Abstract
Two species breeding in sympatry are more likely to coexist if their ecological niches are segregated either in time, space or in trophic habits. Here, we combined GPS-tracking, stable isotope analysis and DNA metabarcoding analysis to understand how the rare Tahiti petrel Pseudobulweria rostrata (TP) copes with the very abundant (i.e. 500,000 breeding pairs) wedge-tailed shearwater Ardenna pacifica (WTS) when breeding in sympatry in a tropical area. WTS foraged in restricted areas along their path, while TP predominantly foraged using extensive search behavior, suggesting a more opportunistic foraging strategy. Interspecific overlap of foraging areas was higher than intraspecific overlap. Breeding seasons largely overlap between species during the study, but TP seems to be asynchronous breeders. TP fed upon prey with higher δ15N values than WTS, and their diet was mainly composed of deep-sea organisms. TP could feed upon dead prey floating at the surface while WTS preyed mainly upon fish species that generally move in schools. Our study highlights several segregating mechanisms (temporal, behavioral and trophic) that could facilitate the coexistence of the two species despite the predominant number of WTS, and provides the very first information on the foraging and trophic ecology of the poorly-known TP.
Similar content being viewed by others
Introduction
The theory of ecological segregation postulates that coexisting species may partition their use of resources, in either time, space or trophic habits to avoid or limit competition, leading to niche divergence1. Among marine predators, seabirds provide good examples of ecologically similar coexisting species, sometimes breeding in sympatry and sharing foraging areas2,3. Inter- and intra-specific competition in seabirds are supposedly higher in tropical areas where marine productivity tends to be lower4 with a more patchy distribution of resources compared to temperate and polar areas5. It is particularly true during chick-rearing when breeding adults acts as central place foragers6. Competition for food resources is therefore expected to be particularly high for tropical seabirds during the breeding season.
The foraging ecology of Procellariiform seabirds (i.e. albatrosses, petrels and shearwaters) has been widely studied thanks to the development of tracking technologies such as miniaturised GPS loggers (e.g. 7,8,9). However, tropical regions remain overall poorly studied, despite being hotspots of seabird species richness10. Tracking data constitute a valuable tool to identify foraging areas, and to characterize foraging trips and at-sea behavior of birds. Information on seabird at-sea movements and behavior can be completed by complementary techniques such as stable isotope analyses (SIA), used to depict trophic ecology (e.g. 11). Carbon stable isotopes (13C/12C) provide information on the feeding habitat and resource use because they reflect the primary carbon sources within a food web12, while nitrogen stable isotopes (15N/14N), showing a stepwise enrichment at each trophic level, are used to estimate the trophic position13. Isotopic niche width (i.e. the isotopic composition of animal tissues in a multivariate space) is a powerful tool to investigate the ecological niche of the species studied14. At the population level, a wide isotope niche is typical of a generalist population, while a narrower isotope niche reveals a species specializing on a more specific trophic level and/or habitat. Additionally, SIA allow the assessment of temporal isotope variance among individuals by carefully selecting tissues with appropriate turn-overs, and examining the consistency of the isotope values among them15. Generalist individuals vary in their resource use, resulting in a wide isotopic niche of the population. However, specialist individuals have a consistent use of resources but variation among individuals would also result in a wide population isotopic niche. Finally, the prevalence of specialist individuals and low variation among individuals result in a specialist population, with a narrow isotope niche. Therefore, individual variation in resource use may influence the population dynamics and ecological interactions within and between species16. However, SIA alone provides a limited understanding of real trophic interactions, not allowing the proper identification of prey. Complementary molecular analyses such as DNA metabarcoding17 have been widely used in recent years to precisely investigate the diet of seabirds, including Procellariiform species, from feces or regurgitate samples. It allows for a semi-quantitative estimation of the food items18,19,20.
The Tahiti petrel (Pseudobulweria rostrata, hereafter TP) and the wedge-tailed shearwater (Ardenna pacifica, hereafter WTS), are two similar-sized Procellariid species breeding in the tropics, sometimes in sympatry. The WTS ranges throughout the Indian and Pacific Oceans, between latitudes 35°N and 35°S21. It is very abundant in the tropical Pacific22. WTS has been shown to use a bi-modal foraging strategy during chick-rearing, alternating a series of short trips close to the colony with longer trips over distant areas when surrounded by low-productivity environments6,8,23,24, supposedly in response to high competition for resources25,26. However, WTS can shift to a unimodal strategy when breeding in richer environments27,28,29. WTS is known to forage in multi species flocks, in association with sub-surface predators such as yellowfin (Thunnus albacares) and skipjack (Katsuwonus pelamis) tuna, which make epi- and meso-pelagic prey available at the surface30,31,32. WTS feed mostly upon cephalopods and fish usually by surface feeding, but can dive up to 66 m deep, with an average of 5–14m22,33,34. In the Southwestern Pacific, the breeding season of the species begins at the end of October with the return of breeding adults to colonies. Adults lay their eggs in December, and chicks fledge at the end of May.
On the other hand, the TP is a poorly known species since it is rarer, and has been poorly studied. Its population sizes are generally imprecise and speculative, estimated between 10.000 and 20.000 mature individuals worldwide22,35. This species is known to breed in French Polynesia, Fiji, American Samoa and New Caledonia (France)35. As TP movements at sea have never been tracked until this study, their foraging ecology is largely unknown. TP diet is suspected to be mostly composed of fish and cephalopods32. This species is an asynchronous breeder, with laying occurring throughout the year, but peaking at various periods of the year depending on the geographical area considered36. When breeding in sympatry, TP and WTS compete fiercely for nests36.
Here, we aim at understanding how the rare TP copes with the much more abundant WTS when breeding and foraging around New Caledonia. For this purpose, we combined GPS tracking data, SIA on blood samples, and DNA metabarcoding on regurgitate samples to depict their foraging and trophic ecology. We hypothesized that either temporal, spatial or trophic segregation would exist to reduce inter-species competition, especially during chick-rearing when the competition is likely to be maximum. If a temporal segregation occurs, we expect both species to have different breeding seasons, and/or to forage at different periods of the day. A spatial segregation would imply at least low overlap of core foraging areas while a trophic segregation would induce small isotopic niche overlap, and a difference in prey composition identified by DNA metabarcoding.
Material and methods
Study area
This study took place in New Caledonia, in the South-west Pacific, which is located in an oligotrophic area with low nutrient and low primary production. This area exhibits a large-scale north–south gradient, with salinity and temperature decreasing from north to south38.
New Caledonia is home of 500,000 WTS breeding pairs nest mostly on sandy islets while TP breeding pairs are rare and scarcely distributed. Indeed, it was tentatively estimated that 1000 TP breeding pairs were distributed across the ~ 300 km-long mountain ranges of the main island39 (Grande Terre) but this figure remains poorly supported by field data. In addition, 11 out of 70 islets surveyed in the southern lagoon houses a total of less than 100 TP breeding pairs40.
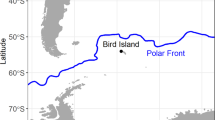
Study colonies are located on three lagoon islets situated off the southern part of Grande Terre (Fig. 1): Mato (22.55°S, 166.80°E), Canard (22.31°S, 166.31°E), and Nemou (20.38°S, 164.04°E). Mato and Canard are two close islets situated off the South-west coast of Grande Terre, while Nemou Islet is located off the South-east coast. Mato Islet hosts 2000 WTS and 20 TP breeding pairs. Canard Islet hosts 340 WTS breeding pairs. On Nemou Islet, 124 TP breeding pairs were recently censused (unpublished data).
