Abstract
The continued use of pyrazinamide in the treatment of tuberculosis in the absence of a rapid, accurate and standardized pyrazinamide drug susceptibility assays is of great concern. While whole genome sequencing holds promise, it is not yet feasible option in low resource settings as it requires expensive instruments and bioinformatic analysis. We investigated the diagnostic performance of a closed-tube Linear-After-The-Exponential (LATE)-PCR assay for pyrazinamide susceptibility in Mycobacterium tuberculosis. Based on a set of 654 clinical Mycobacterium tuberculosis culture isolates with known mutations throughout the pncA gene as determined by Sanger sequencing, the assay displays excellent sensitivity of 96.9% (95% CI: 95.2–98.6) and specificity of 97.9% (95% CI: 96.1–99.7). In a subset of 384 isolates with phenotypic drug susceptibility testing, we also observed high sensitivity of 98.9% (95% CI: 97.5–100) but lower specificity of 91.8% (95% CI: 87.9–95.8) when compared to phenotypic drug susceptibility testing. We conclude that the LATE PCR assay offers both a rapid and accurate prediction of pyrazinamide susceptibility.
Similar content being viewed by others
Introduction
Background
Pyrazinamide (PZA) is an integral component of the treatment regimens for drug susceptible and drug resistant tuberculosis (TB) that are currently recommended by the World Health Organization (WHO)1,2,3,4. Moreover, PZA will likely remain an important anti-TB drug in the future as it is included in multiple ongoing clinical trials for drug susceptible and drug resistant TB due to its unique synergistic properties with new TB drugs (pretomanid and bedaquiline)5,6. In light of the high rates of PZA resistance (39% − 60%) in patients with multi-drug resistant (MDR)-TB and extensively drug resistant (XDR)-TB7,8, rapid assessment of PZA susceptibility is important to enable clinicians to select ≥ 4 effective antibiotics to increase chance of treatment success. Nevertheless, drug susceptibility testing (DST) for PZA is rarely performed because it requires stringent control of pH and inoculum size9,10,11 and is therefore unreliable in most settings12,13,14,15.
Several studies have demonstrated that numerous mutations in the pncA gene are the primary cause of resistance to PZA16,17,18. It is hypothesized that these variants cause decreased PZase activity, thereby limiting the conversion of PZA to the active form of pyrazinoic acid (POA) within M. tuberculosis19. DNA sequencing has identified over 600 different insertion, deletions and single nucleotide polymorphisms (SNPs) across the entire length of the pncA gene, making identification of resistance causing mutations in this gene extremely challenging using methods other than sequencing the entire pncA gene8,20,21,22,23. Sequencing is therefore regarded as essential to detect all possible variants associated with PZA resistance and involves either Sanger sequencing of a PCR amplified pncA gene product24, targeted deep sequencing, or whole genome sequencing (WGS)25. Three recent large studies assessed the performance of molecular assays to predict PZA drug susceptibility26,27,28. A study using targeted sequence data observed an overall sensitivity of 49% and specificity of 98%26. One WGS studies achieved similar specificities but higher sensitivity of 65%27. When also including low-frequency SNPs a sensitivity of 91% was achieved.
WGS currently requires a culture isolate, making this approach unrealistic for routine laboratories in most high burden countries. The closed-tube Linear-After-The-Exponential (LATE)-PCR assay offers a novel and promising alternative. To detect all possible sequence variations in the pncA gene, the LATE-PCR29 method efficiently generates an abundance of one full length pncA gene target strand over the other and employs Lights-On/Lights-Off probes30 in the same closed-tube. LATE-PCR is a non-symmetric PCR method that uses a Limiting Primer and an Excess Primer whose initial melting temperature adhere to the equation Tm0L − Tm0X ≥ 0. This simple rule guarantees efficient amplification of double strands, followed by efficient amplification of a single strand when the Limiting Primer runs out. The Lights-On/Lights-Off probes are comprised of sets of dual-labelled fluorescent probes and adjacent quencher-only probes that coat the entire target sequence when the reaction temperature is dropped at end-point. The unique sequence of the underlying target is revealed by the temperature-dependent pattern in which the Lights-On and Lights-Off probes melt off when the temperature at end-point is slowed increased. This approach allows for highly reliable and robust detection of mutations in a target amplicon in the absence of any form of sequencing and without opening the tube.
