Abstract
Oral cancer (OC) ranked as eleventh malignancy worldwide, with the increasing incidence among young patients. Limited understanding of complications in cancer progression, its development system, and their interactions are major restrictions towards the progress of optimal and effective treatment strategies. The system-level approach has been designed to explore genetic complexity of the disease and to identify novel oral cancer related genes to detect genomic alterations at molecular level, through cDNA differential analysis. We analyzed 21 oral cancer-related cDNA datasets and listed 30 differentially expressed genes (DEGs). Among 30, we found 6 significant DEGs including CYP1A1, CYP1B1, ADCY2, C7, SERPINB5, and ANAPC13 and studied their functional role in OC. Our genomic and interactive analysis showed significant enrichment of xenobiotics metabolism, p53 signaling pathway and microRNA pathways, towards OC progression and development. We used human proteomic data for post-translational modifications to interpret disease mutations and inter-individual genetic variations. The mutational analysis revealed the sequence predicted disordered region of 14%, 12.5%, 10.5% for ADCY2, CYP1B1, and C7 respectively. The MiRNA target prediction showed functional molecular annotation including specific miRNA-targets hsa-miR-4282, hsa-miR-2052, hsa-miR-216a-3p, for CYP1B1, C7, and ADCY2 respectively associated with oral cancer. We constructed the system level network and found important gene signatures. The drug-gene interaction of OC source genes with seven FDA approved OC drugs help to design or identify new drug target or establishing novel biomedical linkages regarding disease pathophysiology. This investigation demonstrates the importance of system genetics for identifying 6 OC genes (CYP1A1, CYP1B1, ADCY2, C7, SERPINB5, and ANAPC13) as potential drugs targets. Our integrative network-based system-level approach would help to find the genetic variants of OC that can accelerate drug discovery outcomes to develop a better understanding regarding treatment strategies for many cancer types.
Similar content being viewed by others
Introduction
Oral Cancer constitutes approximately 90% among all Head and Neck Cancer (HNC) sub-types1. However, it is more prominent in urban areas of South Asia with a ratio of 15–40% among all cancer types2. In Pakistan, it ranked as 2nd most prevalent cancer-type, with increasing incidence in the past few years3,4. The complexity of genetic mechanisms in cancer has been revealed through recent investigations. Many biological systems seem to involved in the development and progression of the cancer. But, the complications in system-interactions are limitedly understood which is a major restriction in developing effective treatments5. The gene expression studies may help to investigate the differential expression of genes in different biological states, cell cycle stages, subjects or tissues. This gene expression analysis is an important pinpoint for investigating biological processes and their functional disorders. cDNA microarrays were used to monitor and reveal the expression level for thousands of the genes which are differentially expressed in tumors simultaneously5. This technique can exploit this valuable information regarding gene expression analysis. This rapidly progressing technique provides comprehensive data for gene expression profiling of thousands of genes to the investigators in one experiment. Many studies demonstrated that this technique is useful to identify novel genes for cancer and its molecular level classification in human6,7. Thus, this novel technique may help us to identify new potential targets for drug development for optimal and effective disease therapies. It may also establish an important link between clinical medicine and gene sequences for humans8. Gene Expression Omnibus (GEO) is a publicly available database that can be used for high-throughput screening of molecular variants. It contains the microarray data including single and multiple channel-based experiments to determine abundance of genomic DNA, mRNA and protein molecules. The scientific community widely uses the gene expression database to analyze and search the molecular components for systems-level investigations. Several microarrays applications for meta-analysis are designed to investigate pathological mechanisms associated with genetic risk factors9,10,11. Therefore, the genome expression analysis may help to identify unusual alterations across the genome using microarray technology12. This study aims to identify genetic causes and some probable genetic variants of oral cancer that will help to modify the therapeutic strategies.
Results
The cDNA datasets used in this study belong to various normal or cancerous oral tissues and cell lines. The list of datasets used in this study are shown in Table 1.
Differential analysis, normalization and cross-validation
We analyzed 21 cDNA datasets specifically associated with oral cancer. Each dataset has a different number of samples and the genes derived through mRNA expression profiling using different Affymetrix platforms for OC. The histograms representing expression after normalization indicate the density estimation of data. The shapes and ranges for arrays distributions are similar indicating the quality of the data. The array’s distribution towards the right shows a high background level. The saturation of signal is specified by bulge which appears at the upper end of intensity range (Fig. 1). An automated mechanism was used to perform a comparison of biologically similar groups in pairs. We let off any subgroup without repetition from comparisons for accuracy and verification of differential analysis, and the generalized linear models' 'cv.glm' method measured the error of the cross-validation prediction. The Gaussian dispersion criterion is 0.00519 which indicates the degree of confidence (Table 2). With K-folds estimation we obtained the same delta value of 0.00515 as we used the LOOCV approach (during raw cross-validation and afterward during modified cross-validation). The substantial codes (0.1, 0.01, 0.001, and 0.05) with residuals of limited deviance suggested the consistency of the differential analysis. cDNA datasets were also analyzed for some necessary factors like RNA quality, sequence biases or RNA degradation. In genomic analysis, the use of low-quality RNA samples in the sequencing of the entire genome is inefficient. It is not clear if transcript degradation occurs reliably in low-quality RNA samples, in which case the effects of degradation can be reversed by data normalization or whether different RNA samples can be degraded at different rates, which could bias expression measurements. So, for differential expression analysis, we assessed the RNA quality. To verify the dataset reliability for identification of variation at the transcriptional level in original samples. The normalization process was used to standardize sample handling techniques and to assess optimal RNA variability threshold by using discrimination measures for statistical and algorithmic analysis. All the probe sets have their individual probes aligned at 5′-end of the target RNA molecule. The competitive binding of a particular probe to its target has been observed to depend upon a 3′/5′ intensity gradient. Due to the poor quality of RNA, a reduced quantity of RNA is hybridized to the array. The low hybridization leads to a decrease in the total signal output level. But if the degree of saturation level increases the 3′/5′ intensity gradient decreases. The 3′-end of the target gene contains a probe set that corresponds to the transcripts. The statistical and function summary for each batch-array is produced by ‘AffyRNAdeg’ to measure RNA degradation level and its significance (Fig. 2).
