Abstract
Small-scale studies have suggested a link between the human gut microbiome and highly prevalent diseases. However, the extent to which the human gut microbiome can be considered a determinant of disease and healthy aging remains unknown. We aimed to determine the spectrum of diseases that are linked to the human gut microbiome through the utilization of its genetic determinants as a proxy for its composition. 180 single nucleotide polymorphisms (SNPs) known to influence the human gut microbiome were used to assess the association with health and disease outcomes in 422,417 UK Biobank participants. Potential causal estimates were obtained using a Mendelian randomization (MR) approach. From the total sample analysed (mean age was 57 ± 8 years), 194,567 (46%) subjects were male. Median exposure was 66-person years (interquartile range 59–72). Eleven SNPs were significantly associated with 28 outcomes (Bonferroni corrected P value < 4.63·10−6) including food intake, hypertension, atopy, COPD, BMI, and lipids. Multiple SNP MR pointed to a possible causal link between Ruminococcus flavefaciens and hypertension, and Clostridium and platelet count. Microbiota and their metabolites might be of importance in the interplay between overlapping pathophysiological processes, although challenges remain in establishing causal relationships.
Similar content being viewed by others
Introduction
It has been suggested that different diseases may share more pathophysiological mechanisms than initially assumed1,2,3,4,5,6. An enhanced understanding of these complex shared disease causing mechanisms is of paramount importance to further improve our strategies to study, prevent and treat diseases.
One of the possible shared systems is the human gut microbiome, which has been suggested as a key inter-player in a variety of individual disease entities7. For example, the gut microbiome has been linked to immune system activation, inflammatory processes and metabolic phenotypes. Moreover, it has been associated with thrombosis and the development of cardiovascular disease (CVD)8,9,10,11,12,13,14,15,16,17, Alzheimer's disease, and cancer18,19,20. Human genetic variants have been linked to the microbiome composition21,22,23 and with this knowledge, rather than actual measurements of the microbiome itself, human genetic variants can be used as a proxy to inform about human gut microbiome composition. This is of interest because large cohorts in which genetic information is available can be studied even if the microbiome itself has not been measured. In addition, the use of genetic determinants can be useful to study the potential long-term health effects of having increased or decreased levels of specific gut microbiota.
Although several associations between the human gut microbiome and individual diseases have been reported, a broad characterization of a large spectrum of health and disease states is lacking but is highly desired as it may help identify potentially common mechanistic pathways17,24,25. Hence, this study aimed to explore the associations between known genetic determinants of the human gut microbiome and the presence of health and its determinants (absence of disease, vital and blood biomarkers and food intake) and disease states in a very large and comprehensive population cohort and investigate possible causal mechanisms using a Mendelian randomization (MR) approach.
Results
Population characteristics
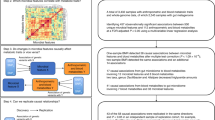
We studied 422,417 unrelated individuals with a mean age of 57 ± 8 years, of which 194,567 (46%) were male (Table 1). Figure 1 presents a flowchart of the study sample selection. Median follow-up was 6 years (interquartile range (IQR) 5–7) and the median total exposure was 66-person years (interquartile range (IQR) 59–72).
Phenome-wide scan on outcomes
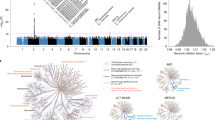
A total of eleven microbiome SNPs was associated with 28 outcomes after a stringent Bonferroni correction (P value < 4.63·10−6) (Fig. 2). The strongest associations with binary traits were observed with diseases related to the following general ICD-10 categories: circulatory system, respiratory system, and respiratory/skin. Complementary information on associations between the SNPs, bacteria, and clinical outcomes is provided in Supplementary Table 1.
Manhattanplot for associations between microbiome SNPs and health and disease outcomes and continuous outcomes. On the x-axis, the health and disease outcomes (according to the ICD 10 code), and continuous are shown. On the y-axis, the P-value is shown. The red line indicates the significance threshold using Bonferroni correction.
An even large number of significant associations with linear traits was discovered; especially health-related measurement including food intake, hemodynamic parameters, anthropometric and laboratory measurements were highlighted (Fig. 2). Extended information on the associations between the SNPs, bacteria, and continuous outcomes is provided in Supplementary Table 2. We identified no significant interactions between sex and genetically determined changes in microbiome.
Mendelian randomization analyses
Single SNP
Single SNP Mendelian randomization (MR) analyses using the Wald estimate were performed for all 28 outcomes. All Wald estimates of all 28 exposure-outcomes were significant (highest P value 4.4·10−6). Results are shown in Fig. 3A, are schematically depicted in Supplementary Figure 1 and full results can be found in Supplementary Table 3. F-statistics were all higher than 10, indicating little chance of weak-instrument bias based on the summary statistics. MR-steiger filtering indicated reversed causation to be likely in the association between Clostridium cellulolyticum and platelet count (P steiger < 5.6·10−119).
(A) Heatmap showing the Wald β’s of the associations between microbiome SNPs and health and disease outcomes, and continuous outcomes. Only significant associations (P value < 4.63·10−6) are shown. BMI body mass index, COPD chronic obstructive pulmonary disease, HDL high density lipoprotein, MAF minor allele frequency. (B) Heatmap showing the inverse variance weighted random effects β’s of the associations between the genetically determined increase of bacteria (family, genus and species level if possible) and health and disease outcomes, and continuous outcomes. Please note that the inverse variance weighted random effects β’s are in some cases liberal, considering the Rucker framework indicated unbalanced horizontal pleiotropy in the estimates as indicated by a significant (P < 0.05) Q–Q′ (and thus the MR-Egger estimate to be a better fit, please see Supplementary Table 5 for these results). BMI body mass index, COPD chronic obstructive pulmonary disease. # indicates suggestively significant.
