Abstract
Rice blast resistance gene, Pi54 provides broad-spectrum resistance against different strains of Magnaporthe oryzae. Understanding the cellular localization of Pi54 protein is an essential step towards deciphering its place of interaction with the cognate Avr-gene. In this study, we investigated the sub-cellular localization of Pi54 with Green Fluorescent Protein (GFP) as a molecular tag through transient and stable expression in onion epidermal cells (Allium cepa) and susceptible japonica cultivar rice Taipei 309 (TP309), respectively. Confocal microscopy based observations of the onion epidermal cells revealed nucleus and cytoplasm specific GFP signals. In the stable transformed rice plants, GFP signal was recorded in the stomata, upper epidermal cells, mesophyll cells, vascular bundle, and walls of bundle sheath and bulliform cells of leaf tissues. These observations were further confirmed by Immunocytochemical studies. Using GFP specific antibodies, it was found that there was sufficient aggregation of GFP::Pi54protein in the cytoplasm of the leaf mesophyll cells and periphery of the epidermal cells. Interestingly, the transgenic lines developed in this study could show a moderate level of resistance to Xanthomonas oryzae and Rhizoctonia solani, the causal agents of the rice bacterial blight and sheath blight diseases, respectively. This study is a first detailed report, which emphasizes the cellular and subcellular distribution of the broad spectrum blast resistance gene Pi54 in rice and the impact of its constitutive expression towards resistance against other fungal and bacterial pathogens of rice.
Similar content being viewed by others
Introduction
Blast disease is one of the dreaded diseases affecting rice (Oryza sativa L.) yield worldwide. The annual loss due to blast disease varies from 10–30%1,2. The blast pathogen M. oryzae is a hemi-biotrophic fungus having dual features of both biotroph and necrotroph. Typically, hemi-biotrophs grow initially as biotroph and then shift to necrotrophic growth, killing the host tissues3. However, M. oryzae maintains both these lifestyles simultaneously, via invasion in foliar tissues4,5. Rice blast is being managed effectively by deploying disease resistance (R) genes, and for this, identification, cloning and functional study of novel blast resistance genes is equally important. Around 26 R genes for blast resistance have been cloned and functionally characterized6,7. Plant disease resistance (R) proteins are key components to activate the defense related downstream pathways through R-Avr (avirulence) protein interaction8,9. The interaction between the host and pathogen is sophisticated and dynamic10. Rice blast pathosystem is one of the best model systems to understand the prominence and dynamics of plant-pathogen interaction. Rice, also a model monocot plant has a vast armor of multi-layered defense mechanisms against various pathogens including fungal pathogen M. oryzae11. The first layer of the defense is through passive structural barriers whereas the other layers are protein-based12. The transmembrane bound pattern recognition receptors (PRRs) mediated second layer of defense play key role in perceiving and combating the evolving pathogen or microbial-associated molecular patterns (PAMPs or MAMPs)2,13. The last layer of defense to counter the pathogen infection employs a pathogen avirulence (Avr)/effector protein triggered resistance response known as effector triggered immunity (ETI) mediated by R-genes of the host plants1,2,7,9,14,15.
Major R gene, Pi54 from rice landrace Tetep was the third R-gene to be cloned after Pib and Pit16 in rice for blast resistance. Pi54 was successfully deployed in the genetic background of rice line Taipei 309 susceptible to blast to complement the blast resistance response against different M. oryzae strains. The transgenic TP309 lines harboring Pi54 conferred high degree of resistance for diverse strains of blast pathogen collected from multiple parts of India17. The transgenic line was found to activate complex set of defense mechanism in rice which includes activation of defense response genes (laccase, PAL, callose and peroxidase), and transcription factors (MAD box, Dof Zinc finger, NAC6, WRKY and bZIP) leading to hypersensitive response and disease resistance18. Two orthologues of Pi54 gene; Pi54rh and Pi54of have also been cloned and functionally validated from wild species of rice7,19. The Pi54 gene was expressed under its native promoter in the transgenic rice lines and it shows constitutive expression at a basal level in both pathogen challenged and mock inoculated transgenic lines, however after 48 hours post inoculation (hpi) Pi54 showed induced expression in pathogen challenged lines with maximum upregulation recorded at 72 hpi18. Subsequently, Pi54 gene is being used by many researchers for blast resistance breeding programmes in rice20,21. The allelic variants of Pi54 from different rice landraces and wild relatives show extensive sequence variation which is found to be responsible for its broad spectrum resistance22,23,24.
Understanding the sub-cellular localization of resistance gene products gives an insight into the site of interaction between the resistance (R-) proteins and cognate pathogen effector (Avr-) protein. These R proteins are localized either to cytoplasm or nucleus or to both. R proteins may sometimes be localized at intracellular and intercellular membranes25. It is reported that cytoplasmic localization of resistance proteins contributes to hypersensitive response (HR), and the nuclear localization helps in achieving the complete resistance response26. Though attempts have been made to understand the sub-cellular localization of Pi54 orthologous proteins from O. rhizomatis and O. officinalis through transient expression studies in onion epidermal cells, no such attempts have been reported with respect to Pi54 protein7,19. Transient expression approach provides important insights into protein localization and possible molecular function. However, stable transformation of fluorescent tagged-Pi54 gene may help in better understanding of the cellular localization of Pi54 protein and to observe cell-cell movement in rice. Therefore, the objectives of this study were to overexpress Pi54 gene tagged with GFP under constitutive CaMV35S promoter in rice to understand sub-cellular localization of Pi54 protein using confocal microscopy and evaluation of transgenic plants for resistance against major pathogens of rice.
