Abstract
Patients that suffer from sepsis exhibit an early hyper-inflammatory immune response which can lead to organ failure and death. In our study, we assessed the immune modulation in the human in vivo endotoxemia model and compared it to ex vivo LPS stimulation using 38 transcriptomic markers. Blood was collected before and after 4 hours of LPS challenge and tested with the Immune Profiling Panel (IPP) using the FilmArray system. The use of IPP showed that markers from the innate immunity dominated the response to LPS in vivo, mainly markers related to monocytes and neutrophils. Comparing the two models, in vivo and ex vivo, revealed that most of the markers were modulated in a similar pattern (68%). Some cytokine markers such as TNF, IFN-γ and IL-1β were under-expressed ex vivo compared to in vivo. T-cell markers were either unchanged or up-modulated ex vivo, compared to a down-modulation in vivo. Interestingly, markers related to neutrophils were expressed in opposite directions, which might be due to the presence of cell recruitment and feedback loops in vivo. The IPP tool was able to capture the early immune response in both the human in vivo endotoxemia model, a translational model mimicking the immune response observed in septic patients.
Similar content being viewed by others

Introduction
The host immune response in sepsis is currently known as an initial phase of a hyper-inflammatory response, that is concomitantly met with an anti-inflammatory response to restore homeostasis1. In the Intensive Care Unit (ICU), mortality attributable to sepsis can reach up to 45%2 due to organ failure – a consequence of a dysregulated host response including inflammatory cytokine storm– and/or acquiring secondary infections - with concurrent sepsis-induced immunoparalysis1. The major challenge in managing septic patients is the high heterogeneity in host responses due to inter-individual variability, the pathogen or source of infection and varying responses to treatment, which lead to different immune trajectories and outcomes3,4. Many clinical trials have been conducted, and are currently ongoing, to test the efficacy of several immune-directed therapies to improve patient outcomes. For instance, anti-inflammatory agents such as Interleukin-1 Receptor antagonist (IL-1Ra)5 and immune stimulatory agents such as interleukin-7 (IL-7)6 and interferon-gamma (IFN-γ)7,8,9 and others with promising results are currently under evaluation. Nonetheless, it is not yet feasible to easily identify and stratify patients with different immune profiles in the ICU that would benefit from such treatment in day-to-day clinical practice10. The availability of an immune profiling tool based on immune biomarkers can help determine the superseding immune dysfunction (hyper inflammation or immune suppression) in septic patients. Such tool can be used in personalized medicine to help stratify and assign patients that are likely to benefit from a targeted immunomodulatory agent and monitor their response to treatment7,11.
Lipopolysaccharide (LPS) is a highly antigenic molecule that is derived from Gram-negative bacteria and activates a plethora of inflammatory cascades via Toll-like receptor-4 (TLR-4)12. LPS stimulation can mimic the initial acute inflammatory response to sepsis, both in vivo and ex vivo. The human in vivo endotoxemia model is a well-established model that can capture the host response from half an hour after the challenge and up to 20 days13. In vivo endotoxemia subjects experience transient symptoms reminiscent of systemic infection in sepsis that include fever, chills, headaches and nausea. These symptoms are accompanied by an increase in blood pressure, heart rate, as well as neutrophilia, lymphocytopenia and monocyte anergy14,15. Other widely used model in immunological studies is the ex vivo LPS stimulation of whole blood. The use of ex vivo models helped unravel many underlying mechanisms of the inflammatory response, and it can also be utilized as a standalone test of the immune function16. The safety, non-invasiveness and flexibility of testing different stimulants make the ex vivo model easy-to-use and appealing to researchers. The ex vivo model has also proved valuable in the diagnostic field, where functional assays such as TNF-α release post-LPS stimulation has been used to test the immune competency of Peripheral Blood Mononuclear Cells (PBMCs) of patients10. Such well-established model can help untangle the complexity of host responses at the transcriptomic and proteomic levels. The conditions of the ex vivo model are easier to control and replicate, while in vivo, the model better mimics the human or patients’ host response to inflammation, as it allows the occurrence of positive and negative feedback loops, in addition to recruitment and mobilization of various subsets of immune cells15,17. However, setting up an in vivo human endotoxemia model is met with several barriers such as the rules and regulations of each country, the need for screening and careful recruitment of volunteers (to minimize heterogeneity), the potential risks for healthy volunteers, and the continuous monitoring directly afterwards.
