Abstract
The transition of the martian climate from the wet Noachian era to the dry Hesperian (4.1–3.0 Gya) likely resulted in saline surface waters that were rich in sulfur species. Terrestrial analogue environments that possess a similar chemistry to these proposed waters can be used to develop an understanding of the diversity of microorganisms that could have persisted on Mars under such conditions. Here, we report on the chemistry and microbial community of the highly reducing sediment of Colour Peak springs, a sulfidic and saline spring system located within the Canadian High Arctic. DNA and cDNA 16S rRNA gene profiling demonstrated that the microbial community was dominated by sulfur oxidising bacteria, suggesting that primary production in the sediment was driven by chemolithoautotrophic sulfur oxidation. It is possible that the sulfur oxidising bacteria also supported the persistence of the additional taxa. Gibbs energy values calculated for the brines, based on the chemistry of Gale crater, suggested that the oxidation of reduced sulfur species was an energetically viable metabolism for life on early Mars.
Similar content being viewed by others
Introduction
Sulfurous and saline waters are proposed to have existed on the surface of Mars during the Noachian–Hesperian transition (4.1–3.0 Gya)1,2,3,4,5,6, whereby in the Noachian period, liquid water formed widespread surface features, such as stream beds and sedimentary deposits, and led to the depositions of clay minerals4. Many locations on Mars feature rock formations rich in sulfur species, with sulfate and sulfide minerals detected by lander missions7,8,9,10,11 and within martian meteorites12,13,14,15,16. At the end of the Noachian and into the beginning of the Hesperian, the presence of water declined and saline-rich brines formed, as evidenced by the jarosite at Gale crater17. On modern day Mars, water is restricted to the subsurface or—potentially—sub-glacial areas at the poles4,18,19.
In sulfur-rich environments on Earth, primary production is typically driven by the oxidation of reduced sulfur species20,21,22,23. The sulfur biogeochemical cycle involves metabolic activity associated with multiple microbial pathways, and molecular and physiological data indicate that these sulfur oxidation–reduction (redox) reactions are an ancient metabolism24. The presence of sulfur species in different redox states on Mars and the detection of suitable electron donors and acceptors (e.g. nitrate and oxygen25,26,27) raises the possibility of whether the sulfur biogeochemical cycle, specifically the oxidation of reduced sulfur species, is plausible on Mars. This is especially important, because Mars offers a wide range of environmental conditions, mostly dominated by basalt-water reactions, with pH varying from alkaline28,29 to acidic (in conjunction with volcanic activity)30. The availability of water and water activity have been invoked as limiting factors for habitability31,32, but predictions suggest a range of water chemistries have existed through martian geological time, from the dilute “groundwater-type”28,29 to the highly concentrated brines associated with the modern day recurring slope lineae occurrences17,33.
Based on the tiered model method of Soare34, Axel Heiberg Island in the Canadian High Arctic represents an ideal analogue to study microbial processes within sulfurous aqueous environments similar to those that existed on Mars, e.g., at Gale crater at the Noachian-Hesperian boundary. The brines were sulfur-rich and became concentrated by processes such as freezing or evaporation29. Figures 1 and 2 illustrate this point, with sulfate-vein forming fluids at Gale crater being most similar to fluids in saline lakes on Earth and in the Axel Heiberg spring. At the same time, the fluids from clay formation events, and more generally the alteration of basaltic material under high water–rock reactions on Mars, matches fluids from terrestrial geologic settings, such as the Deccan traps and Icelandic springs (see Figs. 1 and 2)1,2,3,4,5,35,36,37,38,39.
Axel Heiberg Island lies within the region of continuous permafrost40,41 and is host to eight sulfur–rich, highly saline (2–4 M), and perennially cold (0–7 °C) springs41,42,43. Despite an average air temperature of − 15 °C, which decreases to a minimum of − 40 °C in winter, the springs do not freeze2,3, 36, 44. The water in the island’s spring systems persists as groundwater 600 m below the surface. The water discharges in areas associated with diapiric uplift38, which comprises of gypsum–anhydrite of upper-Mississippian to middle-Pennsylvanian age (224–315 Mya)13. The waters are anoxic upon exiting the diapirs and become rapidly oxidised at the surface2,42. The springs that have been previously characterised with regards to their chemistry and mineralogy are Lost Hammer (LH), Gypsum Hill (GH) and Colour Peak (CP)2,38,42,43. These studies have indicated that the mineralogy of deposits at these spring sites are predominantly calcite (CaCO3), halite (NaCl), thenardite (Na2SO4), mirabilite (Na2SO4·10H2O), gypsum (CaSO4·2H2O) and elemental sulfur (S°)38,42,43, but variations exist between springs as a result of differences in their fluid geochemistry. For example, LH fluids have higher concentrations of sodium than CP fluids, but lower concentrations of sulfide (0.14 mM) and calcium (24.43 mM) than CP (1.8 mM and 33.23 mM respectively)43. The sediments of the Axel Heiberg springs are typically highly reducing, with some sediments containing both anoxic and microaerophilic zones38.
The CP spring system consists of a series of springs that discharge into deep gullies located near the base of the south–facing slope of CP, as shown in Fig. 343. The CP spring waters possess a broadly similar composition and pH (7.3–7.943) to that of a thermodynamically–modelled martian evaporitic fluid (based on the chemistry of Gale crater, Figs. 1 and 2). The CP spring system is therefore a recognised Mars analogue site3, 45 that possesses a chemistry similar to that proposed for the late Noachian era. However, despite several studies investigating the springs on Axel Heiberg Island, specific microbiological studies of the CP spring system have been less forthcoming. The resident microbial community has previously only been characterised through the creation and sequencing of clone libraries (174 clones for bacterial diversity and 164 for archaeal diversity) and no strains have been isolated from this site45.
(A) Map of Axel Heiberg Island and (B) photograph of the Colour Peak (CP) site illustrating the location of CP springs and their sources. (A) Map of Axel Heiberg Island (brown), and icecaps (white) produced by modifying an image from Google Maps (Map Data @2020 Google) using Illustrator Creative Cloud version 21.0.2. (B) Photographs of the CP springs and precipitates along a CP spring channel with annotated spring and channel systems. The site where the sediment was collected is marked with a red dot.
Conversely, LH and GH have been studied using 16S rRNA gene analysis and a multiple amplification enabled metagenome44,45,58,59,60,61,62. At higher taxonomic levels, many of the previous DNA-based studies have shown microbial communities dominated by Gammaproteobacteria, specifically those genera that are associated with sulfur oxidation45,60,61,62, a factor not detected in the cDNA profiles of the LH springs58,59. However, the preservation of environmental DNA under high salinity and cold temperatures63,64,65,66 limits the extent to which the bacterial community within the sediment can be characterised using this approach, since the 16S rRNA gene profile also captures sequences from dead cells. This necessitates the analysis of cDNA produced from RNA extracted from the sediment.
This paper presents the first in-depth characterisation of the viable microbial community at CP springs, determined through DNA and RNA extraction from a CP sediment core. cDNA was produced from the RNA and 16S rRNA amplicons were sequenced using current generation sequencing platforms. Several halophilic bacteria were isolated, representing the first cultivation-dependent characterisation of this site. The viability of metabolisms under thermochemically-modelled martian fluids was also estimated using Gibbs energy (formally called Gibbs free energy) equations to further investigate the habitability of the waters on Mars from the late Noachian.