Results
Genetic transformation of rice using GFP::Pi54 gene construct
The rice transformation vector pRTV containing GFP::Pi54 gene was developed and confirmed by using PCR and restriction enzyme digestion (Fig. S1). The rice line TP309 susceptible to blast was used for genetic transformation using this vector. This construct was used for genetic transformation of a total of 502 calli derived from mature embryos of TP309 by using particle gun approach. After transformation, these calli were cultured on MS-hygromycin media. Healthy and proliferating calli were subcultured to regeneration medium and plantlets obtained were subjected to rooting and hardening before transferring them to the pots. In total, out of 502 calli, 127 plantlets were regenerated after three selection cycle and grown under standard rice growing conditions. Finally, 28 plants belonging to eight independent transformation events could survive up to maturity and these plantlets were further subjected to molecular analysis (Fig. 1). The average efficiency of transformation in this study was found to be 1.59% (Table S1).
Stages of rice genetic transformation using biolistic approach. (a) Induction of embryogenic calli from mature rice seeds, (b) calli subjected to bombardment, (c) proliferating calli on hygromycin selection medium; arrowhead show the actively proliferating calli, (d) shoot regeneration from hygromycin resistant calli, (e) rooting of plantlets (f) transgenic plants grown in pots under controlled conditions.
Molecular analysis of putative transgenic plants
Three sets of primers specific to genes were used to screen putative T0 generation of transgenic rice lines. PCR analysis with CaMV35S specific primers using genomic DNA confirmed the presence of expected 2.8 kb DNA fragment in19 T0 plants. Further, PCR amplification with GFP and hptII specific primers validated the presence of 750 bp and 650 bp DNA fragments, respectively. Another set of primers; GFP-F and Pi54-Rwere also used to amplify GFP and Pi54 gene fragment and a desired 650 bp DNA was amplified in the 19 transgenic lines (Fig. S2).
To confirm the stable integration of transgene, we performed Southern blot analysis. Total gDNA from eight independent PCR confirmed transgenic T1 rice lines (Fig. S3) was digested with HindIII and a DNA probe (620 bp) specific to hptII gene was used for hybridization. The results obtained confirmed the integration of candidate gene cassette in six transgenic lines (Pi54OX1 to 6). Three of the transgenic lines (Pi54OX3, 4 & 5) had single copy insertions (Fig. S4), whereas transgenic rice lines Pi54OX1 and Pi54OX6 (lane 5 and lane 10) had two copies of inserts, each. However, no hybridization was found in non-transgenic NT-TP309 plants. These transgenic lines were grown on hygromycin selection media and used for confocal microscope analysis (Fig. S5).
Expression analysis of Pi54 in transgenic plants
We used RT-PCR as well as qRT-PCR approaches to understand the nature of Pi54 expression in blast resistant transgenic rice lines. GFP::Pi54 specific primers were used for the expression analysis of GFP-Pi54 in transgenic lines Pi54OX1, 3, 4, 5, 6 in comparison to NT- control TP309. These results revealed the abundance of GFP-Pi54 transcripts in transgenic rice lines whereas no expression was found in NT-TP309 plants (Fig. 2a). The internal control gene, ß-actin was used to normalize the expression (Fig. 2b). Among the five lines, maximum expression of GFP-Pi54 transcripts were found in transgenic line Pi54OX6 and minimum expression was observed in transgenic line Pi54OX5 (Fig. 2c).
Reverse transcriptase PCR analysis of transgenic plants. (a) Using GFP:Pi54 specific primers in transgenic Pi54OX and non-transgenic (NT)-control TP 309; Lane 1: 500 bp ladder, Lane 2–6: Pi54OX 1,3,4, 5 and 6, Lane 7: NT- control TP309; Lane 8: pRTV construct, Lane 9: No template control. (b) Internal control; Actin gene amplification in transgenicPi54OX and NT-control TP 309; Lane 1: 500 bp ladder, Lanes 2–6: Pi54OX 1,3,4 5 and 6, Lane 7: NT-control TP309, Lane 8: No template control. (c) CT values during qRT- PCR using GFP::Pi54 specific primers in transgenic Pi54OX and non-transgenic (NT)-control TP 309; Lane 1: NT-control TP309; Lane 2–6: Pi54OX 1,3,4, 5 and 6.
Functional analysis of transgenic lines for rice blast resistance
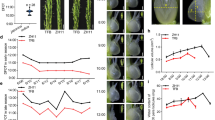
PCR, qPCR, qRT-PCR and Southern blot screened positive transgenic plants were used for functional analysis with M. oryzae infection. Highly blast susceptible rice lines HR12 and non-transgenic (NT) - control TP309 were used as controls, whereas Tetep was used as resistant control. Highly virulent M. oryzae stain; Mo-ei-79 was used for phenotyping. All the transgenic rice plants in T2 generation showed resistance response, and non-transgenic susceptible controls showed typical blast symptoms (Table S2). These results validate the Pi54 mediated resistance reaction in transgenic rice lines against M. oryzae.
Further, we also subjected T3 generation Pi54OX plants to phenotyping by challenging these rice lines with blast pathogen M. oryzae isolates MG-79 and Mo-ni-25. The disease reaction was recorded at 7 dpi. The results indicated complete blast resistance in transgenic lines Pi54OX1, 3, 5 and 6 and a hypersensitive response (HR) in transgenic line Pi54OX4 against M. oryzae strain Mo-ei-79 (Fig. 3a). However, all the five transgenic lines showed high degree of resistance against blast isolate Mo-ni-25 (Fig. 3b; Table S3).
Sub-cellular localization of Pi54 in onion and rice cells
In silico analysis using different online tools predicted cells cytoplasm and nucleus to be the predominant sites of Pi54 protein localization. These in silico observations were further validated both by transient and stable expression using GFP protein as a tag in onion and rice cells, respectively. Confocal microscopic examination of onion epidermal cells revealed that Pi54 protein is localized to cellular compartments like membrane, cytoplasm and nucleus (Fig. 4). To ensure that steric hindrance from the GFP-fusion did not interfere with Pi54 protein activity in transgenic rice lines, we tested the functionality of this protein with N-terminal fluorescent fusion in these rice lines by Immunocytochemical studies. The list of previously reported blast resistance genes with their subcellular localization is given in Table 1.
