Abstract
Iron, an essential element for all organisms, acts as a cofactor of enzymes in bacterial degradation of recalcitrant aromatic compounds. The bacterial family, Sphingomonadaceae comprises various degraders of recalcitrant aromatic compounds; however, little is known about their iron acquisition system. Here, we investigated the iron acquisition system in a model bacterium capable of degrading lignin-derived aromatics, Sphingobium sp. strain SYK-6. Analyses of SYK-6 mutants revealed that FiuA (SLG_34550), a TonB-dependent receptor (TBDR), was the major outer membrane iron transporter. Three other TBDRs encoded by SLG_04340, SLG_04380, and SLG_10860 also participated in iron uptake, and tonB2 (SLG_34540), one of the six tonB comprising the Ton complex which enables TBDR-mediated transport was critical for iron uptake. The ferrous iron transporter FeoB (SLG_36840) played an important role in iron uptake across the inner membrane. The promoter activities of most of the iron uptake genes were induced under iron-limited conditions, and their regulation is controlled by SLG_29410 encoding the ferric uptake regulator, Fur. Although feoB, among all the iron uptake genes identified is highly conserved in Sphingomonad strains, the outer membrane transporters seem to be diversified. Elucidation of the iron acquisition system promises better understanding of the bacterial degradation mechanisms of aromatic compounds.
Similar content being viewed by others
Introduction
Iron is an essential nutrient utilised as a cofactor for enzymes that control various life phenomena such as respiration, dissimilation, and stress response1. Iron exists mainly in ferrous and ferric forms. Ferrous iron is soluble and highly bioavailable; however, the predominant form of iron in the environment is an insoluble ferric form2. Many bacteria secrete specific high-affinity siderophores which form complexes with ferric iron and then uptake these ferric-siderophore complexes to enhance iron acquisition over competitors3,4,5. Besides, pathogenic bacteria can acquire haem and transferrin specifically from their host cells6,7.
Gram-negative bacteria need to transport iron through both the outer and inner membranes. TonB-dependent receptors (TBDRs) mediate transport of ferric complexes (e.g. siderophore, haem, and transferrin) across the outer membrane1,8,9. TBDRs utilise energy derived from the proton motive force transmitted by TonB-ExbB-ExbD complex (Ton complex) localised in the inner membrane (Ton system)10. Beside siderophores, the Ton system is involved in the uptake of vitamin B12, saccharide, aromatic compounds, and metals such as nickel, copper, and lanthanoid11,12,13,14,15,16. The uptake of the ferric complex across the inner membrane is mainly achieved by ATP-binding cassette (ABC) transporters9. A major facilitator superfamily (MFS) transporter, FptX, is also known to mediate the ferric-siderophore (pyochelin) uptake in Pseudomonas aeruginosa17. The ferric iron transported into the cytoplasm is eventually reduced to ferrous iron. In contrast, the transport of ferrous iron across the inner membrane is mainly facilitated by Feo and divalent metal ion transporters such as MntH and ZupT18. The Feo system encoded by feoABC is considered the primary uptake system for ferrous iron18. FeoB is a permease that utilises energy acquired by its N-terminus GTPase domain and transports ferrous iron. FeoA and FeoC are accessory proteins important for FeoB multimer formation19,20. However, feoC is conserved in only gammaproteobacteria, and the organisation of the feo operon varies among bacteria18. The transcription of most of the genes involved in iron uptake and metabolism is regulated by the ferric uptake regulator (Fur)21,22. Excess ferrous iron results in the binding of the ferrous iron-Fur complex to the Fur box in the promoter regions to repress their transcription.
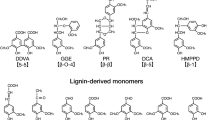
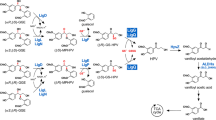
Until now, the iron acquisition pathways of proteobacteria have been mainly investigated in pathogenic bacteria9. To the best of our knowledge, reports regarding the transporters involved in iron acquisition by alphaproteobacteria are limited to Rhizobiales and Caulobacterales17,23,24. The family Sphingomonadaceae in alphaproteobacteria comprises of many unique strains capable of degrading certain recalcitrant aromatic compounds such as lignin-derived aromatic compounds, dibenzo-p-dioxin, polycyclic aromatic hydrocarbons, and pentachlorophenol25,26,27,28. These strains are valuable for bioremediation and the production of industrially useful chemicals from biomass25,26,29. Iron, an essential factor in the bacterial degradation of aromatic compounds is located in the active centre of O-demethylases, aromatic-ring-hydroxylating oxygenases, and ring-cleavage enzymes30,31. Sphingobium sp. SYK-6 produces a promising platform chemical (2-pyrone-4,6-dicarboxylate) that enables the synthesis of functional polymers, during the degradation of lignin-derived aromatics, thereby indicating that the SYK-6 catabolic system is useful to lignin valorisation32,33,34. Iron also has essential roles in the catabolism of lignin-derived aromatic compounds as exemplified by the presence of ferrous iron in the active centres of ring cleavage dioxygenases and a multicomponent O-demethylase35,36,37,38,39. Analysis of the outer membrane transporters of lignin-derived aromatic compounds in SYK-6 has indicated that ddvT, one of the 74 TBDR genes, encodes the outer membrane transporter of a lignin-derived biphenyl compound, 5,5′-dehydrodivanillate, and tonB1 is involved in this transport among the six tonB homologs13. On the other hand, disruption of tonB2 was seen to affect the growth of SYK-6 and decrease the activity of ferrous iron-requiring 5,5′-dehydrodivanillate O-demethylase, thereby suggesting that tonB2 plays a role in the iron acquisition process13.
In this study, we identified the SYK-6 transporters mainly involved in the uptake of iron across the outer and inner membranes through the analyses of mutants of the candidate iron uptake genes, their promoter activities in response to iron, and the binding of Fur to their promoter regions to gain insight into the iron acquisition system of Sphingomonadaceae.
Results
Identification of tonB involved in iron uptake
SYK-6 has six tonB homologs in its genome (Table 1)13. To identify the particular tonB involved in iron uptake among the six tonB homologs, we evaluated the growth of their mutants under iron-replete and -limited conditions. We used vanillate (VA), a major intermediate of lignin biodegradation, and its metabolite, protocatechuate (PCA), as lignin-derived carbon sources. Since SYK-6 cannot grow on single sugars or organic acids, SEMP (10 mM sucrose, 10 mM glutamate, 0.13 mM methionine, and 10 mM proline)40 was used as non-lignin-derived carbon source. We examined the capacity of the wild type and tonB2–6 mutants (a tonB1 mutant was unable to be obtained despite repeated experiments) to grow in a Wx medium (34 µM FeSO4) containing VA, PCA, or SEMP in the presence (the iron-limited condition) and absence (the iron-replete condition) of 100 µM 2,2′-dipyridyl (DIP), an iron chelator41. While ∆tonB2 cells only showed growth retardation when grown on VA and SEMP under iron-replete conditions (Fig. 1), under iron-limited conditions, ∆tonB2 cells showed further growth retardation and almost lost the capacity to grow on VA. However, the growth characteristic of ∆tonB3–6 was mostly the same as that of the wild type. Although under iron-replete conditions, the growth of ∆tonB2–6 cells on PCA matched that of the wild type, iron limiting conditions showed growth retardation of ∆tonB2 cells on PCA. The growth of ∆tonB2 cells on VA, PCA, and SEMP under iron-limited conditions was recovered by the introduction of a tonB2-carrying plasmid, indicating that the growth retardation described above was caused by the disruption of tonB2 (Fig. S1).
