Abstract
The challenge in the treatment of glioblastoma is the failure to identify the cancer invasive area outside the contrast-enhancing tumour which leads to the high local progression rate. Our study aims to identify its progression from the preoperative MR radiomics. 57 newly diagnosed cerebral glioblastoma patients were included. All patients received 5-aminolevulinic acid (5-ALA) fluorescence guidance surgery and postoperative temozolomide concomitant chemoradiotherapy. Preoperative 3 T MRI data including structure MR, perfusion MR, and DTI were obtained. Voxel-based radiomics features extracted from 37 patients were used in the convolutional neural network to train and as internal validation. Another 20 patients of the cohort were tested blindly as external validation. Our results showed that the peritumoural progression areas had higher signal intensity in FLAIR (p = 0.02), rCBV (p = 0.038), and T1C (p = 0.0004), and lower intensity in ADC (p = 0.029) and DTI-p (p = 0.001) compared to non-progression area. The identification of the peritumoural progression area was done by using a supervised convolutional neural network. There was an overall accuracy of 92.6% in the training set and 78.5% in the validation set. Multimodal MR radiomics can demonstrate distinct characteristics in areas of potential progression on preoperative MRI.
Similar content being viewed by others
Introduction
Glioblastoma (GBM) is the most aggressive and the most common primary brain tumour. Despite the extensive standard multimodal treatment regimens including surgical resection followed by concomitant chemo-radiotherapy, the median overall survival is only about 14.9 month1. This poor prognosis largely due to the failure of the local treatment which usually aims the contrast-enhancing lesion as the treatment target. Studies had shown that the GBM cancer cell can extent to the non-contrast enhancing area2,3. Therefore, the recurrence is inevitable and even in patients with a total resection of the contrast enhancing tumour and subsequent chemoradiotherapy, recurrence happens in all patients and in up to 90% in or directly adjacent to the resection area4.
However, the imaging characteristics to this peritumoural non-enhanced invasive area is still not clear. The conventional structure MRI has numerous limitations on the detection of the peritumoural invasive area. Although a high signal in T2 represents vasogenic oedema that associated with the tumour, however, in a biopsy study a normal signal in conventional T2 MRI can have 4% of false negative rate5. Therefore, many different MR techniques have been developed to achieve a better demarcation of the GBM. Apparent diffusion coefficient (ADC) is sensitive to the diffusion of water, and can decrease while the extracellular water diffusion is restricted due to the increase of glioma cellularity6. Diffusion tensor imaging (DTI) can detect subtle white matter change caused by the tumour infiltration7, further decompose DTI in to isotropic p component and anisotropic q component can be used predict the tumour recurrence type by their pattern8. In a recent study focused on the peritumoural abnormal DTI-p areas showed that an increase of the relative cerebral blood volume (rCBV), Choline, Choline/ N-acetyl aspartate (NAA), and the glutamate+ glutathione (Glx)9. Another study focused on the DTI fractional anisotropy (FA) in the peritumoural non-enhancing area showed that a decrease of FA may indicate tumour infiltration which further lead to local tumour recurrence10.
Radiomics is an emerging technique that can provide a more detailed quantification multi-modal MR study. And is defined as the conversion of images to higher dimensional data and the subsequent mining of these data11. In the conventional imaging analysis, only limited semantic features can be extracted, however, by using high-throughput computing, radiomics can quantify more detailed agnostic features including first order, second order or higher order. Hu et al., in 2015 used multi-parametric MRI and texture analysis to predict tumour density in both enhanced and non-enhanced part based on a biopsy study and showed 85% accuracy in training set and 81.8% accuracy in the validation12. Further study conducted by Kim et al., showed that the FA and the CBV radiomics of the peritumoural area, together with the clinical index, can potentially predict the tumour progression13.
Thus, in this study we aimed to characterize the preoperative peritumoural non-enhanced area that demonstrated to have generated tumour progression later on. And further applied these features, together with radiomics features to identify areas of GBM progression on the preoperative MRI by using the convolutional neural network.