In this study, we used a tool we recently described: the Immune Profiling Panel (IPP), a multiplex molecular tool using the FilmArray system18, to assess the host response to LPS stimulation, using 38 immune-related transcriptome markers in vivo. The panel of markers targeted several immune functions, cell surface markers and the two arms of the immune response including inflammatory and anti-inflammatory responses. We compared the gene modulation in vivo versus ex vivo LPS stimulation of whole blood samples from healthy volunteers, in order to better understand the ability and limitations of each model in capturing the complexity of the host response to endotoxemia.
Material and Methods
Healthy volunteers
Eight healthy, non-smoking, Caucasian male subjects (aged 18–25 years) from the Netherlands were examined, screened and recruited for studying the in vivo human endotoxemia model14. The volunteers received an LPS bolus infusion (E. coli O113 Reference Endotoxin, National Institutes of Health, Bethesda, Maryland, USA) at a dose of 2 ng/kg bodyweight. Whole blood (2.5 mL) was collected in PAXgene tubes (Pre-Analytix, Hilden, Germany) at T0 (before LPS injection) and T4 (4 hours after LPS injection). The PAXgene tubes were inverted several times and incubated for 2 hours at room temperature according to the manufacturer’s recommendation, and stored at -80 °C until the transcriptomic testing. The blood cell counts were performed using fully automated hematology analyser XN-9000 (Sysmex, Etten-Leur, The Netherlands). The study was approved by the institutional ethics and research board (Amsterdam University Medical Centers, location AMC; Netherlands Trial Registration number NL4425 (NTR4549); ClinicalTrials.gov identifier NCT02127749). All subjects gave a written informed consent and research was conducted in accordance with the declaration of Helsinki.
For the ex vivo LPS stimulation, whole blood was collected in heparin tubes from 8 healthy volunteers, 6 males and 2 females (aged 21–55 years) obtained from the EFS (Etablissement Français du Sang, French blood bank, Lyon, France). One milliliter of heparinized blood was distributed into pre-warmed TruCulture tubes (Myriad Rbm, Austin, TX, USA) containing 2 ml of medium alone (Nul) or medium with LPS (E. coli O55:B5; 100 ng/mL). The TruCulture tube system was selected for the ex vivo model as it is an easy-to-use culture tube supplied with a standardized culture medium and LPS, thus, designed for use as standardized diagnostic tests (currently available for Research Use Only)19,20. The samples were incubated in a dry block at 37 °C for 4 hours and directly tested in the IPP prototype. Informed consents from the blood donors were obtained and their personal data were anonymized at the time of blood donation and before reaching our laboratory according to EFS institutional regulations.
Transcriptome analysis
Stimulated whole blood samples (PAXgene in vivo samples and TruCulture stimulated ex vivo samples) were tested with IPP prototype18 according to manufacturer’s instructions. Briefly, the pouches were hydrated with the hydration solution supplied with the kit. A 100 μL of either PAXgene mix or TruCulture mix were mixed with approximately 800 µL of the lysis buffer provided with the kit and directly injected into the pouch and ran on FilmArray 2.0 and FilmArray Torch instruments (BioFire, Inc., Salt Lake City, UT, USA). Results were delivered in less than 1 hour and the normalized expression values of markers were computed18.
Statistical analyses
Normalized expression values were analyzed using hierarchical clustering (Euclidean distance) and scaled to demonstrate the differential modulations of genes in a heatmap plot. Normalized data used in Principal Component Analyses (PCA) were scaled and centered. Missing values in the biological parameters and cells count were imputed using missMDA package21, where a PCA model was used to estimate the missing values to perform the PCA- Biplot. Normalized expression levels of genes were expressed as median and interquartile ranges (IQR) box and whisker plots. Samples were compared before and after LPS stimulation using Mann-Whitney U test for unpaired samples (in vivo against ex vivo model), while paired Wilcoxon signed-rank test was used for paired samples (in vivo model T4 against T0. P-values were adjusted using FDR (Benjamini-Hochberg False Detection Rate) to correct for multiple testing22. The level of significance was set at 5% two-sided tests. Statistical analyses were performed and computed using R software version 3.5.1(R core Team, Vienna, Austria)23.
