of Methylobacterium would then be similar to Microcystis and other pathogenic bacteria in these stagnant waters, in that drought zones can lead to the periodic build-up of extremely high levels of toxins or pathogens due to reduced flushing by rainfall and thus render the water of many household wells non-potable. This is particularly of concern considering the number of overall consecutive dry and wet days in Sri Lanka has increased and decreased respectively, with an expansion of the dry zone (average annual precipitation < 1750 mm) observed for the period 1961–1990 compared to 1911–194066. In inland regions, research suggests rainfall during the wet seasons has declined by 40% from 1869 to 199366, leading to increase stagnation of well waters in the central regions. In contrast, the wetter western regions of Sri Lanka are projected to experience increased rainfall during the Southwest Monsoon due to climate change. This is likely to result in a flushing of well waters in areas that are not currently affected by CKDu (Fig. 1).
Limitations and conclusions
Traditional nosography (the classification and description of diseases) proceeds by establishing clinico-pathological correlations between clinical illness presentation, the exposure to one or more pathogen and the presence of histopathological or other biological features. In the case of CKDu/CINAC, this approach is limited by the delayed onset of the disease which forces the observer to back-extrapolate to past pathogen exposure. This approach cannot entirely exclude overlap with the clinical presentation of CKD, and the shared risk factor of low socio-economic status in these regions. It is not fully resolved whether the prevalence of farmworkers amongst the patients is inherent to their low socio-economic status and associated range of other unfavorable health determinants (nutritional and associated developmental deficits), or whether it is an independent risk factor associated with farm work-related pathogen exposure. The latter is supported by the observation in CINAC patients of variable lysosomal anomalies that are consistent with a toxic insult67. A formal link between one or more specific toxins to the lysosomal injury has not yet been established, as these or similar lysosomal anomalies tend to be observable across a range of disease conditions. Given the above, CKDu is now considered a epidemiological problem which is best captured by a pragmatic multilevel clinical case definition1,68.
A complementary approach to the isolation of single or multiple causes of disease, is to view health status as a complex emergent state resulting from adaptive social and biological network interactions68,69. In our analysis of drinking water, which represents a potentially important source of pathogenic exposure, we have therefore described the complete internal microbial ecology of household wells using operational taxonomic units derived from the internal gut microbial ecology of patients with CKD. While we do not have the data to comment on the extent to which the microbial signature of a given household well water might have determined that of the gut microbiome of the consumer, it is a non-trivial observation that there are indeed relevant bacteria in both microbial ecosystems.
This allows the speculation that the microbiome of a household well co-determines the gut microbiome of the user and thus could confer disease risk beyond that associated with the known pathogenic microbes that were also found in our samples. It is noted that the definition of the OTUs was limited by the available knowledge in regard to their presence in kidney disease and that a sequence-based taxonomy alone does not fully capture both genomic or functional-metabolic differences between bacteria70.
The outcomes of this study have led to many more questions around CKDu research. Further work should focus on identifying what the gut microbiome composition is in patients with CKDu compared to those with CKD. Investigations into the gut microbiome of healthy household members who consume the same water source would also provide much needed information. Furthermore, studies designed to understand intake volumes of water for consumers, bacterial densities, the presence and concentrations of cyanotoxins, how the well water microbiome changes seasonally and identifying the range of uses for the water from the household wells are needed. It would also be useful to compare samples from household wells in non-CKDu prevalent areas to the results from this study. Understanding the differences both in the quality of the water and the consumption patterns within individual households71 will also provide further information on the complex interactions between water and gut microbiomes.
It is likely that anticipated increases in water temperatures due to climate change, and increasing nutrients due to current farming practices, will impact on the quality of drinking water and, more speculatively, might change human gut microbial composition in its wake.
Data availability
All data and code are available from the corresponding author upon request.
References
Wijewickrama, E. S., Gunawardena, N., Jayasinghe, S. & Herath, C. CKD of unknown etiology (CKDu) in Sri Lanka: a multilevel clinical case definition for surveillance and epidemiological studies. Kidney Int. Rep. 4, 781–785. https://doi.org/10.1016/j.ekir.2019.03.020 (2019).