Normalization and differential analysis. Histogram (smoothed histograms) shows density estimate of the data. Typically, the distributions of the arrays have similar shapes and ranges. Arrays whose distributions are very different from the others are considered for possible problems. High levels of background shifted an array's distribution to the right. Lack of signal diminishes its right tail. A bulge at the upper end of the intensity range indicated the signal saturation.
Disease-gene curation for differentially expressed genes (DEGs)
From 21 datasets, we found 30 DEGs and David tool was used to retrieve their gene symbol and biological annotation. We selected the most significant ranked genes from the list of differentially expressed genes. For disease-gene curation, these genes were text mined using CTD (Comparative Toxicogenomics Database), PubMed, OMIM, MeSH, and PMC databases to filter disease-specific genes (Supplementary Table 1). We observed CYP1B1, CYP1A1, C7, ADCY2, SERPINB5, and ANAPC13 are the most curated terms in the databases. These shortlisted genes were further analyzed by mapping at (p < 0.00005) through Cancer Genetics and OMIM databases and observed their role in carcinogenesis (Fig. 3).
Enrichment and cluster analysis of DEGs
These genes showed enrichment substantially linked with hydroxylase, P450 pathway, steroid metabolic process, monooxygenase activity, cellular response to organic cyclic compound, and aromatase activity (Table 3). The dysregulation of these genes causes genetic heterogeneity, autosomal recessive disorder, head and neck disorders and other clinical phenotypes (Fig. 4).
The function of the gene, its regulation, subtypes, and cellular processes play a key role in understanding its biology. The functional enrichment analysis showed that shortlisted CYP1A1, CYP1B1 genes are known to involved in xenobiotic metabolic and energy pathways. C7 is involved in immune response whereas SERPINB5 is known for protein metabolism. While ADCY2 plays an important role in cell communication and signal transduction. ANAPC13, have a potential role in class-I MHC-mediated antigens and cell-cycle progression at early tumor stages. The membrane attack complex in the extracellular region for C7, Expression of CYP1A1, SERPINB5, CYP1B1 is found in the endoplasmic reticulum. CYP1A1 is also found in nucleus and microsomes. ANAPC13 is a well-known anaphase-promoting complex. Differential expression of ADCY2, SERPINB5, CYP1B1 is also found in the cytoplasm.
Cluster analysis of selected DEGs helps us in the recognition of functional annotation and significance. The results were observed with the Euclidean distance (Fig. 5). The genetic expression of sample cells is distinguished as cases and control indicating the obvious differences between two of these groups. The analysis showed the down and up-regulated genes based on the p value and fold changes (Table 4).
Mutation analysis
ADCY2 has eight post-translational modification (PTM) sites with 234 recurrent cancer mutations at the chromosome no. 5 positive-strand encoding 1091 protein residues representing 14.02% of the predicted disordered region. The mutation visualization plot shows ADCY2 isoform ADCY2 Q474R direct network-rewiring mutation impact at the position 474, with reference amino acid residue Q and mutated amino acid residue R in the protein. The affected-site at position 472 with S amino acid residue-site enriched with a phosphorylation-type mutation affecting PTMs. Another, ADCY2 isoform ADCY2 S655R, reveals the mutation for this protein at position 655 including amino acid residue S comparison with the mutated amino acid residue R. In position 659, S-amino acid residue site enriched with phosphorylation-type mutation, this shows distal-mutation PTM impact with the affected-site. CYP1B1 showed a 10.5% predicted disordered sequence region with 100 mutations observed at chromosome no 2 on the negative strand with 543 protein residues and 3 PTM sites. CYP1B1 I87S, CYP1B1 Q479H, CYP1B1 T510I isoforms were revealed for CYP1B1 mutational enrichment, at the positions 87,479,510 respectively. The reference amino acid residue for these isoforms was I, Q, T along with mutant amino acid residues S, H, I respectively for each isoform. Similarly, the mutational analysis of C7 showed that 12.57% of the sequence predicted for disease-pathophysiology. Total 244 number of mutations were found on the positive strand of chromosome no. 5 for C7. The number of PTM sites for C7 was eight with 843 protein residues. So, 10 isoforms for C7 were found, among them C7 Q29R and C7 G41D were in distal-mutational PTM impact (Fig. 6). The reference amino acid residues for C7 Q29R, C7 G41D, and C7 T756I were Q, G, T, and mutant amino acid residues reported were I, R, and D, respectively (Supplementary Table 2).
Analysis and exploration of mutations affecting post-translational modification (PTM) sites in human genes/proteins using online ActiveDriverDB database. Needle plots indicate the PTM site mutations in our genes/proteins. The graph shows the outcomes based on the specific type of PTM, cancer type, and mutation subset (presented in legend color codes). Height (y-axis) represents the number of occurrences of the mutation while Horizontal position (x-axis) indicates the position of protein’s amino acid sequence. Pinhead color signifies the mutation impact and X-axis coloring shows the type of PTM associated with the mutation location. Mutational Impacts: Rewiring: mutation-induced gains and losses of kinase-bound sequence motifs (predicted by MIMP software); Distal: mutation affects an amino acid located 4–7 amino acids away from the PTM site; Direct: mutation affects the post transcriptionally modified amino acid; Proximal: mutation affects an amino acid located 1–3 amino acids away from PTM site; Sites: Amino acid sites/ regions enriched for mutations affecting post-translational modifications (PTMs).
