Abstract
Animals may host diverse bacterial communities that can markedly affect their behavioral physiology, ecology, and vulnerability to disease. Fungus-farming ants represent a classical example of mutualism that depends on symbiotic microorganisms. Unraveling the bacterial communities associated with fungus-farming ants is essential to understand the role of these microorganisms in the ant-fungus symbiosis. The bacterial community structure of five species of fungus-farmers (non-leaf-cutters; genera Mycocepurus, Mycetarotes, Mycetophylax, and Sericomyrmex) from three different environments in the Brazilian Atlantic rainforest (lowland forest, restinga forest, and sand dunes) was characterized with amplicon-based Illumina sequencing of 16 S ribosomal RNA gene. Possible differences in bacterial communities between ants internal to the nest (on the fungus garden) and external foragers were also investigated. Our results on the richness and diversity of associated bacteria provide novel evidence that these communities are host- and colony-specific in fungus-farming ants. Indeed, the bacterial communities associated with external foragers differ among the five species, and among colonies of the same species. Furthermore, bacterial communities from internal ants vs. foragers do not differ or differ only slightly within each ant species. This study highlights the importance of describing ant-associated bacterial communities to better understand this host-bacterial interaction in the social environment of insect colonies and provides the foundation for future studies on the ecological and evolutionary processes that drive the success of fungus-farming ants.
Similar content being viewed by others
Introduction
Symbiotic microorganisms play crucial roles in shaping the phenotypes, ecology, and evolution of their hosts1,2. Advances in DNA sequencing techniques and culture-independent genomic tools have resulted in accurate estimates of microbial richness and diversity from host and environmental samples, revealing an abundant and diverse microbiota living in symbiosis with animals1,3. These microorganisms may interact with hosts in many ways, such as preventing pathogenic infections, increasing the host’s ability to cope with stressful environments, or even degrading or synthesizing substances that are important for host nutrition2,4,5. According with the “holobiont” concept, the biological entity on which selection acts is the host and its associated microorganisms together as a unit2.
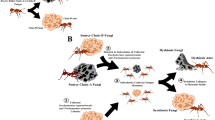
Fungus-farming ants (Formicidae, Myrmicinae, Attina; hereafter referred to as attine ants) are a good example of the tight links between symbiotic microorganisms and host evolutionary success. These ants live in a complex and highly specialized multi-trophic symbiosis6. As an example of obligate mutualism, ants obtain food for the colony by cultivating certain fungal species in their nests, and in return the ants provide the fungus with nourishment, dispersal to new locations, and an environment free of parasites and competitors, such as parasitic fungi from the genus Escovopsis7. Attine ants are divided into two clades, the Paleoattina and the Neoattina8. The latter includes the highly derived attine genera Acromyrmex and Atta, known as “leaf-cutter ants” due their behavior of cutting fresh leaves as the substrate on which they cultivate the fungus garden6. Acromyrmex and Atta have the most derived characteristics within fungus-growers and many studies investigate the bacterial communities associated with these ants9,10,11.
The bacterial community in other attine genera from Paleoattina and Neottina (non-leaf-cutter ants) is poorly known12. Some bacteria present on the ant body surfaces and within the fungus garden are believed to aid in managing the fungiculture and avoiding infection by Escovopsis13,14,15,16, although little is known about their interactions within the nest environment. For instance, these bacteria could be nutritionally important12, play a defensive role against pathogens by producing antibiotic compounds16,17, or even contribute to nest hygiene18. In addition, the fungus garden is constantly exposed to external bacteria that are brought to the nest by ant foragers and/or with items they collect as substrate for fungiculture18, but the influence of this continuous exchange on the nest environment is unknown. Comprehensive studies of these ant- and nest-associated bacterial communities are important to reveal the bacterial community composition of fungus-farming ants and their nests, as a first step in understanding the role of these bacteria in ant-fungus symbiosis.
In this study, we report on the diversity of the bacterial communities associated with five species of fungus-farming ants (non-leaf-cutters) from three different areas in the Atlantic rainforest of Brazil: Mycocepurus smithii (Forel, 1893), Mycetarotes parallelus (Emery, 1906), Mycetophylax morschi (Emery, 1888), Sericomyrmex parvulus (Forel, 1912) and Sericomyrmex saussurei (Emery, 1894). We quantified bacterial species richness and diversity using 16 S ribosomal RNA gene amplicon sequencing. Our objectives were: (i) to characterize the bacterial community composition from five attine species; (ii) to compare the microbiome of the five attine species and investigate possible interspecific differences in bacterial community composition; (iii) to investigate if the microbiota associated with ants whose activity is internal to the nest environment differs from that associated with ants foraging outside the nest; and (iv) to compare the microbiome between different colonies of the same species and investigate possible intraspecific differences in the composition of associated bacterial communities. Although the five ant species investigated cultivate mutualistic fungi, they differ in the size of their colonies, in the material collected for fungiculture, and in the foraging modes19. Thus, we predicted that bacterial communities would differ among the five species of fungus-farming ants. Because the environment outside the nest has several sources for transient bacteria (e.g., soil, plants, other arthropods, material collected for fungiculture), we expected that ant foragers would have different bacterial composition when compared to ants working inside the nest. Our results provide novel evidence that ant-associated bacteria are host- and colony-specific in fungus-farming ants. Bacterial communities associated with external foragers differ among the five species, and among colonies of the same species, suggesting that environment outside the nest has little influence on the associated bacterial community of fungus-farming ants.
