Abstract
Rubylation is a conserved regulatory pathway similar to ubiquitination and essential in the response to the plant hormone auxin. In Arabidopsis thaliana, AUXIN RESISTANT1 (AXR1) functions as the E1-ligase in the rubylation pathway. The gene AXR1-LIKE (AXL), generated by a relatively recent duplication event, can partially replace AXR1 in this pathway. We have analysed mutants deficient for both proteins and complementation lines (with the AXR1 promoter and either AXR1 or AXL coding sequences) to further study the extent of functional redundancy between both genes regarding two processes: meiosis and DNA repair. Here we report that whereas AXR1 is essential to ensure the obligatory chiasma, AXL seems to be dispensable during meiosis, although its absence slightly alters chiasma distribution. In addition, expression of key DNA repair and meiotic genes is altered when either AXR1 or AXL are absent. Furthermore, our results support a significant role for both genes in DNA repair that was not previously described. These findings highlight that AXR1 and AXL show a functional divergence in relation to their involvement in homologous recombination, exemplifying a duplicate retention model in which one copy tends to have more sub-functions than its paralog.
Similar content being viewed by others
Introduction
Ubiquitination is a post-translational modification pathway that affects a wide variety of processes and is highly conserved among eukaryotes1,2,3. Through this process, the small peptide ubiquitin (UBQ) is covalently conjugated to target proteins, altering their stability (by the 26 S proteasome) and activity4,5. The attachment of UBQ is orchestrated by a cascade of three specialized enzymes: a UBQ-activating enzyme (E1), a UBQ-conjugating enzyme (E2) and a UBQ-ligase protein (E3)6,7,8,9,10,11. There are a number of UBQ-like modifiers that utilize similar chemical reactions for covalent ligation to their substrates, such as NEURONAL PRECURSOR CELL EXPRESSED DEVELOPMENTALLY DOWN-REGULATED8 (NEDD8) in animals and fission yeast, known as RELATED TO UBIQUITIN (RUB) in plants. While NEDD8 is encoded by a single copy gene in animals, there are three RUB genes in Arabidopsis thaliana12, where RUB1 and RUB2 encode UBQ-RUB fusion proteins13.
RUB closely resembles UBQ and uses a separate but similar E1-E2-E3 biochemical pathway for activation, conjugation, and ligation. In Arabidopsis, the E1 C-TERMINAL-RELATED1 (ECR1) gene encodes the C-terminal RUB E1 subunit, whereas two paralogous genes, AUXIN RESISTANT1 and AXR1-LIKE (AXR1 and AXL, respectively) encode the N-terminal subunits. Following activation, RUB1 CONJUGATING ENZYME1 (RCE1) and RING-BOX 1 (RBX1) act as RUB-conjugating enzyme E2 and RUB-ligase E3, respectively14,15,16. Rubylation mainly acts as a regulator of cullin-RING E3 UBQ ligases (CRLs). RUB conjugation stabilizes cullins and produces a change of these E3 UBQ ligases that allows the efficient transfer of UBQ to proteins for degradation17. Only a few non-cullin substrates have been recently identified, but the biological relevance of these modifications is not yet known (reviewed in Schwechheimer, 2018)18.
Rubylation regulates a wide range of cellular processes in Arabidopsis, including cell cycle progression, light response, vegetative growth, embryo development, and auxin signaling19,20,21,22,23. In this context, although AXR1 expression is enriched in tissues with high rates of cellular division, in which the importance of DNA repair is critical, the connection of rubylation with DNA damage response has not currently been established in plants24,25. Nevertheless, several publications have described the importance of this pathway in controlling how cells react to DNA breaks in mice and human cell cultures26,27,28.
In Arabidopsis, rubylation is also involved in homologous recombination (HR) during meiosis, the specialised cell division that generates gametes29. HR is a key process during this division, since it ensures bivalent formation. Plants deficient in AXR1 display problems in the formation of the synaptonemal complex (SC), the tripartite structure that intimately connects homologous chromosomes during some stages of prophase I, and also changes in the position of crossovers (COs), the reciprocal DNA exchanges that link homologous chromosomes and ensure their correct segregation during the first meiotic division. The mislocalisation of COs in axr1 leads to the presence of univalents and unbalanced segregations that are the origin of aneuploidies in the gametes29. These errors are probably due partly to a deregulation of modular cullin RING ligase 4 (CRL4) whose meiotic targets have not been characterised. The effect of rubylation (also known as neddylation) and CRL4 on synapsis and CO formation has also been described in Caenorhabditis elegans30,31.
AXR1 and AXL share 80% of amino acid identity. They have redundant functions in RUB activation and show similar biochemical activity in vitro32,33,34,35. Because of the key importance of rubylation in plant growth and development, axr1 axl double mutants are usually lethal at embryonic or early seedling stage. However, whereas axl plants present wild-type (WT) phenotypes, axr1 plants are dwarf and exhibit multiple growth defects, small flowers and short siliques29,33,36. For a long time, it was thought that AXR1 and AXL were functionally equivalent, since Dharmasiri and colleagues33 demonstrated that the phenotype of axr1 plants could be restored by overexpression of AXL. Later, Hotton and colleagues35 proved that the two genes are not equivalent: AXL cannot replace AXR1 when it is expressed at comparable levels to AXR1 in an axr1 background. Trying to delve into this issue, we have analysed the consequences of the absence of AXR1 and AXL, focusing our attention on the HR process, essential for genome maintenance and meiotic chromosomal segregation. Results presented here demonstrate that both proteins are not functionally equivalent during meiotic recombination, although both are essential for efficient DNA repair. Furthermore, the corresponding mutants (axr1 and axl) exhibit a significant reduction in HEI10 expression in meiotic tissues. This is a gene homolog of HUMAN ENHANCER OF INVASION CLONE10 which encodes a SUMO (Small Ubiquitin-like Modifier) E3 ligase necessary for the formation of the majority of COs in Arabidopsis37. We conclude that the functional divergence of AXR1 and AXL is clearly more pronounced in meiotic than in somatic recombination, although AXR1 also plays a more predominant role than AXL in this process. Altogether, these findings suggest that both proteins have probably evolutionary adjusted their preferences for different interactors depending on specific cellular stages or environments.
Results
axr1 and axl mutants show differences in fertility
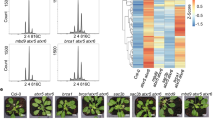
RT-PCR analyses revealed that neither full-length transcripts of AXR1 nor AXL were produced in the T-DNA insertion lines axr1-31 (SALK_013238) and axl-2 (GK-818B10) mutants, respectively (Fig. 1A, Supplementary Fig. S1). However, in agreement with previous observations19,24, homozygous axl plants were indistinguishable from WT plants in terms of germination, vegetative growth, and flowering time, whereas homozygous plants for axr1 alleles displayed severe growth alterations and developmental defects. These plants also showed leaves with irregular shape and slightly jagged edges, and reduced height (maximum ~20 cm in mature plants versus 50 cm in WT plants) (Fig. 1B). In accordance with the phenotype displayed by other axr1 mutants29, axr1-31 showed an increase of irregular pollen grains, smaller than WT, with irregular shape, shrunken and collapsed (26.72%, 144/539 in axr1-31; 1.00%, 6/600 in WT ; χ21 = 164.206, p < 0.001; Fig. 1C) and a severe reduction in the number of seeds per silique (5.67 ± 1.25 in axr1-31; 48.53 ± 1.17 in WT; two-tailed Mann Whitney U-test p < 0.001). Siliques were also shorter than in WT (4.40 ± 0.35 mm in axr1-31; 12.47 ± 0.22 mm in WT; two-tailed Mann Whitney U-test p < 0.001).