Under the ideal scenario that all MR assumptions are met, one could cautiously conclude the following based on these results. Genetically determined higher levels of Clostridium cluster IV were associated with a decreased risk for COPD (β − 0.34, SE 0.05, P = 1.96·10−10) and a decreased risk for atopy (β − 0.54, SE 0.04, P = 1.45·10−37). Genetically determined higher levels of the species Ruminococcus flavefaciens were associated with an increased risk for hypertension (β 0.05, SE 0.01, P = 4.97·10−7).
Genetically determined higher levels of the genera Slackia and Pseudobutyrivibrio were associated with an increase in eosinophil count. Higher levels of the species Clostridium xylanovorans were associated with a decrease in resting heart rate and body mass index (BMI). An increase of the genus Desulfovibrio was also associated with a decrease in resting heart rate. Genetically determined higher levels of Methanobacteria on family level were associated with an increase in alcohol intake frequency. Conversely, it was associated with a decrease in mean arterial blood pressure, systolic blood pressure, diastolic blood pressure, BMI, waist hip ratio, reticulocyte count, and erythrocyte count. Interestingly, genetically determined higher levels of the genus Bifidobacterium were associated with increased levels over cholesterol, apolipoprotein A1, HDL, and grip strength. This might seem paradoxical to previous findings in animals and human studies in which a favourable effect of specific Bifidobacterium strains on obesity and cholesterol levels was observed26,27,28.
A genetically increase in the genus Shigella was associated with an increase in reticulocyte count. Higher levels of Clostridium cluster IV were associated with a decrease in eosinophil count. Genetically determined higher levels of the species Ruminococcus flavefaciens were associated with an increase in systolic blood pressure and mean arterial pressure. Lastly, genetically determined higher levels of the species Lactivibrio alcoholicus were associated with a decrease in lymphocyte count.
Combined SNPs
To investigate whether the MR assumptions were truly met, we increased the amount of SNPs to assess the exposure-outcome association (Supplementary Table 3).
We were able to pool SNPs for a total of 26 exposure-outcome associations (Supplementary Table 4). Heterogeneity was indicated using Cochran’s Q statistic for all exposure-outcome associations (P < 0.05, Supplementary Table 5), except for the Clostridiaceae family and Clostridium genus with platelet count and the genus Ruminococcus with hypertension. This indicates at least balanced pleiotropy to be affecting the results. Given that assumptions of inverse variance weighted (IVW) effects model are violated under this scenario, we took forward the IVW random effects model as most liberal model (Fig. 3B, Supplementary Table 4). Compared to 27 causal estimates provided by the Wald ratio (one excluded by MR-Steiger filtering), now only 3 exposure-outcome associations remained suggestively significant [i.e. P < 0.05, but not significant for multiple testing (0.05/28)]. These were the causal estimates of the Clostridiaceae family and Clostridium genus with platelet count (β 0.43, SE 0.16, P = 7.28·10−3 and β 0.43, SE 0.19, P = 1.99·10−2; respectively) and the genus Ruminococcus with hypertension (β 0.45, SE 0.18, P = 1.11·10−2).
Intriguingly, the association between Clostridiaceae/Clostridium and platelet count remained significant considering the main driver (rs58368959, associated with Clostridium cellulolyticum) was removed as a result of MR-steiger filtering. The results were also robust to several sensitivity analyses, including MR-PRESSO and MR-Lasso, which are more outlier-robust, and the “unweighted” MR, which corrects for potential differences in effect size units as effect sizes depict relative abundance (Supplementary Table 4). The Rucker model showed no differences between the heterogeneity estimates of Cochran’s Q and Rucker’s Q, indicating at the absence of unbalanced horizontal pleiotropy and thus an inverse variance weighted model to be preferred (Supplementary Table 5). The results were not robust to MR-Egger, weighted median and weighted mode analyses (Supplementary Table 4). A forest and scatter plot of the MR between Clostridiaceae/Clostridium and platelet count can be found in Supplementary Figures 2–5.
The association between the genus Ruminococcus with hypertension was also robust to the “unweighted” MR, as expected considering the 2 SNPs included in the MR were obtained from the same study. Further sensitivity analyses could not be performed considering the amount of SNPs. One taxonomic level higher, 11 SNPs were pooled for the Ruminococcaceae family and hence more sensitivity analyses could be performed. The IVW random effect estimate was already insignificant (β = 6.94·10−3, SE = 6.63·10−3, P = 0.29). This suggests that, if the association between genus Ruminococcus and hypertension is true, it is specific for that genus within the Ruminococcaceae family. A forest and scatter plot of the MR between Ruminococcaceae/Ruminococcus and hypertension can be found in Supplementary Figures 6 and 7.
To investigate whether estimated effects of the Methano- and Bifidobacterium on associated outcomes were consistent when considering a community effect of the combined pool of gut bacteria, additional sensitivity analyses were performed (Supplementary Table 6). The results remained similar to the initial single SNP estimates.
Lastly, we performed a look-up in MR-Base, a platform for Mendelian randomization analysis, to explore whether the eleven SNPs were associated with other traits (P-value significance threshold of 5.00·10−8) in other studies than UK Biobank. The variant rs4548017 (Methanobacteria) was associated with ulcerative colitis, inflammatory bowel disease, and psoriasis29,30. The variant rs1446585 (Bifidobacterium) was associated with height31. We additionally performed a look-up in GeneCards for extra information about the genes related to the eleven significantly associated SNPs (Supplementary Table 7).