Material and methods
Samplings
We carried out fieldwork in July 2015 and January 2016 in the Atlantic rainforest at the ‘Parque Estadual Serra do Mar’, Ubatuba municipality, São Paulo State, Brazil (Fig. S1 Supplementary Material 1). We collected workers of five species of fungus-farming ants: Mycocepurus smithii, Mycetarotes parallelus, Mycetophylax morschi, Sericomyrmex parvulus and Sericomyrmex saussurei. Sampled colonies were located in three different areas: (i) S. parvulus and S. saussurei were found in lowland forest (23° 21’ 52.1” S, 44° 49’ 28.5” W), characterized by clay-sandy soils, epiphytes, and trees reaching more than 20 m20; (ii) Mycocepurus smithii, Mycetarotes parallelus and Mycetophylax morschi were found in restinga forest (23° 21’ 28.4” S, 44° 51’ 00.7” W) growing in sandy soils and with trees up to 20 m tall20; (iii) Mycetophylax morschi was also found in the dunes region (23° 21’ 38.8” S, 44° 50’ 58.1” W), this area has a xerophytic vegetation and are under the influence of sea salinity20. Information about the weather of the area is provided in Fig. S1 legend (Supplementary Material 1).
To investigate if bacterial communities differed between foragers in the external environment and ants working inside the nest, for Mycocepurus smithii, Mycetarotes parallelus and Mycetophylax morschi, we collected 16 foraging workers upon their return to the nest and 16 workers from the fungus garden inside the nest. Due to difficulties in finding the nest chambers of Sericomyrmex, only foragers were collected for S. parvulus and S. saussurei. Workers internal to the nest were collected directly from gardens in the field, upon excavation of the colonies. Previous work21 with captive colonies confirmed intracolonial division of labor in these five species. Hence, at any given time, workers performing foraging activities do not sequentially engage in work on the fungus garden, and vice-versa. Thus, in this study we consider two non-mixing worker groups: those collected on the fungus garden and those collected as outside foragers (collected upon their return to the nest).
The following colonies of ants were sampled: three colonies of Mycocepurus smithii, Mycetarotes parallelus, and Mycetophylax morschi (restinga forest); three colonies of S. parvulus (lowland forest); two colonies of Mycetophylax morschi (dune area); and four colonies S. saussurei (lowland forest). All equipment used for collecting ants was sterilized with ethanol 70% and flamed to avoid sample contamination. Ants were stored in separate sterile microcentrifuge vials, in a −20 °C freezer. Since generic names begin with the same letters in the species Mycocepurus smithii, Mycetarotes parallelus and Mycetophylax morschi, the names of these species will be cited in full.
Molecular methods
We extracted Genomic DNA from whole individual ants using the Qiagen DNeasy Blood & Tissue kit, following the manufacturer’s protocol with minor adjustments to increase DNA yield from the bacterial community. Due to the small body length of workers (<5 mm), we extracted DNA from the whole individual4,9,12,16. Ants were not macerated before being placed in AL buffer and incubated at 56 °C with proteinase K for 2 hours to minimize extraction of ant DNA. For the bacterial community profiling, we PCR-amplified the V4 region of the bacterial 16 S rRNA gene using a dual-index approach22 with barcoded-primers 515 F and 806 R23. PCR reactions were performed in duplicates using conditions described in Bletz et al.24, but with Phire® Hot Start II DNA Polymerase (Finnzyme, Espoo, Finland). Negative controls for extraction and each PCR mix were included to check for contamination. PCR products of each sample were combined; samples were pooled and purified using a DNA Gel Extraction Kit (Norgen Biotek Corp, Thorold, ON, Canada). The purified pool was sequenced with paired-end 250 on an Illumina MiSeq sequencer at TUCF Genomics, Boston, MA, USA, using the V4-specific primer sequences and linkers22.
To confirm the identity of the sampled ants of each colony, we sequenced a fragment of the mitochondrial cytochrome c oxidase subunit 1 (COI) using primers LCO1490 and HC0219825 and standard PCR conditions. PCR fragments were purified and sequenced in Macrogen Inc, Seoul, South Korea. Sequences were trimmed and quality-checked using Geneious R6 (http://www.geneious.com 26) and submitted to GenBank (accession numbers MH206536- MH206586). We obtained good sequences from ants from all colonies and found almost no variation between colonies and within species. Mycocepurus smithii, Mycetarotes parallelus and S. parvulus presented only one haplotype in all colonies, and Mycetophylax morschi and S. saussurei presented two haplotypes that differ in only two base pairs. We found no variation between individuals from the same colony.
Sequence analyses
The sequences were first demultiplexed and converted in fastq format using Illumina’s bcl2fastq conversion software (Illumina, San Diego, CA, USA). Then we processed the reads and analyzed the data using QIIME 2 2020.227 on Ubuntu 18.04.4 LTS. We imported the demultiplexed read pairs into QIIME 2, merged pair-ends with VSEARCH28, quality filtered using q-score-joined, and used Deblur29 to denoise data with a trimming length of 250 nt. OTUs comprising less than 10 reads in total were filtered out. Sequences were aligned with mafft30 and the FastTree231 was used to build a phylogenetic tree of OTUs using QIIME 2 standard procedures. We assigned taxonomy to OTUs using the q2-feature-classifier classify-sklearn32 against the Greengenes 13.8, with 99% similarity with the OTUs reference sequences (from 515 F/806 R region of the sequences33). We rarefied all samples to 2000 reads. After filtering, 180 samples remained for analysis out of 243. See Table S1 (Supplementary Material 1) for an overview of sample sizes and numbers of sequences for different subsets of data used in the analysis. One extraction control was retained after quality filtering and rarefaction, however, this control was different from all samples of the dataset and was filtered out from diversity analyses. We also numbered the OTUs that were different, but that had the same classification until taxonomic level of genus (e.g., Pseudonocardia OTU 1 and Pseudonocardia OTU 2 are different OTUs, but from the same genus).