Protein–protein interaction analysis
We retrieved the related nodes and edges of all oral cancer associated DEGs from the HAPPI database to construct the integrated PPI-network (Fig. 7). This interaction analysis helped us to observe the potential functional interaction among OC related DEGs and other associated genes contributing to the disease phenotype. The seeder or source OC associated DEGs CYP1A1_HUMAN, CYP1B1_HUMAN, ADCY2_HUMAN, C7_HUMAN, SERPINB5_HUMAN, ANAPC13_HUMAN interact with the target genes including BRAC1_HUMAN, CO6_HUMAN, ISG15_HUMAN, S1PR3_HUMAN, and other essential proteins. The network topology shows a significant relationship between seeder and target genes. The identified proteins showed a significant association with disease development. The target proteins GST2_HUMAN and GSTK1_HUMAN play important role in phase-II carcinogen metabolism and interacted with source genes CYP1A1_HUMAN and CYP1B1_HUMAN. The target proteins including CO7_HUMAN, CLUS_HUMAN, CO6_HUMAN and CO5_HUMAN involved in laryngeal carcinoma, oral cavity squamous cell carcinoma and oropharyngeal carcinoma, respectively. Oral cancer ADCY2_HUMAN linked with KNG1_HUMAN as potential biomarkers. SPB5_HUMAN is interacted with P53_HUMAN, which is a potential tumor suppressor gene. We have observed that OC related genes APC13_HUMAN is interacting with CDK2_HUMAN, a prognostic indicator of oral cancer.
Pathway modelling of OC associated DEGs
Integrated pathways were modeled to observe the possible role of DEGs in pathophysiological mechanisms. Ras, p53, MEK, SOS, Rb, Bax, PTEN, and Raf are important interacting genes associated with the pathophysiological mechanism of oral cancer. We found that p53 signaling, microRNA signaling, salivary secretion, human papillomavirus, cell cycle, alcoholism, and xenobiotic metabolism-related pathways are interconnected in the progression and development of the disease (Fig. 8).
Toxicogenomic analysis
The toxicogenomic analysis enabled us to explore chemical genotype–phenotype exposomic information that may lead to disease progression. OC associated DEGs were curated in terms of their activity and expression with different environmental chemicals. The data revealed the activity and expression of DEGs, which either increase or decrease the expression, increase or decrease towards gene activity at different cellular events. It may also affect the cotreatment expression leading to disease occurrence. It was revealed that the same chemical exposure may show different reactivity for different genes. In this case, benzo(a)pyrene increases the expression of CYP1A1 but it affects the reactivity of ADCY2. Methylcholanthrene, albendazole, primaquine increases CYP1A1 activity. Resveratrol and tetrachlorodibenzodioxin were found to affect the binding, decrease the reaction, and increase the CYP1A1 and CYP1B1 activity. Similarly, acetaminophen was found to increase the expression of C7, whereas alpha-cobra toxin may account for decrease reaction. Arsenic may affect C7 expression by increasing its abundance. While nickel was found to decrease C7 expression. However, decitabine was found to affect the cotreatment of SERPINB5 and decrease the gene reactivity with trichostatin A. It was observed that fonofos, methapyrilene, parathion increases and affects the ADCY2 reactivity at different cellular events while valproic acid, okadaic acid, doxorubicin, and bisphenol A decreases the ANAPC13 expression (Fig. 9).
De novo prediction of regulatory motifs
Oral cancer-associated DEGs were used for de novo analysis to predict the regulatory motifs. The transcriptional factors include ARNT, AHR, CEBPA, CTCF, HNF1B, ELK4, TCF3, and NR2E3. The conservation cutoff is 0.40 with a matrix-score threshold of 85% were set as default parameters. The parameter settings standardize to analyze oPOSSUM-tool showed how the transcriptional factor controls its related targets (Table 5).
MiRNAs targets for oral cancer-specific genes
miRDB database helped us to predict the miRNA targets based on the algorithms. We found the reliable score (> 80) of oral cancer-specific miRNAs. Mainly miRNAs including hsa-miR-4786-5p, hsa-miR-4282, hsa-miR-2052, hsa-miR-216a-3p, hsa-miR-3148, and hsa-let-7f-1-3p targets were predicted for CYP1A1, CYP1B1, C7, ADCY2, SERPINB5 and ANAPC13 genes respectively. The functional dysregulation of these genes may lead to disease progression. The predicted target scores, total miRNA hits, seed-binding locations, and 3′-UTR length of DEGs were analyzed (Table 6).
Drug-gene interaction analysis
The toxicogenomic approach was used to investigate the drug-genes interaction to explore available treatments. The genes that interact with anticancer drugs are docetaxel, hydroxyurea, bleomycin, daunorubicin, lansoprazole, doxorubicin, liothyronine sodium, risperidone using DGIdb database. We identified sixteen proteins as potential alternative drug targets including CXCL1, FBXO32, PTTG1, CCNB1, ADCY2, NMU, ANAPC13 and others (Fig. 10). The dysregulation of these proteins may affect the normal expression level and could be a potential part of therapeutic strategies.
Discussion
This study focused on the genetic expression and functional enrichment of genetic variants of oral cancer. The six most significantly OC associated genes (CYP1A1, CYP1B1, SERPINB5, ANAPC13, ADCY2, C7) found through a differential analysis were consider as seeder or source genes. This analysis provides us a list of new genes aberrantly expressed in oral cancer including SERPINB5, ANAPC13, ADCY2. We have investigated differential expression between cases and controls of cDNA datasets at cellular level in oral tissues and found the possible association of these genes in oral cancer. We can get more information about the mechanism of human genetic disorder through microarray studies. The expression profiling of these important genes shows obvious differences between cases and controls. Some DEGs were found upregulated while other downregulated in this analysis. These genes are abnormally expressed to affect physiological functions including cellular signaling, replication, mitotic division, and programmed cell death. We have observed that our differentially expressed genes are associated with the cancer pathways including biological oxidations, metabolism, adenylate cyclase-activating pathway, xenobiotics, G alpha signaling events, transcriptional targets of TAp63 isoforms, p53, and IFN-gamma pathway revealed the biological significance of these genes specifically for oral cancer progression. CDC20_HUMAN, HDAC1_HUMAN, CXL10_HUMAN interacting with source genes are potential drug targets13,14,15. The inherent mutations are reported for genes that encode drug-metabolizing enzymes. Such somatic gene mutations are induced chemically that play a vital role in cell differentiation and growth16. The sequencing investigations not only characterizes the genomics but also revealed thousands of SNVs (single nucleotide variants), the alterations in copy number along with many types of genetic variations. Such genomic to phenomic association identification, their molecular-level mechanisms, disease-related variants along with cancer-derived mutations are the current challenges in the biomedical research17,18. Deciphering inter-individual genetic variation is the latest trend in personal genomic era investigations. The interpretation of genomic to proteomic information may integrate the impact of mutations on cellular system-level investigations in the future with a higher magnitude19. Human proteomic data analysis uses PTMs to interpret disease mutations and inter-individual genetic variations. PTMs being important regulators of protein function and signaling pathways facilitate the missense mutational analysis investigations19.