Athuraliya, N. T. C. et al. Uncertain etiologies of proteinuric-chronic kidney disease in rural Sri Lanka. Kidney Int. 80, 1212–1221. https://doi.org/10.1038/ki.2011.258 (2011).
Jayasumana, C. et al. Chronic interstitial nephritis in agricultural communities: a worldwide epidemic with social, occupational and environmental determinants. Nephrol. Dial. Transplant. 32, 234–241. https://doi.org/10.1093/ndt/gfw346%JNephrologyDialysisTransplantation (2016).
Ranasinghe, A. V. et al. The incidence, prevalence and trends of Chronic Kidney Disease and Chronic Kidney Disease of uncertain aetiology (CKDu) in the North Central Province of Sri Lanka: an analysis of 30,566 patients. BMC Nephrol. 20, 338. https://doi.org/10.1186/s12882-019-1501-0 (2019).
Kuruppuarachchi, D. S. P. Impact of agriculture on groundwater: Sri Lankan perspective. J. Food Agric. 5, 1–4. https://doi.org/10.4038/jfa.v5i1-2.5176 (2012).
Kulathunga, M. R. D. L., Ayanka Wijayawardena, M. A., Naidu, R. & Wijeratne, A. W. Chronic kidney disease of unknown aetiology in Sri Lanka and the exposure to environmental chemicals: a review of literature. Environ. Geochem. Health 41, 2329–2338. https://doi.org/10.1007/s10653-019-00264-z (2019).
Nanayakkara, S. et al. Systematic evaluation of exposure to trace elements and minerals in patients with chronic kidney disease of uncertain etiology (CKDu) in Sri Lanka. J. Trace Elem. Med Biol. 54, 206–213. https://doi.org/10.1016/j.jtemb.2019.04.019 (2019).
Friedman, D. & Luyckx, V. A. Genetic and developmental factors in chronic kidney disease hotspots. Semin. Nephrol. 39, 244–255. https://doi.org/10.1016/j.semnephrol.2019.02.002 (2019).
Nanayakkara, I., Dissanayake, R. K. & Nanayakkara, S. The presence of dehydration in paddy farmers in an area with chronic kidney disease of unknown aetiology. Nephrology 25, 156–162. https://doi.org/10.1111/nep.13605 (2019).
O’Callaghan-Gordo, C. et al. Prevalence of and risk factors for chronic kidney disease of unknown aetiology in India: secondary data analysis of three population-based cross-sectional studies. BMJ Open 9, e023353. https://doi.org/10.1136/bmjopen-2018-023353 (2019).
Makehelwala, M., Wei, Y., Weragoda, S. K., Weerasooriya, R. & Zheng, L. Characterization of dissolved organic carbon in shallow groundwater of chronic kidney disease affected regions in Sri Lanka. Sci. Total Environ. 660, 865–875. https://doi.org/10.1016/j.scitotenv.2018.12.435 (2019).
de Silva, M. W. A. Drinking water and chronic kidney disease of unknown aetiology in Anuradhapura, Sri Lanka. Anthropol. Med. 26, 311–327. https://doi.org/10.1080/13648470.2018.1446822 (2018).
Cooray, T. et al. Assessment of groundwater quality in CKDu affected areas of Sri Lanka: implications for drinking water treatment. Int. J. Environ. Res. Public Health 16, 1698 (2019).
Herath, H. M. A. S. et al. Potential risk of drinking water to human health in Sri Lanka. Environ. Forens. 18, 241–250. https://doi.org/10.1080/15275922.2017.1340364 (2017).
Balasooriya, S. et al. Possible links between groundwater geochemistry and chronic kidney disease of unknown etiology (CKDu): an investigation from the Ginnoruwa region in Sri Lanka. Exposure Health https://doi.org/10.1007/s12403-019-00340-w (2019).
Levine, K. et al. Quest to identify geochemical risk factors associated with chronic kidney disease of unknown etiology (CKDu) in an endemic region of Sri Lanka—a multimedia laboratory analysis of biological, food, and environmental samples. Int. J. Devoted Prog. Use Monit. Data Assess. Environ. Risks Man Environ. 188, 1–16. https://doi.org/10.1007/s10661-016-5524-8 (2016).