Insertion sites and phenotypic features of mutant lines for the genes AXR1 and AXL. (a) AXR1 (At1g05180) and AXL (At2g32410) gene structures and localisation of mutant alleles. The T-DNA is located 3,386 nt and 686 nt downstream from the ATG in axr1-31 and axl-2, respectively, and its sequence is represented by white letters on black background. The mutant alleles axr1-30 and axl-1 were described in previous publications33,35. Exons: black boxes; introns: lines joining the boxes; 5′ and 3′ UTRs: white boxes. Orange and blue arrows represent primers used for genotyping and RT-PCR, respectively. (b) Representative pictures of WT, axr1-31 and axl-2 plants (40 days after sowing). (c) Squashed anthers from WT and axr1-31 plants containing mature pollen grains. Arrowheads point to abnormal (small) pollen grains in axr1-31.
AXR1 and AXL are not functionally equivalent during meiosis
AXR1 and AXL genes present high expression levels in flower stage 9, when male meiosis occurs38. It has been reported that reduced fertility in axr1 mutants is related to alterations in meiosis29, but to date no observations of meiosis in pollen mother cells (PMCs) of axl mutants had been reported, perhaps because they display apparent normal fertility33. Figure 2 shows a comparison of different meiotic stages in PMCs of WT and axr1-31 and axl-2 plants, as representative examples of axr1 and axl mutants, respectively. No differences among the genotypes were found at leptotene stage (Fig. 2A,I,Q), but axr1 showed incomplete synapsis in pachytene cells (n = 48; Fig. 2B,J). Problems in synapsis were confirmed using antibodies against the central element protein ZYP1 and the axial element-associated protein ASY1 (Supplementary Fig. S2). Fully synapsed bivalents were observed in axl at this stage (Fig. 2R). Depletion of the function of either AXR1 or AXL produced loosened chromatin at diakinesis (100%, n = 15 and 27.3%, n = 11, respectively; Fig. 2C,K,S). This feature persisted even up to metaphase I in some axr1 cells (Fig. 2L). Only axr1 displayed univalents at metaphase I (1.43 ± 0.13 pairs of univalents per cell, n = 72 vs 0 in WT and axl, Fig. 2L) that lead to abnormal chromosome segregations at anaphase I and unbalanced nuclei at second meiotic division (86.4%, n = 44; Fig. 2M–P). This shortage in bivalent formation observed in axr1-31 was similar to that described for other axr1 alleles (1.0 and 1.3 pair of univalents per cell in axr1-12 and axr1-SAIL_904_E0629). Although five bivalents were always present in axl (Fig. 2T), some anaphase I and telophase I meiocytes showed chromatin bridges connecting the two nuclei (33%, n = 54, Fig. 2U,V). No evidence of chromosome fragmentation was observed in any mutant, suggesting that all programmed double-strand breaks (DSBs) produced at early prophase I were repaired (Fig. 2L–P,T–X).
Meiosis in PMCs from WT (a–h), axr1-31 (i–p), and axl-2 (q–x). (a,i,q) Leptotene meiocyte (b,j,r) Pachytene meiocyte. Arrows indicate incomplete synapsis in axr1-31. (c,k,s) Diakinesis. (d,l,t) Metaphase I. Univalents are only present in axr1-31 (arrows). (e,m,u) Anaphase I. (f,n,v) Telophase I. A bridge connecting the two poles can be distinguished in axl-2 (arrow). (g,o,w) Metaphase II. In axr1-31 the nuclei are unbalanced. (h,p,x) Telophase II. There are chromatids in the organelle band (arrow).
Arabidopsis chromosomes can be easily identified by FISH according to their morphology and the position of 45 S and 5 S rDNA sequences (Fig. 3A–D). To further analyse the meiotic phenotype produced by the axl mutation and determine the possible existence of variations in the behaviour of the different chromosomes, we conducted an in-depth cytological analysis of metaphase I meiocytes to estimate meiotic recombination by measuring chiasmata per cell. As it has been mentioned before, unlike the situation in axr1 (Fig. 3C), we did not observe univalents in axl (Fig. 3D). Table 1 shows the mean chiasma frequencies per cell, per chromosome and per arm in two axl mutant lines (axl-1 and axl-2), and WT plants. Despite dissimilarities in individual chromosomes compared to WT, differences in the overall mean chiasma frequencies were not detected among the different genotypes (Table 1; Fig. 3E). In WT plants, the submetacentric/metacentric group of chromosomes formed by chromosomes 1, 3 and 5 normally accounts for 21–25% of the total chiasmata per cell, whereas small acrocentric chromosomes 2 and 4 contribute to 15–17% of the total chiasmata39. We observed significant decreases in the contribution of chromosome 3 to the total chiasma frequency in axl-1, and of chromosomes 1, 3 and 5 in axl-2. In contrast, in both mutants there was an increase in the contribution of chromosomes 2 and 4 (only statistically significant in chromosome 4) (Table 1; Fig. 3E).
Cytogenetic analysis of metaphase I cells by FISH. Probes against 45 S (green) and 5 S (red) rDNA were used to analyse the behaviour of specific chromosomes. (a) Ideogram showing the location of the probes on the chromosomes. (b) Metaphase I cell from WT. (c) Metaphase I cell from axr1-31. (d) Metaphase I cell from axl-2. E. Contribution of each chromosome pair to the total chiasma frequency. +p ≤ 0.01; #p ≤ 0.001.
We have obtained additional evidence that highlights the different functions of AXR1 and AXL during meiosis using complementation lines of the mutation axr1-30. This mutation (1.57 ± 0.19 pairs of univalents per cell, n = 33; Fig. 4G–L, Supplementary Fig. S3) produces the same defects (in terms of univalent frequency) as axr1-31 (1.43 ± 0.13; two-tailed Mann Whitney U-test p = 0.582). Its complementation with an expression cassette containing the AXR1 promoter, a sequence for an N-terminal 10xMYC epitope tag, and the AXR1 coding sequence (AXR1p::10MYC-AXR1)35, resulted in bivalent frequency restoration (n = 24; Fig. 4M–R, Supplementary Fig. S3). On the other hand, the complementation of axr1-30 with a similar cassette but containing the AXL coding sequence (AXR1p::10MYC-AXL)35 did not restore the WT bivalent frequency (0.72 ± 0.11 pairs of univalents per cell, n = 32; Fig. 4S–X, Supplementary Fig. S3; two-tailed Mann Whitney U-test p = 0.002), although it increased significantly the bivalent frequency in comparison to axr1-30 (two-tailed Mann Whitney U-test p = 0.0012). The restoration was not produced despite the fact that the expression levels of both genes (AXR1 and AXL) were controlled by the same promoter (AXR1p).