Materials and Methods
Patient inclusion criteria
We retrospectively included 57 patients with a newly diagnosed cerebral glioblastoma in this study. Exclusion criteria were previous cranial surgery, previous cerebral radiotherapy or a known other primary tumour. General characteristics were shown in Table 1. 37 patients who had follow-up MRI in Cambridge university hospital were used for training and internal validation. Another 20 patients who received follow-up MRI in other hospital were collected as external validation. There were 5 patients in the training group had radiological pseudo-progression during the follow-up, but all patients had tumour progression later. Tumour location was classified according to a previous classification14. All patients had a Karnofsky performance status ≥70 and were treated with a maximal surgical resection using neuronavigation (StealthStation, Medtronic) and 5-aminolevulinic acid (5-ALA) fluorescence guidance with the aim of maximal tumour resection. This was followed by standard concomitant chemoradiotherapy and adjuvant chemotherapy15.
The study protocol was approved and regulated by the National Research Ethics Service (NRES) Committee East of England - Cambridge Central (ethics reference no. 10/H0308/23). And informed written consents were obtained from all patients and all methods were performed in accordance with the relevant guidelines and regulations.
MRI Data acquisition and imaging processing
Detailed MRI data acquisition and imaging processing protocols were as previous described16,17. In brief, all preoperative MRI data was obtained from the same 3.0 Tesla Siemens MR Magnetron system (Siemens Healthcare, Munich, Germany) with a standard 12-channel head coil. The imaging protocol included anatomical contrast 3D T1-weighted magnetization-prepared rapid acquisition with gradient echo (MPRAGE), T2-weighted and fluid-attenuated inversion recovery (FLAIR) sequences. The preoperative DTI, dynamic susceptibility weighted contrast enhanced (DSC) MR perfusion and 1H MR spectroscopy data were also acquired. DTI data was performed using a single-shot echo-planar sequence scanned in 13 directions.
Follow-up MR protocols included a 2D contrast enhanced T1-weighted, a T2 and a FLAIR sequences. Patients may receive the scanning from different scanner according to the clinical arrangement (a 1.5 T GE Optima, 1.5 or 3.0 T GE Signa or 1.5 T Siemens Avanto).
Imaging processing of the preoperative DTI were done by using tools from the FMRIB Software Library (FSL) version 5.0.0 (http://fsl.fmrib.ox.ac.uk/fsl/fslwiki/). DTI was further decomposed into isotropic (p) component and anisotropic (q) component after eigenvalues were calculated in the DTI data as described previously described18. Perfusion MR was processed using NordicICE (NordicNeuroLab AS, Bergen, Norway) and maps of rCBV were created following contrast agent leakage correction. Spectroscopy data was processed using LCModel19 and the MRS result in each ROIs were obtained by using a previous described 3D voxel-wise approach17.
Regions of interests
Two main regions of interest (ROIs), progression areas and non-progression areas were created in this study (Fig. 1).
All patients had radiological tumour progression in our cohort. The progression area was drawn from the first tumour progression T1 contrast MRI (Fig. 1B) according to the RANO criteria20 by the first authors (J.L.Y.) and approved by the 3rd author (A.H.). In order to avoid pseudoprogression, we examined at least two follow-up scans after the first radiological tumor progression, except for a few patients with rapidly clinical deterioration, and did not receive another follow up scan after the progression.
The areas of the tumour progression (Fig. 1C,D) were created by coregistration the progression contrast enhanced T1 MRI (Fig. 1B) to the pre-operative diagnostic MRI (Fig. 1A). This was done by using a previous described two stage non-linear semi-automatic coregistration21. In short, firstly, we calculated the transformation matrix between preoperative tumour and postsurgical resection cavity by using the linear FLIRT co-registration. Then we applied this transformation matrix to a non-linear FNIRT transformation to coregister the brain. After subtraction of the preoperative contrast enhanced lesion, the peritumoural tumour progression (Fig. 1E) and non-progression areas (Fig. 1F) can be identified on the preoperative MRIs.
The non-progression areas were created from the peritumoural 5, 10, 15, 20 mm excluding the progression areas. In addition, a contralateral area of normal appearing with matter (NAWM) as control representing normal brain tissue.