Statistical analyses
We first performed descriptive analyses of bacterial community structure of the five species of fungus-farming ants. We represented by Venn diagrams (generated by Bioinformatics & Evolutionary Genomics34) the OTUs that were shared between all species, and between ants internal and external to the nest. We also constructed a heatmap to indicate the variation of relative abundance of bacterial OTUs in all species, discriminating the external and internal ants. The heatmap was generated in R version 3.3.335 with the package gplots (function heatmap.2), and the dendrograms were generated with Bray-Curtis distance matrices (vegan package, function vegdist). We removed the OTUs with less than 5% relative abundance.
All statistical analysis of alpha and beta diversity was performed in R version 3.3.335. We used QIIME 2 to calculate the number of observed OTUs, Chao1 and Faith’s Phylogenetic Diversity for all samples as a proxy for bacterial richness and diversity. We used linear mixed-effects models (nlme package, function lme) to investigate the variation in Faith’s Phylogenetic Diversity in OTUs from all species from which we sampled foragers external to the nest; we included species as the main explanatory variable and location as a random factor. For species from which we sampled ants internal and external to the nest (Mycocepurus smithii, Mycetarotes parallelus and Mycetophylax morschi), we included species and internal/external to the nest as main explanatory variables, and location as a random factor. Since Mycetophylax morschi was found in both the restinga forest and in the dunes area, ants from each site were treated separately. We performed pairwise comparisons between all species from which we sampled worker foragers external to the nest (stats package, function pairwise.t.test), and between internal and external ants of each species (lsmeans package, function lsmeans).
For beta-diversity, we used PERMANOVAs with package vegan (function adonis) to quantify the composition similarities between bacterial communities within groups of individuals using abundance data with Bray-Curtis distances. We also conducted this analysis using the unweighted and weighted UniFrac metrics36 for interspecific comparisons and between ants working inside the nest and foragers. For interspecific comparisons, we considered all species from which we sampled worker foragers external to the nest, with species as the main explanatory variable. We separated Mycetophylax morschi data for the restinga forest and the dune area. For comparisons between ants working inside the nest and foragers, we considered only species with internal and external ant samples (Mycocepurus smithii, Mycetarotes parallelus and Mycetophylax morschi from both the restinga forest and the dune area), with species and internal/external to the nest as main explanatory variables. We performed pairwise comparisons with the Bray-Curtis similarity method and Bonferroni adjustment (package EcolUtils, function adonis.pair) between all species for which we sampled worker foragers external to the nest, and between internal and external ants of each species. For intraspecific comparisons, we performed PERMANOVAs for each species, considering the colony as the main explanatory variable. In this analysis, only Mycetophylax morschi from the restinga forest was considered. We used principal coordinate analysis (PCoA) to visualize the bacterial community composition using abundance data with Bray-Curtis distances, with ape package (function pcoa).
Additionally, we identified the OTUs responsible for the patterns in bacterial community internal and external to the nest of each species using similarity percentage analysis (SIMPER), based on Bray-Curtis distances with vegan package (function simper). SIMPER showed the contribution of each OTU to the dissimilarity between the internal and external communities of each species. This analysis was calculated only for species that showed differences in composition of bacterial communities between ants internal and external to the nest environment.
Ethics approval
Samples were collected under permit by SISBIO #45317-3 and this research was registered in the SISGen platform (SISGEN #A210679). All applicable institutional and/or national guidelines for care and use of animals were followed.
Results
General patterns of OTU diversity
The analyzed data included 7,195,350 bacterial sequences from 180 samples of five species of attine ants: Mycetarotes parallelus, Mycetophylax morschi, Mycocepurus smithii, Sericomyrmex parvulus and Sericomyrmex saussurei. Sequences represented 874 OTUs dominated by members of the phyla Actinobacteria, Proteobacteria, Bacteroidetes, and Tenericutes (Fig. 1).
The bacterial taxonomic composition of Mycetarotes parallelus consisted predominantly of Actinobacteria (78.7% ants internal to nest; 75.2% external foragers). External foragers of Mycocepurus smithii were hosted mainly of Bacteroidetes (52.8%), whereas individuals internal to the nest had mainly Proteobacteria (36.4%) and Actinobacteria (53.7%) (Fig. 1). Mycetophylax morschi from restinga forest had predominantly Actinobacteria (88% internal; 70.2% external) and from the dune area had Proteobacteria (72.1% internal; 76.7% external). Tenericutes (31.8%) and Proteobacteria (38.4%) were the most abundant phyla in S. parvulus external foragers, while external ants of S. saussurei hosted mainly Proteobacteria (39.5%) and Actinobacteria (38.3%) (Fig. 1). The phyla Actinobacteria and Proteobacteria were present in the five ant species, both in ants inside the nest and external foragers.
