Abstract
Living organisms require complex signaling interactions and proper regulation of these interactions to influence biological processes. Of these complex networks, one of the most distinguished is the Notch pathway. Dysregulation of this pathway often results in defects during organismal development and can be a causative mechanism for initiation and progression of cancer. Despite previous research entailing the importance of this signaling pathway and the organismal processes that it is involved in, less is known concerning the major Notch downstream targets, especially the onset and sequence in which they are modulated during normal development. As timing of regulation may be linked to many biological processes, we investigated and established a model of temporal patterning of major Notch downstream targets including broad, cut, and hindsight during Drosophila melanogaster egg chamber development. We confirmed the sequential order of Broad upregulation, Hindsight upregulation, and Cut downregulation. In addition, we showed that Notch signaling could be activated at stage 4, one stage earlier than the stage 5, a previously long-held belief. However, our further mitotic marker analysis re-stated that mitotic cycle continues until stage 5. Through our study, we once again validated the effectiveness and reliability of our MATLAB toolbox designed to systematically identify egg chamber stages based on area size, ratio, and additional morphological characteristics.
Similar content being viewed by others
Introduction
Biological organisms require complex signaling interactions and coordination of these interactions to survive and reproduce. Through cellular signaling pathways, cells interact and exchange information with neighboring cells, ultimately determining cell fate and organismal development as a whole1. Of these often complex signaling networks, the Notch signaling pathway is one of the most evolutionarily conserved2. This pathway facilitates local cell-cell interactions to control cell fate and plays a role in many aspects of development, including cell self-differentiation and self-renewal3,4.
As demonstrated by much previous research, the Notch pathway is necessary for proper cellular and organismal development. However, defects or absence of proper functions in this pathway often lead to problems with development, and often certain types of cancers5. Interestingly, the Notch pathway has been linked to not only tissue growth and tumorigenesis, but also cell death and tumor suppression in some circumstances6. Although this highlights the complexity of the function of the Notch pathway, recent research has indicated that the core molecular design is actually rather simple. In canonical Notch signaling, a transmembrane ligand Delta on an adjacent cell interacts with a receptor Notch on the cell receiving the signal, which then results in the receptor being cleaved to release the Notch intracellular domain (NICD). NICD moves to the nucleus of the cell, where it later interacts with DNA-binding proteins to begin transcription of the appropriate genes7.
As with many other aspects of development, recent research has shown that the timing in which this Notch signaling induces expression of its target genes is crucial for many aspects of organismal development. For instance, a study identified that interactions between epithelial cells in mice require proper sequential interaction of signaling pathways, among these most notably, the Notch pathway. This research also showed the transcription and expression patterns of various Notch components, suggesting that timing may play a role in the interactions of downstream targets8. In Drosophila, researchers also reported the importance of temporal regulation of neuroblasts for the cell fate and survival of Notch-mediated neurons9,10. During Drosophila oogenesis, the follicle cells of egg chambers sequentially undergo three different cell cycle programs: the mitotic cycle (stages 1-5), endocycle (stages 6-10a), and gene amplification (stages 10b-14), which are considered as early oogenesis, midoogenesis and late oogenesis, respectively11. The Notch pathway is the main signaling pathway to regulate the switches of cell cycles. There are many important downstream targets of Notch signaling, including cut, hindsight (hnt) and broad (br), during egg chamber development12,13,14. Cut is expressed in the mitotic cycle, then inhibited by Notch signaling in the endocycle. Opposite to the expression pattern of Cut, Hnt and Br are absent in the mitotic cycle, and upregulated by Notch signaling in the endocycle. In addition, Hnt and Br can promote each other, but repress Cut12,13,14. Scientists frequently use Cut expression for the mitotic cycle identification, and Hnt for the endocycle identification. However, an intermediate stage between the mitotic cycle and endocycle has been identified; when Hnt and Cut overlap expression15. We also observed similar overlapping expression patterns, and suggested that anterior follicle cells might undergo the mitotic cycle/endocycle (M/E) switch earlier16. In addition, as a direct downstream target of Notch signaling in follicle cells, the protein level of Br is highly upregulated at stage 6, but we reported that weak Br could be detected at stage 5 egg chambers based on morphology14. In summary, previous studies have shown that certain downstream targets of the Notch signaling pathway respond differently and at different times to stimuli. The findings highlight the sensitivity of the downstream targets br, cut, and hnt to Notch signaling during egg chamber development, indicating that the timing in which gene expression appeared in egg chambers sometimes varied.