We have observed that CYP1A1 belongs to potential and well-preserved phase-I xenobiotic metabolizing gene family which is involved in the activation of procarcinogens. The CYP1A1 enzyme is highly associated with increased risk of tumors in the oral cavity, bronchial and laryngeal regions in smokers4. Similarly, the association of CYP1B1 has found in many cancer types20. Many investigations are reported about the substantial link of HNC with CYP1A1 and CYP1B116,21.
SERPINB5 belongs to serpin encoding serine protease which plays a vital role in tumor metastasis22,23. It is a tumor suppressor in epithelial cells and can suppress cancerous cell invasion and their metastasizing in surrounding tissues24. The paradoxical expression of SERPINB5 has been observed in various types of tumor25,26,27,28. We have seen a highly significant association between SERPINB5 expression and oral carcinoma29. ADCY2 is a membrane-associated enzyme which converts adenosine-5′-triphosphate (ATP) into 3′,5′-adenosine monophosphate (cyclic AMP/cAMP) and pyrophosphate30 involving in the regulation of cAMP synthesis31. This gene catalyzes the signaling molecule cAMP through G-protein beta as well as gamma subunit signaling32,33. Therefore, changes in expression patterns of the gene are mediated through down-streaming of the signaling cascades muscarinic acetylcholine receptors which increases IL6 production. The high regulation of the gene is observed in G-proteins, calcium, calmodulin, pyrophosphate, and post-translational modifications. The signaling pathways include RET signaling, Oocyte meiosis, calcium, and chemokine signaling pathways34,35,36. Aberrant methylation of ADCY2 is observed in colorectal, prostate cancer35,36,37 and urinary bladder cancer38,39,40. It has been studied that ANAPC1341 is a large-sized ubiquitin ligase that controls the cell cycle progression42 and involved at early steps of malignancy in tumor cells43. Similarly, C7 (complement component-7), the terminal component for complement cascade and as a cytolytic effector for complement system, lyses transformed malignant cells44,45,46,47,48,49,50,51. The integration of chemical- gene interaction revealed different environmental chemical exposure to disease progression52,53. This analysis helped to reveal the mechanism of action between the chemical and the related gene products and their effect on human disease influence by environmental exposure53,54,55,56.
The PPI network predicted the important association of these genes with disease. These genes have a potential role in xenobiotic metabolism, tumor progression, suppression, cell cycle, HPV (human papilloma virus), alcoholism, and microRNA signaling pathway30,40,48,49,51,57,58,59,60. The transcriptomic analysis showed expressive transcription factors like JUND, FOXO, STAT1. We found the role of these genes in metabolism of xenobiotics, p53 signaling, salivary secretion, class-I MHC mediated antigens and microRNA cancer pathways. The miRNAs regulate post-transcriptional and translational events and expressional dysregulation in these molecules leads towards the progression of many diseases61,62,63. Therefore, the reliable miRNA target prediction is crucial for the functional annotation of miRNAs64,65.
Recent reports proved that the drug-gene network enables us to understand not only the disease pathophysiology but also important in drug designing or new drug target identification or establishing novel biomedical linkages. More importantly, this network proposed many testable assumptions with the potential of great success, though the real achievement can only be justified by experimental studies.
Conclusion
Our simulation-based systems-level hypothesis is comprehensive and effective to sort out the disease-specific genetic variants from cDNA datasets repositories. Therefore, this approach will support understanding the genetic basis of complex phenotypes including cellular replication, protein signaling, mitotic division, and programmed cell death. Based on genomic to phenomic investigations, we have found new genes including ADCY2, SERPINB5, and ANAPC13 linked with oral cancer that could be potential diagnostic or drug targets. These source genes are clearly interacting with other essential genes affecting cell cycle and apoptosis causing carcinogenesis. These findings can provide a valuable framework for developing new therapeutic strategies against oral cancer.
Methods
Accession to cDNA datasets
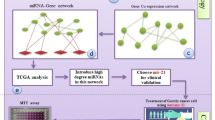
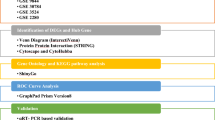
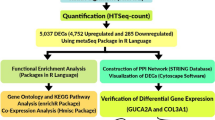
5.2 We downloaded cDNA datasets related to oral cancer from the Gene Expression Omnibus database (GEO) NCBI. The comprehensive framework has been illustrated in Fig. 11 using tools, online servers, and software (Table 7).
Normalization and differential analysis
These datasets were analyzed in identifiable format to easily access pheno-data files and missing values were imputed66. R software version 3.3.3 and Bioconductor packages were used in computational analysis. The normalization and quality control analysis was performed to preprocess the information available by ArrayQuality67,68,69. The background and normalization were aligned by using Robust Multi-Array Analysis (RMA) to detect the PM (perfect matches) and the MM (mismatches) to impute the values for statistical analysis70. RMA is the widely used preprocessing algorithm applied for background correction to remove local artifacts67.
where PM indicates a perfect match, Background by BG and non-specific binding (S); ijk is the signal for probe j of probe set k on array i.
The perfect match involves the combined signals of background (BG) and expression (E). The "ArrayQualityMetrics" software was used to evaluate the quality of dataset that is normalized to each genke's median expression level67,68,71. The gene–gene covariance matrix of each data set was calculated across all arrays while ignoring the missing values. The transformation formula is:
where F1 and F2 represents distribution functions of the actual and reference chips.