Results
Geochemical characterisation
Inductively Coupled Plasma–Optical Emission Spectroscopy (ICP–OES) was used to identity the bioavailable elements in: (1) water from CP; (2) CP water that had CP sediment resuspended in it for seven days; (3) 17% NaCl solution that had CP sediment resuspended in it for seven days, and (4) 17% NaCl solution analysed as a control (Table 1). The CP fluids contained high amounts of sodium (69,600 mg kg−1), calcium (2,850 mg kg−1), sulfur (563 mg kg−1) and potassium (97 mg kg−1). Interaction with the CP sediment increased the concentrations of potassium (threefold), magnesium (tenfold), strontium (ninefold) and sulfur (1.5-fold). The concentrations of sodium and calcium did not alter from contact with the sediment. The high concentrations of sulfur present in the CP waters and sediment is congruent with the prior detection of sulfur oxidising bacteria within the CP springs45, as this represents a highly abundant and bioavailable electron donor and would be expected to impact on the resident microbial community.
Gibbs energy calculations
The Gibbs energy (ΔG) of sulfide oxidation was calculated to assess the energetic feasibility of this reaction within thermochemically-modelled martian brines (with chemistries similar to CP waters)57. The fluid chemistries used for the Gibbs energy calculations were modelled using the concentration of oxygen shown to be viable in the martian near-surface67 and the concentration of nitrate detected in ancient mudstones by the Mars Curiosity rover (e.g., the lower and upper limits of nitrate were 70 and 1,200 ppm respectively26) (Supplementary Table 1). ΔG values indicated that aerobic sulfide oxidation was viable, yielding 2.36 × 10–4 kJ kg−1(fluid) in the fluids modelled with both 70 and 1,200 ppm of nitrate. Denitrification-fuelled sulfide oxidation was also shown to be viable, yielding 2.21 × 10–5 kJ kg-1(fluid) in the modelled fluid chemistries with 70 and 1,200 ppm of nitrate. In terms of cell biomass, this translates to 2.97 × 107 cells kg−1(fluid) supported by aerobic sulfide oxidation and 2.78 × 106 cells kg−1(fluid) supported by anaerobic sulfide oxidation. Other potential metabolisms were viable (Supplementary Table 1), but none were more energy yielding than sulfide oxidation.
Microbial community within the colour peak sediment
DNA and RNA (converted to cDNA) were extracted from replicate sediment samples collected from CP and sequenced using the Ion Torrent PGM platform. 113,278 16S rRNA gene sequences were obtained from the DNA extractions, post-quality control (Supplementary Table 2). Sequences were evenly distributed between the three samples examined (35,201 Sample 1 (S1); 37,206 at Sample 2 (S2); and 40,871 at Sample 3 (S3). cDNA produced from RNA was extracted from the same three sediment samples, pooled, and 41,955 16S rRNA sequences obtained post-quality control.
Alpha diversity metrics were applied to both data sets to assess the community diversity. The number of operational taxonomic units (OTUs, sequences with > 97% sequence identity) in the 16S rRNA gene profile ranged from between 347 and 401 (S1, 401 OTUs; S2, 347 OTUs; S3, 376 OTUs). 94–98% of the OTUs present in the 16S rRNA gene profiles were common to each sample (Fig. 4). Whilst there were over 300 OTUs detected in the DNA profiles, only 33 were detected in the cDNA data. Diversity indices (Faith pd, Shannon, Simpson and Simpson evenness) confirmed that the diversity in the 16S rRNA gene profiles was greater than that of the 16S rRNA profile of the cDNA (Supplementary Table 3). Beta diversity metrics (Euclidian Distance, Dice measure, Chebyshev distance) were applied to identify variation in community structure between the DNA and cDNA profiles and showed that there was less variation within the DNA profiles compared to the cDNA profile (Supplementary Tables 4, 5 and 6).
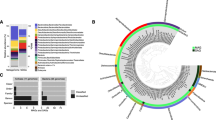
Shared diversity at the genus level between the 16S rRNA gene and 16S rRNA profiles of the Colour Peak sediment. All genera present at greater than 100 reads in the 16S rRNA gene and 16S profiles were compared to identify shared and unique genera between the three replicate sediment samples and the RNA profile of Colour Peak sediment.
Taxonomical assignment demonstrated that the majority of the sequences in the DNA profiles belonged to the phylum Proteobacteria (Fig. 5), which represented between 74% (S1) and 78% (S2) of the relative abundance. Sequences that were assigned to the phylum Proteobacteria, were mainly identified at Class level as Gammaproteobacteria (74–80%). This was followed by the Alphaproteobacteria (8–10%), Bacteroidetes (8%), Firmicutes (6–9%), Betaproteobacteria (4–6%), Epsilonbacteraetota (2–3%) and Cyanobacteria (3%). Halothiobacillus was the most abundant genus (on average 40% of the relative abundance of the total community profile). Other gammaproteobacterial genera associated with sulfur oxidation were also present (Thiobacillus, Thiomicrosospira, Halomonas, Marinobacter and Salinisphaera (> 1% of the total relevant abundance)). With regards to Archaea, the amplicons produced with the universal primers contained no archaeal sequences and the screening of the cDNA and DNA with archaeal specific primers did not produce a successful amplicon. Whilst the lack of detection of Archaeal signatures is unexpected, halophilic bacteria have previously been shown to be capable of outcompeting halophilic archaea under moderately saline (20%) and colder conditions68, which may have resulted in either the competitive exclusion of the archaea or a reduction in their abundance within the site.
16S rRNA gene and 16S rRNA community profiles of Colour Peak (CP) sediment. Sequences were revealed by amplicon sequencing of 16S rRNA gene and 16S rRNA amplicons retrieved by PCR from DNA and RNA extracted from three replicate sediment samples (S1, S2, S3) collected from CP and the pooled 16S rRNA profile of the CP sediment. All genera pictured are present at > 1% relative abundance. DNA refers to an averaged community profile of the three replicate CP 16S rRNA gene profiles.
100% of the cDNA profile was comprised of genera that were also identified in the 16S rRNA gene profiles (Table 2) and was dominated by Proteobacteria (99%), specifically the Gammaproteobacteria (99%). The dominant genus of the 16S rRNA profile was Halothiobacillus, representing 98% of the relative abundance in the cDNA profiles. Other genera present at a read number greater than 100 were Halomonas and Halanaerobium.
Isolation and identification of microbial isolates
19 isolates representing 11 genera were isolated from the sediment (Supplementary Table 7). All of the isolates belonged to the genera Halomonas, Psychrobacter, Marinobacter, Loktanella, Salegentibacter, Sphingopyxis, Sporosarcina, Variovorax, Acidovorax and Nevskia (> 98% identity to known NCBI sequences). Three strains that were isolated from the CP sediment, Psychrobacter (CP4 and CP5) and Sporosarcina (CP16), belong to genera that were not present in the sequencing profiles. This is perhaps a result of the enrichment process which has increased the abundance of strains from genera that were present in the natural community at sufficiently low abundances to preclude their detection in the 16S rRNA gene amplicons.