Results
Gene modulation in the in vivo human endotoxemia model
The IPP tool captured the host immune response in whole blood at the transcriptomic level 4 hours after LPS injection. The PCA illustrates that LPS stimulation drives the variability seen in the dataset (Fig. 1A). All the unstimulated individuals (T0) are grouped on the left-hand side of the plot, and the stimulated individuals (T4) are sparsely grouped in the opposite direction. Principal component-1 (x-axis) explains 78.7% of the variability in the dataset and separates the two time points (before and after the LPS challenge).
Gene modulations captured by IPP in the human in vivo endotoxemia model. (A) Principal component (PC) analysis of the samples before and after stimulation, where the colors differentiate the time points T0 (just before receiving the LPS bolus infusion) and T4 (4 hours after LPS injection). (B) Heatmap showing the hierarchical clustering of the samples (T0 and T4) and genes using Euclidean distance. Normalized gene expression values are color-coded from blue (down-modulation) to red (up-modulation).
In the upper panel of the heatmap (Fig. 1B), an up-modulation can be observed for 20 genes at T4. Markers from the innate immune response dominated the up-modulated genes, which included inflammatory mediators such as NFkB1 (transcription factor involved in inflammatory response) and ALOX5 (involved in leukotrienes synthesis from arachidonic acid associated with inflammation), early pro-inflammatory cytokines TNF and IL-1β, anti-inflammatory cytokines IL-10 and IL1RN, and neutrophil cell surface markers CD177 (neutrophil-specific marker), CD64, TREM1, and S100A9. The latter three are markers expressed on the surface of both neutrophil and monocyte cells, which are the first line of cells that respond to the LPS challenge.
The lower panel highlights 18 genes that were down-modulated at T4. These included monocyte and T-cell surface markers, in addition to factors involved in the activation and differentiation of T-cells. Monocyte surface markers CX3CR1 (fractalkine receptor that has a major role in monocytes survival), CD74 (the invariant chain of HLA-DR) and CIITA (major histocompatibility complex class II transactivator) were all down-modulated. Also, a decreased expression of T-cell markers including IL7R (associated with lymphocyte development), TBX21 (Th1 cell-specific transcription factor), and ZAP70 (part of T cell receptor, and plays a critical role in T-cell signaling) indicating a compromised functionality of lymphocytes. A decrease in CD3D (encodes T-cell surface glycoprotein CD3 delta chain) and GATA3 (regulator of T-cell development) expression was also captured by the IPP tool which might be related to a decline in the lymphocyte count.
Biological parameters and immune cell modulation observed in the in vivo endotoxemia model
The kinetic changes in the absolute count of immune cells before and after the LPS stimulation were recorded in our prior study14. A marked decrease was observed in monocyte, eosinophil and lymphocyte populations while a 2-fold increase in the neutrophil count was detected after 4 hours of LPS stimulation (Supplementary Fig. S1).
The biological parameters measured for the same volunteers were analyzed using PCA to identify and evaluate the contribution of each variable to the separation of T0 and T4. The highest contributing variables to the observed responses were the absolute count of monocytes and lymphocytes, as well as coagulation factors II and X, prothrombin time (PT), Plasminogen activator inhibitor-1 (PAI-1), inflammatory cytokines: IL6, IL8, TNF-α and anti-inflammatory cytokine IL10 (Supplementary Fig. S2). The correlations of various markers that are biologically related to immune cell counts are illustrated in Fig. 2. S100A9 and CD177 had a strong correlation (Pearson coefficient > 0.7) with neutrophils count. Monocytes count was also strongly correlated; positively with CX3CR1 and negatively with TNF expression (Pearson coefficient > 0.8). As for the lymphocytes count, it was strongly correlated with CD3D and GATA3 (Pearson coefficient > 0.7). The rest of the correlations between the biological markers and cell counts with IPP markers were above 0.7 (Pearson coefficient), an indicator for a strong correlation, are shown in Supplementary Fig. S3.
Pearson correlations of IPP markers with the absolute count of immune cells. Six IPP markers were correlated with the absolute count of immune cells at T0 (red cirlces) and T4 (blue triangles), with a strong correlations above 0.7 (Pearson coefficient). All correlations were statistically significant p < 0.001. A positive correlations indicates that the normalized expression of IPP markers increases as the count increases. (A) correlations between S100A9 and CD177 expression with neutrophils count. (B) Correlation between TNF and CX3CR1, and monocytes count. (C) Correlation between CD3D and GATA3, and lymphocytes count.