Jayatissa, L. P., Silva, E. I. L., McElhiney, J. & Lawton, L. A. Occurrence of toxigenic cyanobacterial blooms in freshwaters of Sri Lanka. Syst. Appl. Microbiol. 29, 156–164. https://doi.org/10.1016/j.syapm.2005.07.007 (2006).
Yang, C. W. Leptospirosis renal disease: emerging culprit of chronic kidney disease unknown etiology. Nephron 138, 129–136. https://doi.org/10.1159/000480691 (2018).
Herath, H. M. A. S. et al. Arsenic, cadmium, lead, and chromium in well water, rice, and human urine in Sri Lanka in relation to chronic kidney disease of unknown etiology. J. Water Health 16, 212–222. https://doi.org/10.2166/wh.2018.070 (2018).
Manage, P. Cyanotoxins: a hidden cause of chronic idney disease of unknown etiology (CKDu) in Sri Lanka—a review. Sri Lanka J. Aquat. Sci 24, 1–10 (2019).
Kulasooriya, S. A. Toxin producing freshwater cyanobacteria of Sri Lanka. Ceylon J. Sci. 46, 3–16 (2017).
Zakeel, M. C. M., Weerasinghe, P. A., Kumari, B. A. D. G. & Wickremasinghe, H. A. M. Spatial and temporal dynamics and parameters of growth of toxic cyanobacteria in Nuwara wewa and Nachchaduwa wewa in Sri Lanka. Am. J. Environ. Prot. 4, 23–28 (2015).
Piyathilaka, M. A. P. C. et al. Microcystin-LR-induced cytotoxicity and apoptosis in human embryonic kidney and human kidney adenocarcinoma cell lines. Microbiology (Reading) 161, 819–828. https://doi.org/10.1099/mic.0.000046 (2015).
Liyanage, M. et al. Cyanobacteria and cyanotoxins in well waters of the Girandurukotte, CKDu endemic area in Sri Lanka; do they drink safe water?. J. Ecotechnol. Res. 18, 17–21. https://doi.org/10.11190/jer.18.17 (2016).
Humpage, A. R. & Falconer, I. R. Oral toxicity of the cyanobacterial toxin cylindrospermopsin in male Swiss albino mice: determination of no observed adverse effect level for deriving a drinking water guideline value. Environ. Toxicol. 18, 94–103. https://doi.org/10.1002/tox.10104 (2003).
Kubickova, B., Babica, P., Hilscherová, K. & Šindlerová, L. Effects of cyanobacterial toxins on the human gastrointestinal tract and the mucosal innate immune system. Environ. Sci. Eur. 31, 31. https://doi.org/10.1186/s12302-019-0212-2 (2019).
Yih, W. K. et al. Investigating possible infectious causes of chronic kidney disease of unknown etiology in a nicaraguan mining community. Am. J. Trop. Med. Hyg. 101, 676–683. https://doi.org/10.4269/ajtmh.18-0856 (2019).
de Figueiredo, D. R., Azeiteiro, U. M., Esteves, S. M., Gonçalves, F. J. M. & Pereira, M. J. Microcystin-producing blooms—a serious global public health issue. Ecotoxicol. Environ. Saf. 59, 151–163. https://doi.org/10.1016/j.ecoenv.2004.04.006 (2004).
Johnson, A. J. et al. A guide to diet-microbiome study design. Front. Nutr. https://doi.org/10.3389/fnut.2020.00079 (2020).
Willis, J. R. et al. Citizen science charts two major “stomatotypes” in the oral microbiome of adolescents and reveals links with habits and drinking water composition. Microbiome 6, 218–218. https://doi.org/10.1186/s40168-018-0592-3 (2018).
Barnett, J. A. & Gibson, D. L. H2Oh No! The importance of reporting your water source in your in vivo microbiome studies. Gut Microbes 10, 261–269. https://doi.org/10.1080/19490976.2018.1539599 (2019).