Discussion
In the present study, we used a data-driven approach to identify human health and diseases parameters that are associated with genetic variants thought to influence the microbiome and identified 28 associations with 11 genetic variants. We found relevant associations with food intake, hypertension, BMI, lipids, atopy, and COPD. The possibility of a causal nature of these associations was tested using a MR approach. Although 27 out of 28 associations were indicated to be causal using a single SNP approach, only three of these associations were consistent across some, but not all sensitivity analyses.
Our findings highlight diseases that have been investigated in previous observational studies32,33,34,35,36,37,38,39,40,41. However, contrary to earlier observational research on the human gut microbiome and CVD, associations with CAD and MI did not reach statistical significance in our phenome-wide scan of microbiome associated SNPs8,34. While this might be due to a lack of study power or a small effect size of the evaluated genetic variants, associations with pathways via which the human gut microbiome may influence the development of CAD or MI (i.e. inflammation, arterial blood pressure, and circulating lipoproteins) were documented. As these factors are strongly related to CVD development, it is highly interesting to further unravel their, by microbiome influenced, effect in the long-term.
The association between a genetically determined increase of Bifidobacterium and elevated lipids is notable, since Bifidobacterium is considered to exert beneficial effects in human health42. Small sample sizes (n = 19–32 humans), differences in microbiome behaviour between obese and lean individuals, and the hypothesis that probiotic supplements may disturb the original composition of the microbiome could be possible explanations for this discrepancy43,44,45. In fact, a previous MR between genetically determined Bifidobacterium and HDL and LDL also found a positive causal estimate46. It is likely that total composition of the gut microbiome and bacteria ratios may eventually tell us more than single particular increases or decreases.
Strengths and limitations
This study is the first to investigate the association between previously established microbiome SNPs and a wide variety of health and disease states simultaneously. The major strengths are the considerable sample size, the variety of explored outcomes without an a priori hypothesis, the prospective design of the UK Biobank study. This study could lead other researchers in certain directions to further explore the role of the gut microbiome in health and disease. In addition, we performed a large amount of MR sensitivity analyses (when enough SNPs were available). Another strength is that the MR was performed at different taxonomic levels. This allowed for the investigation of specific strains (species level) while also being able to increase the number of instruments when investigating at genus or family level. The potential importance was illustrated by the consistent estimates between Clostridiaceae family and Clostridium genus with platelet count, and the inconsistent estimates between the family Ruminococcaceae and genus Ruminococcus with hypertension. We believe this might provide insights how to perform and interpret MR estimates within the microbiome research.
We first address limitations most pertinent to a MR approach within microbiome research. It is of importance to note that the human gut microbiome is shaped for a large part by environmental factors47. However, the importance of a genetic component has been established as well23,48. In addition, the genetic approach allowed us to identify new pathways that are interesting to further unravel in the large cohort of the UK Biobank without real stool samples. A second limitation includes the lack of reliable SNPs associated with the gut microbiome. Although we adapted a fairly stringent P value threshold (5.00·10−8) for inclusion criteria and all F statistics were larger than 10, our results could still influenced by weak-instrument bias. “Winner’s curse”, i.e. overestimation of genetic associations within the dataset in which the SNPs were first identified, is likely considering the failure of replication of previously established SNPs21,48,49,50,51 in newer MWAS47,52. Since a two-sample approach was used without sample overlap, any bias due to weak instruments is directed to the null and therefore does not lead to false positive findings53. Another limitation is that the biological function through which the SNPs influence the gut microbiome is unsure and complex, especially considering the possible bilateral nature of the association within the GWAS exposure (i.e. the human microbiome influencing health status and health status influencing the human microbiome). We therefore performed an array of sensitivity analyses to assess whether the results of the single SNP estimates were robust to pleiotropic effects; this was not the case for most exposure-outcome associations. However, whether associations in multiple SNP MR analyses due to pleiotropic effects or weak instrument bias through the “winner’s curse” cannot be differentiated with the current set of analyses. In addition, we applied MR-Steiger filtering to evaluate potential reversed causation. We were unable to validate that the traits investigated share the same causal variant at a particular locus for both the exposure and outcome using colocalization methods, as we did not have the full Linkage Disequilibrium structure of the SNPs investigated. We did use MR-Base to check whether the SNPs were already known to be associated with other traits.
Considering the drawbacks of the current approach, we are very cautious in labelling the associations as “causal”. The current study should be considered as a broad hypothesis free scan in which causality of some associations is strengthened by contextualisation with previously described possible mechanisms (for example, the known risk factor hypertension in the development of CVD). In addition, it highlights the need for strict MR analyses in the microbiome context, as more extensively discussed previously54.
We believe that current study is of additive value to the multi-omics approach required to further dissect the role of the human microbiome in human health and disease55.
Future perspectives
Our study could constitute a useful tool to identify bacteria of interest in order to thoroughly investigate the mechanisms between these bacteria and clinical outcomes; not only in the field of cardiology, but also in other fields. Second, the human gut microbiome might be used as a biomarker in disease risk stratification. Third, the observed associations can point towards new possibilities for therapeutic treatment. Furthermore, the abundance of particular bacteria or their metabolites could influence therapeutic treatment efficacy, as recently demonstrated in the treatment of Parkinson’s disease56.
Conclusions
Human genetic determinants of the gut microbiome are associated with 28 specific health and disease outcomes including hypertension, atopy, COPD, lipids, and BMI. Multiple SNP MR pointed to a possible causal link between Ruminococcus flavefaciens and hypertension, and Clostridium and platelet count. Microbiota and their metabolites might be of importance in the interplay between overlapping pathophysiological processes, and could serve as potential therapeutic targets for the maintenance of health and prevention and treatment of diseases. However, many challenges remain in establishing causal relationships using current genetic data and approaches.