Discussion
In this study, high-throughput sequencing was utilised to study the microbial community of CP, a saline and sulfate-rich spring system located in the Canadian High Arctic, which is chemically similar to waters proposed to have been present on the surface of Mars during the late Noachian1,2,3,4,5. 16S rRNA gene data demonstrated that the sediment was dominated by members of gammaproteobacterial genera that are comprised solely of obligate chemolithoautotrophic sulfur oxidisers69,70,71, specifically Halothiobacillus (35–43% relative abundance) and Thiomicrosopsira (2–3% relative abundance), or contained species that were capable of complete or partial sulfur oxidation, for example Marinobacter (4–6%) and Halomonas (2–3%)72,73. Sequences belonging to Sulfurovum and Sulfurospirillum (members of the Epsilonbacteraetota Phylum) together represented 3% of the community profile. The dominance of diversity associated with sulfur-related metabolisms is presumably due to the high concentrations of bioavailable sulfur detected within the sediment and waters of the CP spring system. The genera associated with sulfur oxidation that are most abundant in the CP sediment are most commonly associated with the oxidation of sulfide to sulfite by sulfite reductase and subsequent transformation of the sulfite to thiosulfate by thiosulfate sulfurtransferase. The thiosulfate is then oxidised to sulfate via the enzymes of the Sox pathway69,70,71. Species of Thiobacillus that were also detected in the CP sediment have also been shown to complete sulfur oxidation via the oxidation of sulfite to sulfate by a sulfite dehydrogenase and via the reverse dissimilatory sulfate reduction pathway71. Sulfate is the end product of these sulfur oxidation pathways, but with the accumulation of thiosulfate, sulfur and sulfite as metabolic intermediates shown to occur within the diversity detected71.
The results from this study are consistent with previous 16S rRNA gene profiles of the LH, GH and CP spring sediments that showed a dominance of Gammaproteobacterial genera associated with sulfur oxidation45,58,60,62. However, previous 16S rRNA profiling of the LH springs showed a dominance of either Chloroflexi and Alphaproteobacteria and a minimal presence of Gammaproteobacteria40 or showed abundant Gammaproteobacteria but this was diversity not associated with sulfur oxidation38. This study, however, showed that the CP sediment was dominated by Gammaproteobacteria, specifically the genus Halothiobacillus (98% relative abundance). This result indicates that chemolithoautotrophic sulfur oxidation is an active process within the CP sediment.
As well as water chemistry, another key environmental parameter of this site that is analogous to Mars is the low temperature, which models have shown exist at atmospheric pressures between 100 mbar and 4 bar that can be constrained through the investigation of carbonates and sulfates18,74,75. Climate models indicate that, during the Noachian, Mars would have experienced seasonal temperature variations around the freezing point of water at low latitudes. This would have allowed surface water features, such as rivers and lakes, to form and be sustained18,76, but on modern day Mars water reservoirs are expected to be found in the subsurface77,78,79,80. This is consistent with the temperatures observed in CP, with air temperatures consistently below freezing. This is reflected in the fact that the majority of the isolates (e.g. Marinobacter, Halomonas, Loktanella and Psychrobacter) obtained in this study are > 98% homologous to isolates previously detected in the Arctic or other cryoenvironments, including GH and LH springs44,45,81,82,83,84. Further, the 16S rRNA profile, 16S rRNA gene sequence data and isolates gained from this study reinforce the similarities in community composition between the separate sites on Axel Heiberg island identified in previous studies59,60,62. This could be explained by the sites having a common, or related, water source42, but determining the relative roles of stochastic and deterministic factors that might control community composition in the sediments would require a larger and more rigorous sampling effort85. The survival and growth of these organisms within the CP sediment suggests that they are suitable candidates for studies simulating the martian chemical environment.
Within the CP sediment, functional guilds associated with both obligate anaerobism (e.g., fermentative metabolisms) and aerobism (e.g., sulfur-oxidising metabolisms) were identified in the 16S rRNA profile. The continued viability of obligate anaerobes within the CP sediment proves that conditions enabling their survival exist, possibly within anaerobic microenvironments within the heterogeneous sediment38. Based on the chemistry and the community profiling of the CP spring sediment, the comparison with sulfur-rich Mars raises the potential for the oxidation of reduced sulfur species to be an energy yielding metabolism that could fuel primary production within ancient martian environments, or even in modern subsurface environments. Further, sulfide-bearing minerals, including pyrite and pyrrhotite, have been detected in martian meteorites and via in-situ measurements from the martian surface14,15, which could be used as electron donors for this metabolism.
On Earth, the oxidation of reduced sulfur compounds by sulfur oxidising bacteria can be coupled to either oxygen (under aerobic conditions) or nitrate (under microaerophilic or anaerobic conditions) as electron acceptors. For example, this occurs in marine sediments and hydrothermal vents, where there is limited light and a gradient in the availability of oxygen and nitrate86,87, and in artificial environments, such as wastewater treatment plants, which have higher concentrations of nitrate88,89. On Mars, the oxidation of sulfur species could be coupled to these electron acceptors in a similar manner90. For example, oxygen has been detected in the martian atmosphere by the Curiosity rover25, with thermodynamic models indicating that subsurface environments on Mars could possess sufficient O2 to allow for aerobic metabolisms to be viable67. The Curiosity rover has also detected nitrates in ancient mudstones analysed at Gale crater at 70–1,100 ppm26. These values were used in the Gibbs energy calculations presented here and showed that, in the presence of modelled Mars-relevant brines, both aerobic and anaerobic sulfide oxidation are thermodynamically viable in a martian chemical environment.
Using the concentrations of oxygen shown to be thermodynamically feasible in brines under near surface conditions67, aerobic sulfide oxidation was shown to support a greater number of cells than denitrification-fuelled sulfide oxidation (2.97 × 107 cells kg−1(fluid) and 2.78 × 106 cells−1(fluid), respectively). Phototrophic sulfur oxidation may have also been a plausible metabolism in the late Noachian. However, as phototrophy is dependent on light, it is restricted to the surface and not viable in the sub-surface sediment environments considered here. Denitrification-fuelled sulfide oxidation was shown to be viable with both the higher and lower concentrations of nitrate detected in the mudstone from Gale crater, suggesting that this metabolism would have been feasible within a broad range of potential environments on Mars during the late Noachian.
If sulfur-oxidising microorganisms existed on Noachian Mars, evidence of their activity might be preserved in Noachian-aged martian sediments. The entombment of microbial lipids within iron sulfates and iron oxides might be used to identify the presence of life, however there is ambiguity with regards to the identity and metabolisms of the microbes associated with the lipids91. Biosignatures more specific to sulfur oxidising bacteria include the enhanced formation and altered composition of specific biominerals92,93, e.g., the enhanced production of gypsum38 or the substitution of calcium by barium in gypsum94. Specific sulfur oxidising bacteria also possess unique budding and filamentous cell morphologies that could be preserved in specific environmental systems95. However, the concentration of sulfur within a system cannot be used as a reliable biosignature, since different sulfur oxidising bacteria are known to either accumulate or enhance the removal of extracellular sulfur92,96,97,98. Sulfur isotopic fractionation patterns, however, could be utilised since the oxidation of reduced sulfur compounds by sulfur oxidising bacteria has recently been shown to enrich its oxidation products with 34S99. Although variable sulfur isotopic compositions have been observed between sediments at Gale crater on Mars, this does not allow the conclusion for a biological origin because it is at present not possible to discount all non-biological reasons for these differences16.
In addition to autotrophic sulfur oxidisers, the cDNA profile of the CP sediment included strains of fermentative bacteria and Halomonas, a genus that includes heterotrophic strains100 that require an exogenous source of organic carbon. Carbon has been detected on the surface of Mars and in martian meteorites101. Sutter101 postulated that < 1% of the carbon detected at Gale crater would support the biomass requirements for 1 × 105 cells g−1 sediment101. In addition, if sulfur biogeochemical cycling occurred on Mars, the organic carbon could be supplied via the secretion and necrophagy of sulfur oxidising bacteria102. The role of sulfur oxidising bacteria as primary producers within environments raises the issue of whether Halothiobacillus could therefore be considered a keystone species or sulfur oxidation a keystone function in the CP sediment85, enabling the viability of additional metabolisms. If so, it could be extrapolated that this could also occur under proposed martian conditions (Supplementary Fig. 1). Syntrophy and co-cultivation have been shown to be highly influential to the persistence of microbial populations103,104,105. Therefore, in a community-dependent context, a greater diversity of metabolisms might be viable, which could have profound implications for the formation and preservation of biosignatures under martian chemical conditions.