Location of the three study sites in New Caledonia: Nemou islet, a Tahiti petrel (TP) colony, Canard islet, a wedge-tailed shearwater (WTS) colony, and Mato, where WTS and TP breed in sympatry. The insert shows the position of New Caledonia in the South West Pacific. Loyalty Islands are visible at the top of the map. Bathymetry map was obtained from https://carto.gouv.nc/arcgis/services/fond_relief/MapServer/WMSServer. The map was created using QGIS version 2.1841 (URL: https://qgis.org/).
Field work
GPS-loggers were fitted on breeding adults during the chick-rearing period at the three study sites (Fig. 1). Phenology of individuals was determined by checking the presence of a chick in the burrow. On Mato Islet, 7 and 2 TP and 22 and 7 WTS were equipped in 2017 and 2018, respectively. On Canard Islet, 11 WTS were equipped in 2017. On Nemou Islet 3 and 9 TP were equipped in 2018 and 2019, respectively. Due to logistical and manpower constraints, the two species could not be tracked simultaneously, but 12 out of 27 TP were tracked during the WTS breeding season (mainly before the WTS chick-rearing phase; Fig. 2).
Periods of tracking for each species, and the extent of the breeding period of wedge-tailed shearwater (WTS) in New Caledonia, from the return of breeding adults to colonies (end of October) to chick fledging (May). Breeding and chick-rearing periods for Tahiti petrel (TP) are not shown as little prior information is available. Pictures are © Tubenoses Project, Hadoram Shirihai.
Breeding adults were fitted with either 4.5 g ECOTONE, 6 g LOTEK, or 12.5 g TECHNOSMART GPS-loggers, representing less than 3% of WTS and TP body weight (mean weight ± standard error: 413 ± 40 g and 430 ± 43 g in this study, respectively), i.e. below the limit commonly accepted to limit behavior modification42. The lightest GPS-loggers (ECOTONE and LOTEK) were attached to three tail feathers using TESA tape, while the heaviest GPS-loggers (TECHNOSMART) were back-mounted with 4 stripes of TESA tape to ensure that balance during flight would not be affected43. GPS were set to record location every 15 min. Birds were captured by hand at their burrow entrance before feeding their chick. Colonies were monitored every night for 15–20 days, to recapture birds for logger recovery. On recapture, blood was collected (maximum volume of 0.4 mL) from the tarsal vein using a 0.5 mL 29G syringe. Blood samples were centrifuged within 1 h from collection to separate plasma and blood cells that were then stored separately in 70% ethanol, and preserved in a cooler until return to the lab. During bird handling on Mato islet, spontaneous regurgitates were collected from 3 TP and 6 WTS and stored frozen at − 20 °C as soon as possible for DNA analyses. However, due to logistical constraints in the field, regurgitates were sometimes stored in a cooler for 1–3 days before being frozen. To estimate a possible impact of GPS-tracking on fledging success, we compared the proportion of TP chick-fledging in Nemou islet between 8 burrows were individuals were fitted with loggers and 45 control burrows. A chi-square test with Yate’s continuity correction indicated that GPS-equipment did not significantly impact the fledging success (Χ-squared = 0.023, p = 0.879).
Phenology of Tahiti petrels
One hundred TP burrows were monitored every 2 months from July 2018 to March 2020 on Nemou Islet. Nest contents were checked using a burrowscope to determine the breeding status (i.e. egg, or stage of the chick). Camera traps placed in front of 15 burrows allowed to obtain the precise emergence and fledging dates of chicks. Combined with known incubation and chick-rearing duration (i.e. 55 days and 110–120 days respectively, according to Villard36), this allowed us to estimate egg-laying, hatching, and fledging dates with an accuracy ranging from 6 days to 2 weeks. Hatching dates were estimated by subtracting the average duration of chick-rearing to the fledging date and using the stage of development of the chicks. Stage of development was determined based on our knowledge of TP chick growth from prior monitoring (unpublished data) and coupled with information from prior studies36. Laying dates were estimated by subtracting the average incubation duration from the estimated hatching date. Egg-laying, hatching, and fledging dates were estimated similarly in burrows without camera traps, but with less accuracy, ranging from 2 to 3 weeks. For burrows containing a chick during our last visit (i.e. March 2020), the laying date was estimated by subtracting the incubation duration from the estimated hatching date. A total of 86 breeding events were observed during the study period. Egg-laying, hatching, and fledging dates were estimated and visually represented using 1d Kernel Density Estimate with the stat_density function implemented in the package ‘ggplot2’44.
Foraging trip characteristics
Data for a total of 40 TP foraging trips were collected in 2017 (number of trips: n = 8), 2018 (n = 12), and 2019 (n = 20) from 21 GPS-tracked individuals, and 57 WTS foraging trips were obtained in 2017 (n = 45), and 2018 (n = 12) from 38 individuals. Multiple trips were sometimes recorded from the same individuals (Table 1).
The following metrics were calculated for each trip, using the R package “trip”45: (1) foraging trip duration from the departure to the return to the colony, (2) cumulative distance travelled between all locations assuming straight-line Euclidean distances between 2 successive locations, (3) maximum distance from the colony (hereafter “maximum range”), (4) average travel speed along the trip at sea (i.e. total distance travelled divided by the trip duration), and (5) maximum travel speed during the trip, computed between two successive locations, and assuming straight-line Euclidean distances. When tracks were incomplete (i.e. when the battery stopped before the individual started to return to the colony, n = 9), trip duration was estimated using the individual return date to the burrow surveyed by the field team. Total durations of six incomplete tracks were impossible to estimate, and these tracks were therefore removed from the trip duration analysis, resulting in a total of 53 and 24 trips being considered for WTS and TP respectively. Incomplete trips were also removed from maximal distance travelled analysis. To compare trip parameters between species, we constructed an ANOVA model including trip parameters as response variables (i.e. trip duration, distance travelled, maximum range, average speed and maximum speed) and species, colony type (i.e. uni-species or sympatric), interaction between species and colony type, and interaction between species and year as explanatory variables. Given the large-scale travelling capability of Procellariids seabirds22 and the proximity between colonies (i.e. less than 180 km apart by travelling over the sea), we did not expect the flight characteristics to be impacted by the fact that the two species are breeding in sympatry or in separate islets. However, colony type was kept in the analyses to make sure this assumptions are true. Pairwise Tukey HSD post-hoc tests were then conducted to test for differences between species. We examined the foraging trip length of breeding individuals using a frequency distribution of foraging trip duration for both species. Silverman’s test were performed to test if the distributions significantly differed from unimodal distribution. Moreover, as 3-day trip represents the same time than three 1-day trips, the frequency distribution plot underestimates the importance of long trips. Following Congdon et al.24, we calculated the proportion of time spent on trips of different length by each individual to remove this bias.
Behavioral assessment and activity pattern
We determined TP and WTS at-sea behaviors using the Expectation Maximization binary Clustering (EMbC) algorithm46 for each species separately. EMbC is a robust multivariate clustering algorithm using speed and turning angle of the animal trajectory to identify four main behaviors. GPS tracks with intervals shorter than 30 min were analyzed using this method, and then each GPS location was assigned to a cluster (i.e. one of the four behaviors). Low speed and high turns were interpreted as intensive foraging, high speeds and high turns as extensive searching, low speeds and low turns as resting on the water and high speeds and low turns as travelling-commuting movements. This method has been used to assess ecologically meaningful behaviors from geolocation data for a range of seabird species, including Procellariiforms47,48,49,50,51.
In order to estimate daily activity patterns of each species, the total number of each behavioral type identified by the EMbC was summed per hour of the day, and divided by the total number of behaviors per hour, thus representing the relative proportion of each behavior according to the time of the day. Daily distribution of each behavior were compared using Watson-Wheelers test for homogeneity on two samples of circular data, implemented in the ‘circular’ package52. Proportion of each behavior was also calculated per individual in order to compare the time spent on each behavior between species. This comparison was computed using a multivariate analysis of variance (MANOVA), including the proportional use of each behavior per individual as response variables, and species, colony type (i.e. colonies where individuals breed in sympatry or where only one of the species breed) and their interaction as response variable. Finally, the proportion of each behavior were compared between daytime and nighttime for each species using a chi-squared test.