Methods
Study population
Data from the UK Biobank was accessed and analysed under an approved research proposal (application ID: 12006). The UK Biobank is a large community-based prospective study in the United Kingdom that recruited approximately 500,000 participants aged 40 to 69 years old with the general objective to improve the prevention, diagnosis, and treatment of diseases. All methods were performed in accordance with the relevant guidelines and regulations.The study design and population have been extensively described elsewhere57. All participants provided informed consent to participate58. The UK Biobank obtained approval from the relevant institutional review boards, that is, the North West Multi-centre Research Ethics Committee for the UK, the National Information Governance Board for Health and Social Care for England and Wales, and the Community Health Index Advisory Group for Scotland59.
Identification of single nucleotide polymorphisms
Single nucleotide polymorphisms associated with the gut microbiome were collected from 6 previous GWASes21,22,50,51,52,60, using a genome-wide significance threshold of P < 5 × 10−8 for their inclusion in the current study. If the same SNPs and effect sizes were found across different taxonomic level within a single study, we took forward the association with the most differentiated taxonomic level to reduce multiple testing burden. SNPs were clumped within study per bacterium (on the lowest taxonomic level available) using the PLINK (version 1.961) clumping procedure to prune genetic variants at a stringent linkage disequilibrium (LD) of R2 < 0.005 within a five megabase window. We did not clump the 25 SNPs from Goodrich et al. due to the character of the study (twin study instead of GWAS). In the end, this resulted in a total of 180 SNPs associated with the gut microbiome. For more information on the SNP selection, please see Table 2.
Genotyping and imputation
The genotyping process and arrays used in the UK Biobank study have been previously described59. Briefly, participants were genotyped using the custom UK Biobank Lung Exome Variant Evaluation Axiom (Affymetrix; n = 49,949), which includes 807,411 SNPs, or the UK Biobank Axiom array (Affymetrix; n = 452,713), which includes 820,967 SNPs59,62. The arrays share over 95% of insertion and deletion markers59,62. Imputed genotype data were provided by the UK Biobank, based on merged UK10K and 1000 Genomes phase 3 panels63. Participants were excluded in case of missing genotype or sex mismatch (n = 378). Participants with high missingness or excess heterozygosity were also excluded (n = 963). Furthermore, 64,535 participants were excluded because of familial relatedness and being of non-Caucasian British descent. Lastly, participants without information available on the previously reported SNPs associated with the composition of the gut microbiome were also excluded (n = 14,243)21,22. Figure 1 presents a flowchart of the study sample selection.
Definition of (cardiovascular) health, prevalent disease and new-onset disease
Two “health state” variables were generated. Cardiovascular health was defined as the absence of type 2 diabetes mellitus (DM2), stroke, and myocardial infarction (MI)64. Total health was based on the WHO top 10 causes of deaths in high-income countries. It was defined as the absence of coronary artery disease (CAD), stroke, Alzheimer’s disease, lung cancer, chronic obstructive pulmonary disorder (COPD), pneumonia, colon cancer, rectum cancer, DM, kidney diseases, and breast cancer at baseline and during follow-up65. The diseases was defined using self-reported diagnoses and medication (Hospital Episode Statistics data) as well as ICD-9 and ICD-10 codes as described previously66. Definitions of prevalent (established) and incident (new-onset) disease (outcomes) are shown in Supplementary Table 8.
Vital signs, blood count, and food intake
Vital signs and biological samples were collected during the baseline visit together with data of self-completed questionnaires (including food questionnaires), interviews, and physical measurements. Blood pressure was measured twice for consistency and the average value was used. Automated measurements were corrected according to previously described methodology67.
Phenome-wide scan on outcomes
Outcomes with a prevalence of ≤ 1% were excluded from the analyses. Logistic regression modelling was performed to assess the effect of microbiome SNPs on combined prevalent and new-onset diseases, and health outcomes in term of the resulting odds ratios (OR) and corresponding 95% confidence intervals (CIs). Genotypes were coded according to a dose–response model (0, 1, 2), where the allele coded with a 2 corresponded with an increased abundance of a certain bacterium. Linear regression analyses were conducted to assess the effect of microbiome SNPs on continuous outcome measures (anthropometric, hemodynamic and laboratory variables as well as longevity and food intake at baseline) in terms of the resulting β coefficients and associated standard errors (SE). Since sex differences exist regarding the human gut microbiome, we tested for sex-interactions in the association between the SNPs and all outcomes. All regression analyses were adjusted for age (at the moment of the last follow-up or age at baseline visit), sex (except when sex-interactions were tested), genotyping chip, and the first 30 principal components (to adjust for population structure) provided by the UK Biobank. A stringent Bonferroni correction was used to correct for multiple testing (180 SNPs, 60 outcomes; 0.05/10,800 = P value < 4.63·10−6). Outcomes reaching statistical significance after Bonferroni correction were considered of interest for further analyses (see Mendelian randomization section below). Analyses described above are performed using Stata version 15 (StataCorp, Texas, United States).
Mendelian randomization analyses
In order to explore the possibility of a causal relationship between the exposure (the gut microbiome) and the outcomes reaching statistical significance after Bonferroni correction, we first performed a single SNP Mendelian randomization (MR) using the Wald-estimate68. Weak instrument bias was assessed per SNP using the F-statistic, calculated with the following formula: F = R2(n − 2)/(1 − R2). In this formula, n is the sample size of the exposure and R2 is the amount of variance of the exposure explained by the SNP69. R2 was calculated using a previously established formula70. Allele frequencies and SNP position were obtained for the SNPs obtained from the study of Wang et al.71. In order to explore the possibility of reversed causation to be a driver of the current results, we applied MR-Steiger filtering. MR-Steiger filtering calculates the R2 for the exposure and outcome and removes variants if the R2 of the exposure is significantly lower (P value < 0.05) than the R2 of the outcome72. R2 for linear traits was calculated as mentioned above70. R2 for binary outcomes was calculated on the liability scale according to previously established methods73. These results can be found in Supplementary Table 3.