Conclusion
Constraining the parameters concerning biosignature formation in the former and potentially extant waters of Mars requires the identification of organisms capable of surviving in sites that represent appropriate analogue environments. The sulfidic, sulfurous and saline conditions of the CP spring system on Axel Heiberg Island represent such an environment, as an analogue for waters on the surface of Mars during the Noachian–Hesperian transition. This study shows that the microbial community within the CP sediment was dominated by bacteria associated with the oxidation of reduced sulfur species. Based on thermochemical models for the sulfur-rich brines of the Noachian–Hesperian period, conditions could have been thermodynamically viable for similar biotic sulfur oxidation to occur. The potential role of chemolithoautotrophic sulfur oxidation as a keystone function that drives primary production and helps to maintain diversity in terrestrial environments has implications for our understanding of the habitability of martian environments by non-chemolithoautotrophic metabolisms and the subsequent impacts on biosignature formation. The relationship and dependencies between metabolisms require further exploration under controlled laboratory conditions, with simulation studies presenting an appropriate method for achieving this future goal.
Methods
Site sampling and description
Samples were collected during the summer field season in 2017. Sediment samples were aseptically collected from a sediment-rich pool (79.381359°, − 91.272664°) for molecular analysis and culturing and were stored at ambient arctic temperatures whilst in the field. All tools used to collect the samples were cleaned with 95% ethanol and rinsed with autoclaved ddH2O between sampling. Approximately 150 g of sediment was collected and stored in sterile 50 ml tubes (for molecular work) and 125 ml Nalgene bottles (for culturing). The samples were shipped to the UK chilled (4 °C) and on return to the laboratory were stored at – 80 °C (for molecular analysis) and 4 °C (for culturing). Temperature, pH and dissolved oxygen (DO) concentrations were measured in situ using a Mettler Toledo FiveGo probe.
Nucleic acid extraction
The extraction process was performed in a clean hood (PURAIR, Air Science) used exclusively for low biomass samples. Prior to use, the hood was sterilized with 2% chemgene and RNaseZap (ThermoFisher) and then UV sterilized for 72 h. The extractions were performed using the XS buffer extraction technique106. All reagents, except the potassium ethyl xanthogenate, were UV sterilized and after preparation the XS buffer was filter–sterilized through a 0.22 µm filter. For each stage of the extraction process, nuclease–free water (Sigma) was introduced as an additional negative control. All controls were processed in parallel with the samples and used as negative extraction controls in the PCRs.
10 g from each sediment sample was suspended in 30 ml of PCR grade molecular water (Sigma). The sediment was vortexed for 20 min prior to centrifugation at 1,000×g. After 5 min, the supernatant was removed and centrifuged for a further 5 min at 4,700×g through a 15 ml 30 kDa UV sterilised filter (Merck). The tube was emptied, the filters were washed with 400 µL of 1 M pH 8 Tris HCL and then 400 µl of XS buffer was added to the wash solution106. DNA was extracted from the filtered samples using the XS buffer DNA extraction technique, with the addition of a freeze–thaw step, with the samples being stored at – 80 °C for 30 min after 30 min of incubation at 65 °C106. Nucleic acids were precipitated with one volume of ethanol and 4 µl of GlycoBlue Coprecipitant (ThermoFisher), as per manufacturer’s instructions. The supernatant was discarded and the pellet was washed in 200 µl of 70% ice cold ethanol and air dried as in Green et al.107.
To remove excess salts, the nucleic acids were re-suspended in 1.5 mL of PCR grade molecular water and then centrifuged at 12,000×g for five mins through a 500 µl volume UV sterilised 30 kDA filter (Merck). The filter was inverted, transferred to a new tube, and centrifuged as described previously. The eluted volume was adjusted to a final volume of 40 µL. DNA was quantified using 1 μL with the high sensitivity DNA assay for Qubit fluorometric quantitation (ThermoFisher). Ten microlitres of the nucleic acid suspension was stored for DNA analysis whilst the remaining volume of nucleic acids extracted from each sample was pooled prior to being treated with DNase using the TURBO DNA-free™ Kit (Thermofisher) according to manufacturer’s instructions. Samples were pooled to ensure sufficient yield for successful reverse transcription. To prepare cDNA via reverse transcription, the PCR BIOSYSTEMS qPCRBIO cDNA Synthesis Kit was used according to manufacturer’s instructions.
PCR amplification and Ion torrent sequencing
Both DNA and cDNA extracts were PCR amplified using a set of primers (515F-806R) specific to the V4 hypervariable region of the bacterial 16 s rRNA gene108. The PCR reaction mixture contained (per 25 µL): 1 × PCRBIO Ultra Polymerase red mix (PCR BIOSYSTEMS, United Kingdom), 0.4 μM forward primer and 0.4 μM reverse primer. The PCR conditions were an initial denaturation at 95 °C for 5 min, followed by 30 cycles of: denaturing 30 s at 95 °C, annealing at 1 min 56 °C, elongation at 1 min 72 °C and final elongation for 5 min at 72 °C. PCR products were precipitated as previously decribed107, and re-suspended in 20 µl of molecular grade water. Purified PCR products were quantified using Qubit fluorometric quantitation (ThermoFisher) and sequenced using the Ion Torrent PGM platform by the company Molecular Research LP (Texas, USA).
Bioinformatics analysis
The raw sequencing data was processed using the QIIME2 pipeline109. The amplicons were demultiplexed and primers and barcodes removed from all reads. The DADA2 noise removal algorithm was used to remove all chimeric sequences, sequences above 270 bp and the first 15 bp of all sequences. Sequences were then clustered into amplicon sequences variants (ASVs) using the DADA2 algorithm. Phylogeny was assigned to the amplicon sequence variants using Scikit–learn classifier, which compared the ASVs against the Greengenes database110,111 with a confidence threshold of p = 0.7. The ASVs were aligned using MAFFT112 and a rooted tree produced. All of the amplicons were normalised by rarefaction to 35,000 reads and alpha and beta diversity metrics calculated from the normalised data using QIIME2109.
Isolation and identification of bacterial isolates
Microorganisms were isolated from the sample site using a range of media (Supplementary Table 8). The inoculum was prepared by adding 5 g of sediment to 5 ml of 2 M NaCl solution. A 1% inoculum was used to inoculate a series of dilutions (10–2–10–6), which were incubated at 4–22 °C, for 7–40 days. For isolation, the cultures were plated onto solid media (1.5% agarose) or/and semi-solid media. The individual cells were identified using sequencing of the 16S rRNA gene, using the primers and protocol described previously113. Purified PCR products were diluted to 1 ng/μL per 100 bp sequence length and sequenced using Sanger sequencing by MWG Eurofins (Germany). Chromatograms of sequences were analysed using Bioedit (7.0.5)114,115 to assess sequence quality. 16S rRNA gene sequences were analysed using the SILVA alignment, classification and tree service to identify the species to which these strains were most closely related116.