LATE-PCR and Lights-On/Lights-Off probes have already been used before to detect mutations in the rpoB, katG, inhA promoter genes/regions responsible for resistance to rifampicin and isoniazid in M. tuberculosis31. The present report demonstrates for the first time that LATE-PCR can be used to efficiently amplify the GC-rich target pncA gene that is 685 nucleotides long. Lights-On/Lights-Off probe pairs can be designed in standardized sets comprised of shorter Off-probes and longer On-probes whose placement along the single-stranded target sequence does not need to be fitted to the target sequence, which may or may not fold into temperature-dependent hairpins. In addition, the innovative probe design used here did not use the three different colors to partition the target into separate sections, as described previously30,31. Instead, probes labelled with Quasar 670, Cal Red 610, and Cal Orange 560 are interspersed along the length of the target with the result that the fluorescent signature in each color is a composite fluorescent signature of separate positions along the length of the target.
Results
Comparison of PZA susceptibility using Sanger sequencing vs lCosed-Tube Assay
Examples of the fluorescence signatures for wildtype and variants in the different fluorescent colors are shown in Fig. 1, along with additional signatures in supplemental data (supplemental figures). In Fig. 1A and B we observe fluorescent signatures for the reference strain and the test strain to overlap in Q670 and CR610, however the fluorescent signature shifts by 1 °C around the 60 °C in CO560 (Fig. 1C) which suggests a variant is present in the test strain compared to the reference strain. Supplemental table 2 lists the pncA genotypes detected using Sanger sequencing, the PZA phenotypes, and the Closed-Tube Assay Scores for all isolates. Among the 654 Mtb isolates, Sanger sequencing identified 239 (36.5%) as wild type pncA sequences while the Closed-Tube Assay scored 234 isolates as wildtype, for a concordance of 97.9%. Sanger sequencing identified 415 (63.5%) as pncA variant sequences. Among a total of 181 different pncA mutations, the genetic alteration included: insertions (n = 20), deletions (n = 18), SNPs (n = 123), promoter SNPs (n = 7), and double SNPs (n = 13). Although the Closed-Tube Assay does not distinguish these different classes of mutations, its concordance with Sanger sequencing for overall mutation detection was 97.2% (636/654). Of the 415 isolates identified as mutations by sequencing, 402 (96.9%) were also classified as pncA variants by the Closed-Tube Assay.
(A) Lights-On/Lights-Off probe fluorescence signature for Quasar 670. This figure shows the first derivative of the fluorescence profile for Quasar 670. The red fluorescence signature is the control of H37Rv; the blue fluorescence signature is the test/unknown sample and the black fluorescence signature is the no template control (NTC). No observed difference in this figure. (B) Lights-On/Lights-Off probe fluorescence signature for Cal Fluor Red 610. This figure shows the first derivative of the fluorescence profile for Cal Fluor Red 610. The red fluorescence signature is the control of H37Rv; the blue fluorescence signature is the test/unknown sample and the black fluorescence signature is the no template control (NTC). No observed difference in this figure. (C) Lights-On/Lights-Off probe fluorescence signature for Cal Fluor Orange 560. This figure shows the first derivative of the fluorescence profile for Cal Fluor Orange 560. The red fluorescence signature is the control of H37Rv; the blue fluorescence signature is the test/unknown sample and the black fluorescence signature is the no template control (NTC). We observe a shift in the fluorescence signature (indicated at the arrow), revealing that this strain harbors a variant.
Overall the comparison of Sanger sequencing and the Closed-Tube Assay demonstrate that the novel approached used here had a sensitivity of 96.9% (95% CI: 95.2–98.6) and specificity of 97.9% (95% CI: 96.1–99.7) for the detection of pncA variants.