To assess the impact of immune cell count changes on gene expression, we adjusted the expression level of each marker to the absolute number of the corresponding immune cell. Most of the markers remained differentially expressed between T0 and T4, with the exception of CD64, GATA3 and CD3D (Fig. S4). In the case of CD177 and CD74, we observed a clear differential expression that did not reach statistical significance, which might be due to the limited number of samples.
Comparison of the in vivo Human Endotoxemia Model with the ex vivo LPS Stimulation in TruCulture System
IPP markers were used to study the difference between the in vivo endotoxemia model and LPS blood stimulation ex vivo. The majority of the IPP markers (68%) showed a similar pattern of expression in both models post-LPS stimulation. PCA analysis was performed on the entire dataset (Fig. 3), and it can be observed that the unstimulated (NUL) ex vivo samples clustered closely to the in vivo control samples on the first PCA axis, while stimulated samples (T4) were grouped on the other side of the same axis (PC-1). Principal component-1 (x-axis) represents 47% of the dataset variability and separates the LPS response after 4 hours from unstimulated samples with an equal distance. Of particular interest is PC-2 (y-axis) in this analysis, which separates the two LPS responses according to the type of model whether ex vivo or in vivo, explaining 41.2% of the variability of the whole dataset (Fig. 3). Interestingly, Fig. 3 shows that the direction of variance is opposite in the two models. The genes that mostly contributed to this difference along PC-2 axis were TDRD9, GSN, S100A9, CD177, TREM1, CD64, ALOX5, ADGRE3 and IL1RN (up-regulated genes in vivo), and CTLA4, TIM3, GATA3, IL7R, CCNB1IP1 and ZAP70 genes (down-regulated in vivo).
Comparison between in vivo and ex vivo LPS stimulation using IPP. PC analysis was used to compare the difference of expression between the human in vivo endotoxemia model (blue) and the ex vivo stimulation model (red) before (NUL, circles) and after the LPS challenge (LPS, triangles). The contribution of each gene to the plot is represented as arrows.
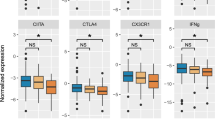
We compared the markers that showed an opposite and different magnitude of expression patterns between the two models, by subtracting the unstimulated expression from the LPS stimulation for each marker (Fig. 4). The comparison revealed that the expression of ADGRE3, ALOX5, CD177, CD64, GSN, S100A9 and TREM1 were up-modulated only in the in vivo model, which are chiefly associated with the early innate immune response (Fig. 4A). In contrast, some genes were down-modulated in vivo only and were mainly related to the adaptive immune response (IL7R, CTLA4, GATA3, TIM3, and GNLY) (Fig. 4B). Interestingly, some markers had different magnitudes of expression in the two models, such as IFN-γ, IL-1β, and TNF, and were strongly up-regulated ex vivo compared to the in vivo model (Fig. 4C). The comparison between the two models highlighted that the majority of the IPP markers were expressed in a similar pattern, while others, interestingly, were mostly modulated in vivo only, and clearly discriminated the two models.
Boxplots showing the fold change of IPP markers observed between the in vivo and the ex vivo models after LPS stimulation. On the y-axis is the fold change of both models (normalized expression in the stimulated condition (LPS) minus the unstimulated condition) presented as boxplots, in vitro stimulation (blue) and in vivo endotoxemia (green). (A) Fold change of markers associated with the early innate immune response. (B) Fold change of markers associated with the adaptive immune response. (C) Fold change of markers associated with inflammatory and anti-inflammatory cytokines. Fold changes were compared using Mann-Whitney U test and P values were adjusted using FDR (Benjamini-Hochberg), where NS: p > 0.05, *p < 0.05, **p < 0.01 and ***p < 0.001.
