Abstract
We present the largest and most representative study of the serological diversity of Dichelobacter nodosus in England. D. nodosus causes footrot and is one of the top five globally important diseases of sheep. The commercial vaccine, containing nine serogroups, has low efficacy compared with bivalent vaccines. Our aim was to investigate the prevalence and distribution of serogroups of D. nodosus in England to elucidate whether a bivalent vaccine could protect the national flock. Farmers from 164 flocks submitted eight interdigital swabs from eight, preferably diseased, sheep. All serogroups, A–I, were detected by PCR in 687/1150 D. nodosus positive swabs, with a prevalence of 2.6–69.3% of positive swabs per serogroup. There was a median of two serogroups per flock (range 0–6). Serogroups were randomly distributed between, but clustered within, flocks, with 50 combinations of serogroups across flocks. H and B were the most prevalent serogroups, present in > 60% of flocks separately but in only 27% flocks together. Consequently, a bivalent vaccine targeting these two serogroups would protect 27% of flocks fully (if only H and B present) and partially, if more serogroups were present in the flock. We conclude that one bivalent vaccine would not protect the national flock against footrot and, with 50 combinations of serogroups in flocks, flock-specific vaccines are necessary.
Similar content being viewed by others
Introduction
Dichelobacter nodosus is a gram negative, anaerobic bacterium and the causal agent of footrot1,2,3, the principal cause of lameness in sheep in the UK4. Footrot presents as an interdigital dermatitis which sometimes progresses to severe footrot with destruction of the epidermis resulting in separation of the hoof horn from the underlying tissue5. Transmission of D. nodosus occurs between sheep via pasture6. The survival of D. nodosus in soil is influenced by soil type, temperature and moisture6. Footrot is endemic in the UK and is present in over 90% of flocks in England, with a mean prevalence of 4.5% and 3.1% for interdigital dermatitis and severe footrot lesions respectively, accounting for 68% of foot lesions4. Footrot is estimated to cost the UK sheep industry about £80 million per annum in treatments and lost production7 and is of considerable concern to sheep farmers8,9.
D. nodosus is subdivided into ten serogroups (A–I and M) which differ in their fimbrial antigens10. All ten serogroups have been detected in the UK11,12. To date the prevalence of serogroups has been reported in studies with only 34–58 non-randomly selected flocks per study and serogroups H and B have dominated11,12,13,14. It is common for several serogroups of D. nodosus to be detected concurrently in flocks and on feet in the UK, with a median of two and maximum of four serogroups per flock, with > 1 serogroup detected from 10% of sheep and 60% of flocks using culture methods12,13,14. Multiple serogroups are also detected on feet and in flocks in other countries, e.g. McPherson et al15. Previous UK studies have used culture and slide agglutination to detect serogroups, but PCR is more sensitive15 consequently the number of serogroups per flock is likely to be higher than previous estimates.
A vaccine that provides effective protection against footrot is highly desirable because it would prevent sheep from becoming lame and so improve welfare and productivity7. It would also reduce the need for treatment with antibiotics, which would significantly reduce their use in sheep farming because lameness accounts for over 60% of antibiotic use in sheep16. Despite the poor natural immune response of sheep to footrot, vaccination does initiate immunity17 to homologous serogroups18,19. Individual sheep differ in their response to the vaccine, both in the total amount of antibody produced and the relative proportion of antibodies against each serogroup in the vaccine18,20. Serogroup antigens also differ in their immunodominance20,21. Because several serogroups are often present in flocks, researchers have tested the efficacy of bivalent vaccines that target the specific serogroups in a flock22. Sequential administration of tailored mono- and bivalent vaccination over time has had some success in eliminating severe footrot from flocks in Australia23. However, as the number of serogroups present in a flock increases, increased testing and vaccination effort is required to eliminate severe footrot23. Multivalent vaccines against D. nodosus confer lower immunity per strain compared with monovalent vaccines because of antigenic competition24. Sheep vaccinated with multivalent vaccines, compared with monovalent vaccines, have significantly lower antibody titres and these decline more rapidly, therefore the sheep has both less protection and for a shorter duration24. Alternatives to a fimbrial vaccine that provide cross-protection between serogroups would be advantageous, but these have not been marketed, and research is ongoing in this area25,26. The commercial multivalent fimbrial vaccine, Footvax, is protective for up to 5 months27 and field studies indicate only partial protection28,29. In a recent observational study of 1260 flocks in England, Footvax reduced the proportional prevalence of lameness by an average of 20%, resulting in an absolute mean reduction in lameness of 1%4, indicating that vaccination might not be cost effective. Consequently, a bivalent vaccine could be of major benefit to the control of footrot in sheep in the UK if it protected a large proportion of the national flock for longer than six months with a higher magnitude of protection than Footvax.
The aims of the study were to investigate the national distribution, geographical co-location and flock prevalence of nine serogroups (A–I) of D. nodosus in England and to test the hypotheses that one bivalent vaccine could be used across the national flock, or several could be used with regional variation in the selected serogroups. We also hypothesised that serogroups in flocks were influenced by use of Footvax or biosecurity practices. Farmers were recruited from 722 compliant respondents to a previous study who were originally from a random sample of 4000 farmers4. The 164 /722 farmers who participated in the current study were representative of the original random sample30. Participants were asked to complete a questionnaire30 and send eight swabs from eight, preferably footrot-affected, feet of eight sheep.