Analysis of bioavailable elements in the Colour Peak sediment
Major and trace elements in the CP waters and the bioavailable elements in the sediment were measured by Inductively Couple Plasma-Optical Emission Spectroscopy (ICP–OES) using an Agilent 5110 at the Open University. 5 g aliquots of each sediment were mixed in either 5 ml of sterilised 17% NaCl solution (to simulate the salinity of the CP spring water43) or in CP spring water. The samples were incubated at 7 °C for 7 days prior to analysis. For controls, the 17% NaCl solution was analysed in parallel. The accuracy of results was estimated using a 28 component multi-element standard solution for ICP (Fisher Chemical MS102050). The specified wavelength for each element (e.g., Ca 317.933 nm) was selected for repeatability and performance. Check standards for ppm (0.5, 1, 2.5) and ppb (10, 100, 250, 500), blanks and drifts checks were all run to ensure quality control and repeatability of data. Minimum detection limits for individual elements can be found in Table 1 and were derived from three times the standard deviation of the blanks.
Gibbs energy calculations
Gibbs energy calculations were conducted to determine the feasibility of potential metabolic reactions using Eq. (1)117:
where ΔG is the Gibbs energy of the reaction, ΔG° is the Gibbs energy of the reaction under standard conditions, R is the universal gas constant, T is the temperature in Kelvin and Q is the activity quotient. Activities were determined using the program Spec8 (Geochemist Workbench) and fugacity of atmospheric gases25,118. ΔG° values were determined using the online SUPCRT programme GEOPIG, which uses the slop07 database. ΔG can then be multiplied by the concentration of the limiting reactant (considering the stoichiometry of the reaction) to determine the potential energy available per kg of fluid. To contextualise this, cell densities were estimated using the amount of adenosine triphosphate (ATP) that could be generated from the available energy119, where 41.8 kJ is required to make 1 mol of ATP. It was assumed 10% of the available energy would be used to generate new cells119,120, 0.02 mol of ATP is required to produce 1 g of biomass and one cell has a mass of 9.50E−13 g. These calculations used brine chemistries determined from thermochemical modelling reported in Bridges and Schwenzer57. Concentrations of oxygen modelled as thermodynamically viable within the modern martian near-surface detected in the martian atmosphere and the upper and lower values of nitrates detected in ancient martian sediments by the Curiosity rover were used in these calculations25,26,67.
Data availability
Amplicon sequence data generated in this study were deposited to sequence read archives (SRA) under project number PRJNA558950, and Sanger sequence data were deposited to NCBI GenBank under accession numbers MN326776 to MN326790.
References
Fairen, A. G. et al. Astrobiology through the ages of Mars: the study of terrestrial analogues to understand the habitability of Mars. Astrobiology 10, 821–843 (2010).
Pollard, W. H., Omelon, C., Andersen, D. T. & McKay, C. P. Perennial spring occurrence in the expedition fiord area of western Axel Heiberg Island, Canadian High Arctic. Can. J. Earth Sci. 36, 105–120 (1999).
Pollard, W. et al. Overview of analogue science activities at the McGill Arctic Research Station, Axel Heiberg Island. Can. High Arctic. Planet. Sp. Sci. 57, 646–659 (2009).
Carr, M. H. & Head, J. W. Geologic history of Mars. Earth Planet. Sci. Lett. 294, 185–203 (2010).
Warner, N. et al. Late Noachian to Hesperian climate change on Mars: evidence of episodic warming from transient crater lakes near ares vallis. J. Geophys. Res. E Planets 115, 9002 (2010).
Rapin, W. et al. An interval of high salinity in ancient Gale crater. Nat. Geosci. 12, 889–895 (2019).
Wanke, H., Bruckner, J., Dreibus, G., Rieder, R. & Ryabchikov, I. Chemical composition of rocks and soils at the Pathfinder site. Space Sci. Rev. 96, 317–330 (2001).
Clark, B. C. et al. Inorganic analyses of martian surface samples at the Viking landing sites. Science. 194, 1283–1288 (1976).
Connell-Cooper, C. O., Spray, J., Thompson, L. & Berger, J. A. APXS-derived chemistry of the Bagnold dune sands: comparisons with Gale crater soils and the global martian average: APXS—Bagnold Sands and Gale Soils. J. Geophys. Res. Planets 122, 1–21 (2017).
Gellert, R. et al. Chemistry of rocks and soils in Gusev crater from the alpha particle X-ray spectrometer. Science. 305, 829–832 (2014).
Ehlmann, B. L. & Edwards, C. S. Mineralogy of the martian surface. Annu. Rev. Earth Planet. Sci. 42, 291–315 (2014).
Burgess, R., Wright, I. P. & Pillinger, C. T. Distribution of sulphides and oxidised sulphur components in SNC meteorites. Earth Planet. Sci. Lett. 93, 314–320 (1989).
Gooding, J. L. Soil mineralogy and chemistry on Mars: possible clues from salts and clays in SNC meteorites. Icarus. 1, 28–41 (1992).
Burns, R. G. & Fisher, D. S. Evolution of sulfide mineralization on Mars. J. Geophys. Res. 95, 14169–14173 (1990).
Morris, R. V. et al. Iron mineralogy and aqueous alteration from Husband Hill through Home Plate at Gusev crater, Mars: results from the Mössbauer instrument on the Spirit Mars exploration rover. J. Geophys. Res. E Planets 113, 1 (2008).
Franz, H. B. et al. Large sulfur isotope fractionations in martian sediments at Gale crater. Nat. Geosci. 10, 658–662 (2017).
Martin, P. E. et al. A two-step K-Ar experiment on Mars: dating the diagenetic formation of jarosite from Amazonian groundwaters. J. Geophys. Res. Planets 122, 2803–2818 (2017).
Lasue, J., Clifford, S. M., Conway, S. J., Mangold, N. & Butcher, F. E. The hydrology of Mars including a potential cryosphere. in Filiberto, J., Schwenzer S. P., (Eds.), Volatiles in the Martian Crust 185–246 (2019).
Orosei, R. et al. Radar evidence of subglacial liquid water on Mars. Science. 361, 448–449 (2018).
Jannasch, H. W. The chemosynthetic support of life and the microbial diversity at deep-sea hydrothermal vents. Proc. R. Soc. 225, 277–297 (1985).
Visser, J. A. N. M., Robertson, L. A. & Verseveld, H. W. V. A. N. Sulfur production by obligately chemolithoautotrophic Thiobacillus species. Appl. Environ. Microbiol. 63, 2300–2305 (1997).
Meier, D. V. et al. Niche partitioning of diverse sulfur-oxidizing bacteria at hydrothermal vents. ISME J. 11, 1545–1558 (2017).
McNichol, J. et al. Primary productivity below the seafloor at deep-sea hot springs. Proc. Natl. Acad. Sci. 115, 6756–6761 (2018).
Fike, D. A., Bradley, A. S. & Rose, C. V. Rethinking the ancient sulfur cycle. Annu. Rev. Earth Planet. Sci. 43, 593–622 (2015).
Mahaffy, P. R. et al. Abundance and isotopic compoisition of gases in the martian atmosphere from the Curiosity rover. Science. 341, 263–266 (2013).
Stern, J. C. et al. Evidence for indigenous nitrogen in sedimentary and aeolian deposits from the Curiosity rover investigations at Gale crater, Mars. Proc. Natl. Acad. Sci. 112, 4245–4250 (2015).
Jakosky, B. M. et al. Mars’ atmospheric history derived from upper-atmosphere measurements of 38Ar/36Ar. Science. 355, 1408–1410 (2017).
Schwenzer, S. P. & Kring, D. A. Impact-generated hydrothermal systems capable of forming phyllosilicates on Noachian Mars. Geology 37, 1091–1094 (2009).
Schwenzer, S. P. et al. Fluids during diagenesis and sulfate vein formation in sediments at Gale crater. Mars. Meteorit. Planet. Sci. 51, 2175–2202 (2016).
McAdam, A. C., Zolotov, M. Y., Mironenko, M. V. & Sharp, T. G. Formation of silica by low-temperature acid alteration of martian rocks: Physical-chemical constraints. J. Geophys. Res. E Planets 113, 1–8 (2008).