Phenotypic PZA DST vs Closed-Tube detection for determination of PZA susceptibility
Among the isolates there was a subset of 384 for which pDST results were available: 188 (48.9%) were resistant to PZA and 196 (51.1%) were susceptible to PZA. The Closed-Tube Assay identified 202 (52.6%) of these isolates as pncA variants and 182 (47.4%) as pncA wildtype. Thus, the overall concordance between the Closed-Tube Assay and the pDST culture method was 95.3% (366/384). Of the 196 phenotypically PZA susceptible isolates, 180 (91.8%) were also classified as PZA susceptible by the Close-Tube Assay. Of the 188 phenotypically PZA resistant isolates, 186 (98.9%) were correctly classified as drug resistant by Closed-Tube Assay. Thus, diagnosis of PZA resistance using the Closed-Tube Assay has a sensitivity of 98.9% (95% CI: 97.5–100) and specificity of 91.8% (95% CI: 87.9–95.8).
Discussion
The results presented here demonstrate that the closed-tube method that was previously described for the detection of mutations with relatively short amplicons from the rpoB, katG and inhA promotor regions31 is a flexible technology that can be adapted for detection of virtually all sequence variants in a large set of purified Mtb DNA isolates from patients in South Africa and the United States. Moreover, these sequence variants are spread throughout the full length pncA gene target of 685 nucleotides. The strains tested included pncA SNPs widely distributed across the entire pncA gene (Fig. 2), providing a good representation of the distribution of mutations observed globally8. The performance of the Closed-Tube Assay was excellent, with a sensitivity of 96.9% and specificity of 97.9% compared to pncA Sanger sequencing, and a sensitivity of 98.9% and specificity of 91.8% when compared to pDST testing. The assays utility was not affected by the position in the gene that was analysed. The observed performance is similar to that reported for Sanger sequencing of the pncA gene, a line probe-based assay (Genoscholar PZA-TB II line probe assay, NIPRO Corporation, Japan) and WGS24,32. The line probe assay showed high 97.6% concordance with a composite reference standard pncA Sanger and Illumina sequencing plus phenotypic susceptibility testing. A pncA Sanger sequencing study observed a sensitivity of 90.9% and specificity were of 100% when using pDST as the reference standard24. Both, Sanger sequencing and pDST require additional steps, which are not required in the close-tube method. The Sanger sequencing, while returning a result within 48 hours, still requires a sequencing instrument. The line probe-based assay avoids the need for a sequencing instrument, but both assays require transfer of DNA (opening the tube) either to load onto a sequencer or for hybridization onto a strip membrane, this creates the opportunity for environmental contamination. The sensitivity and specificity achieved using the closed-tube approach are comparable to what was observed in the 100,000 genomes project (91% and 97% sensitivity and specificity), but did not require the use of costly WGS28. The cost of goods of this single tube assay is under $0.50, this includes the plastics, primers, probes, and PCR reagents, but excludes the cost for DNA extraction and culture of the isolate.
Of the 18 isolates which showed discordant results, 5 were scored as pncA wildtype using Sanger sequencing, but showed a shift in their fluorescent signatures using the Closed-Tube Assay and 13 contained a pncA variant using Sanger sequencing, but showed no shift in fluorescent signatures. Of the 5 isolates scored a wildtype on Sanger sequencing, 4 had changes in the Cal Fluor Red 610 fluorophore channel. The fluorescent signature shifts can be detected down to 90:10 mixtures of different genotypes (unpublished data). Thus, 4 of the 5 isolates scored as wildtype by Sanger sequencing may have been due to the presence of hetero-resistance that is below the level of detectability by Sanger sequencing33,34. The pncA variants in isolates where no shift in the fluorescent signatures were not clustered and observed between codon 27 and codon 187 as well as in the promoter region. It is also possible that there is a shift in the fluorescent signatures that is extremely subtle and is not detected.