Tierney, B. T. et al. The predictive power of the microbiome exceeds that of genome-wide association studies in the discrimination of complex human disease. bioRxiv. https://doi.org/10.1101/2019.12.31.891978 (2020).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat. Methods 7, 335–336. https://doi.org/10.1038/nmeth.f.303 (2010).
Piper, A. M. A graphic procedure in the geochemical interpretation of water-analyses. Eos Trans. AGU 25, 914–928. https://doi.org/10.1029/TR025i006p00914 (1944).
WHO. Guidelines for Drinking-Water Quality, 4th edn. 0-564 (2011).
SLSI. Specification for Potable Water (First Revision) SLS 614:2013. (Colombo, Sri Lanka, 2013).
Li, F., Wang, M., Wang, J., Li, R. & Zhang, Y. Alterations to the gut microbiota and their correlation with inflammatory factors in chronic kidney disease. Front. Cell Infect. Microbiol. 9, 206–206. https://doi.org/10.3389/fcimb.2019.00206 (2019).
Vaziri, N. D. et al. Chronic kidney disease alters intestinal microbial flora. Kidney Int. 83, 308–315. https://doi.org/10.1038/ki.2012.345 (2013).
Xu, K.-Y. et al. Impaired renal function and dysbiosis of gut microbiota contribute to increased trimethylamine-N-oxide in chronic kidney disease patients. Sci. Rep. 7, 1445–1445. https://doi.org/10.1038/s41598-017-01387-y (2017).
Barth, J. A. C., Mader, M., Nenning, F., van Geldern, R. & Friese, K. Stable isotope mass balances versus concentration differences of dissolved inorganic carbon—implications for tracing carbon turnover in reservoirs. Isot. Environ. Health Stud. 53, 413–426. https://doi.org/10.1080/10256016.2017.1282478 (2017).
Jeffrey, A. W. A., Pflaum, R. C., Brooks, J. M. & Sackett, W. M. Vertical trends in particulate organic carbon 13C: 12C ratios in the upper water column. Deep Sea Res. Part A Oceanogr. Res. Pap. 30, 971–983. https://doi.org/10.1016/0198-0149(83)90052-3 (1983).
Martinelli, L. A. et al. Landcover changes and δ13C composition of riverine particulate organic matter in the piracicaba river basin (southeast region of Brazil). Limnol. Oceanogr. 44, 1826–1833. https://doi.org/10.4319/lo.1999.44.7.1826 (1999).
Borges, A. V. et al. Carbon cycling of Lake Kivu (East Africa): net autotrophy in the epilimnion and emission of co2 to the atmosphere sustained by geogenic inputs. PLoS ONE 9, e109500. https://doi.org/10.1371/journal.pone.0109500 (2014).
Mori, N., Kanduč, T., Opalički Slabe, M. & Brancelj, A. groundwater drift as a tracer for identifying sources of spring discharge. Ground Water 53, 123–132. https://doi.org/10.1111/gwat.12314 (2015).
Kendall, C., Silva, S. R. & Kelly, V. J. Carbon and nitrogen isotopic compositions of particulate organic matter in four large river systems across the United States. Hydrol. Process. 15, 1301–1346. https://doi.org/10.1002/hyp.216 (2001).
Gu, B. & Alexander, V. Stable carbon isotope evidence for atmospheric CO2 uptake by cyanobacterial surface scums in a eutrophic lake. Appl. Environ. Microbiol. 62, 1803–1804 (1996).
Corbel, S., Mougin, C. & Bouaïcha, N. Cyanobacterial toxins: Modes of actions, fate in aquatic and soil ecosystems, phytotoxicity and bioaccumulation in agricultural crops. Chemosphere 96, 1–15. https://doi.org/10.1016/j.chemosphere.2013.07.056 (2014).
Boopathi, T. & Ki, J.-S. Impact of environmental factors on the regulation of cyanotoxin production. Toxins (Basel) 6, 1951–1978. https://doi.org/10.3390/toxins6071951 (2014).