Venn diagrams between the five fungus-farming ant species from which we sampled external foragers showed that 88 OTUs were common among these species (Fig. S2 Supplementary Material 1). Comparisons of the bacterial composition between ants in the internal and external nest environment, for each ant species, showed that in Mycocepurus smithii internal and external ants shared 35.7% of the total bacterial OTUs, Mycetarotes parallelus shared 44% OTUs, and Mycetophylax morschi from the restinga forest and dune area, 29.4% and 32.1% OTUs, respectively (Fig. S3 Supplementary Material 1).
The average relative abundance of bacterial OTUs in external foragers showed that different ant species were dominated by different genera of bacteria (Fig. 2). For instance, in Mycetarotes parallelus more than 50% of the bacterial community consisted of Pseudonocardia OTU 2 (Fig. 2). Pseudonocardia OTU 2, Chryseobacterium and Luteimonas were the main OTUs found in Mycocepurus smithii (Fig. 2). Workers of Mycetophylax morschi in the internal and external nest environment from the restinga forest and dune area showed that OTUs with higher average relative abundance were only found in this species. Finally, Sericomyrmex parvulus was dominated by Entomoplasmatales and S. saussurei by Actinomycetales OTU 2.
Heatmap indicating variation in relative abundance of different bacteria in the five species of fungus-farming ants from which external foragers and internal ants (on the fungus garden) were sampled in the Atlantic rainforest of Brazil. Colors indicate the relative abundance of bacterial OTUs, ranging from 0% (light yellow) to 100% (dark blue). Dendrograms were generated with Bray-Curtis distance matrices. For clarity, we removed the OTUs with less than 5% relative abundance.
Bacterial alpha-diversity
Within the five species for which we sampled worker foragers external to the nest, the diversity of bacterial communities of ants was higher in S. saussurei (followed by S. parvulus) as seen by richness (number of OTUs), Chao 1 diversity index and Faith’s Phylogenetic Diversity index (Table 1). For workers sampled inside the nest, Mycetarotes parallelus had the most diverse bacterial community, with more OTUs per sample, and higher values of Chao 1 and Faith’s Phylogenetic Diversity (Table 1).
Faith’s Phylogenetic Diversity differed between ant species for which we sampled worker foragers external to the nest (F = 2.81, df = 5, P = 0.019; Fig. 3a). Sericomyrmex saussurei differed from all other species and S. parvulus differed from Mycetophylax morschi (dune and restinga) and Mycocepurus smithii (Fig. 3a). In species for which we sampled individuals internal and external to the nest, Faith’s Phylogenetic Diversity differed between internal and external workers only in Mycetophylax morschi from restinga (P = 0.014; Fig. 3b).
Faith’s Phylogenetic Diversity among: (a) the five species of fungus-farming ants from which ants external to the nest were sampled, and (b) the three species of fungus-farming ants from which external foragers (white) and internal ants (on the fungus garden; dark grey) were sampled. Note that data for Mycetophylax morschi are presented separately for the restinga forest and the dune area.
Composition of bacterial communities: interspecific comparisons and differences between forager and internal ants
The composition of the bacterial communities in individuals external to the nest differed among the attine species (Table 2). Similar patterns were observed when using unweighted and weighted UniFrac distance (Table S2 Supplementary Material 1). We characterized the differences in bacterial composition with the first two axes of a PCoA (Fig. 4a). We also observed that the bacterial composition of external individuals of Mycetophylax morschi from restinga forest were different from the dune area (Table 2, Fig. 4a).
Two-dimensional plot of principal coordinates analysis (Bray-Curtis distance) showing differences in microbiota composition: (a) of five species of fungus-farming ants from which ants external to the nest were sampled, and (b) of three species of fungus-farming ants from which external and internal ants were sampled. Note that data for Mycetophylax morschi are presented separately for the restinga forest and the dune area in (a) and (b). The samples were rarefied at 2000 reads. In (b), triangles represent ants internal to the nest (on the fungus garden), and circles represent ants external to the nest (foragers).
When we considered the species of ants for which we sampled individuals internal (on the fungus garden) and external (foragers) to the nest (Mycocepurus smithii, Mycetarotes parallelus, Mycetophylax morschi from the restinga forest, and Mycetophylax morschi from the dune area), the bacterial community composition differed among the host species, and between foragers and internals within some ant species (Table 2, Fig. 4b. The same pattern was observed with unweighted and weighted UniFrac distance, see Table S2 Supplementary Material 1). In Mycocepurus smithii and Mycetophylax morschi (restinga forest), the bacterial communities of forager and internal workers differed slightly (Table 2, Fig. 4b). Bacterial communities of Mycetarotes parallelus and Mycetophylax morschi (dune area), however, did not differ between foragers and ants internal to the nest (Table 2, Fig. 4b).
SIMPER results showed that the main OTUs responsible for the dissimilarity between bacterial communities in internal and external workers of Mycocepurus smithii were Chryseobacterium, Pseudonocardia OTU 2, and Luteimonas; and in Mycetophylax morschi from restinga were Pseudonocardia OTU 2, Arthrobacter woluwensis, Intrasporangiaceae OTU 1, Chitinophagaceae OTU 1, and Nocardioidaceae OTU 1 (See Table S3 Supplementary Material 1). The bacterial communities of internal and external workers of Mycetarotes parallelus and Mycetophylax morschi from the dune area did not differ significantly.