Foraging areas and their overlap
To deal with data heterogeneity related to the use of various GPS devices with differences in the acquisition frequency of GPS locations (between 15 and 60 min, average 23.7 min for WTS, 27.0 min for TP), all tracks were interpolated at a regular interval of 15 min using the function redisltraj from the R package adehabitatLT53. Interpolated tracks were used to identify the main foraging areas of both species, by computing the Kernel Utilization Distribution of the GPS locations identified as “intensive foraging” or “extensive search” with a smoothing parameter h = 0.2° to avoid over-fragmentation, using the R package adehabitatHR53. Main foraging areas were defined as 90% Utilization Distributions (UDs), representing the 90 percent volume contours of the Kernel Utilization Distribution. Spatial overlap of colony foraging areas was determined by overlapping the UDs 90% of the different colonies for each species, and calculating the percent of area shared, ranging from 0–100% following this equation54:
With A0 = the area of 90% UD intersection between colonies, and AColony = the area of the 90% UD of the colony, calculated with the package rgeos55.
Depth of foraging bouts
The depth of the ocean where intensive foraging or extensive search bouts were performed was determined using the ‘ETOPO180’ variable downloaded from https://www.ngdc.noaa.gov/mgg/global/global.html. Values were compared between species using Wilcoxon–Mann–Whitney tests.
Stable isotope analyses
Values of carbon (δ13C) and nitrogen (δ15N) were analyzed in plasma and red blood cells of GPS-tracked TP and WTS. Since lipids can affect plasma δ13C values, they were removed using 2:1 chloroform: methanol mixture56. Between 0.5 and 5 mg of dried plasma were repeatedly shaken (2–3 treatments) for 1 h in 4 ml of the solvent mixture. The sample was then centrifuged at 4000 g for 5 min and the supernatant containing the lipids was discarded. Lipid-free pellets were then dried at 60 °C overnight. Sub-samples of plasma and red blood cells were weighed (0.3 mg) with a microbalance, and packed into tin cups. Relative abundances of C and N isotopes were determined with a continuous flow mass spectrometer (THERMO SCIENTIFIC Delta V Advantage) coupled to an elemental analyzer (THERMO SCIENTIFIC Flash EA 1112). Replicate measurements of internal laboratory standards (acetanilide) indicated measurement errors < 0.10‰ for both δ13C and δ15N values. Stable isotope ratios are reported in δ (Delta) notation as parts per thousand (‰) deviation from the international standards δ13CPDB and δ15Nair according to the equation \(\updelta {\text{X }} = \left[ {{\text{Rsample }} / {\text{Rstandard}}} \right){-}1] \times 1000\) where X is 13C or 15N and Rsample and Rstandard are the corresponding ratio 13C ⁄12C or 15N⁄14N of samples and international standards.
δ15N values are generally used as a proxy of the trophic level of consumers, and indirectly inform on the type of prey eaten13. We thus used them to test if TP and WTS forage at different trophic levels. δ13C values mainly reflect the carbon source food and habitat type of consumers14,57, and were used to look at a possible difference in foraging habitats (e.g. neritic vs. oceanic for instance) between the two seabird species. Stable isotope values were compared between species using a PERMANOVA, with δ15N and δ13C values as response variables, and species, colony type, year and as explanatory variables. As mentioned above, we did not expect an impact of colony type on stable isotope values, but this variable was kept in the analyses to ensure the veracity of our assumptions. The PERMANOVA was performed using the function adonis from the vegan58 package. SIA of carbon and nitrogen were used together to estimate and compare isotopic niche width between the two species using Stable Isotope Bayesian Ellipses In R (R package SIBER59). The standard ellipse area corrected for small sample sizes (SEAC), containing 40% of the bivariate δ13C and δ15N data, and the convex hull areas (TA) were computed for each species, giving an estimation of their isotopic niche width. For statistical comparison, we calculated the Bayesian standard ellipse areas (SEAB) from 10,000 iterations of Markov-chain Monte Carlo simulation59.
Isotopic niche consistency
Plasma isotope values reflect diet integrated 3–4 days prior to sampling, whereas red blood cells isotopic values reflect longer term integrated diet (i.e. several weeks)56. Because of the different turn-over time between tissues, we used them to investigate the short-term consistency in the isotopic niche (i.e. consistency in trophic level and carbon sources) of both species by regressing stable isotope values in plasma on those in red blood cells60. Because δ13C has a trophic component, we used the studentized residuals of the relationships with δ15N in the same tissue to determine the degree of repeatability in δ13C, independently of trophic effects61,62. A significant result from the linear regression model would indicate the use of constant habitat (δ13C) or trophic level of prey consumed (δ15N) over time by individuals.
DNA metabarcoding
Regurgitate samples were used for DNA metabarcoding dietary analyses. Only a brief overview of the DNA metabarcoding protocol is given below with the full details being described in the supplementary information. DNA was extracted from the regurgitate samples using a modified Qiagen DNeasy Blood & Tissue protocol. DNA extracts were screened for PCR inhibitors and the optimal working solution (i.e. undiluted DNA extracts or a 1:10 dilution) was used for the construction of high-throughput sequencing (HTS) libraries.
HTS libraries for each sample were constructed using fish (MiFish-U63), cephalopod (CephMLS64), and crustacean (CrustMLS65) specific primers with three PCR replicates performed for each sample by primer combination. Amplicon libraries were pooled and cleaned prior to paired-end sequencing at the Ramaciotti Centre for Genomics on the MiSeq platform using the v2 2 × 300 bp sequencing kit to obtain approximately 50,000–60,000 reads for each sample by primer combination. The Trimmomatic v.0.3666 and OBITOOLS software package67 were used for subsequent filtering of the reads following the general workflow described in De Barba et al.68. Taxonomic assignments were performed using both the approach available within the OBITOOLS pipeline and a BLAST search on the NCBI nucleotide database. A consensus taxonomic assignment was obtained considering only family and genus level assignments and used for further analyses. Negative extraction and PCR controls were used and carried through the workflow to assess potential cross-contamination and set minimal threshold values for species detections. Each prey taxa identified was categorized according to its habitat, depth range and migrating behavior, i.e., pelagic, benthic, benthopelagic, reef-associated, based on Fishbase69 and SeaLifeBase70 information.
The R packages tidyverse71 and vegan58 were used to summarize the data. A community matrix was constructed based on the presence-absence detection for the genus and family level assignments. A Bray–Curtis dissimilarity matrix was constructed, and used to produce a non-metric multidimensional scaling (NMDS) plot showing the dissimilarities in dietary composition between WTS and TP.
Data were compiled and analysed using R v3.4.072.
Ethical statement
All animal experimentation met the ABS/ASAB guidelines for ethical treatment of animals37. Permits to handle birds at studied sites were delivered and approved by New Caledonia’s Province Sud (permits nos. 609-2014/ARR/DENV, 2903-2015/ARR/DENV and 2695-2016/ARR/DENV).
Results
Phenology of Tahiti petrels
TP breeding period on Nemou Islet took place throughout the year, with egg-laying recorded during every visit to the islet. However, a first egg-laying peak occurred in December 2018, and a second one between September and October 2019 (Fig. 3). This led to peaks of hatching in February 2019 and November 2019. This implies that the two main chick-rearing periods spread from February to June 2019, and from December 2019 to April 2020. Therefore, TP main chick-rearing period largely overlapped with the WTS one in 2019 and, to a lesser extent, in 2020.