The excellent sensitivity and specificity coupled with a rapid turnaround time for a result (same day), and relatively inexpensive costs of the reaction components means that the Closed-Tube method holds promise for a diagnostic to identify pncA mutations. Nevertheless, there are some limitations of the study described here. First, the samples used in this study were purified DNA, which requires culturing for 4–6 weeks as well as DNA extraction. This could be resolved by validating the assay using a “raw” specimen type such as the Hain Lifescience FluoroLyse Kit for Genomic Bacterial DNA (Hain Lifescience, GmbH, Nehren, Germany) from both sputum as well as culture. This would demonstrate the ease of use at point of care, where no culturing is required. Second, while we evaluated an extensive list of different clinical pncA mutations the full spectrum of mutations has not been evaluated. Finally, the assay focuses on any mutation in the entire pncA gene, which may result in identification of mutations not associated with PZA resistance26,35,36. The next version of the assay could be expanded to include targets from the rpsA gene, since mutations in this gene have been associated with PZA resistance and to develop an algorithm to relate specific mutations to fluorescent signatures for pncA mutations not associated with PZA resistance37,38.
In summary, we have successfully designed and tested a single Closed-Tube Assay that is able to identify and distinguish the wildtype pncA gene from strains with pncA sequence variants that cause PZA resistance. This assay is for detection of PZA resistance accurate and rapid, and relatively inexpensive compared to both genotyping by sequencing and phenotyping by culture. Additional refinements, including use of PCR thermal cyclers that are more precise than the Stratagene MX3005p, will improve identification of the rare synonymous mutations and the few non-synonymous mutations in pncA that were not correctly scored as PZA resistance. This virtual sequencing method is a promising advance in novel diagnostics for PZA and could be integrated alongside the GeneXpert (Cepheid, Sunnyvale, CA, USA) or Line Probe Assay to determine the patient’s resistance profile so that the treatment can be optimised. Parallel comparisons using these different methods would likely also improve understand of resistance to PZA, which is such an important drug in the fight against tuberculosis.
Methods
Ethics statement
Methods and protocols were carried out in accordance with the relevant guidelines and regulations. The isolates used from South Africa was approved by the Health Research Ethics Committee of Stellenbosch University (S12/01/020). The isolates used from the Center for Discovery and Innovation were selected from an archived collection of isolates all of which are de-identified.
Clinical isolates
Clinical Mycobacterium tuberculosis (Mtb) culture isolates (n = 654) were selected from a culture bank at Stellenbosch University (n = 424) and from the global Mtb strain collection archived at the Center for Discovery and Innovation (n = 230). Isolates were selected to have as many different pncA variants as possible and where possible biological replicates of different pncA variants as well as wild type. DNA was purified from each isolate as per the procedure described by Warren et al.39. The pncA gene was Sanger sequenced for all samples according to the method of Streicher et al.24.
The clinical isolates from Stellenbosch University included 115 RIF-mono resistant; 38 poly-resistant; 158 MDR-TB and 113 XDR-TB isolates. Unfortunately, the drug resistant profile was not available for the clinical isolates from the archived collection at the Center for Discovery and Innovation.
Phenotypic susceptibility to PZA was performed using the non-radiometric BACTEC MGIT 960 method as per the manufacturer’s instructions. This system makes use of modified test media to support the growth at a pH of 5.9. To distinguish between resistant and susceptible isolates, a critical concentration of 100 µg/ml PZA is utilized (BD Diagnostic Systems, NJ, USA). PZA drug susceptibility testing (DST) results were available from a subset of isolates, 384 of 654 (59%).
Closed-tube assay for detection of pncA gene variants
LATE-PCR29,40,41 primers (pncA_LP3: 5’ – GCGGCGTCATGGACCCTATATCTGTGGCTGCCGCGTC – ‘3 and pncA_XP3 5’ – TTGCTCCACCGCCGCCAACAGTTCAT – ‘3) were designed to amplify a 685 bp amplicon encompassing the entire coding region of the pncA gene, the 37 bp of the upstream sequence and 24 bp of the downstream sequence. In total, 17 pairs of Lights-On/Lights-Off probes were designed, according to Rice et al. protocol30, to cover the entire pncA gene including upstream and downstream flanking regions (Supplemental table 1). Lights-On probes of between 20–26 bp were designed to be complementary to a distinct region of the single-stranded DNA of pncA generated by the LATE-PCR. The primers and probes were designed using Visual-OMP software (DNA Software Inc., Ann Arbor, MI, USA). Each Lights-On probe is a low-temperature dual-labelled molecular beacon with a two base pair self-complementary stem and was labelled with a fluorophore on its 5’end and a Black Hole quencher on its 3’end. Each of the 17 Lights-Off probes serve to absorb energy from the fluorophore of its adjacent Lights-On probe when both are bound to the target strand. Lights-Off probes of 15 bp were labelled with a Black Hole Quencher I (BHQ-1) or Black Hole Quencher 2 (BHQ-2) quencher. Some probes included nucleotide mismatches to their wildtype target sequences to adjust the probe melting temperature. Probes designed in this way will nevertheless change Tm when hybridized to a target sequence that includes a mutation.
