Abstract
MOG-antibody associated disease (MOG-AAD) is a recently recognized demyelinating disorder predominantly affecting children but also occurs in adults, with a relapsing course in approximately 50% of patients. We evaluated peripheral blood mononuclear cells from MOG-AAD patients by flow cytometry and found a strong antigen specific central memory cell (CMC) response with increased Th1 and Th17 cells at the time of a relapse. Transcriptomic analysis of CMCs by three independent sequencing platforms revealed TNFAIP3 as a relapse biomarker, whose expression was down regulated at a relapse compared to remission in MOG-AAD patients. Serum in an additional cohort of patients showed decreased TNFAIP3 levels at relapse compared to remission state in MOG-AAD patients. Our studies suggest that alterations in TNFAIP3 levels are associated with relapses in MOG-AAD patients, which may have clinical utility as a disease course biomarker and therapeutic target.
Similar content being viewed by others
Introduction
Myelin oligodendrocyte glycoprotein antibody associated diseases (MOG-AAD) have been recently described in approximately 25% of children and 5% of adults with demyelinating disorders1,2. MOG is a component of myelin expressed in the central nervous system (CNS), and MOG antibodies can be detected by cell-based assays in the serum3,4. MOG-AAD exhibits an age-related array of clinical phenotypes ranging from acute disseminated encephalomyelitis (ADEM), clinically isolated syndrome (CIS), optic neuritis (ON), recurrent forms of ADEM and ON, transverse myelitis (TM) and neuromyelitis spectrum disorder (NMO-SD)2,3,5,6. Approximately 50% of patients experience a multiphasic disease course5,7,8. There is an increasing body of literature demonstrating that patients with MOG-AAD have distinct clinical and MRI features from multiple sclerosis (MS)9,10,11. Treatment response varies between MOG-AAD patients, and immunosuppressive treatments including rituximab or mycophenolate mofetil often result in incomplete control of disease12. Moreover, standard multiple sclerosis medications such as beta interferon may exacerbate disease13,14.
Individual patients with MOG-antibodies experience varying disease courses, some with frequent relapses, which can result in significant disability13. One of the main challenges in clinical care is to identify patients who will develop a multiphasic disease course, and require chronic immunotherapy. The persistent presence of high titers of serum MOG antibody titers has been studied in the prognostication of a multiphasic disease and relapses, although with inconsistent results15,16. Patients with MOG-antibodies have also been shown to have a higher frequency of MOG-reactive T cells suggesting an antigen-specific response, which may play a key role in disease initiation17, however have not been studied as disease biomarkers. In classic autoimmune disease, antigen-presenting cells, such as dendritic cells present a specific antigen to T cells to initiate an immune response. In the case of diseases characterized by self-antigen specific antibodies, it is likely that plasma cells and B cells are involved and cooperate with T cells to elicit autoimmunity18. In this study, we evaluate serological and T cell transcriptomic profiling in blood samples from patients with MOG-AAD at the time of a relapse versus remission with the goal of identifying mechanisms and biomarkers of relapse in this disease.
Results
Flow cytometric analysis of MOG-reactive CMC T cells show an increased percentage of IL17+, IFNγ+ and IL17+/IFNγ+ cells
Prior studies have demonstrated an increased T cell response to MOG peptides in MOG-AAD patients compared to controls.17 Here, we sought to evaluate the percentage of IL17+ and IFNγ+ cells in CMCs of untreated MOG-AAD patients (n = 8) and age and sex matched PHCs (n = 7) (Table 1 and Supplementary Table 1a), when stimulated with peptides MOG p1-20, MOG p35-55, MOG p119-130, MOG p181-195, MOG p186-200 and myelin peptides. We found an increase in the percentage of CMCs, as well as IL17+ and IL17+/IFNγ+ (double positive) CMCs in MOG-AAD patients when compared to PHCs (Fig. 1). (CMC-MOG 1–20, P = 0.05, CMC-MOG 119–130, P = 0.03; CMC-IL17+-MOG 35–55, P = 0.02, CMC-IL17+-MOG 119–130, P = 0.03; CMC++MOG 119–130, P = 0.04).
Cytokine expression in MOG-reactive central memory cells (CMCs) from untreated MOG-AAD patients and pediatric healthy controls (PHCs): PBMCs were stimulated and cultured with 10 μg/ml of MOG peptides (p1-20, p35-55, p119-130, p181-195 and p186-200) and cocktail of myelin peptides (Myelin Phospholipid Protein 139–154 (PLP), Myelin Basic Protein (MBP) 13–32, MBP 111–129 and MBP 146–170) for 7 days followed by staining with antibodies and flow cytometry as described in Materials and Methods. Data was analyzed using Flowjo vX.0.7 and the graphs were made using GraphPadPrism version 8.4.2 (464) A Percentage of CMCs (CD4+CCR7+CD45RA−). B Percentage of IL17+producing CMCs (CD4+CCR7+CD45RA-, IL17+). C Percentage of IFNγ+producing CMCs (CD4+CCR7+CD45RA−,IFNγ +). D Percentage of IL17+ and IFNγ+ producing CMCs (CD4+CCR7+CD45RA−,IL17+,IFNγ+). MOG-AAD n = 8, PHC n = 7, Mann–Whitney test, *P < 0.05.
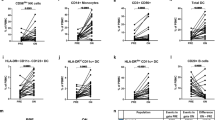
We next identified a group of MOG-AAD patients with a blood sample within 30 days prior to a relapse (n = 9) and during a remission time point (non-relapse, n = 6) (Supplementary Table 1a). Interestingly, we found an increased proportion of IL17+, IFNγ+ and IL17+IFNγ+ (double positive) CMCs after stimulation with several individual MOG peptides in MOG-AAD patients at the time of relapse as compared to remission time point (Fig. 2a–c).
Cytokine expression in MOG-reactive central memory cells (CMCs) from MOG-AAD patients at the time of relapse and remission: PBMCs were stimulated and cultured with 10 μg/ml of MOG peptides (p1-20, p35-55, p119-130, p181-195 and p186-200) and cocktail of myelin peptides (Myelin Phospholipid Protein 139–154, Myelin Basic Protein (MBP) 13–32, MBP 111–129 and MBP 146–170) for 7 days followed by staining with antibodies and flow cytometry as described in Materials and Methods. Data was analyzed using Flowjo vX.0.7 and the graphs were made using GraphPadPrism version 8.4.2 (464) A Percentage of IL17+producing CMCs (CD4+CCR7+CD45RA−, IL17+). B Percentage of IFNγ+producing CMCs (CD4+CCR7+CD45RA−,IFNγ+). C Percentage of IL17+ and IFNγ+ producing CMCs (CD4+CCR7+CD45RA−,IL17+,IFNγ+). Relapse n = 9, Remission n = 6 D CMCs (CD4+CCR7+CD45RA–), CMC IL17+ (CD4+CCR7+CD45RA−,IL17 +), CMC IFNγ+ (CD4+CCR7+CD45RA−, IFNγ+), CMC++ (CD4+CCR7+CD45RA−,IL17+, IFNγ+) analysis from a single pediatric MOG-AAD patient#1 with longitudinal samples, (1) Rm (remission, MOG-AAD#1.2), (2) PR (pre-relapse, MOG-AAD#1.3), (3) R (relapse, MOG-AAD#1.4) and (4) Rm/S (remission treated with steroid, MOG-AAD#1.5).
Furthermore, PBMCs from MOG-AAD patient#1 with 4 longitudinal samples, (1) remission (MOG-AAD#1.2) 2) pre-relapse (MOG-AAD#1.3) time point which was 2 months prior to a relapse during which the patient experienced headache and mild visual symptoms, (3) relapse (MOG-AAD#1.4) characterized by bilateral weakness and multiple new MRI lesions and (4) remission (MOG-AAD#1.5) on treatment with steroid were stimulated with individual MOG peptides. We found a dramatic increase in CMC IL17+ and CMC IFNγ+cells at a relapse. Specifically, there was an increase in CMC IL17+cells responsive to MOG p35-55 in the 2 months prior to fulminant relapse, when the patient had mild visual symptoms (pre-relapse state) (Fig. 2d).