Top panel: Kernel density estimate of the various breeding periods observed for the Tahiti petrel (TP) on Nemou Islet from July 2018 to March 2020. The top colored rectangles represent the breeding season of wedge-tailed shearwater (WTS). Bottom panel: breeding periods of TP monitored on Nemou Islet. Each bar represents a burrow.
Foraging trip characteristics
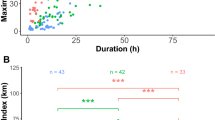
Trip duration (Anova: p = 0.600, Tukey post-hoc test: p.adj = 0.814) and maximum speed (Anova: p = 0.924, Tukey: p.ajd = 0.633) did not differ significantly between the two species (Table 2). In contrast, TP travelled significantly faster (mean speed, Anova: p < 0.001, Tukey: p.adj = 0.001), on longer trips (Anova: p = 0.021, Tukey: p.ajd = 0.005) and further from the colony (Anova: p = 0.031, Tukey: p.adj = 0.005) than WTS (Fig. 4).
Foraging trips of GPS-tracked wedge-tailed shearwaters from Mato (red) and Canard (pink), and Tahiti petrel from Nemou (yellow) and Mato (orange). Blue dots represent the three study colonies. The three islands close to New Caledonia north coast are the Loyalty Islands, the islands further north are part of the Vanuatu archipelago. Bathymetry data were extracted from ‘ETOPO1 Global ReliefModel’ from ‘National Oceanic and Atmospheric Administration’. The map was created using the package ggplot2 version 3.3.244 in R version 3.672 (URL: https://www.r-project.org/index.html).
The frequency distribution of TP foraging trip duration suggests a unimodal foraging strategy with the majority of trips lasting 2 or 3 days (Fig. 5A), and most of the time spent at-sea being during 3-day trips (Fig. 5C). The frequency distribution of WTS trip duration indicates foraging trip duration ranging from 1 to 11 days with ~ 40% of 1-day trips (Fig. 5B). Silverman’s test indicated that TP and WTS foraging trip duration did not significantly differed from a unimodal distribution. However, when taking into account the time spent in each trip, WTS foraging trip duration visually appears bi-modal with most of the time spent at-sea during 1–3 and 7–8 days foraging trips (Fig. 5D).
Behavioral assessment and activity pattern
The proportion of time allocated to each behavior as identified by EMbC, differed strongly between species according to the multivariate analysis of variance (p < 0.001), but they were no significant effects of the colony type (p = 0.64) or the interaction between species and colony type (p = 0.15; Fig. 6). TP spent significantly more time commuting (F = 20.9, p < 0.001) and searching extensively (F = 34.7, p < 0.001) but less time foraging intensively (F = 71.3, p < 0.001) and resting (F = 7.15, p = 0.009) compared with WTS.
The daily activity pattern also differed between species (Fig. 7). Watson–Wheeler tests indicated indicated that daily patterns of behavioural mode use were significantly different between species (foraging: W = 52.231, p < 0.001; extensive search: W = 104.14, p < 0.001; commuting: W = 109.96, p < 0.001; resting: W = 551.84, p < 0.001).
WTS behavior was clearly related to day/night cycle. The species performed intensive foraging mainly during the day (X-squared = 86.650, p < 0.001), rested mostly at night (X-squared = 2014.900, p < 0.001), and commuted at dawn and dusk. In contrast, TP showed a more consistent activity pattern between day and night. TP extensive search pattern was undifferentiated between day and night (X-squared = 0.416, p = 0.51), and intensive foraging was slightly more important during the day (X-squared = 24.487, p < 0.001).
Foraging areas and overlap
All the TP trips from Mato Islet headed towards the south, while TP from Nemou Islet headed towards the north east, mostly targeting the coasts of Loyalty and Vanuatu islands (Fig. 4). WTS from Mato Islet mostly headed towards the east coast of New Caledonia, and WTS from Canard islet either towards the south or the north.
The inter-specific overlap of the foraging areas represented by the 90% Kernel Utilization Distribution was higher (mean: 15.1%) than the intra-specific (i.e. for individuals of the same species but from different colonies) overlap (mean: 9.7%, Fig. 8). However, where species are breeding in sympatry (Mato Islet), the overlap of foraging areas was low (7.8%) between species. Intra-specific overlap was higher in the WTS (16.2%) than in the TP (3.2%) (Table 3).
90% Utilization distributions (UDs) of foraging GPS locations of Tahiti petrels (TP) and wedge-tailed shearwater (WTS). The overlap between UDs 90% is represented in purple. Bathymetry data were extracted from ‘ETOPO1 Global ReliefModel’ from ‘National Oceanic and Atmospheric Administration’. The maps were created using the package ggplot2 version 3.3.244 in R version 3.672 (URL: https://www.r-project.org/index.html).
Depth of foraging bouts
Foraging locations of WTS were situated over significantly deeper areas than TP (mean depth: 2177 ± 22 m and 1900 ± 24 m, respectively, Wilcoxon test: W = 499,910, p < 0.001, Fig. 9). This difference remains significant (Wilcoxon test: W = 1,582,300, p < 0.001) when considering only birds breeding in sympatry (i.e. from Mato islet).
Stable isotope analyses
The PERMANOVA indicated a significant effect of species (R2 = 0.86, p = 0.001), Year (R2 = 0.06, p = 0.001), and interaction between year and species (R2 = 0.02, p = 0.001) on δ13C and δ15N values. Colony type was not significant in this analysis (R2 = 0.003, p = 0.06), and neither was the interaction between species and the colony type.
Mean δ13C values in red blood cells were -17.93 ‰ (± 0.07, n = 23) for TP, and − 17.79 ‰ (± 0.04, n = 46) for WTS (Fig. 10), and did not differ significantly (W = 641.0, p = 0.156). Mean δ13C values for TP were − 17.772 ‰ ± 0.076 in 2017, − 18.227 ‰ ± 0.357 in 2018, and − 17.953 ‰ ± 0.099 in 2019. Mean δ13C values for WTS were − 17.812 ‰ ± 0.033 in 2017 and − 17.732 ‰ ± 0.091 in 2018.
In contrast, δ15N values were significantly different between species (W = 0.0, p < 0.001), with mean δ15N values in TP 5 ‰ higher than those in WTS (12.84‰ ± 0.07, n = 23 and 7.86‰ ± 0.13, n = 46; respectively). Mean δ15N values for TP were 12.872‰ ± 0.104 in 2017, 12.800‰ ± 0.215 in 2018, and 12.837‰ ± 0.107 in 2019. Mean δ15N values for WTS were 7.325‰ ± 0.072 in 2017 and 9.0.21‰ ± 0.147 in 2018.
Mean δ13C values in plasma were − 17.48 ‰ (± 0.06, n = 16) for TP and − 16.93 ‰ (± 0.03, n = 43) for WTS. Mean δ15N values in plasma were 13.53 ‰ (± 0.18, n = 16) for TP and 8.77‰ (± 0.11, n = 43) for WTS.
During chick-rearing, both species exhibited narrow, non-overlapping isotopic niches (Fig. 10, Table 4). However, WTS had a wider breadth of δ15N values, which resulted in a wider isotopic niche for this species (SEAB: p = 0.03).
Isotopic niche consistency
No significant relationship was found in δ15N values between red blood cells and plasma of TP (R2 = 0.207, p = 0.051, Fig. 11A), but a positive significant relationship was found in δ13C values (R2 = 0.299, p = 0.020, Fig. 11B), indicating short-term consistency of carbon source in TP. In contrast, no significant relationship was observed in WTS δ13C values (R2 = 0.007, p = 0.261), while δ15N values in WTS red blood cells and plasma were significantly positively related (R2 = 0.794, p < 0.001), indicating a short-term consistency in the trophic level of WTS prey.