AXR1 and AXL are involved in DNA repair
During the cell cycle, DSBs are produced by endogenous or exogenous sources, but during meiosis, recombination events are initiated by programmed DSBs40. Indeed, many of the genes involved in meiotic recombination have a role during somatic recombination and mutants with altered CO frequency have also reduced ability to repair DNA damage produced from a variety of exogenous sources. For this reason, we decided to determine the DNA repair capability of axr1 and axl mutants in relation to that of WT plants in response to the following agents: γ rays, mitomycin C (MMC), cisplatin (CDDP), and UV-C radiation.
Gamma rays are a powerful inductor of complex DSBs that can be repaired through different pathways, one of them being HR41. Both axr1-31 and axl-2 mutants were hypersensitive to this agent, though axr1-31 to a greater extent, and showed a reduced mean number of true leaves per plant in the different doses applied (Fig. 5A,B; Supplementary Table S2). Similar results were obtained analysing the relative dry weight per plant (Supplementary Fig. S4; Table S3). MMC mainly forms crosslinks (CLs) between complementary strands of DNA. These lesions are highly cytotoxic, and their repair is primarily triggered by stalled replication forks and depends on HR42,43,44. Only axr1-31 plants displayed hypersensitivity after MMC treatment in comparison with WT plants at higher concentrations (>6 μg/mL) (Fig. 5C and Supplementary Fig. S5; Table S2). Data corresponding to relative dry weight showed similar behaviour (Supplementary Fig. S4; Table S3). CDDP differs from MMC since it is a chemotherapy drug that induces higher relative levels of intra-strand CLs versus inter-strand CLs in the DNA45,46. These adducts, which affect only one strand of the DNA, are mainly removed by nucleotide excision repair (NER). However, HR is also critical to manage CDDP toxicity, probably because this agent can eventually produce inter-strand CLs47,48. As in the case of gamma rays, both axr1-31 and axl-2 were hypersensitive to CDDP, axr1-31 being even more affected by the treatment (Fig. 5D, Supplementary Fig. S4; Tables S2 and S3).
DNA damage sensitivity assays in axr1-31 and axl-2. (a) Representative images of WT, axr1-31 and axl-2 seedlings after exposure to 0, 200 and 300 Gy. (b) Number of leaves per plant in WT, axr1-31, and axl-2 seedlings after the exposure to different doses of gamma radiation. (c) Number of leaves per plant in WT, axr1-31, and axl-2 seedlings after the exposure to different concentrations of MMC. (d) Number of leaves per plant in WT, axr1-31, and axl-2 seedlings after the exposure to different concentrations of CDDP. (e) Root length per plant in WT, axr1-31, and axl-2 seedlings during different days after the exposure to 500 J/m2 of UV-C light irradiation. All the results were normalised to standard grown conditions. *p ≤ 0.05; +p ≤ 0.01; #p ≤ 0.001.
UV-C radiation mainly causes a photo-chemical effect in adjacent thymines that leads to their dimerization49. Photoreactivation is thought to be the major DNA repair pathway for these pyrimidine dimers in plants50. Enzymes involved in this pathway remove DNA lesions by absorbing light. Repair without a light source includes NER and other DNA repair pathways such as base excision repair (BER) or mismatch repair (MMR)50,51. By covering the plants with aluminium foil after irradiation we avoided photorreactivation, favouring the repair by NER and the other DNA repair pathways mentioned above. Unlike the previous treatments, no hypersensitivity response was detected in any of the mutants. Moreover, axr1-31 plants exhibited a slightly better response than the control (Fig. 5E; Supplementary Table S2).
The expression of genes involved in HR is disturbed in axr1
Given the differences between axr1 and axl in their behaviour during meiotic recombination and DNA repair, we considered evaluating the expression of several genes by qRT-PCR. Specifically, we analysed transcription levels of AXR1, AXL, Topoisomerase II (TOPII, involved in chromosome condensation52 and interlock resolution53), and sixteen key genes for HR (Supplementary Table S1). Since the mutations axr1 and axl affect auxin response, it is important to note that several of the analysed genes present an auxin-responsive element AuxRE (Auxin-Responsive Element) in their promoters: ATM, ATR, BRCA2A, BRCA2B, MLH3, MND1, MSH4, MSH5, RAD51C, RMI1, and XRI154,55 (Promomer, http://bar.utoronto.ca/ntools/cgi-bin/BAR_Promomer.cgi). We have conducted qRT-PCR parallel analyses on samples from flower buds and young seedlings to determine whether or not the variations in the gene expression levels are meiosis-specific.
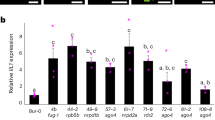
A noteworthy result was the overexpression of each corresponding paralogous gene in flower bud samples from both mutants: AXL expression was up-regulated in axr1 flower buds, and AXR1 was in axl (Fig. 6A,B), revealing a possible functional relationship between both genes. The observed overexpression seems to be meiosis-specific, since it was not detected in seedlings, in which even AXR1 was down-regulated in axl-2 samples (Fig. 6D). The reduced expression of HEI10 in flower bud samples from both mutants (and in seedlings from axl-2) was also relevant (Fig. 6). This gene encodes a conserved meiotic ubiquitin E3 ligase required for the formation of most COs in Arabidopsis37,56. TOPII was also down-regulated in flower bud samples from both mutants (although not statistically significantly in axl-2) (Fig. 6A,B). In the case of samples obtained from seedlings it is noteworthy that the expression levels of the genes analysed were normal in axl-2 (in which only HEI10 and AXR1 were under-expressed), whereas several genes were up-regulated in axr1-31: BRCA2B, DMC1, RAD51, FANCM and MSH4 (Fig. 6C,D).
Discussion
Duplicated genes have played a critical role in the evolution of all major eukaryotic linages, especially in plants. Their preponderance is consequence of the high rate of duplication occurrence along evolution together with preferential retention of paralogous genes (retention bias). After duplication, maintenance of a perfect equivalency in gene function is unusual and most models of duplicate retention assume various degrees of functional redundancy because of changes in function, expression, or interactions with other gene products57. Among Angiosperms, the existence of distinguishable AXR1 and AXL genes seems to be limited to Arabidopsis species. In other plant species there are either well preserved copies, virtually identical, or only one AXR1 gene35. Why are two functional copies necessary in the Arabidopsis genome? Hotton and colleagues35 suggested that the two corresponding proteins could interact with different factors. Alternatively, AXR1 and AXL could have different roles during certain cellular environments or developmental stages. Here we report new evidence that point in this direction.