Growth of tonB mutants on VA, PCA, and SEMP. Cells of SYK-6, ∆tonB2, ∆tonB3, ∆tonB4, ∆tonB5, and ∆tonB6 were cultured in Wx medium containing 5 mM VA, 5 mM PCA, or SEMP in the presence or absence of 100 µM DIP. Cell growth was monitored by measuring the OD660. Each value is the average ± the standard deviation of three independent experiments.
∆tonB2 retained the capacity to grow on PCA and SEMP under iron-limited conditions, implying the involvement of other tonB in iron acquisition. We examined the growth of ∆tonB3456 and ∆tonB23456 cells on VA, PCA, and SEMP under iron-limited conditions (Fig. S2). The growth of ∆tonB3456 cells on VA and PCA was somewhat retarded compared with that of the wild type. Besides, the growth of ∆tonB23456 cells on PCA and SEMP was lower than that of ∆tonB2 cells indicating that any of the tonB3–6 appears to have some involvement in iron acquisition. To evaluate the involvement of tonB1 in iron acquisition, we introduced a plasmid carrying tonB1 into ∆tonB2 cells. While the growth of the tonB2-complemented ∆tonB2 on VA, PCA, and SEMP under iron-limited conditions was seen to recover, the introduction of tonB1 did not have a positive effect on the growth of ∆tonB2 (Fig. S1). These results indicate that tonB1 could not replace the function of tonB2.
We assessed the cellular localisation of TonB2 by performing western blot analysis using anti-TonB2 antibodies against a cell extract and a total membrane fraction prepared from SYK-6 grown on LB (Fig. S3). A clear signal was observed in the total membrane fraction, suggesting that TonB2 is localised in the cell membrane. The production of TonB2 in the cell membrane was also confirmed in tonB2-complemented ∆tonB2 cells (Fig. S4). All these results suggest that TonB2, a component of the Ton complex, plays a vital role in growth under iron-limited conditions.
Characterisation of a TBDR gene downstream of tonB2
A previous phylogenetic analysis has indicated that SLG_34550 just downstream of tonB2 is classified into a clade comprising of known iron uptake TBDRs (Table 1)13. To identify the TBDR gene involved in iron uptake, SLG_34550, designated as fiuA, was deleted to obtain a fiuA mutant (∆fiuA), and the capacity of the mutant to grow on VA, PCA, and SEMP was measured (Fig. S5). ∆fiuA cells showed growth retardation on VA and SEMP under iron-replete conditions, similar to ∆tonB2 cells (Fig. 2) which increased further under iron-limited conditions ∆fiuA not growing at all on VA and growth retardation also seen on PCA. The growth of ∆fiuA cells on VA, PCA, and SEMP under iron-limited conditions was recovered by the introduction of a fiuA-carrying plasmid (Fig. S6).
Growth of a fiuA mutant on VA, PCA, and SEMP. Cells of SYK-6 and ∆fiuA were cultured in Wx medium containing 5 mM VA, 5 mM PCA, or SEMP in the presence or absence of 100 µM DIP. Cell growth was monitored by measuring the OD660. Each value is the average ± the standard deviation of three independent experiments.
We measured the intracellular iron concentrations of ∆tonB2, ∆fiuA, and wild-type cells grown in SEMP to evaluate whether a reduction in the intracellular iron concentration resulted in the decrease in the growth of ∆tonB2 and ∆fiuA (Fig. 3). The intracellular iron concentrations of ∆tonB2 and ∆fiuA cells reduced to approximately 37% and 61% of that of the wild type, respectively. These results indicate the involvement of tonB2 and fiuA in iron acquisition.
Intracellular iron concentrations of ∆tonB2 and ∆fiuA. Intracellular iron concentrations of wild type, ∆tonB2, and ∆fiuA were determined as described in the Methods. Each value is the average ± the standard deviation of three independent experiments. **P < 0.01, ***P < 0.001 (one-way ANOVA with Dunnett’s multiple comparisons).
Promoter activities of tonB2 and fiuA under iron-limited conditions
Rodionov et al. discovered a 19-bp Fur box sequence conserved among alphaproteobacteria using comparative genomic analysis42. Incomplete inverted repeat sequences similar to this Fur box sequence were found upstream of each of tonB2 and fiuA (Fig. 4A, Table S1). We evaluated promoter activities of SYK-6 cells harbouring the reporter plasmid carrying a transcriptional fusion of a Fur box-containing promoter region of tonB2 or fiuA with lacZ (Fig. 4B–D). Promoter activities of the cells carrying the tonB2 and fiuA promoter regions were seen to increase 1.6-fold and 8.0-fold, respectively, under iron-limited conditions compared to iron-replete conditions. The addition of Fe2+ reduced the activities to levels comparable with those of the cells under iron-replete conditions, indicating that the expression of tonB2 and fiuA was induced under iron-limited conditions. However, the promoter activity of tonB2 was significantly higher (61-fold) than that of the fiuA promoter under iron-replete conditions. These findings suggest that tonB2 is expressed at a high level, even under iron-replete conditions. Western blot analysis using anti-TonB2 antibodies against the total membrane fractions obtained from SYK-6 grown with and without DIP demonstrated the production of almost equal amount of TonB2 between both membrane samples, suggesting that expression of tonB2 is not greatly influenced by iron-limitation (Fig. S7). In contrast, the promoter activity of tonB1 did not demonstrate any change regardless of the presence or absence of DIP (Fig. S8).
Promoter activities of tonB2 and fiuA under iron-replete or limited conditions. (A) Gene organisation of tonB2 and fiuA. Fur box-like sequences upstream of tonB2 and fiuA are indicated by red squares. Genes: SLG_34530, hypothetical protein; tonB2, TonB-like protein; fiuA, TBDR; SLG_34560, putative hydroxylase; SLG_34570, putative oxidoreductase; SLG_34580, putative oxidoreductase. (B–D) β-galactosidase activities of SYK-6 cells harbouring pS-t2 (B), pS-fiuA (C), and pSEVA225 (D) grown in Wx-SEMP with or without 100 µM DIP and 100 µM FeCl2. The DNA fragments used for the promoter analysis are shown on the left. Each value is the average ± the standard deviation of three independent experiments. ns, P > 0.05, ***P < 0.001, ****P < 0.0001 (one-way ANOVA with Dunnett’s multiple comparisons). (E) RT-PCR analysis of tonB2 and fiuA. Total RNA used for cDNA synthesis was isolated from SYK-6 cells grown in Wx-SEMP with 100 µM DIP. The regions to be amplified are indicated by black bars below the genetic map. Lanes: M, molecular size markers; g, control PCR with the SYK-6 genomic DNA; ‘ + ’ and ‘−’, RT-PCR with and without reverse transcriptase, respectively.
Although an independent promoter region was found upstream of each of tonB2 and fiuA, both genes were likely to be transcribed in the same transcription unit under iron-limited conditions (Fig. 4). Reverse transcription (RT)-PCR analysis of tonB2 and fiuA was performed using cDNA obtained from total RNA isolated from SYK-6 cells grown under iron-limited conditions. An amplification product between tonB2 and fiuA was observed (Fig. 4E), suggesting that fiuA is mainly transcribed from the more active tonB2 promoter under iron-limited conditions. These results led to the conclusion that tonB2 and fiuA play significant roles in iron acquisition.