Here, we present a model of the temporal patterning of Notch downstream targets br, hnt and cut during Drosophila melanogaster egg chamber development. Investigation and establishment of this signaling model during normal development may serve as a baseline for comparison of additional research findings, as modulation of such downstream targets is often accompanied by cell-cycle switches. In addition, we further confirmed the validity of a MATLAB toolbox for identification of egg chamber stages based on area size, ratio, and additional morphological characteristics that improves accuracy of stage identification, reduces reliance on visual determination, and can be useful for many areas of research.
Results
Gene expression pattern during transitional stages
Our previous findings with Drosophila follicle cells showed the expression of Hnt and Cut overlapped and weak Br could be detected at stage 5 egg chambers, suggesting varied sensitivity of br, cut, and hnt to Notch signaling11,14,16. As indicated in Fig. 1, we frequently observed strong gene expression based on immunohistochemical staining using antibodies. There is an inherent emphasis on the strong levels of gene expression when interrogating gene function or developmental patterning. In fact, from no expression to strong expression, or vice versa, there are transitional stages of gene expressions, during which the genes are weakly or partially expressed. Transitional stages are frequently unnoticed, but can be informative to indicate signaling onset. Focusing on more nuanced transitional patterns of expression might also provide additional mechanistic insight into genetic regulation.
It is known that br and hnt are activated by Notch signaling during the mitotic cycle, while cut is suppressed by Notch in the later endocycle. This marks the mitotic to endocycle transition (also called the M/E switch). However, the exact stages of their upregulation/downregulation are still debatable, partially because of different staging methods and inconsistent individual criteria. Previously, we created a toolbox to unbiasedly identify the stages based on morphological characteristics marked by DAPI staining. We concluded that follicle cells underwent the mitotic cycle at stage 5, and endocycle is from stage 6 without mitosis11,17.
We were particularly interested in the transitional expression patterns, because they indicated the sensitive response of gene activation/inactivation and signaling onset. To characterize these patterns, we specifically selected egg chambers during the transitional stages, which showed weak/partial upregulation of Br and Hnt, and downregulation of Cut (Fig. 2).
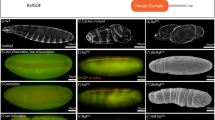
Confocal images of Drosophila egg chambers with stages identified. No expression, weak/partial expression, and full expression are shown for each target of interest. Downstream targets Br and Hnt are shown in earlier and later stage egg chambers corresponding with no expression, transitional, and later stage full expression. Cut is shown with early stage full expression, transitional, and later stage no expression. Scale bar =10 µm.
Early gene upregulation/downregulation during transitional stages
Previously, it was known that Notch signaling starts from stage 518,19, which then activates Br at stage 5/614, and Hnt at stages 6/713,20. Downregulation of Cut happens at stages 6/712,21. However, surprisingly, we found that early activation of both Br and Hnt could be observed as early as stage 4 (Figs. 3A,B), which was clearly indicated by the 5-blob phenotype of nurse cells in the egg chambers. During the Drosophila egg chamber development, the chromosomes of nurse cells appear polytene in stages 2–4 but later on dissociate22. For a stage 5 egg chamber, nurse cells no longer contain polytene chromosomes11,17. Prior to the breakdown of polytene chromosomes, the nurse cell chromosomes present a striking 5-blob phenotype, which can be used as a hallmark for stage 4 egg chambers11,17. In addition, Cut was not downregulated at stage 4, but its early downregulation could be found at stage 5 (Fig. 3C).