Radiomics analysis and machine learning
The multimodal MRI characteristics preoperatively are extracted using Matlab (MathWorks Inc., Natick, MA, USA). First order and second order radiomics features were extracted from 7 different MR sequences by using Matlab. The first order features include mean, standard deviation, median, minimum, maximum, variance, skewness, kurtosis, energy, entropy, uniformity, root mean square, mean gray level. The second order features include GLCM (gray level co-occurrence matrix which includes, auto-correlation, contrast, correlation-M, correlation-P, cluster-Prominence, cluster-shad, dissimilarity, energy, entropy, homogeneity-M, homogeneity-P, sum of square, Sum of variance, sum of entropy, difference of Variance, Difference of entropy, Information measure if correlation 1, Information measure of correlation 2, and inversed difference normalised) and GLRLM (gray level run length matrix which includes, short run length, long run length, gray level non-uniformity, run percentage, run length non-uniformity, low gray level emphasis, and high gray level emphasis). A total number of 294 features were extracted.
Identification of the progression areas were done by using supervised convolutional neural network from the Matlab neural network toolbox (nntool). The training of the machine learning was done voxel by voxel. Voxel-wise feature extraction from 37 patients (Table 1, Training group) who had follow-up MRI in Cambridge university hospital were used for training and internal validation. A total number of 2,589,473 peritumoural voxel with 294 radiomics features were input for training. The output was binary, either progression or non-progression. The input data was divided randomly with 70% used for training, 15% for validation, and 15% for testing. There was 10 hidden layers in our neural network, and we used scaled conjugated gradient back propagation in our training. The network performance was calculated by using cross-entropy. The trained model was further tested in another 20 patients who received follow-up MRI in other hospital were collected as external validation (Table 1, validation group). The results were tested 3 repeated time. Further identification of the areas of progression were drawn by using a probability heat map in Matlab.
Statistical analysis
General patients’ characteristics were tested for group differences with a t-test or Mann-Witney U test for continuous variable depending on the normality of the data. A chi-square test was used for categorical data. Differences between MRI characteristics of the preoperative area later showing tumour progression and non-progression areas were done using paired t-test. Two-sided p-values were used. All statistical tests were performed using SPSS version 22 (IBM Inc., New York, USA).
Results
Multimodal MRI characteristics
Multimodal MRI characteristics were shown in Fig. 2. The ADC values in the progression area were lower than 5 mm and 10 mm peri-tumoural non-progression area (Fig. 2A, p < 0.001, 0.029). Fractional anisotropy (FA) showed lower in progression area than 5 mm non-progression area (Fig. 2B, p = 0.041). The isotropic p component showed a significant decrease in progression area than 5 mm and 10 mm of non-progression area (Fig. 2D, p < 0.001). In areas of progression, anisotropic q did not show difference between progression and non-progression area (Fig. 2E). In areas of later progression, there was significant increase in the FLAIR signal (Fig. 2C, p = 0.020 ~ <0.001) and contrast enhanced T1 MRI (Fig. 2G, p = 0.026 ~ 0.0004) compared to non-progression area. Relative cerebral blood volume was increased in the progression areas than 15–20 mm non-progression area (Fig. 2F, p = 0.038 ~ 0.042). A higher Cho/NAA and lower NAA can be seen in the progression areas but without statistical significance (Fig. 2H)
Showed MRI characteristics (A–G) of contrast enhanced, progression, non-progression (NP) area in 5–20 mm peritumourtumoural area and contralateral normal appearing white matter (NAWM, control). (H) Showed MRS of Cho/NAA, Choline, NAA + NAAG and Glu + Gln of progression (P), non-progression (NP) and NAWM. *p < 0.05; **p < 0.01; ***p < 0.001.
Radiomics features
Thirty-five out of the 91 first order radiomics features had significant difference between progression area and non-progression area (Fig. 3). Most distinct features were in ADC, p and contrast enhanced T1 MRI (Fig. 3A,C,F). In the 203 second order radiomics features, 77 were found to have significant difference between progression and non-progression areas (Fig. 4). Total 112 radiomics features were identified in the progression areas.