TBDR genes other than fiuA required for normal growth under iron-limited conditions
∆fiuA retained the capacity to grow on PCA and SEMP under iron-limited conditions (Fig. 2), suggesting that other TBDR genes were also involved in iron uptake. In the phylogenetic tree of SYK-6 TBDRs constructed in our previous study, SLG_04340, SLG_04380, SLG_10860, and SLG_17010 in addition to FiuA were classified into two phylogenetic clades, containing siderophore and haem uptake TBDRs13. Therefore, we focused on these TBDR genes as candidate iron uptake genes other than fiuA (Table 1, S2). A Fur box-like sequence was found just upstream of each gene except SLG_04340 (Table S1). RT-PCR analysis revealed that SLG_04320-SLG_04360 constituted an operon, and a Fur box-like sequence was found just upstream of SLG_04320 (Fig. S9). We evaluated promoter activities of SYK-6 cells harbouring a reporter plasmid carrying a transcriptional fusion of a Fur box-containing promoter region of SLG_04340, SLG_04380, SLG_10860, or SLG_17010 with lacZ (Fig. S10A-D). Promoter activities were detected in all the cells except the one harbouring a plasmid carrying the upstream region of SLG_17010. Promoter activities of the cells carrying the SLG_04340 and SLG_04380 promoter regions were seen to increase 1.6-fold and 12-fold, respectively, under iron-limited conditions (Fig. S10A, B). Next, mutants of SLG_04340, SLG_04380, and SLG_10860 were constructed and their growth was compared with the wild type on VA, PCA, and SEMP under iron-limited and replete conditions (Fig. S5, S11). Since disruption of these genes did not show significant effect on the growth of SYK-6, we constructed ∆fiuA 4340, ∆fiuA 4380, and ∆fiuA 10860 and evaluated their growth under iron-limited conditions (Fig. S12). The growth of these double mutants on SEMP was further retarded as compared to that of the ∆fiuA. The growth of the former two mutants was also delayed on PCA. Thus, not only fiuA but also SLG_04340, SLG_04380, and SLG_10860 are involved in iron acquisition. Further, we constructed ∆fiuA 4340 4380 and ∆fiuA 4340 4380 10860 and evaluated their capacity to grow under iron-limited conditions (Fig. S13). When grown on SEMP, ∆fiuA 4340 4380 cells exhibited almost the same level of growth as that of ∆fiuA 4380, however, ∆fiuA 4340 4380 10860 cells showed substantial growth retardation as compared to ∆fiuA 4340 4380. By contrast, multiple mutations did not affect the capacity of these cells to grow on PCA, implying that different TBDRs are involved in the iron uptake during growth on PCA.
Identification of an inner membrane iron transporter
To identify the inner membrane iron transporters, we searched for SYK-6 genes showing similarity with known inner membrane transporters. The SYK-6 genome consists of four genes (SLG_06990, SLG_13630, SLG_36840, and SLG_p-00340) showing similarity with known inner membrane transporter genes involved in the uptake of siderophore and ferrous iron (Table 1, S2). To evaluate their involvement in iron acquisition, we constructed mutants of these genes and measured their growth on VA, PCA, and SEMP under iron-limited and replete conditions (Fig. 5A). ∆36840 showed growth retardation on VA and PCA under iron-limited conditions. SLG_36840 has 28–29% amino acid sequence identity with the ferrous iron inner membrane transporter gene (feoB) of Escherichia coli K-12 (AAC76434) and Pseudomonas aeruginosa PAO1 (AAG07746); thus SLG_36840 was designated feoB. The introduction of a feoB-carrying plasmid into ∆36840 (∆feoB) recovered the growth of ∆feoB cells on VA and PCA under iron-limited conditions (Fig. S14). In addition, the intracellular iron concentration of ∆feoB cells grown on SEMP was reduced to approximately 52% of that of the wild type (Fig. 5B). There is a feoA-like gene (SLG_36850), just upstream of feoB that encodes an important factor for FeoB multimer formation18. RT-PCR analysis showed that feoA and feoB comprise an operon (Fig. 5C, D). Since a Fur box was found upstream of feoA, the promoter activities of a feoA promoter region containing the Fur box were evaluated (Fig. 5E). Promoter activity was observed to be increased 1.5-fold under iron-limited conditions. All these results indicate that feoAB is involved in the uptake of ferrous iron across the inner membrane. However, it is not clear why the growth of SYK-6 on SEMP remaining unaffected by the disruption of feoB.
Identification of a transporter gene involved in the iron uptake across the inner membrane. (A) Growth of mutants of putative inner membrane iron transporter genes on VA, PCA, and SEMP. Cells of SYK-6, ∆6990, ∆13630, ∆feoB (∆36840), and ∆p-00340 were cultured in Wx medium containing 5 mM VA, 5 mM PCA, or SEMP in the presence or absence of 100 µM DIP. Cell growth was monitored by measuring the OD660. (B) Intracellular iron concentrations of wild type and ∆feoB. **P < 0.01 (two-tailed, unpaired t-test). (C) Gene organisation of feoAB. Genes: SLG_36830, putative single-stranded DNA-binding protein; feoB, putative ferrous iron transporter protein B; feoA, putative ferrous iron transporter protein A; SLG_36860, putative ubiquinone biosynthesis protein. (D) RT-PCR analysis of feoAB. Total RNA used for cDNA synthesis was isolated from SYK-6 cells grown in Wx-SEMP with 100 µM DIP. The region to be amplified is indicated by a bar below the genetic map (C). Lanes: M, molecular size markers; g, control PCR with the SYK-6 genomic DNA; ‘ + ’ and ‘−’, RT-PCR with and without reverse transcriptase, respectively. (E) β-galactosidase activities of SYK-6 cells harbouring pS-feoA grown in Wx-SEMP with or without 100 µM DIP and 100 µM FeCl2. The DNA fragment used for the promoter analysis is shown on the left. Each value is the average ± the standard deviation of three independent experiments. ns, P > 0.05, **P < 0.01 (one-way ANOVA with Dunnett’s multiple comparisons).
Identification of Fur involved in the regulation of iron uptake genes
SYK-6 has two fur-like genes, fur1 (SLG_29410) and fur2 (SLG_05570), which showed 21% amino acid sequence identity with each other and exhibited 35% and 21%, 38% and 21%, and 68% and 20% identity with Fur of P. aeruginosa PAO1 (AAG08150), E. coli K-12 (AAC73777), and Caulobacter crescentus NA1000 (ACL93522), respectively (Table 1). The involvement of fur1 and fur2 in the transcriptional regulation of tonB2, fiuA, SLG_04340, SLG_04380, and feoAB was evaluated by attempting to disrupt these genes, which resulted in only a fur2 mutant being obtained (Fig. S5). We assessed the promoter activities of ∆fur2 harbouring a reporter plasmid carrying a transcriptional fusion of each Fur box-containing promoter region of the above genes with lacZ (Fig. S15). However, ∆fur2 cells harbouring each plasmid showed almost the same level of promoter activities with wild type under iron-replete conditions, indicating that fur2 is not involved in their transcriptional regulation. Next, we examined whether Fur1 could bind these promoter regions using purified Fur1 obtained from fur1-expressing E. coli BL21(DE3) (Fig. S16). Electrophoretic mobility shift assay revealed that Fur1 was bound to the promoter regions of tonB2, fiuA, SLG_04340, SLG_04380, and feoA and was not bound to the promoter regions without the Fur box (Fig. 6A–E). These results strongly suggest that Fur1 regulates the expression of these genes by binding to the Fur box.
