Abstract
Pericentromeric heterochromatin is generally composed of repetitive DNA forming a transcriptionally repressive environment. Dozens of genes were embedded into pericentromeric heterochromatin during evolution of Drosophilidae lineage while retaining activity. However, factors that contribute to insusceptibility of gene loci to transcriptional silencing remain unknown. Here, we find that the promoter region of genes that can be embedded in both euchromatin and heterochromatin exhibits a conserved structure throughout the Drosophila phylogeny and carries motifs for binding of certain chromatin remodeling factors, including insulator proteins. Using ChIP-seq data, we demonstrate that evolutionary gene relocation between euchromatin and pericentric heterochromatin occurred with preservation of sites of insulation of BEAF-32 in evolutionarily distant species, i.e. D. melanogaster and D. virilis. Moreover, promoters of virtually all protein-coding genes located in heterochromatin in D. melanogaster are enriched with insulator proteins BEAF-32, GAF and dCTCF. Applying RNA-seq of a BEAF-32 mutant, we show that the impairment of BEAF-32 function has a complex effect on gene expression in D. melanogaster, affecting even those genes that lack BEAF-32 association in their promoters. We propose that conserved intrinsic properties of genes, such as sites of insulation near the promoter regions, may contribute to adaptation of genes to the heterochromatic environment and, hence, facilitate the evolutionary relocation of genes loci between euchromatin and heterochromatin.
Similar content being viewed by others
Introduction
Eukaryotic genomes are packaged in chromatin consisting of DNA, RNA and associated proteins. Typically, chromatin can be divided into two basic forms, euchromatin and heterochromatin1. These types of chromatin are distinguished by several distinctive properties, including DNA sequence composition, specific histone modifications and binding proteins, nuclear and chromosomal localization, rate of DNA replication and frequency of meiotic recombination1,2. One of the major subtypes of heterochromatin in Drosophila is marked by heterochromatin protein 1 (HP1a) and di- or trimethylated H3K93,4. This subtype of heterochromatin covers large genomic segments primarily around centromeres and, in association with the protein POF (painting of fourth), the entire dot chromosome 4 in D. melanogaster3,4,5. Pericentric heterochromatin is mainly composed of repetitive sequences, including remnants of various transposable elements (TEs) and satellite DNAs6. A distinctive feature of heterochromatin is the ability to silence euchromatic genes placed within heterochromatic environment due to chromosomal inversions or transposition events, a phenomenon called position effect variegation (PEV)7,8,9,10,11,12. It is generally believed that transcriptional silencing of euchromatic genes in PEV is mediated by spreading of heterochromatin-associated marks HP1a and H3K9me3 across the gene loci transferred to heterochromatin8,10.
Despite the repressive environment, dozens of essential genes were identified in the pericentric heterochromatin of D. melanogaster13,14,15,16. Interestingly, genes embedded in pericentric heterochromatin in D. melanogaster may occupy distinct genomic regions, euchromatic and heterochromatic, in other Drosophila species17. For instance, two adjacent genes RpL15 and Dbp80 located in the pericentric region of chromosome 3L in D. melanogaster reside in a euchromatic region in D. pseudoobscura18. A similar pattern was demonstrated for genes light and Yeti located in pericentric regions in D. melanogaster, while in D. virilis they are found within euchromatin on the same chromosomal elements19,20. Recently, it was shown that most of the pericentric genes found at both arms of chromosome 2 of D. melanogaster are located in euchromatic region in the D. virilis genome21. However, although relocation of genes between euchromatin and heterochromatin during genome evolution is not unusual in the Drosophilidae lineage, the insusceptibility of heterochromatic genes to transcriptional silencing remains paradoxical and unexplained. It is still not clear whether pericentric gene loci have undergone adaptation to heterochromatic environment or originally had some intrinsic properties permitting local adaptation.
Chromatin insulator elements and associated proteins were originally defined by their ability to protect transgenes from PEV22,23,24,25. Numerous studies demonstrated that insulator proteins are responsible for a vast number of genomic functions, including stimulation of gene transcription, enhancer-blocking and barrier insulation partitioning of eukaryotic genomes into independently regulated domains26,27,28. Hence, one may hypothesize that gene loci capable of adaptation to heterochromatin probably share specific sites of insulation that ensure their expression in the repressive environment.
To address this issue, we initially investigated the molecular evolution of Myb and Ranbp16 genes which were relocated between euchromatic and heterochromatic environment in the Drosophilidae lineage. Further, we examined the regulatory factors that contribute to normal functioning of genes relocated into heterochromatic locations in distant Drosophila species, e.g. D. melanogaster and D. virilis. Myb is an essential gene encoding a transcription factor involved in transition from G2 to M phase of the cell cycle29,30. Ranbp16 encodes a RanGTP-binding protein belonging to the importin-β superfamily and mediates translocation of proteins into the nucleus. Both genes are located in euchromatic region of the D. melanogaster X-chromosome, while in other studied Drosophila species belonging to Sophophora and Drosophila subgenus they are found in genomic regions with a high density of repetitive DNA elements, suggesting their localization in heterochromatin. We found that regardless of the euchromatic or heterochromatic surroundings, the promoter region of Myb displays a high degree of sequence homology among Drosophila species studied so far. The conserved motifs in the promoter sequence of Myb serve as a binding site for the chromatin insulator protein BEAF-32 (Boundary element associated factor of 32 kDa) and transcriptional factor Dref (The DNA replication-related element (DRE) binding factor). Using ChIP-seq data, we demonstrate that the insulator protein BEAF-32 occupies promoters of the same genes which are located in contrasting chromatin types in D. melanogaster and D. virilis, denoting the boundary of the nucleosome-free region available for RNA polymerase II recruitment and the surrounding heterochromatin. Moreover, our analysis revealed that promoters of practically all protein-coding genes located in heterochromatin in D. melanogaster are enriched with insulator proteins BEAF-32, GAF (GAGA factor) and dCTCF (Drosophila homolog of CCCTC-binding factor). Exploring available RNA-seq data of mutant BEAF-32 function in Drosophila cells, we show that deficiency of BEAF-32 has a complex effect on expression of most genes in the genome, including heterochromatic ones. Overall, we propose that insulator proteins, in particular BEAF-32, are linked to expression of heterochromatic genes and may facilitate their normal function after evolutionary relocation into transcriptionally repressive genomic environment.
Results
Evolutionary relocation of Myb and Ranbp16 genes between euchromatin and heterochromatin in Drosophila phylogeny
In order to determine whether Myb and Ranbp16 gene locations have been rearranged on the evolutionary timescale, we first mapped these genes onto genomic scaffolds of Drosophila species separated by evolutionary distances from 5 to 40 million years31,32,33,34,35. These include species of the melanogaster group (D. melanogaster and D. yakuba) and the obscura group (D. persimilis and D. miranda), with both groups belonging to the Sophophora subgenus, along with the virilis group (D. virilis and D. novamexicana) and the repleta group (D. mojavensis and D. hydei) that belong to the Drosophila subgenus (Table S1). Next, we performed comparative analysis of Myb and Ranbp16 genes, as well as the intergenic regions between these genes. As indicated in Fig. 1, the coding sequences of Myb and Ranbp16 genes are highly homologous between the species studied (Fig. 1). However, the regions between Myb and Ranbp16 genes differ significantly, exhibiting sequence similarity only within the related groups. For instance, while Myb and Ranbp16 genes of D. melanogaster and D. yakuba are embedded within a large gene cluster, their orthologues in other Drosophila species reside in the genomic regions mostly occupied by repetitive sequences, including the introns of Ranbp16 gene (Fig. 1). Thus, Myb and Ranbp16 genes in the species belonging to the melanogaster group are located in euchromatin, in the region containing a large gene cluster. On the contrary, the localization of the studied genes in the environment packed with repetitive elements is typical for most other Drosophila species studied here including the virilis, obscura and repleta groups (Fig. 1).
Similarities and differences of genomic regions comprising Myb and Ranbp16 in Drosophila species. On the left—the phylogenetic tree indicating the relationships among studied species with estimated times of divergence according to Clark et al.31, Gao et al.32 for the obscura group, O’Grady et al.35 for the virilis group and Gibbs et al.33 for the repleta group. On the right—circular plot demonstrating similarity of Myb (red nodes) and Ranbp16 (blue nodes) and diversity of intergenic regions between these genes consisting of repeats (light blue nodes) and protein-coding genes (gray nodes) among the species of the Sophophora and Drosophila subgenera. Tracks of the plot indicate the region of comparison, coordinates of genes including Myb and Ranbp16, as well as the content of annotated repeats in the plotted region. Flanking regions of Myb and Ranbp16 (20 Kb upstream and downstream from the gene location) were used instead of intergenic regions, due to the long distance between these genes in D. miranda (> 17 Mb) and low scaffold contiguity around these genes for D. persimilis and D. hydei. These contigs borders are shown by lines (signed c.b., contig borders).
Next, we studied in more detail the genomic loci containing Myb and Ranbp16 genes, focusing on two evolutionarily distant species, D. melanogaster and D. virilis, separated by 40 million years of evolution31. The single copies of Myb and Ranbp16 genes map to the chromosome X of D. melanogaster at the cytogenetic areas 13F14 and 14A1 of salivary gland polytene chromosomes, respectively (Fig. 2a). These regions are significantly less enriched with heterochromatic marks H3K9me3 and HP1a than telomeric and pericentric regions of the chromosome and located at the distance more than 6 Mb from the heterochromatin–euchromatin border delineated in Riddle et al.4. Hence, these regions represent typical euchromatin in this species. As mentioned above, Myb and Ranbp16 genes in D. melanogaster are located at a distance ~ 80 Kb from each other within a large protein-coding gene cluster, which includes only a few repetitive sequences (Fig. 2a).