Despite AXR1 and AXL are highly expressed in the ninth stage of flower development, axr1 and axl mutants showed differences in fertility that could be related to conspicuous differences in meiotic chromosome behaviour (Fig. 2). Meiocytes from plants deficient in AXR1 did not display complete synapsis during pachytene stage and presented a high frequency of univalents at metaphase I that led to abnormal chromosome segregations at anaphase I and unbalanced second division nuclei29 (Fig. 2I–P). On the other hand, as expected from their full fertility, meiosis was normal in axl plants, with the exception of the presence of chromosome bridges at anaphase I (Fig. 2Q–X). Although axl and WT plants exhibited a similar mean cell chiasma frequency, there were slight differences in the contribution of each chromosome to the total chiasma frequency (Fig. 3E; Table 1). Particularly, in comparison with the situation observed in WT cells, the decrease in the contribution of long chromosomes 1, 3, and 5 was compensated by an increase in the contribution of the acrocentric chromosomes (2 and 4). These last chromosomes are the smallest and bear the nucleolar organising regions (NORs) (Fig. 3A). Under altered CO distribution circumstances, chromosomes 2 and 4 could have more chances to recombine due to the physical proximity of the homologous pairs through their nucleolar domains58,59.
Jahns and colleagues29 reported that the conspicuous decrease in the frequency of bivalents showed by axr1 mutants was not due to a decrease in CO frequency but rather due to changes in CO distribution. The phenotype we observed in axl might be reflecting slight variations in the distribution of COs, although not as drastic as those observed in axr1. In this context, altered chiasma distribution in axr1-31 and axl-2 might be related to impaired expression of the meiotic E3 ligase HEI10, since this gene was down-regulated in both genetic backgrounds (Fig. 6A,B). Supporting this idea, HEI10 (as well as its orthologues RNF212 and ZIP337,60,61) is postulated to participate in designating HR intermediates as COs and has been described as a dosage-sensitive regulator of class I COs during Arabidopsis meiosis56. We also detected decreased expression levels of TOPII (Fig. 6A,B), which has been involved in chromosome condensation and meiotic interlock resolution in A. thaliana53. In other organisms, mutations affecting this gene are related to alterations in chromosome condensation, defects in chromosome segregation, and reduced strength of CO interference62,63,64. Abnormal chromosome condensation, altered CO distribution and chromosome segregation problems were observed in axr1 meiocytes and also in axl to a lesser extent (Fig. 2), suggesting that part of their phenotypes could be due to an impaired activity of TOPII. Nevertheless, neither the absence of AXR1 nor AXL appear to alter the expression of the other meiotic genes tested in our experiments, even to those with an auxin-related sequence motif (AuxRE) motif on their promoters.
We have obtained more results indicating that AXR1 and AXL are not functionally equal in meiosis, the former playing a more active role than the later. The presence of the cassette AXR1p::10MYC-AXR1 was capable to complement the strong axr1-30 mutant phenotype in bivalent formation, while AXR1p::10MYC-AXL was only able to do it partially: univalents were still present, but with a lower frequency (Fig. 4, Supplementary Fig. S3). This result is in line with that obtained in the analyses by qRT-PCR. AXR1 displayed a three-fold change compared to WT levels in flower bud samples from axl-2 and AXL was also overexpressed in flower bud samples from axr1-31 (Fig. 6). One possible explanation is that the increased expression of AXR1 in axl-2 is sufficient to ensure normal meiosis, while the presence of a greater amount of AXL does not allow it in axr1-31. These findings indicate that the functions of AXR1 and AXL on meiotic recombination are not redundant.
Most proteins that participate in meiotic recombination are also active during somatic DNA damage repair. However, their somatic functions might differ from their meiotic roles through interactions with different factors or post-translational modifications. Presumably, according to the results discussed above, AXR1 and AXL diverged in relation to their meiotic recombination functions. In this study we find for the first time a role for both rubylation E1 subunits AXR1 and AXL. Nevertheless, as far as their role in DNA repair, although with subtle differences, the function of both genes appears to be more redundant. Both axr1-31 and axl-2 plants showed hypersensitivity to gamma-rays and CDDP, although only axr1-31 plants were hypersensitive to MMC (Fig. 5; Supplementary Table S2). In the latter, AXR1 might act as a back-up in axl-2 mutant background, whereas its presence is not enough to avoid the hypersensitivity phenotype displayed after the treatment with the other DNA damage agents. These results suggest that although both AXR1 and AXL are involved in DNA repair they are not equally important in the repair of the different CLs produced by the treatments. One possible explanation is that AXL has a more prominent role in the repair through NER, since this is the main repair pathway for intra-strand CLs created by CDDP, whereas AXR1 would have a more direct involvement in HR repair, the process required to remove the inter-strand CLs formed by MMC. In agreement with this idea, genes involved in HR were up-regulated in axr1-31 seedlings in relation to WT, whereas they presented normal levels in axl-2 seedlings (Fig. 6D). In this context, it has been reported that expression of AXR1 after exposure to a mix of bleomycin and MMC is slightly higher than in normal conditions, whereas AXL levels do not change65. CDDP also induces inter-strand CLs (although to a lesser extent than intra-strand CLs) and this could be the reason why both mutants were hypersensitive to this agent.
The results corresponding to the UV radiation experiment did not indicate hypersensitivity in any of the mutants (Fig. 5E; Supplementary Table S2). Furthermore, axr1-31 showed some tolerance to the treatment. In previous studies, the rubylation pathway has been indirectly linked to UV-C sensitivity through the ubiquitin-mediated degradation of photomorphogenic factors, interconnecting light signalling and DNA repair66. It is tempting to speculate that the efficiency of other DNA repair pathways different from HR could be improved in this genetic background in which HR is deficient, although future experiments should be performed to test this hypothesis. Because of the lethality of the axr1 axl double mutant, it is not possible to conduct sensitivity assays in this background. Nevertheless, the results obtained indicate that the two proteins could have non-overlapping functions and cannot complement each other, presenting more defects when AXR1 is absent. The lethality of the double mutant might even reflect that both proteins are involved in parallel DNA repair pathways.