Pleiotropy within MR analyses refers to a genetic variant having multiple effects and distorts MR estimates when genetic variants affect the outcome independently. Given the uncertainty of the biological mechanisms underlying the SNP-exposure association, and hence uncertainty on whether the genetic variants affect the outcome independently, we aimed to gain additional insight in pleiotropy by increasing the amount of SNPs to assess the exposure-outcome association. SNPs associated with the same exposure and their effect estimates on the outcome (possibly not reaching Bonferroni significance) were obtained from the pool of all 180 SNPs. This resulted in an additional 153 exposure-outcome associations, for which single SNP MR estimates, F-statistics69 and MR-Steiger72 results can be found in Supplementary Table 3 as well. By definition, effect sizes used to obtain pooled estimates should reflect the same unit. We therefore only took forward SNPs with effect sizes reflecting relative abundance, given the fact that (a) only one SNP (rs4548017) reached statistical significance on the tested outcomes after Bonferroni correction, (b) 169 out of 180 SNPs reflected relative abundance and c) the relative effects better reflect the biology of the milieu intérieur. Due to difference in in-depth classification of the taxonomic rank in the SNP-exposure association between and within study, SNPs were pooled on species, genus and family level. Again, SNPs were clumped (between-studies) for every bacterium per level. Detailed information on the amount of SNPs per bacterium per taxonomic level and the outcomes tested can be found in Supplementary Table 4. Three methods were applied based on the amount of genetic variants available: (a) if after pooling only one SNP remained available, only Wald ratio, MR-Steiger filtering72 and F-statistics69 were calculated as explained above (28 exposure-outcome associations, see Supplementary Table 3), (b) if two SNPs were available, inverse variance weighted fixed effects and random effects were calculated; I2- index74 and Cochran’s Q75were used to assess heterogeneity and differentiate between models (6 exposure-outcome associations) (Supplementary Tables 4 and 5) and (c) if more than two SNPs were available, a more extensive pallet of analyses was performed (22 exposure-outcome associations) (Supplementary Tables 4 and 5). First of all, this extended pallet included the Rucker framework. Heterogeneity was and thus potential pleiotropy was first assessed in the IVW effect estimate using Cochran’s Q statistic. A P value of < 0.05 was considered as an indication of at least balanced horizontal pleiotropy, indicating the IVW random effects model to be preferred over a fixed effects model. heterogeneity and, as a consequence, of pleiotropy. Next, the MR-Egger test was performed76. The MR-Egger relies on different assumptions than the IVW method, as it does not assume all instruments to be valid by allowing for a non-zero intercept. The intercept therefore represents unbalanced horizontal pleiotropy. The MR-Egger intercept, the standard error and p-value were therefore calculated. A MR-Egger intercept P value of < 0.05 was considered to be an indicator of unbalanced horizontal pleiotropy. Heterogeneity was assessed within the MR-Egger by calculating Rucker’s Q. A significant difference between these two heterogeneity statistics (Q–Q′, P < 0.05) was considered indicative of unbalanced horizontal pleiotropy as well and thus the MR-Egger test to be the preferred method over the IVW. I2GX was calculated to assess weak-instrument bias within MR-Egger estimates77. An I2GX of >> 95% was considered to indicate a low risk of weak-instrument bias within the MR-Egger test. In addition, MR-Lasso78 and MR-PRESSO79 (the latter when 4 or more SNPs were available) were performed, two tests that are robust to outliers due horizontal pleiotropy when the SNPs used are valid instrumental variables. Lastly, weighted median80 and weighed mode81 effect estimates were calculated. These methods allow for estimation of causal effect estimates when the majority (> 50%) or the plurality (no larger subset of invalid instruments estimating the same causal parameter) of SNPs are valid instruments.
Under scenario (b) and (c), we additionally performed an “unweighted” MR by calculated an inverse variance weighted fixed and random effects estimate after equally weighting all genetic instruments (i.e. negative and positive beta’s were set to − 1 and 1 respectively) and were assumed to be estimated with infinite precision (i.e. the stand error was set to 0). This was performed considering that relative abundance might still reflect different biological implications between studies/populations. We recalculated Cochran’s Q using these fixed weights to differentiate between the fixed and random effects model.
MR analyses were performed using R (version 3.6.3), the TwoSampleMR package 0.5.382, MR-PRESSO79 (version 1.0), and the source code for MR-Lasso78 in a Two-Sample MR setting as proposed in the article of Rees et al.
Conditional analyses on the methano- and bifidobacterium
To investigate whether estimated effects of the Methano- and Bifidobacterium on associated outcomes were consistent when considering a community effect of the combined pool of gut bacteria, we performed additional sensitivity analyses. First, we took forward SNPs containing (a) taxonomic information up to family level and (b) effect sizes depicting relative abundance, totalling to 136 SNP. After clumping on family level, 32 unweighted GRS for 32 different bacterial families were constructed. These families included the Acidaminococcaceae, Bacteroidaceae, Barnesiellaceae, Bifidobacteriaceae, Clostridiaceae, Coriobacteriaceae, Desulfovibrionaceae, Enterobacteriaceae, Erysipelotrichaceae, Eubacteriaceae, Firmicutes, Lachnospiraceae, Lactobacillaceae, Leuconostocaeae, Marinilabiliaceae, Micrococcaceae, Mogibacteriaceae, Moraxellaceae, Odoribacteraceae, Paenibacillaceae, Pasteurellaceae, Peptococcaceae, Peptoniphilaceae, Porphyromonadaceae, Rhodospirillaceae, Rikenellaceae, Ruminococcaceae, Streptococcaceae, Synergistaceae, Veillonellaceae, Victivallaceae and the Bifidobacteriaceae/Methanobacteriaceae family (for the association of the Methanobacteriaceae/Bifidobacteriaceae respectively). We opted for an unweighted GRS based on the same rationale as the choice for performing an “unweighted” MR as sensitivity analysis. These were subsequently used as covariates in the SNP (rs4548017 and rs1446585 for the Methano- and Bifidobacteriaceae, respectively)-outcome association in the UK Biobank.