Sequential gene expression pattern during transitional stages
Immunostaining is mainly used to examine the expression patterns of target proteins. Primary antibodies are raised in various species, and those raised in the same host species can’t be stained simultaneously because the secondary antibodies can’t distinguish them. Host species of primary antibodies mouse anti-Cut (2B10)23, mouse anti-Br-Core (25E9)24, and mouse anti-Hnt (1G9)25 from Development Studies Hybridoma Bank were the same, therefore co-staining of them cannot be achieved to show the sequential gene expression patterns in the same egg chambers. Our previously developed toolbox can automatically identify egg chamber stages based on egg chamber size and ratio11. We systematically analyzed all the results of the transitional expression patterns based on confocal images using the toolbox. We found that transitional Br was mostly expressed in stage 4 egg chambers (64.8%, n = 91), while transitional Hnt was only found in 40.6% (n = 96) stage 4 egg chambers (Fig. 4A). Our findings suggested that Br was upregulated earlier than Hnt. In addition, the transitional expression patterns of Cut primarily occurred in stage 5 (96.4%, n = 84), indicating Cut was downregulated after the upregulation of Br and Hnt (Fig. 4A).
The DAPI images were collected from the largest cross-sectional area of the egg chambers in the middle plane, and this area size was used for our stage identification11. We found that egg chambers with transitional Hnt-expressing were significantly larger than those of transitional Br-expressing egg chambers, while the area sizes of both Hnt- and Br-expressing egg chambers were significantly smaller than those of transitional Cut-expressing egg chambers (Fig. 4B), further confirming the Br-Hnt-Cut sequential order of expression. We also examined the ratio of the major and minor axes of egg chambers, and found that transitional Cut-expressing egg chambers were more significantly elongated than Hnt- and Br-expressing egg chambers (Fig. 4C), indicating transitional Cut-expressing egg chambers were at later stages. Hnt- and Br-expressing egg chambers were similar in ratio (Fig. 4C), which is consistent with our previous findings that ratio differences were not evident until stage 511.
Cell cycle pattern during transitional stages
We identified and focused on the specific transitional stages with weak or partial expression of Notch target genes. We would ask whether these transitional egg chambers have already entered endocycle, ceasing mitotic cycle. Phospho-histone H3 (PH3) has frequently been utilized as a mitotic marker14,18, which shows oscillating patterns in follicle cells during the mitotic cycle, but disappears once they enter the endocycle. We co-stained PH3 with Br, Hnt, and Cut, respectively. We systematically analyzed the co-staining results at the transitional stages based on confocal images using the toolbox. Consistent with prior findings (Fig. 4), We found that transitional Br was mostly expressed in stage 4 egg chambers (59%, n = 39), while transitional Hnt was only found in 46.7% (n = 30) stage 4 egg chambers (Fig. 5). The transitional downregulated expression of Cut primarily occurred in stage 5 (91.9%, n = 37), further confirming Cut was downregulated after the upregulation of Br and Hnt (Fig. 5). In addition, we found 100% of both Hnt- and Br-expressing egg chambers in stages 4 and 5 still showed mitotic marker PH3 (Fig. 5), suggesting early weak upregulation of Br or Hnt is not robust enough to cease the mitosis to enter endocycle. Further observation showed that some Hnt- and Br-expressing follicle cells were still in mitotic cycle (Figs. 5A,B), further confirming that mitotic cycle continues until stage 5.
Co-staining of PH3 with Br, Hnt and Cut. (A–C) Confocal images showed the co-stainings, Br, Hnt and Cut antibody staining in green, PH3 in red and DAPI in blue. Co-expressing follicle cells were indicated by arrowheads. (D) Statistical analysis of data obtained from egg chamber staging of transitional Br, Hnt, and Cut expression with PH3 co-staining. Percentage of PH3-positive egg chambers were labeled on top of each column. Scale bar =10 µm.