Gene expression analysis revealed and confirmed a relapse biomarker in MOG-AAD patients
Since our initial results demonstrated that CMC T cells might play a role in relapse and in multiphasic disease course in MOG-AAD patients, we sought to further evaluate the transcriptomic profile of these cells. Single cell RNA sequencing (inDrop) was performed on an untreated MOG-AAD patient#1 with 3 longitudinal samples during remission (MOG-AAD#1.2), pre-relapse (MOG-AAD#1.3) and at a relapse (MOG-AAD#1.4). The relapse sample failed sequencing, however the pre-relapse and remission samples were evaluable. Given the evolving understanding that in MOG-AAD, clinical symptoms can occur prior to the presence of new MRI lesions19, moving forward we re-classified the pre-relapse sample as relapse. Analysis was performed as described in “Materials and methods”.
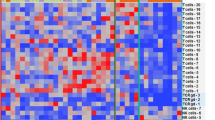
Cluster analysis of all the cells from relapse and remission samples using Seurat indicated that gene expression was homogeneous and there were no obvious differences between cells of relapse and remission samples (Fig. 3a). However, to identify genes that were differentially expressed in relapse and remission samples, we calculated average gene expression between the relapse and remission samples and subtracted gene expression values from relapse and remission samples for each gene. From this analysis, we chose the top 5 differentially expressed genes that were up regulated in the relapse and remission samples. We demonstrated differential expression of TNFAIP3 (Tumor Necrosis Factor Inducible Protein A20), which was higher in the remission sample as compared to relapse sample of a MOG-AAD patient (Fig. 3b). The other top genes identified to be up regulated during remission were HILPDA (Hypoxia Inducible Lipid Droplet-Associated), MXI-1 (MAX Interactor 1, Dimerization Protein), BNIP3 (BCL2 Interacting Protein 3), FAM162A (Family With Sequence Similarity 162 Member A); while genes up regulated during relapse included SNRPG (Small Nuclear Ribonucleoprotein Polypeptide G), EWSR1 (Ewing Sarcoma RNA Binding Protein 1), AL137058.2 (Clone-based Ensemble gene), HMGN1 (High Mobility Group Nucleosome Binding Domain 1), LRRC75A-AS1/ SNHG29 (Small Nucleolar RNA Host Gene 29) (Supplementary Fig. 1).
Gene expression analysis in PBMCs from MOG-AAD patients: Single cell RNA sequencing (inDrop) was performed on an untreated MOG-AAD patient#1 with 2 longitudinal samples, (1) Remission (MOG-AAD#1.2) and (2) Pre-relapse (MOG-AAD#1.3) as described in Materials and Methods. cDNA libraries were sequenced using the Illumina NextSeq 500 platform and analyzed following V3 Indrop criteria. After sequencing the raw BCL files were demultiplexed using bcl2fastq software by illumina (https://support.illumina.com/sequencing/sequencing_software/bcl2fastq-conversion-software.html). Reads obtained from bcl2fastq were further processed using the single-cell RNA-seq pipeline of the bcbio-nextgen (https://bcbio-nextgen.readthedocs.io/en/latest/contents/pipelines.html#single-cell-rna-seq) software suite. The scaled data was further clustered using Seurat and visualized using TSNE (https://www.biorxiv.org/content/early/2018/11/02/460147). A Cluster analysis of a relapse and remission sample. B Differential expression of TNFAIP3 in the relapse and remission sample by single cell sequencing. Digital Gene Expression (DGE) sequencing was performed on an untreated MOG-AAD patient#1 with 3 longitudinal samples, (1) Rm (remission, MOG-AA#1.2), (2) PR (pre-relapse, MOG-AAD#1.3), and (3) R (relapse, MOG-AAD#1.4) as described in Materials and Methods. Raw BCL files generated through sequencing were further de-multiplexed using Picard (https://github.com/broadinstitute/picard) and the resulting FASTQ files where aligned to the human reference genome (GRCh38) using the STAR v2.4.2a52 aligner. Further QC was done using the RNA-seQC53 and transcript counts were produced using feature Counts function of the Subread package54. Data was normalized using the DESeq2 package55 and the graphs were made using GraphPadPrism version 8.4.2 (464). C TNFAIP3 and NFκβ1 expression by DGE sequencing. NanoString Gene Expression Assay was performed on a MOG-AAD patient#2 with 3 longitudinal samples, (1) untreated R (relapse, MOG-AAD#2.1), (2) 8 months Rm/Ct (mycophenolate mofetil treated at remission, MOG-AAD#2.2), and (3) 11 months Rm/Ct (mycophenolate mofetil treated at remission, MOG-AAD#2.3) as described in Materials and Methods. Data were normalized and analyzed using nSolver software via the geometric mean of included housekeeping genes. The graphs were made using GraphPadPrism version 8.4.2 (464). D TNFAIP3 and TNF-α expression by NanoString Gene Expression Assay. qPCR was performed on CD4+ T cells from 7 MOG-AAD patients with longitudinal samples as described in Materials and Methods. It also included MOG-AAD patient#2 with 3 longitudinal samples as previously used for NanoString gene expression assay. The graphs were made using GraphPadPrism version 8.4.2 (464). E TNFAIP3 expression in MOG-AAD patient#2 by qPCR. F Grouped analysis of TNFAIP3 expression in relapse samples, remission samples and samples treated with corticosteroids by qPCR. Relapse n = 5, Remission n = 5, Steroid n = 4, Ordinary 1-way ANOVA; P = 0.0137.
In order to validate our single cell RNA sequencing findings, we sought to evaluate the same MOG-AAD patient#1 with 3 longitudinal samples during remission (MOG-AAD#1.2), pre-relapse (MOG-AAD#1.3) and at a relapse (MOG-AAD#1.4) using DGE sequencing. DGE sequencing is a cost-effective method that offers low background noise, increased sensitivity and reproducibility24. For this, PBMCs were stimulated with individual MOG peptides, p35-55, p119-130 and p181-195. Total RNA from CMC T cells was isolated and sequenced. Broad Genomics Platform sequenced the libraries. Consistent with the single cell RNA sequencing findings, we found that gene expression of TNFAIP3 was decreased at both the pre-relapse and relapse sample compared to the remission sample. TNFAIP3 is known to be a regulator of NFκβ. We found that the gene expression of NFκβ1 was increased at relapse compared to remission time point, the inverse of TNFAIP3 expression (Fig. 3c). The other genes identified by single cell RNA sequencing, namely HILPDA, MXI-I, BNIP3 and FAM162A showed a similar trend by DGE sequencing. They were up regulated during remission as compared to pre-relapse and relapse time points (Supplementary Fig. 2a). However, SNRPG, EWSR1, HMGN1 and LRRC75A-AS1, that were shown to be up regulated during pre-relapse time point by single cell RNA sequencing did not follow the same trend by DGE sequencing (Supplementary Fig. 2b). Gene expression for AL137058.2 was below the detection limit by DGE sequencing.
To further confirm our RNA sequencing results, we decided to test another MOG-AAD patient#2 with 3 longitudinal samples, 1 at relapse (MOG-AAD#2.1) and 2 mycophenolate mofetil treated samples at remission (MOG-AAD#2.2 and MOG-AAD#2.3), using NanoString’s differential gene expression platform. NanoString is a high throughput technique that allows simultaneous gene expression of more than 700 genes. We analyzed gene expression in CD4+ T cells, CD19+ B cells and CD14+monocytes using the nCounter software. There was a distinct increase in the TNFAIP3 expression from CD4+ T cells in remission samples as compared to relapse sample thereby indicating that CD4+ T cells play an important role in TNFAIP3 regulation (Fig. 3d). In contrast, TNF-α expression from CD14+ monocytes, principal source of TNF-α in humans was increased in the relapse sample as compared to remission. CD4+ T cells followed a similar trend, where in TNF-α expression was higher at relapse as compared to remission samples.