Relationship between δ15N (A) or residuals δ13C (B) values in red blood cells and plasma for Tahiti petrels (TP, orange) and wedge-tailed shearwaters (WTS, red). Lines indicate linear regressions and grey shadows their 95% confidence interval. R2 and p values of the models are represented in the boxes of corresponding colors. Significant p values are indicated in bold.
DNA metabarcoding
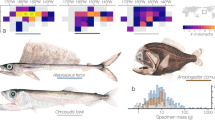
When considering the reads assigned to a family or genus level taxonomy, we obtained a total of 249,918 and 478,790 fish sequences, 146,962 and 55,068 cephalopod sequences and 94,046 and 284,686 crustacean sequences for TP and WTS respectively (see Supplementary Information S1 for summary data of the DNA metabarcoding analyses). A total of 18 taxa were identified in the 3 TP regurgitates with 6 fish families, 4 cephalopod families and 1 crustaceans family. Most of the prey were deep pelagic organisms migrating at the surface at night (Gempylidae, Myctophidae, Chiroteuthidae, Enoploteuthidae, Histopteuthidae, Onychoteuthidae, Stylocheiron sp.), but some of the prey were non-migrating deep pelagic organisms (Neoscopelidae, Sternoptychidae, Euphausia sp.), and three taxa of deep benthopelagic organisms were also identified (Macrouridae, Trichiuridae). Sequences obtained from the 6 WTS regurgitates allowed to identify 25 taxa including 5 fish families, 3 cephalopod families and 9 crustacean families. Fish prey were mainly pelagic species found close to the surface, noticeably anchovies Encrasicholina sp. found in the 6 samples, but also Spratelloides sp. (Clupeidae), Decapterus sp. (Carangidae), and the skipjack tuna Katsuwonus sp. (Scombridae). One fish (Gempylus sp.), the four cephalopod taxa observed (Abralia sp., Enoploteuthis sp., Stenoteuthis sp. and Pterygioteuthis sp.) and two crustaceans of the Euphausiidae family are deep pelagic organisms migrating at the surface at night. Most of the other crustaceans found in the regurgitates are benthic organisms (e.g. Automate sp., Axiidae) and some of them are reef-associated (e.g. Alpheus sp., Dynomene sp., Atergatis sp.). Of the limited number of regurgitate samples available, and over a total of 41 taxa identified overall, only 2 taxa (the cephalopod Abralia sp. and the krill Euphausia sp.) were found in the 2 seabird species regurgitates; the NMDS-plot showed a clear absence of overlap in prey species composition between TP and WTS (Fig. 12). The two-dimensional Bray Curtis dissimilarity index indicated a stress value of 9.72 × 10–5.
Non-Metric Multidimensional Scaling (NMDS) plot based on the presence/absence data derived from the DNA metabarcoding analyses of wedge-tailed shearwaters (WTS) and Tahiti petrels (TP) regurgitate samples. Dots represent the samples, ellipses show the clustering of the different samples according to species, and lines represent the distance of samples to the centre of the ellipse.
Discussion
By combining information from GPS tracking, stable isotope analyses and metabarcoding, this study revealed how the rare and poorly known TP copes with the abundance of WTS foraging in oligotrophic tropical waters surrounding New Caledonia. These two similar-sized sympatric pelagic seabirds differ in their foraging strategy through behavioral differences and trophic segregation. In addition, this is the first comprehensive study to date on TP trophic ecology and foraging strategy, providing the first GPS tracking, isotopic niche and regurgitate metabarcoding data essential for the conservation of this species.
TP bred asynchronously on Nemou Islet, with breeding occurring throughout the year, and a chick-rearing period often largely overlapping the WTS chick-rearing season. Inter-specific breeding season overlap in seabirds has been previously shown to lead to competition for nests when breeding in sympatry, as previously evidenced in the New Caledonian Southern lagoon36. Such an overlap can also lead to competition for resources at-sea when sharing the same diet. Therefore, asynchronous breeding of TP implies only a partial seasonal segregation with WTS. At-sea activity patterns indicated that WTS mainly foraged by day, while TP foraged by day and by night, as previously suggested by Spear et al.32 from observations at sea. This partial temporal segregation of foraging activity may limit the competition for resources to some extent but could also be related to prey differences between the two species. Therefore, despite being not fully elucidated, partial seasonal and daily segregation could occur between species.
Overall, mean foraging trip duration did not differ between species, but the frequency of trip duration was differently distributed. This suggests a unimodal foraging strategy for the TP, and despite non-significant differences from uni-modality, a more bi-modal strategy for the WTS. The bi-modal foraging strategy of WTS breeding in New Caledonia has been previously demonstrated, and linked to poor local biological productivity, and potentially to strong intra-specific competition8. Indeed, studies conducted on breeding WTS proposed that adult foraging strategy is adjusted accordingly to the productivity of near colony areas29. Bimodal foraging strategies are assumed to be used by populations as a means of overcoming intra-specific limitations associated with insufficient resource availability near the colony24,73,74. In contrast, unimodal foraging strategies have been attributed to birds using only highly productive areas close to the colony27,75. Therefore, the unimodal foraging strategy exhibited by TP would suggest that they either access prey more efficiently, or have a different mode of foraging (travelling more, at higher speed).
Despite finding similar maximum flight speed between the species, TP travelled on average faster during their trips than WTS and travelled on longer distances suggesting contrasted behavior time allocations during foraging trips between species. Indeed, during foraging trips, TP spent more time commuting or extensively searching for food, and less time resting than WTS which mainly performed intensive foraging. Extensive search is characterized by high speed movements to forage in a large area in low detail in order to locate prey, while intensive foraging consists of low speed movements with high turning, covering a small area in detail once the individuals encountered areas where resources are plentiful (defined as Area Restricted Search76). This suggests that WTS forage mainly upon patchily distributed prey such as schools of fish, while TP exhibit a more opportunistic foraging behavior, travelling rapidly over longer distances to target more isolated prey.
Moreover, significant and substantial differences found in nitrogen isotope values between these two species suggest the consumption of different prey (e.g. different species and/or different size77). Despite the small sample size, DNA metabarcoding confirmed that WTS targeted schooling prey, such as Engraulidae or Clupeidae, which were the most frequent prey identified in regurgitates. The predation of such schooling species and their mainly diurnal foraging reinforce the hypothesis that WTS could have foraged in association with diurnal sub-surface predators78, making prey accessible to this shallow-diving species (mean maximum diving depth: 5–14 m33,34). Indeed, in Australia, WTS were documented foraging in association with yellowfin (Thunnus albacares) and skipjack (Katsuwonus pelamis) tuna31,32, which are mainly diurnal feeders79. WTS breeding in New Caledonia were shown to forage mostly over deep waters or more rarely at the edge of the barrier coral reef8. The presence of benthic crustaceans identified in their stomach could be secondary consumption, i.e. prey of species preyed by WTS, or could be the consumption of the pelagic larvae of those benthic adults. Moreover, as WTS can sometimes take advantage of the presence of moonlight to feed also at night51, it is possible that they caught some of the deep pelagic organisms (e.g. Thysanopoda sp., Enoploteuthis sp.) migrating to the surface at night.