Discussion
Overall, the importance of the Notch pathway for many aspects of development has clearly been demonstrated by numerous publications, and the pathway itself has been studied extensively. The Notch pathway is essential for many diseases, including Zika virus disease26, cardiovascular diseases27, Down syndrome28, and various cancer types29,30,31. Therefore, better understanding of the complex Notch pathway from all aspects are needed for potential therapeutic values. While there has been extensive research entailing what the Notch signaling pathway does and what processes it may be directly and indirectly involved in, the temporal expression of its major targets relative to one another has not been extensively studied.
During Drosophila oogenesis, Notch signaling plays a vital role in cell cycle switches. Previous research has discovered several important downstream targets in this pathway, but comparatively little is known about the timing in which they are activated in normal developmental processes. As downstream targets may respond to Notch signaling at different times, insight into this activation sequence may have many implications for the field. We previously studied some important downstream targets of Notch signaling, including cut, hindsight (hnt) and broad (br), during egg chamber development. In addition, the three targets also showed varied sensitivity to Notch. Therefore, we decided to examine these three important Notch targets to understand the temporal patterning. Immunostaining is the most popular way to examine the expression patterns of certain proteins. However, primary antibodies raised in the same host species can’t be stained simultaneously. In our case, we found the publicly available primary antibodies from Developmental Studies Hybridoma Bank. Unfortunately, the host species of primary antibodies of mouse anti-Cut (2B10), mouse anti-Br-Core (25E9), mouse anti-Hnt (1G9) were the same. To overcome this barrier, we applied a previously developed MATLAB toolbox to automatically and unbiasedly identify egg chamber stages based on egg chamber morphology. Using the toolbox, we were able to decipher the temporal expression patterns of Br, Hnt and Cut. The effectiveness of the toolbox also enabled our scientific community to study egg chamber temporal expression patterns, even though scientists might not have primary antibodies from different host species.
Surprisingly, we found that both Br and Hnt were upregulated by Notch signaling as early as stage 4, and Br was upregulated earlier than Hnt (Fig. 4). Previously, it was well known that Notch signaling was activated by Delta ligand binding to Notch receptor at stage 518,19. We speculate that there was a biased focus on strong expression of Notch signaling components to define the activation stages. Here, we focused on the specific transitional stages with weak or partial expression of target genes. With systematic analyses, we showed that Notch signaling could be activated at stage 4, one stage earlier than the stage 5, a previously long-held belief. In addition, downregulation of Cut takes place at stage 5, earlier than previously believed stages 6/7. Taken together, for the first time, we discovered that Notch signaling is activated at stage 4. As a direct transcriptionally regulated target of Notch, Br is upregulated at stages 4/5. Later than Br, Hnt is activated at stages 4/5. Due to the upregulation of Notch signaling, Cut expression is downregulated at stage 5. Our further mitotic marker analysis re-stated that mitotic cycle continues until stage 5 (Fig. 6). The observed variability in the weak/partial expression patterns at transitional stages might be caused by the unsynchronized Notch activation in the follicle cell. It might take time for inherent property of the cell cycle regulation system to finally transit the egg chamber into the endocycle at stage 6. It will also be interesting to study how the Notch signaling downstream targets interact with each other to boost the cell cycle transition at this special transitional stage. Further research into this aspect of Notch signaling may serve as a basis for additional investigation of developmental issues and defects. Better understanding of the complex Notch pathway and its downstream network could shed new lights on developmental regulations and cancer mechanisms.
Model for broad, cut, and hindsight expression patterning during Drosophila egg chamber development. The model is based on statistical analysis of data acquired from egg chamber staging. Transitional stages for each target appear near stages 4, 5, and 6 of development with broad upregulation happening first, hindsight upregulation next, then cut downregulation occurring from stage 5. Our further mitotic marker analysis re-stated that mitotic cycle continues until stage 5.