Transient expression and localization analysis of GFP::Pi54 in onion epidermal cells. Specimens were observed under Leica SP5 Confocal microscope at 488 nm excitation and 560 nm emission spectra. (a,b) GFP::Pi54 fusion protein expressing in nucleus, cytoplasmic and membrane; Scale bar: 50 µm; (c) Positive control: CaMV 35S-GFP; Scale bar: 75 µm.
Nuclear observation of GFP signals in rice calli
To further examine the nuclear localization of GFP::Pi54, we performed another set of experiment using confocal microscopy. Calli derived from mature embryo of transgenic rice seeds was used to prepare slide and observed under GFP fluorescence filter. The GFP signals were found both in the cytoplasm and also in nucleus of the cells (Fig. S6).
Membrane analysis of Pi54 protein
The membrane localization of Pi54 protein was examined in the epidermal sheath cells of transgenic rice plant. The FM4–64 FX staining technique which specifically binds to membranes4,27,28 indicated that Pi54 protein is localized to cellular membrane of rice cells (Fig. S7).
Localization analysis of Pi54 protein in different cells of leaf of transgenic rice lines
The hand cut fine transverse sections (TS) of transgenic rice leaves were observed under the confocal microscopy. These results showed GFP signals in the stomata (Fig. 5a). Further observation of the epidermal peel of the transgenic leaf tissue indicated the GFP signals in silica cells (Fig. 5b). Besides, GFP signal was also observed in upper epidermal cells, mesophyll cells, vascular bundles, walls of bundle sheath and bulliform cells (Fig. 6).
Horizontal sections of rice leaves. (a) Transgenic rice leaf showing GFP signal in Guard cells (GC), Long cells (LC) and Short cell (ShC) in epidermis; scale bar 10 µm. (b) Peeled epidermis from transgenic rice leaf showing GFP expression in dumbbell shaped Guard cells (GC) & Silica cells (SC); scale bar 10 µm. (c) Non-transgenic rice leaf showing no GFP signal in Guard cells (GC), Long cells (LC) and Short cell (ShC) in epidermis; scale bar 10 μm.
Transverse section of transgenic rice leaf observed under Leica SP5 confocal microscope at 488 nm excitation and 560 nm emission spectra. (a) transgenic plant leaf showing GFP signals in mesophyll cell (MC), upper (UE) and lower epidermis (LE), vascular bundle (VB), cell wall of bundle sheath cells (BSC) and bulliform cells (BC); Scale bar: 25 µm. (b) Non-transgenic leaf showing no GFP signal; Scale bar: 25 µm.
Immunofluorescence of GFP::Pi54 protein in rice tissues
Immunofluorescence study was performed to further validate the localization of GFP::Pi54 proteins in the Pi54OX transgenic rice leaves and confirmed the Pi54 protein localization in different parts of rice leaf tissue. Majorly, the Pi54 protein was found to be aggregated in the mesophyll cells, followed by cells in upper epidermis, lower epidermis, and walls of vascular bundle, bundle sheath and bulliform cells (Fig. 7).
Immunocytochemistry analysis of transgenic rice leaf. (a) Immunofluorescence (green fluorescence) patters obtained in mesophyll cell (MC), upper (UE) and lower epidermis (LE), vascular bundle cell wall (VB), cell wall of bundle sheath cells (BSC) and bulliform cells (BC); Scale bar: 25 µm.(b) Non-transgenic leaf; no binding immunofluorescence of primary secondary antibody; Scale bar: 25 µm.
Overexpression of Pi54 activates general defense response with Xanthomonas oryzae and Rhizoctonia solani
Upon infection with M. oryzae, Pi54 activates the downstream defense mechanism in rice. To confirm whether this mechanism also provide resistance to other pathogens of rice, the transgenic rice plants were first inoculated with avirulent strain of M. oryzae followed by inoculation with X. oryzae and R. solani, separately. The average size of lesions in transgenic lines infected with X. oryzae was significantly smaller than the control plants TP309 (Fig. S8a). Similarly, the percentage disease incidence was less than 20% in Pi54OXplants whereas it was more than 40% in NT-TP309 (Fig. S8b).
In this study, we have also observed that the transgenic Pi54OX lines offer a moderate level of resistance for sheath blight disease. The lesion length was calculated in Pi54OX and NT plant and we found the less disease severity in transgenic plant in comparison to NT plant (Fig. S9a). At 36 hpi, the lesion size recorded on transgenic plant leaves was 30% to 40% lesser than the lesion size of NT-TP309 plant (Fig. S9b). These results demonstrate the Pi54 mediated activation of general defense pathways in transgenic plants against necrotrophic pathogen R. solani and bacterial blight (BLB) pathogen X. oryzae.