On the other hand, the most frequent species in the stomach content of TP were deep pelagic fish and cephalopod species, migrating at the surface at night, and benthopelagic fish. Spear et al.32 found similar types of prey when visually analyzing and identifying stomach contents of TP caught at sea in the Eastern Tropical Pacific (i.e. presence of Sternoptychidae, Myctophidae, Macrouridae, Onychoteuthidae, Gempylidae, Trichiuridae and Histioteuthidae). The presence of benthopelagic prey in the stomach contents of TP (e.g.; Macrouridae, Evoxymetopon sp.) could partly explain the high δ15N values found in their red blood cells, as δ15N values of marine organisms, in some particular areas, may increase with the depth of their habitat80. Since TP do not dive, we can assume that deep prey performing diel vertical migration (e.g. Myctophidae) were captured at night, when coming closer to the surface. However, some deep-sea prey species of the TP do not undergo diel vertical migrations, and are thus unlikely to be present at the sea surface. An alternative explanation could thus be that TP scavenge on dead organisms floating at the surface, a behavior observed in many Procellariform species81. The high δ15N values and the high proportion of extensive search behavior in TP provide further support to this explanation. This is consistent with Spear & Ainley82 previous at-sea observations showing that TP obtained 100% of their prey by surface-seizing. This also supports the assumption of Spear et al.32 that 70% of squids eaten by TP were obtained by scavenging. Indeed, TP morphology (i.e. robust bill, extremely long tarsus, short tail, small wing area) is quite distinct from many other tropical petrel species, and Spear and Ainley82 supposed these characteristics to be morphological adaptations for ripping flesh from dead prey too large to be swallowed whole. They also considered their small wing area and short tail as additional adaptations allowing to efficiently search for non-active prey over large areas. Finally, higher δ15N values found in TP red blood cells might reflect the consumption of decomposing prey (since decomposing tissues undergo biochemical changes leading to higher δ15N values83), or alternatively, the consumption of larger prey, as δ15N values most often increase with organism size77, or prey situated at higher trophic levels13, all these hypotheses not being exclusive.
Our results also documented short-term consistency (within ca. 1 month) in the δ15N values of WTS, implying a greater variation in the δ15N values of prey consumed among individuals than within individuals. Along with the wider isotopic niche found in WTS compared to TP, and the broad range of δ15N values detected in their red blood cells, these results suggest that the New Caledonian WTS population is rather generalist, feeding on a variety of trophic levels, but composed of individuals differently specialized, which may reduce intra-specific competition84. In contrast, short-term consistency in δ15N values was not significant for TP. However, δ15N encompassed a shorter range, which could explain the absence of significant results, and suggest a specialist population in terms of trophic levels and/or origin of the prey consumed.
Short-term consistency in δ13C values was not significant in WTS, suggesting the use of variable habitats by the studied population. These results are consistent with their dual foraging strategy, alternating short trips over shallower waters and long trips over deep oceanic waters8, resulting in different δ13C values. In contrast, short-term consistency in δ13C values was significant in TP, suggesting the use of consistent foraging habitats within individuals, in line with the unimodal distribution of their trip duration. The use of constant foraging habitats could also partly explain the narrower isotopic niche width of the species. However, the model testing the δ13C values consistency poorly explained the deviance in these values, and these results have to be taken with precaution.
The high intra-specific segregation observed between WTS colonies found in this study seems to shape the at-sea distribution of the WTS population, most likely in response to the abundance of the species in the region8. Similarly, the two TP colonies are well segregated whereas inter-specific overlap is overall more important. This makes unclear the picture of a possible spatial segregation between TP and WTS. However, the depth of areas foraged is contrasted between the two species, with TP foraging over shallower waters. These results show that TP also forage in coastal areas, particularly individuals from Nemou Islet which travel along the coast of Loyalty Islands and Vanuatu, possibly in relation to their different diet. Tracking both species simultaneously and comparing other environmental variables such as sea surface temperature, productivity or distance to seamounts would be necessary to assess more finely the use of different habitats. This work is in progress and will be the subject of a forthcoming publication.
Finally, we showed that TP and WTS foraged in different habitats, but without a clear spatial segregation. Seasonal segregation occurs during a part of the year, as TP are asynchronous breeders, but most individuals from Nemou Islet were breeding during the WTS breeding season. Temporal segregation also takes place on a daily scale, TP foraging by day and by night, while WTS concentrating their activity during the day. Their diets differed widely, as shown by metabarcoding and stable isotope analyses. WTS mainly foraged on patchily distributed prey, possibly in association with sub-surface predators, while TP had a more opportunistic foraging behavior, and possibly often scavenged on dead prey floating to the surface. They could therefore be associated with fisheries discards. Thus, trophic segregation could facilitate TP access to food resources and the coexistence of the two species, despite the oligotrophic environment surrounding New Caledonia38, and the high abundance of WTS breeding in the area (i.e. > 500,000 breeding pairs40). By analyzing TP trophic ecology and foraging behavior for the first time, this study provides important information about this species relationship with prey, and crucial data for its conservation.
Limitation of the study
Despite providing new insights in the TP ecology and the segregation occurring between species when breeding in sympatry, the present study shows some limitations. First of all, due to manpower constraints, the asynchronous breeding and the rarity of the Tahiti petrel, both species have not been tracked simultaneously. Thus, we cannot exclude that behavior and isotopic values were influenced by environmental conditions at the time of monitoring. Furthermore, we did not take the chick age into account in the models analyzing the trips characteristics. Seabirds can modify the length of their trips according to the breeding stage85. Thus, we cannot exclude that the very long trip (13 days) of one of the TP was linked to the advanced chick age. Finally, one of the limitation of the statistical analyses is that we could not include individuals as a random factor in the models. Given the low number of repeated tracks by individuals, including it as a random factor result in a singular fit of most of the models. However, precautions were taken to analyse the data in the most meaningful way possible, and clear patterns are emerging.
Data availability
Data are available on reasonable request made to the corresponding author.
References
Hutchinson, G. E. Homage to Santa Rosalia or why are there so many kinds of animals?. Am. Nat. 93, 145–159 (1959).
Navarro, J. et al. Ecological segregation in space, time and trophic niche of sympatric planktivorous petrels. PLoS ONE 8, e622897 (2013).
Dehnhard, N. et al. High inter- and intraspecific niche overlap among three sympatrically breeding, closely related seabird species: generalist foraging as an adaptation to a highly variable environment?. J. Anim. Ecol. 89, 104–119 (2020).
Longhurst, A. R. & Pauly, D. Ecology of Tropical Oceans (Academic Press, New York, 1987).
Weimerskirch, H. Are seabirds foraging for unpredictable resources?. Deep Res. Part II Top. Stud. Oceanogr. 54, 211–223 (2007).
McDuie, F., Weeks, S. J. & Congdon, B. C. Oceanographic drivers of near-colony seabird foraging site use in tropical marine systems. Mar. Ecol. Prog. Ser. 589, 209–225 (2018).
Grémillet, D. et al. Irreplaceable area extends marine conservation hotspot off Tunisia: insights from GPS-tracking Scopoli’s shearwaters from the largest seabird colony in the Mediterranean. Mar. Biol. 161, 2669–2680 (2014).
Weimerskirch, H. et al. At-sea movements of wedge-tailed shearwaters during and outside the breeding season from four colonies in New Caledonia. Mar. Ecol. Prog. Ser. 663, 225–238 (2020).
de Grissac, S., Börger, L., Guitteaud, A. & Weimerskirch, H. Contrasting movement strategies among juvenile albatrosses and petrels. Sci. Rep. 6, 26103 (2016).
Mott, R. & Clarke, R. H. Systematic review of geographic biases in the collection of at-sea distribution data for seabirds. Emu 118, 235–246 (2018).
Cherel, Y. et al. Resource partitioning within a tropical seabird community: new information from stable isotopes. Mar. Ecol. Prog. Ser. 366, 281–291 (2008).
France, R. L. & Peters, R. H. Ecosystem differences in the trophic enrichment of13C in aquatic food webs. Can. J. Fish. Aquat. Sci. 54, 1255–1258 (1997).
Minagawa, M. & Wada, E. Stepwise enrichment of 15N along food chains: further evidence and the relation between δ15N and animal age. Geochim. Cosmochim. Acta 48, 1135–1140 (1984).