Analysis of genomic regions comprising Myb and Ranbp16 genes in D. melanogaster and D. virilis. (a) Genomic map of whole assembled chromosome X of D. melanogaster with mapped ChIP-seq reads of heterochromatic markers H3K9me3 and HP1a and the region depicting Myb and Ranbp16 gene location (marked with bold font). (b) DNA in situ hybridization of Myb gene probe to polytene chromosomes of D. virilis. Black arrow indicates the hybridization signal at the base of chromosome X of D. virilis. (c) Genomic map of scaffold_13050 of D. virilis and the proximal scaffold_12970 which is attributed to chromosome X of D. virilis according to Schaeffer et al.36 with mapped ChIP-seq reads of H3K9me3 and the region containing Myb and Ranbp16 genes (marked with bold font) on the larger scale. ChIP-seq reads are shown in RPMs (reads per million) normalized to input samples.
To confirm that Myb and Ranbp16 reside in a heterochromatic region in D. virilis, we performed DNA in situ hybridization on polytene chromosomes of this species using the unique sequence of Myb gene as a probe. In situ hybridization indicates that Myb is located at the base of chromosome X near the chromocenter in D. virilis (Fig. 2b). Mapping analysis of Myb and Ranbp16 genes on the genomic scaffolds of D. virilis reveals that both genes reside in one scaffold_13050 at a distance ~ 120 Kb from each other (Fig. 2c). In contrast to D. melanogaster, the region between these genes in D. virilis does not contain any other protein-coding genes and consists of remnants of TEs and other repeats diverged to varying degrees (Fig. S1). We used annotation of known genomic scaffolds of D. virilis made by Schaeffer et al.36 to assign the proximal scaffold of D. virilis genome r1.06 to the centromeric region of chromosome X. According to specific marker genes (para and tRNA:S7 located at the cytogenetic locus 19B), we retrieved scaffold_12970 and extended it with scaffold_13050 containing Myb and Ranbp16 genes, keeping some space unassembled between these scaffolds (Fig. 2c). Enrichment profile of H3K9me3 clearly indicates that the putative euchromatin-heterochromatin border lies in the proximal 1 Mb of scaffold_12970 (Fig. 2c). The whole scaffold_13050, including the region where Myb and Ranbp16 are localized, is heavily occupied by the H3K9me3 mark in comparison with the most contiguous fragment of scaffold_12970 (Fig. 2c).
To evaluate the possible impact of heterochromatic location on molecular evolution of Myb and Ranbp16, we examined whether their coding sequences underwent negative (purifying) or positive selection during Drosophila evolution. To this end, we first estimated the number of base substitutions per site in their coding sequences (Fig. S2a,b), and then calculated the ratio of non-synonymous to synonymous substitutions (dN/dS) (Fig. S2c,d) using eight aforementioned representatives of the Sophophora and Drosophila subgenera, as well as the sequences from Anopheles gambiae as an outgroup (Fig. S2). The results suggest that both Myb and Ranbp16 genes are under purifying (negative) selection (dN/dS < 0.2 in all pairs of comparison) when present either in heterochromatin or euchromatin.
Insulator protein BEAF-32 is enriched near the promoters of Myb and Ranbp16 genes in both D. melanogaster and D. virilis
To reveal specific properties of gene loci allowing the essential Myb and Ranbp16 genes to be actively transcribed in both euchromatin and heterochromatin, we studied the promoter region of Myb by searching for common motifs in Drosophila species. For this purpose, we expanded the list of analyzed species to 19 representatives of the Sophophora and 11 representatives of the Drosophila subgenera. As mentioned above, due to the absence of gene annotation for a range of studied species (D. miranda, D. guanche, D. subobscura and all virilis group species with the exception of D. virilis) the putative TSS was set as the first mapped nucleotide of 5′UTR of Myb gene of related species. Using the MEME suite37, we were able to identify three motifs that are present in the promoter of Myb gene in all analyzed species of Drosophila (Fig. 3a, b). Search through OnTheFly38 and REDfly v5.639,40 databases of known transcription factors and their binding sites indicates that one highest-scoring motif contains a potential binding site for insulator protein BEAF-32 and transcriptional factor Dref (1st motif, Fig. 3a). The two other motifs show limited similarity to the additional BEAF-32 motif and binding site of undescribed C2H2-type zinc finger transcription factor (Fig. 3a).
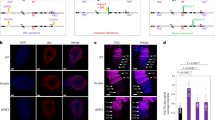
Promoter analysis of Myb and Ranbp16 in D. melanogaster and D. virilis. (a) Sequence logos of the common motifs among the studied Drosophila species with matches between motifs and binding sites of known transcription factors. Logos for binding sites of the factors are shown with the corresponding p- and q-values. (b) Distribution of common motifs and their orientation (shown by colored arrows) in the promoter of Myb. Prior to the analysis, orientation of all sequences was adjusted so that the transcription start site (TSS) would be on the right. The colors of motifs in A and B correspond to each other. Bootstrap consensus phylogenetic tree is given according to Clark et al.31, Gao et al.32, O’Grady et al.35 and Jezovit et al.34. (c) and (d) Enrichment of ChiP-seq reads within Myb and Ranbp16 genic loci in D. melanogaster, respectively. (e) The enrichment profile within the Myb and Ranpb16 genic loci and the intergenic regions between these genes in D. virilis. Mapped ChIP-seq reads without normalization (mapped reads), calculated areas of enrichment relative to the input data (peaks; P < 0.05) are shown for BEAF-32, Pol II, H3K9me3 and ATAC-seq data. Summits of the enriched reads are shown for BEAF-32 (peaks summits). RNA-seq reads were normalized to the sequence depth (RPM, reads per million).
To confirm that the insulator protein BEAF-32 binds to the promoter region of Myb gene, we used ChIP-seq data to profile BEAF-32 occupancy in the gene loci containing Myb and Ranbp16 genes in D. melanogaster and D. virilis. Additionally, we applied ChIP-seq to profile RNA polymerase II (Pol II) distribution in the promoters of Myb and Ranbp16 genes as well as ATAC-seq data to correlate BEAF-32 and Pol II enrichment with the nucleosome-free conformation of chromatin in the promoters of these genes, indicative of intensive transcription. Mapping of RNA-seq and H3K9me3 ChIP-seq data also provides valuable information regarding expression levels of these genes and the heterochromatic profile of the analyzed gene loci, respectively.
As seen in Fig. 3c, in D. melanogaster the locus containing Myb gene and the adjacent AlkB gene is highly enriched with BEAF-32, with the peak summit of mapped BEAF-32 reads in the promoter of the Myb gene (Fig. 3c). The enrichment of Pol II and mapped ATAC-seq reads reside downstream from the peak of BEAF-32 binding of the promoter region of Myb, probably indicating that BEAF-32 binds to DNA at the boundary of the Myb gene locus (Fig. 3c). Likewise, the promoter of Ranbp16 gene of D. melanogaster is enriched with BEAF-32, located slightly upstream from the Pol II binding site and open chromatin region (Fig. 3d).
A similar pattern of BEAF-32 occupancy in the promoters of heterochromatic Myb and Ranbp16 genes is observed in D. virilis (Fig. 3e). However, the enrichment profile of BEAF-32 upstream of the TSS of Ranbp16 gene in D. virilis and D. melanogaster has one difference which is worth mentioning. In contrast to D. melanogaster, where BEAF-32 is enriched in close proximity to the TSS of Ranbp16, the binding of BEAF-32 to DNA in D. virilis is observed only at a distance of ~ 6.5 Kb from the TSS of Ranbp16 (Fig. 3e). Notably, within this range, three copies of Jockey transposon are located. According to the data obtained by CAGE-seq (Cap Analysis Gene Expression), Ranbp16 gene in D. melanogaster has two TSS located at a distance of ~ 200 bp from each other, giving rise to slightly distinct transcripts in terms of the length of their 5′UTR (Fig. S3). Given that BEAF-32 is enriched within the upstream promoter of Ranbp16 in D. melanogaster, we assumed that in D. virilis the distance to the upstream promoter has been extended due to the transposon insertions that may be spliced in the course of transcription. However, we failed to observe more than a single TSS by 5′RACE analysis at the larval and imago stages as well as in the gonads of D. virilis, suggesting either loss of the second promoter or its extremely low efficacy. Interestingly, H3K9me3 is present within the gene bodies of Myb and Ranbp16 genes including exons in D. melanogaster (Fig. 3c,d). In turn, in D. virilis H3K9me3 is lacking in Myb and present only in introns of Ranbp16 that are enriched with repetitive elements (Fig. 3e).
These results indicate that the insulator protein BEAF-32 is enriched in the vicinity of the promoters of Myb gene embedded in distinct chromatin structures (euchromatic vs heterochromatic) in D. melanogaster and D. virilis. Notably, while binding of BEAF-32 is observed in the promoter of Ranbp16 gene in D. melanogaster, in the case of D. virilis the active binding site of BEAF-32 has moved upstream of the proximal observed TSS of Ranbp16 gene apparently due to transposon insertions.
Binding of BEAF-32 near the transcription start sites is preserved in the course of evolutionary relocation of gene loci between euchromatin and heterochromatin
Considering the above results, two important questions may be asked. First, do the BEAF-32 binding sites in the vicinity of promoter regions represent a peculiar property of Myb and Ranbp16 genes, or are they a common feature of heterochromatic genes? Second, do the BEAF-32 binding sites found in these promoters emerge in the course of adaptive evolution of genes transposed to heterochromatin? Alternatively, they might represent an ancestral feature which may contribute to the adaptation of genes relocated to the repressive environment without any deleterious impact on fitness.
To address these questions, we analyzed a representative set of more than 30 genes that reside in the pericentric heterochromatin of both arms of chromosome 2 in D. melanogaster, while in D. virilis the orthologs of these genes are located in different euchromatic regions of the same chromosomal elements (Table S2)21. Genes demonstrating the opposite scenario of relocation in these two species were also included in the analysis. Among them we considered as heterochromatic the genes Stim and Rrp47 that are located on the same scaffold_13050 as Myb and Ranbp16 of D. virilis, as well as the genes RpL15, Calr, Atg2 and CG40228 that are embedded in scaffold_12736 located near the chromocenter of D. virilis (Table S2)18. In contrast to D. virilis, most of these genes, with the exception of RpL15 and CG40228, are located in euchromatic regions of different chromosomal elements in D. melanogaster (Table S2). It is of note that localization of all of the selected genes was confirmed by in situ hybridization technique in this and other studies18,21. Also, due to the low annotation of 5′UTRs in D. virilis we have extended the analyzed region to 10 Kb upstream the annotated gene loci in order to identify the nearest binding of BEAF-32.