Discussion
Plant resistance (genes are categorized into two sub-group on the basis of the presence of Toll-Interleukin Receptor like domain containing Nucleotide-Binding Site Leucine-Rich Repeat (TIR-NBS-LRR) or Coiled Coil domain (CC-NBS-LRR). Among the 26 blast R genes cloned and functionally characterized, eight of them (Pi37, pi21, Pid2, Pita, Pi64, Pb1, Pi54rh and Pi54of ) were examined, though by transient expression for their sub-cellular localization7,19,29,30,31,32,33,34. Plant R proteins typically have an NBS domain in the middle and LRR domains towards their carboxyl terminal end. However, among the 26-rice blast R genes, Pid2 and pi21 belong to B-lectin receptor kinase and proline containing protein, respectively (Table 1). Pi54 was the third major rice blast R gene to be cloned and functionally studied from indica rice landrace Tetep, and confers broad spectrum resistance for multiple isolates of M. oryzae across different parts of the world16,17,24,35,36,37,38. Recently, the AVR counterpart of Pi54 has also been cloned from the avirulent strain of M. oryzae39. Though attempts have been made to understand the molecular mechanism of Pi54 mediated resistance7,15, but there are no studies related to cellular localization of Pi54 gene in rice plant cells. Interestingly, there are no published reports on cellular localization of blast resistance proteins in rice cells and tissues, except a recent study which shows the localization of blast R protein; Ptr using rice protoplasts40. However, attempts were made to understand the localization of these proteins in onion epidermal cells through transient assays (Table 1). In this study, we describe the overexpression of GFP::Pi54 gene in rice for the analysis of cellular and subcellular localization of Pi54 protein in onion and rice cells through transient and stable plant expression, respectively. Taipei 309, a highly susceptible line to M. oryzae infection has been commonly used for overexpression of many blast R genes, including rice blast resistance genes Pid2, Pi54, Pi54of, Pi54rh7,17,19,31,41. Previously, this line was also used for genetic transformation of Pi54, Pi54rh and Pi54of 7,17,19,42. In this study, we used TP309 for the genetic transformation of 35 S::GFP::Pi54 gene fusion construct for overexpression and cellular localization studies. The transgenic Pi54OX rice lines were validated using PCR, Southern blot hybridization, RT-PCR and qRT-PCR for gene integration and expression analysis. Previously, Pi54 and its orthologues; Pi54rh and Pi54of were reported to show maximum expression between 72–96 hours post inoculation (hpi) with M. oryzae7,17,19. We found the abundance of GFP-Pi54 transcripts in the transgenic lines (Pi54OX) from 0 to 96 hours post inoculation (hpi). The transgenic rice lines (Pi54OX) obtained in this study were subjected to phenotypic studies in T2 and T3 generations by challenging with two highly virulent strains of M. oryzae MG-79 and Mo-ni-25. We found that Pi54 imparts complete resistance against blast pathogen similar to that observed in rice line Tetep, the donor line from which Pi54 gene was originally mapped and cloned16,17,18,41. Previously, it has already been reported that pathogen induced expression of Pi54 gene is responsible for wide-ranging resistance against various virulent isolates of blast pathogen from different parts of the world17,24,35,36,37,38,42. Also, Pi54 is responsible for the activation of complex defense response pathways in rice mediated by transcription factors like WRKY6, WRKY45, NAC6 and a battery of defense response genes18. The upregulation of these transcription factors is responsible for induced defense response in transgenic plants. The ability of Pi54 to provide moderate resistance against multiple diseases was confirmed by challenging Pi54OX transgenic lines with highly virulent strains of sheath blight and bacterial blight pathogens. The moderate level of resistance response against R. solani and X. oryzae could be attributed to the Pi54 mediated activation of complex defense response pathways leading to general resistance response. However, further research is needed to explore the importance of Pi54 mediated downstream signaling and their potential cross-talks during establishment of broad spectrum defense response for M. oryzae and moderate resistance to other rice pathogens. Priming of general resistance response using chemical agents against plant pathogens had been reported in various crops belonging to both monocot and dicot family43. Previously, Sugano et al. (2010)44 reported benzothiadiazole (BTH) priming of resistance response against M. oryzae and X. oryzae in rice. Further, over-expression of WRKY45 transcription factor (TF) could prime the resistance response in transgenic rice lines against both rice blast and bacterial blight infection, but these lines were susceptible to R. solani infection45,46. These studies together with the present analysis indicate the existence of general disease resistance mechanism which target biotrophic, hemibiotrophic and necrotrophic rice pathogens.
Cellular and subcellular localization study of resistance (R)-protein is imperative for its proper functional interpretation in plant disease resistance. In case of R proteins (NBS-LRR or NLR), cytoplasmic and nuclear localizations is very important for the hypersensitive response47,48,49,50. Generally, R proteins are localized to cytoplasm and nucleus51. Further, different NLR proteins function in different sub-cellular locations52. For instance, barley MLA10 is initially localized in the cytoplasm, however, its nuclear localization is responsible for development of hypersensitive responses (HR)26. Several plant disease resistance proteins had been found to localize to both the nucleus and the cytoplasm, even if no nuclear localization signals are found53,54. Previous reports on cellular localization of rice blast R- proteins indicate that they are localized either in cytoplasm, nucleus or cell membrane7,31. It is also known that small protein molecule (30–60 kDa) passively move from cytoplasm into nucleus55,56. Further, nuclear localization in response to pathogen infection is well documented in many R proteins54 and in some of them like, Arabidopsis RPS4 and RPS6, tobacco N, barley mildew A (MLA) 10 and rice blast resistance Pb1, it is a prerequisite to localize these protein in nucleus to provide intended resistance response57,58. Previously, the blast resistance protein Pb1 showed both cytoplasmic and nuclear localization, and localization to nucleus was necessary for resistance response49.
In the present study, both transient and stable expression in onion and rice cells, respectively, revealed that Pi54 protein being localized in the cytoplasm, nucleus and in membrane. This cytoplasmic localization of Pi54 might facilitate its interaction with Avr-Pi54 protein. However, the nuclear localization of Pi54 might be necessary for Pi54 mediated resistance response or it is simple passive transport without any functional significance as GFP::Pi54 protein is small enough (~64 kDa) to pass through nuclear pore complex. In potato, Potato virus X NLR resistance protein Rx is activated in cytoplasm and subsequently translocated to the nucleus to induce resistance response59,60. Further confocal analysis of transgenic rice leaves pre-stained with membrane specific FM4–64 stain proves the presence of this protein in cell membrane of transgenic rice leaves. These finding indicates that Pi54 protein is localized to the membrane either alone or along with other member protein of plant defensome complex. Interestingly to the best of our knowledge only maize disease resistance protein, Rp1-dp2 is known to localize both in cytoplasm and cell membrane52,61. However, there is need for further studies to know precisely how Pi54 is localized to the membrane and to identify and characterize its interacting proteins.