Identification of the progression areas from the pre-operative MRIs
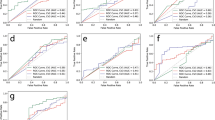
In the training set (n = 37) of the supervised convolutional neural network, the overall accuracy 92.6% (Fig. 5). The training, testing and validation accuracy were 92.7%, 92.4% and 92.4% respectively. The overall sensitivity was 80% and the overall specificity was 97.7%.
Further application of the trained model can be used to generate predicted areas of progression on the preoperative MRI, and the example case was shown in Fig. 6.
A representative result of the areas of progression (color map) drawn automatically with machine learning and compared to its actual images at progression. The color map showed the probability of progression (red: high, blue: low). White arrows indicate that a high chance of progression at periventricular area.
External validation in the 20 cases were done by overlapping the resulted progression map drawn by the convolutional neural network model and the actual progression MRI. 3 repeated validation results were shown in Table 2. The overall accuracy in the validation group was 78.0%. The positive predict value and the negative predict value were 16.8% and 78.5%.
`Discussion
In this study, we showed distinctive imaging characteristics in the peritumoural areas that potentially progress. Lower ADC, p and higher FLAIR, contrast enhanced T1 signal were noted in these areas. Further comparison by using radiomics features showed more difference supports the identification of these peritumoural invasive areas.
We found a significant lower ADC pixel value in the progression areas this may due to the increase cancer cellularity in this invasive peritumoural areas. ADC is usually used to quantify the diffusivity of the water. In the circumstances of the normal brain tissue or increase extracellular fluid, ADC may increase due to the un-restriction of the water diffusion. On the other hand, ADC may decrease due to increase cellularity which resulted to the restriction of the extracellular fluid diffusion22. ADC can be seen in higher grade glioma6 resulted in a worse prognosis. In addition, an increase in ADC after treatment can be a favourable prognostic predictor23.
DTI p is another representative form of the mean diffusivity. Therefore, similar results with the ADC were expected. Previous biopsy had shown an increase of 10% in the p can show tumour infiltration and further MRS and rCBV study also showed that in these p abnormal peritumoural area can have higher Cho/NAA ratio and higher rCBV which refers to high cellular turnover rate and angiogenesis. This further indicated cancer cell activity. Our results showed that q also features contributes to the identification of the progression area. In the previous biopsy study, q was found largely represents cancer itself and was found larger than the contrast enhancing lesion in approximate 50% of the GBM. Extent of resection showed that a larger resection area of the abnormal q can result a better prognosis. Further study in these peritumoural abnormal q areas showed that there were increase rCBV and increase Choline which also indicates cancer activity. In this study, we found less features by using DTI-q, this may because there was another half of the cases that had q abnormal areas smaller or equal to the contrast enhancing areas that may potentially masked the difference.
FLAIR and contrast enhanced T1C were the two conventional MRI that was found to have distinct features in our study. Although FLAIR represents the vasogenic edema of the brain and is non-specific. Many studies used the FLAIR to define the non-enhancing peritumoural area and showed that abnormal cancer cell. Lemée et al., found that about 1/3 of the histology analysis in the FLAIR areas had cancer cell infiltration. However, the FLAIR may be affected by steroid treatment or antiangiogenic therapy such as Avastin. In our study, we found that although the peritumoural area were non-enhanced, there was higher signal in the progression areas. This can be due to the subtle blood brain barrier interference by the cancer infiltration that beyond human visual detection capability.
In addition to the difference of pixel value extracted from difference MRI sequences. We used quantitative radiomics feature to characterise the peritumoural progression area from the preoperative MRIs. We found a total 112 out of 294 features that are different between progression and non-progression areas. Most studies focused on the features of the contrast enhancing lesion. Gevaert et at, in 2014, showed the radiomics features can correlate with manual radiologist’s Visually Accessible Rembrandt Images features, patient clinical survival. Another study used quantified radiomics data from 121 GBM patients to cluster 3 distinct MR phenotypes, the peri-multifocal, spherical and rim-enhanced. They found each cluster had its different pathway which possibly explain the distinct prognosis.