To get a description of intensities, we used the RMA-algorithm to measure averages between probes in a sample set. During this analysis RNA quality was evaluated in samples. The RNA degradation analysis was performed by using AffyRNAdeg, summary AffyRNAdeg, and plot AffyRNAdeg packages72. The DEGs were observed using the LIMMA package, that process the information based on modified statistics which is proportional to sample variance offsets. The LIMMA package measured the duplicate spots and quality weights. The statistical analysis was performed to categorize the genes based on the significant cutoffs values logFC greater than 1, FDR less than 0.05, AEL ≥ 40% and p value ≤ 0.0573.
K-fold cross-validation
We used K-Fold Cross-validation and Bootstrap test to estimate accuracy in the differential analysis74 and this approach has the advantage that all the samples in the dataset can ultimately be used for both training and research. This technique is usually easier to calculate estimated average error and has been used to validate the shortlisted differentially expressed genes using the 'Boot' package of Bioconductor. Boots trapping is used effectively in molecular analysis to correct biases75. In such cases, we applied the generalized linear Gaussian models and used the 'cv.glm' method to test the k-fold cross-validation. It estimates the true error as the average rate error:
Leave-one-out-cross-validation (LOOCV) continued to trail the Gaussian rule. The LOOCV approach is instinctively termed as the test set is left out and the rest of the data is used as the training-set75. We used N − 1 subsets for training and the rest for testing. Increasing the number of folds would make the bias of the true error rate estimator low and valid75,76.
The true error is assessed as the average error rate on test cases:
Disease-gene curation of differentially expressed genes (DEGs)
The text mining is important in biomedical research to extract useful information77. This analysis is designed to identify the most significant DEGs, all 30 genes from 21 datasets were curated from the DAVID database to retrieve their gene symbol, gene name, Uniprot_ID. These genes were curated using Comparative Toxicogenomics Database (CTD), Online Mendelian Inheritance in Man (OMIM), PubMed and MeSH databases to observe their role in oral cancer. This screening further shortlisted the significant DEGs78.
Enrichment and cluster analysis
The biological functions of the genes help us to understand the cellular level signaling network. We performed enrichment analysis using the DAVID tool78,79,80. FunRich tool was used to observe the biological functions of oral cancer-related DEGs at molecular level81. The list of DEGs were analyzed for their p- and FDR values79. For cluster analysis, gene expression values of cases and controls of each dataset were studied to observe genetic variations and expression profiling using One Matrix CIMminer tool82,83.
Mutation analysis
Mutations resulting from cancer and the inherited-disease process can be understood to decode the genetic variation by associations of genotype-phenotype. The human genome contains thousands of SNVs (single nucleotide variants) and many are known for the progression of the disease. Approximately 21% of amino acid substitutions are known to be associated with disease-progression in correspondence with missense single nucleotide variants located at PTM protein sites (post-translation modifications). The chemical modification of the amino acid thus basically extends the functionality of the associated protein19.
Mutation of differentially expressed genes were analyzed using online ActiveDriverDB database19. The needle plot mutations analysis provides a visual overview of the position, frequency, and functional significance of all identified mutations in our DEGs. PTM sites with all mutations and the predicted disordered region of protein sequences were observed. Placing the pins corresponds to the position along the sequence of the genes and protein, whereas the related mutation effect and PTM are explained in the figure legend.
Protein–protein interaction
The biological functions are mainly carried out by protein–protein interactions30,84. The interaction of proteins reveals that each protein interacts with one or more genes related to their molecular functions85. The biological networks indicate altered activity in normal or disease conditions. This gene-network aims to identify potentially OC associated gene signatures whose dysfunction directly contributes to disease phenotype are functionally associated. The gene signatures related to each source protein was measured. Human Annotated and Predicted Protein Interaction (HAPPI) and String databases were used to analyze gene–gene/protein–protein interactions of microarray dataset DEGs86. This database annotates and mine comprehensive physical as well as genetic mapping and includes experimentally validated data to simulate biological networks. We have mentioned the threshold for PPI network from HAPPI database. We used high-confidence interactions in our network (the five stars are equivalent to high score (0.90–1). The role and association of these source and target genes in oral cancer were evaluated from Cancer Genetics Web, National Cancer and OMIM database. The molecular networks were visualized by Cytoscape software (version 3.6.0)87. The Cytoscape Network Analyzer calculates topological properties of networks. The degree of annotation between the gene and disease is categorized by nodes in the network.
Pathway analysis of oral cancer linked genes
Reactomic analysis enable us to explore all metabolic networks of DEGs regarding their molecular mechanism. We analyzed these pathways to inter-connect DEGs to show the pathological mechanism of oral cancer. The KEGG and Wiki pathways databases were used to map target genes88,89. PathVisio tool was used to reconstruct the pathway model for understanding system-level analysis90.
Toxicogenomic analysis
The toxicogenomic analysis is carried out by a comparative toxicogenomic database (CTD) to retrieve exposome data. The exposome data helps investigate chemical-genome to phenome relationships to interpret the functional pathway cellular signaling-mechanism towards disease progression influenced by environmental exposures. It provides information regarding chemical-gene/protein and disease interactions which may reveal the particular gene-activity or expression regarding gene-disease connections. The curation of environment-disease exposure helps to analyze the available toxicogenomic information55.
Prediction of regulatory motifs
Cancer has a complex mechanism that can be explored by understanding the biological functions at transcription and post-transcriptional level. oPOSSUM version 3.0 was used to analyze promoter region target motifs like transcription factor binding sites (TFBS) or the overexpression of target matrices91,92. This information helps to understand the functional role of gene targets and eventually gene ontology93.
Prediction of oral cancer-associated miRNA targets
Numerous genes are involved in the biological signaling cascade. These cascades are influenced through small noncoding RNAs as post-transcriptional regulators, known as microRNAs (miRNAs). The function and expression of miRNA play a significant role in understanding gene etiology5,93. miRNA target prediction helps to explore the functional and molecular annotation of disease-specific DEG’s5,94. Therefore, oral cancer associated DEG’s miRNA targets were predicted by miRDB, an online database for functional microRNA target prediction. The target prediction data involves specie specific 3′-UTR sequences, 3′-UTR region length, miRNA seed binding-sites, miRNA-candidate target pairs along with target prediction scores, miRNA-target sequences, and other important description95,96. The MiRNA target predictive score is ranked and > 80 was considered as a reliable score95,96.