Fur1 binds to the Fur box sequences upstream of the iron uptake genes. The panels on the left show the DNA fragments used for EMSA. The panels on the right show the results of the EMSA of the binding of purified Fur1 to DNA probes of promoter regions of tonB2 (A), fiuA (B), SLG_04340 (C), SLG_04380 (D), and feoA (E). The uncropped gel images are shown in Fig. S21.
The results of this study demonstrated that the transcription of fiuA was driven from the tonB2 promoter under iron-limited conditions (Fig. 4E), and Fur1 was able to bind to the Fur box upstream of fiuA (Fig. 6B). Based on these results, it was hypothesised that the transcriptional regulation of fiuA was as described below. Under iron-replete conditions, Fur1 binds to the Fur box upstream of fiuA, interrupting the transcription from the tonB2 promoter that is active even under iron-replete conditions. Under iron-limited conditions, Fur1 is released from the Fur box, and then fiuA is strongly co-transcribed with tonB2. This hypothesis was verified by constructing a plasmid carrying a transcriptional fusion of a tonB2 promoter region and fiuA promoter region with lacZ (pS-t2-fiuA) and evaluating the promoter activities of SYK-6 cells harbouring pS-t2-fiuA (Fig. S17A). Under iron-limited conditions, the promoter activities between the cells harbouring pS-t2-fiuA and the cells harbouring pS-t2, which carries a tonB2 promoter-lacZ fusion, were almost the same. Under iron-replete conditions, while the activity of SYK-6 cells harbouring pS-t2 decreased to only ca. 63% of the activity under iron-limited conditions, the activity of SYK-6 cells harbouring pS-t2-fiuA was drastically reduced (ca. 11%). These observations, thus, support the hypothesis (Fig. S17B).
PCA is a potential siderophore used for SYK-6
SYK-6 cells were grown on Wx-SEMP agar medium containing chrome azurol S (CAS) to examine whether SYK-6 secretes siderophores (Fig. S18). SYK-6 cells grown for 144 h formed a slight halo around the colony, suggesting that SYK-6 cells weakly secrete siderophores. However, no genes showed similarity with known siderophore synthetase genes in the SYK-6 genome. Besides, the size of the halo did not change when ∆tonB2 and ∆fiuA were assayed.
As shown in Figs. 1 and 2, ∆tonB2 and ∆fiuA did not exhibit severe growth retardation on PCA as compared to VA and SEMP. This fact may imply that the TonB2-FiuA-independent iron acquisition system functions in SYK-6 during its growth on PCA. Since PCA is known to form a complex with iron43,44, we examined whether the addition of PCA improves the growth of ∆tonB2 and ∆fiuA on SEMP under iron-limited conditions (Fig. 7, Fig. S19). Interestingly, the addition of 100 µM PCA improved their growth after 40 h as seen from the OD660 values of the cultures which was 1.3- to 1.4-fold higher than that without PCA with high concentrations of PCA (500 to 1,000 µM) showing more effect on promoting growth (1.6- to 1.8-fold). We measured the growth of a fiuA ligAB double mutant (∆fiuA ligAB) to confirm that PCA did not contribute to these growth improvements as a carbon source. ligAB encodes PCA 4,5-dioxygenase which is essential for growth of SYK-6 on PCA45. ∆fiuA ligAB cells exhibited increased growth (1.8- to 2.5-fold) when PCA (100 to 1,000 µM) was added (Fig. 7). These results suggest that SYK-6 cells utilise PCA as a siderophore or secrete unknown siderophore(s) induced by PCA. Since ∆tonB2 and ∆fiuA showed reduced growth on VA which is metabolised via PCA45 and the presence of PCA did not promote halo formation in CAS assay (Fig. S18), PCA appears to act as a siderophore. Notably, the growth improvement of SYK-6 and ∆ligAB cells on SEMP was modest with the addition of PCA, unlike ∆fiuA ligAB cells (Fig. 7). These results suggest that TonB2 and FiuA are mainly involved in the iron acquisition and that the pathway utilising PCA as a siderophore is ancillary.
PCA enhances the growth of ∆tonB2 and ∆fiuA under iron-limited conditions. Cells of SYK-6, ∆tonB2, ∆fiuA, ∆fiuA ligAB, and ∆ligAB were cultured in Wx-SEMP containing 100 µM DIP with or without PCA (100 µM, 500 µM, or 1,000 µM). Cell growth was monitored by measuring the OD660. Each bar shows a relative value of OD660 at 40 h of cultures in the presence of PCA when the OD660 at 40 h of culture in the absence of PCA was set to 1.0 (leftmost bars). Each value is the average ± the standard deviation of three independent experiments. ns, P > 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001 (one-way ANOVA with Dunnett’s multiple comparisons). Individual growth curves are shown in Fig. S19.
Discussion
We conclude that the TonB2-FiuA system plays a significant role in iron acquisition in SYK-6 based on the following observations (Fig. 8): (i) tonB2 and fiuA are essential for normal growth on various carbon sources, (ii) the promoter activities of tonB2 and fiuA were activated under iron-limited conditions and suggested to be regulated by Fur1, and (iii) intracellular iron levels in ∆tonB2 and ∆fiuA cells were significantly reduced. tonB2 and fiuA constituted an operon, and a Fur box sequence was found upstream of each of tonB2 and fiuA (Fig. 4). The transcription of fiuA is tightly repressed by binding of Fur1 to its Fur box under iron-replete conditions and strongly activated from the tonB2 promoter under iron-limited conditions (Fig. S17). This transcriptional control is a sophisticated system that regulates the transcription of fiuA from the tonB2 promoter, which shows strong activities regardless of iron-replete and -limited conditions. There are examples of co-transcription of tonB and TBDR genes46; however, to our knowledge, this regulation system for tonB2-fiuA has not been documented. Considering the tonB2 expression profile, which is highly expressed regardless of iron-replete and -limited conditions, TonB2 likely interacts with not only FiuA but also other TBDRs to acquire iron and other nutrients.
Proposed iron acquisition pathways in SYK-6. Ferric iron acquisition across the outer membrane is mediated by the TonB2-FiuA system. Other TBDRs (SLG_04340, SLG_04380, and SLG_10860) are also involved in the outer membrane transport of ferric iron. In the inner membrane, FeoB, plays a vital role in the uptake of ferrous iron, together with unidentified ferric iron transporter(s). SLG_04360 showing similarity with ferric reductase FprA of Pseudomonas putida KT2440 (48% identity) may be involved in ferric reduction in the cytoplasm. Fur1 represses the expression of iron uptake genes under iron-replete conditions.
Bacteria living in unstructured environments, such as an open ocean, utilise amphipathic siderophores in their cell membrane to prevent loss of siderophores5,47. In contrast, bacteria commonly produce highly diffusive siderophores in structured environments, such as soil5,47. SYK-6 displayed weak halo formation in the CAS assay (Fig. S18) and had no genes showing similarity with known siderophore biosynthesis genes. The halo formation of ∆tonB2 and ∆fiuA was mostly the same as the wild type (Fig. S18), suggesting that SYK-6 does not promote producing and secreting siderophores even under iron-limited conditions, unlike other siderophore-producing bacteria9. A recent study has suggested that Synechocystis sp. strain PCC 6803 does not produce siderophores, and its TBDRs mediate the uptake of free iron and ferric-siderophores produced by other bacteria (xenosiderophore)48. In addition to all of the above, considering there are 74 TBDR-like genes in the SYK-6 genome, TBDRs of SYK-6 may mediate the uptake of free iron and xenosiderophore. Although we revealed that SYK-6 could utilise PCA as an ancillary siderophore (Fig. 7), the TonB2-FiuA system, mainly playing a vital role in the iron acquisition of SYK-6, did not involve uptake of the PCA-iron complex. In future, we need to clarify whether TonB2-FiuA takes up free iron or ferric complexes.