Materials and Methods
Fly strains and genetics
Wild-type (w1118) Drosophila melanogaster flies were obtained from Bloomington Drosophila Stock Center (#5905), cultured with standard Bloomington medium and fed with yeast paste two days before dissection to enlarge ovaries. Female Drosophila were anesthetized and dissected according to standard techniques32.
Immunohistochemistry and image acquirement
Immunohistochemistry was performed according to previous description32. The primary and secondary antibodies used in this study were as follows: mouse anti-Cut (2B10) (1:50; Developmental Studies Hybridoma Bank, USA), mouse anti-Br-Core (25E9) (1:30; Developmental Studies Hybridoma Bank, USA), mouse anti-Hnt (1G9) (1:15; Developmental Studies Hybridoma Bank, USA), rabbit anti-PH3 1:200 (Upstate Biotechnology, NY, USA) and Alexa Fluor secondary antibodies (1:400; Invitrogen). To stain nuclei, DAPI (1:500; Invitrogen) was applied. All samples were imaged using a Zeiss LSM 710 confocal microscope at Georgia Southern University. The acquired DAPI images represented the largest cross-sectional area of the egg chambers in the middle plane. Images were obtained of egg chambers showing partial, weak, strong and no expression of the targets of interest.
Image processing methods
Images of DAPI staining were formatted using the Fiji package from Image J, and the images were cropped to display a single egg chamber. Egg chamber images were further analyzed using scripts written in MATLAB, as previously described11. This toolbox was used to automatically identify egg chamber stages based on egg chamber size and ratio. Egg chamber size was the size of the largest cross-sectional area of the egg chambers in the middle plane, and was measured in μm2. The egg chamber ratio was the ratio of the major and minor axes. Visual determination of egg chamber stage based on morphological criteria was used to confirm toolbox findings11,17.
Quantitative methods
Statistical analyses were performed based on egg chamber staging data. Boxplots were used to display the distribution of data in Fig. 4. Mean values of each group were indicated by the red dots in the figure. The stage distribution plot in the Fig. 4 showed the frequency of specimens in each stage (from 4–6) in Br, Hnt and Cut groups. Beside the visualization, pairwise permutation tests were also conducted to assess the differences of the group-wise means. To control the familywise error rate, Bonferroni correction was further applied33. Since there were 3 tests in each table, instead of setting the critical P value for significance to 0.05, we use a lower critical value 0.017 (0.05/the number of tests = 0.017). In the boxplot, we would only consider individual tests with P < 0.017 to be statistically significant.
Data availability
The authors declare that the main data of this study are available within the article.
References
Radhakrishnan, K., Halasz, A., Vlachos, D. & Edwards, J. S. Quantitative understanding of cell signaling: the importance of membrane organization. Curr Opin Biotechnol 21, 677–682, https://doi.org/10.1016/j.copbio.2010.08.006 (2010).
Kopan, R. & Ilagan, M. X. The canonical Notch signaling pathway: unfolding the activation mechanism. Cell 137, 216–233, https://doi.org/10.1016/j.cell.2009.03.045 (2009).
Artavanis-Tsakonas, S., Rand, M. D. & Lake, R. J. Notch signaling: cell fate control and signal integration in development. Science 284, 770–776, https://doi.org/10.1126/science.284.5415.770 (1999).
Borggrefe, T. & Oswald, F. The Notch signaling pathway: Transcriptional regulation at Notch target genes. Cell Mol Life Sci 66, 1631–1646, https://doi.org/10.1007/s00018-009-8668-7 (2009).
Yuan, X. et al. Notch signaling: an emerging therapeutic target for cancer treatment. Cancer letters 369, 20–27, https://doi.org/10.1016/j.canlet.2015.07.048 (2015).