Newsome, S. D., Martinez del Rio, C., Bearhop, S. & Phillips, D. L. A niche for isotope ecology. Front. Ecol. Environ. 5, 429–436 (2007).
Bearhop, S., Adams, C. E., Waldron, S., Fuller, R. A. & Macleod, H. Determining trophic niche width: a novel approach using stable isotope analysis. J. Anim. Ecol. 73, 1007–10012 (2004).
Bolnick, D. I., Svanbäck, R., Araújo, M. S. & Persson, L. Comparative support for the niche variation hypothesis that more generalized populations also are more heterogeneous. Proc. Natl. Acad. Sci. USA 104, 10075–10079 (2007).
Symondson, W. O. C. Molecular identification of prey in predator diets. Mol. Ecol. 11, 627–641 (2002).
Komura, T., Ando, H., Horikoshi, K., Suzuki, H. & Isagi, Y. DNA barcoding reveals seasonal shifts in diet and consumption of deep-sea fishes in wedge-tailed shearwaters. PLoS ONE 13, e0195385 (2018).
Carreiro, A. R. et al. Metabarcoding, stables isotopes, and tracking: unraveling the trophic ecology of a winter-breeding storm petrel (Hydrobates castro) with a multimethod approach. Mar. Biol. 167, 1–13 (2020).
Alonso, H. et al. An holistic ecological analysis of the diet of Cory’s shearwaters using prey morphological characters and DNA barcoding. Mol. Ecol. 23, 3719–3733 (2014).
del Hoyo, J., Elliott, A. & Sargatal, J. Handbook of the Birds of the World. Ostrich to Ducks Vol. 1 (Lynx Editions, Barcelona, 1992).
Brooke, M. Albatrosses and Petrels Across the World (Oxford University Press, Oxford, 2004).
McDuie, F., Weeks, S. J., Miller, M. G. R. & Congdon, B. C. Breeding tropical shearwaters use distant foraging sites when self-provisioning. Mar. Ornithol. 43, 123–129 (2015).
Congdon, B. C., Krockenberger, A. K. & Smithers, B. V. Dual-foraging and co-ordinated provisioning in a tropical Procellariiform, the wedge-tailed shearwater. Mar. Ecol. Prog. Ser. 301, 293–301 (2005).
Furness, R. W. & Birkhead, T. R. Seabird colony distributions suggest competition for food supplies during the breeding season. Nature 311, 655–656 (1984).
Lewis, S., Sherratt, T. N., Hamer, K. C. & Wanless, S. Evidence of intra-specific competition for food in a pelagic seabird. Nature 412, 816–819 (2001).
Baduini, C. L. Parental provisioning patterns of wedge-tailed shearwaters and their relation to chick body condition. Condor 104, 823 (2002).
Cecere, J. G., Calabrese, L., Rocamora, G. & Catoni, C. Movement patterns and habitat selection of wedge-tailed shearwaters (Puffinus pacificus) breeding at Aride Island, Seychelles. Waterbirds 36, 432–437 (2013).
Peck, D. R. & Congdon, B. C. Colony-specific foraging behaviour and co-ordinated divergence of chick development in the wedge-tailed shearwater Puffinus pacificus. Mar. Ecol. Prog. Ser. 299, 289–296 (2005).
Jaquemet, S., Le Corre, M. & Weimerskirch, H. Seabird community structure in a coastal tropical environment: importance of natural factors and fish aggregating devices (FADs). Mar. Ecol. Prog. Ser. 268, 281–292 (2004).
Miller, M. G. R., Carlile, N., Phillips, J. S., Mcduie, F. & Congdon, B. C. Importance of tropical tuna for seabird foraging over a marine productivity gradient. Mar. Ecol. Prog. Ser. 586, 233–249 (2018).
Spear, L. B., Ainley, D. G. & Walker, W. A. Foraging dynamics of seabirds in the eastern tropical Pacific Ocean. Stud. Avian Biol. 35, 1–99 (2007).
Burger, A. E. Diving depths of shearwaters. Auk 118, 755–759 (2001).
Peck, D. R. & Congdon, B. C. Sex-specific chick provisioning and diving behaviour in the wedge-tailed shearwater Puffinus pacificus. J. Avian Biol. 37, 245–251 (2006).
IUCN 2020. The IUCN Red List of Threatened Species. Version 2020-1. https://www.iucnredlist.org. Downloaded on 19 March 2020.
Villard, P., Dano, S. & Bretagnolle, V. Morphometrics and the breeding biology of the Tahiti petrel Pseudobulweria rostrata. Ibis 148, 285–291 (2006).
Guidelines for the treatment of animals in behavioural research and teaching. Anim. Behav. (2020).
Menkes, C. E. et al. Seasonal oceanography from physics to micronekton in the south-west pacific. Deep Res. Part II Top. Stud. Oceanogr. 113, 125–144 (2015).
Bretagnolle, V. Le Pétrel de la Chaîne Pterodroma (leucoptera) caledonica: Statut et Menaces. Unpubl. Rep. Prov. Sud, Nouméa, New Caledonia (2001).
Pandolfi, M. & Bretagnolle, V. Seabirds of the Southern Lagoon of New Caledonia; distribution, abundance and threats. Waterbirds 25, 202–214 (2002).
QGIS Development Team. QGIS geographic information system. Open Source Geospatial Foundation Project (2018).
Phillips, R. A., Xavier, J. C. & Croxall, J. P. Effects of satellite transmitters on albatrosses and petrels. Auk 120, 1082–1090 (2003).
Vandenabeele, S. P. et al. Excess baggage for birds: inappropriate placement of tags on gannets changes flight patterns. PLoS ONE 9, e92657 (2014).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer, Berlin, 2016).
Michael, A., Sumner, D., Luque, S. & Fischbach, A. Trip : Tools for the Analysis of Animal Track Data. R package version 1.5.0. (2016).
Garriga, J., Palmer, J. R. B., Oltra, A. & Bartumeus, F. Expectation–maximization binary clustering for behavioural annotation. PLoS ONE 11, e0151984 (2016).
de Grissac, S., Bartumeus, F., Cox, S. L. & Weimerskirch, H. Early-life foraging: behavioral responses of newly fledged albatrosses to environmental conditions. Ecol. Evol. 7, 6766–6778 (2017).
Bennison, A. et al. Search and foraging behaviors from movement data: a comparison of methods. Ecol. Evol. 8, 13–24 (2018).
Clay, T. A. et al. Divergent foraging strategies during incubation of an unusually wide-ranging seabird, the Murphy’s petrel. Mar. Biol. 166, 8 (2019).
Mendez, L., Prudor, A. & Weimerskirch, H. Ontogeny of foraging behaviour in juvenile red-footed boobies (Sula sula). Sci. Rep. 7, 13886 (2017).
Ravache, A. et al. Flying to the moon: lunar cycle influences trip duration and nocturnal foraging behavior of the wedge-tailed shearwater Ardenna pacifica. J. Exp. Mar. Biol. Ecol. 525, 151322 (2020).
Lund, U. et al. R package ‘circular’: circular Statistics (version 0.4-93). R Packag (2017).
Calenge, C. The package ‘adehabitat’ for the R software: a tool for the analysis of space and habitat use by animals. Ecol. Modell. 197, 516–519 (2006).
Hedd, A. et al. Foraging areas, offshore habitat use and colony overlap by incubating leach’s storm-petrels Oceanodroma leucorhoa in the Northwest Atlantic. PLoS ONE 13, 1–18 (2018).
Bivand, R. & Rundel, C. Interface to Geometry Engine - Open Source ('GEOS’) (2018).
Hobson, K. A. & Clark, R. G. Turnover of 13 C in cellular and plasma fractions of blood: implications for nondestructive sampling in avian dietary studies. Auk 110, 638–641 (1993).