Results
Descriptive summary of the dataset
In total 1150 swabs from 164 flocks throughout England (Dataset I; Fig. 1) were tested for load and serogroup of D. nodosus (3450 qPCR tests and 6183 PCR tests). There were 156 (95.1%) flocks with at least one sample from a footrot-affected foot and 153 (93.3%) with at least one D. nodosus positive swab (Dataset II). Flock size and prevalence of lameness and severe footrot were not associated with the number of D. nodosus positive swabs submitted (Supplementary Table 1). There were 138 (84%) flocks with 566 swabs from footrot-affected feet and 687 (59.7%) swabs positive for D. nodosus. All 687 D. nodosus positive swabs were positive for aprV2 and 74 swabs were also positive for aprB2. The median usable number of swabs per flock was 7 (range 3–8) with a median of 4 (range 0–8) D. nodosus positive swabs per flock (Table 1).
The geographical location of 164 flocks with the number of serogroups of D. nodosus detected per flock (range 0–6). The map was created in R statistical software (version 3.5.1)48.
Number of serogroups by feet and flock
Not all D. nodosus positive samples were serogroup positive, swabs were more likely to be serogroup negative when the load of D. nodosus was low (Supplementary Fig. 1). The number of serogroups per foot and flock was skewed with a median of one (range 0–4) per foot (Fig. 2) and two (range 0–6) per flock (Fig. 3). There was no significant difference (Fisher’s exact test, p = 0.55) between the observed and expected number of serogroups per foot in Dataset III when data were simulated assuming no clustering of serogroups using the 566 D. nodosus positive, footrot-affected feet, indicating that the number of serogroups per foot was randomly distributed. However, again using simulated data, the distribution of number of serogroups per flock was significantly lower (Fisher’s exact test, p ≤ 0.01) than expected by chance in the 138 D. nodosus positive flocks, indicating that serogroups were clustered within flocks (Fig. 3b). There was evidence that the number of serogroups per flock was potentially higher than that detected, especially when there were fewer than three D. nodosus positive swabs from a flock (Table 2).
The number and percentage of feet by number of serogroups detected on feet with exact binomial 95% confidence intervals (solid error bars) from (a) all 1150 feet from 164 flocks and (b) from 566 D. nodosus positive footrot-affected feet from 138 flocks with the simulated expected percentage (point) and 95% distribution of the simulated data (dashed error bars) in blue.
The number and percentage of flocks by number of serogroups detected with exact binomial 95% confidence intervals (solid error bars) from (a) all 1150 feet from 164 flocks and (b) from 566 D. nodosus and footrot positive feet from 138 flocks with the expected percentage (point) and 95% distribution of the simulated data (dashed error bars) in blue.
Prevalence and diversity of serogroups on feet and in flocks
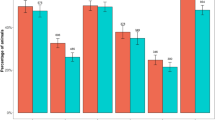
The prevalence of each serogroup varied from 2.6 to 69.3% per flock. Exact 95% confidence intervals around the point estimates indicate that serogroups H and B were distinctly the most commonly detected serogroups in > 60% of flocks (Fig. 4) and > 30% of feet (Fig. 5). This was followed by A and C and then serogroups D, E, F, G and I were detected at a similar prevalence in < 20% flocks and < 10% feet. The distribution of serogroups in flocks and on feet were similar, although there were fewer feet positive for all serogroups (Fig. 4 compared with Fig. 5) indicating that no serogroup was present in many flocks but on few feet, or vice versa. As with the number of serogroups, fewer flocks were positive for each serogroup than expected by chance (Fig. 4) indicating that serogroups were clustered within flocks. This is biologically plausible given that D. nodosus is infectious and so likely to amplify within flocks, but it might be explained in part by the limit of detection of a serogroup which was influenced by the test sensitivity31 and the number of D. nodosus positive swabs per flock (Table 2).
The number and percentage of flocks positive for each serogroup of D. nodosus with exact binomial 95% confidence intervals (solid error bars) for (a) 687 D. nodosus positive feet from 153 flocks and (b) 566 D. nodosus and footrot positive feet from 138 flocks with expected (point) and 95% distribution of the simulated data (dashed error bars) in blue.
The number and percentage of feet positive for each serogroup of D. nodosus with exact binomial 95% confidence intervals (solid error bars) for (a) 687 D. nodosus positive feet from 153 flocks and (b) 566 D. nodosus and footrot-affected feet from 138 flocks with the expected (point) and 95% distribution of the simulated data (dashed error bars) in blue.
Communities of serogroups in flocks
There were 50 combinations of serogroups across flocks. Using Raup-Crick analysis, only 7 pairs of flocks had communities of serogroups that were more different than would be expected by chance (βRC > 0.95), and there were no pairs that were more similar than would be expected by chance (βRC < − 0.95), out of 10,296 pairwise comparisons. The mean βRC for all pairwise comparisons was − 0.16 (range − 0.93 to 0.99), therefore community assembly was highly stochastic. There were no regional clusters in a principal coordinates analysis of the data (Supplementary Fig. 2). Therefore, the communities of serogroups within flocks is random, with no geographical clustering.