Our results in the radiomics provide reasonable size of the features to the application of the convolutional neural networks. And total 294 features coded in every voxel generated a large number of the sample size that can be used to train the model. Although the optimal features number and the sample size were not tested in this study. Our attempt to use the convolutional neural network to draw the possible progression map from the preoperative MRI had shown moderately fair results. Although the positive predict value and the sensitivity were lower, however, the specificity and negative predict were good.
We have several limitations in this study. We are not able to take the MRI sampling time into this study. GBM progression may change from time to time, our progression area was created by coregistration of the first true progression image to the reference image. This may underestimate the area of progression. In addition, the follow-up MRI protocols were different in two groups. Although the preoperative MR radiomics were obtained by the same MR protocols, different MR protocol at the time of tumour progression may have slightly effects on the spatial co-registration. Another limitation is the limited sample size. As most of the radiomics study require a large sample size in the machine learning in order to reach a stable training model. Our initial training accuracy was 92.7% then drop to 78% in the external validation. This can be over-fitted in the initial training due to inadequate sample size. Or may due to the difference between training and the external validation groups. The overall survival and progression free survival were both shorter in the external validation group, we cannot eliminate the potential bias between this two groups.
Conclusion
Multimodal quantitative MR imaging analysis, including structure MRI, perfusion MR and diffusion tensor imaging can demonstrate distinct characteristics in areas of potential later progression on preoperative MRI. Moreover, the application of these imaging features, site of tumour progression can be potentially identified via a trained machine learning model which provides a direction for future study and treatment target.
Change history
11 August 2020
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Brodbelt, A. et al. Glioblastoma in England: 2007-2011. Eur. J. Cancer 51, 533–542, https://doi.org/10.1016/j.ejca.2014.12.014 (2015).
Lemee, J. M. et al. Characterizing the peritumoral brain zone in glioblastoma: a multidisciplinary analysis. J. Neurooncol 122, 53–61, https://doi.org/10.1007/s11060-014-1695-8 (2015).
Lemee, J. M., Clavreul, A. & Menei, P. Intratumoral heterogeneity in glioblastoma: don’t forget the peritumoral brain zone. Neuro Oncol. 17, 1322–1332, https://doi.org/10.1093/neuonc/nov119 (2015).
Petrecca, K., Guiot, M. C., Panet-Raymond, V. & Souhami, L. Failure pattern following complete resection plus radiotherapy and temozolomide is at the resection margin in patients with glioblastoma. J. neuro-oncology 111, 19–23, https://doi.org/10.1007/s11060-012-0983-4 (2013).
Price, S. J. & Gillard, J. H. Imaging biomarkers of brain tumour margin and tumour invasion. Br. J. Radiol. 84(Spec No 2), S159–167, https://doi.org/10.1259/bjr/26838774 (2011).
Sugahara, T. et al. Usefulness of diffusion-weighted MRI with echo-planar technique in the evaluation of cellularity in gliomas. J. magnetic Reson. imaging: JMRI 9, 53–60 (1999).
Price, S. J. Improved delineation of glioma margins and regions of infiltration with use of diffusion tensor imaging: an image-guided biopsy study. Am. J. neuroradiology 27, 1969–1974 (2006).
Price, S. J. et al. Predicting patterns of glioma recurrence using diffusion tensor imaging. Eur. Radiol. 17, 1675–1684, https://doi.org/10.1007/s00330-006-0561-2 (2007).
Price, S. J. et al. Multimodal MRI can identify perfusion and metabolic changes in the invasive margin of glioblastomas. J. Magn. Reson. Imaging 43, 487–494, https://doi.org/10.1002/jmri.24996 (2016).
Bette, S. et al. Local Fractional Anisotropy Is Reduced in Areas with Tumor Recurrence in Glioblastoma. Radioloy 283, 499–507 (2017).
Gillies, R. J., Kinahan, P. E. & Hricak, H. Radiomics: Images Are More than Pictures, They Are Data. Radiology 287, 563–577 (2016).
Hu, L. S. et al. Multi-Parametric MRI and Texture Analysis to Visualize Spatial Histologic Heterogeneity and Tumor Extent in Glioblastoma. PLoS One 10, e0141506, https://doi.org/10.1371/journal.pone.0141506 (2015).