Using these sets of genes and specified parameters, we performed enrichment analysis of BEAF-32, Pol II, H3K9me3 and ATAC-seq reads upstream and downstream of the TSS in all of these genes (Fig. 4). Given the opposite chromatin state of the studied genes, we subdivided gene sets into two groups, group 1 includes genes that reside in heterochromatic regions in D. melanogaster but located in euchromatin in D. virilis (Fig. 4). Group 2 consists of genes located within euchromatin in D. melanogaster and heterochromatin in D. virilis. As indicated in Fig. 4, the insulator protein BEAF-32 is highly enriched in the vicinity of TSS of virtually all considered heterochromatic genes in both D. melanogaster (group 1) and D. virilis (group 2) (Fig. 4). However, binding of BEAF-32 in the promoters of heterochromatic genes does not necessarily coincide with the TSS of their euchromatic orthologs (group 1 of D. virilis and group 2 of D. melanogaster) (Fig. 4, Table S2). Overall, we observed 27/38 shared genes comprising both groups (euchromatic and heterochromatic) of D. virilis and D. melanogaster which exhibit similar BEAF-32 binding in their promoters (Fig. 4, Table S2). The binding area of BEAF-32 is in strong association with the enrichment profile of Pol II and ATAC-seq reads in the proximity of TSS for most of the studied genes (Fig. 4). Importantly, the mean enrichment value of BEAF-32 in the vicinity of the promoter does not differ significantly between heterochromatic genes (median: 8) and euchromatic ones (median: 9.25) with P = 0.4 (Mann–Whitney U test) indicating that the enrichment of BEAF-32 does not depend on the local chromatin environment.
Enrichment of BEAF-32 near the TSS of genes repositioned between euchromatin and heterochromatin during Drosophila evolution. (a) and (b) show the enrichment profiles of ChIP-seq reads of genic loci of D. melanogaster and D. virilis, respectively. Note that the analyzed area for BEAF-32 enrichment includes 10 Kb upstream and 2 Kb downstream from TSS, while for RNA Pol II, H3K9me3 and ATAC-seq reads area includes 2 Kb both upstream and downstream from the TSS. Color codes below the heatmaps indicate fold enrichment (treat vs input).
Motif CGATA is a hallmark of BEAF-32 genomic binding sites41,42. However, recently it was shown that even though BEAF-32 can bind DNA directly, a large subset of BEAF-32 peaks that does not share BEAF-32 consensus motif and apparently mediates functional long-range contacts among distinct insulator sites through indirect binding with a co-factor CP19043. Given this, we have analyzed group 1 and group 2 genes and observed that 27/32 of heterochromatic genes and 4/6 of euchromatic genes in D. melanogaster comprising group 1 and 2, respectively, share motif CGATA in their promoters (within 200 nt upstream from TSS) demonstrating the direct binding of BEAF-32, while others do not and thus exhibit the indirect binding of BEAF-32 (Table S2). In D. virilis we have defined 15/30 of euchromatic genes (group 1) and 8/8 of heterochromatic genes (group 2) exhibiting a direct binding of BEAF-32. In agreement with the previous findings43, direct peaks exhibit greater enrichment than indirect ones (median of direct peaks: 11, median of indirect peaks: 5, P < 0.01 by Mann–Whitney U test).
Taken together, these results indicate that BEAF-32 binds predominantly directly the promoters of genes that were juxtaposed with heterochromatin during the evolution of the genus Drosophila. However, while the promoters of heterochromatic genes of D. melanogaster are occupied by BEAF-32, their euchromatic orthologs in D. virilis have partially lost their BEAF-32 binding sites in the vicinity of promoter regions.
Promoters of most heterochromatic genes are occupied by insulator proteins BEAF-32, GAF and dCTCF in D. melanogaster
It is known that among the described insulator proteins, BEAF-32, GAF and dCTCF are widespread upstream of the TSS of actively transcribed genes in the genome of D. melanogaster44,45,46,47,48. Indeed, distribution of BEAF-32, GAF and dCTCF has a similar pattern in terms of genomic features (Figs. S4–S6). Most of the binding sites of these insulator proteins are located predominantly within 1 Kb upstream the TSS (BEAF-32 ~ 64% of all sites, GAF ~ 66% and dCTCF ~ 60%). Notably, these proteins also bind intronic (BEAF-32 ~ 7% of all sites, GAF ~ 7% and dCTCF ~ 8%) and intergenic (BEAF-32 ~ 10% of all sites, GAF ~ 6% and dCTCF ~ 17%) regions in D. melanogaster genome (Figs. S4–S6).
In order to elucidate whether the promoters of all heterochromatic genes are occupied by insulator proteins, we analyzed the enrichment profile of BEAF-32, GAF and dCTCF within 2 Kb upstream and downstream of TSS of all heterochromatic genes of D. melanogaster (Fig. 5a). For this purpose, we sorted pericentric protein-coding genes located downstream of the euchromatin-heterochromatin border defined by a gradual increase of H3K9me3 and HP1a enrichment in D. melanogaster. We have also included genes from the dot chromosome 4 considering it as entirely heterochromatic5. Next, we selected only those genes that show significant enrichment of RNA Pol II and expression level not less than 10 RPM (reads per million), according to ChIP-seq and RNA-seq data, respectively. A final list of heterochromatic protein-coding genes includes 135 genes located at different chromosomes of D. melanogaster (Table S3).
The enrichment profile of insulator proteins around the promoter regions of heterochromatic genes in D. melanogaster. (a) The enrichment profiles of ChIP-seq reads for BEAF-32, GAF, dCTCF, RNA Pol II, H3K9me3 and ATAC-seq reads on heterochromatic genes of D. melanogaster. The analyzed area includes 2 Kb upstream and 2 Kb downstream from TSS. Color codes below the heatmaps indicate fold enrichment (treat vs input). (b) Venn diagram indicates the number of gene loci enriched with all three, two and only one studied insulator protein BEAF-32, GAF and dCTCF. (c) Diagram shows the percentage of genes whose promoters are enriched with BEAF-32, GAF or dCTCF. Euchromatic and heterochromatic genes are indicated separately according to the number of genes comprising these regions.
The performed analysis of the enrichment of insulator proteins shows that although BEAF-32 occupies the majority of gene promoters it is not present ubiquitously (Fig. 5a). However, the lack of BEAF-32 near the promoters of heterochromatic genes is usually compensated by the presence other insulator proteins GAF and dCTCF, keeping the promoters of > 93% (126/135) heterochromatic genes enriched with insulator proteins (Table S3). Importantly, the binding area of insulator proteins is strongly correlated with the area of decreased levels of methylated H3K9 and nucleosome-free regions defined by ATAC-seq, which can be seen at a distance of 2 Kb and 1 Kb around TSS (Fig. S7). Notably, all three studied insulator proteins are present together in more than a half of the promoters of heterochromatic genes (78/135 genes, range of the promoter was defined as 200 nt upstream the TSS) (Fig. 5b). A lot of genes are occupied by a pair of the insulator proteins, e. g. BEAF-32 and dCTCF (29 genes) or BEAF-32 and GAF (5 genes) (Fig. 5b). Among the number of BEAF-32 binding sites, ~ 84% (104/121 genes) share CGATA motif indicating the predominantly direct binding of this insulator proteins to the promoters of heterochromatic genes. Consistent with the previous observation, the enrichment of BEAF-32 in direct peaks are ~ 2.5 fold greater than in indirect ones (median of direct peaks: 23, median of indirect peaks: 11, P < 0.01, Mann–Whitney U test). Notably, there is a small subset (9 of 135) of heterochromatic genes that lack the studied insulator proteins in their promoters. Part of them (CG17450, CG33502, CG32857, CG32820, CG32819, and CG32500) are located within a gene cluster which includes eight closely spaced genes and covers 30 Kb of the genome (Fig. S8). Such gene cluster is not typical for the pericentric heterochromatin where genes are widely spaced by repetitive sequences. Interestingly, the promoters of the flanking genes (GCS2α and DIP1) of this cluster are occupied by the studied insulator proteins (Fig. S8). It can be speculated that such a placement could allow the formation of a loop between the flanking genes to form an actively transcribed region, which eliminates the need to keep insulators in the promoters of each gene included in this structure.
It is evident that insulator proteins occupy a higher number of genes in heterochromatin than euchromatin (Fig. 5c). Specifically, BEAF-32 is the most prominent insulator that is enriched in the promoters of heterochromatic genes (> 90% (122/135) of all heterochromatic genes), while GAF is prevalent in euchromatic genes (49% (6,717/13,709) of all euchromatic genes) (Fig. 5c).
Taken together, these data show that insulator proteins BEAF-32, GAF and dCTCF, solely or in combination with each other, are present in the promoters of virtually all heterochromatic genes of D. melanogaster studied so far.
BEAF-32 and Dref binding overlaps in the promoters of a subset of heterochromatic genes implicated in gene transcription in D. melanogaster
Dref is described as a master regulator of cell proliferaton in Drosophila49. The DNA recognition motif for BEAF-32 (CGATA) is contained within Dref binding sequence (TATCGATA), and the binding of Dref to DNA has been shown to antagonize the binding of BEAF-32 in vitro50. In order to demonstrate to what extent BEAF-32 binding overlaps with Dref in the promoters of heterochromatic genes, we have analyzed the enrichment of these proteins in the promoters of previously defined heterochromatic genes of D. melanogaster using ChIP-seq data from Kc167 cell line (see “Methods”). Notably, genes embedded into heterochromatin in D. melanogaster are implicated in a variety of biological processes that mediate many aspects of cellular function, including protein phosphorylation, transcription, translation, development and recombination processes (Fig. 6a).
Enrichment of BEAF-32 and Dref in the promoters of heterochromatic genes in D. melanogaster. (a) Gene Ontology (GO) analysis by biological processes of all heterochromatic genes (P < 0.05 of presented GO terms). Green color bars denote the processes in which genes occupied by BEAF-32 are involved, red bars: genes enriched with BEAF-32 and Dref. (b) Enrichment plots of BEAF-32 and Dref in heterochromatic genes of D. melanogaster. Plots for heterochromatic genes are given separately for each chromosome. (c) Venn diagram indicates the number of heterochromatic genes whose promoter regions are occupied solely by BEAF-32 or Dref as well as overlapped genes.