Jaeger, A., Lecomte, V. J., Weimerskirch, H., Richard, P. & Cherel, Y. Seabird satellite tracking validates the use of latitudinal isoscapes to depict predators’ foraging areas in the Southern Ocean. Rapid Commun. Mass Spectrom. 24, 3456–3460 (2010).
Oksanen, J. et al. Package vegan. R Packag ver (2013).
Jackson, A. L., Inger, R., Parnell, A. C. & Bearhop, S. Comparing isotopic niche widths among and within communities: SIBER - stable isotope Bayesian ellipses in R. J. Anim. Ecol. 80, 595–602 (2011).
Ceia, F. R., Paiva, V. H., Garthe, S., Marques, J. C. & Ramos, J. A. Can variations in the spatial distribution at sea and isotopic niche width be associated with consistency in the isotopic niche of a pelagic seabird species?. Mar. Biol. 161, 1861–1872 (2014).
Votier, S. C. et al. Individual responses of seabirds to commercial fisheries revealed using GPS tracking, stable isotopes and vessel monitoring systems. J. Appl. Ecol. 47, 487–497 (2010).
Bearhop, S. et al. Stable isotopes indicate sex-specific and longer-term individual foraging specialisation in diving seabirds. Mar. Ecol. Prog. Ser. 311, 157–164 (2006).
Miya, M. et al. MiFish, a set of universal PCR primers for metabarcoding environmental DNA from fishes: detection of more than 230 subtropical marine species. R. Soc. Open Sci. 2, 150088 (2015).
Jarman, S. N., Redd, K. S. & Gales, N. J. Group-specific primers for amplifying DNA sequences that identify Amphipoda, Cephalopoda, Echinodermata, Gastropoda, Isopoda, Ostracoda and Thoracica. Mol. Ecol. Notes 6, 268–271 (2006).
Braley, M., Goldsworthy, S. D., Page, B., Steer, M. & Austin, J. J. Assessing morphological and DNA-based diet analysis techniques in a generalist predator, the arrow squid Nototodarus gouldi. Mol. Ecol. Resour. 10, 466–474 (2010).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30, 2114–2120 (2014).
Boyer, F. et al. obitools: A unix-inspired software package for DNA metabarcoding. Mol. Ecol. Resour. 16, 176–182 (2016).
De Barba, M. et al. DNA metabarcoding multiplexing and validation of data accuracy for diet assessment: application to omnivorous diet. Mol. Ecol. Resour. 14, 306–323 (2014).
Froese, R. & Pauly, D. Fishbase. World Wide Web electronic publication. FishBase (2019).
Palomares, M. L. D. & Pauly, D. SeaLifeBase. World Wide Web Electronic Publication. www.sealifebase.org, version (2014).
Wickham, H. tidyverse: Easily Install and Load ‘Tidyverse’ Packages. R package version 1.0.0 (2016).
R Core Team. R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, Vienna, Austria, 2018).
Baduini, C. L. & Hyrenbach, K. D. Biogeography of procellariiform foraging strategies: does ocean productivity influence provisioning?. Mar. Ornithol. 31, 101–112 (2003).
Weimerskirch, H. et al. Alternate long and short foraging trips in pelagic seabird parents. Anim. Behav. 47, 472–476 (1994).
Granadeiro, J., Nunes, M., Silva, M. & Furness, R. Flexible foraging strategy of Cory’s shearwater, Calonectris diomedea, during the chick-rearing period. Anim. Behav. 56, 1169–1176 (1998).
Kareiva, P. & Odell, G. Swarms of predators exhibit ‘preytaxis’ if individual predators use area-restricted search. Am. Nat. 130, 233–270 (1987).
Jennings, S., Pinnegar, J. K., Polunin, N. V. C. & Warr, K. J. Linking size-based and trophic analyses of benthic community structure. Mar. Ecol. Prog. Ser. 226, 77–85 (2002).
Clua, É & Grosvalet, F. Mixed-species feeding aggregation of dolphins, large tunas and seabirds in the Azores. Aquat. Living Resour. 14, 11–18 (2001).
Roger, C. Relationships among yellowfin and skipjack tuna, their prey-fish and plankton. Fish. Oceanogr. 3, 133–141 (1994).
Choy, C. A., Popp, B. N., Hannides, C. C. S. & Drazen, J. C. Trophic structure and food resources of epipelagic and mesopelagic fishes in the north pacific subtropical Gyre ecosystem inferred from nitrogen isotopic compositions. Limnol. Oceanogr. 60, 1156–1171 (2015).
Lipinski, M. R. & Jackson, S. Surface-feeding on cephalopods by procellariiform seabirds in the southern Benguela region, South Africa. J. Zool. 218, 549–563 (1989).
Spear, L. B. & Ainley, D. G. Morphological differences relative to ecological segregation in petrels (family: Procellariidae) of the Southern Ocean And Tropical Pacific. Auk 115, 1017–1033 (1998).
Keenan, S. W. & DeBruyn, J. M. Changes to vertebrate tissue stable isotope (δ15N) composition during decomposition. Sci. Rep. 9, 1–12 (2019).
Bolnick, D. I. et al. The ecology of individuals: Incidence and implications of individual specialization. Am. Nat. 161, 1–28 (2003).
Mendez, L., Cotté, C., Prudor, A. & Weimerskirch, H. Variability in foraging behaviour of red-footed boobies nesting on Europa Island. Acta Oecol. 72, 87–97 (2016).
Acknowledgements
This study was mainly supported by the European Union BEST 2.0 project “Biopelagos” (Grant Number 1070–2015 to SPC), the Province Nord (Grant Number 15C493) and the DAFE-NC (Grant Number 2019/DAFE/873), while the “Ecole Doctorale du Pacifique” and “Université de la Nouvelle-Calédonie” provided A.R. with a PhD fellowship. CPER (Contrat de Projet Etat-Région) and the FEDER (Fonds Européen de Développement Régional) provided additional funding to the IRMS of LIENSs laboratory and the IUF (Institut Universitaire de France) is acknowledged for its support to P. Bustamante as a Senior Member. We wants to warmly thank the many collaborators and students involved in the fieldwork in New Caledonia, especially M. Thibault, J. Allemand, E.Bourguet, M. Brisset, A. Bodin, G. Chagneau, J. Dijoux, M. Mathivet, A. Pujapujane, and H. de Méringo. We also thank for field work facilities and permits, the environmental services of the Province Nord and Province Sud along with customary traditional authorities for allowing us working on Nemou islet. We appreciated the support of M. Brault-Favrou from the Plateforme Analyses Elémentaires of the LIENSs laboratory for proceeding the samples for the stable isotope analyses, and of G. Guillou from the Plateforme Analyses Isotopiques of the LIENSs laboratory for running stable isotope analyses. We thank Jack Rojahn and Sumaiya Quasimf from the University of Canberra for undertaking the trials required to get the extractions from the regurgitate samples and their laboratory contribution to the DNA metabarcoding analyses.
Author information
Authors and Affiliations
Contributions
A.R., K.B. and E.V. conceived and designed the research. A.R., K.B., A.P., S.D. and S.dG. conducted the fieldwork. A.L., P.B., J.B. and D.G. analyzed the samples. A.R. wrote the manuscript, with inputs from K.B., H.W., S.dG., M.M., V.A., J.B., Y.L. and E.V. All authors commented on manuscript drafts.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ravache, A., Bourgeois, K., Weimerskirch, H. et al. Behavioral and trophic segregations help the Tahiti petrel to cope with the abundance of wedge-tailed shearwater when foraging in oligotrophic tropical waters. Sci Rep 10, 15129 (2020). https://doi.org/10.1038/s41598-020-72206-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-72206-0
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.