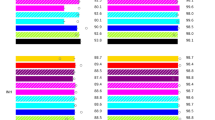
The probes were catalogued into three groups according to their fluorophore labels (Quasar 670, Cal Fluor Red 610 and Cal Fluor Orange 560) (Fig. 3). Multiple pairs of probes of the same color or different colors can be used to analyse sequences several hundred nucleotides long. During development the probes Tm and binding were checked utilizing Visual-OMP software (DNA Software Inc., Ann Arbor, MI, USA). Each fluorescent contour was transformed into its first derivative curve, which we refer to as a fluorescent signature30. This design results in a unique fluorescent signature for each probe target sequence, making it possible to detect the presence or absence of any nucleotide change in the pncA gene and flanking regions.
Lights-On/Lights-Off probe placement on the coding region of the pncA gene. pncA probes in the three fluorescence colors showing each probes respective Tm. The dashed bars separate different fluorescent colors (Quasar 670; Cal Fluor Red 610; Cal Fluor Orange 560). The black bars indicate the ON probes while the grey bars indicate the OFF probes. The probe name indicates the gene (pncA), the nucleotide position in the pncA gene, and lastly whether the probe is an ON or OFF probe.
PCR amplification of extracted Mtb DNA was carried out using a Stratagene MX3005p device (Agilent Technologies, Santa Clara, CA, USA). Each reaction mixture (25 μl) contained 1 to 2.5 μl of template DNA (purified DNA 25 ng/ μl), 0.05 μM of the pncA_LP3 and 1.0 μM of the pncA_XP3 PCR primers, 1× PCR buffer, 3.5 mM magnesium, 0.6 μM of PrimeSafe40 (Smiths Detection, London, UK), 0.3 mM deoxynucleoside triphosphates, 0.05 μM of each of the 34 Lights-On probes and 0.15 μM of each of the 34 Lights-Off probes, 25 nM Internal Marker labelled with Quasar 670, 75 nM Internal Marker labelled with BHQ-2, and 0.125 μl (1.5 U) of Taq polymerase (Thermo Fisher Scientific, MA, USA). All reagents were added in a PCR preparation room. The PCR mix was then taken to another clean room where the DNA (up to 25 ng/µl) for each sample was added under sterile conditions and the tube sealed to minimize the risk of contamination. Water was included as a negative control and H37Rv DNA as a positive control. The amplification protocol consisted of 60 cycles of 98 °C for 10 seconds and 75 °C for 40 seconds, followed by a final elongation step of 75 °C for 10 minutes. After amplification, the temperature of the reaction was lowered to 25 °C for 10 minutes. This results in the coating of the single-stranded DNA templates with 17 Lights-On/Lights-Off fluorescent probe pairs labeled in three colors. The temperature was then increased from 25 °C to 95 °C at a rate of 1 °C for 30 seconds while concurrently measuring fluorescence. Fluorescent signatures were compared to the signature of a pan-susceptible strain (H37Rv). Deviations in the signature pattern were interpreted (visual inspection) to reflect the presence of genetic variants in the pncA gene as they would alter the melting temperature of the complementary probe.
Statistical analysis
The sensitivity and specificity of the virtual sequencing method were determined with R: Foundation for Statistical Computing, https://www.R-project.org (Vienna, Austria).
Data availability
The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request.
References
World Health Organization. Global Tuberculosis Report, http://apps.who.int/medicinedocs/documents/s23098en/s23098en.pdf (2016).
Organization, W. H. Global Tuberculosis Report, http://apps.who.int/iris/bitstream/10665/259366/1/9789241565516-eng.pdf?ua=1 (2017).