Discussion
The in vivo human endotoxemia model is a translational model that can be used to mimic the acute inflammatory profile reported in septic patients15. Once the volunteers are challenged with LPS, the innate immune response is triggered initiating several pro-inflammatory cascades and the secretion of inflammatory cytokines, quickly followed by an increase in neutrophil count24. The peak of neutrophilia, mono-, eosino- and lympho-cytopenia occurred at 2–4 hours after the endotoxin challenge in vivo25,26,27,28. This initial systemic inflammatory response is concomitantly met with an anti-inflammatory response to restore homeostasis. In sepsis, this immune response can become dysregulated causing detrimental consequences. The use of the Immune Profiling Panel (IPP) helped capture the gene modulations observed in the in vivo endotoxemia model. An increase in neutrophil-associated markers (S100A9, TREM1 and neutrophil specific marker CD17729) reflects the increased recruitment and activation of neutrophils, while the increased expression of CD64 might indicate an increase in neutrophil count. A down-modulation in monocyte surface markers (CX3CR1, CIITA and CD74, a surrogate marker for HLA-DR30) may indicate monocyte deactivation. The down-modulation of the adaptive immune markers (GNLY, TBX21, RORγt, CTLA4) highlights an alteration in lymphocyte function, signaling and the activation of Th1, Th17 and Treg cells. When CD3D and GATA3 were adjusted to the lymphocyte count they seem to be non-differentially expressed indicating that their expression might mainly reflect lymphocytopenia. An overall increase was observed in both pro- and anti-inflammatory cytokines expression such as TNF, IL-1β, IFN-γ, IL10 and IL1RN after stimulation (Fig. 1B). Each of these modulations pinpoints to a functional alteration, that resembles the acute inflammatory phase observed in septic patients31. These results underscore future possibilities of using novel molecular tools such as IPP at the bedside in the ICU to help characterize the immune profile of patients at risk, enabling personalized medicine based on immune markers.
In the second part of our study, we compared the same 38 IPP markers with the ex vivo LPS stimulation of whole blood after 4 hours. Most of the markers (~70%) were expressed in a similar pattern and direction with minor non-significant differences in magnitude (data not shown). This slight difference in magnitude between the two models might be due to the use of different healthy volunteers or the type and concentration of LPS. Recently, two markers S100A9 and TREM1 which are included in our panel were identified as LPS dose-dependent, where their expression increased with an increase in the LPS dose32. Curiously, some markers such as TNF were highly expressed ex vivo compared to the in vivo model, while IL10 had the same expression level in both models (Fig. 4C). These results differ from the protein data reported by Dorresteijn et al., where TNF and IL10 levels were higher in vivo compared to ex vivo17. In our data, IL-1β and IFN-γ were highly expressed ex vivo compared to in vivo after 4 hours of stimulation, which corresponds with the findings of Dorresteijn et al. at the protein level 24 hours post-stimulation17. Nonetheless, the difference in timing regimens and the nature of markers (transcriptome versus protein) might also contribute to the differences reported above. Sometimes, the local production of cytokines in tissues exert a “spillover effect”, where the tissues can pour cytokines into peripheral blood, thus increasing the serum levels33. Although transcriptomic tests are more sensitive than protein detection, the post-transcriptional regulation mechanisms can vary across tissues, and the detection of mRNA does not necessarily imply its translation17. The stability of transcripts can also affect their detection in blood, for instance, IL1RN mRNA tends to remain effective and detectable in blood after 4 hours compared to the mRNA of IL-1β which decays rapidly (T1/2 ~1.8 hours)34. All the previous points should be considered when extrapolating results from the ex vivo model to the in vivo model.
Interestingly, some markers showed an opposite expression pattern between the two models. Most of these markers were related to neutrophils such as CD177 as well as CD64, S100A9 and TREM1 that are highly expressed on the surface of both monocytes and neutrophils. This phenomenon might be observed in vivo due to the direct stimulation of circulating neutrophils by LPS, and the recruitment of new neutrophils from the bone marrow25,29. Besides, the ex vivo model is mainly dependent on peripheral blood, while in vivo has both tissue and blood compartments, actively contributing to the detected response. However, this is not the case for the ex vivo model that lacks feedback loops, cell recruitment and communication with the tissue compartment. Our data also showed that the down-modulation in the adaptive immune response markers (CTLA4, GATA3, GNLY, IL7 and TIM3) are linked to the function and activation of T-cells in the in vivo model. This interesting observation might be due to the reduced capacity of antigen-presenting cells to activate the T-cells post-LPS challenge35.