Intraspecific differences in bacterial community composition
The composition of the bacterial community within each species of fungus-farming ant differed among colonies (Table 3, Fig. 5). Specifically, the differences were observed for Mycocepurus smithii, Mycetophylax morschi from restinga, S. parvulus and S. saussurei (Table 3, Fig. 5). Only Mycetarotes parallelus did not show differences in bacterial community composition among colonies (Table 3, Fig. 5).
Two-dimensional plot of principal coordinates analysis (Bray-Curtis distance) showing differences in microbiota composition between colonies from each species of fungus-farming ant. The samples were rarefied at 2000 reads. Each color represents one colony. Four nests were sampled for Sericomyrmex saussurei. Three nests were sampled for the other species.
Discussion
The current study provided data on the richness and diversity of bacterial communities associated with species of fungus-farming ants (non-leaf-cutters) from the Atlantic rainforest, with novel evidence that microbiota in these ants are host-specific and colony-specific. Additionally, we investigated the potential differences in bacterial communities of ants internal vs. external (foragers) to the nest environment. Our results revealed clear differences in bacterial communities associated with external foragers from the species of fungus-growers, indicating host specificity of these communities. On the other hand, the bacterial communities from internal (on the fungus garden) versus external ants did not differ within species (Mycetarotes parallelus, Mycetophylax morschi from the dune area) or differed only slightly (Mycocepurus smithii, Mycetophylax morschi from the restinga forest). Moreover, the composition of the bacterial community differed among colonies of the same species (with exception of Mycetarotes parallelus), which indicates that the bacterial microbiota may be colony-specific.
We showed that the five species of attine ants were predominantly associated with members of the phyla Actinobacteria and Proteobacteria. In a recent study by Sapountzis et al.37 with 17 species of attine ants (including non-leaf-cutters), the abdominal microbiomes were dominated by Actinobacteria, Proteobacteria and Mollicutes. Actinobacteria and Proteobacteria are frequently present in the fungus garden of leaf-cutters (e.g., Acromyrmex echinatior, Atta cephalotes, and Atta colombica), suggesting a role by these bacteria in nutrient cycling in the fungus-garden4,11,38. Actinobacteria comprise a diverse group commonly found in soil-dwelling insects39, and are known in attine ants because some of their representatives (e.g., Pseudonocardia) produce secondary metabolites with antibiotic activity that protects the fungus garden against pathogens7,40. Li et al.16 recently showed that Pseudonocardia bacteria are the dominant filamentous Actinobacteria on the exoskeleton of several species of fungus-farming ants. In our study, the association of Mycocepurus smithii, Mycetarotes parallelus, and Mycetophylax morschi with Actinobacteria (especially with Pseudonocardia in Mycocepurus smithii and Mycetarotes parallelus) suggests a protective role by these microorganisms that, together with grooming and other hygienic behavior21, could reduce pathogenic infection in the fungus garden.
In our study, few OTUs were shared among the five species of fungus-farming ants and each species was dominated by specific OTUs. The microbiota is often host-specific12,41, which may be a mechanism to prevent host colonization by pathogenic microorganisms42. In fungus-farming ants, the microbiota could be also related with the phylogeny of these ants, as demonstrated by Sapountzis et al.37. Moreover, bacterial communities in attines could be shaped by ant behaviors, such as grooming and licking, which remove the unwanted bacteria from the cuticle of the ants and from the fungus garden18,43. We also suggest that the items collected for fungiculture may influence the bacteria associated with these ants, since different species use variable proportions of vegetable matter and arthropod feces to cultivate the symbiotic fungi19. Kellner et al.12 suggested that some groups of bacteria associated with Mycocepurus smithii are influenced by the insect feces collected by the ants. A previous study focusing solely on abdominal microbiomes showed that bacterial communities of Mycocepurus smithii vary markedly between field and captive colonies, suggesting that even if host specialization occurs, it is influenced by other factors, such as diet37. Overall, we found differences in the phylogenetic diversity of bacteria among species, with Sericomyrmex saussurei hosting the highest bacterial diversity. Since S. saussurei traveled the greatest foraging distances (up to 3.60 m) among the five studied species19, foragers in this species may be exposed to an increased bacterial diversity from the external environment.
Fungus-farming ants species host different bacterial communities, indicating that species identity is an important factor shaping the associated microbiota. Indeed, several studies have shown that host-associated bacterial communities are species-specific12,37,41,44. Our findings indicate that bacterial communities differ between host species even when they occur in the same area, suggesting that the ant identity, rather than local microorganism assemblages, drives the composition of associated bacteria. This pattern was also observed for Megalopta bees45 and Camponotini ants46,47. Furthermore, the environment outside the nest seems to have little influence on the associated bacterial community of fungus-farming ants, since the composition of ants on the fungus garden versus outside foragers did not differ within species (Mycetarotes parallelus, Mycetophylax morschi from the dune area) or differed only slightly (Mycocepurus smithii, Mycetophylax morschi from the restinga forest), reinforcing the hypothesis of specificity for this host-bacterial interaction. Similar results were obtained by Kellner et al.12 for the attine ant Mycocepurus smithii, whose associated microbiota was distinct from the community in the soil adjacent to the nest. Ishak et al.48 also did not observe differences in bacterial community composition between workers of the fungus-farming ant Trachymyrmex septentrionalis sampled inside and outside the nest.