Drug-gene interaction analysis
In our study, the drug-gene network analysis was performed to correlate our shortlisted DEGs with FDA approved commercially available anti-cancer drugs. CTD database was used to investigate the relationship between the chemical and disease at default parameters. In this analysis, DEGs were directly linked to anticancer drugs. All drugs, used in this interaction, were verified through the Drug Bank database to check approval status by the FDA.
Data availability
All the other data that support the findings of this study are available from the corresponding author upon request.
References
Lozano, R. et al. Global and regional mortality from 235 causes of death for 20 age groups in 1990 and 2010: A systematic analysis for the Global Burden of Disease Study 2010. The Lancet 380, 2095–2128. https://doi.org/10.1016/S0140-6736(12)61728-0 (2010).
Khawaja, M. R. et al. Chewing of betel, areca and tobacco: Perceptions and knowledge regarding their role in head and neck cancers in an urban squatter settlement in Pakistan. Asian Pac. J. Cancer Prev. 7, 95–100 (2006).
Bhurgri, Y., Bhurgri, A. & Usman, A. Epidemiological review of head and neck cancers in Karachi. Asian Pac. J. 7, 195–200 (2006).
Masood, N., Kayani, M. A., Malik, F. A., Baig, R. M. & Faryal, R. Genetic variations in carcinogen metabolizing genes associated with oral cancer in pakistani population. Asia. Pac. J. Cancer Prev. 12, 491–495 (2011).
Muhammad, S. A. et al. Cellular signaling pathways in insulin resistance-systems biology analyses of microarray dataset reveals new drug target gene signatures of type 2 diabetes mellitus. Front. Physiol. 8, 1–15 (2017).
Golub, T. R. et al. Molecular classification of cancer: Class discovery and class prediction by gene expression monitoring. Science 286(80), 531–527 (1999).
Alizadeh, A. A. et al. Distinct types of diffuse large B-cell lymphoma identified by gene expression profiling. Nature 403, 1–9 (1909).
Okabe, H. et al. Genome-wide analysis of gene expression in human hepatocellular carcinomas using cDNA microarray. Cancer Res. 61, 2129–2137 (2001).
Hong, F. & Breitling, R. A comparison of meta-analysis methods for detecting differentially expressed genes in microarray experiments. Bioinformatics 24, 374–382 (2008).
Irizarry, R. A. et al. Multiple-laboratory comparison of microarray platforms. Nat. Methods 2, 345–349 (2005).
Wheeler, E. & Barroso, I. Genome-wide association studies and type 2 diabetes. Brief. Funct. Genomics 10, 52–60 (2011).
Siddiqui, A. S. et al. Sequence biases in large scale gene expression profiling data. Nucleic Acids Res. 34, e83 (2006).
Wagner, J. M., Hackanson, B., Lübbert, M. & Jung, M. Histone deacetylase (HDAC) inhibitors in recent clinical trials for cancer therapy. Clin. Epigenet. https://doi.org/10.1007/s13148-010-0012-4 (2010).
Wang, Z. et al. Cdc20: A potential novel therapeutic target for cancer treatment. Curr. Pharma. Des. 19, 3210–3214 (2013).
Muller, A. et al. Chemokine receptors in head and neck cancer: Association with metastatic spread and regulation during chemotherapy. Int. J. Cancer https://doi.org/10.1002/ijc.21514 (2014).
Ko, Y. et al. Association of CYP1B1 codon 432 mutant allele in head and neck squamous cell cancer is reflected by somatic mutations of p53 in tumor tissue. Cancer Res. 61, 4398–4404 (2001).
Gonzalez-perez, A., Mustonen, V., Reva, B. & Ritchie, G. R. S. NIH Public Access. 10, 723–729 (2014).
Macarthur, D. G. et al. Guidelines for investigating causality of sequence variants in human disease. Nature 508, 469–476 (2014).
Krassowski, M. et al. ActiveDriverDB: Human disease mutations and genome variation in post-translational modification sites of proteins. 46, 901–910 (2018).
Shimada, T. et al. Metabolic activation of polycyclic aromatic hydrocarbons and other procarcinogens by cytochromes p450 1a1 and p450 1b1 allelic variants and other human cytochromes p450 in Salmonella typhimurium NM2009. Drug Metab. Dispos. 29, 1176–1182 (2001).
Boccia, S. et al. Polymorphisms in metabolic genes, their combination and interaction with tobacco smoke and alcohol consumption and risk of gastric cancer: A case–control study in an Italian population. BMC Cancer 7, 3–10 (2007).
Luo, J. et al. Nuclear cytokine-activated IKK a controls prostate cancer metastasis by repressing. Maspin https://doi.org/10.1038/nature05656 (2007).
Khalkhali-ellis, Z. Maspin. New Front. 12, 7279–7284 (2006).
Zou, Z. et al. Maspin, a serpin with tumor-suppressing activity in human mammary epithelial cells. Science 80(263), 526–529 (1994).
Sood, A. K. et al. The paradoxical expression of Maspin in ovarian carcinoma. Clin Cancer Res 1(8), 2924–2932 (2015).
Achtens, S. M. et al. Expression of the p53 and Maspin protein in primary prostate cancer: Correlation with clinical features. Int. J. Cancer 342, 337–342 (2001).
Ohike, N., Maass, N., Mundhenke, C. & Biallek, M. Clinicopathological significance and molecular regulation of maspin expression in ductal adenocarcinoma of the pancreas. Cancer Lett. 199, 193–200 (2003).
Yu, M., Zheng, H. & Tsuneyama, K. Paradoxical expression of Maspin in gastric carcinomas: Correlation with carcinogenesis and progression B. Hum. Pathol. 38, 1248–1255. https://doi.org/10.1016/j.humpath.2006.11.025 (2007).