Indirect proofs about the relationship between rubylation and DNA repair have been obtained in mouse models and human cells. In these materials the combination of either MMC or CDDP and MLN4924, the inhibitor of NEDD8/RUB-activating enzyme, synergistically enhances the cytotoxicity of both drugs through increased DNA damage67,68. Moreover, the N-terminal neddylation of histone H4 in response to DNA damage in mammalian cells suggests that this modification could break inter-nucleosome interactions facilitating the access to the repair machinery26. Neddylation has also been shown to regulate DSB pathway choice and the balance between non-homologous end-joining (NHEJ) and HR in somatic DNA repair by inhibiting CtIP-mediated DNA end resection27 and promoting Ku ubiquitination and its release from damage sites69. In A. thaliana, the complex CUL4-DDB1-DDB2 (CULLIN4 together with DNA DAMAGE BINDING PROTEINS 1 and 2), a substrate of AXR1 rubylation pathway, ubiquitylates a high variety of proteins during DNA repair by NER in an ATR-dependent manner70. CUL4 levels also increase after genotoxic damage65. In relation to meiotic recombination, CUL4 is apparently involved in the deregulation of CO localisation observed in axr1 mutants29. However, cul1 and the hypomorphic cul3a/3b mutants, defective for cullins that are also rubylated, display normal meiotic behaviour29. Interestingly, CUL1 partner ASK1 (Arabidopsis SKP1-like1) is required to ensure proper chromosome condensation, synapsis and chromosome segregation in male Arabidopsis meiosis71. Our results showing DNA repair deficiencies in mutants for RUB-activating enzyme confirm more directly the connection between rubylation and DNA repair in plants. However, further studies with mutants defective for other proteins involved in rubylation will be necessary to reveal a more mechanistic view of the link between this post-translational modification and both somatic and meiotic HR.
Based on all the results presented here it seems that the asymmetry in the pattern of functional partitioning between these paralogs is different during meiosis and DNA repair. These genes are predicted to have duplicated after the more recent whole-genome duplication event (α)72. After duplication, random genetic drift might lead to an uneven contribution of the two copies, with AXR1 taking on a leading role, this being more pronounced in meiosis. This model seems to be intermediate among Angiosperms species, in which either there are two identical copies in sequence, and therefore with the same function, or one of the copies has been lost. The existence of distinguishable AXR1 and AXL proteins could influence their preferences for different interactors depending on the developmental stage or the cellular environment. This difference could in turn lead to some specialisation (such as the repair of one of the types of CLs or involvement in a particular DNA repair pathway). This situation has been also described for other Arabidopsis genes such as OSD1 (OMISSION OF SECOND DIVISION 1), a gene essential to enter into the second meiotic division, and its paralog UVI4 (UV-B-INSENSITIVE 4), without a meiotic function but required for the cell cycle73,74. The study of these genes in species close to Arabidopsis, not yet characterized, might shed some light on the possible consequences of these divergences.
Methods
Plant material
Several transfer DNA (T-DNA) mutant lines defective for the genes AXR1 (At1g05180) and AXL (At2g32410) were analysed (Fig. 1A). The lines axr1-31 (SALK_013238) and axl-2 (GK_818B10) were identified by “Signal T-DNA Express Arabidopsis Gene Mapping Tool” (http://signal.salk.edu/cgi-bin/tdnaexpress) and obtained from Nottingham Arabidopsis Stock Centre (NASC). Numbers were assigned to each of these new alleles in concordance to previous references33,75,76. Line axl-1 was provided in homozygous condition by Prof. Mark Estelle (University of California San Diego, USA). Lines axr1-30 (SAIL_904_E06), AXR1p:10MYC-AXR1 and AXR1p::10MYC-AXL (all homozygous for axr1-30) were kindly donated by Prof. Judy Callis (University of California Davis, USA). Arabidopsis Col-0 accession was used as a control.
Growth conditions and plant genotyping
Seeds were sown in pots containing a soil mixture of vermiculite and commercial soil (3:1). Plants were maintained in a growth chamber (temperature 18–20 °C; photoperiod 16 h light/8 h dark).
Genomic DNA was extracted mashing young leaves in extraction buffer (100 mM Tris pH 9.5; 250 mM KCl: 10 mM EDTA). After 10 minutes at 95 °C, samples were diluted in BSA 3%. PCR amplifications of mutant alleles were performed using a specific T-DNA primer and a T-DNA flanking primer (RP). Specific T-DNA primers were: LBa1 (5′TGGTTCACGTAGTGGGCCATCG3′) for SALK lines; and o8409 (5′ATATTGACCATCATACTCATT3′) for the GK line. T-DNA flanking primers used were: 5′TCATGTGGAGAATGGGCTTAC3′ (RP-axr1-31) and 5′AAGTATCACCATTGTCGACGG3′ (RP-axl-2). WT alleles were amplified with primers LP-axr1-31 (5′TGTGATTGAATATTGCAGGAGC3′) and RP-axr1-31; LP-axl-2 (5′ TCTTCATCCGCTGATACCATC3′) and RP-axl-2.
Identification of abnormal pollen grains and fertility assessment
Pollen grains were screened using the pollen squash method. Mature flowers were dissected, and the anthers were stained with 2% aceto-carmine (Sigma) and fixed to the slide through a flame. Slides were observed by phase-contrast microscopy. To evaluate fertility, data corresponding to the number of seeds per silique and silique length were collected from five randomly selected siliques (once they completely developed but before they had dropped the seeds) belonging to three plants of each genotype.
Cytological techniques
Flower buds were fixed in Carnoy (60% ethanol, 30% chloroform, 10% acetic acid) at room temperature during at least 24 h and kept at −20 °C until used. Chromosome spreads and fluorescence in situ hybridization (FISH) were performed according to the methodology previously described (the most recent version of the protocol is reported in77). More than 50 cells per genotype were analyzed at each stage. The following DNA probes were used: pTa71 [45 S ribosomal DNA (rDNA), pTa71 of Triticum aestivum78] and pCT4.2 (5 S rDNA79).
Immunolocalisation procedures were conducted as detailed in Armstrong and colleagues80, with slight modifications81. The primary antibodies used were kindly provided by Prof. Chris Franklin (University of Birmingham): anti-AtASY1 (rat; 1:1,000 dilution), anti-AtZYP1 (rabbit; 1:500).
Slide preparations were observed with an Olympus BX61 microscope equipped with epifluorescence optics. Images were captured with an Olympus DP70 digital camera controlled by DP Controller software version 2.2.1.227 (Olympus).
DNA damage sensitivity analyses
Seeds were surface sterilized for 5 min in 2.5% sodium hypochlorite and rinsed with sterilized distilled water at least five times. After leaving them overnight at 4 °C, the seeds were plated in plastic Petri plates containing germination media (GM82: salt mixture (1×) in 1% sucrose, and 1% agarose) to conduct the genotoxic assays following the protocol reported in Martinez-Garcia and Pradillo (2017)77. To test the sensitivity to γ-rays, sterilized seeds were kept overnight in sterile water at 4 °C and then exposed to 0, 100, 200, 300, 400, and 500 Gy doses (2.94 Gy/min) from a 137Cs source (IBL 437 C; CIS Bio). After irradiation, seeds were germinated on GM agar medium. Different concentrations for mitomycin C (MMC, Duchefa Biochemie: 2, 4, 6, 8 and 10 µg/mL) and cisplatin [cis-diamminedichloroplatinum (II), CDDP, Sigma: 0, 15, 30, 50, and 75 μM] were also applied. Solutions were prepared and then added directly into MS media after cooling down. Plates were maintained under standard growth conditions (temperature 22–25 °C; photoperiod 16 h light/8 h dark) until analysis. The effects of the DNA damage agents on plant growth (number of true leaves) were evaluated 12 days (MMC) or 14 days (γ-rays, CDDP) after sowing. For the UV-C damage assay, 5-day-old seedlings grown in vertical square plates were irradiated for 5 minutes with a dose of 500 J/m2 from a Sanyo Denki g15T8 lamp. Irradiated plants were grown in dark conditions to avoid photorreactivation, favouring DNA repair by nucleotide excision repair (NER). Root lengths were measured 1, 2, 3, and 6 days after the treatment. Statistical comparison was performed using non-parametric two-tailed Mann Whitney U-test. P-values are available in Supplementary Table S2.