Despite the importance of host species identity in predicting associated bacterial communities, environmental characteristics such as temperature, salinity, and pH can also influence the structure of these communities49,50. Indeed, we showed that the diversity and composition of bacterial communities associated with Mycetophylax morschi differed between restinga forest and the dune area. This result may indicate an environmental effect on the bacterial community composition in this species, since Mycetophylax morschi from the dune area is constantly under influence of the sea and salinity. Additionally, the dune area has little canopy cover when compared with restinga forest (canopy up to 20 m tall)20, which could lead to variation in microclimatic conditions near the nests (temperature, humidity). The sampling of a large number of colonies of Mycetophylax morschi from the restinga forest and dune area coupled with abiotic data (e.g., temperature, humidity, salinity, pH), could clarify if interhabitat difference in associated bacterial communities in this species is affected by the environment or results from intracolonial variation, as detected in the restinga forest.
Our results demonstrate that colonies of most species of fungus-farming ants harbor specific bacterial communities, except for Mycetarotes parallelus. Ants are social insects that live in populous colonies of related individuals sharing the same space and food51, which favors the exchange of microorganisms among nestmates52. In other social insects such as bumblebees and termites, the gut microbiota varies among colonies and suggests a colony-specific signature in bacterial communities53,54,55. Hu et al.56 suggest that differences in bacterial communities between colonies of the same ant species can be explained by host genetic variability, which would lead to a colony-level natural selection of the microorganisms. In addition, the microbiome is related to group-specific odors, suggesting that the members of the same social group harbor similar odor-producing bacterial communities, which is potentially important for within group recognition (e.g., termites53, hyenas57, meerkats58, and the leaf-cutter ant Acromyrmex echinatior59).
In conclusion, our results show that host species is a strong predictor of the bacterial composition in fungus-farming ants, supporting host species specificity in associated bacterial microbiota. We also detected colony-specificity in the composition of bacterial communities, demonstrating intraspecific variation in the microbiota associated with the attine species investigated. Our study unravels the richness and diversity of bacterial communities that live with fungus-farming ants in the Atlantic rainforest and highlights the importance of describing ant-associated bacterial communities to better understand this host-bacterial interaction in the social environment of insect colonies. The results provide key information for future studies on the ecological and evolutionary processes that drive the success of fungus-farming ants in tropical habitats.
Data availability
Ant voucher specimens were deposited at the Museu de Zoologia da Universidade Estadual de Campinas (ZUEC, Campinas, Brazil; registration numbers ZUEC4236 to ZUEC4240). Sequences of all barcoded specimens were deposited in GenBank (accession numbers MH206536- MH206586). Sequences were deposited in the NCBI Sequence Read Archive (accession number PRJNA575362). The dataset supporting this article is available as electronic supplementary material (Supplementary Material 2).
References
McFall-Ngai, M. et al. Animals in a bacterial world, a new imperative for the life sciences. Proceedings of the National Academy of Sciences of the United States of America 110, 3229–3236 (2013).
Koskella, B., Hall, L. J. & Metcalf, C. J. E. The microbiome beyond the horizon of ecological and evolutionary theory. Nature Ecology & Evolution 1, 1606–1615 (2017).
Xu, J. Microbial ecology in the age of genomics and metagenomics: concepts, tools, and recent advances. Molecular Ecology 15, 1713–1731 (2006).
Aylward, F. O. et al. Convergent Bacterial Microbiotas in the Fungal Agricultural Systems of Insects. mBio 5 (2014).
Parfrey, L. W., Moreau, C. S. & Russell, J. A. Introduction: The host-associated microbiome: Pattern, process and function. Molecular Ecology 27, 1749–1765 (2018).
Hölldobler, B. & Wilson, E. O. The leafcutter ants: civilization by instinct. (Norton (2011).
Currie, C. R., Bot, A. N. M. & Boomsma, J. J. Experimental evidence of a tripartite mutualism: bacteria protect ant fungus gardens from specialized parasites. Oikos 101, 91–102 (2003).
Sosa-Calvo, J., Ješovnik, A., Vasconcelos, H. L., Bacci, M. & Schultz, T. R. Rediscovery of the enigmatic fungus-farming ant ‘Mycetosoritis’ asper Mayr (Hymenoptera: Formicidae): Implications for taxonomy, phylogeny, and the evolution of agriculture in ants. PLOS ONE 12, e0176498 (2017).
Van Borm, S., Billen, J. & Boomsma, J. J. The diversity of microorganisms associated with Acromyrmex leafcutter ants. BMC Evolutionary Biology 2, 9 (2002).
Pinto-Tomás, A. A. et al. Symbiotic Nitrogen Fixation in the Fungus Gardens of Leaf-Cutter Ants. Science 326, 1120–1123 (2009).
Aylward, F. O. et al. Metagenomic and metaproteomic insights into bacterial communities in leaf-cutter ant fungus gardens. The ISME Journal 6, 1688–1701 (2012).
Kellner, K., Ishak, H. D., Linksvayer, T. A. & Mueller, U. G. Bacterial community composition and diversity in an ancestral ant fungus symbiosis. FEMS Microbiology Ecology 91, fiv073 (2015).
Currie, C. R. et al. Coevolved Crypts and Exocrine Glands Support Mutualistic Bacteria in Fungus-Growing Ants. Science 311, 81–83 (2006).
Little, A. E. F. & Currie, C. R. Symbiotic complexity: discovery of a fifth symbiont in the attine ant–microbe symbiosis. Biology Letters 3, 501–504 (2007).