Look-ups
We performed a look-up in MR-Base, a platform for Mendelian randomization analysis, to explore whether the SNPs would be associated with other traits in order to gain insights in potential pleiotropic effects82. Furthermore, we consulted GeneCards for extra information about the genes related to the analysed SNPs83.
Data availability
Data is available on request.
References
Valdes, A. M., Walter, J., Segal, E. & Spector, T. D. Role of the gut microbiota in nutrition and health. BMJ 361, k2179 (2018).
Ruscitti, P. et al. Adipocytokines in rheumatoid arthritis: the hidden link between inflammation and cardiometabolic comorbidities. J. Immunol. Res. 2018, 8410182 (2018).
Zhao, T. X. & Mallat, Z. Targeting the immune system in atherosclerosis: JACC state-of-the-art review. J. Am. Coll. Cardiol. 73, 1691–1706 (2019).
Dick, S. A. & Slava, E. Chronic heart failure and inflammation. Circ. Res. 119, 159–176 (2016).
Lu, Y. et al. Sex differences in inflammatory markers and health status among young adults with acute myocardial infarction: results from the VIRGO (Variation in Recovery: Role of Gender on Outcomes of Young Acute Myocardial Infarction Patients) Study. Circ. Cardiovasc. Qual. Outcomes 10, e003470 (2017).
Yu, L. et al. Pattern recognition receptor-mediated chronic inflammation in the development and progression of obesity-related metabolic diseases. Mediators Inflamm. 2019, 5271295 (2019).
Clemente, J. C., Manasson, J. & Scher, J. U. The role of the gut microbiome in systemic inflammatory disease. BMJ 360, j5145 (2018).
Wang, Z. et al. Gut flora metabolism of phosphatidylcholine promotes cardiovascular disease. Nature 472, 57–65 (2011).
Zhou, X. et al. The gut microbiome contributes to a substantial proportion of the variation in blood lipids. Nature 271, 1689–1699 (2018).
Allayee, H. & Hazen, S. L. Contribution of gut bacteria to lipid levels: another metabolic role for microbes?. Circ. Res. 117, 750–754 (2015).
Falony, G. et al. Population-level analysis of gut microbiome variation. Science 352, 560–564 (2016).
Zununi Vahed, S., Barzegari, A., Zuluaga, M., Letourneur, D. & Pavon-Djavid, G. Myocardial infarction and gut microbiota: an incidental connection. Pharmacol. Res. 129, 308–317 (2018).
Kasselman, L. J., Vernice, N. A., DeLeon, J. & Reiss, A. B. The gut microbiome and elevated cardiovascular risk in obesity and autoimmunity. Atherosclerosis 271, 203–213 (2018).
Vinjé, S., Stroes, E., Nieuwdorp, M. & Hazen, S. L. The gut microbiome as novel cardio-metabolic target: the time has come!. Eur. Heart J. 35, 883–887 (2014).
Brown, J. M. & Hazen, S. L. Microbial modulation of cardiovascular disease. Nat. Rev. Microbiol. https://doi.org/10.1038/nrmicro.2017.149 (2018).
Tang, W. H. W., Li, D. Y. & Hazen, S. L. Dietary metabolism, the gut microbiome, and heart failure. Nat. Rev. Cardiol. https://doi.org/10.1038/s41569-018-0108-7 (2018).
Young, V. B. The role of the microbiome in human health and disease: an introduction for clinicians. BMJ 356, j831 (2017).
Zhuang, Z.-Q. et al. Gut microbiota is altered in patients with Alzheimer’s disease. J. Alzheimers Dis. 63, 1337–1346 (2018).
Jiang, C., Li, G., Huang, P., Liu, Z. & Zhao, B. The gut microbiota and Alzheimer’s disease. J. Alzheimers. Dis. 58, 1–15 (2017).
Zhu, J. et al. Breast cancer in postmenopausal women is associated with an altered gut metagenome. Microbiome 6, 136 (2018).
Bonder, M. J. et al. The effect of host genetics on the gut microbiome. Nat. Genet. 48, 1407–1412 (2016).
Goodrich, J. K. et al. Genetic determinants of the gut microbiome in UK twins. Cell Host Microbe 19, 731–743 (2016).
Goodrich, J. K. et al. Human genetics shape the gut microbiome. Cell 159, 789–799 (2014).
van den Munckhof, I. C. L. et al. Role of gut microbiota in chronic low-grade inflammation as potential driver for atherosclerotic cardiovascular disease: a systematic review of human studies. Obes. Rev. https://doi.org/10.1111/obr.12750 (2018).
Brown, J. M. & Hazen, S. L. Targeting of microbe-derived metabolites to improve human health: the next frontier for drug discovery. J. Biol. Chem. 292, 8560–8568 (2017).
An, H. M. et al. Antiobesity and lipid-lowering effects of Bifidobacterium spp. in high fat diet-induced obese rats. Lipids Health Dis. 10, 116 (2011).
Xiao, J. Z. et al. Effects of milk products fermented by Bifidobacterium longum on blood lipids in rats and healthy adult male volunteers. J. Dairy Sci. 86, 2452–2461 (2003).
Cerdo, T., Garcia-Santos, J. A., Bermudez, M. G. & Campoy, C. The role of probiotics and prebiotics in the prevention and treatment of obesity. Nutrients 11, 635 (2019).