To validate NanoString gene expression assay results, we isolated CD4+ T cells from 7 additional MOG-AAD patients with longitudinal samples (longitudinal samples n = 5/7, relapse n = 7 and remission n = 8, total n = 15). It also included MOG-AAD patient#2 with 3 longitudinal samples as previously used for NanoString gene expression assay. Quantitative real-time polymerase chain reaction (qPCR) was performed using FAM-labeled primer for TNFAIP3. GAPDH gene was used as an endogenous control to normalize for differences in the amount of total RNA in each sample. All values are shown as relative expression. The qPCR data replicated the results from NanoString gene expression assay. There was an increase in the relative expression of TNFAIP3 at remission time points and a decrease at relapse in the CD4+ T cells of MOG-AAD patient#2 (Fig. 3e). We next conducted grouped analysis of CD4+ TNFAIP3 expression levels from patients in relapse or remission states on disease modifying therapies, and samples from patients receiving high dose of corticosteroids, which are known to induce TNFAIP3 through binding of the glucocorticoid receptor20. Grouped analysis comparing relapse samples (n = 5), remission samples (n = 5) and samples from patients treated with high dose of corticosteroids (n = 4) showed a significant difference between the three groups (Fig. 3f) (Ordinary 1-way ANOVA; P = 0.013).
Protein expression analysis demonstrated decreased TNFAIP3 expression in MOG-AAD patient at relapse timepoint
As the sequencing data on three independent platforms consistently suggested the differential expression of TNFAIP3 in MOG-AAD patients, we evaluated protein expression of TNFAIP3 in whole PBMCs from an untreated MOG-AAD patient#3 (relapse MOG-AAD#3.1 and non-relapse MOG-AAD#3.2). Ligation of the TCR has been shown to induce TNFAIP3 expression21 and corticosteroids are commonly used as treatment for MOG-AAD which can aid in the resolution of relapses and potentially prevent new relapses5,7. Hence, we cultured PBMCs with 2 conditions: MOG antigen stimulation (1 μg/ml) and MOG+dexamethasone stimulation (1 μg + 100 nm) at 4 timepoints: 4, 8, 16 and 24 h. Addition of MOG peptide did not increase TNFAIP3 expression in relapse or non-relapse samples (Fig. 4a, full-length blots/gels are presented in Supplementary Fig. 5a) but in the presence of both, MOG antigen and dexamethasone, which is a synthetic glucocorticoid, TNFAIP3 expression was comparable at 4, 8 and 16 h, however expression increased after 24 h of stimulation in the relapse sample as compared to non-relapse sample thereby partially rescuing the expression of TNFAIP3 in the relapse sample (Fig. 4b, full-length blots/gels are presented in Supplementary Fig. 5b).
TNFAIP3 protein expression in relapse and non-relapse samples from an untreated MOG-AAD patient: PBMCs from a MOG-AAD patient#3 at a relapse (MOG-AAD#3.1) and non-relapse (MOG-AAD#3.2) time point were cultured under different conditions, analyzed by SDS-PAGE and followed by western blot for TNFAIP3 at the indicated time points (hours) as described in Materials and Methods. A MOG antigen stimulation at 1 μg/ml (peptide cocktail comprising of MOG p1-20, p35-55, p119-130, p181-195 and p186-200). B MOG antigen + Dexamethasone stimulation at 1 μg + 100 nm. Representative Western blot images are shown on left and protein bands quantified using ImageJ version 1.53b and normalized to β-actin are shown on the right. The graphs were made using GraphPadPrism version 8.4.2 (464). Full-length blots/gels for TNFAIP3 and β-Actin are presented in Supplementary Fig. 5.
We also studied a dose response at lower and higher concentration of MOG antigen and dexamethasone stimulations in a MOG-AAD patient#2 (untreated, relapse MOG-AAD#2.1 and treated with mycophenolate mofetil, non-relapse MOG-AAD#2.3). Here we cultured PBMCs with 5 conditions: exvivo (unstimulated), MOG antigen stimulation (lower dose at 1 μg/ml and higher dose at 10 μg/ml) and MOG + dexamethasone stimulation (lower dose at 1 μg + 100 nm and higher dose at 10 μg + 1000 nm) at 3 timepoints: 4, 16 and 24 h. TNFAIP3 expression increased in presence of dexamethasone as compared to MOG antigen alone stimulation, more so in the non-relapse MOG-AAD sample as compared to relapse and with higher dose of MOG + dexamethasone (Supplementary Fig. 3a–e, full-length blots/gels are presented in Supplementary Fig. 6). TNFAIP3 expression was also increased in the non-relapse MOG-AAD sample as compared to relapse MOG-AAD sample in the exvivo (unstimulated) condition (Supplementary Fig. 3f, full-length blots/gels are presented in Supplementary Fig. 6).
Since TNFAIP3 is a well-known regulator of NFκβ. We studied its correlation with NFκβ subunits p50 and p65 in the same MOG-AAD patient#2 at a lower dose of MOG antigen stimulation (1 μg/ml) at 4 timepoints: 4, 8, 16 and 24 h. There was a negative correlation of TNFAIP3 expression with NFκβ subunits p50 and p65 (Supplementary Fig. 4, full-length blots/gels are presented in Supplementary Fig. 7).
Serum analysis showed decreased levels of TNFAIP3 at relapse and increased levels at remission in MOG-AAD patients and in healthy controls
Previous studies measuring TNFAIP3 in the serum using ELISA assays have been performed in the context of viral infections such as chronic Hepatitis B infections22. To confirm that alterations in TNFAIP3 transcription corresponded with translation into the TNFAIP3 protein, we tested serum levels of TNFAIP3 in MOG-AAD patients. We found a strong correlation of TNFAIP3 serum level decrement with the onset of relapse and increase following intravenous steroids in MOG-AAD patient#1.1–1.5 (Fig. 5a). Serum from other MOG-AAD patients, #5–8 (Supplementary Table 1a), with longitudinal samples showed similar trends (Fig. 5b–e). Mixed model paired analysis between relapse (n = 8) and remission (n = 22) samples from MOG-AAD patients (6 pairs) showed a significant reduction in TNFAIP3 levels in paired relapse samples as compared to remission samples (P = 0.006) (Fig. 5f).
Serum levels of TNFAIP3 in indicated longitudinal samples of MOG-AAD patients determined by ELISA: The graphs were made using GraphPadPrism version 8.4.2 (464) A TNFAIP3 levels in MOG-AAD patient#1 with 5 longitudinal samples, (1) Rm (remission, MOG-AAD#1.1), (2) Rm (remission, MOG-AAD#1.2), (3) PR (pre-relapse, MOG-AAD#1.3), (4) R (relapse, MOG-AAD#1.4), and (5) Rm/S (remission treated with steroid, MOG-AAD#1.5). B TNFAIP3 levels in MOG-AAD patient#5 with 8 longitudinal samples, (1) Rm (remission, MOG-AAD#5.1), (2) Rm (remission, MOG-AAD#5.2), (3) Rm (remission, MOG-AAD#5.3), (4) Rm (remission, MOG-AAD#5.4), (5) PR (pre-relapse, MOG-AAD#5.5), (6) Rm (remission, MOG-AAD#5.6), (7) Rm/Ct (mycophenolate mofetil treated at remission, MOG-AAD#5.7), and (8) Rm/Ct (mycophenolate mofetil treated at remission, MOG-AAD#5.8). C TNFAIP3 levels in MOG-AAD patient#6 with 3 longitudinal samples, (1) R/S (relapse treated with steroid, MOG-AAD#6.1), (2) Rm/Ct (mycophenolate mofetil treated at remission, MOG-AAD#6.2), and (3) Rm/Ct (mycophenolate mofetil treated at remission, MOG-AAD#6.3). D TNFAIP3 levels in MOG-AAD patient#7 with 2 longitudinal samples, (1) R/S (relapse treated with steroid, MOG-AAD#7.1), and (2) Rm/Rebif (remission treated with interferon beta-1a, MOG-AAD#7.2). E TNFAIP3 levels in MOG-AAD patient#8 with 2 longitudinal samples, (1) Rm (remission, MOG-AAD#8.1) and (2) R/S/Co (remission treated with steroid and Glatiramer acetate, MOG-AAD#8.2). F Mixed model paired analysis compared MOG-AAD patients, who had, both, relapse and remission samples. Relapse n = 8, Remission n = 22. The comparison between the relapse and remission group was done using Nonlinear Mixed-Effects Models library in R (nlme).