Organization, W. H. Treatment guidelines for drug-resistant tuberculosis 2016 update (October 2016 Revision), http://apps.who.int/iris/bitstream/handle/10665/250125/9789241549639-eng.pdf;jsessionid=3CBE6F3CC2F3C756D62EE6D9153B877B?sequence=1 (2016).
Organization, W. H. The Shorten MDR-TB Regimen, http://www.who.int/tb/Short_MDR_regimen_factsheet.pdf (2016).
Dawson, R. et al. Efficiency and safety of the combination of moxifloxacin, pretomanid (PA-824), and pyrazinamide during the first 8 weeks of antituberculosis treatment: a phase 2b, open-label, partly randomised trial in patients with drug-susceptible or drug-resistant pulmonary tuberculosis. The Lancet 385, 1738–1747, https://doi.org/10.1016/S0140-6736(14)62002-X (2015).
Diacon, A. H. et al. Randomized Pilot Trial of Eight Weeks of Bedaquiline (TMC207) Treatment for Multidrug-Resistant Tuberculosis: Long-Term Outcome, Tolerability, and Effect on Emergence of Drug Resistance. Antimicrob. Agents Chemother. 56, 3271–3276, https://doi.org/10.1128/aac.06126-11 (2012).
Whitfield, M. G. et al. Prevalence of pyrazinamide resistance across the spectrum of drug resistant phenotypes of Mycobacterium tuberculosis. Tuberculosis 99, 128–130, https://doi.org/10.1016/j.tube.2016.05.003 (2016).
Whitfield, M. G. et al. A Global Perspective on Pyrazinamide Resistance: Systematic Review and Meta-Analysis. PLoS One 10, e0133869, https://doi.org/10.1371/journal.pone.0133869 (2015).
Fonseca, Ld. S. et al. Correlation between resistance to pyrazinamide and resistance to other antituberculosis drugs in Mycobacterium tuberculosis strains isolated at a referral hospital. J. Bras. Pneumol. 38, 630–633 (2012).
Zhang, Y., Permar, S. & Sun, Z. H. Conditions that may affect the results of susceptibility testing of Mycobacterium tuberculosis to pyrazinamide. J. Med. Microbiol. 51, 42–49 (2002).
Zhang, Y. & Mitchison, D. The curious characteristics of pyrazinamide: a review. Int. J. Tuberc. Lung Dis. 7, 6–21 (2003).
World Health Organization. Global Tuberculosis Report, http://apps.who.int/iris/bitstream/10665/91355/1/9789241564656_eng.pdf (2014).
World Health Organization. Global Tuberculosis Report, http://apps.who.int/iris/bitstream/10665/75938/1/9789241564502_eng.pdf (2012).
World Health Organization. Guidelines for the programmatic management of drug-resistant tuberculosis: emergency update 2008, http://whqlibdoc.who.int/publications/2011/9789241501583_eng.pdf (2008).
World Health Organization. Global Tuberculosis Report, http://apps.who.int/iris/bitstream/10665/191102/1/9789241565059_eng.pdf (2015).
Scorpio, A. et al. Characterization of pncA mutations in pyrazinamide-resistant Mycobacterium tuberculosis. Antimicrob. Agents Chemother. 41, 540–543 (1997).
Scorpio, A. & Zhang, Y. Mutations in pncA, a gene encoding pyrazinamidase/nicotinamidase, cause resistance to the antituberculous drug pyrazinamide in tubercle bacillus. Nat. Med. 2, 662–667, https://doi.org/10.1038/nm0696-662 (1996).
Cheng, S. J., Thibert, L., Sanchez, T., Heifets, L. & Zhang, Y. pncA mutations as a major mechanism of pyrazinamide resistance in Mycobacterium tuberculosis: Spread of a monoresistant strain in Quebec, Canada. Antimicrob. Agents Chemother. 44, 528–532, https://doi.org/10.1128/aac.44.3.528-532.2000 (2000).
Sandgren, A. et al. Tuberculosis Drug Resistance Mutation Database. PLoS Med. 6, 132–136, https://doi.org/10.1371/journal.pmed.1000002 (2009).