Multinomial model of disease state, number of swabs and biosecurity on the number of serogroups detected in flocks
A multivariable multinomial model was used to investigate factors associated with the number of serogroups detected in the 153 D. nodosus positive flocks (Dataset I). As the number of D. nodosus positive swabs per flock increased ≥ 3 serogroups were more likely to be detected than 1–2 or 0 serogroups (p < 0.01) and flocks with a stocking density of ≥ 4 ewes/acre compared with < 4 ewes/acre were more likely to have ≥ 3 serogroups than 1–2 serogroups (p = 0.017; Table 2). No other biosecurity variable was correlated with stocking density.
Probability of detection of a serogroup in a flock
When serogroups were investigated from swab data from 270 sheep from 11 flocks in a clinical trial32, when there were 16–60 swabs per flock compared with up to 8 as in Dataset I, a mean of one extra serogroup was detected (Table 3). In total, 1–3 more serogroups were identified per flock from 8 (72.7%) flocks, 1 fewer serogroup was identified in 1 (9.1%) flock, and identical serogroups were detected in 3 (27.3%) flocks (Table 3). This confirms the finding that there was probably under detection of serogroups in some flocks in the current study.
The number of swabs analysed per flock to be 95% confident of detecting a serogroup that was truly present was calculated using the observed prevalence of each serogroup. There was a 95% likelihood of detecting serogroups that were present in 32% and 53% of samples with eight and four D. nodosus positive swabs (the median number of D. nodosus positive swabs per flock in this study) respectively (Supplementary Table 2). Therefore, serogroups with < 53% prevalence would not always have been detected.
Theoretical protection offered by bivalent vaccines in England
Our results indicate that approximately 27% of flocks would be fully protected and 94% of flocks would be partially protected by a bivalent vaccine containing serogroups H and B (where full protection was because only serogroups H and B were present in the flock and partial protection was because other serogroups were in the flock). A total of 16 bivalent vaccines (out of the 36 possible combinations) against the two most common serogroups in all flocks would have fully protected 3–27% of flocks in the study and partially protected 39–94%. Sequential application of a further 2, 3, 4 and 5 bivalent vaccines in descending order of national serogroup prevalence (i.e. HB + CA + GD + IE + F) would have fully protected 65%, 83%, 97% and 100% of flocks, and partially protected 98%, 98%, 99% and 100% of flocks.
Discussion
This is the largest and most representative study of the national prevalence and diversity of serogroups of D. nodosus in England to date. Key findings are that serogroups A–I are present and that the prevalence of serogroups varies. Serogroups are randomly distributed across England, clustered within flocks but not geographically co-located, with 50 combinations of serogroups across flocks, therefore one bivalent vaccine would not control footrot nationally.
The national prevalence of each serogroup varied considerably (Fig. 5) with three distinct prevalence groups, there is 95% probability of a true difference in the prevalence of B and H, A and C and then the remaining less prevalent serogroups. Serogroups H and B have been reported as the two most prevalent serogroups in the UK previously as far back as 30 years11,14, so the prevalence of each serogroup appears reasonably stable nationally over time. The high prevalence of serogroup B is also consistent with other countries where it is often the most prevalent serogroup33,34,35,36,37,38, however, serogroup H is not reported to be the most common serogroup in any other country. Despite the high prevalence of serogroups H and B a bivalent vaccine would protect only 27% of flocks in England and we conclude that one bivalent vaccine would not be effective in protecting the national flock.
The strength of the current study is the number of flocks investigated nationally which provides the robust estimates of the prevalence of serogroups. The percentage of flocks and feet with multiple serogroups and the maximum number of serogroups per flock are higher than reported in previous UK studies12,13,14,39 this is probably partly due to the increased sensitivity of PCR directly from DNA without culture15. Despite this, the number of serogroups detected in flocks is likely to be underestimated. Hill et al.40 calculated that over 40 swabs were required for a 95% chance of identifying all serogroups in Australian flocks, however this is likely to be lower with the smaller size of flocks in England, and with the increased sensitivity of the methods used here. If the aim is eradication of footrot by flock-specific vaccination, more intensive strategies are required to ensure that no serogroups are omitted. The farmers were not requested to ensure that the sheep that they took swabs from were representative of the flock, however flocks are typically kept in one group in the UK so it is expected that most samples were representative of the flock. Figure 3 indicates a slight left shift in distribution of observed than expected number of serogroups and that flocks would have approximately one more serogroup than observed. This was probably because some flocks submitted < 4 D. nodosus positive swabs which was associated with fewer serogroups detected (Table 2). The number of serogroups per swab might also be underestimated because of the limit of detection of the PCR test and because there is no specific PCR primer for serogroup M so it was not possible to estimate the prevalence of M. There is no reason to expect serogroup M to behave differently from other serogroups and so we would expect it to be randomly distributed across England, clustered within flocks and to increase the number of serogroup combinations further.
Few studies have investigated geographical distribution of serogroups in England11,13 and there is little published evidence of geographical co-location of serogroups in other countries, except possibly in India41, where breeds and managements are very different by region and fewer serogroups have been detected than in England and there is some evidence that the prevalence of serogroups is geographically clustered. In the current study, between flock serogroup communities were highly stochastic, with 50 combinations of serogroups detected in flocks and no clusters of flocks with more similar communities of serogroups, even in flocks within the same region (Supplementary Fig. 2) no two flocks had communities of serogroups that were more similar than would be expected by chance. Therefore, no bivalent candidate vaccines could protect flocks grouped geographically.