Silicon (Si) plays an active role in plant for disease resistance. Cereals, including rice, wheat, maize and barley are the major accumulators of silicon, amounting to more than 10% of dry weight. In naturally Si accumulating plants, Si is deposited over the epidermal layer beneath the cuticle in the form of silica cells. The development of silica cells has been studied in rice leaves62,63 and these silica cells present on the epidermal layer of the rice leaf are found to increase the resistance response of rice against M. oryzae by acting as a mechanical barrier for fungal penetration64. Further, high number of silica cells are found on the leaf blades of rice lines resistant to rice leaf folder attack65,66,67. The report says that silica cells affect the response of plant at biochemical level against the blast pathogen68. Hence, the findings of present investigation which showed localized GFP::Pi54 signals in the silica cells of transgenic rice plants indicate that Pi54 might be involved in imparting resistance response at these sites involving an unknown mechanism.
We also performed horizontal tissues sectioning and observed the localization of GFP::Pi54 under confocal microscopy. The GFP::Pi54 signals were recorded in the periphery of long cells, short cells and stomata of the epidermal layer cells. During the infection process, excessive transpiration is reported in stomata of the leaves69. Hence, the localization of Pi54 around stomata cells could explain accumulation of this protein as a defense response against M. oryzae infection. Plant pathogens interfere with the normal translocation process of plant by blocking the translocation of nutrients and water. The GFP::Pi54 signal in bulliform cells of plant epidermis which is a first line of defense indicates its probable role in providing resistance against M. oryzae infection as these cells are specialized in rolling and unrolling during water stress and leaf damage. We have also observed the GFP::Pi54 signals in the mesophyll cell and it indicates the localization of Pi54 protein in the mesophyll cells might have direct role in inhibiting M. oryzae infection in mesophyll cells, which are vital compartment of the storage tissue. The GFP dots in the mesophyll cell represent the cell-cell movement of GFP fusion protein through plasmodesmata as cell-cell movement of GFP fusion proteins need little energy during the process70. Previously, the role of Pi54 in fortification of plasmodesmata has been reported in transgenic rice expressing Pi54 gene18. Therefore, the present finding of localization of Pi54 correlates with the callose mediated fortification of plasmodesmata in Pi54OX rice lines. Plant vascular bundle (VB) is a central system of the plant which is responsible for the transportation of water, food and nutrients and M. oryzae is very well known for colonizing plant VB71. Therefore, GFP::Pi54 expression in the vascular bundle indicates the functioning of Pi54 in VB for providing resistance against M. oryzae colonization. Though, no published reports were found on localization of blast R proteins in different rice tissues, Ji et al. (2016)72 while studying the cellular localization of BPH18 have reported strong signals of GFP bound-BPH18 at the site of infection in the VB of rice leaf cells. Hence, the localization of GFP::Pi54 protein in different cells, tissues, and periphery of cells attributes the resistance response of transgenic rice plants to the expression and localization of Pi54 protein in these parts of the rice plant.
The present investigation is a first attempt to understand the cellular and subcellular localization of major rice blast R- gene, Pi54. We observed that Pi54 is largely localized in the cell cytoplasm where it might interact with its Avr- counterpart and subsequently move into nucleus where it might act on different regulatory genes. We also noticed Pi54 protein accumulation in mesophyll cells, vascular bundle cells and in plasmodesmata through which it probably moves between the cells which might activate signaling mechanism in neighboring cells. Therefore, the present study along with the existing resources will help in understanding the mechanism of Pi54 mediated resistance in a better way.
Materials and Methods
Rice seed materials and pathogen cultures
Seeds of blast susceptible japonica rice variety Taipei 309 (TP309), resistant indica rice variety Tetep and highly blast susceptible line HR12 were available at ICAR-NRCPB, New Delhi. The cultures of M. oryzae; MG-79 (Hazaribagh, Jharkhand, India) and Mo-ni-25 (Dehradun, Uttarakhand, India), Xanthomonas oryzae and Rhizoctonia solani were also available at ICAR-NRCPB, New Delhi. The oligonucleotides used are listed in the Table S4.
In silico prediction of subcellular location of Pi54 protein
Sub-cellular localization of Pi54 protein was analyzed in silico using different online software, like Multiloc2 (https://abi.inf.uni-tuebingen.de/Services/MultiLoc2), WOLFPSORT (http://www.genscript.com/wolf-psort.html), PSORT II Prediction73, MEMSAT-SV, Cell-Ploc74,75 and BacelLO76.
Development of GFP::Pi54 gene fusion construct for plant transformation
Pi54 gene was amplified using gene specific primers Pi54_F & Pi54_R from Tetep and cloned at KpnI and BamHI sites of pRT100 vector. The mGFP gene PCR amplified from pCAMBIA1302.5 was cloned N-terminal to the Pi54 at SacI site of modified pRT100 vector. For the development of rice transformation vector (pRTV), DNA fragment comprising of 35S-GFP::Pi54-nos (2.5 kb) was PCR amplified from modified pRT100 using primers CaMV35S_F & NOS_R and cloned at SmaI restriction enzyme site of modified pBSK + II vector having hptII gene. All these clones were confirmed through restriction digestion and PCR amplification and final vector was names as pRTV.
Generation of transgenic rice overexpressing GFP::Pi54 gene
Rice line TP309 was used for gene gun mediated genetic transformation using pRTV plasmid construct77. Scutellar calli derived from matured TP309 seeds were used for PDS-1000/He gene gun (Bio-Rad, USA) mediated transformed and later selected using hygromycin (50 mgL−1) containing MS media. Further, regeneration and rooting were induced using the protocol given in Table S5. After hardening, plants were grown at 16/8 h light/dark regime at National Phytotron Facility (NPF), ICAR- IARI, New Delhi.
Molecular screening of putative transgenics plants
Putative transgenic plants were analyzed by using PCR with primers specific to CaMV35S promoter, GFP::Pi54, GFP and hptII genes (Table S4). The putative PCR positive transgenic plants were further evaluated through Southern blotting using DIG labeled probe specific to hptII gene, following standard protocol78.