Gene expression analyses
For reverse transcription PCR (RT-PCR) and quantitative RT-PCR (qRT-PCR) experiments, total RNA was extracted from 10-day-old seedlings and flower buds using RNeasy PlantMini kit (Qiagen) and DNase enzymatic treatment (RNase-Free DNase Set, Qiagen). mRNAs from AXR1 and AXL were retrotranscribed with One Step RT-PCR Kit (Qiagen). GAPC (glyceraldehyde-3-phosphate dehydrogenase, cytosolic gene) was used as constitutive control. Primers used in the RT-PCR were: F-AXR1: 5′ATGCAAGCAGTAAAAAGATCCAG3′ and R-AXR1: 5′CTACAATTTCAATAACTGAGAC3′; F-AXL: 5′TGGTAGCAGCACTCAAGGAG3′ and R-AXL: 5′CCGTCACAACCCATTTCACT3′; N-GAPC: 5′CTTGAAGGGTGGTGCCAAGAAGG3′ and C-GAPC: 5′CCTGTTGTCGCCAACGAAGTCAG3′.
qRT-PCR was performed with the Transcriptor First Strand cDNA Synthesis kit and the FastStart TaqMan Probe Master kit (Roche). Primers were designed using the tool UPL Assay Design Centre (Roche) and their sequences are listed in Supplementary Table S1. Data were analyzed considering fold variation over a calibrator using the ΔΔCt method and the ACTIN2 gene as endogenous control. At least three experimental replicates were performed. We considered a fold-change variation significant when there was a minimum of two-fold change compared to the WT levels: RQ higher than 2 or lower than 0.5 and non-overlapping 95% confidence intervals.
Database resources and bioinformatic tools
The identification of T-DNA insertion lines, analysis of sequences and primer design were performed using databases and tools from National Centre for Biotechnology Information (http://www.ncbi.nlm.nih.gov/), The Arabidopsis Information Resource (http://www.arabidopsis.org/), Nottingham Arabidopsis Stock Centre (http://arabidopsis.info/) and The Salk Institute Genome Analysis Laboratory (http://signal.salk.edu/cgi-bin/tdnaexpress/). Gene interactions were analysed using Promomer (http://bar.utoronto.ca/ntools/cgi-bin/BAR_Promomer.cgi). Statistical analyses were conducted with Microsoft Excel and the software IBM SPSS 22. Data in the text with their mean as measures of center, is followed by ± their standard error as a measure of variability. Statistical comparisons between values were made using non-parametric two-tailed Mann Whitney U-test, whether comparisons between frequencies were made using χ21 test. Adobe Photoshop CS4, Microsoft Excel, SigmaPlot, and ImageJ were used for graph and image composition.
Data availability
All data generated or analysed during this study are included in this published article (and its Supplementary Information files). Biological materials will be shared upon request to the corresponding author.
References
Ozkaynak, E., Finley, D. & Varshavsky, A. The yeast ubiquitin gene: head-to-tail repeats encoding a polyubiquitin precursor protein. Nature 312, 663–666 (1984).
Zuin, A., Isasa, M. & Crosas, B. Ubiquitin signaling: extreme conservation as a source of diversity. Cells 3, 690–701 (2014).
Miricescu, A., Goslin, K. & Graciet, E. Ubiquitylation in plants: signaling hub for the integration of environmental signals. J. Exp. Bot. 69, 4511–4527 (2018).
Hershko, A. & Ciechanover, A. The ubiquitin system. Annu. Rev. Biochem. 67, 425–479 (1998).
Voges, D., Zwickl, P. & Baumeister, W. The 26S proteasome: a molecular machine designed for controlled proteolysis. Annu. Rev. Biochem. 68, 1015–1068 (1999).
Callis, J., Carpenter, T., Sun, C. W. & Vierstra, R. D. Structure and evolution of genes encoding polyubiquitin and ubiquitin-like proteins in Arabidopsis thaliana ecotype Columbia. Genetics 139, 921–939 (1995).
Vierstra, R. D. Proteolysis in plants: mechanisms and functions. Plant Mol. Biol. 32, 275–302 (1996).
Hatfield, P. M., Gosink, M. M., Carpenter, T. B. & Vierstra, R. D. The ubiquitin-activating enzyme (E1) gene family in Arabidopsis thaliana. Plant. J. Cell Mol. Biol. 11, 213–226 (1997).
Smalle, J. & Vierstra, R. D. The ubiquitin 26S proteasome proteolytic pathway. Annu. Rev. Plant Biol. 55, 555–590 (2004).
Vierstra, R. D. The ubiquitin-26S proteasome system at the nexus of plant biology. Nat. Rev. Mol. Cell Biol. 10, 385–397 (2009).
Sadanandom, A., Bailey, M., Ewan, R., Lee, J. & Nelis, S. The ubiquitin-proteasome system: central modifier of plant signalling. New Phytol. 196, 13–28 (2012).
Rao-Naik, C. et al. The rub family of ubiquitin-like proteins. Crystal structure of Arabidopsis rub1 and expression of multiple rubs in Arabidopsis. J. Biol. Chem. 273, 34976–34982 (1998).
Callis, J. The Ubiquitination Machinery of the Ubiquitin System. Arab. Book 12, e0174 (2014).
Hotton, S. K. & Callis, J. Regulation of Cullin RING Ligases. Annu. Rev. Plant Biol. 59, 467–489 (2008).
Duda, D. M. et al. Structural regulation of cullin-RING ubiquitin ligase complexes. Curr. Opin. Struct. Biol. 21, 257–264 (2011).
Mergner, J. & Schwechheimer, C. The NEDD8 modification pathway in plants. Front. Plant Sci. 5 (2014).
Duda, D. M. et al. Structural Insights into NEDD8 Activation of Cullin-RING Ligases: Conformational Control of Conjugation. Cell 134, 995–1006 (2008).
Schwechheimer, C. NEDD8 — its role in the regulation of Cullin-RING ligases. Curr. Opin. Plant Biol. 45, 112–119 (2018).
Dharmasiri, N., Dharmasiri, S., Jones, A. M. & Estelle, M. Auxin Action in a Cell-Free System. Curr. Biol. 13, 1418–1422 (2003).
Dreher, K. & Callis, J. Ubiquitin, Hormones and Biotic Stress in Plants. Ann. Bot. 99, 787–822 (2007).