Lack of geographical clustering of serogroups indicates that serogroups of D. nodosus are randomly distributed across England. Footrot has been reported in the UK for over 200 years42 and over 90% of flocks have footrot4. D. nodosus persists on feet6 and so it seems likely that movement of infected sheep and poor biosecurity between flocks has contributed to its highly endemic profile. There is evidence for this in the current study where most flocks were open, 83.5% of the 164 farmers purchased sheep, often from distant locations30 and only 50% quarantined sheep for at least 3 weeks30, and less than 25% treated sheep with footrot or interdigital dermatitis on arrival4. It is also possible that within flock seroconversion of a serogroup occurred, this has been demonstrated in the laboratory43 and possibly in a UK flock39. Although the continuing dominance of serogroups H and B indicate national stability, the current study was limited to a single timepoint on each farm and so we could not determine stability of serogroups within flocks over time. Some longitudinal studies have reported that new serogroups ‘appear’ in flocks, although it is not possible to determine whether this is due to limits of detection or targeted vaccination altering the dominance of serogroups within a flock, or seroconversion23,39.
Flocks that were stocked at a density of ≥ 4 ewes/acre were more likely to have ≥ 3 serogroups detected than the 1–2 serogroups detected in less densely stocked flocks. This is a novel finding. This could be a true difference in number of serogroups in those flocks or it might be explained by increased transmission of D. nodosus between sheep within a densely stocked flock that increases the prevalence of rarer serogroups within a flock and therefore that they were more likely to be detected in the current study. Vaccination with Footvax, the nine-serogroup commercial vaccine, and biosecurity managements were not associated with number of serogroups or specific serogroups.
We conclude that the lack of dominance of the same two serogroups in flocks nationally or regionally make one national or a few regional bivalent vaccines unfeasible. With 50 communities of serogroups across 164 flocks we conclude that flock-specific bivalent vaccines would be essential.
Materials and methods
This project was carried out according to approval from The University of Warwick Biomedical and Scientific Research Ethics Committee (REGO-2016-1758 AMO1) and Animal Welfare and Ethical Review Body (AWERB.24/15-16). Approval was granted on 23/02/16 and 08/04/16 respectively. All experiments were performed in accordance with relevant guidelines and regulations.
Selection of farmers to the study
An invitation letter and reply slip were posted to 722 farmers in England in February and April 2016. These farmers had completed detailed questionnaires on lameness in their flock in 20134, and also, for the majority, in 201444, and indicated that they were willing to participate in further research. Replies were received between 09/02/2016–30/06/2016 from 192/722 (26.6%) farmers who agreed to participate. An additional 18 farmers who had participated in the original study4 and were participating in a separate clinical trial32 were recruited. There was no difference in the proportion of D. nodosus positive swabs or the number of serogroups detected from the swabs from the surveyed farms and the clinical trial (Table 4), so data were combined and are presented together.
Swab sample and questionnaire collection
Farmers participating by post were sent eight sterile swabs with charcoal amies gel transport medium (Thermo Scientific) and illustrated instructions on how to swab the feet31. Participants were asked to swab the interdigital skin of one foot of each of eight sheep ideally lame with footrot, that had not received treatment in the previous 2 weeks and to complete the sampling with healthy sheep if eight lame sheep were not present. They were not given any further instructions on which sheep to swab. Farmers recorded all lesions on the swabbed foot. Swabs were posted immediately at ambient temperature or were refrigerated overnight and arrived within 24 h of posting. On arrival, swabs, together with the amies medium, were immediately transferred to − 20 °C storage. Swabs were received between 02/03/16 and 08/09/16 from 146 farmers. The farmers who returned swabs also returned a completed three-page questionnaire regarding their flock in 2015. The 2015 questionnaire included questions on the presence and prevalence of four foot lesions, treatment and control managements for footrot, biosecurity practices, and some information about the flock in 2015.
The 18 flocks in the clinical trial had swabs from the interdigital skin of up to 15 lame sheep collected between November 2015 and August 2016. Eight swabs per flock were selected for the current project using the same criteria as for the farmers contacted by post. Sheep were selected in the following order of preference: lame with severe footrot or interdigital dermatitis (locomotion score ≥ 245), lame with another lesion (locomotion score ≥ 2), not lame (locomotion score < 2). Feet were selected in the following order of preference: severe footrot lesion, interdigital dermatitis lesion, another lesion, no lesion. Swabs were stored in phosphate buffered saline. The liquid was spun off in a centrifuge and split into two aliquots, with one aliquot used for this project. The responses to the questionnaire for the farmers participating in the clinical trial32 were retrieved from a more detailed questionnaire conducted as part of that study.