Further, using Nonlinear Mixed-Effects Models library in R (nlme), we evaluated all available relapse n = 10 and remission samples n = 40 from MOG-AAD patients and compared them with PHC n = 44 samples (Supplementary Table 1a and 1b). We found a significant reduction in TNFAIP3 serum levels in relapse samples as compared to remission samples (P = 0.04). There was also significant reduction in TNFAIP3 serum levels in relapse samples as compared to PHC (P = 0.0001). This indicated that low levels of TNFAIP3 are associated with the onset and subsequent relapses in MOG-AAD patients whereas patients at remission and healthy controls show high levels of TNFAIP3.
In order to assess if there were other proteins associated with relapse in MOG-AAD, we evaluated serum samples of 49 MOG-AAD patients (relapse samples n = 10 and remission samples n = 38) using a multiplex inflammatory panel made up of 92 proteins. Data was presented as normalized protein expression (NPX) values. After adjusting for age, sex and treatments, we found 7 biomarkers: FLT3 (Fms Related Tyrosine Kinase 3 Ligand, P = 0.0061), CDCP1 (CUB Domain Containing Protein 1, P = 0.008), IL12B (Interleukin 12B, P = 0.0284), NTF3 (Neurotrophin 3, P = 0.036), EIF4EBP1 (Eukaryotic Translation Initiation Factor 4E Binding Protein 1, P = 0.04), KITLG (KIT Ligand, P = 0.042), CCL2 (C–C Motif Chemokine Ligand 2, P = 0.046) and CCL25 (C–C Motif Chemokine Ligand 25, P = 0.05) that were significantly down regulated in relapse samples. However, these were no longer significant when corrected for multiple comparisons.
Discussion
In this study we evaluated PBMC and serum samples from MOG-AAD patients at relapse and remission time points. We assessed for biomarkers by transcriptomic analysis. Utilizing 3 independent gene expression platforms, we identified TNFAIP3 as a relapse biomarker in MOG-AAD patients. Further we showed that TNFAIP3 protein level is reduced at relapse time point and increased at remission time point in MOG-AAD patient as well as in healthy controls. This was later confirmed in serum samples of MOG-AAD patients and healthy controls, where we found TNFAIP3 serum levels were decreased at relapse and increased at remission and in healthy controls.
MOG-AAD is a unique autoimmune demyelinating disease with a strong antigen-specific central memory Th1 and Th17 T cell response, and associated antibody production. TNFAIP3 gene, encoding the A20 protein, is emerging as a pivotal checkpoint in autoimmune diseases such as MS, rheumatoid arthritis and Crohn’s disease38. We found through transcriptomic analysis of central memory T cells (CMC) that TNFAIP3 gene expression is decreased during a relapse, and that this decrease correlates with a subsequent activation of NFκβ signaling. Protein analysis demonstrates that MOG-AAD relapse and remission samples respond differently to antigen stimulation, suggesting a dysregulation of TNAIP3 response, which may be dependent on cell state. We found that TNFAIP3 serum level decrements are associated with the onset of a relapse in individual MOG-AAD patients and thus has significant potential as a biomarker and therapeutic target for MOG-AAD and other autoimmune diseases.
The human TNFAIP3 gene is located on chromosome 6, and is also known as A20. It encodes the 790 amino acid protein, A20 that is, made of an N-terminal protease domain and seven Cys2-Cys2 zinc finger C-terminal domains. A20 is an ubiquitin-editing enzyme belonging to the ovarian tumor (OTU) proteases family of deubiquitinating (DUB) enzymes23. A20 functions as an E3 ubiquitin ligase as a result of its fourth zinc finger motif in the C-terminal domain TNFAIP3 is a key regulator of cellular processes including NFκβ activation and apoptosis24,25,26. TNFAIP3 suppresses cellular processes by down regulating NFκβ activation in part through DUB activity, ubiquitin-binding activity, and/or E3 ligase activity of critical signaling components including RIP1, TRAF6 and NEMO, upstream of the IKK complex27,28,29,34. It also binds directly to the C-terminus of IL-17RA30, and can decrease IL-17 responses through inhibition of p3831. Several stimuli, including TNFα, LPS, TLR and IL-1 through activation of NFκβ via the non-canonical pathway, increase TNFAIP3 mRNA expression25,32, thus initiating a negative feedback loop to regulate NFκβ33 via TRAF1/TRAF234. TNFAIP3-deficient cells fail to terminate TNF-α-induced NFκβ responses25. Mice deficient in TNFAIP3 develop severe cachexia and inflammation in the liver, kidneys, intestines, joints, and bone marrow, and die prematurely25. Astrocytic expression of TNFAIP3 in EAE protects from CNS immune-mediated demyelination through suppression of chemokines35. Deletion of TNFAIP3 in microglia increases microglial cell number and affects microglial regulation of neuronal synaptic function and worsens demyelination in EAE through hyperactivation of the Nlrp3 inflammasome36. Thus loss of TNFAIP3 function and/or polymorphisms in the TNFAIP3 gene encoding for A20 protein is related to reduced A20 expression which thereby causes immune mediated inflammation and autoimmune diseases in humans36.
Activation of NFκβ through the canonical pathway related to TCR activation requires MALT1 signaling, which is suppressed by TNFAIP3 through deubiquitination37. In turn, MALT1 mediates rapid proteolytic cleavage and inactivation of TNFAIP3 after TCR stimulation, thus fine-tuning TCR activation21. MALT1 is also required for BCR activation and TNFAIP3 deletion is frequently observed in B cell lymphomas38.
Our results demonstrate that CD4+ T cell and serum decrements of TNFAIP3 are associated with clinical relapses in MOG-AAD patients and also associated with relative increases in NF-κβ expression, which is consistent with the role of TNFAIP3 in modulating NF-κβ activation. However these results could be due to the effects of strong antigen-specific activation of T cells in MHC-matched patients, or because of aberrant regulation of TNFAIP3 in this condition. Indeed, genetic polymorphisms of the TNFAIP3 gene have been described in several autoimmune diseases including rheumatoid arthritis, psoriasis, type 1 diabetes, inflammatory bowel disease, systemic lupus erythematosus (SLE), coronary artery disease and celiac disease39,40,41,42,43,44. Several TNFAIP3 intergenic polymorphisms have also been associated with MS susceptibility45. MOG-AAD is closely related to MS, however it has distinct clinical and radiological features. A comprehensive large-scale genomic analysis of MOG-AAD has not been reported thus far.
MOG-AAD, which is largely a pediatric disease, is highly sensitive to glucocorticoid treatment7. Glucocorticoids bind to the glucocorticoid receptor (GR) leading to immune suppression. TNFAIP3, which is an anti-inflammatory target of TNF-α, inhibitor of NF-κβ, is regulated by steroids including estrogen46 and glucocorticoids47. While stimulation with MOG antigen failed to induce TNFAIP3, dexamethasone stimulation, a potent glucocorticoid, partially rescued the expression of TNFAIP3 protein and transcriptome levels in the relapse sample thereby suggesting that the induction of TNFAIP3 by glucocorticoids can improve relapses in MOG-AAD patients.