Balasubramanya, S., Stifel, D., Horbulyk, T. & Kafle, K. Chronic kidney disease and household behaviors in Sri Lanka: historical choices of drinking water and agrochemical use. Econ. Hum. Biol. 37, 100862. https://doi.org/10.1016/j.ehb.2020.100862 (2020).
Sidelev, S. I. & Babanazarova, O. V. Detection of cyanobacterial toxins in water supply sources and tap water in some Russian cities: searching producers and testing removal methods. Water Resour. 47, 304–314. https://doi.org/10.1134/S0097807820020189 (2020).
Haake, D. A. & Levett, P. N. Leptospirosis in humans. Cur. Top. Microbiol. Immunol. 387, 65–97. https://doi.org/10.1007/978-3-662-45059-8_5 (2015).
Peters, B. A. et al. Association of coffee and tea intake with the oral microbiome: results from a large cross-sectional study. Cancer Epidemiol. Biomark. Prev. 27, 814–821. https://doi.org/10.1158/1055-9965.EPI-18-0184 (2018).
Liu, Y.-C., Li, X.-Y. & Shen, L. Modulation effect of tea consumption on gut microbiota. Appl. Microbiol. Biotechnol. 104, 981–987. https://doi.org/10.1007/s00253-019-10306-2 (2020).
Boling, L. et al. Dietary prophage inducers and antimicrobials: toward landscaping the human gut microbiome. Gut Microbes https://doi.org/10.1080/19490976.2019.1701353 (2020).
Nicholson, J. K. et al. Host-gut microbiota metabolic interactions. Science 336, 1262. https://doi.org/10.1126/science.1223813 (2012).
Al Khodor, S. & Shatat, I. F. Gut microbiome and kidney disease: a bidirectional relationship. Pediatr. Nephrol. 32, 921–931. https://doi.org/10.1007/s00467-016-3392-7 (2017).
Hu, J. et al. Location-specific oral microbiome possesses features associated with CKD. Kidney Int. Rep. 3, 193–204. https://doi.org/10.1016/j.ekir.2017.08.018 (2017).
Yahiro, T. et al. Long-term potable effects of alkalescent mineral water on intestinal microbiota shift and physical conditioning. Evid. Based Complement Altern. Med. 2019, 2710587–2710587. https://doi.org/10.1155/2019/2710587 (2019).
Lokmer, A. et al. Response of the human gut and saliva microbiome to urbanization in Cameroon. Sci. Rep. 10, 2856–2856. https://doi.org/10.1038/s41598-020-59849-9 (2020).
Burcham, Z. M. et al. Patterns of oral microbiota diversity in adults and children: a crowdsourced population study. Sci. Rep. 10, 2133–2133. https://doi.org/10.1038/s41598-020-59016-0 (2020).
Sirisena, D. & Suriyagoda, L. D. B. Toward sustainable phosphorus management in Sri Lankan rice and vegetable-based cropping systems: a review. Agric. Nat. Resour. 52, 9–15. https://doi.org/10.1016/j.anres.2018.03.004 (2018).
Wang, Y. et al. Response of soil microbes to a reduction in phosphorus fertilizer in rice-wheat rotation paddy soils with varying soil P levels. Soil Tillage Res. 181, 127–135. https://doi.org/10.1016/j.still.2018.04.005 (2018).
de Cal, M. et al. Methylobacterium radiotolerans bacteremia in hemodialysis patients. G. Ital. Nefrol. 26, 616–620 (2009).
Truant, A. L., Gulati, R., Giger, O., Satishchandran, V. & Caya, J. G. Methylobacterium species: an increasingly important opportunistic pathogen. Lab. Med. 29, 704–710. https://doi.org/10.1093/labmed/29.11.704%JLaboratoryMedicine (1998).
Kovaleva, J., Degener, J. E. & van der Mei, H. C. Methylobacterium and its role in health care-associated infection. J. Clin. Microbiol. 52, 1317. https://doi.org/10.1128/JCM.03561-13 (2014).
Eriyagama, N., Smakhtin, V., Chandrapala, L. & Fernando, K. Impacts of Climate Change on Water Resources and Agriculture in Sri Lanka: A Review and Preliminary Vulnerability Mapping (International Water Management Institute, Colombo, 2010).