Martín-Torres, F. J. et al. Transient liquid water and water activity at Gale crater on Mars. Nat. Geosci. 8, 357–361 (2015).
Fox-Powell, M. G., Hallsworth, J. E., Cousins, C. R. & Cockell, C. S. Ionic strength is a barrier to the habitability of Mars. Astrobiology 16, 427–442 (2016).
McEwen, A. S. et al. Supporting material for seasonal flows on warm martian slopes. Science 333, 740–743 (2011).
Soare, R., Pollard, W. & Green, D. Deductive model proposed for evaluating terrestrial analogues. Eos. 82, 501 (2001).
Malin, M. C. & Edgett, K. S. Evidence for recent groundwater seepage and surface runoff on Mars. Science. 288, 2330–2336 (2000).
Andersen, D. T. & Pollard, W. H. Cold springs in permafrost on Earth and Mars. J. Geophys. Res. 107, 5015 (2002).
Malin, M. C., Edgett, K. S., Posiolova, L. V., Mccolley, S. M. & Dobrea, E. Z. N. Present-day impact cratering rate and contemporary gully on Mars. Science. 314, 1573–1578 (2006).
Battler, M. M., Osinski, G. R. & Banerjee, N. R. Mineralogy of saline perennial cold springs on Axel Heiberg Island, Nunavut, Canada and implications for spring deposits on Mars. Icarus 224, 364–381 (2013).
Rossi, A. P. et al. Large-scale spring deposits on Mars? J. Geophys. Res. 113, 1–17 (2008).
Brown, R. J. E. Permafrost in the Canadian Arctic archipelago. Geomorphol. Suppl. 13, 102–130 (1972).
Pollard, W. H. Icing processes associated with high Arctic perennial springs, Axel Heiberg Island, Nunavut Canada. Permafr. Periglac. Process. 16, 51–68 (2005).
Omelon, C. R., Pollard, W. H. & Andersen, D. T. A geochemical evaluation of perennial spring activity and associated mineral precipitates at Expedition Fjord, Axel Heiberg Island. Can. High Arctic. Appl. Geochem. 21, 1–15 (2006).
Fox-Powell, M. G. et al. Natural analogue constraints on Europa’s non-ice surface material. Geophys. Res. Lett. 46, 5759–5767 (2019).
Niederberger, T. D. et al. Novel sulfur-oxidizing streamers thriving in perennial cold saline springs of the Canadian High Arctic. Environ. Microbiol. 11, 616–629 (2009).
Perreault, N. N., Andersen, D. T., Pollard, W. H., Greer, C. W. & Whyte, L. G. Characterization of the prokaryotic diversity in cold saline perennial springs of the Canadian High Arctic. Appl. Environ. Microbiol. 73, 1532–1543 (2007).
Eckardt, F. D., Bryant, R. G., McCulloch, G., Spiro, B. & Wood, W. W. The hydrochemistry of a semi-arid pan basin case study: Sua Pan, Makgadikgadi, Botswana. Appl. Geochem. 23, 1563–1580 (2008).
Moiseenko, T. I., Gashkina, N. A., Dinu, M. I., Kremleva, T. A. & Khoroshavin, V. Y. Aquatic geochemistry of small lakes: effects of environment changes. Geochem. Int. 51, 1 (2013).
Minissale, A., National, I., Vaselli, O. & Chandrasekharam, D. Origin and evolution of `intracratonic’ thermal fluids from central-western peninsular India. Earth Planet. Sci. Lett. 181, 377–394 (2000).
Das, A., Rishnaswami, S. K., Arin, M. M. S. & Ande, K. P. Chemical weathering in the Krishna Basin and Western Ghats of the Deccan Traps India. Geochim. Cosmochim. Acta 69, 2067–2084 (2005).
Bullock, M. A., Moore, J. M. & Mellon, M. T. Laboratory simulations of Mars aqueous geochemistry. Icarus 170, 404–423 (2004).
Catling, D. C. A chemical model for evaporites on early Mars’ possible sedimentary tracers of the early climate and implications for exploration. J. Geophys. Res. 104, 16453–16469 (1999).
Moore, J. M. & Bullock, A. Experimental studies of Mars-analog brines. J. Geophys. Res. 104, 21925–21934 (1999).
Schwenzer, S. P. & Kring, D. A. Alteration minerals in impact-generated hydrothermal systems: exploring host rock variability. Icarus 226, 487–496 (2013).
Filiberto, J. et al. A review of volatiles in the martian interior. Meteorit. Planet. Sci. 1958, 1935–1958 (2016).
Melwani Daswani, M., Schwenzer, S. P., Reed, M. H., Wright, I. P. & Grady, M. M. Alteration minerals, fluids, and gases on early Mars: predictions from 1-D flow geochemical modeling of mineral assemblages in meteorite ALH 84001. Meteorit Planet Sci. 2174, 2154–2174 (2016).
Tosca, N. J., Mclennan, S. M., Lamb, M. P. & Grotzinger, J. P. Physicochemical properties of concentrated martian surface waters. J. Geophys. Res. 116, 1–16 (2011).
Bridges, J. C. & Schwenzer, S. P. The nakhlite hydrothermal brine on Mars. Earth Planet. Sci. Lett. 359–360, 117–123 (2012).
Niederberger, T. D. et al. Microbial characterization of a subzero, hypersaline methane seep in the Canadian High Arctic. ISME J. 4, 1326–1339 (2010).
Lay, C. Y. et al. Microbial diversity and activity in hypersaline High Arctic spring channels. Extremophiles 16, 177–191 (2012).
Lay, C. Y. et al. Defining the functional potential and active community members of a sediment microbial community in a high-arctic hypersaline subzero spring. Appl. Environ. Microbiol. 79, 3637–3648 (2013).
Lamarche-Gagnon, G., Comery, R., Greer, C. W. & Whyte, L. G. Evidence of in situ microbial activity and sulphidogenesis in perennially sub-0 °C and hypersaline sediments of a high Arctic permafrost spring. Extremophiles 19, 1–15 (2015).
Sapers, H. M. et al. Biological characterization of microenvironments in a hypersaline cold spring Mars analog. Front. Microbiol. 8, 2527 (2017).
Willerslev, E., Hansen, A. J. & Poinar, H. N. Isolation of nucleic acids and cultures from fossil ice and permafrost. Trends Ecol. Evol. 19, 141–147 (2004).
Willerslev, E. et al. Long-term persistence of bacterial DNA. Curr. Biol. 14, 13–14 (2004).
Pietramellara, G. et al. Extracellular DNA in soil and sediment: Fate and ecological relevance. Biol. Fertil. Soils 45, 219–235 (2009).
Li, R. et al. Comparison of DNA-, PMA-, and RNA-based 16S rRNA Illumina sequencing for detection of live bacteria in water. Sci. Rep. 7, 1–11 (2017).
Stamenković, V., Ward, L. M., Mischna, M. & Fischer, W. W. O2 solubility in martian near-surface environments and implications for aerobic life. Nat. Geosci. 11, 905–909 (2018).
Ventosa, A., Nieto, J. J. & Oren, A. Biology of moderately halophilic aerobic bacteria. Microbiol. Mol. Biol. Rev. 62, 504–544 (1998).
Kelly, D. P. Halothiobacillus. Bergey’s Man Syst. Archaea Bact. 1, 1–3. https://doi.org/10.1002/9781118960608.gbm01133 (2015).
Brinkhoff, T., Kuever, J., Muyzer, G. & Jannasch, H. W. Thiomicrospira. Bergey’s Man Syst. Archaea Bact 1, 1–10. https://doi.org/10.1002/9781118960608.gbm01221 (2015).