DNA extraction from swabs
Swabs were randomised for processing by assigning a random number generated in Microsoft Excel 2016 stratified by flock. DNA extraction was performed using a Nucleospin® Tissue kit (Macherey–Nagel) with modifications. Swabs were thawed at room temperature, transferred to microcentrifuge tubes and lysis buffer T1 (400 μl) and proteinase K (40 μl) were added. Samples were vortexed for 5 s and incubated for 10 min at 56 °C. Post-incubation, lysis buffer B3 (400 μl) was added. Each sample was vortexed for 5 s and incubated at 70 °C for 5 min. Samples were cooled at room temperature for 5 min before being centrifuged at 12,000g for 1 min. The supernatant was added to 100% ethanol (400 μl) and centrifuged at 11,000g for 1 min and then loaded onto spin columns and centrifuged at 11,500g for 1 min. Flow-through was discarded and BW wash buffer (500 μl) was added to the spin columns and centrifuged at 11,000g for 1 min. Flow-through was discarded and B5 wash buffer (600 μl) was added to the spin columns and centrifuged at 11,000g for 1 min. Flow-through was discarded and spin columns were centrifuged for 1 min at 11,000g to dry the membrane. Spin columns were placed in microcentrifuge tubes and BE elution buffer (45 μl) heated to 70 °C was added and allowed to stand for 2 min. Tubes were spun at 11,000g for 1 min to elute DNA and extracted DNA was stored at − 20 °C.
Screening for Dichelobacter nodosus with qPCR
To identify which samples contained D. nodosus and could therefore be tested for presence of each serogroup, each DNA sample was tested in triplicate using a qPCR that detected the aprV2/aprB2 gene using the method published by Frosth et al.46 with some modifications. qPCR tests were used because they are more sensitive than the PCR tests. Modifications were made to the probe labelling (aprV2probe 6FAM-BHQ1, aprB2probe TxRd-BHQ2), Klearkall master mix (LGC Group) was used and only 1 μl of template DNA was used. The initial denaturation step was extended to 15 min as required by the master mix. D. nodosus strain VCS1703A was the positive control for aprV2, and strain C305 was the positive control for aprB2 (Supplementary Table 3). Each qPCR plate contained a no template control in triplicate to differentiate noise from true positives and in addition, only samples that were 3/3 positive for either aprV2 or aprB2 were classed as positive which was a conservative way of defining positive samples and limiting the risk of false positive results. Total D. nodosus load in a sample was calculated as the sum of the number of genome copies of aprB2 and aprV2. Most samples that were positive for D. nodosus in only 1 or 2 of the 3 replicate qPCR tests had low loads.
Detection of serogroups from DNA using PCR
Samples positive for D. nodosus by qPCR were tested separately for nine serogroups (A–I) of D. nodosus using single serogroup PCRs. Serogroup M was not investigated because there is no published PCR test. Multiplex serogroups PCRs were tested but these were less sensitive31. The primers and program were published by Dhungyel et al.47. Each reaction (25 μl) contained MyTaq Red Mix (Bioline) (12.5 μl), forward primer (0.5 μM), reverse primer (0.5 μM) and bovine serum albumin (0.5 mg/ml). Products were electrophoresed on 3% agarose gels stained with ethidium bromide and visualised under ultraviolet light. Positive controls were normalised to 10 ng/μl and are in Supplementary Table 3.
Data entry and analysis
Data was entered manually into Microsoft Excel and rechecked once to ensure accuracy. All cases where farmers did not answer the question were coded as No Response. There was no difference in the number of serogroups detected per swab between the post and clinical trial farmers (Table 4). Data analysis was conducted in R statistical software (version 3.5.1)48. The significance level used for all tests was 0.05. Kruskal–Wallis tests were used to investigate whether flock size, the prevalence of lameness or the prevalence of severe footrot in ewes, were associated with the number of swabs submitted and the number of swabs that were D. nodosus positive per flock because this data was not normally distributed.
Detection of serogroups of Dichelobacter nodosus by swab and flock
Three datasets were created:
-
(I)
1150 swabs from 164 flocks.
-
(II)
566 D. nodosus positive swabs from footrot-affected feet from 138 flocks.
-
(III)
A randomly simulated dataset of Dataset II.
For simulated Dataset III, the proportion of swabs positive for each serogroup in Dataset II was calculated. Each swab was then assigned as positive or negative for each serogroup (A–I) weighted by the observed proportion of swabs positive. Only swabs from footrot-affected feet were simulated to remove the heterogeneity in probability of detection of D. nodosus from other lesions, and because the majority of D. nodosus positive swabs were from footrot-affected feet (Table 1). The simulation from the observed data was run 1000 times assuming a random distribution of serogroups present within and between flocks. Differences in the number of serogroups in feet and flocks between the observed and simulated data were tested with Fisher’s exact tests. Post hoc analysis of chi-square and Fisher’s exact tests were conducted with the fifer (version 1.1)49 and rcompanion (version 2.1.1)50 packages respectively.
Multinomial model of disease state, number of swabs and biosecurity managements on the number of serogroups detected
Unordered multinomial logistic regression models51 with ≥ 3, 1–2, or 0 serogroups per flock were used to investigate whether the number of swabs submitted/analysed and biosecurity managements in the flock were associated with the number of serogroups detected per flock. The nnet package (version 7.3-12)52 was used. The univariable model results of variables considered for inclusion in the model are in Supplementary Tables 4 and 5. The models were built using a manual forward stepwise procedure by sequentially testing each term using variables with the lowest Akaike’s Information Criterion (AIC) at each iteration. Variables were retained in the final multivariable models if p ≤ 0.05 and > 1 flock was in each significantly different category (Supplementary Table 5). All remaining variables were retested in the final model53. Chi-square tests and Fisher’s exact tests were used to test for correlations between the biosecurity terms in the final model and all other tested biosecurity terms. The models took the form:
where \(logit({\pi }_{1k}/{p}_{i0k})\) is the probability of having 1–2 serogroups detected versus ≥ 3 and \(logit({\pi }_{2k}/{p}_{i0k})\) is the probability of having 0 serogroups detected versus ≥ 3, \({\beta }_{0k}\) and \({\beta }_{1k}\) are constants for 1–2 serogroups and 0 serogroups, \({\beta }_{0}X\) and \({\beta }_{1}X\) are coefficients for number of swabs and biosecurity variables X for 1–2 serogroups and 0 serogroups, and \({e}_{k}\) is the residual random error that follows a binomial distribution.