Ferric iron is probably reduced to ferrous iron in the periplasm, and then it is taken up by the Feo system (Fig. 8). In addition to the Feo system, ferric iron appears to be incorporated into the cytoplasm using unidentified transporters (e.g. ABC transporters). The reduction of ferric iron in the periplasm is essential for ferrous iron uptake by the Feo system, however, genes similar to periplasmic reductase such as vciB of V. cholerae were not found in the SYK-6 genome49,50. On the other hand, SLG_04360, which constituted an operon with SLG_04340 (Fig. S9), showed 48% identity with NADPH-dependent ferric reductase FprA (AAN67259) of Pseudomonas putida KT244051, implying that SLG_04360 is involved in ferric reduction in the cytoplasm (Fig. 8).
SYK-6 has two genes belonging to the ferric uptake regulator family (fur1 and fur2). We found that Fur1, showing 68% identity with Fur of C. crescentus NA1000 (ACL93522), regulates the transcription of iron uptake genes (Fig. 6, S15). Because Fur2 shows around only 20% amino acid sequence identity with Fur1 and other known Fur variants, Fur2 may regulate the transcription of other metal uptake genes (e.g. zinc and manganese). In alphaproteobacteria, transcriptional regulators other than Fur that respond to iron, such as RirA and Irr, are known to regulate the iron acquisition and storage genes in Rhizobiales and Rhodobacterales52. However, comparative genomic analyses revealed that iron acquisition in Sphingomonadaceae is regulated by Fur, consistent with our finding42,52.
We examined whether the genes involved in iron uptake in SYK-6 are conserved in ten Sphingomonad strains shown in Table S3. It has been reported that approximately 40% of Gram-negative bacteria with known genomes have more than two tonB-like genes53. The Sphingomonad strains compared here have 3–8 tonB-like genes, however, there was no gene showing > 40% amino acid sequence identity with tonB2. The proportion of Gram-negative bacteria, which have more than 30 TBDR genes in their genomes, is only ca. 16%12. Since the Sphingomonad strains have a large number of TBDR genes (from 39 to 153), they may play important roles not only in iron acquisition but also in other functions13,54. Five of the ten strains investigated showed the presence of TBDR-like genes, which showed 49–54% amino acid sequence identity with fiuA. Although their similarities with fiuA were somewhat low, a phylogenetic tree of all TBDRs of the five strains demonstrated the formation of a specific clade containing fiuA and its homologs mentioned above with known iron uptake TBDRs (Fig. S20). Among them, there was a tonB homolog in the vicinity of the fiuA-like gene of Novosphingobium nitrogenifigens DSM 19370, Novosphingobium sp. PP1Y, and Novosphingobium pentaromativorans US6-1. Thus, these fiuA and tonB homologs are likely to participate in the outer membrane iron uptake. In contrast, every Sphingomonad strain contained genes whose amino acid sequence showed 32–60% and 62–72% identities with those of SYK-6 feoA and feoB. FeoAB may therefore play an important role in the inner membrane ferrous iron uptake in Sphingomonadaceae.
Methods
Bacterial strains, plasmids, culture conditions, and substrates
The strains and plasmids used in this study are listed in Table S4. Sphingobium sp. SYK-6 (NBRC 103272/JCM 17495) and its mutants were grown at 30 °C with shaking (160 rpm) in LB or Wx minimal medium (containing 34 µM FeSO4) with SEMP40. Media for SYK-6 transformants and mutants was supplemented with 50 mg l−1 kanamycin (Km). E. coli strains were cultured in LB at 37 °C. Media for E. coli transformants carrying antibiotic resistance markers was supplemented with 25 mg l−1 km or 100 mg l−1 ampicillin (Amp). VA and PCA were purchased from Sigma-Aldrich and the Tokyo Chemical Industry Co., Ltd., respectively.
Mutant construction
Plasmids for gene disruption were constructed by amplifying ca. 1-kb fragments carrying upstream and downstream regions of each gene by PCR with SYK-6 genome DNA as a template and the primer pairs as shown in Table S5. The resultant fragments were inserted into the BamHI site in pAK405 by In-Fusion cloning (Takara Bio, Inc.). These plasmids were independently introduced into SYK-6 cells and its mutants by triparental mating, and candidate mutants were isolated as previously described55. Disruption of the genes was confirmed by colony PCR using primer pairs (Table S5). The plasmids for gene complementation of ∆tonB2, ∆fiuA, and ∆feoB (Table S4) were introduced into the mutants by electroporation.
Sequence analysis
Sequence analysis was performed using the MacVector program version 15.5.2. Sequence similarity searches, multiple alignments, and pairwise alignments were performed using the BLAST program56, Clustal Omega program57, and the EMBOSS program58, respectively. A phylogenetic tree was generated using the FigTree program (https://tree.bio.ed.ac.uk/software/figtree/).
RT-PCR analysis
SYK-6 cells grown in LB were harvested and washed twice with Wx medium. The cells were resuspended to an optical density at 600 nm (OD600) of 0.2 in Wx-SEMP and cultured at 30 °C until OD600 of the culture reached 0.5, after which they were incubated in the presence of 100 µM DIP for 2 h. Total RNA was isolated from the cells using an Illumina RNAspin Mini RNA isolation kit (GE Healthcare). The samples were treated with DNase I to remove any contaminating genomic DNA. Total RNA (4 µg) was reverse transcribed using SuperScript IV reverse transcriptase (Invitrogen) with random hexamer primers. The cDNA was purified using a NucleoSpin Gel and PCR Clean-up kit (Takara Bio, Inc.). PCR was performed with the cDNA, specific primers (Table S5), and Gflex DNA polymerase (Takara Bio, Inc.). The DNA obtained was electrophoresed on a 0.8% agarose gel.
Growth measurement
SYK-6 cells, its mutants, and complemented strains were grown in LB for 24 h. The cells were harvested by centrifugation at 4,800×g for 5 min, washed twice with Wx medium, and resuspended in 3 ml of the same medium. The cells were then inoculated in Wx medium containing SEMP, 5 mM VA, or 5 mM PCA to an OD660 of 0.2 with or without 100 µM DIP. Since SYK-6 exhibits auxotrophy for methionine when grown in a methoxy-group-free substrate, 0.13 mM methionine was added to the medium to grow on PCA. Cells were incubated at 30 °C with shaking (60 rpm) and cell growth was monitored every hour by measuring the OD660 with a TVS062CA biophotorecorder (Advantec Co., Ltd.). The complemented strains of ∆feoB, ∆fiuA, and ∆tonB2 were analysed by growing cells in Wx medium containing Km and 1 mM m-toluate (an inducer of the Pm promoter in pJB861).