Zimic, M., Sheen, P., Quiliano, M., Gutierrez, A. & Gilman, R. H. Peruvian and globally reported amino acid substitutions on the Mycobacterium tuberculosis pyrazinamidase suggest a conserved pattern of mutations associated to pyrazinamide resistance. Infection Genetics and Evolution 10, 346–349, https://doi.org/10.1016/j.meegid.2009.11.016 (2010).
Werngren, J. et al. Reevaluation of the Critical Concentration for Drug Susceptibility Testing of Mycobacterium tuberculosis against Pyrazinamide Using Wild-Type MIC Distributions and pncA Gene Sequencing. Antimicrob. Agents Chemother. 56, 1253–1257, https://doi.org/10.1128/aac.05894-11 (2012).
Stoffels, K., Mathys, V., Fauville-Dufaux, M., Wintjens, R. & Bifani, P. Systematic Analysis of Pyrazinamide-Resistant Spontaneous Mutants and Clinical Isolates of Mycobacterium tuberculosis. Antimicrob. Agents Chemother. 56, 5186–5193, https://doi.org/10.1128/aac.05385-11 (2012).
Somoskovi, A. et al. Sequencing of the pncA gene in members of the Mycobacterium tuberculosis complex has important diagnostic applications: Identification of a species-specific pncA mutation in “Mycobacterium canettii” and the reliable and rapid predictor of pyrazinamide resistance. J. Clin. Microbiol. 45, 595–599, https://doi.org/10.1128/jcm.01454-06 (2007).
Streicher, E. M. et al. Rapid Sequencing of the Mycobacterium tuberculosis pncA Gene for Detection of Pyrazinamide Susceptibility. J. Clin. Microbiol. 52, 4056–4057, https://doi.org/10.1128/jcm.02438-14 (2014).
Ioerger, T. R. et al. Genome Analysis of Multi- and Extensively-Drug-Resistant Tuberculosis from KwaZulu-Natal, South Africa. PLoS One 4, https://doi.org/10.1371/journal.pone.0007778 (2009).
Miotto, P. et al. A standardised method for interpreting the association between mutations and phenotypic drug resistance in Mycobacterium tuberculosis. Eur. Respir. J. 50 (2017).
Coll, F. et al. Genome-wide analysis of multi- and extensively drug-resistant Mycobacterium tuberculosis. Nat. Genet. 50, 307–316, https://doi.org/10.1038/s41588-017-0029-0 (2018).
Prediction of Susceptibility to First-Line Tuberculosis Drugs by DNA Sequencing. N. Engl. J. Med. 379, 1403–1415, https://doi.org/10.1056/NEJMoa1800474 (2018).
Pierce, K. E., Sanchez, J. A., Rice, J. E. & Wangh, L. J. Linear-After-The-Exponential (LATE)-PCR: primer design criteria for high yields of specific single-stranded DNA and improved real-time detection. Proc. Natl. Acad. Sci. USA 102, 8609–8614, https://doi.org/10.1073/pnas.0501946102 (2005).
Rice, J. E., Reis, A. H. Jr., Rice, L. M., Carver-Brown, R. K. & Wangh, L. J. Fluorescent signatures for variable DNA sequences. Nucleic Acids Res. 40, e164–e164, https://doi.org/10.1093/nar/gks731 (2012).
de Vos, M. et al. Diagnostic Accuracy and Utility of FluoroType MTBDR, a New Molecular Assay for Multidrug-Resistant Tuberculosis. J. Clin. Microbiol. 56, e00531–00518, https://doi.org/10.1128/JCM.00531-18 (2018).
Driesen, M. et al. Evaluation of a novel line probe assay to detect resistance to pyrazinamide, a key drug used for tuberculosis treatment. Clin. Microbiol. Infect. 24, 60–64, https://doi.org/10.1016/j.cmi.2017.05.026 (2018).
Pholwat, S., Stroup, S., Foongladda, S. & Houpt, E. Digital PCR to Detect and Quantify Heteroresistance in Drug Resistant Mycobacterium tuberculosis. PLoS One 8, e57238, https://doi.org/10.1371/journal.pone.0057238 (2013).