Serum TNFAIP3 levels may be related to secretion by immune cells. The decrement in TNFAIP3 may be associated with antigen-stimulation of the TCR, thus associated with activation of antigen-specific T cells. In this study, we analyzed gene expression only in CMC T cells, which are integral in initiating antigen-specific immune response. However, B cells, monocytes and other cells also express TNFAIP3, and may be involved in eliciting TNFAIP3 serum levels. It is possible that antigen-specific responses in other diseases may be associated with similar findings. The serum level of TNFAIP3 varies between individual patients, and only decrements observed in longitudinal samples, but not absolute values were associated with relapses.
The utility of TNFAIP3 as a biomarker for relapses will need to be validated in additional MOG-AAD patients with age and gender-normalized comparisons in healthy controls. Here we demonstrate that decreased TNFAIP3 levels are associated with relapses in MOG-AAD patients. TNFAIP3 may not only indicate disease severity but also serve as a therapeutic target and a prognostic biomarker in MOG-AAD patients. Thus, further studies are required to evaluate the effect of increased expression of TNFAIP3 in patients with MOG-AAD and other related diseases.
Strengths of this study include carefully phenotyped cohorts of pediatric patients with MOG-AAD, and detailed cellular, transcriptomic and proteomic analysis. Limitations are the small number of subjects investigated, which reflects the challenges of obtaining multiple biological samples in pediatric patients. Steps are underway to further validate these results in separate cohorts of subjects, and to develop a set of multivariate proteomic and/or transcriptomic biomarkers for potential use in the clinical setting.
Materials and methods
Subjects and blood samples
All methods in this study were carried out in accordance with relevant guidelines and regulations. Subjects were selected from an ongoing biomarker study at the Partners Pediatric MS Center at Massachusetts General Hospital, which is approved by Partners Human Research Committee/Institutional Review Board for the use of human material. Parents of children signed an informed consent form. Peripheral venous blood was collected in lithium heparin blood collection tubes (Becton Dickinson, NJ, USA) from subjects after obtaining informed consent. We included 20 pediatric MOG-AAD patients and 44 age and sex-matched pediatric healthy controls (PHCs). MOG antibody testing was performed at the sample collection site by cell-based assay6, or at the Mayo Clinic as part of clinical care. The 20 MOG-AAD patients had the following diagnoses at the time of sample collection according to International Pediatric MS Study Group diagnostic criteria: 7 with MS, 7 with ADEM-ON, 1 with multiphasic ADEM, 1 with ADEM-TM, 1 with CIS, 1 with a demyelinating neurological disorder and 2 with NMO-SD48. Patients were diagnosed with NMO-SD if presenting with ON, TM and at least two of these 3 criteria: MRI evidence of a continuous spinal cord lesion, brain MRI that was non-diagnostic of MS, and NMO IgG seroposivity48,49. Out of the 20 MOG-AAD patients, 15 patients had longitudinal samples, including treated/untreated and relapse/non-relapse samples. Untreated samples were defined as no steroids or intravenous immunoglobulin nor disease modifying therapies within 30 days prior to sample collection. Samples within 30 days of a clinical relapse with new MRI lesions were defined as “relapse” samples or within 64 days of ongoing relapse symptoms, while samples at a non-relapse time point were defined as remission samples. One sample (#1.3) was identified as a pre-relapse sample, since the patient reported new mild clinical symptoms, but no radiological correlation was found (Table 1 and Supplementary Table 1 and b). A second sample (#5.5) was also termed pre-relapse since the patient had significant new headache not responsive to standard medication. Neurologists specialized in pediatric demyelinating disorders validated all clinical and radiological data (TC and GG).
Cell stimulation assay and FACS analysis
Peripheral blood mononuclear cells (PBMCs) were isolated by Ficoll-Paque Plus (GE healthcare Biosciences AB, Sweden) density gradient centrifugation. Cells were cultured using Hybridomas and Lymphoid cells (HL-1, Lonza, MD, USA) media containing 5% of human serum (Valley Biomedical INC, VA, USA), L-Glutamine (Fisher Scientific, NH, USA), Penicillin–Streptomycin (Fisher Scientific, NH, USA), HEPES, non-essential amino acids (NEAA) and sodium pyruvate (all 3 from Lonza, MD, USA). PBMCs from 7 MOG-AAD patients of which 3 patients had longitudinal samples and 7 age and sex matched PHCs (Table 1) were plated at a density of 500,000 cells/well in a 96-well plate (Corning, ME, USA). Cells were stimulated by either one of the following 6 peptide conditions, (1) MOG p1-20, (2) MOG p35-55 (Immune Tolerance Network, USA), (3) MOG p119-130 (4) MOG p181-195 (5) MOG p186-200 (Genemed INC, CA, USA), (6) cocktail of myelin peptides consisting of Proteolipid Protein (PLP) 139–154, Myelin Basic Protein (MBP) 13–32, MBP 111–129 and MBP 146–170 (Immune Tolerance Network, USA) at 10 μg/ml. Cells were cultured for 7 days at 37 °C, 5% CO2 and 90% humidity.
To evaluate cytokine production, cells were further stimulated with Phorbol 12-myristate 13-acetate (PMA) at 33 ng/ml, Ionomycin at 166 ng/ml (Sigma Aldrich, MO, USA) and Golgi stop (BD Bioscience, CA, USA) for 4 h. After incubation cells were stained with the following antibodies, human anti-CD4 APC (RPA-T4, Biolegend Inc, CA, USA), anti-CD45RA-AF700 (HI100, Biolegend Inc, CA, USA), anti-CCR7-PE (GO43H7, Biolegend Inc, CA, USA) and live/dead fixable violet dead cell stain kit (Thermo Fischer Scientific, USA). Cells were then fixed and permeabilized with BD fixation and permeabilization buffer (BD Biosciences, CA, USA) and stained with the following intracellular cytokine antibodies: anti-IL17-PeCy7 (BL168, Biolegend Inc, CA, USA) and anti- IFNγ-PerCp (4S.B3, Biolegend Inc, CA, USA). Flow cytometry was performed on BD LSR II (BD Biosciences, CA, USA); data was analyzed using Flowjo vX.0.7 and the graphs were made using GraphPadPrism version 8.4.2 (464).
Live CD4+ T cells were selected from all lymphocytes and analyzed for expression of central memory cells (CMCs: CCR7+CD45RA−), effector memory cells (EMCs: CCR7−CD45RA−), and effector cells (CD45RA+). Intracellular pro-inflammatory cytokines, IL17+, IFNγ+, and IL17+IFNγ+ (double positive) were gated within CMCs and EMCs.
Gene expression assays and data analysis
Single cell RNA sequencing (inDrop)
Single cell RNA sequencing was conducted in CMCs from CD4+ T cells isolated from MOG-AAD patient#1 with 3 untreated longitudinal samples, (1) at remission (MOG-AAD#1.2) (2) at a pre-relapse (MOG-AAD#1.3), two months prior to a relapse in which the patient experienced headache and mild visual symptoms with no radiological correlate and (3) at a relapse (MOG-AAD#1.4) characterized by bilateral weakness and multiple new MRI lesions (Table 1) following the inDrop technique as previously described20,21. InDrop single cell library sequencing was performed at the Single Cell Core (SCC), laboratory of Dr. Allon Klein, Department of Systems Biology at Harvard Medical School (HMS). The relapse sample failed sequencing, however the remission and pre-relapse samples were evaluable. For simplicity, moving forward we reassigned the pre-relapse sample to be a relapse sample.