Promoter assay
SYK-6 cells and ∆fur2 harbouring each plasmid (Table S4) grown in LB containing Km for 20 h were harvested by centrifugation at 4,800×g for 5 min, washed twice with Wx medium, and resuspended in 1 ml of the same medium. The cells were then inoculated in Wx-SEMP containing Km to an OD600 of 0.2. Samples were incubated at 30 °C until OD600 of the culture reached 0.5. Then, the cells were further incubated with or without 100 µM DIP and 100 µM FeCl2 for 2 h. β-galactosidase activity of the cells was measured using 2-nitrophenyl-β-D-galactopyranoside as described previously13 and expressed as Miller units.
Western blot analysis
A peptide corresponding to residues 247–266 (HGPDPRDRPLSDGQIKTIET) of TonB2 was synthesised and used as an antigen to obtain antisera against TonB2 in rabbits (Cosmo Bio, Inc.). Anti-TonB2-peptide antibodies were obtained by purification of the antiserum using peptide affinity column chromatography (Cosmo Bio, Inc.). Total membrane fractions were prepared as described previously from SYK-6 cells incubated in LB for 20 h with or without 100 µM DIP13. When total membrane fractions were prepared from the tonB2-complemented ∆tonB2, cells were incubated in LB containing Km and 1 mM m-toluate. TonB2 was detected by western blot analysis using anti-TonB2 antibodies (0.09 µg/ml) as described previously13. Horseradish peroxidase-conjugated goat anti-rabbit IgG antibodies (Invitrogen, 0.2 µg/ml) were used as the secondary antibodies. Protein concentrations were determined by the Bradford method using a Bio-Rad protein assay kit or Lowry’s assay with a DC protein assay kit (Bio-Rad Laboratories). TonB2 was detected using the ECL Western Blotting Detection System (GE Healthcare) with a LumiVision PRO image analyser (Aisin Seiki Co., Ltd).
Intracellular iron concentration measurement
SYK-6 cells and its mutants were grown in LB for 20 h, harvested by centrifugation at 4,800 × g for 5 min, washed twice with Wx medium, and resuspended in 1 ml of the same medium. The cells were then inoculated in Wx medium containing SEMP to an OD600 of 0.2. Samples were incubated at 30 °C for 6 h. The cells were harvested by centrifugation, washed twice with 50 mM Tris–HCl buffer (pH 7.5), and resuspended in 200 µl of the same buffer. The cells were disrupted by sonication to obtain cell lysates. The cell lysates were then centrifuged at 18,800 × g for 10 min and the protein concentration of the resulting supernatants (cell extracts) was determined. The iron concentration of cell extracts was determined using an Iron Assay Kit LS (Metallogenics Co., Ltd.) based on the ferrozine chromogenic method. Protein concentrations were determined using a Bio-Rad protein assay kit.
Expression of fur1 in E. coli and purification of Fur1
A fur1-coding region was PCR-amplified from SYK-6 genome DNA using primers listed in Table S5. A 0.4-kb NdeI-BamHI fragment carrying fur1 was inserted into the corresponding sites of pET-16b (pET-fur1) by In-Fusion cloning (Takara Bio, Inc.). E. coli BL21(DE3) harbouring pET-fur1 was grown in LB containing Amp at 30 °C until the OD600 of the culture reached 0.5, and then the expression of fur1 was induced for 4 h at 30 °C by addition of 1 mM isopropyl-β-D-thiogalactopyranoside. The cells were harvested by centrifugation at 4,800×g for 5 min, washed twice with 50 mM Tris–HCl buffer (pH 7.5) containing 100 mM NaCl, and resuspended in 200 µl of the same buffer. The cells were disrupted by sonication and cell lysate was obtained. The cell lysate was then centrifuged at 18,800×g for 10 min and the resulting supernatant was applied to a His Spin Trap (GE Healthcare). After centrifugation (100×g, 1 min, 4 °C), samples were washed 3 times with 50 mM Tris–HCl buffer (pH 7.5) containing 100 mM NaCl and 50 mM imidazole, and Fur1 was eluted with 50 mM Tris–HCl buffer (pH 7.5) containing 100 mM NaCl and 500 mM imidazole. Purified Fur1 was subjected to desalting and concentrating by centrifugal filtration using an Amicon Ultra 3 k (Merck Millipore). The purity of Fur1 was analysed by sodium dodecyl sulfate-15% polyacrylamide gel electrophoresis. Protein concentrations were determined by a Bio-Rad protein assay kit.
Electrophoretic mobility shift assay
DNA probes were PCR-amplified from SYK-6 genome DNA using the primers listed in Table S5. The DNA–protein binding reactions were performed at 20 °C for 30 min in 10 µl of binding buffer (50 mM Tris–HCl, 5 mM dithiothreitol, 50 mM MgCl2, 200 mM KCl, and 0.5% [wt./vol.] Tween 20, pH 7.5) containing 20 fmol DNA probe, 500 ng of purified Fur1, 1 µg of poly(dI-dC), and 100 mM MnSO4. The resulting samples were separated by electrophoresis on 2.5% agarose gel and signals were detected using SYBR Gold Nucleic Acid Gel Stain (Invitrogen).
CAS assay
Fifty millilitres of 1.2 g l−1 chrome azurol S solution was mixed with 10 ml of 1 mM FeCl3 (dissolved in 10 mM HCl) and 40 ml of 5 mM hexadecyltrimethylammonium bromide (CAS solution). Eighteen millilitres of a Wx-SEMP agar medium with or without PCA (final conc. 1 mM) was mixed with 2 ml of the CAS solution to prepare CAS assay plates. The cells of SYK-6, ∆tonB2, and ∆fiuA were grown in LB for 20 h, harvested by centrifugation at 4,800 × g for 5 min, washed twice with Wx medium, and resuspended in 1 ml of the same medium. Ten microlitres of the culture (OD600 = 10) was inoculated on a cellulose filter (12 mm) on a CAS assay plate and incubated for 6 days at 30 °C.
Statistics and reproducibility
All results were obtained from n = 3 independent experiments. Statistical tests were performed using GraphPad Prism8 (GraphPad software). One-way ANOVA with Dunnett’s multiple comparisons and unpaired, two-tailed t-test were used as shown in figure legends. P < 0.05 was considered statistically significant.
Data availability
All data supporting this study are available within the article and its Supplementary Information or are available from the corresponding author upon request.
Change history
02 October 2020
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Andrews, S. C., Robinson, A. K. & Rodríguez-Quiñones, F. Bacterial iron homeostasis. FEMS Microbiol. Rev. 27, 215–237. https://doi.org/10.1016/S0168-6445(03)00055-X (2003).
Melton, E. D., Swanner, E. D., Behrens, S., Schmidt, C. & Kappler, A. The interplay of microbially mediated and abiotic reactions in the biogeochemical Fe cycle. Nat. Rev. Microbiol. 12, 797–808. https://doi.org/10.1038/nrmicro3347 (2014).
Crosa, J. H. & Walsh, C. T. Genetics and assembly line enzymology of siderophore biosynthesis in bacteria. Microbiol. Mol. Biol. Rev. 66, 223–249. https://doi.org/10.1128/mmbr.66.2.223-249.2002 (2002).
Saha, M. et al. Microbial siderophores and their potential applications: a review. Environ. Sci. Pollut. Res. 23, 3984–3999. https://doi.org/10.1007/s11356-015-4294-0 (2016).
Kramer, J., Özkaya, Ö & Kümmerli, R. Bacterial siderophores in community and host interactions. Nat. Rev. Microbiol. 18, 152–163. https://doi.org/10.1038/s41579-019-0284-4 (2019).