Vervaet, B. A. et al. Chronic interstitial nephritis in agricultural communities is a toxin-induced proximal tubular nephropathy. Kidney Int. 97, 350–369. https://doi.org/10.1016/j.kint.2019.11.009 (2020).
Finzer, P. How we become ill. EMBO Rep. 18, 515–518. https://doi.org/10.15252/embr.201743948 (2017).
Sturmberg, J. P. et al. Health and disease-emergent states resulting from adaptive social and biological network interactions. Front. Med. (Lausanne) 6, 59–59. https://doi.org/10.3389/fmed.2019.00059 (2019).
Lladó-Fernández, S., Větrovský, T. & Baldrian, P. The concept of operational taxonomic units revisited: genomes of bacteria that are regarded as closely related are often highly dissimilar. Folia Microbiol. 64, 19–23. https://doi.org/10.1007/s12223-018-0627-y (2019).
Kafle, K., Balasubramanya, S. & Horbulyk, T. Prevalence of chronic kidney disease in Sri Lanka: a profile of affected districts reliant on groundwater. Sci. Total Environ. https://doi.org/10.1016/j.scitotenv.2019.133767 (2019).
Acknowledgements
The authors would like to thank Rohana Chandrajith from the University of Peradeniya for various support. We would also like to thank Tharange Senevirathna and Lakruwan Alwis from the National Water Supply and Drainage Boards of Anuradhapura and Mahesh Karunarathne and Dilki Ekanayake from the University of Peradeniya for their assistance in field sampling. We thank various ANSTO personnel including Barbora Gallagher, Jennifer Van Holst, for 13CPOC, stable water isotopes and 13CDIC analyses, Chris Dimovski for assistance with fieldwork preparation and sample organization and Henri Wong for anion/cation analyses. We also thank Laurence Macia from University of Sydney for advice in microbiome analysis. This research was supported by the Australian Department of Foreign Affairs and Trading (DFAT) and Australian Nuclear Science Technology Organisation (ANSTO). We acknowledge the leadership of Dr. Adi Paterson (CEO of ANSTO, 2009-2020) who initiated ANSTO’s investigation into CKDu, water management and environmental health.
Author information
Authors and Affiliations
Contributions
Sampling design and the suite of water quality analyses were organised by C.N. and K.T.M. Field sampling was undertaken by C.N., A.V.R. and R.B.B. R.J.M. performed microbiome experiments and provided text for the methods section of the manuscript regarding the microbial analyses and post-processing of the data. K.T.M., R.J.M. and J.K.T. performed preliminary microbiome and water quality analyses. L.K.M. drafted up the main manuscript, undertook statistical associations between water quality parameters and microbiome, and prepared figures under the guidance of K.T.M. and R.B.B. R.B.B. and F.S. contributed to study design and implications for health-related studies. C.N. generated GIS files for Fig. 1 and provided input regarding the analysis of water chemistry data. All authors reviewed and provided comments on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
McDonough, L.K., Meredith, K.T., Nikagolla, C. et al. The water chemistry and microbiome of household wells in Medawachchiya, Sri Lanka, an area with high prevalence of chronic kidney disease of unknown origin (CKDu). Sci Rep 10, 18295 (2020). https://doi.org/10.1038/s41598-020-75336-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-75336-7
This article is cited by
-
Molecular Linkage of Dissolved Organic Matter in Groundwater with Prevalence of Chronic Kidney Disease with Unknown Etiology
Exposure and Health (2023)
-
Hydrogeochemical factors controlling the occurrence of chronic kidney disease of unknown etiology (CKDu)
Environmental Geochemistry and Health (2023)
-
Citizen-science reveals changes in the oral microbiome in Spain through age and lifestyle factors
npj Biofilms and Microbiomes (2022)
-
Chronic kidney disease of unknown etiology in India: a comparative study with Mesoamerican and Sri Lankan nephropathy
Environmental Science and Pollution Research (2022)
-
The influence of water–rock interactions on household well water in an area of high prevalence chronic kidney disease of unknown aetiology (CKDu)
npj Clean Water (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.