Cafaro, M. J. et al. Specificity in the symbiotic association between fungus-growing ants and protective Pseudonocardia bacteria. Proceedings of the Royal Society B: Biological Sciences 278, 1814–1822 (2011).
Li, H. et al. Convergent evolution of complex structures for ant–bacterial defensive symbiosis in fungus-farming ants. Proceedings of the National Academy of Sciences of the United States of America 115, 10720–10725 (2018).
Kaltenpoth, M. Actinobacteria as mutualists: general healthcare for insects? Trends in Microbiology 17, 529–535 (2009).
Currie, C. R. A Community of Ants, Fungi, and Bacteria: A Multilateral Approach to Studying Symbiosis. Annual Review of Microbiology 55, 357–380 (2001).
Ronque, M. U. V., Feitosa, R. M. & Oliveira, P. S. Natural history and ecology of fungus-farming ants: a field study in Atlantic rainforest. Insectes Sociaux 66, 375–387 (2019).
Joly, C. A. et al. As parcelas permanentes do projeto temático Biota Gradiente Funcional: composição florística, estrutura e funcionamento da floresta ombrófila densa dos Núcleos Picinguaba e Santa Virgínia do Parque Estadual da Serra do Mar, Estado de São Paulo, Brasil. In: Experiências de monitoramento no Bioma Mata Atlântica com uso de parcelas permanentes (ed Sanquetta, C. R.) (Funpar, Curitiba, pp 109-129 (2008).
Ronque, M. U. V. Ecology, behavior and microbiology of fungus-farming ants (Formicidae, Myrmicinae, Attini, Attina) in Atlantic rainforest. PhD dissertation, Institute of Biology, University of Campinas, Campinas, Brazil (2018).
Kozich, J. J., Westcott, S. L., Baxter, N. T., Highlander, S. K. & Schloss, P. D. Development of a Dual-Index Sequencing Strategy and Curation Pipeline for Analyzing Amplicon Sequence Data on the MiSeq Illumina Sequencing Platform. Applied and Environmental Microbiology 79, 5112–5120 (2013).
Caporaso, J. G. et al. Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proceedings of the National Academy of Sciences of the United States of America 108, 4516–4522 (2011).
Bletz, M. C. et al. Amphibian gut microbiota shifts differentially in community structure but converges on habitat-specific predicted functions. Nature Communications 7, 13699 (2016).
Folmer, O., Black, M., Hoeh, W., Lutz, R. & Vrijenhoek, R. DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Molecular Marine Biology and Biotechnology 3, 294–299 (1994).
Kearse, M. et al. Geneious Basic: An integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28, 1647–1649 (2012).
Bolyen, E. et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nature Biotechnology 37, 852–857 (2019).
Rognes, T., Flouri, T., Nichols, B., Quince, C. & Mahé, F. VSEARCH: a versatile open source tool for metagenomics. PeerJ 4, e2584 (2016).
Amir, A. et al. Deblur Rapidly Resolves Single-Nucleotide Community Sequence Patterns. mSystems 2, e00191–16 (2017).
Katoh, K. & Standley, D. M. Mafft multiple sequence alignment software version 7: improvements in performance and usability. Molecular biology and evolution 30, 772–780 (2013).
Price, M. N., Dehal, P. S. & Arkin, A. P. FastTree 2 – Approximately Maximum-Likelihood Trees for Large Alignments. PLOS ONE 5, e9490 (2010).
Bokulich, N. A. et al. Optimizing taxonomic classification of marker‐gene amplicon sequences with QIIME 2’s q2‐feature‐classifier plugin. Microbiome 6, 90 (2018).
McDonald, D. et al. An improved Greengenes taxonomy with explicit ranks for ecological and evolutionary analyses of bacteria and archaea. The ISME Journal 6, 610–618 (2012).
Bioinformatics & Evolutionary Genomics. http://http://bioinformatics.psb.ugent.be/webtools/Venn/. Accessed 08 April 2020.
R Core Team. R: A language and environment for statistical computing. Vienna, Austria: R Foundation for Statistical Computing (2017).
Lozupone, C., Lladser, M. E., Knights, D., Stombaugh, J. & Knight, R. UniFrac: an effective distance metric for microbial community comparison. The ISME Journal 5, 169–172 (2011).
Sapountzis, P., Nash, D. R., Schiøtt, M. & Boomsma, J. J. The evolution of abdominal microbiomes in fungus-growing ants. Molecular Ecology 28, 879–899 (2019).
Suen, G. et al. Ant insect herbivore microbiome with high plant biomass-degrading capacity. PLOS ONE 6, e1001129 (2010).
Goodfellow, M. & Williams, S. T. Ecology of Actinomycetes. Annual Review of Microbiology 37, 189–216 (1983).
Poulsen, M. et al. Variation in Pseudonocardia antibiotic defence helps govern parasite-induced morbidity in Acromyrmex leaf-cutting ants. Environmental Microbiology Reports 2, 534–540 (2010).
Muletz Wolz, C. R., Yarwood, S. A., Campbell Grant, E. H., Fleischer, R. C. & Lips, K. R. Effects of host species and environment on the skin microbiome of Plethodontid salamanders. Journal of Animal Ecology 87, 341–353 (2017).
Dillon, R. J. & Dillon, V. M. The gut bacteria of insects: nonpathogenic interactions. Annual Review of Entomology 49, 71–92 (2004).
Fernandez-Marin, H., Zimmerman, J. K., Nash, D. R., Boomsma, J. J. & Wcislo, W. T. Reduced biological control and enhanced chemical pest management in the evolution of fungus farming in ants. Proceedings of the Royal Society B: Biological Sciences 276, 2263–2269 (2009).