Barber, M. F. & Elde, N. C. Escape from bacterial iron piracy through rapid evolution of transferrin. Science 346, 1362–1366. https://doi.org/10.1126/science.1259329 (2014).
Burkhard, K. A. & Wilks, A. Characterization of the outer membrane receptor ShuA from the heme uptake system of Shigella dysenteriae. Substrate specificity and identification of the heme protein ligands. J. Biol. Chem. 282, 15126–15136. https://doi.org/10.1074/jbc.M611121200 (2007).
Nikaido, H. Molecular basis of bacterial outer membrane permeability revisited. Microbiol. Mol. Biol. Rev. 67, 593–656. https://doi.org/10.1128/mmbr.67.4.593-656.2003 (2003).
Porcheron, G., Garénaux, A., Proulx, J., Sabri, M. & Dozois, C. M. Iron, copper, zinc, and manganese transport and regulation in pathogenic Enterobacteria: correlations between strains, site of infection and the relative importance of the different metal transport systems for virulence. Front. Cell. Infect. Microbiol. 3, 90. https://doi.org/10.3389/fcimb.2013.00090 (2013).
Celia, H., Noinaj, N. & Buchanan, S. K. Structure and stoichiometry of the Ton molecular motor. Int. J. Mol. Sci. 21, E375. https://doi.org/10.3390/ijms21020375 (2020).
Noinaj, N., Guillier, M., Barnard, T. J. & Buchanan, S. K. TonB-dependent transporters: regulation, structure, and function. Annu. Rev. Microbiol. 64, 43–60. https://doi.org/10.1146/annurev.micro.112408.134247 (2010).
Blanvillain, S. et al. Plant carbohydrate scavenging through TonB-dependent receptors: a feature shared by phytopathogenic and aquatic bacteria. PLoS ONE 2, e224. https://doi.org/10.1371/journal.pone.0000224 (2007).
Fujita, M. et al. A TonB-dependent receptor constitutes the outer membrane transport system for a lignin-derived aromatic compound. Commun. Biol. 2, 432. https://doi.org/10.1038/s42003-019-0676-z (2019).
Schauer, K., Rodionov, D. A. & de Reuse, H. New substrates for TonB-dependent transport: do we only see the “tip of the iceberg”?. Trends. Biochem. Sci. 33, 330–338. https://doi.org/10.1016/j.tibs.2008.04.012 (2008).
Han, Y. et al. A Pseudomonas aeruginosa type VI secretion system regulated by CueR facilitates copper acquisition. PLoS Pathog. 15, e1008198. https://doi.org/10.1371/journal.ppat.1008198 (2019).
Ochsner, A. M. et al. Use of rare-earth elements in the phyllosphere colonizer Methylobacterium extorquens PA1. Mol. Microbiol. 111, 1152–1166. https://doi.org/10.1111/mmi.14208 (2019).
Cuív, P. O., Clarke, P., Lynch, D. & O’Connell, M. Identification of rhtX and fptX, novel genes encoding proteins that show homology and function in the utilization of the siderophores rhizobactin 1021 by Sinorhizobium meliloti and pyochelin by Pseudomonas aeruginosa, respectively. J. Bacteriol. 186, 2996–3005. https://doi.org/10.1128/jb.186.10.2996-3005.2004 (2004).
Lau, C. K., Krewulak, K. D. & Vogel, H. J. Bacterial ferrous iron transport: the Feo system. FEMS Microbiol. Rev. 40, 273–298. https://doi.org/10.1093/femsre/fuv049 (2016).
Stevenson, B., Wyckoff, E. E. & Payne, S. M. Vibrio cholerae FeoA, FeoB, and FeoC interact to form a complex. J. Bacteriol. 198, 1160–1170. https://doi.org/10.1128/JB.00930-15 (2016).
Weaver, E. A., Wyckoff, E. E., Mey, A. R., Morrison, R. & Payne, S. M. FeoA and FeoC are essential components of the Vibrio cholerae ferrous iron uptake system, and FeoC interacts with FeoB. J. Bacteriol. 195, 4826–4835. https://doi.org/10.1128/JB.00738-13 (2013).
McHugh, J. P. et al. Global iron-dependent gene regulation in Escherichia coli. A new mechanism for iron homeostasis. J. Biol. Chem. 278, 29478–29486. https://doi.org/10.1074/jbc.M303381200 (2003).
Troxell, B. & Hassan, H. M. Transcriptional regulation by ferric uptake regulator (Fur) in pathogenic bacteria. Front. Cell. Infect. Microbiol. 3, 59. https://doi.org/10.3389/fcimb.2013.00059 (2013).
Balhesteros, H. et al. TonB-dependent heme/hemoglobin utilization by Caulobacter crescentus HutA. J. Bacteriol. https://doi.org/10.1128/JB.00723-16 (2017).
Sankari, S. & O’Brian, M. R. The Bradyrhizobium japonicum ferrous iron transporter FeoAB is required for ferric iron utilization in free living aerobic cells and for symbiosis. J. Biol. Chem. 291, 15653–15662. https://doi.org/10.1074/jbc.M116.734129 (2016).
Kamimura, N. et al. Bacterial catabolism of lignin-derived aromatics: new findings in a recent decade: update on bacterial lignin catabolism. Environ. Microbiol. Rep. 9, 679–705. https://doi.org/10.1111/1758-2229.12597 (2017).
Chai, B. et al. Sphingomonas wittichii strain RW1 genome-wide gene expression shifts in response to dioxins and clay. PLoS ONE 11, e0157008. https://doi.org/10.1371/journal.pone.0157008 (2016).
Sohn, J. H., Kwon, K. K., Kang, J. H., Jung, H. B. & Kim, S. J. Novosphingobium pentaromativorans sp. Nov., a high-molecular-mass polycyclic aromatic hydrocarbon-degrading bacterium isolated from estuarine sediment. Int. J. Syst. Evol. Microbiol. 54, 1483–1487. https://doi.org/10.1099/ijs.0.02945-0 (2004).
Copley, S. D. et al. The whole genome sequence of Sphingobium chlorophenolicum L-1: insights into the evolution of the pentachlorophenol degradation pathway. Genome Biol. Evol. 4, 184–198. https://doi.org/10.1093/gbe/evr137 (2012).
Perez, J. M. et al. Funneling aromatic products of chemically depolymerized lignin into 2-pyrone-4-6-dicarboxylic acid with Novosphingobium aromaticivorans. Green Chem. 21, 1340–1350. https://doi.org/10.1039/C8GC03504K (2019).
Ladino-Orjuela, G., Gomes, E., da Silva, R., Salt, C. & Parsons, J. R. Metabolic pathways for degradation of aromatic hydrocarbons by bacteria. Rev. Environ. Contam. Toxicol. 237, 105–121. https://doi.org/10.1007/978-3-319-23573-8_5 (2016).
Mallinson, S. J. B. et al. A promiscuous cytochrome P450 aromatic O-demethylase for lignin bioconversion. Nat. Commun. 9, 2487. https://doi.org/10.1038/s41467-018-04878-2 (2018).
Higuchi, Y. et al. Discovery of novel enzyme genes involved in the conversion of an arylglycerol-β-aryl ether metabolite and their use in generating a metabolic pathway for lignin valorization. Metab. Eng. 55, 258–267. https://doi.org/10.1016/j.ymben.2019.08.002 (2019).