Whole PBMCs were stimulated with anti-CD3 (OKT3, Ebioscience, CA, USA) and anti-CD28 (CD28.2, Ebioscience, CA, USA) at 0.5 μg/ml for 3 days. Cells were stained with human anti-CD4 APC (RPA-T4, Biolegend Inc, CA, USA), anti-CD45RA-AF700 (HI100, Biolegend Inc, CA, USA), anti-CCR7-PE (GO43H7, Biolegend Inc, CA, USA) and live/dead fixable violet dead cell stain kit (Thermo Fischer Scientific, USA). CMCs (CCR7+CD45RA−) were sorted from CD4+ T cells using BD FACS ARIA (BD Biosciences, CA, USA). 3,000 cells from a cell suspension comprising of CMCs from the 2 samples were isolated into droplets that contained lysis buffer. cDNA libraries were sequenced using the Illumina NextSeq 500 platform and analyzed following V3 Indrop criteria. After sequencing the raw BCL files were manually demultiplexed using the bcl2fastq (https://support.illumina.com/sequencing/sequencing_software/bcl2fastq-conversion-software.html) software by illumina. Reads obtained from bcl2fastq were further processed using the single-cell RNA-seq pipeline of the bcbio-nextgen (https://bcbio-nextgen.readthedocs.io/en/latest/contents/pipelines.html#single-cell-rna-seq) software suite. The single-cell RNA-seq pipeline inspected each read, performed alignment using RapMap50 and produced transcript level count matrix. This matrix was further processed with Seurat (https://www.biorxiv.org/content/early/2018/11/02/460147) where QC, filtering, log normalization and scaling was performed. The scaled data was further clustered using Seurat and visualized using TSNE51.
The TSNE plot was labeled with sample state (Relapse/Remission) to identify cluster differences between states. Since the clusters showed homogeneity between relapse and remission sample states, gene specific differences between the samples were assessed. “AverageExpression” function in Seurat was used to calculate average gene expression for the relapse and remission samples separately. The difference in the gene expression was calculated by subtracting the expression values from relapse and remission samples for each gene. The obtained list was further sorted to identify genes that were up regulated in relapse and remission samples respectively. Based on the difference in the gene expression, the top 5 genes were further evaluated.
Digital gene expression (DGE) sequencing
DGE sequencing was conducted in CMCs from CD4+ T cells isolated from the same MOG-AAD patient#1 with 3 untreated longitudinal samples at remission (MOG-AAD#1.2), pre-relapse (MOG-AAD#1.3) and relapse (MOG-AAD#1.4) as previously described (Table 1)24,25,26. PBMCs were stimulated with MOG peptides, p35-55, p119-130 and p181-195 at 10 μg/ml for 7 days. Cells were stained with human anti-CD4 APC (RPA-T4, Biolegend Inc, CA, USA), anti-CD45RA-AF700 (HI100, Biolegend Inc, CA, USA), anti-CCR7-PE (GO43H7, Biolegend Inc, CA, USA) and live/dead fixable violet dead cell stain kit (Thermo Fischer Scientific, USA). CMCs (CCR7+CD45RA−) were sorted from CD4+ T cells using BD FACS ARIA. Total RNA was isolated using Total RNA purification kit following manufacturer`s guidelines (Norgen, MA, USA). Based on the guidelines provided by the Broad institute for library preparation, RNA concentration for all samples was normalized to 5 ng/μl l in 10 μl of nuclease free water. Library sequencing was performed at The Broad Genomics Platform.
Raw BCL files generated through sequencing were further de-multiplexed using Picard (https://github.com/broadinstitute/picard) and the resulting FASTQ files where aligned to the human reference genome (GRCh38) using the STAR v2.4.2a52 aligner. Further QC was done using the RNA-seQC53 and transcript counts were produced using feature Counts function of the Subread package54. Before running the analysis, genes with low overall expression were removed from the analysis and the data was normalized using the DESeq2 package55. The graphs were made using GraphPadPrism version 8.4.2 (464).
Nanostring ncounter gene expression assay
Cell subsets were isolated from MOG-AAD patient#2 with 3 longitudinal samples, 1 at relapse (MOG-AAD#2.1) and 2 mycophenolate mofetil treated samples at remission (MOG-AAD#2.2 and MOG-AAD#2.3) (Table 1) were evaluated by NanoString array as previously described31. RNA expression of 770 genes was detected by nCounter XT Code-Set Gene Expression Assay, Human Autoimmune kit. CD4+ T cells, CD19+ B cells and CD14+monocytes were positively selected using micro beads and magnetically isolated using MACSQuant columns placed in the magnetic field of a MACS separator (Miltenyi Biotec, CA, USA). In order to achieve maximum purity, staining antibodies for anti-CD4-PE (OKT4, Biolegend Inc, CA, USA) anti-CD19-PE (HIB19, Biolegend Inc, CA, USA) and anti-CD14-FITC (M5E2, BD Biosciences, CA, USA) were added during magnetic separation. 7-AAD viability staining solution was used to separate the live cells from the dead. Further CD4+, CD19+ and CD14+ cells were sorted using BD FACS ARIA (BD Biosciences, CA, USA). Total RNA was isolated using Total RNA purification kit following manufacturer`s guidelines (Norgen, MA, USA). RNA concentration for all samples was normalized to 30 ng/μl l in 5 μl of nuclease free water. Hybridization protocol for nCounter XT Code-Set Gene Expression Assay was performed following the manufacturer`s instructions. Data were normalized and analyzed using nSolver software via the geometric mean of included housekeeping genes. The graphs were made using GraphPadPrism version 8.4.2 (464).
Quantification by real time PCR
CD4+ T cells from 7 MOG-AAD patients (longitudinal n = 5/7, relapse n = 7, remission n = 8, total n = 15, Table 1) were isolated as previously described. Total RNA was isolated using Total RNA purification kit following manufacturer`s guidelines (Norgen, MA, USA). RNA concentration for all samples was determined using NanoDrop 2000/2000c Spectrophotometer (Thermo fischer scientific, USA). First-strand cDNA synthesis was performed for each RNA sample from 50 ng of total RNA using SuperScript IV VILO Master Mix (Thermo fischer scientific, USA) following manufacturer’s guidelines. qPCR was performed using FAM-labeled primers for TNFAIP3, Hs00234713_m1 and GAPDH, Hs99999905_m1 with TaqMan Fast Universal qPCR Master Mix (No ampErase Uracil N-Glycosylase, Thermo fischer scientific, USA). Samples were run in duplicates on QuantStudio 7 Flex (Applied Biosystems, Life Technologies, USA). GAPDH gene was used as an endogenous control to normalize for differences in the amount of total RNA in each sample, and all values are shown as relative expression. The graphs were made using GraphPadPrism version 8.4.2 (464).