One of our limitations is the use of different E. coli LPS serotypes and concentration, which could contribute to the difference in the host response observed between the two models. However, we wanted to assess the ability of ex vivo and in vivo models to capture the complexity of the host response to LPS and understand their limitations in a standardized manner. This will help provide insights and data about the tool’s ability to capture such modulations which will make it comparable to future studies and tool users. Another limitation was the use of a mixed-gender population for the ex vivo model in comparison to an all-male population in vivo, which might influence the differences observed in immune response between the two models. Wegner et al. observed a higher pro-inflammatory cytokine response to LPS in females compared to males, which was significant only in vivo but not ex vivo36. Nonetheless, we performed a preliminary analysis on the healthy volunteers population (n = 180) in the REALISM project37 and so far we didn’t observe a significant gender-related difference for the transcriptomic markers included in the IPP tool (unpublished data).
To the best of our knowledge, this is the first study to report a direct comparison between the in vivo and ex vivo models after only 4 hours of LPS stimulation using transcriptomic immune markers. This study underscores that caution must be taken when interpreting and extrapolating the results of certain markers from the ex vivo model. Our study also highlights how employing an immune profiling panel can capture and describe the host immune responses in vivo after 4 hours of LPS stimulation. The use of the in vivo human endotoxemia model showed gene modulations and alterations similar to those observed in the early inflammatory response associated with sepsis syndrome and other critical conditions in the ICU. However, in sepsis, a complex dysregulation in the immune response can occur such as the early onset of immunosuppression making it hard to predict the disease trajectory or outcomes. More than 100 clinical trials were conducted on novel immunotherapies in sepsis, those trials mainly targeted the modulation of the systemic inflammatory response in patients with acute clinical manifestations. However, almost none of these trials have resulted in new treatments available in the market. Those clinical trials might have failed as all patients were treated in the same manner, and few patient stratification approaches were adopted11,38. Septic patients have heterogeneous responses and a patient stratification strategy based on a panel of immune biomarkers can help identify an individual patient’s dysfunction11. An immune profiling tool can guide the use of targeted immunotherapies and contribute to the success of future clinical trials. Testing IPP in human in vivo endotoxemia model showed the potential value of such tool to determine the immune status of critically-ill patients. The immune profiling panel has 5-mins only hands-on, immune markers are measured directly from whole blood and results can be achieved in less than an hour. The ease and rapid turnaround time of IPP suggest its future use at the bedside compared to the currently available techniques that offer limited information about the host response and require long and complicated technical procedures. Our next step is to evaluate the capability and the predictive performances of the IPP markers in a dedicated cohort of critically-ill patients to stratify patients at high risk of stratify patients at high risk of deterioration and enable the use of new immunotherapies and personalized medicine.
References
Hotchkiss, R. S., Monneret, G. & Payen, D. Sepsis-induced immunosuppression: from cellular dysfunctions to immunotherapy. Nat. Rev. Immunol. 13, 862–874, https://doi.org/10.1038/nri3552 (2013).
Fleischmann, C. et al. Assessment of Global Incidence and Mortality of Hospital-treated Sepsis. Current Estimates and Limitations. Am. J. Resp. Crit. Care Med. 193, 259–272, https://doi.org/10.1164/rccm.201504-0781OC (2016).
Scicluna, B. P. et al. Classification of patients with sepsis according to blood genomic endotype: a prospective cohort study. Lancet Resp. Med. 5, 816–826, https://doi.org/10.1016/s2213-2600(17)30294-1 (2017).
Monneret, G., Gossez, M., Aghaeepour, N., Gaudilliere, B. & Venet, F. How Clinical Flow Cytometry Rebooted Sepsis Immunology. Cytometry Part. A. 95, 431–441, https://doi.org/10.1002/cyto.a.23749 (2019).
Shakoory, B. et al. Interleukin-1 Receptor Blockade Is Associated With Reduced Mortality in Sepsis Patients With Features of Macrophage Activation Syndrome: Reanalysis of a Prior Phase III Trial. Crit. Care Med. 44, 275–281, https://doi.org/10.1097/ccm.0000000000001402 (2016).
Francois, B. et al. Interleukin-7 restores lymphocytes in septic shock: the IRIS-7 randomized clinical trial. JCI insight. 3, e98960, https://doi.org/10.1172/jci.insight.98960 (2018).