Spatial analysis of the diversity of serogroups of Dichelobacter nodosus in England
Raup-Crick analysis of community diversity was conducted using null models following the method described by Chase et al.54. Principal coordinates analysis55 of the Raup–Crick analysis was used to visually assess for geographical clusters using the ape package (version 5.2)56.
Probability of detection of a serogroup from the swabs
In the current study there were up to eight swabs per flock. For 11 of the clinical trial flocks, > 8 swabs (range 16–60) were pooled and the number of serogroups per pooled sample was analysed in a separate study (Monaghan et al., unpublished data). For these flocks each foot of 15 sheep was swabbed and pooled into one of ten footrot disease severity states (Supplementary Material 1). The presence of each serogroup was investigated using the same PCR method as used in this study. The number of serogroups detected per flock when eight (the current study) and more than eight swabs (clinical trial) were collected were calculated to investigate whether the number of serogroups detected per flock was associated with the number of D. nodosus positive swabs analysed.
Prevalence and probability of detection of a serogroup
In order to calculate the probability of detecting common and rare serogroups in a flock, the minimum numbers of swabs required to have a 95% probability of detection of serogroups by prevalence X was calculated using the formula:
where n is the number of swabs.
Data availability
The datasets supporting the conclusions of this study are available on request.
References
Beveridge, W. I. B. Foot-Rot in Sheep: A Transmissable Disease due to Infection with Fusiformis nodosus (n. sp.) Studies on its Cause, Epidemiology and Control, Vol. 140 (Commonwealth of Australia Council for Scientific and Industrial Research, Canberra, 1941).
Kennan, R. M., Dhungyel, O. P., Whittington, R. J., Egerton, J. R. & Rood, J. I. The type IV fimbrial subunit gene (fimA) of Dichelobacter nodosus is essential for virulence, protease secretion, and natural competence. J. Bacteriol. 183, 4451–4458 (2001).
Witcomb, L. A. et al. A longitudinal study of the role of Dichelobacter nodosus and Fusobacterium necrophorum load in initiation and severity of footrot in sheep. Prev. Vet. Med. 115, 48–55 (2014).
Winter, J. R., Kaler, J., Ferguson, E., KilBride, A. L. & Green, L. E. Changes in prevalence of, and risk factors for, lameness in random samples of English sheep flocks: 2004–2013. Prev. Vet. Med. 122, 121–128 (2015).
Egerton, J. R., Roberts, D. S. & Parsonson, I. M. The aetiology and pathogenesis of ovine foot-rot I. A histological study of the bacterial invasion. J. Comp. Pathol. 79, 207–216 (1969).
Clifton, R., Giebel, K., Liu, N. L. B. H., Purdy, K. J. & Green, L. E. Sites of persistance of Fusobacterium necrophorum and Dichelobacter nodosus: a paradigm shift in understanding the epidemiology of footrot in sheep. Sci. Rep. 9, 14429. https://doi.org/10.1038/s41598-019-50822-9 (2019).
Wassink, G. J. et al. A within farm clinical trial to compare two treatments (parenteral antibacterials and hoof trimming) for sheep lame with footrot. Prev. Vet. Med. 96, 93–103 (2010).
Goddard, P., Waterhouse, T., Dwyer, C. & Stott, A. The perception of the welfare of sheep in extensive systems. Small Rumin. Res. 62, 215–225 (2006).
O’Kane, H., Ferguson, E., Kaler, J. & Green, L. Associations between sheep farmer attitudes, beliefs, emotions and personality, and their barriers to uptake of best practice: the example of footrot. Prev. Vet. Med. 139, 123–133 (2017).
Claxton, P. D., Ribeiro, L. A. & Egerton, J. R. Classification of Bacteroides nodosus by agglutination tests. Aust. Vet. J. 60, 331–334 (1983).
Thorley, C. M. & Day, S. E. J. In Footrot in Ruminants, Proceedings of a Workshop (eds Stewart, D. J. et al.) 135–142 (CSIRO Australia and Australian Wool Corporation, Melbourne, 1986).
Hindmarsh, F. & Fraser, J. Serogroups of Bacteroides nodosus isolated from ovine footrot in Britain. Vet. Rec. 116, 187–188 (1985).
Kingsley, D. F., Hindmarsh, F. H., Liardet, D. M. & Chetwin, D. H. in Footrot in Ruminants: Proceedings of a Workshop (CSIRO Division of Animal Health, 1986).