Expression analysis of GFP::Pi54 transcript in transgenic rice lines
Total RNA was extracted using the leaf tissue of transgenic and non-transgenic (NT) rice plants using Spectrum Plant Total RNA Kit, following standard protocol given by the manufacturer (Sigma-Aldrich, USA). Extracted total RNA was subjected to on-column DNase treatment to remove leftover DNA (Sigma-Aldrich, USA). ProtoScript First Strand cDNA Synthesis kit (NEB, USA) was for cDNA synthesis using 1 µg total RNA as the template. The expression analysis of GFP::Pi54 transcript was then performed both by RT-PCR as well as quantitative real-time PCR (qRT-PCR) approach using primers, GFP_F and Pi54_R (Table S4). The conditions for PCR amplification using Taq DNA polymerase (Vivantis, USA) were; initial denaturation (2 min at 94 °C); followed by 35 cycles of denaturation (30 s at 94 °C), annealing (30 s at 60 °C) and extension (30 s at 72 °C), and final amplification for 7 min at 72 °C. PCR fragments were resolved in agarose gel (2%; w/v) electrophoresis.
Real-Time quantitative PCR (qRT-PCR) experiment was done with three replications using cDNA prepared from total RNA of both transgenic and (NT) plants and experiment was conducted using Light cycler® SYBR green I master Kit on Light Cycler 480 II PCR system (Roche, USA) platform following standard guidelines. qRT-PCR cycling parameters were: initial denaturation for 5 min at 95 °C, followed by 50 cycles of denaturation (95 °C for 10 s), annealing (60 °C for 15 s) and extension (72 °C for 15 s).
Functional complementation assay of transgenic plants
Transgenic plants of T2 generation along with non-transgenic (NT) - control TP309, blast resistant control Tetep and susceptible line HR12 were inoculated with M. oryzae isolates Mo-ni-25 and MG-79 by following standard procedure described by Sharma et al.79. For this we have used fifteen days old rice seedlings for spray inoculated with the M. oryzae spore suspension (1 × 106 spores/ml). These plants were initially kept at 28 ±1 °C and a relative humidity of 95% under dark for 24 h at the beginning and later transferred to 16/8 h of light/dark regime with same temperature and humidity. After seven days of pathogen challenge, the disease reaction was recorded on a 0–5 disease rating scale80. Similarly, phenotyping of T3 generation of transgenic plants were also performed in NPF, ICAR-IARI, New Delhi.
Cellular localization analysis of Pi54 in onion cells
Prior to bombardment, peeled lower epidermis of onion bulb was dark incubated for 4 h in an osmotic medium [MS + 0.2 M each of mannitol and sorbitol + gelrite (3 gL−1)]. The CaMV35S-GFP and CaMV35S-GFP::Pi54 gene construct was coated separately (2.0 μg each) on gold particles (0.6 μm diameter) and bombarded into osmotically adjusted onion epidermal cells by using a gene gun (PDS-1000/He, BioRad, USA). Bombardment conditions were kept as 25 mm Hg vacuum, 1,100 psi He and target distance of 6 cm from the launch assembly. Epidermal tissue was incubated in the same medium for 24 h at 22 °C under dark and subsequently examined under confocal microscope (TCS SP5, Leica, Wetzlar, Germany) using a filter set providing 455–490 nm excitation and emission above 507 nm and the images were captured using Leica AF Lite software.
Confocal imaging of GFP in transgenic plants
Live rice plant imaging was done on a Leica SP5 laser scanning confocal microscope using a 40X C-Apochromat (NA = 1.2) water immersion objective lens. Leaf samples from both transgenic and non-transgenic plants were obtained to get transverse sections using clean double-sided razor blade, under a sliding motion to avoid crushing. Transverse as well as horizontal sections were used for imaging. The 458 and 514 nm laser lines of a 25-mW argon laser with appropriate emission filters were used for imaging GFP fluorescence.
We also performed FM4–64 staining of leaf tissue for detecting membrane localization of Pi54 protein. Leaf sheath of the transgenic plants were placed horizontally on the glass slide. These trimmed sheath segments were incubated for 2 h in 17 µM FM4–64 working solution (Invitrogen, USA) and results obtained were recorded by confocal imaging after the incubation.
Preparation of specimens for Immunocytochemistry analysis
The leaf tissues of transgenic plant were fixed in 4% paraformaldehyde (pH 7.2) using PBS solution at 65 °C for 10 min and subsequently incubated in 10 mM EDTA for 3–5 h at 65 °C. Fixed plant material was dehydrated using a series of ethanol treatment [50, 70, 90 and 100% (v/v)], infiltrated with series of polyethylene glycol (PEG) solution of 30, 50, 70% (v/v) and finally embedded in 100% PEG of 1500 grade (CDH, India). The tissue samples embedded in PEG were sectioned by using Microm HM500 (Thermo scientific Ultra microtomy), spread on frosted slides (Himedia, India). Thin sections of 10 mm were cut and collected on 1% gelrite-coated frosted slides.
Immunocytochemistry of transgenic plant tissue
Leaf sections embedded in PEG were washed with double distilled H2O for 15 min and pre-incubated in phosphate buffered saline (PBS) with 1% (w/v) BSA (pH 7.4) for 30 min, then incubated in PBS containing 1% (w/v) BSA, 0.05% (w/v) Triton-X and primary antibody (monoclonal anti-green fluorescent protein, Sigma-Aldrich diluted 1:500) for 40 min. Later slides were washed three times with PBS–Triton (0.5%) for 5 min each, then incubated for 2 h with secondary Anti-Mouse antibody (IgG-Atto 488 antibody, Sigma-Aldrich) diluted 1:100 in PBS, 1% (w/v) BSA, 0.5% (w/v) Tween-20, under dark. Slides were again rinsed with PBS–Triton-X followed with water. Processed samples were finally analyzed with Leica SP5 laser scanning microscope and observations were documented.