Enrichment analysis shows that BEAF-32 binds DNA within the promoters of 88% (119/135) of heterochromatic genes in Kc167 cell line (Table S4). Among them, 102 genes exhibit CGATA motif within their promoter regions indicating the predominantly direct binding of BEAF-32 to the heterochromatic gene loci as was indicated previously (Table S4, Fig. 5). In contrast to BEAF-32, the enrichment of Dref is significantly lower in the promoters of heterochromatic genes of D. melanogaster (Fig. 6b). Thus, binding of Dref in the promoters of heterochromatic genes overlaps with BEAF-32 for 46 genes among 119 genes occupied by BEAF-32 (Fig. 6c, Table S4). Most of these genes (e.g. pan, hcf, Sox102F) are implicated in regulation of transcription and developmental processes (red bar in Fig. 6a). Moreover, in contrast to BEAF-32, the binding of Dref does not occur in the absence of BEAF-32 (Fig. 6c). Importantly, a half (23/46) of genes those promoters are occupied by Dref do not share the canonical Dref motif TATCGATA (Table S4) indicating that Dref binding may be weaker in the promoters of heterochromatic genes. Indeed, the overall enrichment of BEAF-32 is 3–fourfold higher than Dref (median of BEAF-32 peaks: 21, median of Dref peaks: 5, P < 0.01 by Mann–Whitney U test). Notably, there are 16 genes involved in developmental processes occupied neither by BEAF-32 nor by Dref in Kc167 cell line (Table S4).
These data indicate that Dref is present in the promoters of heterochromatic genes together with BEAF-32, suggesting that BEAF-32 may be required for Dref binding to the promoters of these genes.
Disruption of BEAF-32 has a complex effect on genome expression affecting even those genes that lack this insulator protein in their promoter regions
As seen from the results above, BEAF-32 is the most prevalent insulator protein in promoters of genes in the pericentric heterochromatin in D. melanogaster. Drosophila BEAF-32 gene encodes two isoforms, BEAF-32A and BEAF-32B. Both proteins are essential, but BEAF-32B alone is sufficient for the viability of flies51. Homozygous mutation of BEAF32 is characterized by disorders in oogenesis, resulting in drastically reduced fertility of females51. To find out to what degree BEAF-32 contributes to expression of genes, we analyzed RNA-seq data of stably transfected Drosophila Schneider S2 cells expressing mutant BEAF-32 in the absence of endogenous protein43.
The analyzed data indicate that impairment of BEAF-32 strongly affects gene expression (767 differentially expressed genes (DEG), P ≤ 0.05; Fig. 7a). Notably, the disruption of BEAF-32 function has a complex effect on transcription, which includes not only downregulation but also upregulation of gene expression levels (Fig. 7a). Given that insulator proteins BEAF-32, dCTCF and GAF may overlap in promoters of genes, we sorted DEG according to the association of these insulators with DEG. We found that among 580 DEG the largest groups comprise all three insulator proteins (180 DEG), and 162 DEG contain only GAF (Fig, 7b), whereas BEAF-32-associted DEG include only 42 genes. Furthermore, 64 and 23 DEG exhibit overlapping with GAF and dCTCF, respectively (Fig. 7b). Interestingly, among differentially expressed genes 187 genes (P ≤ 0.05) are not occupied by any of the three proteins (BEAF-32, dCTCF and GAF) (Fig. 7b).
Mutant BEAF-32 strongly affects gene expression both in euchromatin and heterochromatin in D. melanogaster. (a) Volcano plot demonstrating genes with expression changes in Drosophila S2 cells expressing mutant BEAF-32 in comparison with control cells (767 genes with P ≤ 0.05). Genes with expression level < 1 Log10 CPM (counts per million) were discarded. (b) Pie charts show the number of differentially expressed genes whose promoters are enriched with one, two or simultaneously occupied by all three insulator proteins BEAF-32, dCTCF and GAF (580 differentially expressed genes, P ≤ 0.05). Genes whose promoters are free of these insulator proteins are also shown (187 differentially expressed genes, P ≤ 0.05). (c) Box plots demonstrate trends of gene expression changes upon the impairment of BEAF-32 function. All differentially expressed genes (P ≤ 0.05) are divided into 4 groups according to the presence of insulator protein in their promoter regions—group 1 (one of three proteins), group 2 (combination of two of three proteins), group 3 (combination of all three insulator proteins) and group 4 (genes that are not occupied by these proteins). *Indicates P ≤ 0.05 (Fisher exact test). d) Bar plots demonstrate overall expression changes in S2 cells expressing mutant BEAF-32 separately for euchromatic genes (left) and heterochromatic genes (right).
Next, we estimated trends of gene expression changes analyzing separately DEG showing association with BEAF-32, dCTCF, and GAF or a combination of these proteins. Unexpectedly, BEAF-32 associated DEG tend to be upregulated upon disruption of BEAF-32 function (left box in group 1, Fig. 7c). On the other hand, downregulation is observed for GAF-associated genes (right box in group 1, Fig. 7c) and for the genes that are not occupied by any of the studied insulator proteins (group 4, Fig. 7c). Other groups of genes, in particular the ones associated with dCTCF only (middle box in group 1), a pair of insulator proteins in various combinations (group 2) and DEG associated with all three insulator proteins (group 3) exhibit a complex pattern of gene expression, with upregulated and downregulated changes (Fig. 7c). Moreover, the upregulated expression profile of BEAF-32 and dCTCF-associated genes are significantly different from GAF-associated DEG (p ≤ 0.05). The same applies to DEG that are associated with all three proteins in comparison with genes that are not occupied with any of them (group 3 vs group 4, P ≤ 0.05).
Overall, effect of mutation disrupting BEAF-32 is more pronounced for euchromatic genes than heterochromatic ones (Fig. 7d). Among euchromatic genes more than 25% (P ≤ 0.05) were affected, while only 15% of heterochromatic genes show expression changes higher than 1.5 fold (P ≤ 0.05, Fig. 7d), suggesting that expression of heterochromatic genes is more robust to the disruption of the function of only one of the insulator proteins.
Therefore, these data show that the impairment of BEAF-32 strongly affects expression of both euchromatic and heterochromatic genes, including those that lack this insulator protein in their promoter regions.
Discussion
During speciation, the genomes of Drosophila species underwent multiple chromosome rearrangements that disrupt gene order, modifying or preserving gene function17,52. In this study, we show that in the course of evolution of Drosophila species the essential Myb and Ranbp16 genes have been relocated into different chromatin types, i.e. euchromatin or heterochromatin. Despite the contrasting chromatin structure and local repressive environment of heterochromatic regions enriched with repetitive DNA, both genes were shown to be under purifying selection due to their highly conserved and vital function.
According to studies of PEV, genes that are juxtaposed with heterochromatin by chromosomal rearrangements or transposition events exhibit a variegating phenotype resulting in silencing of genes due to heterochromatin environment8. Given that, how might the evolutionary relocation of essential genes into the pericentric heterochromatin be explained? What determines the insusceptibility of regulatory regions of heterochromatic genes to the repressive surroundings? Considering the peculiarities of heterochromatic genes, such as accumulation of TEs within their introns, one may suggest that heterochromatic genes have evolved to adapt to the heterochromatic environment and became dependent on heterochromatin specific proteins4,53,54,55,56. Previously, it was shown that functioning of heterochromatic genes depends on repetitive environment and heterochromatin factors, such as HP1a, to facilitate their expression and probably long-distance interactions between enhancers and promoters11,57,58,59. Moreover, evolutionary relocation preferentially occurred only with genes exhibiting close association with HP1a, suggesting that HP1a binding to these genes existed in the progenitor21. An analysis conducted by Yasuhara et al.20 demonstrated that promoters of heterochromatic genes have not undergone major alterations after relocation into the repetitive environment of heterochromatin, which excludes the existence of heterochromatin-specific promoters. Together, these observations allow one to suggest that certain gene loci were probably pre-adapted or had acquired adaptive properties in the ancestral species for relocation between euchromatin to heterochromatin in the course of evolution. If this is the case, gene loci relocated to heterochromatin probably should retain the transcriptionally active euchromatin-like structure of chromatin capable of efficient transcription in the heterochromatin. Indeed, the proximal regulatory regions of heterochromatic genes are not occupied by the heterochromatic mark H3K9me3, forming a nucleosome-free binding platform for transcriptional factors and RNA polymerase II. Recently, a remarkable peculiarity of HP1a binding at several gene loci has been described, whereby HP1a can be recruited to gene promoters independently of H3K9 methylation60. Along these lines, the observed binding of BEAF-32 in the vicinity of gene promoters which underwent relocation between euchromatin and heterochromatin in evolutionarily distant species of Drosophila is very intriguing. In cooperation with HP1a, the presence of BEAF-32 and probably other insulator proteins such as dCTCF and GAF at gene promoters probably contributed to the formation of “proto-heterochromatic” gene loci in the ancestral species of Drosophila and thus facilitated their normal functioning in the heterochromatic environment. Hence, one may hypothesize that insulator proteins may block the spreading of heterochromatin to the promoter regions of heterochromatic genes, while HP1a maintains a proper chromatin structure at and around such gene loci that might be controlled epigenetically.