Peters van Ton, A. M., Kox, M., Abdo, W. F. & Pickkers, P. Precision Immunotherapy for Sepsis. Front. Immunol. 9, 1926–1926, https://doi.org/10.3389/fimmu.2018.01926 (2018).
Delsing, C. E. et al. Interferon-gamma as adjunctive immunotherapy for invasive fungal infections: a case series. BMC Infect. Dis. 14, 166, https://doi.org/10.1186/1471-2334-14-166 (2014).
Docke, W. D. et al. Monocyte deactivation in septic patients: restoration by IFN-gamma treatment. Nat. Med. 3, 678–681, https://doi.org/10.1038/nm0697-678 (1997).
Albert-Vega, C. et al. Immune Functional Assays, From Custom to Standardized Tests for Precision Medicine. Frontiers Immol. 9 https://doi.org/10.3389/fimmu.2018.02367 (2018).
Davies, R., O’Dea, K. & Gordon, A. Immune therapy in sepsis: Are we ready to try again? JICS. 19, 326–344, https://doi.org/10.1177/1751143718765407 (2018).
Rosadini, C. V. & Kagan, J. C. Early innate immune responses to bacterial LPS. Curr. Opin. Immunol. 44, 14–19, https://doi.org/10.1016/j.coi.2016.10.005 (2017).
Rodriguez-Rosales, Y. A. et al. Long-Term Effects of Experimental Human Endotoxemia on Immune Cell Function: Similarities and Differences With Sepsis. Shock. 51, 678–689, https://doi.org/10.1097/shk.0000000000001222 (2019).
Lankelma, J. M. et al. Antibiotic-induced gut microbiota disruption during human endotoxemia: a randomised controlled study. Gut. 66, 1623, https://doi.org/10.1136/gutjnl-2016-312132 (2017).
van Lier, D., Geven, C., Leijte, G. P. & Pickkers, P. Experimental human endotoxemia as a model of systemic inflammation. Biochimie. 159, 99–106, https://doi.org/10.1016/j.biochi.2018.06.014 (2019).
Segre, E. & Fullerton, J. N. Stimulated Whole Blood Cytokine Release as a Biomarker of Immunosuppression in the Critically Ill: The Need for a Standardized Methodology. Shock. 45, 490–494, https://doi.org/10.1097/SHK.0000000000000557 (2016).
Dorresteijn, M. J., Draisma, A., van der Hoeven, J. G. & Pickkers, P. Lipopolysaccharide-stimulated whole blood cytokine production does not predict the inflammatory response in human endotoxemia. Innate Imm. 16, 248–253, https://doi.org/10.1177/1753425909339923 (2010).
Tawfik, D. M. et al. Immune Profiling Panel: a proof of concept study of a new multiplex molecular tool to assess the immune status of critically-ill patients. bioRxiv, 636522 (2019).
Brunet, L. R., LaBrie, S. & Hagemann, T. Immune monitoring technology primer: immunoprofiling of antigen-stimulated blood. J. Immunotherapy Cancer. 4, 18, https://doi.org/10.1186/s40425-016-0122-4 (2016).
Duffy, D. et al. Functional analysis via standardized whole-blood stimulation systems defines the boundaries of a healthy immune response to complex stimuli. Immunity. 40, 436–450, https://doi.org/10.1016/j.immuni.2014.03.002 (2014).
Josse, J. & Husson, F. missMDA: A Package for Handling Missing Values in Multivariate Data Analysis. J STAT SOFTW. 1 (2016).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. STAT. SOC. B . 57, 289–300 (1995).
R Core Team. R: A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria (2018). https://www.R-project.org/.
Murphy, K. & Weaver, C. in Janeway’s Immunobiology Ch. Part V, 533-748 (Garland Science Taylor & Francis Group (2018).
de Kleijn, S. et al. Transcriptome Kinetics of Circulating Neutrophils during Human Experimental Endotoxemia. Plos One. 7, e38255, https://doi.org/10.1371/journal.pone.0038255 (2012).
Tak, T., van Groenendael, R., Pickkers, P. & Koenderman, L. Monocyte Subsets Are Differentially Lost from the Circulation during Acute Inflammation Induced by Human Experimental Endotoxemia. J. Innate Immun. 9, 464–474, https://doi.org/10.1159/000475665 (2017).