Tsoi, L. C. et al. Identification of 15 new psoriasis susceptibility loci highlights the role of innate immunity. Nat. Genet. 44, 1341–1348 (2012).
Liu, J. Z. et al. Association analyses identify 38 susceptibility loci for inflammatory bowel disease and highlight shared genetic risk across populations. Nat. Genet. 47, 979–986 (2015).
Wood, A. R. et al. Defining the role of common variation in the genomic and biological architecture of adult human height. Nat. Genet. 46, 1173–1186 (2014).
Branchereau, M., Burcelin, R. & Heymes, C. The gut microbiome and heart failure: a better gut for a better heart. Rev. Endocr. Metab. Disord. https://doi.org/10.1007/s11154-019-09519-7 (2019).
Kummen, M. et al. Gut microbiota signature in heart failure defined from profiling of 2 independent cohorts. J. Am. Coll. Cardiol. 71, 1184–1186 (2018).
Wilson Tang, W. H., Kitai, T. & Hazen, S. L. Gut microbiota in cardiovascular health and disease. Circ Res. 120, 1183–1196 (2018).
Larson, N. B. et al. ABO blood group associations with markers of endothelial dysfunction in the multi-ethnic study of atherosclerosis. Atherosclerosis 251, 422–429 (2016).
Tang, W. H. W., Li, D. Y. & Hazen, S. L. Dietary metabolism, the gut microbiome, and heart failure. Nat. Rev. Cardiol. 16, 137–154 (2019).
Fu, J. et al. The gut microbiome contributes to a substantial proportion of the variation in blood lipids. Circ. Res. 117, 817–824 (2015).
Huart, J. et al. Gut microbiota and fecal levels of short-chain fatty acids differ upon 24-hour blood pressure levels in men. Hypertension 74, 1005–1013 (2019).
Boer, C. G. et al. Intestinal microbiome composition and its relation to joint pain and inflammation. Nat. Commun. 10, 4881 (2019).
Li, F., Wang, M., Wang, J., Li, R. & Zhang, Y. Alterations to the gut microbiota and their correlation with inflammatory factors in chronic kidney disease. Front. Cell. Infect. Microbiol. 9, 206 (2019).
Sun, S. et al. Gut microbiota composition and blood pressure. Hypertension 73, 998–1006 (2019).
Turroni, F., Berry, D. & Ventura, M. Editorial: bifidobacteria and their role in the human gut microbiota. Front. Microbiol. 7, 2148 (2017).
Jones, R. B. et al. Probiotic supplementation increases obesity with no detectable effects on liver fat or gut microbiota in obese Hispanic adolescents: a 16-week, randomized, placebo-controlled trial. Pediatr. Obes. 13, 705–714 (2018).
Bouter, K. E., van Raalte, D. H., Groen, A. K. & Nieuwdorp, M. Role of the gut microbiome in the pathogenesis of obesity and obesity-related metabolic dysfunction. Gastroenterology 152, 1671–1678 (2017).
Turnbaugh, P. J. et al. A core gut microbiome in obese and lean twins. Nature 457, 480–484 (2009).
Yang, Q., Lin, S. L., Kwok, M. K., Leung, G. M. & Schooling, C. M. The roles of 27 genera of human gut microbiota in ischemic heart disease, type 2 diabetes mellitus, and their risk factors: a mendelian randomization study. Am. J. Epidemiol. 187, 1916–1922 (2018).
Rothschild, D. et al. Environment dominates over host genetics in shaping human gut microbiota. Nature 555, 210–215 (2018).
Goodrich, J. K. et al. Genetic determinants of the gut microbiome in UK twins. Cell Host Microbe 19, 731–743 (2017).
Wang, J., Thingholm, L. B., Skiecevičienė, J., Rausch, P. & Kummen, M. Genome-wide association analysis identifies variation in vitamin D receptor and other host factors influencing the gut microbiota. Nat Genet 48, 1396–1406 (2016).
Turpin, W. et al. Association of host genome with intestinal microbial composition in a large healthy cohort. Nat. Genet. 48, 1413–1417 (2016).
Davenport, E. R. et al. Genome-wide association studies of the human gut microbiota. PLoS ONE 10, e0140301 (2015).
Scepanovic, P. et al. A comprehensive assessment of demographic, environmental, and host genetic associations with gut microbiome diversity in healthy individuals. Microbiome 7, 130 (2019).
Pierce, B. L. & Burgess, S. Efficient design for Mendelian randomization studies: subsample and 2-sample instrumental variable estimators. Am. J. Epidemiol. 178, 1177–1184 (2013).
Wade, K. H. & Hall, L. J. Improving causality in microbiome research: can human genetic epidemiology help?. Wellcome Open Res. 4, 199 (2019).
Cani, P. D. Human gut microbiome: hopes, threats and promises. Gut 67, 1716–1725 (2018).
van Kessel, S. P. et al. Gut bacterial tyrosine decarboxylases restrict levels of levodopa in the treatment of Parkinson’s disease. Nat. Commun. 10, 310 (2019).
UK Biobank. UK Biobank: Protocol for a Large-Scale Prospective Epidemiological Resource (2007).
UK Biobank. UK Biobank Ethics and Governance Framework 2007 (2007).
Bycroft, C. et al. Genome-wide genetic data on ~500,000 UK Biobank participants. bioRxiv https://doi.org/10.1101/166298 (2017).
Wang, J. et al. Genome-wide association analysis identifies variation in vitamin D receptor and other host factors influencing the gut microbiota. Nat. Genet. 48, 1396–1406 (2016).
Purcell, S. et al. PLINK: a tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 81, 559–575 (2007).