Kelly, D. P., Wood, A. P. & Stackebrandt, E. Thiobacillus. Bergey’s Man. Syst. Archaea Bact 1, 1–10. https://doi.org/10.1002/9781118960608.gbm00969 (2015).
Choi, B.-R. et al. Characterization of facultative sulfur-oxidizing Marinobacter sp. BR13 isolated from marine sediment of Yellow Sea, Korea. J. Korean Soc. Appl. Biol. Chem. 52, 309–314 (2009).
Sorokin, D. Y. Oxidation of inorganic sulfur compounds by obligately organotrophic bacteria. Microbiology 72, 641–653 (2003).
Bridges, J. C., Hicks, L. J. & Treiman, A. H. Carbonates on Mars. in Filiberto, J. Schwenzer S. P., (Eds.), Volatiles in the Martian Crust, pp. 89–118 (2019).
Franz, H. B., King, P. L. & Gaillard, F. Sulfur on Mars from the atmosphere to the core. in Filiberto, J. Schwenzer S. P., (Eds.), Volatiles in the Martian Crust, pp. 119–184 (2019).
Grotzinger, J. P. et al. A habitable fluvio-lacustrine environment at Yellowknife Bay, Gale Crater Mars. Science. 343, 1242777 (2014).
Fastook, J. L., Head, J. W., Marchant, D. R., Forget, F. & Madeleine, J. B. Early Mars climate near the Noachian-Hesperian boundary: Independent evidence for cold conditions from basal melting of the south polar ice sheet (Dorsa Argentea Formation) and implications for valley network formation. Icarus 219, 25–40 (2012).
Weiss, D. K. & Head, J. W. Crater degradation in the Noachian highlands of Mars: assessing the hypothesis of regional snow and ice deposits on a cold and icy early Mars. Planet. Space Sci. 117, 401–420 (2015).
Abramov, O. & Mojzsis, S. J. Thermal effects of impact bombardments on Noachian Mars. Earth Planet. Sci. Lett. 442, 108–120 (2016).
Fairén, A. G. A cold and wet Mars. Icarus 208, 165–175 (2010).
Perreault, N. N. et al. Heterotrophic and autotrophic microbial populations in cold perennial springs of the High Arctic. Appl. Environ. Microbiol. 74, 6898–6907 (2008).
Liu, C. et al. Marinobacter antarcticus sp. nov., a halotolerant bacterium isolated from Antarctic intertidal sandy sediment. Int. J. Syst. Evol. Microbiol. 62, 1838–1844 (2012).
Moghadam, M. S. et al. Isolation and genome sequencing of four Arctic marine Psychrobacter strains exhibiting multicopper oxidase activity. BMC Genomics 17, 1–14 (2016).
Bakermans, C. et al. Psychrobacter cryohalolentis sp. nov. and Psychrobacter arcticus sp. nov., isolated from Siberian permafrost. Int. J. Syst. Evol. Microbiol. 56, 1285–1291 (2006).
Antwis, R. E. et al. Fifty important research questions in microbial ecology. FEMS Microbiol. Ecol. 93, 1 (2017).
Sogin, M. L. et al. Microbial diversity in the deep sea and the underexplored ‘rare biosphere’. Proc. Natl. Acad. Sci. 103, 12115–12120 (2011).
Larsen, S., Nielsen, L. P. & Schramm, A. Cable bacteria associated with long-distance electron transport in New England salt marsh sediment. Environ. Microbiol. Rep. 7, 175–179 (2015).
Huber, B., Herzog, B., Drewes, J. E., Koch, K. & Müller, E. Characterization of sulfur oxidizing bacteria related to biogenic sulfuric acid corrosion in sludge digesters. BMC Microbiol. 1, 1–11. https://doi.org/10.1186/s12866-016-0767-7 (2016).
Krishnakumar, B. & Manilal, V. B. Bacterial oxidation of sulphide under denitrifying conditions. Biotechnol. Lett. 21, 437–440 (1999).
Price, A., Pearson, V. K., Schwenzer, S. P., Miot, J. & Olsson-Francis, K. Nitrate-dependent iron oxidation: a potential Mars metabolism. Front. Microbiol. 9, 1–15 (2018).
Blumenberg, M., Seifert, R., Petersen, S. & Michaelis, W. Biosignatures present in a hydrothermal massive sulfide from the Mid-Atlantic Ridge. Geobiology 5, 435–450 (2007).
Banfield, J. F., Moreau, J. W., Chan, C. S., Welch, S. A. & Little, B. Search for life on Mars. Astrobiology 1, 448–465 (2001).
Douglas, S. Mineralogical footprints of microbial life. Am. J. Sci. 305, 503–525 (2015).
Glamoclija, M. et al. Biosignatures and bacterial diversity in hydrothermal deposits of Solfatara Crater, Italy. Geomicrobiol. J. 21, 529–541 (2010).
Chan, C. S., Fakra, S. C., Emerson, D., Fleming, E. J. & Edwards, K. J. Lithotrophic iron-oxidizing bacteria produce organic stalks to control mineral growth: implications for biosignature formation. ISME J. 5, 717–727 (2010).
Taylor, C. D., Wirsen, C. O. & Gaill, F. Rapid microbial production of filamentous sulfur mats at hydrothermal vents. Appl. Environ. Microbiol. 65, 2253–2255 (1999).
Engel, A. S., Lichtenberg, H., Prange, A. & Hormes, J. Speciation of sulfur from filamentous microbial mats from sulfidic cave springs using X-ray absorption near-edge spectroscopy. FEMS Microbiol. Lett. 269, 54–62 (2007).
Seager, S., Schrenk, M. & Bains, W. An astrophysical view of Earth-based metabolic biosignature gases. Astrobiology 12, 62–82 (2012).
Pellerin, A. et al. Large sulfur isotope fractionation by bacterial sulfide oxidation. Sci. Adv. 5, 1–7 (2019).
Dobson, V. P., Vreeland, R. H. & Chester, W. Halomonas. Bergey’s Man. Syst. Archaea Bact. https://doi.org/10.1002/9781118960608.gbm01190 (2015).
Sutter, B. et al. The sample analysis at Mars (SAM) detections of CO2 and CO insedimentary material from Gale crater, Mars: implications for the presence of organic carbon and microbial habitability on Mars. in Proceedings of the AGU Fall Meeting (2016).
Chatzigiannidou, I., Props, R. & Boon, N. Drinking water bacterial communities exhibit specific and selective necrotrophic growth. NPJ Clean Water 1, 22 (2018).
An, D. et al. Metagenomics of hydrocarbon resource environments indicates aerobic taxa and genes to be unexpectedly common. Environ. Sci. Technol. 47, 10708–10717 (2013).
Schmidt, O., Hink, L., Horn, M. A. & Drake, H. L. Peat: home to novel syntrophic species that feed acetate- and hydrogen-scavenging methanogens. ISME J 1, 1–13. https://doi.org/10.1038/ismej.2015.256 (2016).
Timmers, P. H. A. et al. Metabolism and occurrence of methanogenic and sulfate-reducing syntrophic acetate oxidizing communities in haloalkaline environments. Front. Microbiol. 9, 1–18 (2018).
Tillett, D. & Neilan, B. A. Xanthogenate nucleic acid isolation from cultured and environmental cyanobacteria. J. Phycol. 36, 251–258 (2000).
Green, M. R. & Sambrook, J. Precipitation of DNA with ethanol. Cold Spring Harb. Protoc. 2016, 1116–1120 (2016).