Bray, S. J. Notch signalling in context. Nat Rev Mol Cell Biol 17, 722–735, https://doi.org/10.1038/nrm.2016.94 (2016).
Andersson, E. R., Sandberg, R. & Lendahl, U. Notch signaling: simplicity in design, versatility in function. Development 138, 3593–3612, https://doi.org/10.1242/dev.063610 (2011).
Murta, D. et al. Notch signaling in the epididymal epithelium regulates sperm motility and is transferred at a distance within epididymosomes. Andrology 4, 314–327, https://doi.org/10.1111/andr.12144 (2016).
Li, X. et al. Temporal patterning of Drosophila medulla neuroblasts controls neural fates. Nature 498, 456–462, https://doi.org/10.1038/nature12319 (2013).
Bertet, C. et al. Temporal patterning of neuroblasts controls Notch-mediated cell survival through regulation of Hid or Reaper. Cell 158, 1173–1186, https://doi.org/10.1016/j.cell.2014.07.045 (2014).
Jia, D., Xu, Q., Xie, Q., Mio, W. & Deng, W. M. Automatic stage identification of Drosophila egg chamber based on DAPI images. Scientific reports 6, 18850, https://doi.org/10.1038/srep18850 (2016).
Sun, J. J. & Deng, W. M. Notch-dependent downregulation of the homeodomain gene cut is required for the mitotic cycle/endocycle switch and cell differentiation in Drosophila follicle cells. Development 132, 4299–4308, https://doi.org/10.1242/dev.02015 (2005).
Sun, J. & Deng, W. M. Hindsight mediates the role of notch in suppressing hedgehog signaling and cell proliferation. Developmental cell 12, 431–442, https://doi.org/10.1016/j.devcel.2007.02.003 (2007).
Jia, D., Tamori, Y., Pyrowolakis, G. & Deng, W. M. Regulation of broad by the Notch pathway affects timing of follicle cell development. Developmental biology 392, 52–61, https://doi.org/10.1016/j.ydbio.2014.04.024 (2014).
Jouandin, P., Ghiglione, C. & Noselli, S. Starvation induces FoxO-dependent mitotic-to-endocycle switch pausing during Drosophila oogenesis. Development 141, 3013–3021, https://doi.org/10.1242/dev.108399 (2014).
Jia, D. et al. A large-scale in vivo RNAi screen to identify genes involved in Notch-mediated follicle cell differentiation and cell cycle switches. Scientific reports 5, 12328, https://doi.org/10.1038/srep12328 (2015).
Spradling, A. Developmental genetics of oogenesis. In The development of Drosophila melanogaster. Cold Spring Harbor Lab. Press, NY (1993).
Lopez-Schier, H. & St Johnston, D. Delta signaling from the germ line controls the proliferation and differentiation of the somatic follicle cells during Drosophila oogenesis. Gene Dev 15, 1393–1405, https://doi.org/10.1101/Gad.200901 (2001).
Roth, S. Drosophila oogenesis: Coordinating germ line and soma. Curr Biol 11, R779–R781, https://doi.org/10.1016/S0960-9822(01)00469-9 (2001).
Tseng, C. Y. & Hsu, H. J. Decreased expression of lethal giant larvae causes ovarian follicle cell outgrowth in the Drosophila Scutoid mutant. Plos One 12, doi:ARTN e018891710.1371/journal.pone.0188917 (2017).
Blochlinger, K., Jan, L. Y. & Jan, Y. N. Postembryonic patterns of expression of cut, a locus regulating sensory organ identity in Drosophila. Development 117, 441–450 (1993).
Dej, K. J. & Spradling, A. C. The endocycle controls nurse cell polytene chromosome structure during Drosophila oogenesis. Development 126, 293–303 (1999).