Hassani, M. et al. Differentiation and activation of eosinophils in the human bone marrow during experimental human endotoxemia. J Leukoc Biol. n/a, https://doi.org/10.1002/jlb.1ab1219-493r (2020).
Brinkhoff, A. et al. Pro-Inflammatory Th1 and Th17 Cells Are Suppressed During Human Experimental Endotoxemia Whereas Anti-Inflammatory IL-10 Producing T-Cells Are Unaffected. Front Immunol. 9 https://doi.org/10.3389/fimmu.2018.01133 (2018).
Demaret, J. et al. Identification of CD177 as the most dysregulated parameter in a microarray study of purified neutrophils from septic shock patients. Immunol. Lett. 178, 122–130, https://doi.org/10.1016/j.imlet.2016.08.011 (2016).
Cazalis, M. A. et al. Decreased HLA-DR antigen-associated invariant chain (CD74) mRNA expression predicts mortality after septic shock. Crit. Care. 17, R287, https://doi.org/10.1186/cc13150 (2013).
Davies, M. G. & Hagen, P.-O. Systemic inflammatory response syndrome. BJS. 84, 920–935, https://doi.org/10.1002/bjs.1800840707 (1997).
Khan, H. N. et al. Leukocyte transcriptional signatures dependent on LPS dosage in human endotoxemia. J Leukoc Biol. 0 https://doi.org/10.1002/jlb.4a0219-050r (2019).
Janicki-Deverts, D., Cohen, S., Doyle, W. J., Turner, R. B. & Treanor, J. J. Infection-induced proinflammatory cytokines are associated with decreases in positive affect, but not increases in negative affect. Brain Behav. Immun. 21, 301–307, https://doi.org/10.1016/j.bbi.2006.09.002 (2007).
Mueller, L. P. et al. Endotoxin-adapted septic shock leukocytes selectively alter production of sIL-1RA and IL-1beta. Shock. 16, 430–437 (2001).
Poujol, F. et al. Lymphocyte Proliferation upon Lipopolysaccharide Challenge Ex Vivo. Plos One. 10, e0144375, https://doi.org/10.1371/journal.pone.0144375 (2015).
Wegner, A. et al. Sex differences in the pro-inflammatory cytokine response to endotoxin unfold in vivo but not ex vivo in healthy humans. Innate Immun. 23, 432–439, https://doi.org/10.1177/1753425917707026 (2017).
Rol, M.-L. et al. The REAnimation Low Immune Status Markers (REALISM) project: a protocol for broad characterisation and follow-up of injury-induced immunosuppression in intensive care unit (ICU) critically ill patients. BMJ Open. 7 (2017).
Marshall, J. C. Why have clinical trials in sepsis failed? Trends Mol. Med. 20, 195–203, https://doi.org/10.1016/j.molmed.2014.01.007 (2014).
Acknowledgements
We acknowledge Francois Mallet and Sophie Blein for their advices and sharing their expertise to enrich our study. D.M.T., L.V., A.P., W.J.W. and J.T. are part of the European Sepsis Academy, funded by the European Union’s Horizon-2020 research and innovation program under grant agreement No. 676129 from the Marie Skłodowska-Curie Innovative Training Networks (ITN).
Author information
Authors and Affiliations
Contributions
D.M.T., L.V., W.J.W. and J.T. made substantial contributions to conception and design, and acquisition of samples. D.M.T., J.M.L. and E.C. performed the experiments. D.M.T. performed the statistical analyses and drafted the manuscript. D.M.T., J.M.L., L.V., E.C., A.P., W.J.W. and J.T. contributed to the manuscript structuring, critical revision for important intellectual content, read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
D.M.T., L.V., E.C., A.P. and J.T. are employed by an in-vitro diagnostic company, bioMérieux. The remaining authors have no conflict of interest to declare.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tawfik, D.M., Lankelma, J.M., Vachot, L. et al. Comparison of host immune responses to LPS in human using an immune profiling panel, in vivo endotoxemia versus ex vivo stimulation. Sci Rep 10, 9918 (2020). https://doi.org/10.1038/s41598-020-66695-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-66695-2
This article is cited by
-
Non-invasive ventral cervical magnetoneurography as a proxy of in vivo lipopolysaccharide-induced inflammation
Communications Biology (2024)
-
Cytokines and tryptophan metabolites can predict depressive symptoms in pregnancy
Translational Psychiatry (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.