Xia, W. et al. High tumoral Maspin expression is associated with improved survival of patients with oral squamous cell carcinoma. Oncogene 19, 2398–2403 (2000).
Li, Y. X. et al. Molecular cloning, sequence identification, and gene expression analysis of bovine ADCY2 gene. Mol. Biol. Rep. 41, 3561–3568 (2014).
Ding, Q. et al. Raf kinase activation of adenylyl cyclases: Isoform-selective regulation. Mol. Pharmacol. 66, 921–928 (2004).
Drozdov, I. et al. Gene network inference and biochemical assessment delineates GPCR pathways and CREB targets in small intestinal neuroendocrine neoplasia. PLoS ONE 6, e22457 (2011).
Iyengar, R. Molecular and functional diversity of mammalian GS – stimulated adenylyl cyclases. The FASEB J. 7, 768–775 (1993).
Plaza-menacho, I., Mologni, L. & Mcdonald, N. Q. Mechanisms of RET signaling in cancer: Current and future implications for targeted therapy. Cell Signal. 26, 1743–1752 (2014).
Liang, B., Li, C. & Zhao, J. Identification of key pathways and genes in colorectal cancer using bioinformatics analysis. Med. Oncol. 33, 1–8 (2016).
Liu, J. et al. Aberrantly methylated—differentially expressed genes and pathways in colorectal cancer. Cancer Cell Int. https://doi.org/10.1186/s12935-017-0444-4 (2017).
Sun, Y., Jia, X., Hou, L. & Liu, X. Screening of differently expressed miRNA and mRNA in prostate cancer by integrated analysis of transcription data. Urology https://doi.org/10.1016/j.urology.2016.04.041 (2016).
Dolićanin, Z., Lj, J. V. & Katić, V. Biomarkers for detection treatment decision and prognosis ontogeneses telomerase tumor suppressor genes genomic studies in bladder cancer. Facta Universitatis 14, 1–5 (2007).
Hardin, M. et al. Severe chronic obstructive pulmonary disease IN Poland. Am. J. Respir. Cell Mol. Biol. 47(2), 203–208. https://doi.org/10.1165/rcmb.2012-0011OC (2012).
Yu, S. J. et al. SPARCL1, Shp2, MSH2, E-cadherin, p53, ADCY-2 and MAPK are prognosis-related in colorectal cancer. World J. Gastroenterol. 17, 2028–2036 (2011).
Yoon, H. et al. Proteomics analysis identifies new components of the fission and budding yeast anaphase-promoting complexes. 12, 2048–2054 (2002).
Peters, J. The anaphase-promoting complex Proteolysis in mitosis and beyond. 9, 931–943 (2002).
Aparecida, C. et al. Translational oncology down-regulation of ANAPC13 and CLTCL1: Early events in the progression of preinvasive ductal carcinoma of the breast 1, 2. Transl. Oncol. 5, 113–123 (2012).
Ricklin, D., George, H., Kun, Y. & John D. L. Complement – a key system for immnue surveillance and homeostasis. Nat. Immunol. 11, 785–797 (2010).
Serna, M., Giles, J. L., Morgan, B. P. & Bubeck, D. Structural basis of complement membrane attack complex formation. Nat. Commun. 7, 10587 (2016).
Podack, E. R. Molecular composition of the tubular structure of the membrane attack complex of complement. J. Biol. Chem. 259, 8641–8647 (1984).
Botto, M. et al. Complement in human diseases: Lessons from complement deficiencies. Mol. Immunol. 46, 2774–2783 (2009).
Gelderman, K. A., Tomlinson, S., Ross, G. D. & Gorter, A. Complement function in mAb-mediated cancer immunotherapy. Trends Immunol. 25, 158–164 (2004).
Ying, L. et al. Complement component 7 (C7), a potential tumor suppressor, is correlated with tumor progression and prognosis. Oncotarget 7, 86536–86546 (2016).
Shen, Y. B. et al. A 425 T > C polymorphism in complement C7 association with resistance to Aeromonas hydrophila in grass carp. Genet Mol. Res. 1, 1–9 (2016).
Ying, L. et al. Complement component 7 (C7), a potential tumor suppressor, is correlated with tumor progression and prognosis. Oncotarget 7, 86536 (2016).
Mortensen, H. M. & Euling, S. Y. Integrating mechanistic and polymorphism data to characterize human genetic susceptibility for environmental chemical risk assessment in the 21st century. Toxicol. Appl. Pharmacol. 271(3), 395–404. https://doi.org/10.1016/j.taap.2011.01.015 (2011).
Mattingly, C. J. et al. The comparative toxicogenomics database: A cross-species resource for building chemical-gene interaction networks. Toxicol. Sci. 92, 587–595 (2006).
Davis, A. P. et al. The comparative toxicogenomics database: Update 2011. Nucleic Acids Res. 92, 1067–1072 (2011).
Davis, A. P. et al. The comparative toxicogenomics database: Update 2019. Nucleic Acids Res. 92, 948–954 (2019).
Davis, A. P. et al. Comparative toxicogenomics database: A knowledgebase and discovery tool for chemical—gene—disease networks. Nucleic Acids Res. 37, 786–792 (2009).
Lynam-Lennon, N., Maher, S. G. & Reynolds, J. V. The roles of microRNA in cancer and apoptosis. Biol. Rev. 84, 55–71 (2009).
Mamidi, S., Höne, S. & Kirschfink, M. The complement system in cancer: Ambivalence between tumour destruction and promotion. Immunobiology 222, 45–54 (2017).
Suryawanshi, S. et al. Complement pathway is frequently altered in endometriosis and endometriosis-associated ovarian cancer. Clin. Cancer Res. 20, 6163–6174 (2014).
Pio, R. Tumor Microenvironment and Cellular Stress (Springer, Berlin, 2014), 772.
Thierfelder, N., Fang, Z., Khanin, R., Rajewsky, N. & Selbach, M. Widespread changes in protein synthesis induced by microRNAs. Nature 455, 58–63 (2008).
Lim, L. P., Lau, N. C., Garrett-engele, P. & Grimson, A. Microarray analysis shows that some microRNAs downregulate large numbers of target mRNAs. Nature 292, 288–292 (2005).