Originally, insulator proteins were defined as regulators of interaction between enhancers and promoters able to block silencing effect of PEV22,23,24,25. A growing body of evidence suggests that insulator proteins exercise diverse roles, including barrier function, and mediate short and long-distance chromosomal contacts at the genome-wide scale45,61,62,63,64. Our enrichment analysis of ChIP-seq data indicated that the insulator protein BEAF-32 is enriched upstream of the TSS of heterochromatic genes in D. melanogaster and D. virilis, probably demarcating the euchromatin-heterochromatin border between the promoter and the surrounding heterochromatin. Furthermore, using the same set of genes that reside in different types of chromatin in D. melanogaster and D. virilis, we show that BEAF-32 binding is predominantly preserved in the promoter regions of heterochromatic genes during evolution of different Drosophila species, suggesting that BEAF-32 binding is an ancestral property of these genes, rather than adaptation to the heterochromatic environment. However, the binding of BEAF-32 in the vicinity of TSS is not always preserved in euchromatic genes in comparison to their heterochromatic orthologs in distant species (Fig. 4). One may suggest that due to the high density of genes in euchromatic regions of the genome and lower occupancy of repressive histone mark H3K9me3, not every gene requires barrier insulation of BEAF-32 for functioning. In contrast to euchromatic regions, genes in the pericentric heterochromatin are largely dispersed, separated by numerous TEs and other repeats in intergenic regions enriched with H3K9me3 and HP1a. In such an environment barrier function of insulator proteins might have become one of the major determinants that contribute to the proper function of heterochromatic genes. Importantly, the binding sites of BEAF-32 near the promoters could be indirect even in the heterochromatic regions and result in weaker binding of insulator protein. As was shown previously, insulator proteins BEAF-32 and dCTCF may facilitate long-range contacts of the chromatin through CP19043. However, how this machinery works remains an unresolved question. Obviously the interactions of cis-regulatory elements and trans-acting factors involved are more complex and include a variety of chromatin-remodeling factors and insulators acting to facilitate continuous gene expression and chromosomal architecture of heterochromatic gene loci.
Along with BEAF-32, insulator proteins dCTCF and GAF are also enriched at the TSS of heterochromatic genes. Moreover, a combination of BEAF-32, GAF and dCTCF covers virtually all promoters of protein-coding genes located in the pericentric chromosome regions and heterochromatic dot chromosome 4 in D. melanogaster, suggesting that proper functioning of heterochromatic gene loci requires insulators (Fig. 5). However, there is a subset of genes whose promoters lack the binding site of these three insulator proteins suggesting that their functioning is mediated by other insulator proteins such as Pita that also belongs to a class of insulator proteins that preferentially bind to promoters near the TSS65. Alternatively, the architecture of certain gene loci especially of gene clusters probably allows functioning without these regulatory elements resulting in their eventual loss in the promoter regions during evolution.
The insulator proteins GAF and especially dCTCF have plenty of overlapping binding sites with BEAF-32 and Dref in the Drosophila genome66. Moreover, Dref co-localizes at the same genomic sites as BEAF-32 and other insulator proteins and is enriched at the boundaries of topologically associated domains (TAD)66. To this end, we observed that the promoters of heterochromatic genes do not appear to have Dref without binding BEAF-32 (Fig. 6). Notably, cis-acting elements that exercise the transcriptional control of genes by Dref, as well as protein sequence of Dref, are conserved between such evolutionarily distant species as D. melanogaster and D. virilis 46,67. Together, these data probably suggest that Dref function in heterochromatin is mediated by and might depend on insulator proteins on an evolutionary timescale.
It is of note that a direct impact of insulator presence on gene expression has been established for the D. melanogaster GAGA factor (GAF) that resides in the hsp70 promoter. GAF mediates the recruitment of chromatin remodeling factors, including SWI/SNF, the CHD, and the ISWI family complexes, that ensure formation of nucleosome-free region in the hsp70 promoter68,69,70. Knockout mutation showed that BEAF-32 is important for both oogenesis and development51. Furthermore, it was shown that PEV of the wm4h allele and different y transgenes was enhanced by the BEAF-32 KO, suggesting that BEAF-32 function affects chromatin structure or dynamics51. Other studies of BEAF-32 demonstrated that most BEAF-associated genes are transcriptionally active or even highly expressed and are associated with negative elongation factor Nelf that stimulates transcription levels by inhibiting promoter-proximal nucleosome assembly61,71. This provides evidence that BEAF-32 facilitates high levels of gene expression. Indeed, the mutation of BEAF-32 which abrogates BEAF-32 function results in misregulation of hundreds of genes43. Surprisingly, most of the affected genes show an association predominantly with GAF (162 genes) but not with BEAF-32 (42 genes) (Fig. 7). Moreover, the downregulation of gene expression was observed mostly for genes that lack direct association with BEAF-32 protein. According to this complex pattern of gene expression, one may suggest that deficiency of BEAF-32 disrupts chromosomal contacts, resulting in misregulation of genome-wide expression.
While it is clear that further studies are needed to elucidate all the factors required for normal gene functioning in the heterochromatic surroundings, our results suggest a possible evolutionary path that can be utilized by heterochromatic genes to maintain their expression in the repressive environment.
Conclusions
Heterochromatin in Drosophila is generally associated with transcriptional silencing. Nevertheless, dozens of essential genes have been identified in the pericentric heterochromatin of D. melanogaster and other species. In this study, we investigated the molecular evolution of the essential genes that were relocated between euchromatin and pericentric heterochromatin in the phylogeny of Drosophila. By surveying factors necessary for normal functioning of genes relocated into heterochromatin in distant Drosophila species, e.g. D. melanogaster and D. virilis, we conclude that certain insulator proteins (i.e. BEAF-32) may contribute to the successful adaptation of genes to the pericentric heterochromatin by facilitating normal gene expression in the repressive surrounding.
Methods
Drosophila genomes and sequence analyses
Drosophila genomes and gene sequences for comparative analysis were extracted from FlyBase and NCBI databases. Sequences of genomic regions containing Myb gene of virilis group species (D. lacicola, D. littoralis, D. borealis, D. flavomontana, D. lummei, D. ezoana) were fetched from unpublished data of Dr. Venera Tyukmaeva and Prof. Michael Ritchie from the University of St. Andrews, UK (personal communication). Orthologous genes were retrieved from OrthoDB v9.172. In the case of absence of gene annotation (e.g. for D. guanche, D. subobscura, most of the virilis group species), orthologs were retrieved with TblastN73 using protein sequence of the most closely related species (i.e. D. obscura for D. subobscura and D. guanche, D. virilis for D. lacicola and other virilis species) as queries. All the query subjects mapped on the same DNA strand adjacent to each other with E-value > e-80 were considered as valid. To perform reciprocal BLAST, obtained hits were aligned back to the original genome. Aligned hits were considered as the best reciprocal hits and used for reconstruction of coding sequences. Sequences between mapped subjects were considered as introns. Putative transcriptional start sites (TSS) of poorly annotated genes were identified with blastN74 using 1st exon sequence of related species. Blast results with E-value > e-60 and adjacent to annotated coding sequence at a distance not exceeding 600 nt were considered as true and the 1st mapped nucleotide as TSS. All essential information, including genes IDs, genomes IDs, and genomic coordinates of Myb and Ranbp16 in all studied species, is listed in Table S1. Orthologous sequences of Myb and Ranbp16 genes of Anopheles gambiae were extracted from VectorBase (https://www.vectorbase.org/) by the numbers AGAP008160 – Myb and AGAP004535 – Ranbp16. Protein sequences of Myb and Ranbp16 (also known as Xpo7) of mouse and human were extracted from UniProt (https://www.uniprot.org/). Protein motifs were scanned using the PROSITE database and methodology75,76.
Estimation of repeat content in intergenic regions and within studied genes was performed using RepeatMasker (https://www.repeatmasker.org) and computationally predicted libraries of TEs generated with ReAS77 that are available in FlyBase (ftp://ftp.flybase.net/genomes/aaa/transposable_elements/ReAS/v2/consensus_fasta/). For repeat masking of D. miranda genome, we used consensus sequences of TEs of D. pseudoobscura and D. persimilis, and for D. hydei we applied the library of D. mojavensis. TEs were classified using RepeatClassifier implemented in RepeatModeler software78.
Multiple sequence alignment was performed with ClustalW79 and Clustal Omega80 programs (https://www.ebi.ac.uk/Tools/msa/) for nucleotide and amino acid alignments, respectively. Multiple protein alignments were visualized with Jalview81.
Circular plot was made using Circos visualization tool82. Flanking regions of Myb and Ranbp16 (20 Kb upstream and downstream from the gene location) were used instead of intergenic regions, due to the long distance between these genes in D. miranda (> 17 Mbp) and low scaffold contiguity around these genes for D. persimilis and D. hydei.
ChIP-seq, RNA-seq and ATAC-seq analyses
Raw data of genome binding/occupancy (ChIP-seq), transcriptome (RNA-seq) and nucleosome (ATAC-seq) profiling were obtained from GEO database and used in the analyses. They include: GSE59965—contains data for D. virilis including RNA-seq, ChIP-seq of H3K9me3 and RNA polymerase II performed using commercially available anti-H3K9me3 (ab8898, Abcam) and anti-RNA Pol II (ab5408, Abcam) antibodies; GSE35648—contains data for both D. melanogaster and D. virilis including ChIP of BEAF-32 performed using antibodies generated against amino acids 1–83 of the major highly conserved isoform BEAF-32B in D. melanogaster42; GSE43829—contains RNA-seq as well as ChIP-seq of H3K9me3 and RNA polymerase II for D. melanogaster performed using aforementioned commercially available antibodies ab8898 and ab5408; GSE56347—includes ChIP-seq of HP1a for D. melanogaster performed with polyclonal anti-HP1 (PRB-291C, Covance innovative); GSE102439—includes ATAC-seq data for D. melanogaster and D. virilis; GSE62904 – ChIP-seq for Dref and BEAF-32 in Kc167 cells of D. melanogaster; finally, GSE85404 and GSE70632—contains ChIP-seq data of dCTCF and GAF for D. melanogaster. Comparative analysis of each deep sequencing data was conducted on the same type of tissue of D. melanogaster and D. virilis.
For analysis of sequence data, we used genome sequence and annotations released in FlyBase, D. melanogaster r.6.19 and D. virilis r1.06. Prior to mapping, all libraries were subjected to adapter clipping, filtering by length (> 20 nt) and quality (80% of nt must have at least 20 Phred quality) using TrimGalore (https://github.com/FelixKrueger/TrimGalore). Then, sequences were aligned to corresponding genomes using Bowtie83 with the following settings: “–best –strata -m 1”, retaining only uniquely mapped reads. Output sequence alignment map (SAM) files were converted to binary (BAM) format using SAMtools84. Aligned reads normalized to input samples in wig format were visualized using the Integrative Genome Viewer (IGV)85.