Analysis of transgenic plants for their reaction to Xanthomonas oryzae and Rhizoctonia solani
For bacterial blight (BLB) assay, rice plants at tillering stage were used. Initially X. oryzae culture grown on peptone sucrose agar at 28 °C for 3 days was inoculated into 250 ml of potato dextrose broth (PDB) and cultured by incubating at 28 °C under continuous shaking for 3 days. This inoculum was used for bacterial blight (BLB) assay. The uppermost 3 to 5 fully expanded rice leaves from each plant were pre inoculated with M. oryzae (Mo-ni-25) spore suspension as described in the previous section and after 48hpi they were challenged with broth cultures of X. oryzae by leaf clipping method as described by Kauffman et al.81. The disease response was recorded 14 days post inoculation (dpi) by measuring the lesion length and total leaf length (cm) of infected leaves of Pi54OX and NT-plants, based on which the mean and standard deviation were calculated. Percent disease incidence was calculated according to formula given below:
In another set of experiment, transgenic and NT- control TP309 plants were evaluated for sheath blight resistance using detached leaf bioassay as described by Richa et al.82. The R. solani culture was grown on PDA plates for 10 days to obtain mycelial mat. Detached rice leaves from three- month-old transgenic and NT- plants, previously sprayed with M. oryzae suspension were kept on the moist filter paper in a Petri-plate. Equal sized agar plugs containing mycelia was obtained from fungal plates and inoculated on leaf surface kept in wet Petri plates. The plates were then sealed by using parafilm to retain humid conditions. Phenotypic observation was noted as degree of R. solani infection at 36 hpi. Statistical analysis was performed on three replicates for each of the transgenic plant.
References
Skamnioti, P. & Gurr, S. J. Against the grain: safeguarding rice from rice blast disease. Trends Biotechnol. 27, 141–50, https://doi.org/10.1016/j.tibtech.2008.12.002 (2009).
Guo, M., Kim, P., Li, G., Elowsky, C. & Alfano, J. R. A Bacterial Effector Co-opts Calmodulin to Target the Plant Microtubule Network. Cell Host Microbe 19(1), 67–78, https://doi.org/10.1016/j.chom.2015.12.007 (2016).
Perfect, S. E. & Green, J. R. Infection structures of biotrophic and hemibiotrophic fungal plant pathogens. Mol. Plant Pathol. 2, 101–108, https://doi.org/10.1046/j.1364.3703.2001.00055.x (2001).
Kankanala, P., Czymmek, K. & Valent, B. Roles for Rice Membrane Dynamics and Plasmodesmata during Biotrophic Invasion by the Blast Fungus. Plant Cell 19, 706–724, https://doi.org/10.1105/tpc.106.046300 (2007).
Marcel, S., Sawers, R., Oakeley, E., Angliker, H. & Paszkowski, U. Tissue-Adapted Invasion Strategies of the Rice Blast Fungus Magnaporthe oryzae. Plant Cell 22, 3177–3187, https://doi.org/10.1105/tpc.110.078048 (2010).
Sharma, T. R. et al. Rice blast management through host-plant resistance: retrospect and prospects. Agric. Res. 1, 37–52 (2012).
Devanna, N. B., Vijayan, J. & Sharma, T. R. The Blast Resistance Gene Pi54of Cloned from Oryza officinalis Interacts with Avr-Pi54 through Its Novel Non-LRR Domains. PLoS One https://doi.org/10.1371/journal.pone.0104840 (2014).
Flor, H. H. Current status of the gene-for-gene concept. Annu. Rev. Phytopathol. 9, 275–296 (1971).
Jones, J. D. G. & Dangl, J. L. The plant immune system. Nature 444, 323–329 (2006).
Coll, N. S., Epple, P. & Dangl, J. L. Programmed Cell Death in plant immune system. Cell Death Differ. 18, 1247–1256 (2011).
Dean, R. A. et al. The genome sequence of the rice blast fungus Magnaporthe grisea. Nature 434, 980–986 (2005).
Nurnberger, T., Brunner, F., Kemmerling, B. & Piater, L. Innate immunity in plants and animals: striking similarities and obvious differences. Immunol. Rev. 198, 249–266 (2004).
García, A.V. & Hirt, H. Salmonella enterica induces and subverts the plant immune system. Front. Microbiol. https://doi.org/10.3389/fmicb.00141 (2014).
de Wit, P. J. G. M. How plants recognize pathogens and defend themselves. Cell Mol. Life Sci. 64, 2726–2732, https://doi.org/10.1007/s00018-007-7284-7 (2007).
Gupta, S. K., Rai. A. K., Kanwar, S. S. & Sharma, T. R. Comparative Analysis of Zinc Finger Proteins Involved in Plant Disease Resistance. PLoS One https://doi.org/10.1371/journal.pone.0042578 (2012).
Sharma, T. R. et al. High resolution mapping, cloning and molecular characterization of the Pi-kh gene of rice, which confers resistance to M. grisea. Mol. Genet. Genomics 274, 569–578 (2005).
Rai, A. K. et al. Functional complementation of rice blast resistance gene Pi-kh (Pi54) conferring resistance to diverse strains of Magnaporthe oryzae. J. Plant Biochem. Biotechnol. 20, 55–65 (2011).
Gupta, S. K. et al. The single functional blast resistance gene Pi54 activates a complex defence mechanism in rice. J. Exp. Bot. 63, 757–772, https://doi.org/10.1093/jxb/err297 (2012).
Das, A. et al. A novel blast resistance gene, Pi54rh cloned from wild species of rice, Oryza rhizomatis confers broad spectrum resistance to Magnaporthe oryzae. Funct. Integr. Genomics 12, 215–228 (2012).
Singh, A. K. et al. Marker assisted selection: a paradigm shift in Basmati breeding. Indian J. Genet. Pl. Br. 71, 1–9 (2011).