Shin, C. et al. The impact of microRNAs on protein output. Nature 455, 64–71 (2008).
Wong, N. & Wang, X. miRDB: An online resource for microRNA target prediction and functional annotations. Nucleic Acids Res. 43, 146–152 (2015).
Wang, X. & Wang, X. miRDB: A microRNA target prediction and functional annotation database with a wiki interface miRDB: A microRNA target prediction and functional annotation database with a wiki interface. RNA 14, 1012–1017. https://doi.org/10.1261/rna.965408.was (2008).
Tibshirani, R. et al. Missing value estimation methods for DNA microarrays. Bioinformatics 17, 520 (2001).
Bolstad, B. M., Irizarry, R. A., Astrand, M. & Speed, T. P. Gene expression omnibus a comparison of normalization methods for high density oligonucleotide array data based on bias and variance. Bioinformatics 19, 185–193 (2003).
Fujita, A., Sato, J. R., de Oliveira Rodrigues, L., Ferreira, C. E. & Sogayar, M. C. Evaluating different methods of microarray data normalization. BMC Bioinform. 7, 1–11 (2006).
Obenchain, V. et al. Variant annotation: A bioconductor package for exploration and annotation of genetic variants. Bioinformatics 30, 2076–2078 (2014).
Yoon, D., Yi, S. G., Kim, J. H. & Park, T. Two-stage normalization using background intensities in cDNA microarray data. BMC Bioinform. 5, 1–12 (2004).
Obenchain, V. et al. Variant annotation: A bioconductor package for exploration and annotation of genetic variants. Bioinformatics 30, 2076–2078 (2014).
Affymetrix® Microarray Suite User’s Guide. Version 5.0. (2000, 2001).
Jin, Y. & Da, W. Screening of key genes in gastric cancer with DNA microarray analysis (retraction of vol 18, 37, 2013). Eur. J. Med. Res. 20, 30 (2015).
Geisser, S. The predictive sample reuse method with application. J. Am. Stat. Assoc. 70, 320–328 (1975).
Picard, R. R. & Cook, R. D. Cross-validation of regression models. J. Am. Stat. Assoc. 79, 575–583 (1984).
MAQC Consortium. The MicroArray Quality Control (MAQC)-II study of common practices for the development and validation of microarray-based predictive models. Nat. Biotechnol. 8, 827–838 (2010).
Clematide, S. & Rinaldi, F. Ranking relations between diseases, drugs and genes for a curation task. J. Biomed. Semant. 3, S5 (2012).
Nam, D. & Kim, S. Y. Gene-set approach for expression pattern analysis. Brief. Bioinform. 9, 189–197 (2008).
Muhammad, S. A. et al. MicroRNA expression profiling of human respiratory epithelium affected by invasive candida infection. PLoS ONE 10, 1–14 (2015).
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Pathan, M. et al. FunRich: An open access standalone functional enrichment and interaction network analysis tool. Proteomics 15, 2597–2601 (2015).
Weinstein, J. N. et al. An information-intensive approach to the molecular pharmacology of cancer. Science 80(275), 343–349 (1997).
Scherf, U. et al. A gene expression database for the molecular pharmacology of cancer. Nat. Genet. 24, 236–244 (2000).
Muhammad, S. A. et al. Prioritizing drug targets in Clostridium botulinum with a computational systems biology approach. Genomics 104, 24–35 (2014).
Rachlin, J., Cohen, D. D., Cantor, C. & Kasif, S. Biological context networks: A mosaic view of the interactome. Mol. Syst. Biol. 2, 1–12 (2006).
Chen, J. Y., Mamidipalli, S. R. & Huan, T. HAPPI: An online database of comprehensive human annotated and predicted protein interactions. BMC Genomics 10, 1–11 (2009).
Cline, M. S. et al. Integration of biological networks and gene expression data using cytoscape. Nat. Protoc. 2, 2366–2382 (2007).
Bergholdt, R. et al. Identification of novel type 1 diabetes candidate genes by integrating genome-wide association data, protein–protein interactions, and human pancreatic islet gene expression. Diabetes 61, 954–962 (2012).
Califano, A., Butte, A. J., Friend, S., Ideker, T. & Schadt, E. Leveraging models of cell regulation and GWAS data in integrative network-based association studies. Nat. Genet. 44, 841–847 (2012).
Kutmon, M. et al. PathVisio 3: An extendable pathway analysis toolbox. PLoS Comput. Biol. 11, 1–14 (2015).
Ho Sui, S. J. et al. oPOSSUM: Identification of over-represented transcription factor binding sites in co-expressed genes. Nucleic Acids Res. 33, 3154–3164 (2005).
Pavesi, G., Mereghetti, P., Mauri, G. & Pesole, G. Weeder web: Discovery of transcription factor binding sites in a set of sequences from co-regulated genes. Nucleic Acids Res. 32, 199–203 (2004).
Alshalalfa, M. & Alhajj, R. Using context-specific effect of miRNAs to identify functional associations between miRNAs and gene signatures. BMC Bioinform. 14, 1–13 (2013).
Ambros, V. R. The function of animal MicroRNAs. Nature https://doi.org/10.1038/nature02871 (2019).
Liu, W. & Wang, X. Prediction of functional microRNA targets by integrative modeling of microRNA binding and target expression data. Genome Biol. 20, 1–10 (2019).
Chen, Y. & Wang, X. miRDB: An online database for prediction of functional microRNA targets. Nucleic Acids Res. 48, 127–131 (2020).
Author information
Authors and Affiliations
Contributions
SZA: Collected materials & interpreted the data; MIQ: Designed and write the manuscript; SAM: Directed the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Abbas, S.Z., Qadir, M.I. & Muhammad, S.A. Systems-level differential gene expression analysis reveals new genetic variants of oral cancer. Sci Rep 10, 14667 (2020). https://doi.org/10.1038/s41598-020-71346-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-71346-7
This article is cited by
-
Artificial intelligence in cancer target identification and drug discovery
Signal Transduction and Targeted Therapy (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.