Peak calling was performed using MACS software86 with the recommended parameters for narrow (PolII, BEAF-32, dCTCF, GAF) and broad peak calling (H3K9me3, HP1a) as well as normalization on input chromatin controls. Enrichment analysis was performed using pipelines implemented in deepTools package87 with the parameters including ignoring of duplicates.
ChIP-indirect and direct peaks of BEAF32 were identified as described in Liang et al.43. Briefly, identified peaks from MACS that overlap with promoter regions (200 nt upstream TSS) were scanned for DNA-binding motif of BEAF-32 (extracted from JASPAR database88) using TFBSTools package89. If motif exists, binding is considered as direct, and in the absence of appropriate DNA motif in peak, binding is considered indirect.
For ATAC-seq analysis, reads that mapped on mitochondrial genomes were discarded, and peak calling was performed using Genrich (https://github.com/jsh58/Genrich) with the following settings: “-j -y -r -d 50”, including removal of PCR duplicates.
The analysis of enriched gene ontology (GO) terms was performed using DAVIDWebService package for R with a P = 0.05 (Fisher exact test)90,91.
Gene expression analysis of mutant BEAF-32 by RNA-seq
Raw data for stably transfected Drosophila Schneider S2 cell line expressing synthetic WT/mutant BEAF-32 in the absence of endogenous BEAF-32 were fetched from NCBI GEO (GSE52887)43. Processing of data included the adapter, length and quality trimming by Trimmomatic, mapping of reads to the genome (release GRCm38) by STAR aligner, counting the overlap of reads with genes by featureCounts, implemented in PPLine script92,93,94,95. Differential gene expression analysis was performed with the edgeR package using a Fisher exact test between experimental groups96. The genes with expression level ≥ 1 Log10 CPM (counts per million) and P ≤ 0.05 were considered as differentially expressed. Three biological replicates were analyzed for each sample.
Differential expression analysis, data visualization, and GSEA (Gene Set Enrichment Analysis) were performed using R project for statistical computing91. Visualization of experimental data was made with ggplot2 and GOplot R packages97.
Promoter analysis
Because of insertion of DAIBAM MITE at a distance of 92 bp upstream from TSS of Myb in D. virilis, the promoter regions of Myb in studied species were shortened to 100 bp. After sequence extraction, promoter regions of Myb and Ranbp16 were searched for common motifs using MEME98 and identification of matches to known transcription factors was performed by Tomtom99 using OnTheFly38 and REDfly v5.639,40 databases implemented in MEME Suite 5.0.537.
Sequence evolution and testing for selection
Analysis of nucleotide substitutions per site was conducted in MEGA X100 using the Tamura-Nei model101. Rate variation among sites was modeled with a gamma distribution (shape parameter = 1). All positions containing gaps and missing data were eliminated (complete deletion option).
Ratio of nonsynonymous and synonymous substitutions (dN/dS) was estimated using PAL2NAL software102 by converting multiple sequence alignment of proteins and the corresponding nucleotide sequences into a codon alignment, and the calculation of synonymous (dS) and non-synonymous (dN) substitution rates using codeml program implemented in PAML package103.
Cytology and DNA in situ hybridization
D. virilis larvae were grown at 180C on standard medium supplemented with live yeast solution for 2 days before dissection. Salivary glands from 3rd instar larvae were dissected in 45% acetic acid and squashed. DNA probes corresponding to D. virilis Myb (Dvir\GJ18431; FlyBase ID: FBgn0205590) were prepared by PCR using gene-specific primers (Forward_ GCAAGTGCGAGCACTGAAAA; Reverse_TGCATACTGAGGTGTGCCAG). Then, DNA probe was biotinylated by nick translation using Biotin-14-dATP (Thermo Fisher Scientific, USA) as described in104. Localization of the probe was made using the cytological map of D. virilis chromosomes105. Images were obtained by binocular microscope Nikon Alphaphot-2 YS2 (Japan).
RNA isolation, RT-PCR and 5′-RACE analysis
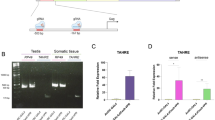
Total RNA from 3rd instar larvae, adult females and gonads was isolated using Extract RNA reagent (Evrogen, Russia). Synthesis of the first strand of cDNA from total RNA and subsequent amplification of regions of interest were performed using MINT cDNA kit (Evrogen, Russia) following manufacturer’s instructions. For specific rapid amplification of cDNA 5′-end (5′-RACE) analysis, we applied two outward primers (primer1 5′-AGTAGTTGTGCGTAGCTGGA-3′; primer2 5′-GCTGCTTGCACAATGTTTCTA-3′) corresponding to the annotated 5′-fragment of D. virilis Ranbp16 gene (Dvir\GJ18467; FlyBase ID: FBgn0205626). PCR reaction was conducted using Encyclo DNA polymerase (Evrogen, Russia). The resulting PCR fragments were cloned into pAL2-T vector (Evrogen, Russia) and sequenced using plasmid-specific primers. In all RT-PCR experiments, probes containing all components but lacking reverse transcriptase were used as negative controls. The obtained sequence of 5′UTR of D. virilis Ranbp16 gene was deposited in GenBank under the number MN481598.
Data availability
The datasets supporting the conclusions of this article are available in the NCBI GEO repository: GSE59965; GSE35648; GSE43829; GSE56347; GSE102439; GSE85404; GSE70632; GSE36737; GSE62904; GSE52887. The obtained sequence of 5′UTR of D. virilis Ranbp16 gene was deposited in GenBank under the number MN481598.
References
Berger, S. L. The complex language of chromatin regulation during transcription. Nature 447, 407–412. https://doi.org/10.1038/nature05915 (2007).
Eissenberg, J. C. & Reuter, G. Cellular mechanism for targeting heterochromatin formation in Drosophila. Int. Rev. Cell Mol. Biol. 273, 1–47. https://doi.org/10.1016/S1937-6448(08)01801-7 (2009).
Filion, G. J. et al. Systematic protein location mapping reveals five principal chromatin types in Drosophila cells. Cell 143, 212–224. https://doi.org/10.1016/j.cell.2010.09.009 (2010).
Riddle, N. C. et al. Plasticity in patterns of histone modifications and chromosomal proteins in Drosophila heterochromatin. Genome Res 21, 147–163. https://doi.org/10.1101/gr.110098.110 (2011).
Riddle, N. C. et al. Enrichment of HP1a on Drosophila chromosome 4 genes creates an alternate chromatin structure critical for regulation in this heterochromatic domain. PLoS Genet 8, e1002954. https://doi.org/10.1371/journal.pgen.1002954 (2012).
Pimpinelli, S. et al. Transposable elements are stable structural components of Drosophila melanogaster heterochromatin. Proc. Natl. Acad. Sci. USA 92, 3804–3808. https://doi.org/10.1073/pnas.92.9.3804 (1995).
Abramov, Y. A. et al. The differences between cis- and trans-gene inactivation caused by heterochromatin in Drosophila. Genetics 202, 93–106. https://doi.org/10.1534/genetics.115.181693 (2016).
Elgin, S. C. & Reuter, G. Position-effect variegation, heterochromatin formation, and gene silencing in Drosophila. Cold Spring Harb. Perspect. Biol. 5, a017780. https://doi.org/10.1101/cshperspect.a017780 (2013).
Sentmanat, M. F. & Elgin, S. C. Ectopic assembly of heterochromatin in Drosophila melanogaster triggered by transposable elements. Proc. Natl. Acad. Sci. USA 109, 14104–14109. https://doi.org/10.1073/pnas.1207036109 (2012).
Shatskikh, A. S., Abramov, Y. A. & Lavrov, S. A. Trans-inactivation: repression in a wrong place. Fly (Austin) 11, 96–103. https://doi.org/10.1080/19336934.2016.1225634 (2017).
Wakimoto, B. T. & Hearn, M. G. The effects of chromosome rearrangements on the expression of heterochromatic genes in chromosome 2L of Drosophila melanogaster. Genetics 125, 141–154 (1990).
Wheeler, B. S., Blau, J. A., Willard, H. F. & Scott, K. C. The impact of local genome sequence on defining heterochromatin domains. PLoS Genet 5, e1000453. https://doi.org/10.1371/journal.pgen.1000453 (2009).
Coulthard, A. B. et al. Essential loci in centromeric heterochromatin of Drosophila melanogaster I: the right arm of chromosome 2. Genetics 185, 479–495. https://doi.org/10.1534/genetics.110.117259 (2010).
Dimitri, P. Cytogenetic analysis of the second chromosome heterochromatin of Drosophila melanogaster. Genetics 127, 553–564 (1991).
Syrzycka, M. et al. Genetic and Molecular Analysis of Essential Genes in Centromeric Heterochromatin of the Left Arm of Chromosome 3 in Drosophila melanogaster. G3 (Bethesda) 9, 1581–1595. https://doi.org/10.1534/g3.119.0003 (2019).
Yasuhara, J. C. & Wakimoto, B. T. Oxymoron no more: the expanding world of heterochromatic genes. Trends. Genet 22, 330–338. https://doi.org/10.1016/j.tig.2006.04.008 (2006).
Bhutkar, A. et al. Chromosomal rearrangement inferred from comparisons of 12 Drosophila genomes. Genetics 179, 1657–1680. https://doi.org/10.1534/genetics.107.086108 (2008).
Schulze, S. R. et al. Heterochromatic genes in Drosophila: a comparative analysis of two genes. Genetics 173, 1433–1445. https://doi.org/10.1534/genetics.106.056069 (2006).
Moschetti, R., Celauro, E., Cruciani, F., Caizzi, R. & Dimitri, P. On the evolution of Yeti, a Drosophila melanogaster heterochromatin gene. PLoS ONE 9, e113010. https://doi.org/10.1371/journal.pone.0113010 (2014).
Yasuhara, J. C., DeCrease, C. H. & Wakimoto, B. T. Evolution of heterochromatic genes of Drosophila. Proc. Natl. Acad Sci. USA 102, 10958–10963. https://doi.org/10.1073/pnas.0503424102 (2005).