Blochlinger, K., Bodmer, R., Jan, L. Y. & Jan, Y. N. Patterns of Expression of Cut, a Protein Required for External Sensory Organ Development in Wild-Type and Cut Mutant Drosophila Embryos. Gene Dev 4, 1322–1331, https://doi.org/10.1101/Gad.4.8.1322 (1990).
Emery, I. F., Bedian, V. & Guild, G. M. Differential Expression of Broad-Complex Transcription Factors May Forecast Tissue-Specific Developmental Fates during Drosophila Metamorphosis. Development 120, 3275–3287 (1994).
Yip, M. L. R., Lamka, M. L. & Lipshitz, H. D. Control of germ-band retraction in Drosophila by the zinc-finger protein HINDSIGHT. Development 124, 2129–2141 (1997).
Ferraris, P. et al. Zika virus differentially infects human neural progenitor cells according to their state of differentiation and dysregulates neurogenesis through the Notch pathway. Emerging microbes & infections 8, 1003–1016, https://doi.org/10.1080/22221751.2019.1637283 (2019).
Fortini, F. et al. Estrogen-mediated protection against coronary heart disease: The role of the Notch pathway. The. Journal of steroid biochemistry and molecular biology 189, 87–100, https://doi.org/10.1016/j.jsbmb.2019.02.008 (2019).
Yusof, H. H. et al. Expression Profiling of Notch Signalling Pathway and Gamma-Secretase Activity in the Brain of Ts1Cje Mouse Model of Down Syndrome. Journal of molecular neuroscience: MN 67, 632–642, https://doi.org/10.1007/s12031-019-01275-2 (2019).
Chatterjee, S. & Sil, P. C. Targeting the crosstalks of Wnt pathway with Hedgehog and Notch for cancer therapy. Pharmacological research 142, 251–261, https://doi.org/10.1016/j.phrs.2019.02.027 (2019).
Leonetti, A. et al. Notch pathway in small-cell lung cancer: from preclinical evidence to therapeutic challenges. Cellular oncology 42, 261–273, https://doi.org/10.1007/s13402-019-00441-3 (2019).
Cui, Q. et al. GXYLT2 accelerates cell growth and migration by regulating the Notch pathway in human cancer cells. Experimental cell research 376, 1–10, https://doi.org/10.1016/j.yexcr.2019.01.023 (2019).
Jia, D., Huang, Y. C. & Deng, W. M. Analysis of Cell Cycle Switches in Drosophila Oogenesis. Methods in molecular biology 1328, 207–216, https://doi.org/10.1007/978-1-4939-2851-4_15 (2015).
Simes, R. J. An Improved Bonferroni Procedure for Multiple Tests of Significance. Biometrika 73, 751–754, https://doi.org/10.2307/2336545 (1986).
Acknowledgements
We would like to thank the Biology Microscope Facility at Georgia Southern University for technical help, the Bloomington Stock Center for fly stocks, and the Developmental Studies Hybridoma Bank for antibodies. D. J. is supported by Start-Up Funding, College of Science and Mathematics Research Grant, Summer Research Session Award, and Research Seed Funding Award from Georgia Southern University. M.R. is supported by the Undergraduate Research Travel Award from Georgia Southern University. M.R.’s thesis contributes to part of the work. L.P. is supported by Undergraduate Research Assistantship Funding from Georgia Southern University.
Author information
Authors and Affiliations
Contributions
D.J. designed the experiments; M.R., L.P., and F.T. collected the data; D.J., M.R., L.P., Q.Xu., Q.Xie. and M.L. participated in the data analysis; M.R. and D.J. drafted the paper; D.J. and A.J. created the figures; all authors contributed to the paper revision. All authors approved the manuscript for submission.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Rowe, M., Paculis, L., Tapia, F. et al. Analysis of the Temporal Patterning of Notch Downstream Targets during Drosophila melanogaster Egg Chamber Development. Sci Rep 10, 7370 (2020). https://doi.org/10.1038/s41598-020-64247-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-64247-2
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.