Moore, L. J., Wassink, G. J., Green, L. E. & Grogono-Thomas, R. The detection and characterisation of Dichelobacter nodosus from cases of ovine footrot in England and Wales. Vet. Microbiol. 108, 57–67 (2005).
McPherson, A. S., Dhungyel, O. P. & Whittington, R. J. Detection and serogrouping of Dichelobacter nodosus infection by use of direct PCR from lesion swabs to support outbreak-specific vaccination for virulent footrot in sheep. J. Clin. Microbiol. 56, e01730-01717. https://doi.org/10.1128/jcm.01730-17 (2018).
Davies, P. et al. Quantitative analysis of antibiotic usage in British sheep flocks. Vet. Rec. 181, 1–7 (2017).
Egerton, J. R. & Burrell, D. H. Prophylactic and therapeutic vaccination against ovine foot-rot. Aust. Vet. J. 46, 517–522 (1970).
Hunt, J. D., Jackson, D. C., Brown, L. E., Wood, P. R. & Stewart, D. J. Antigenic competition in a multivalent foot rot vaccine. Vaccine 12, 457–464 (1994).
Stewart, D. J. The role of various antigenic fractions of Bacteroides nodosus in eliciting protection against foot-rot in vaccinated sheep. Res. Vet. Sci. 24, 14–19 (1978).
Raadsma, H. W., Omeara, T. J., Egerton, J. R., Lehrbach, P. R. & Schwartzkoff, C. L. Protective antibody titres and antigenic competition in multivalent Dichelobacter nodosus fimbrial vaccines using characterised rDNA antigens. Vet. Immunol. Immunopathol. 40, 253–274 (1994).
Dhungyel, O. P. & Whittington, R. J. Modulation of inter-vaccination interval to avoid antigenic competition in multivalent footrot (Dichelobacter nodosus) vaccines in sheep. Vaccine 28, 470–473 (2010).
Egerton, J. R. Surface and somatic antigens of Fusiformis nodosus. J. Comp. Pathol. 83, 151–159 (1973).
Dhungyel, O. et al. Outbreak-specific monovalent/bivalent vaccination to control and eradicate virulent ovine footrot. Vaccine 31, 1701–1706 (2013).
Schwartzkoff, C. L. et al. The effects of antigenic competition on the efficacy of multivalent footrot vaccines. Aust. Vet. J. 70, 123–126 (1993).
Humbert, M. V., Jackson, A., Orr, C. M., Tews, I. & Christodoulides, M. Characterization of two putative Dichelobacter nodosus footrot vaccine antigens identifies the first lysozyme inhibitor in the genus. Sci. Rep. 9, 10055. https://doi.org/10.1038/s41598-019-46506-z (2019).
Myers, G. S. A. et al. Genome sequence and identification of candidate vaccine antigens from the animal pathogen Dichelobacter nodosus. Nat. Biotechnol. 25, 569–575 (2007).
MSD Animal Health. Footvax. https://www.msd-animal-health.co.uk/products_public/footvax/overview.aspx.
Allworth, M. B. & Egerton, J. R. Comparison of footbathing and vaccination to control ovine footrot in an experimentally infected flock. Aust. Vet. J. 96, 395–399 (2018).
Duncan, J. S. et al. Impact of footrot vaccination and antibiotic therapy on footrot and contagious ovine digital dermatitis. Vet. Rec. 170, 462 (2012).
Prosser, N. S., Purdy, K. J. & Green, L. E. Increase in the flock prevalence of lameness in ewes is associated with a reduction in farmers using evidence-based management of prompt treatment: A longitudinal observational study of 154 English sheep flocks 2013–2015. Prev. Vet. Med. 173, 104801. https://doi.org/10.1016/j.prevetmed.2019.104801 (2019).
Prosser, N. S. The prevalence of lameness and diversity of serogroups of Dichelobacter nodosus in sheep in England. Ph.D. thesis, University of Warwick (2019).
Witt, J. & Green, L. Development and assessment of management practices in a flock-specific lameness control plan: a stepped-wedge trial on 44 English sheep flocks. Prev. Vet. Med. 157, 125–133 (2018).
Gurung, R. B., Tshering, P., Dhungyel, O. P. & Egerton, J. R. Distribution and prevalence of footrot in Bhutan. Vet. J. 171, 346–351 (2006).
Best, N., Gwozdz, J., Suter, R., Rawlin, G. & Beddoe, T. Direct serogrouping of Dichelobacter nodosus from Victorian farms using conventional multiplex polymerase chain reaction. BMC Res. Notes 11, 108. https://doi.org/10.1186/s13104-018-3229-5 (2018).
Olson, M. E., Card, M. S., Gradin, J. & Morck, D. W. Serological classification and virulence determination of Dichelobacter nodosus isolated from Alberta and British Columbia sheep. Can. J. Vet. Res. Rev. Can. De Rech. Vet. 62, 33–37 (1998).
Wani, S. A. et al. Determination of prevalence, serological diversity, and virulence of Dichelobacter nodosus in ovine footrot with identification of its predominant serotype as a potential vaccine candidate in J&K, India. Trop. Anim. Health Prod. 51, 1089–1095 (2019).
Zhou, H. & Hickford, J. G. H. Extensive diversity in New Zealand Dichelobacter nodosus strains from infected sheep and goats. Vet. Microbiol. 71, 113–123 (2000).