SDS-PAGE and western blot
PBMCs from an untreated MOG-AAD patient#3 with 2 longitudinal samples at relapse (MOG-AAD#3.1) and non-relapse (remission, MOG-AAD#3.2) (Table 1) were cultured at a density of 250,000 cells/well in a 96-well plate (Corning, ME, USA) for 4, 8, 16 and 24 h under the following 2 conditions: (1) MOG antigen at 1 μg/ml (peptide cocktail comprising of MOG p1-20, p35-55, p119-130, p181-195 and p186-200, Sigma Aldrich, MO, USA), and (2) MOG antigen at 1 μg/ml with Dexamethasone at 100 nM (Sigma Aldrich, MO, USA). PBMCs for an additional MOG-AAD patient#2 with 2 longitudinal samples, untreated at relapse (MOG-AAD#2.1) and treated with mycophenolate mofetil at non-relapse (remission, MOG-AAD#2.3) (Table 1) were cultured similarly under the following 5 conditions: (1) Exvivo (unstimulated) (2) MOG antigen stimulation at a lower dose of 1 μg/ml, (3) MOG antigen stimulation at a higher dose of 10 μg/ml (4) MOG antigen stimulation with dexamethasone at a lower dose of 1 μg + 100 nM, and (5) MOG antigen stimulation with dexamethasone at a higher dose of 10 μg + 1000 nM. Cells were lysed with RIPA lysis buffer (Thermo Fisher Scientific, MA, USA) supplemented with protease and phosphatase inhibitor cocktails (Thermo Fisher Scientific, MA, USA). Total protein was determined by Pierce BCA protein assay following manufacturer’s instructions (Thermo Fisher Scientific, MA, USA). Samples were prepared with 30 μg of cell lysate, loading buffer (Life Technologies, OR, USA), and reducing agent (Life Technologies, OR, USA) and then heated for 10 min at 95 °C before use. Samples were run on 4–12% Bis Tris gels (Invitrogen, CA, USA) and transferred to polyvinylidene difluoride membranes (PVDF) (Immobilon-P, Millipore Sigma, MA, USA). Membranes were blocked using 5% BSA (Sigma Aldrich, MO, USA) and blotted overnight at 4 °C with antibodies against anti-TNFAIP3 (ab92324, 1:1,000), anti- NFκβ p50 (ab32360, 1:2000) Abcam, MA, USA, anti-phospho NFκβ p65 (3033L, 1:2000), anti- NFκβ p65 (8242S, 1:2000), and anti- β-actin (4970S, 1:4,000) Cell signaling, MA, USA. Membranes were incubated with secondary antibody anti-rabbit IgG HRP linked (7074P2, 1:10,000, Cell signaling, MA, USA) for 1 h at 37 °C. Immunoblots were developed using KwikQuant western blot detection kit (Kindle biosciences, CT, USA). Protein bands were quantified using ImageJ version 1.53b and normalized to their respective β-actin. The graphs were made using GraphPadPrism version 8.4.2 (464).
ELISA
TNFAIP3 concentration in serum samples of 50 MOG-AAD patients (Table 1 and Supplementary Table 1) with relapse samples n = 10, remission samples n = 40 and PHC n = 44 was assessed by commercially available TNFAIP3 ELISA kit (MyBiosource, CA, USA) following manufacturer’s instructions. All samples were tested in duplicates. Detection range of the assay was 23.5 pg/ml-1500 pg/ml. The Intra-assay precision CV was < 8% and Inter-assay precision CV was < 10%. The optical density was determined at 450 nm and the reference wavelength was set at 560 nm. Nonlinear Mixed-Effects Models library in R (nlme) was used for statistical analysis. The graphs were made using GraphPadPrism version 8.4.2 (464).
Multiplex panel comprising of 92 protein biomarkers (Olink Proteomics, Uppsala, Sweden) was used to assess serum samples of 49 MOG-AAD patients (Table 1 and Supplementary Table 1) with relapse samples n = 11 and remission samples n = 38 following manufacturer’s instructions. The Intra-assay precision CV was < 15% and Inter-assay precision CV was < 25%. Data from the analyzed protein biomarkers was presented in normalized protein expression (NPX) values, Olink Proteomics’s arbitrary unit on log2 scale.56,57.
Disclosures
SS-none, HL-none, GG-none, RR-none, TR- none, TC has received personal compensation for advisory board/consulting for Biogen-Idec, Merck Serono, Novartis, Sanofi, Bayer, Celgene, Alexion and has received financial support for research activities from Merck Serono and Novartis Pharmaceuticals, the Department of Defense, National MS Society, the Guthy-Jackson Charitable Foundation and the Peabody Foundation.
References
Reindl, M., Di Pauli, F., Rostasy, K. & Berger, T. The spectrum of MOG autoantibody-associated demyelinating diseases. Nat. Rev. Neurol. 9, 455–461 (2013).
Jarius, S. et al. MOG-IgG in NMO and related disorders: a multicenter study of 50 patients. Part 1: Frequency, syndrome specificity, influence of disease activity, long-term course, association with AQP4-IgG, and origin. J. Neuroinflamm. 13, 279 (2016).
McLaughlin, K. A. et al. Age-dependent B cell autoimmunity to a myelin surface antigen in pediatric multiple sclerosis. J. Immunol. 183, 4067–4076 (2009).
Waters, P. et al. MOG cell-based assay detects non-MS patients with inflammatory neurologic disease. Neurol. Neuroimmunol. Neuroinflamm. 2, e89 (2015).
Jurynczyk, M. et al. Clinical presentation and prognosis in MOG-antibody disease: a UK study. Brain 140, 3128–3138 (2017).
Fernandez-Carbonell, C. et al. Clinical and MRI phenotype of children with MOG antibodies. Mult. Scler. 22, 174–184 (2016).
Ramanathan, S. et al. Clinical course, therapeutic responses and outcomes in relapsing MOG antibody-associated demyelination. J. Neurol. Neurosurg. Psychiatry 89, 127–137 (2018).
Lopez-Chiriboga, A. S. et al. Association of MOG-IgG serostatus with relapse after acute disseminated encephalomyelitis and proposed diagnostic criteria for MOG-IgG-associated disorders. JAMA Neurol. 75, 1355 (2018).
Hacohen, Y. et al. Diagnostic algorithm for relapsing acquired demyelinating syndromes in children. Neurology 89, 269–278 (2017).
Baumann, M. et al. MRI of the first event in pediatric acquired demyelinating syndromes with antibodies to myelin oligodendrocyte glycoprotein. J. Neurol. 265, 845–855 (2018).
Hyun, J.W., et al. Evaluation of brain lesion distribution criteria at disease onset in differentiating MS from NMOSD and MOG-IgG-associated encephalomyelitis. Mult Scler, 1352458518761186 (2018).
Hacohen, Y. & Banwell, B. Treatment approaches for MOG-Ab-associated demyelination in children. Curr. Treat. Opt. Neurol. 21, 2 (2019).
Jarius, S. et al. MOG-IgG in NMO and related disorders: a multicenter study of 50 patients. Part 2: Epidemiology, clinical presentation, radiological and laboratory features, treatment responses, and long-term outcome. J. Neuroinflamm. 13, 280 (2016).
Hacohen, Y. et al. Disease course and treatment responses in children with relapsing myelin oligodendrocyte glycoprotein antibody-associated disease. JAMA Neurol. 75, 478–487 (2018).
Hennes, E. M. et al. Prognostic relevance of MOG antibodies in children with an acquired demyelinating syndrome. Neurology 89, 900–908 (2017).
Probstel, A. K. et al. Antibodies to MOG are transient in childhood acute disseminated encephalomyelitis. Neurology 77, 580–588 (2011).
Varrin-Doyer, M. et al. MOG transmembrane and cytoplasmic domains contain highly stimulatory T-cell epitopes in MS. Neurol. Neuroimmunol. Neuroinflamm. 1, e20 (2014).
Weber, M. S. et al. B-cell activation influences T-cell polarization and outcome of anti-CD20 B-cell depletion in central nervous system autoimmunity. Ann. Neurol. 68, 369–383 (2010).
Sechi, E., et al. Frequency and characteristics of MRI-negative myelitis associated with MOG autoantibodies. Mult Scler, 1352458520907900 (2020).
Altonsy, M. O., Sasse, S. K., Phang, T. L. & Gerber, A. N. Context-dependent cooperation between nuclear factor kappaB (NF-kappaB) and the glucocorticoid receptor at a TNFAIP3 intronic enhancer: a mechanism to maintain negative feedback control of inflammation. J. Biol. Chem. 289, 8231–8239 (2014).
Coornaert, B. et al. T cell antigen receptor stimulation induces MALT1 paracaspase-mediated cleavage of the NF-kappaB inhibitor A20. Nat. Immunol. 9, 263–271 (2008).
Xu, H. et al. Elevated serum A20 is associated with severity of chronic hepatitis B and A20 inhibits NF-kappaB-mediated inflammatory response. Oncotarget 8, 38914–38926 (2017).
Abbasi, A., Forsberg, K. & Bischof, F. The role of the ubiquitin-editing enzyme A20 in diseases of the central nervous system and other pathological processes. Front. Mol. Neurosci. 8, 21 (2015).