Feuerriegel, S. et al. PhyResSE: a Web Tool Delineating Mycobacterium tuberculosis Antibiotic Resistance and Lineage from Whole-Genome Sequencing Data. J. Clin. Microbiol. 53, 1908–1914, https://doi.org/10.1128/jcm.00025-15 (2015).
Whitfield, M. G. et al. Mycobacterium tuberculosis pncA Polymorphisms That Do Not Confer Pyrazinamide Resistance at a Breakpoint Concentration of 100 Micrograms per Milliliter in MGIT. J. Clin. Microbiol. 53, 3633–3635, https://doi.org/10.1128/jcm.01001-15 (2015).
Yadon, A. N. et al. A comprehensive characterization of PncA polymorphisms that confer resistance to pyrazinamide. Nat Commun 8, 588–588, https://doi.org/10.1038/s41467-017-00721-2 (2017).
Tan, Y. et al. Role of pncA and rpsA Gene Sequencing in Detection of Pyrazinamide Resistance in Mycobacterium tuberculosis Isolates from Southern China. J. Clin. Microbiol. 52, 291–297, https://doi.org/10.1128/jcm.01903-13 (2014).
Khan, M. T. et al. Pyrazinamide resistance and mutations in pncA among isolates of Mycobacterium tuberculosis from Khyber Pakhtunkhwa, Pakistan. BMC Infect. Dis. 19, 116–116, https://doi.org/10.1186/s12879-019-3764-2 (2019).
Warren, R. et al. Safe Mycobacterium tuberculosis DNA extraction method that does not compromise integrity. J. Clin. Microbiol. 44, 254–256, https://doi.org/10.1128/JCM.44.1.254-256.2006 (2006).
Rice, J. E. et al. Monoplex/multiplex linear-after-the-exponential-PCR assays combined with PrimeSafe and Dilute-‘N’-Go sequencing. Nat. Protoc. 2, 2429, https://doi.org/10.1038/nprot.2007.362 (2007).
Sanchez, J. A., Pierce, K. E., Rice, J. E. & Wangh, L. J. Linear-after-the-exponential (LATE)-PCR: an advanced method of asymmetric PCR and its uses in quantitative real-time analysis. Proc. Natl. Acad. Sci. USA 101, 1933–1938, https://doi.org/10.1073/pnas.0305476101 (2004).
Acknowledgements
This study was funded by the National Institute of Allery and Infectious Disease, National Institutes of Health (#R21 AI106551–01) awarded to B.N.K. This study was funded by the Division of AIDS, National Institutes of Health (#R01 AI099026) and funding by the FWO (Centre for Whole Genome Sequencing of Mycobacterium tuberculosis) awarded to A.V.R. This work was supported by the National Institutes of Health (NIH) under award number #R01 A1099532 awarded to R.M.W. S.A.E.M is partly supported by a grant from the National Cancer Institute, National Institutes of Health (#R01 CA227291). This research was supported by the South African Medical Research Council. The content is solely the responsibility of the authors and does not necessarily represent the official views of the NIH, the FWO and the SAMRC. The funders had no role in study design, data collection and interpretation, or the decision to submit the work for publication.
Author information
Authors and Affiliations
Contributions
B.N.K., L.J.W. and R.M.W. conceived the experiments. M.G.W. and S.A.E.M. conducted the experiments. B.N.K. and A.V.R. provided the specimens. M.G.W., S.A.E.M., L.J.W. and J.R. analysed and interpreted the data. M.G.W. wrote the main manuscript text, prepared figures, tables and supplemental data. All authors reviewed the manuscript and provided critical input.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Whitfield, M.G., Marras, S.A.E., Warren, R.M. et al. Rapid Pyrazinamide Drug Susceptibility Testing using a Closed-Tube PCR Assay of the Entire pncA gene. Sci Rep 10, 4234 (2020). https://doi.org/10.1038/s41598-020-61286-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-61286-7
This article is cited by
-
In silico evaluation of WHO-endorsed molecular methods to detect drug resistant tuberculosis
Scientific Reports (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.