Gradin, J. L., Sonn, A. E. & Petrovska, L. Serogrouping of Bacteroides nodosus isolates from 62 sources in the United States. Am. J. Vet. Res. 54, 1069–1073 (1993).
Smith, E. M. et al. Within-flock population dynamics of Dichelobacter nodosus. Front. Vet. Sci. 4, 58. https://doi.org/10.3389/fvets.2017.00058 (2017).
Hill, A. E., Dhungyel, O. P. & Whittington, R. J. Diagnostic sampling strategies for virulent ovine footrot: simulating detection of Dichelobacter nodosus serogroups for bivalent vaccine formulation. Prev. Vet. Med. 95, 127–136 (2010).
Sreenivasulu, D. et al. Prevalence of ovine footrot in the tropical climate of southern India and isolation and characterisation of Dichelobacter nodosus. Rev. Sci. et Tech. Off. Int. des Epizoot. 32, 869–877 (2013).
Peall, T. A Treatise on the Foot-Rot in Sheep (Royal Dublin Society, Leinster, 1822).
Kennan, R. M., Dhungyel, O. P., Whittington, R. J., Egerton, J. R. & Rood, J. I. Transformation-mediated serogroup conversion of Dichelobacter nodosus. Vet. Microbiol. 92, 169–178 (2003).
Grant, C., Kaler, J., Ferguson, E., O’Kane, H. & Green, L. E. A comparison of the efficacy of three intervention trial types: postal, group, and one-to-one facilitation, prior management and the impact of message framing and repeat messages on the flock prevalence of lameness in sheep. Prev. Vet. Med. 149, 82–91 (2018).
Kaler, J., Wassink, G. J. & Green, L. E. The inter- and intra-observer reliability of a locomotion scoring scale for sheep. Vet. J. 180, 189–194 (2009).
Frosth, S., König, U., Nyman, A.-K., Pringle, M. & Aspán, A. Characterisation of Dichelobacter nodosus and detection of Fusobacterium necrophorum and Treponema spp. in sheep with different clinical manifestations of footrot. Vet. Microbiol. 179, 82–90 (2015).
Dhungyel, O. P., Whittington, R. J. & Egerton, J. R. Serogroup specific single and multiplex PCR with pre-enrichment culture and immuno-magnetic bead capture for identifying strains of D. nodosus in sheep with footrot prior to vaccination. Mol. Cell. Probes 16, 285–296 (2002).
R Core Team. R: A language and environment for statistical computing (R Foundation for Statistical Computing, Vienna, 2017).
Fife, D. A Biostatisticians Toolbox for Various Activities, Including Plotting, Data Cleanup, and Data Analysis v. 1.1 (2017).
Magniafico, S. rcompanion: Functions to Support Extension Education Program Evaluation v. 2.1.1 (2019).
Dohoo, I., Martin, W. & Stryhn, H. Veterinary Epidemiologic Research 1st edn, 126–130 (AVC Inc, Naagarabhaavi, 2003).
Venables, W. N. & Ripley, B. D. Modern Applied Statistics with S 4st edn (Springer, New York, 2002).
Cox, D. R. & Wermuth, N. Multivariate Dependencies: Models, Analysis and Interpretation (Chapman & Hall, London, 1996).
Chase, J. M., Kraft, N. J. B., Smith, K. G., Vellend, M. & Inouye, B. D. Using null models to disentangle variation in community dissimilarity from variation in alpha-diversity. Ecosphere 2, art24. https://doi.org/10.1890/ES10-00117.1 (2011).
Gower, J. C. Some distance properties of latent root and vector methods used in multivariate analysis. Biometrika 53, 325–338 (1966).
Paradis, E. & Schliep, K. ape 5.0: an environment for modern phylogenetics and evolutionary analyses in R. Bioinformatics 35, 526–528 (2018).
Acknowledgements
Naomi Prosser was funded by a BBSRC CASE studentship with AHDB. Laura Green and Kevin Purdy were funded by BBSRC grant BB/M012980/1. The authors thank the participating farmers for submitting swabs and completing the questionnaires, Dr. Zoë Willis for illustrating the farmer swabbing instructions, Jessica Witt for contributing questionnaire data, Prof. Matt Keeling and Dr. Chris O’Grady for statistical advice, and Dr. Claire Russell (Bristol University), and Prof. Julian Rood and Dr. Ruth Kennan (Monash University) for providing D. nodosus isolates.
Author information
Authors and Affiliations
Contributions
N.P. recruited farmers to the study and collected the samples and questionnaires and carried out the laboratory and data analysis. K.P. and L.G. devised and supervised the project and advised on methods, data analysis and discussion. E.M. contributed swabs from the clinical trial, analysed the pooled clinical trial swabs and advised on laboratory methods. N.P. drafted the manuscript, K.P. and L.G. reviewed and revised the manuscript, the final version was approved by all authors.
Corresponding author
Ethics declarations
Competing of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Prosser, N.S., Monaghan, E.M., Green, L.E. et al. Serogroups of Dichelobacter nodosus, the cause of footrot in sheep, are randomly distributed across England. Sci Rep 10, 16823 (2020). https://doi.org/10.1038/s41598-020-73750-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-73750-5
This article is cited by
-
Differences in composition of interdigital skin microbiota predict sheep and feet that develop footrot
Scientific Reports (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.