UK Biobank. Genotyping and Quality Control of UK Biobank, A Large-Scale, Extensively Phenotyped Prospective Resource (2018).
Marchini, J. UK Biobank Phasing and Imputation Documentation. 13 (2018).
Di Angelantonio, E. et al. Association of cardiometabolic multimorbidity with mortality. JAMA 314, 52–60 (2015).
WHO. The top 10 causes of death (2018). https://www.who.int/news-room/fact-sheets/detail/the-top-10-causes-of-death. Accessed 10th Dec 2018.
Abdullah Said, M., Eppinga, R. N., Lipsic, E., Verweij, N. & van der Harst, P. Relationship of arterial stiffness index and pulse pressure with cardiovascular disease and mortality. J. Am. Heart Assoc. https://doi.org/10.1161/JAHA.117.007621 (2018).
Stang, A. et al. Algorithms for converting random-zero to automated oscillometric blood pressure values, and vice versa. Am. J. Epidemiol. 164, 85–94 (2006).
Wald, A. The fitting of straight lines if both variables are subject to error. Ann. Math. Stat. 11, 284–300 (1940).
Palmer, T. M. et al. Using multiple genetic variants as instrumental variables for modifiable risk factors. Stat. Methods Med. Res. 21, 223–242 (2012).
Teslovich, T. M. et al. Biological, clinical and population relevance of 95 loci for blood lipids. Nature 466, 707–713 (2010).
Ward, L. D. & Kellis, M. HaploReg: a resource for exploring chromatin states, conservation, and regulatory motif alterations within sets of genetically linked variants. Nucleic Acids Res. 40, D930–D934 (2012).
Hemani, G., Tilling, K. & Davey Smith, G. Orienting the causal relationship between imprecisely measured traits using GWAS summary data. PLoS Genet. 13, e1007081 (2017).
Lee, S., Goddard, M., Wray, N. & Visscher, P. A better coefficient of determination for genetic profile analysis. Genet. Epidemiol. 36, 214–224 (2012).
Del Greco, M. F., Minelli, C., Sheehan, N. A. & Thompson, J. R. Detecting pleiotropy in Mendelian randomisation studies with summary data and a continuous outcome. Stat. Med. 34, 2926–2940 (2015).
Bowden, J. et al. A framework for the investigation of pleiotropy in two-sample summary data Mendelian randomization. Stat. Med. 36, 1783–1802 (2017).
Bowden, J., Davey Smith, G. & Burgess, S. Mendelian randomization with invalid instruments: effect estimation and bias detection through Egger regression. Int. J. Epidemiol. 44, 512–525 (2015).
Bowden, J. et al. Assessing the suitability of summary data for two-sample Mendelian randomization analyses using MR-Egger regression: the role of the I2 statistic. Int. J. Epidemiol. 45, 1961–1974 (2016).
Rees, J. M. B., Wood, A. M., Dudbridge, F. & Burgess, S. Robust methods in Mendelian randomization via penalization of heterogeneous causal estimates. PLoS ONE 14, 1–24 (2019).
Verbanck, M., Chen, C.-Y., Neale, B. & Do, R. Detection of widespread horizontal pleiotropy in causal relationships inferred from Mendelian randomization between complex traits and diseases. Nat. Genet. 50, 693–698 (2018).
Bowden, J., Davey Smith, G., Haycock, P. C. & Burgess, S. Consistent estimation in Mendelian randomization with some invalid instruments using a weighted median estimator. Genet. Epidemiol. 40, 304–314 (2016).
Hartwig, F. P., Davey Smith, G. & Bowden, J. Robust inference in summary data Mendelian randomization via the zero modal pleiotropy assumption. Int. J. Epidemiol. 46, 1985–1998 (2017).
Hemani, G. et al. The MR-base platform supports systematic causal inference across the human phenome. eLife 7, e34408 (2018).
Stelzer, G. et al. GeneCards—The Human Gene Database. https://www.genecards.org/. Accessed 6 Jan 2020.
Acknowledgements
We would like to thank the Centre for Information Technology of the University of Groningen for their support and for providing access to the Peregrine high-performance computing cluster. We thank Ruben N. Eppinga, MD PhD, Tom Hendriks, MD, M. Abdullah Said,MD, M. Yldau van der Ende, MD PhD, Yanick Hagemeijer, MSc, Luis Juarez Orozco, MD PhD, Jan-Walter Benjamins, BEng, and Ming W. Yeung, MSc (Department of Cardiology, University of Groningen, University Medical Center Groningen, Groningen, the Netherlands), for their contributions to the extraction and processing of data in the UK Biobank. None of the contributors received compensation except for their employment at the University Medical Center Groningen. NV is supported by NWO VENI Grant (016.186.125). The funding agencies had no role in the study design, analysis, or interpretation of data; the writing of the manuscript; or in the decision to submit the article for publication.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection and analysis were performed by H.E.G. and Y.J.V. The first draft of the manuscript was written by H.E.G. and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript. All authors take full responsibility for all aspects of the reliability and freedom from bias of the data presented and their discussed interpretation.
Corresponding author
Ethics declarations
Competing interests
All authors have completed and submitted the ICMJE Form for Disclosure of Potential Conflicts of Interest and none were reported. Author NV is an employee of Genomics plc.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Groot, H.E., van de Vegte, Y.J., Verweij, N. et al. Human genetic determinants of the gut microbiome and their associations with health and disease: a phenome-wide association study. Sci Rep 10, 14771 (2020). https://doi.org/10.1038/s41598-020-70724-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-70724-5
This article is cited by
-
The gut microbiota and depressive symptoms across ethnic groups
Nature Communications (2022)
-
Improvement of intestinal flora: accompany with the antihypertensive effect of electroacupuncture on stage 1 hypertension
Chinese Medicine (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.