Mockaitis, K. & Estelle, M. Auxin Receptors and Plant Development: A New Signaling Paradigm. Annu. Rev. Cell Dev. Biol. 24, 55–80 (2008).
Hua, Z. & Vierstra, R. D. The Cullin-RING Ubiquitin-Protein Ligases. Annu. Rev. Plant Biol. 62, 299–334 (2011).
Vain, T. et al. Selective auxin agonists induce specific AUX/IAA protein degradation to modulate plant development. Proc. Natl. Acad. Sci. USA 116, 6463–6472 (2019).
Lincoln, C., Britton, J. H. & Estelle, M. Growth and development of the axr1 mutants of Arabidopsis. Plant Cell 2, 1071–1080 (1990).
del Pozo, J. C. et al. AXR1-ECR1–Dependent Conjugation of RUB1 to the Arabidopsis Cullin AtCUL1 Is Required for Auxin Response. Plant Cell 14, 421–433 (2002).
Ma, T. et al. RNF111-Dependent Neddylation Activates DNA Damage-Induced Ubiquitination. Mol. Cell 49, 897–907 (2013).
Jimeno, S. et al. Neddylation inhibits CtIP-mediated resection and regulates DNA double strand break repair pathway choice. Nucleic Acids Res. 43, 987–999 (2015).
Meir, M. et al. The COP9 signalosome is vital for timely repair of DNA double-strand breaks. Nucleic Acids Res. 43, 4517–4530 (2015).
Jahns, M. T. et al. Crossover Localisation Is Regulated by the Neddylation Posttranslational Regulatory Pathway. PLoS Biol. 12, e1001930 (2014).
Brockway, H., Balukoff, N., Dean, M., Alleva, B. & Smolikove, S. The CSN/COP9 Signalosome Regulates Synaptonemal Complex Assembly during Meiotic Prophase I of Caenorhabditis elegans. PLoS Genet. 10, e1004757 (2014).
Alleva, B., Clausen, S., Koury, E., Hefel, A. & Smolikove, S. CRL4 regulates recombination and synaptonemal complex aggregation in the Caenorhabditis elegans germline. PLoS Genet. 15, e1008486 (2019).
del Pozo, J. C. The Ubiquitin-Related Protein RUB1 and Auxin Response in Arabidopsis. Science 280, 1760–1763 (1998).
Dharmasiri, N. et al. AXL and AXR1 have redundant functions in RUB conjugation and growth and development in Arabidopsis: AXL1 and AXR1 function redundantly. Plant J. 52, 114–123 (2007).
Singh, S. K., Roy, S., Choudhury, S. & Sengupta, D. N. DNA repair and recombination in higher plants: insights from comparative genomics of arabidopsis and rice. BMC Genomics 11, 443 (2010).
Hotton, S. K., Eigenheer, R. A., Castro, M. F., Bostick, M. & Callis, J. AXR1-ECR1 and AXL1-ECR1 heterodimeric RUB-activating enzymes diverge in function in Arabidopsis thaliana. Plant Mol. Biol. 75, 515–526 (2011).
Estelle, M. A. & Somerville, C. Auxin-resistant mutants of Arabidopsis thaliana with an altered morphology. 7.
Chelysheva, L. et al. The Arabidopsis HEI10 Is a New ZMM Protein Related to Zip3. PLoS Genet. 8, e1002799 (2012).
Schmid, M. et al. A gene expression map of Arabidopsis thaliana development. Nat. Genet. 37, 501–506 (2005).
Sanchez-Moran, E., Armstrong, S. J., Santos, J. L., Franklin, F. C. H. & Jones, G. H. Variation in Chiasma Frequency Among Eight Accessions of Arabidopsis thaliana. 8.
de Massy, B. Distribution of meiotic recombination sites. Trends Genet. 19, 514–522 (2003).
Benítez-Bribiesca, L. & Sánchez-Suárez, P. Oxidative Damage, Bleomycin, and Gamma Radiation Induce Different Types of DNA Strand Breaks in Normal Lymphocytes and Thymocytes: A Comet Assay Study. Ann. N. Y. Acad. Sci. 887, 133–149 (1999).
Tomasz, M. et al. Isolation and structure of a covalent cross-link adduct between mitomycin C and DNA. Science 235, 1204–1208 (1987).
Sasaki, M. S., Takata, M., Sonoda, E., Tachibana, A. & Takeda, S. Recombination repair pathway in the maintenance of chromosomal integrity against DNA interstrand crosslinks. Cytogenet. Genome Res. 104, 28–34 (2004).
Deans, A. J. & West, S. C. DNA interstrand crosslink repair and cancer. Nat. Rev. Cancer 11, 467–480 (2011).
Eastman, A. Interstrand cross-links and sequence specificity in the reaction of cis-dichloro(ethylenediamine)platinum(II) with DNA. Biochemistry 24, 5027–5032 (1985).
Jamieson, E. R. & Lippard, S. J. Structure, Recognition, and Processing of Cisplatin−DNA Adducts. Chem. Rev. 99, 2467–2498 (1999).
De Silva, I. U. Defects in interstrand cross-link uncoupling do not account for the extreme sensitivity of ERCC1 and XPF cells to cisplatin. Nucleic Acids Res. 30, 3848–3856 (2002).
Wang, Q.-E. et al. Differential contributory roles of nucleotide excision and homologous recombination repair for enhancing cisplatin sensitivity in human ovarian cancer cells. Mol. Cancer 10, 24 (2011).
Rastogi, R. P., Richa, Kumar, A., Tyagi, M. B. & Sinha, R. P. Molecular mechanisms of ultraviolet radiation-induced DNA damage and repair. J. Nucleic Acids 2010, 592980 (2010).
Kimura, S. DNA repair in higher plants; photoreactivation is the major DNA repair pathway in non-proliferating cells while excision repair (nucleotide excision repair and base excision repair) is active in proliferating cells. Nucleic Acids Res. 32, 2760–2767 (2004).
Manova, V. & Gruszka, D. DNA damage and repair in plants – from models to crops. Front. Plant Sci. 6 (2015).
Uemura, T. et al. DNA topoisomerase II is required for condensation and separation of mitotic chromosomes in S. pombe. Cell 50, 917–925 (1987).
Martinez-Garcia, M. et al. TOPII and chromosome movement help remove interlocks between entangled chromosomes during meiosis. J. Cell Biol. 217, 4070–4079 (2018).
del Pozo, J. C. & Manzano, C. Auxin and the ubiquitin pathway. Two players–one target: the cell cycle in action. J. Exp. Bot. 65, 2617–2632 (2014).
Franco-Zorrilla, J. M. et al. DNA-binding specificities of plant transcription factors and their potential to define target genes. Proc. Natl. Acad. Sci. 111, 2367–2372 (2014).
Ziolkowski, P. A. et al. Natural variation and dosage of the HEI10 meiotic E3 ligase control Arabidopsis crossover recombination. Genes Dev. 31, 306–317 (2017).
Panchy, N., Lehti-Shiu, M. & Shiu, S.-H. Evolution of Gene Duplication in Plants. Plant Physiol. 171, 2294–2316 (2016).