Walters, W. A. et al. Improved bacterial 16S rRNA gene (V4 and V4–5) and fungal internal transcribed spacer marker gene primers for microbial community surveys. mSystems 1, 1–10 (2012).
Boylen, E. et al. QIIME 2: reproducible, interactive, scalable, and extensible microbiome data science. PeerJ https://doi.org/10.7287/peerj.preprints.27295 (2018).
DeSantis, T. Z. et al. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl. Environ. Microbiol. 72, 5069–5072 (2006).
McDonald, D. et al. An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. ISME J. 6, 610–618 (2012).
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: Improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Lane, D.J. 16S/23S rRNA Sequencing. in Nucleic Acid Techniques in Bacterial Systematics (ed. Stackebrandt, E. and Goodfellow, M.) 115–175 (John Wiley and Sons, 1991).
Hall, A. T. BioEdit: an important software for molecular biology. GERF Bull. Biosci. 2, 60–61 (2011).
Hall, T. A. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp. Ser. No. 41, 95–98 (1999).
Pruesse, E., Peplies, J. & Glöckner, F.O. SINA: accurate high-throughput multiple sequence alignment of ribosomal RNA genes. Bioinformatics 28, 1823–1829 (2012).
McCollom, T. M. Geochemical constraints on sources of metabolic energy for chemolithoautotrophy in ultramafic-hosted deep-sea hydrothermal systems. Astrobiology 7, 933–950 (2007).
Nier, A. & Mcelroy, M. B. Structure of the neutral upper atmosphere of Mars. Science. 194, 28–30 (1976).
Jakosky, B. M. & Shock, E. L. The biological potential of Mars, the early Earth, and Europa. J. Geophys. Res. E Planets 103, 19359–19364 (1998).
McCollom, T. M. & Amend, J. P. A thermodynamic assessment of energy requirements for biomass synthesis by chemolithoautotrophic micro-organisms in oxic and anoxic environments. Geobiology 3, 135–144 (2005).
Kaye, J. Z., Márquez, M. C., Ventosa, A. & Baross, J. A. Halomonas neptunia sp. nov., Halomonas sulfidaeris sp. nov., Halomonas axialensis sp. nov. and Halomonas hydrothermalis sp. nov.: halophilic bacteria isolated from deep-sea hydrothermal-vent environments. Int. J. Syst. Evol. Microbiol. 54, 499–511 (2004).
Lee, J. C. et al. Halomonas taeanensis sp. nov., a novel moderately halophilic bacterium isolated from a solar saltern in Korea. Int. J. Syst. Evol. Microbiol. 55, 2027–2032 (2005).
Yumoto, I. et al. Psychrobacter piscatorii sp. nov., a psychrotolerant bacterium exhibiting high catalase activity isolated from an oxidative environment. Int. J. Syst. Evol. Microbiol. 60, 205–208 (2010).
VanTrappen, S., Mergaert, J. & Swings, J. Loktanella salsilacus gen. nov., sp. Nov., Loktanella fryxellensis sp. Nov. and Loktanella vestfoldensis sp. Nov., new members of the Rhodobacter group isolated from microbial mats in Antarctic lakes. Int. J. Syst. Evol. Microbiol. 54, 1263–1269 (2004).
Hahnke, R. L., Meier-kolthoff, J. P. & García-lópez, M. Genome-based taxonomic classification of Bacteroidetes. Front. Microbiol. 7, 1–37 (2016).
Godoy, F. A., Swings, J. & Rehm, B. Sphingopyxis chilensis sp. Nov., a chlorophenol-degrading bacterium that accumulates polyhydroxyalkanoate, and transfer of Sphingomonas alaskensis to Sphingopyxis alaskensis comb. Nov.. Int J. Syst. Evol. Microbiol. 53, 473–477 (2003).
Yu, Y. et al. Sporosarcina antarctica sp. Nov., a psychrophilic bacterium isolated from the Antarctic. Int. J. Syst. Evol. Microbiol. 3, 2114–2117 (2008).
Satola, B., Wübbeler, J. H. & Steinbüchel, A. Metabolic characteristics of the species Variovorax paradoxus. Appl. Microbiol. Biotechnol. 97, 541–560 (2013).
Morohoshi, T. et al. Biofilm formation and degradation of commercially available biodegradable plastic films by bacterial consortiums in freshwater environments. Microbes Environ. 33, 332–335 (2018).
Glockner, F. O., Babenzien, H. & Amann, R. Phylogeny and identification in situ of Nevskia ramosa. Appl. Environ. Microbiol. 64, 1895–1901 (1998).
Sorokin, D. Y. et al. Discovery of extremely halophilic, methyl-reducing euryarchaea provides insights into the evolutionary origin of methanogenesis. Nat. Microbiol. 2, 1 (2017).
Reasoner, D. J. et al. A new medium for the enumeration and subculture of Bacteria from potable water. Appl. Environ. Microbiol. 49, 1–7 (1985).
Bertani, G. Studies on lysogenesis. I. The mode of phage liberation by lysogenic Escherichia coli. J. Bacteriol. 62, 293–300 (1951).
Macey, M. C., Pratscher, J., Crombie, A. & Murrell, J. C. Draft genome sequences of obligate methylotrophs Methylovorus sp. strain MM2 and Methylobacillus sp. Strain MM3, isolated from grassland soil. Microbiol. Resour. Announc. 7, 1–2 (2018).
Sievert, S. M., Heidorn, T. & Kuever, J. Halothiobacillus kellyi sp. Nov., a mesophilic, obligately chemolithoautotrophic, sulfur-oxidizing bacterium isolated from a shallow-water hydrothermal vent in the Aegean Sea, and emended description of the genus Halothiobacillus. Int. J. Syst. Evol. Microbiol. 50, 1229–1237 (2000).
Hobbs, G., Frazer, C. M., Gardner, D. C. J., Cullum, J. A. & Oliver, S. G. Dispersed growth of Streptomyces in liquid culture. Appl. Microbiol. Biotechnol. 31, 272–277 (1989).
Acknowledgements
We would like to acknowledge Hugo Moors from the Belgian Nuclear Research Center for his advice on handling nucleic acids extracted from saline environments. We would like to thank Gordon Osinski from Western University, Ontario for leading the sampling trip to Axel Heiberg island in 2017. We would like to thank Bea Baharier from the Open University for assisting with the Spec8 models. We would like to acknowledge funding from the Science and Technology Facilities Council from the Grant ST/P000657/1. We would also like to acknowledge funding from a Leverhulme Trust Research Project Grant (RPG-2016-153) and thank the Polar Continental Shelf Program (Natural Resources Canada) for logistical field support in Nunavut. This article was funded by Science and Technology Facilities Council (Grant no. ST/P000657/1) and Leverhulme Trust (Grant nos. RPG-2016-153, RPG-2016-200).
Author information
Authors and Affiliations
Contributions
M.F.P. collected the environmental materials from Axel Heiberg Island. M.C.M. and B.P.S. performed the molecular work. T.B. performed the ICP–OES work. M.C.M., K.O.F., M.F.P. and C.R.C. devised the series of experiments. N.K.R. performed the speciation models and Gibbs energy calculations. S.P.S. produced Fig. 2 and 3. M.C.M., K.O.F., N.K.R., V.K.P. and S.P.S. performed analysis and interpretation of the data. All authors contributed to the manuscript and approved the final version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Macey, M.C., Fox-Powell, M., Ramkissoon, N.K. et al. The identification of sulfide oxidation as a potential metabolism driving primary production on late Noachian Mars. Sci Rep 10, 10941 (2020). https://doi.org/10.1038/s41598-020-67815-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-67815-8
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.