Caizzi, R. et al. Comparative genomic analyses provide new insights into the evolutionary dynamics of heterochromatin in Drosophila. PLoS Genet 12, e1006212. https://doi.org/10.1371/journal.pgen.1006212 (2016).
Cho, D. H. et al. Antisense transcription and heterochromatin at the DM1 CTG repeats are constrained by CTCF. Mol. Cell. 20, 483–489. https://doi.org/10.1016/j.molcel.2005.09.002 (2005).
Pikaart, M. J., Recillas-Targa, F. & Felsenfeld, G. Loss of transcriptional activity of a transgene is accompanied by DNA methylation and histone deacetylation and is prevented by insulators. Genes Dev 12, 2852–2862. https://doi.org/10.1101/gad.12.18.2852 (1998).
Yusufzai, T. M. & Felsenfeld, G. The 5’-HS4 chicken beta-globin insulator is a CTCF-dependent nuclear matrix-associated element. Proc. Natl. Acad. Sci. USA 101, 8620–8624. https://doi.org/10.1073/pnas.0402938101 (2004).
Recillas-Targa, F. et al. Position-effect protection and enhancer blocking by the chicken beta-globin insulator are separable activities. Proc. Natl. Acad. Sci. USA 99, 6883–6888. https://doi.org/10.1073/pnas.102179399 (2002).
Kyrchanova, O. & Georgiev, P. Chromatin insulators and long-distance interactions in Drosophila. FEBS Lett. 588, 8–14. https://doi.org/10.1016/j.febslet.2013.10.039 (2014).
Phillips-Cremins, J. E. & Corces, V. G. Chromatin insulators: linking genome organization to cellular function. Mol. Cell. 50, 461–474. https://doi.org/10.1016/j.molcel.2013.04.018 (2013).
Schwartz, Y. B. et al. Nature and function of insulator protein binding sites in the Drosophila genome. Genome Res 22, 2188–2198. https://doi.org/10.1101/gr.138156.112 (2012).
Manak, J. R., Wen, H., Van, T., Andrejka, L. & Lipsick, J. S. Loss of Drosophila Myb interrupts the progression of chromosome condensation. Nat. Cell Biol 9, 581–587. https://doi.org/10.1038/ncb1580 (2007).
Okada, M., Akimaru, H., Hou, D. X., Takahashi, T. & Ishii, S. Myb controls G(2)/M progression by inducing cyclin B expression in the Drosophila eye imaginal disc. EMBO J. 21, 675–684. https://doi.org/10.1093/emboj/21.4.675 (2002).
Drosophila 12 Genomes, C. et al. Evolution of genes and genomes on the Drosophila phylogeny. Nature 450, 203–218. https://doi.org/10.1038/nature06341 (2007).
Gao, J. J., Watabe, H. A., Aotsuka, T., Pang, J. F. & Zhang, Y. P. Molecular phylogeny of the Drosophila obscura species group, with emphasis on the Old World species. BMC Evol. Biol 7, 87. https://doi.org/10.1186/1471-2148-7-87 (2007).
Gibbs, A. G. & Matzkin, L. M. Evolution of water balance in the genus Drosophila. J Exp Biol 204, 2331–2338 (2001).
Jezovit, J. A., Levine, J. D. & Schneider, J. Phylogeny, environment and sexual communication across the Drosophila genus. J Exp Biol 220, 42–52. https://doi.org/10.1242/jeb.143008 (2017).
O’Grady, P. M. & DeSalle, R. Phylogeny of the Genus Drosophila. Genetics 209, 1–25. https://doi.org/10.1534/genetics.117.300583 (2018).
Schaeffer, S. W. et al. Polytene chromosomal maps of 11 Drosophila species: the order of genomic scaffolds inferred from genetic and physical maps. Genetics 179, 1601–1655. https://doi.org/10.1534/genetics.107.086074 (2008).
Bailey, T. L. et al. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res 37, W202-208. https://doi.org/10.1093/nar/gkp335 (2009).
Shazman, S., Lee, H., Socol, Y., Mann, R. S. & Honig, B. OnTheFly: a database of Drosophila melanogaster transcription factors and their binding sites. Nucleic Acids Res 42, D167-171. https://doi.org/10.1093/nar/gkt1165 (2014).
Bergman, C. M., Carlson, J. W. & Celniker, S. E. Drosophila DNase I footprint database: a systematic genome annotation of transcription factor binding sites in the fruitfly Drosophila melanogaster. Bioinformatics 21, 1747–1749. https://doi.org/10.1093/bioinformatics/bti173 (2005).
Rivera, J., Keranen, S. V. E., Gallo, S. M. & Halfon, M. S. REDfly: the transcriptional regulatory element database for Drosophila. Nucleic Acids Res 47, D828–D834. https://doi.org/10.1093/nar/gky957 (2019).
Nègre, N. et al. A comprehensive map of insulator elements for the Drosophila genome. PLoS Genet. 6, e1000814–e1000814. https://doi.org/10.1371/journal.pgen.1000814 (2010).
Bushey, A. M., Ramos, E. & Corces, V. G. Three subclasses of a Drosophila insulator show distinct and cell type-specific genomic distributions. Genes Dev 23, 1338–1350. https://doi.org/10.1101/gad.1798209 (2009).
Liang, J. et al. Chromatin immunoprecipitation indirect peaks highlight long-range interactions of insulator proteins and Pol II pausing. Mol. Cell 53, 672–681. https://doi.org/10.1016/j.molcel.2013.12.029 (2014).
Hou, C., Li, L., Qin, Z. S. & Corces, V. G. Gene density, transcription, and insulators contribute to the partition of the Drosophila genome into physical domains. Mol. Cell 48, 471–484. https://doi.org/10.1016/j.molcel.2012.08.031 (2012).
Gomez-Diaz, E. & Corces, V. G. Architectural proteins: regulators of 3D genome organization in cell fate. Trends Cell Biol 24, 703–711. https://doi.org/10.1016/j.tcb.2014.08.003 (2014).
Yang, J., Ramos, E. & Corces, V. G. The BEAF-32 insulator coordinates genome organization and function during the evolution of Drosophila species. Genome Res. 22, 2199–2207. https://doi.org/10.1101/gr.142125.112 (2012).
Tsai, S. Y., Chang, Y. L., Swamy, K. B., Chiang, R. L. & Huang, D. H. GAGA factor, a positive regulator of global gene expression, modulates transcriptional pausing and organization of upstream nucleosomes. Epigenet. Chromatin 9, 32. https://doi.org/10.1186/s13072-016-0082-4 (2016).
Fuda, N. J. et al. GAGA factor maintains nucleosome-free regions and has a role in RNA polymerase II recruitment to promoters. PLoS Genet 11, e1005108. https://doi.org/10.1371/journal.pgen.1005108 (2015).
Matsukage, A., Hirose, F., Yoo, M. A. & Yamaguchi, M. The DRE/DREF transcriptional regulatory system: a master key for cell proliferation. Biochim. Biophys. Acta 1779, 81–89. https://doi.org/10.1016/j.bbagrm.2007.11.011 (2008).
Hart, C. M., Cuvier, O. & Laemmli, U. K. Evidence for an antagonistic relationship between the boundary element-associated factor BEAF and the transcription factor DREF. Chromosoma 108, 375–383. https://doi.org/10.1007/s004120050389 (1999).
Roy, S., Gilbert, M. K. & Hart, C. M. Characterization of BEAF mutations isolated by homologous recombination in Drosophila. Genetics 176, 801–813. https://doi.org/10.1534/genetics.106.068056 (2007).
Ranz, J. M. et al. Principles of genome evolution in the Drosophila melanogaster species group. PLoS Biol 5, e152. https://doi.org/10.1371/journal.pbio.0050152 (2007).
de Wit, E., Greil, F. & van Steensel, B. High-resolution mapping reveals links of HP1 with active and inactive chromatin components. PLoS Genet 3, e38. https://doi.org/10.1371/journal.pgen.0030038 (2007).
Hearn, M. G., Hedrick, A., Grigliatti, T. A. & Wakimoto, B. T. The effect of modifiers of position-effect variegation on the variegation of heterochromatic genes of Drosophila melanogaster. Genetics 128, 785–797 (1991).
Lu, B. Y., Emtage, P. C., Duyf, B. J., Hilliker, A. J. & Eissenberg, J. C. Heterochromatin protein 1 is required for the normal expression of two heterochromatin genes in Drosophila. Genetics 155, 699–708 (2000).
Lundberg, L. E., Stenberg, P. & Larsson, J. HP1a, Su(var)3–9, SETDB1 and POF stimulate or repress gene expression depending on genomic position, gene length and expression pattern in Drosophila melanogaster. Nucleic Acids Res. 41, 4481–4494. https://doi.org/10.1093/nar/gkt158 (2013).
Eberl, D. F., Duyf, B. J. & Hilliker, A. J. The role of heterochromatin in the expression of a heterochromatic gene, the rolled locus of Drosophila melanogaster. Genetics 134, 277–292 (1993).
Yasuhara, J. C. & Wakimoto, B. T. Molecular landscape of modified histones in Drosophila heterochromatic genes and euchromatin-heterochromatin transition zones. PLoS Genet 4, e16. https://doi.org/10.1371/journal.pgen.0040016 (2008).
Howe, M., Dimitri, P., Berloco, M. & Wakimoto, B. T. Cis-effects of heterochromatin on heterochromatic and euchromatic gene activity in Drosophila melanogaster. Genetics 140, 1033–1045 (1995).
Figueiredo, M. L., Philip, P., Stenberg, P. & Larsson, J. HP1a recruitment to promoters is independent of H3K9 methylation in Drosophila melanogaster. PLoS Genet 8, e1003061. https://doi.org/10.1371/journal.pgen.1003061 (2012).
Jiang, N., Emberly, E., Cuvier, O. & Hart, C. M. Genome-wide mapping of boundary element-associated factor (BEAF) binding sites in Drosophila melanogaster links BEAF to transcription. Mol. Cell. Biol. 29, 3556–3568. https://doi.org/10.1128/MCB.01748-08 (2009).