Pecinka, A. et al. Chromosome territory arrangement and homologous pairing in nuclei of Arabidopsis thaliana are predominantly random except for NOR-bearing chromosomes. Chromosoma 113, 258–269 (2004).
Berr, A. et al. Chromosome arrangement and nuclear architecture but not centromeric sequences are conserved between Arabidopsis thaliana and Arabidopsis lyrata. Plant. J. Cell Mol. Biol. 48, 771–783 (2006).
Reynolds, A. et al. RNF212 is a dosage-sensitive regulator of crossing-over during mammalian meiosis. Nat. Genet. 45, 269–278 (2013).
Agarwal, S. & Roeder, G. S. Zip3 provides a link between recombination enzymes and synaptonemal complex proteins. Cell 102, 245–255 (2000).
Gómez, R. et al. Cohesin removal precedes topoisomerase IIα-dependent decatenation at centromeres in male mammalian meiosis II. Chromosoma 123, 129–146 (2014).
Zhang, L. et al. Topoisomerase II mediates meiotic crossover interference. Nature 511, 551–556 (2014).
Shintomi, K., Takahashi, T. S. & Hirano, T. Reconstitution of mitotic chromatids with a minimum set of purified factors. Nat. Cell Biol. 17, 1014–1023 (2015).
Kilian, J. et al. The AtGenExpress global stress expression data set: protocols, evaluation and model data analysis of UV-B light, drought and cold stress responses: AtGenExpress global abiotic stress data set. Plant J. 50, 347–363 (2007).
Castells, E. et al. det1-1-induced UV-C hyposensitivity through UVR3 and PHR1 photolyase gene over-expression: det1-induced UV-C hyposensitivity. Plant J. 63, 392–404 (2010).
Garcia, K. et al. Nedd8-Activating Enzyme Inhibitor MLN4924 Provides Synergy with Mitomycin C through Interactions with ATR, BRCA1/BRCA2, and Chromatin Dynamics Pathways. Mol. Cancer Ther. 13, 1625–1635 (2014).
Ho, I.-L. et al. MLN4924 Synergistically Enhances Cisplatin-induced Cytotoxicity via JNK and Bcl-xL Pathways in Human Urothelial Carcinoma. Sci. Rep. 5, 16948 (2015).
Brown, J. S. et al. Neddylation Promotes Ubiquitylation and Release of Ku from DNA-Damage Sites. Cell Rep. 11, 704–714 (2015).
Molinier, J., Lechner, E., Dumbliauskas, E. & Genschik, P. Regulation and Role of Arabidopsis CUL4-DDB1A-DDB2 in Maintaining Genome Integrity upon UV Stress. PLoS Genet. 4, e1000093 (2008).
Zhao, D. et al. ASK1, a SKP1 homolog, is required for nuclear reorganization, presynaptic homolog juxtaposition and the proper distribution of cohesin during meiosis in Arabidopsis. Plant Mol. Biol. 62, 99–110 (2006).
Bowers, J. E., Chapman, B. A., Rong, J. & Paterson, A. H. Unravelling angiosperm genome evolution by phylogenetic analysis of chromosomal duplication events. Nature 422, 433–438 (2003).
Hase, Y., Trung, K. H., Matsunaga, T. & Tanaka, A. A mutation in the uvi4 gene promotes progression of endo-reduplication and confers increased tolerance towards ultraviolet B light. Plant J. 46, 317–326 (2006).
Cromer, L. et al. OSD1 Promotes Meiotic Progression via APC/C Inhibition and Forms a Regulatory Network with TDM and CYCA1;2/TAM. PLoS Genet. 8, e1002865 (2012).
Larsen, P. B. & Cancel, J. D. A recessive mutation in the RUB1-conjugating enzyme, RCE1, reveals a requirement for RUB modification for control of ethylene biosynthesis and proper induction of basic chitinase and PDF1.2 in Arabidopsis. Plant. J. Cell Mol. Biol. 38, 626–638 (2004).
Gilkerson, J. et al. Isolation and Characterization of cul1-7, a Recessive Allele of CULLIN1 That Disrupts SCF Function at the C Terminus of CUL1 in Arabidopsis thaliana. Genetics 181, 945–963 (2009).
Martinez-Garcia, M. & Pradillo, M. Functional Analysis of Arabidopsis ARGONAUTEs in Meiosis and DNA Repair. in Plant Argonaute Proteins (ed. Carbonell, A.) vol. 1640 145–158 (Springer New York, 2017).
Gerlach, W. L. & Bedbrook, J. R. Cloning and characterization of ribosomal RNA genes from wheat and barley. Nucleic Acids Res. 7, 1869–1885 (1979).
Campell, B. R., Song, Y., Posch, T. E., Cullis, C. A. & Town, C. D. Sequence and organization of 5S ribosomal RNA-encoding genes of Arabidopsis thaliana. Gene 112, 225–228 (1992).
Armstrong, S. J., Caryl, A. P., Jones, G. H. & Franklin, F. C. Asy1, a protein required for meiotic chromosome synapsis, localizes to axis-associated chromatin in Arabidopsis and Brassica. J. Cell Sci. 115, 3645–55 (2002).
Varas, J. & Pradillo, M. Immunolabeling Protocols for Studying Meiosis in Plant Mutants Defective for Nuclear Envelope Components. Methods Mol. Biol. Clifton NJ 1840, 237–247 (2018).
Murashige, T. & Skoog, F. A Revised Medium for Rapid Growth and Bio Assays with Tobacco Tissue Cultures. Physiol. Plant. 15, 473–497 (1962).
Acknowledgements
We are very grateful to Prof. Mark Estelle, Dr. Michael Prigge, Prof. Judy Callis, Prof. Claus Schwechheimer and Dr. Julia Mergner for providing seeds. We also thank Bianca Martín for technical assistance and M. Cruz Valladares for helping us with the gamma-rays treatment. Authors acknowledge the support of the Ministry of Economy and Competitiveness of Spain (by grants AGL2012-38852 and AGL2015-67349-P) and of the European Union (Marie Curie ITN, COMREC 606956). MP is part of the International Plant Nucleus Consortium (IPNC, https://radar.brookes.ac.uk) and COST Action n° CA 16212 ‘In Depth’ http://www.cost.eu/COST_Actions/ca/CA16212. NFJ is a PhD fellow funded by the FPU programme of Spanish Ministry of Education (FPU16/02772).
Author information
Authors and Affiliations
Contributions
M.M.G., N.F.J. and M.P. completed the experiments and performed data analyses. M.M.G., J.L.S. and M.P. conceived and designed the experiments, wrote and reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Martinez-Garcia, M., Fernández-Jiménez, N., Santos, J.L. et al. Duplication and divergence: New insights into AXR1 and AXL functions in DNA repair and meiosis. Sci Rep 10, 8860 (2020). https://doi.org/10.1038/s41598-020-65734-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-65734-2
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.