Lu, T. T. et al. Dimerization and ubiquitin mediated recruitment of A20, a complex deubiquitinating enzyme. Immunity 38, 896–905 (2013).
Lee, E. G. et al. Failure to regulate TNF-induced NF-kappaB and cell death responses in A20-deficient mice. Science 289, 2350–2354 (2000).
Onizawa, M. et al. The ubiquitin-modifying enzyme A20 restricts ubiquitination of the kinase RIPK3 and protects cells from necroptosis. Nat. Immunol. 16, 618–627 (2015).
Wertz, I. E. et al. De-ubiquitination and ubiquitin ligase domains of A20 downregulate NF-kappaB signalling. Nature 430, 694–699 (2004).
Boone, D. L. et al. The ubiquitin-modifying enzyme A20 is required for termination of Toll-like receptor responses. Nat. Immunol. 5, 1052–1060 (2004).
Garg, A. V., Ahmed, M., Vallejo, A. N., Ma, A. & Gaffen, S. L. The deubiquitinase A20 mediates feedback inhibition of interleukin-17 receptor signaling. Sci. Signal. 6, 44 (2013).
Garg, A. V., Ahmed, M., Vallejo, A. N., Ma, A. & Gaffen, S. L. The deubiquitinase A20 mediates feedback inhibition of interleukin-17 receptor signaling. Sci. Signal. 6, 44 (2013).
Urbano, P. C. M. et al. TNF-alpha-induced protein 3 (TNFAIP3)/A20 acts as a master switch in TNF-alpha blockade-driven IL-17A expression. J. Allergy Clin. Immunol. 142, 517–529 (2018).
Luo, H. et al. A20 regulates IL-1-induced tolerant production of CXC chemokines in human mesangial cells via inhibition of MAPK signaling. Sci. Rep. 5, 18007 (2015).
Krikos, A., Laherty, C. D. & Dixit, V. M. Transcriptional activation of the tumor necrosis factor alpha-inducible zinc finger protein, A20, is mediated by kappa B elements. J. Biol. Chem. 267, 17971–17976 (1992).
Song, H. Y., Rothe, M. & Goeddel, D. V. The tumor necrosis factor-inducible zinc finger protein A20 interacts with TRAF1/TRAF2 and inhibits NF-kappaB activation. Proc. Natl. Acad. Sci. USA 93, 6721–6725 (1996).
Wang, X. et al. Astrocytic A20 ameliorates experimental autoimmune encephalomyelitis by inhibiting NF-kappaB- and STAT1-dependent chemokine production in astrocytes. Acta Neuropathol. 126, 711–724 (2013).
Voet, S. et al. A20 critically controls microglia activation and inhibits inflammasome-dependent neuroinflammation. Nat. Commun. 9, 2036 (2018).
Duwel, M. et al. A20 negatively regulates T cell receptor signaling to NF-kappaB by cleaving Malt1 ubiquitin chains. J. Immunol. 182, 7718–7728 (2009).
Kato, M. et al. Frequent inactivation of A20 in B-cell lymphomas. Nature 459, 712–716 (2009).
Nair, R. P. et al. Genome-wide scan reveals association of psoriasis with IL-23 and NF-kappaB pathways. Nat. Genet. 41, 199–204 (2009).
Trynka, G. et al. Coeliac disease-associated risk variants in TNFAIP3 and REL implicate altered NF-kappaB signalling. Gut 58, 1078–1083 (2009).
Fung, E. Y. et al. Analysis of 17 autoimmune disease-associated variants in type 1 diabetes identifies 6q23/TNFAIP3 as a susceptibility locus. Genes Immun. 10, 188–191 (2009).
Wang, K. et al. Comparative genetic analysis of inflammatory bowel disease and type 1 diabetes implicates multiple loci with opposite effects. Hum. Mol. Genet. 19, 2059–2067 (2010).
Boonyasrisawat, W. et al. Tag polymorphisms at the A20 (TNFAIP3) locus are associated with lower gene expression and increased risk of coronary artery disease in type 2 diabetes. Diabetes 56, 499–505 (2007).
Lodolce, J. P. et al. African-derived genetic polymorphisms in TNFAIP3 mediate risk for autoimmunity. J. Immunol. 184, 7001–7009 (2010).
International Multiple Sclerosis Genetics , C. et al. Analysis of immune-related loci identifies 48 new susceptibility variants for multiple sclerosis. Nat. Genet. 45, 1353–1360 (2013).
Vendrell, J. A. et al. A20/TNFAIP3, a new estrogen-regulated gene that confers tamoxifen resistance in breast cancer cells. Oncogene 26, 4656–4667 (2007).
Sasse, S. K. et al. Glucocorticoid and TNF signaling converge at A20 (TNFAIP3) to repress airway smooth muscle cytokine expression. Am. J. Physiol. Lung Cell Mol. Physiol. 311, L421-432 (2016).
Krupp, L. B. et al. International Pediatric Multiple Sclerosis Study Group criteria for pediatric multiple sclerosis and immune-mediated central nervous system demyelinating disorders: revisions to the 2007 definitions. Mult. Scler. 19, 1261–1267 (2013).
Gombolay, G. Y. & Chitnis, T. Pediatric neuromyelitis optica spectrum disorders. Curr. Treat. Opt. Neurol. 20, 19 (2018).
Srivastava, A., Sarkar, H., Gupta, N. & Patro, R. RapMap: a rapid, sensitive and accurate tool for mapping RNA-seq reads to transcriptomes. Bioinformatics 32, i192–i200 (2016).
van der Maaten, L. & Hinton, G. Visualizing data using t-SNE. J. Mach. Learn. Res. 9, 2579–2605 (2008).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
DeLuca, D. S. et al. RNA-SeQC: RNA-seq metrics for quality control and process optimization. Bioinformatics 28, 1530–1532 (2012).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30, 923–930 (2014).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Assarsson, E. et al. Homogenous 96-plex PEA immunoassay exhibiting high sensitivity, specificity, and excellent scalability. PLoS ONE 9, e95192 (2014).
Lundberg, M., Eriksson, A., Tran, B., Assarsson, E. & Fredriksson, S. Homogeneous antibody-based proximity extension assays provide sensitive and specific detection of low-abundant proteins in human blood. Nucleic Acids Res. 39, e102 (2011).
Acknowledgements
The authors would like to thank all the patients who participated in this research. The Peabody Foundation supported this work. The Partners Pediatric MS Center at Massachusetts General Hospital is supported in part by the National MS Society. Authors would like to thank the Single Cell Core (SCC), laboratory of Dr. Allon Klein, Department of Systems Biology at Harvard Medical School (HMS) for sequencing inDrop single cell library. Authors would also like to thank The Broad institute’s Genomic service for sequencing DGE libraries.
Author information
Authors and Affiliations
Contributions
T.C. conceived and designed the research studies; G.G. and T.C. validated all clinical and radiological patient data; S.S. performed all experiments and analyzed data for flow cytometry, western blot and ELISA; S.S. and H.L. performed transcriptomic data analysis; S.S. and T.C. interpreted results of experiments; R.R. assisted in experiments for western blot; S.S.; R.R. and H.L. prepared figures; S.S. and T.C. drafted manuscript; S.S.; T.C.; R.R.; G.G.; H.L. and T.R. edited, revised and approved final version of manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Saxena, S., Lokhande, H., Gombolay, G. et al. Identification of TNFAIP3 as relapse biomarker and potential therapeutic target for MOG antibody associated diseases. Sci Rep 10, 12405 (2020). https://doi.org/10.1038/s41598-020-69182-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-69182-w
This article is cited by
-
miRNA 548a-3p as biomarker of NEDA-3 at 2 years in multiple sclerosis patients treated with fingolimod
Journal of Neuroinflammation (2023)
-
Association of rare variants in genes of immune regulation with pediatric autoimmune CNS diseases
Journal of Neurology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.