Otsuka, Y. et al. Efficient production of 2-pyrone 4,6-dicarboxylic acid as a novel polymer-based material from protocatechuate by microbial function. Appl. Microbiol. Biotechnol. 71, 608–614. https://doi.org/10.1007/s00253-005-0203-7 (2006).
Qian, Y. et al. Engineered microbial production of 2-pyrone-4,6-dicarboxylic acid from lignin residues for use as an industrial platform chemical. BioResources 11, 6097–6109. https://doi.org/10.15376/biores.11.3.6097-6109 (2016).
Sugimoto, K. et al. Crystal structure of an aromatic ring opening dioxygenase LigAB, a protocatechuate 4,5-dioxygenase, under aerobic conditions. Structure 7, 953–965. https://doi.org/10.1016/S0969-2126(99)80122-1 (1999).
Sugimoto, K. et al. Molecular mechanism of strict substrate specificity of an extradiol dioxygenase, DesB, derived from Sphingobium sp. SYK-6. PLoS ONE 9, e92249. https://doi.org/10.1371/journal.pone.0092249 (2014).
Kasai, D., Masai, E., Miyauchi, K., Katayama, Y. & Fukuda, M. Characterization of the 3-O-methylgallate dioxygenase gene and evidence of multiple 3-O-methylgallate catabolic pathways in Sphingomonas paucimobilis SYK-6. J. Bacteriol. 186, 4951–4959. https://doi.org/10.1128/JB.186.15.4951-4959.2004 (2004).
Peng, X. et al. Cloning of a Sphingomonas paucimobilis SYK-6 gene encoding a novel oxygenase that cleaves lignin-related biphenyl and characterization of the enzyme. Appl. Environ. Microbiol. 64, 2520–2527 (1998).
Yoshikata, T. et al. Three-component O-demethylase system essential for catabolism of a lignin-derived biphenyl compound in Sphingobium sp. strain SYK-6. Appl. Environ. Microbiol. 80, 7142–7153. https://doi.org/10.1128/AEM.02236-14 (2014).
Kasai, D. et al. Characterization of FerC, a MarR-type transcriptional regulator, involved in transcriptional regulation of the ferulate catabolic operon in Sphingobium sp. strain SYK-6. FEMS Microbiol. Lett. 332, 68–75. https://doi.org/10.1111/j.1574-6968.2012.02576.x (2012).
Runci, F. et al. Contribution of active iron uptake to Acinetobacter baumannii pathogenicity. Infect. Immun. https://doi.org/10.1128/IAI.00755-18 (2019).
Rodionov, D. A., Gelfand, M. S., Todd, J. D., Curson, A. R. & Johnston, A. W. Computational reconstruction of iron- and manganese-responsive transcriptional networks in α-proteobacteria. PLoS Comput. Biol. 2, e163. https://doi.org/10.1371/journal.pcbi.0020163 (2006).
Liebl, W., Klamer, R. & Schleifer, K.-H. Requirement of chelating compounds for the growth of Corynebacterium glutamicum in synthetic media. Appl. Environ. Microbiol. 32, 205–210. https://doi.org/10.1007/BF00165889 (1989).
Andjelković, M. et al. Iron-chelation properties of phenolic acids bearing catechol and galloyl groups. Food Chem. 98, 23–31. https://doi.org/10.1016/j.foodchem.2005.05.044 (2006).
Masai, E., Katayama, Y. & Fukuda, M. Genetic and biochemical investigations on bacterial catabolic pathways for lignin-derived aromatic compounds. Biosci. Biotechnol. Biochem. 71, 1–15. https://doi.org/10.1271/bbb.60437 (2007).
Tong, Y. & Guo, M. Bacterial heme-transport proteins and their heme-coordination modes. Arch. Biochem. Biophys. 481, 1–15. https://doi.org/10.1016/j.abb.2008.10.013 (2009).
Kümmerli, R., Schiessl, K. T., Waldvogel, T., McNeill, K. & Ackermann, M. Habitat structure and the evolution of diffusible siderophores in bacteria. Ecol. Lett. 17, 1536–1544. https://doi.org/10.1111/ele.12371 (2014).
Qiu, G. W. et al. Outer membrane iron uptake pathways in the model Cyanobacterium Synechocystis sp. strain. PCC 6803. Appl. Environ. Microbiol. https://doi.org/10.1128/AEM.01512-18 (2018).
Peng, E. D. & Payne, S. M. Vibrio cholerae VciB mediates iron reduction. J. Bacteriol. 199, e00874-e916. https://doi.org/10.1128/JB.00874-16 (2017).
Schroder, I., Johnson, E. & de Vries, S. Microbial ferric iron reductases. FEMS Microbiol. Rev. 27, 427–447. https://doi.org/10.1016/S0168-6445(03)00043-3 (2003).
Yeom, J., Jeon, C. O., Madsen, E. L. & Park, W. Ferredoxin-NADP+ reductase from Pseudomonas putida functions as a ferric reductase. J. Bacteriol. 191, 1472–1479. https://doi.org/10.1128/JB.01473-08 (2009).
O’Brian, M. R. Perception and homeostatic control of iron in the Rhizobia and related bacteria. Annu. Rev. Microbiol. 69, 229–245. https://doi.org/10.1146/annurev-micro-091014-104432 (2015).
Chu, B. C., Peacock, R. S. & Vogel, H. J. Bioinformatic analysis of the TonB protein family. Biometals 20, 467–483. https://doi.org/10.1007/s10534-006-9049-4 (2007).
Samantarrai, D., Lakshman Sagar, A., Gudla, R. & Siddavattam, D. TonB-dependent transporters in Sphingomonads: unraveling their distribution and function in environmental adaptation. Microorganisms 8, 359. https://doi.org/10.3390/microorganisms8030359 (2020).
Kaczmarczyk, A., Vorholt, J. A. & Francez-Charlot, A. Markerless gene deletion system for Sphingomonads. Appl. Environ. Microbiol. 78, 3774–3777. https://doi.org/10.1128/AEM.07347-11 (2012).
Johnson, M. et al. NCBI BLAST: a better web interface. Nucleic Acids Res. 36, W5-9. https://doi.org/10.1093/nar/gkn201 (2008).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539. https://doi.org/10.1038/msb.2011.75 (2011).
Li, W. et al. The EMBL-EBI bioinformatics web and programmatic tools framework. Nucleic Acids Res. 43, W580-584. https://doi.org/10.1093/nar/gkv279 (2015).
Acknowledgements
We thank Yudai Higuchi and Aya Takeuchi for assistance with the construction of the SLG_06990, SLG_13630, and SLG_p-00340 mutants. This work was supported by JSPS KAKENHI Grant Numbers 15H04473, 19H02867, and 19J11312.
Author information
Authors and Affiliations
Contributions
E.M. supervised the project. M.F., N.K., and E.M. designed the study and wrote the manuscript. M.F. performed data analysis, western blot analysis, intracellular iron concentration measurement, purification of Fur1, EMSA, and CAS assay. M.F. and T.S. constructed plasmids and performed growth measurement and promoter assay. T.S., M.F., H.Y. and K.M. constructed mutants. K.T. performed RT-PCR analysis and helped to perform EMSA. T.S., K.T., H.Y., and K.M. helped to interpret the data and discussed the results. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fujita, M., Sakumoto, T., Tanatani, K. et al. Iron acquisition system of Sphingobium sp. strain SYK-6, a degrader of lignin-derived aromatic compounds. Sci Rep 10, 12177 (2020). https://doi.org/10.1038/s41598-020-68984-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-68984-2
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.