Maksimenko, O. & Georgiev, P. Mechanisms and proteins involved in long-distance interactions. Front. Genet 5, 28. https://doi.org/10.3389/fgene.2014.00028 (2014).
Ramirez, F. et al. High-resolution TADs reveal DNA sequences underlying genome organization in flies. Nat. Commun 9, 189. https://doi.org/10.1038/s41467-017-02525-w (2018).
Shrestha, S., Oh, D. H., McKowen, J. K., Dassanayake, M. & Hart, C. M. 4C-seq characterization of Drosophila BEAF binding regions provides evidence for highly variable long-distance interactions between active chromatin. PLoS ONE 13, e0203843. https://doi.org/10.1371/journal.pone.0203843 (2018).
Zolotarev, N. et al. Architectural proteins Pita, Zw5, and ZIPIC contain homodimerization domain and support specific long-range interactions in Drosophila. Nucleic Acids Res 44, 7228–7241. https://doi.org/10.1093/nar/gkw371 (2016).
Gurudatta, B. V., Yang, J., Van Bortle, K., Donlin-Asp, P. G. & Corces, V. G. Dynamic changes in the genomic localization of DNA replication-related element binding factor during the cell cycle. Cell Cycle 12, 1605–1615. https://doi.org/10.4161/cc.24742 (2013).
Kwon, E. et al. Transcription control of a gene for Drosophila transcription factor, DREF by DRE and cis-elements conserved between Drosophila melanogaster and virilis. Gene 309, 101–116. https://doi.org/10.1016/s0378-1119(03)00493-1 (2003).
Guertin, M. J., Petesch, S. J., Zobeck, K. L., Min, I. M. & Lis, J. T. Drosophila heat shock system as a general model to investigate transcriptional regulation. Cold Spring Harb. Symp. Quant. Biol 75, 1–9. https://doi.org/10.1101/sqb.2010.75.039 (2010).
Lebedeva, L. A. et al. Occupancy of the Drosophila hsp70 promoter by a subset of basal transcription factors diminishes upon transcriptional activation. Proc. Natl. Acad. Sci. USA 102, 18087–18092. https://doi.org/10.1073/pnas.0509063102 (2005).
Lomaev, D. et al. The GAGA factor regulatory network: Identification of GAGA factor associated proteins. PLoS ONE 12, e0173602. https://doi.org/10.1371/journal.pone.0173602 (2017).
Gilchrist, D. A. et al. NELF-mediated stalling of Pol II can enhance gene expression by blocking promoter-proximal nucleosome assembly. Genes Dev 22, 1921–1933. https://doi.org/10.1101/gad.1643208 (2008).
Kriventseva, E. V. et al. OrthoDB v8: update of the hierarchical catalog of orthologs and the underlying free software. Nucleic Acids Res. 43, D250–D256. https://doi.org/10.1093/nar/gku1220 (2015).
States, D. J. & Gish, W. Combined use of sequence similarity and codon bias for coding region identification. J. Comput. Biol. 1, 39–50. https://doi.org/10.1089/cmb.1994.1.39 (1994).
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410. https://doi.org/10.1016/S0022-2836(05)80360-2 (1990).
Sigrist, C. J. et al. PROSITE: a documented database using patterns and profiles as motif descriptors. Brief Bioinform 3, 265–274. https://doi.org/10.1093/bib/3.3.265 (2002).
Sigrist, C. J. et al. New and continuing developments at PROSITE. Nucleic Acids Res 41, D344-347. https://doi.org/10.1093/nar/gks1067 (2013).
Li, R. et al. ReAS: Recovery of ancestral sequences for transposable elements from the unassembled reads of a whole genome shotgun. PLoS Comput Biol 1, e43. https://doi.org/10.1371/journal.pcbi.0010043 (2005).
Smit, A.F. & Hubley, R. RepeatModeler Open-1.0. Available from http://www.repeatmasker.org (2008).
Thompson, J. D., Higgins, D. G. & Gibson, T. J. CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22, 4673–4680. https://doi.org/10.1093/nar/22.22.4673 (1994).
Sievers, F. & Higgins, D. G. Clustal Omega, accurate alignment of very large numbers of sequences. Methods Mol. Biol. 1079, 105–116. https://doi.org/10.1007/978-1-62703-646-7_6 (2014).
Waterhouse, A. M., Procter, J. B., Martin, D. M., Clamp, M. & Barton, G. J. Jalview Version 2–a multiple sequence alignment editor and analysis workbench. Bioinformatics 25, 1189–1191. https://doi.org/10.1093/bioinformatics/btp033 (2009).
Krzywinski, M. et al. Circos: an information aesthetic for comparative genomics. Genome Res. 19, 1639–1645. https://doi.org/10.1101/gr.092759.109 (2009).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S. L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25. https://doi.org/10.1186/gb-2009-10-3-r25 (2009).
Li, H. et al. The sequence alignment/map format and SAMtools. Bioinformatics 25, 2078–2079. https://doi.org/10.1093/bioinformatics/btp352 (2009).
Thorvaldsdottir, H., Robinson, J. T. & Mesirov, J. P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief Bioinform 14, 178–192. https://doi.org/10.1093/bib/bbs017 (2013).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol 9, R137. https://doi.org/10.1186/gb-2008-9-9-r137 (2008).
Ramirez, F., Dundar, F., Diehl, S., Gruning, B. A. & Manke, T. deepTools: a flexible platform for exploring deep-sequencing data. Nucleic Acids Res 42, W187-191. https://doi.org/10.1093/nar/gku365 (2014).
Fornes, O. et al. JASPAR 2020: update of the open-access database of transcription factor binding profiles. Nucleic Acids Res. 48, D87–D92. https://doi.org/10.1093/nar/gkz1001 (2020).
Tan, G. & Lenhard, B. TFBSTools: an R/bioconductor package for transcription factor binding site analysis. Bioinformatics (Oxford, England) 32, 1555–1556. https://doi.org/10.1093/bioinformatics/btw024 (2016).
Fresno, C. & Fernandez, E. A. RDAVIDWebService: a versatile R interface to DAVID. Bioinformatics 29, 2810–2811. https://doi.org/10.1093/bioinformatics/btt487 (2013).
R: A language and environment for statistical computing (R Foundation for Statistical Computing, 2010).
Krasnov, G. S. et al. PPLine: an automated pipeline for SNP, SAP, and splice variant detection in the context of proteogenomics. J. Proteome Res. 14, 3729–3737. https://doi.org/10.1021/acs.jproteome.5b00490 (2015).
Liao, Y., Smyth, G. K. & Shi, W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics (Oxford, England) 30, 923–930. https://doi.org/10.1093/bioinformatics/btt656 (2014).
Bolger, A. M., Lohse, M. & Usadel, B. Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics (Oxford, England) 30, 2114–2120. https://doi.org/10.1093/bioinformatics/btu170 (2014).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics (Oxford, England) 29, 15–21. https://doi.org/10.1093/bioinformatics/bts635 (2013).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics (Oxford, England) 26, 139–140. https://doi.org/10.1093/bioinformatics/btp616 (2010).
Walter, W., Sánchez-Cabo, F. & Ricote, M. GOplot: an R package for visually combining expression data with functional analysis. Bioinformatics (Oxford, England) 31, 2912–2914. https://doi.org/10.1093/bioinformatics/btv300 (2015).
Bailey, T. L. & Elkan, C. Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proc. Int. Conf. Intell. Syst. Mol. Biol. 2, 28–36 (1994).
Gupta, S., Stamatoyannopoulos, J. A., Bailey, T. L. & Noble, W. S. Quantifying similarity between motifs. Genome Biol 8, R24. https://doi.org/10.1186/gb-2007-8-2-r24 (2007).
Kumar, S., Stecher, G., Li, M., Knyaz, C. & Tamura, K. MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol. Biol. Evol 35, 1547–1549. https://doi.org/10.1093/molbev/msy096 (2018).
Tamura, K. & Nei, M. Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Mol. Biol. Evol 10, 512–526. https://doi.org/10.1093/oxfordjournals.molbev.a040023 (1993).
Suyama, M., Torrents, D. & Bork, P. PAL2NAL: robust conversion of protein sequence alignments into the corresponding codon alignments. Nucleic Acids Res. 34, W609-612. https://doi.org/10.1093/nar/gkl315 (2006).
Yang, Z. PAML 4: phylogenetic analysis by maximum likelihood. Mol. Biol. Evol 24, 1586–1591. https://doi.org/10.1093/molbev/msm088 (2007).
Lim, J. In situ hybridization with biotinylated DNA. Dros. Inf. Serv 72, 73–77 (1993).
Gubenko, I. S. & Evgen’ev, M. B. Cytological and linkage maps of Drosophila virilis chromosomes. Genetica 65, 127–139. https://doi.org/10.1007/bf00135277 (1984).
Acknowledgements
We are grateful to Drs. Pavel Georgiev and Anton Golovnin from the Institute of gene biology RAS for ideas and critical comments on the manuscript. We thank Alexei M. Kulikov from the Koltzov Institute of developmental biology RAS for helpful advice and technical assistance. The bioinformatics was performed using the computational facilities of Engelhardt Institute of Molecular Biology RAS Genome center (https://www.eimb.ru/rus/ckp/ccu_genome_c.php). This work was supported by the Russian Foundation for Basic Research (Grant number 19-04-00337). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
Conceptualization: S.Y.F., I.R.A., M.B.E. Data curation: L.N.C., L.A.P., E.S.Z. Formal analysis: L.A.P., V.T., A.P.R., L.N.C. Funding acquisition: S.Y.F. Investigation: S.Y.F., D.A.K., E.S.Z., L.A.P. Methodology: S.Y.F., A.P.R., I.R.A. E.S.Z. Project administration: M.B.E. Software: A.P.R. Supervision: M.B.E. Writing—original draft: S.Y.F., M.B.E. Writing—review and editing: I.R.A., E.S.Z.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Funikov, S.Y., Rezvykh, A.P., Kulikova, D.A. et al. Adaptation of gene loci to heterochromatin in the course of Drosophila evolution is associated with insulator proteins. Sci Rep 10, 11893 (2020). https://doi.org/10.1038/s41598-020-68879-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-68879-2
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.