Council, S. E. et al. Diversity and evolution of the primate skin microbiome. Proceedings of the Royal Society B: Biological Sciences 283, 20152586 (2016).
McFrederick, Q. S., Wcislo, W. T., Hout, M. C. & Mueller, U. G. Host species and developmental stage, but not host social structure, affects bacterial community structure in socially polymorphic bees. FEMS Microbiology Ecology 88, 398–406 (2014).
Ramalho, M. O., Bueno, O. C. & Moreau, C. S. Microbial composition of spiny ants (Hymenoptera: Formicidae: Polyrhachis) across their geographic range. BMC Evolutionary Biology 17 (2017).
Ramalho, M. O., Bueno, O. C. & Moreau, C. S. Species-specific signatures of the microbiome from Camponotus and Colobopsis ants across developmental stages. PLOS ONE 12, e0187461 (2017).
Ishak, H. D. et al. Microbiomes of ant castes implicate new microbial roles in the fungus-growing ant Trachymyrmex septentrionalis. Scientific Reports 1 (2011).
Lokmer, A. & Mathias Wegner, K. Hemolymph microbiome of Pacific oysters in response to temperature, temperature stress and infection. The ISME Journal 9, 670–682 (2015).
Schmidt, V. T., Smith, K. F., Melvin, D. W. & Amaral-Zettler, L. A. Community assembly of a euryhaline fish microbiome during salinity acclimation. Molecular Ecology 24, 2537–2550 (2015).
Hölldobler, B. & Wilson, E. O. The ants. (Belknap Press of Harvard University Press (1990).
Archie, E. A. & Theis, K. R. Animal behaviour meets microbial ecology. Animal Behaviour 82, 425–436 (2011).
Minkley, N., Fujita, A., Brune, A. & Kirchner, W. H. Nest specificity of the bacterial community in termite guts (Hodotermes mossambicus). Insectes Sociaux 53, 339–344 (2006).
Koch, H., Cisarovsky, G. & Schmid-Hempel, P. Ecological effects on gut bacterial communities in wild bumblebee colonies. Journal of Animal Ecology 81, 1202–1210 (2012).
Boucias, D. G. et al. The hindgut lumen prokaryotic microbiota of the termite Reticulitermes flavipes and its responses to dietary lignocellulose composition. Molecular Ecology 22, 1836–1853 (2013).
Hu, Y., Łukasik, P., Moreau, C. S. & Russell, J. A. Correlates of gut community composition across an ant species (Cephalotes varians) elucidate causes and consequences of symbiotic variability. Molecular Ecology 23, 1284–1300 (2014).
Theis, K. R., Schmidt, T. M. & Holekamp, K. E. Evidence for a bacterial mechanism for group-specific social odors among hyenas. Scientific Reports 2 (2012).
Leclaire, S., Jacob, S., Greene, L. K., Dubay, G. R. & Drea, C. M. Social odours covary with bacterial community in the anal secretions of wild meerkats. Scientific Reports 7, 3240 (2017).
Teseo, S. et al. The scent of symbiosis: gut bacteria may affect social interactions in leaf-cutting ants. Animal Behaviour 150, 239–254 (2019).
Acknowledgements
We thank Cameron Currie, Sasha Greenspan, Alexander Christianini, Paula de Omena, and Elizabeth Bilsland for valuable comments on the manuscript. The Parque Estadual Serra do Mar (Núcleo Picinguaba) and the Instituto Florestal do Estado de São Paulo provided logistic support for field work. We thank Ana Ješovnik and Rodrigo Feitosa for identifying the ant species, Sérgio Kakazu and Renata Zani for helping with lab work, and Guilherme Becker for valuable discussions at early stages of the study. M.U.V.R. and G.H.M. were funded by the Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES; Finance code 001). M.L.L. (2017/26162-8) and M.B.Jr. (2014/24407-3) were supported by the Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP). P.S.O was supported by the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq 306115/2013-1, 302219/2017-0) and the Fundação de Amparo à Pesquisa do Estado de São Paulo (2014/23141-1, 2017/16645-1).
Author information
Authors and Affiliations
Contributions
M.U.V.R., M.L.L., M.B. and P.S.O. conceived the study. M.U.V.R. and G.H.M. conducted the field work. M.U.V.R. and M.L.L. conducted the molecular work, and M.B. provided the facilities and equipment to perform the molecular work. M.U.V.R., G.H.M. and M.L.L. analyzed the data. M.U.V.R. wrote the paper with the assistance of P.S.O. All authors read and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Ronque, M.U.V., Lyra, M.L., Migliorini, G.H. et al. Symbiotic bacterial communities in rainforest fungus-farming ants: evidence for species and colony specificity. Sci Rep 10, 10172 (2020). https://doi.org/10.1038/s41598-020-66772-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-66772-6
This article is cited by
-
Symbiosis, dysbiosis and the impact of horizontal exchange on bacterial microbiomes in higher fungus-gardening ants
Scientific Reports (2024)
-
Environments and Hosts Structure the Bacterial Microbiomes of Fungus-Gardening Ants and their Symbiotic Fungus Gardens
Microbial Ecology (2023)
-
Gram-negative bacteria associated with a dominant arboreal ant species outcompete phyllosphere-associated bacteria species in a tropical canopy
Oecologia (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.