Kim, J. Y. et al. Radiomics in peritumoral non-enhancing regions: fractional anisotropy and cerebral blood volume improve prediction of local progression and overall survival in patients with glioblastoma. Neuroradiology 61, 1261–1272, https://doi.org/10.1007/s00234-019-02255-4 (2019).
Sawaya, R. E. et al. Neurosurgical outcomes in a modern series of 400 craniotomies for treatment of parenchymal tumors. Neurosurgery 42, 1044–1055 (1998).
Hegi, M. E. et al. MGMT gene silencing and benefit from temozolomide in glioblastoma. N. Engl. J. Med. 352, 997–1003, https://doi.org/10.1056/NEJMoa043331 (2005).
Yan, J. L. et al. Extent of resection of peritumoral diffusion tensor imaging-detected abnormality as a predictor of survival in adult glioblastoma patients. J. Neurosurg. 126, 234–241, https://doi.org/10.3171/2016.1.JNS152153 (2017).
Yan, J.-L. et al. Multimodal MRI characteristics of the glioblastoma infiltration beyond contrast enhancement. Therapeutic Advances in Neurological Disorders 12, https://doi.org/10.1177/1756286419844664 (2019).
Pena, A. et al. Enhanced visualization and quantification of magnetic resonance diffusion tensor imaging using the p:q tensor decomposition. Br. J. Radiol. 79, 101–109, https://doi.org/10.1259/bjr/24908512 (2006).
Provencher, S. W. Estimation of metabolite concentrations from localized in vivo proton NMR spectra. Magn. Reson. Med. 30, 672–679 (1993).
Wen, P. Y. et al. Response Assessment in Neuro-Oncology Clinical Trials. J. Clin. Oncol. 35, 2439–2449, https://doi.org/10.1200/JCO10.1200/JCO.2017 (2017).
van der Hoorn, A. et al. Validation of a semi-automatic co-registration of MRI scans in patients with brain tumors during treatment follow-up. NMR Biomed. 29, 882–889, https://doi.org/10.1002/nbm.3538 (2016).
Deng, Z. et al. Quantitative analysis of glioma cell invasion by diffusion tensor imaging. J. Clin. Neurosci. 17, 1530–1536, https://doi.org/10.1016/j.jocn.2010.03.060 (2010).
Hilario, A. et al. A prognostic model based on preoperative MRI predicts overall survival in patients with diffuse gliomas. AJNR. Am. J. neuroradiology 35, 1096–1102, https://doi.org/10.3174/ajnr.A3837 (2014).
Acknowledgements
This research was funded by the grants from Keelung Chang Gung Memorial Hospital (CMRPG2H0371 J.L.Y.), and the UK National Institute of Health Research Clinician Scientist Fellowship (S.J.P.).
Author information
Authors and Affiliations
Contributions
Yan J.L. wrote the main article, developed the hypothesis and carried out the study. Li C. verified the analysis methods, helped writing the manuscript. van der Hoorn A. helped to develop the method and supported on the manuscript writing. Boonzaire N. supported the manuscript writing. Matys T. supervised the study and helped to collect study materials. Price S. developed the idea of the study, supervised the entire work and verified the study results. All authors reviewed the manuscript
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yan, JL., Li, C., van der Hoorn, A. et al. A Neural Network Approach to Identify the Peritumoral Invasive Areas in Glioblastoma Patients by Using MR Radiomics. Sci Rep 10, 9748 (2020). https://doi.org/10.1038/s41598-020-66691-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-66691-6
This article is cited by
-
Discrimination Between Glioblastoma and Solitary Brain Metastasis Using Conventional MRI and Diffusion-Weighted Imaging Based on a Deep Learning Algorithm
Journal of Digital Imaging (2023)
-
Qualitative MR features to identify non-enhancing tumors within glioblastoma’s T2-FLAIR hyperintense lesions
Journal of Neuro-Oncology (2023)
-
Radiomics for precision medicine in glioblastoma
Journal of Neuro-Oncology (2022)
-
Quantitative mapping of individual voxels in the peritumoral region of IDH-wildtype glioblastoma to distinguish between tumor infiltration and edema
Journal of Neuro-Oncology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.