Ellur, R. K. et al. Marker-aided Incorporation of Xa38, a Novel Bacterial Blight Resistance Gene, in PB1121 and Comparison of its Resistance Spectrum with xa13 + Xa21. Sci. Rep. https://doi.org/10.1038/srep29188(2016).
Kumari, A. et al. Mining of rice blast resistance gene Pi54 shows effect of single nucleotide polymorphisms on phenotypic expression of the alleles. Eur. J. Plant Pathol. 137, 55–65, https://doi.org/10.1007/s10658-013-0216-5 (2013).
Thakur, S. et al. Extensive sequence variation in rice blast resistance gene Pi54 makes it broad spectrum in nature. Front. Plant Sci. https://doi.org/10.3389/fpls.2015.00345 (2015).
Vasudevan, K., Gruissem, W. & Bhullar, N. K. Identification of novel alleles of the rice blast resistance gene Pi54. Sci. Rep. 5, 15678, https://doi.org/10.1038/srep15678 (2015).
Berkey, R. et al. Homologues of RPW8 resistance protein are localized to the extrahaustorial membrane that is likely synthesized De Novo. Plant Physiol. 173, 600–613 (2017).
Bai, S. et al. Structure-function analysis of barley NLR immune receptor MLA10 reveals its cell compartment specific activity in cell death and disease resistance. PLoS Pathog. https://doi.org/10.1371/journal.ppat.1002752 (2012).
Nandety, R. S. et al. The Role of TIR-NBS and TIR-X Proteins in Plant Basal Defense Responses. Plant Physiol. 162, 1459–72, https://doi.org/10.1104/pp.113.219162 (2013).
Jelinkova, A. et al. Probing plant membranes with FM dyes: tracking, dragging or blocking? Plant J. 61, 883–892, https://doi.org/10.1111/j.1365-313X.2009.04102.x (2010).
Lin, F. et al. The blast resistance gene Pi37 encodes a nucleotide binding site–leucine-rich repeat protein and is a member of a resistance gene cluster on rice chromosome 1. Genetics. 177(3), 1871–1880, https://doi.org/10.1534/genetics.107.080648 (2007).
Fukuoka, S. et al. Loss of function of a proline containing protein confers durable disease resistance in rice. Science. 325, 998–1001 (2009).
Chen, X. et al. A B-lectin receptor kinase gene conferring rice blast resistance. Plant J. 46, 794–804 (2006).
Bryan, G. T. et al. A single amino acid difference distinguishes resistant and susceptible alleles of the rice blast resistance gene Pi-ta. Plant Cell. 12, 2033–2045 (2000).
Ma, J. et al. Pi64, Encoding a Novel CC-NBS-LRR Protein, Confers Resistance to Leaf and Neck Blast in Rice. Mol. Plant Microbe Interact. 28(5), 558–568 (2015).
Hayashi, N. et al. Durable panicle blast-resistance gene Pb1 encodes an atypical CC-NBS-LRR protein and was generated by acquiring a promoter through local genome duplication. Plant J. 64, 498–510, https://doi.org/10.1111/j.1365-313X.2010.04348.x (2010).
Xiao, N. et al. Improving of rice blast resistances in japonica by pyramiding major R genes. Front Plant. Sci 7, 1918, https://doi.org/10.3389/fpls.2016.01918 (2017).
Abhilash, K. V. et al. Development of gene-pyramid lines of the elite restorer line, RPHR-1005 possessing durable bacterial blight and blast resistance. Front Plant Sci 7, 1195, https://doi.org/10.3389/fpls.2016.01195 (2016).
Wu, Y. et al. Combination patterns of major R genes determine the level of resistance to the M. oryzae in Rice (Oryza sativa L.). PLoS One. 10, e0126130 (2015).
Azizi, P. et al. Over-expression of the Pikh gene with a CaMV 35S promoter leads to improved blast disease (Magnaporthe oryzae) tolerance in rice. Front Plant. Sci 7, 773, https://doi.org/10.3389/fpls.2016.00773 (2016).
Ray, S. et al. Analysis of Magnaporthe oryzae Genome Reveals a Fungal Effector, Which Is Able to Induce Resistance Response in Transgenic Rice Line Containing Resistance Gene, Pi54. Front. Plant Sci. https://doi.org/10.3389/fpls.2016.01140 (2016).
Zhao, H. et al. The rice blast resistance gene Ptr encodes an atypical protein required for broad-spectrum disease resistance. Nat. Commun. 9, 2039 (2018).
Kumari, M. et al. Co-transformation mediated stacking of blast resistance genes Pi54 and Pi54rh in rice provides broad spectrum resistance against Magnaporthe oryzae. Plant Cell Rep. 36(11), 1747–1755 (2017).
Arora, K. et al. Phenotypic expression of blast resistance gene Pi54 is not affected by its chromosomal position. Plant Cell Rep. https://doi.org/10.1007/s00299-014-1687-3 (2014).
Takatsuji H. Development of disease-resistant rice using regulatory components of induced disease resistance. Front Plant Sci. https://doi.org/10.3389/fpls.2014.00630 (2014).
Sugano, S. et al. Role of OsNPR1 in rice defense program as revealed by genome-wide expression analysis. Plant Mol. Biol. 74, 549–562, https://doi.org/10.1007/s11103-010-9695-3 (2010).
Shimono, M. et al. Rice WRKY45 plays a crucial role in benzothiadiazole-inducible blast resistance. Plant Cell 19, 2064–2076, https://doi.org/10.1105/tpc.106.046250 (2007).
Shimono, M. et al. Rice WRKY45 plays important roles in fungal and bacterial disease resistance. Mol. Plant Pathol. 13, 83–94, https://doi.org/10.1111/j.1364-3703.2011.00732.x (2012).
Engelhardt, S. et al. Relocalization of late blight resistance protein R3a to endosomal compartments is associated with effector recognition and required for the immune response. Plant Cell 12, 5142–58, https://doi.org/10.1105/tpc.112.104992 (2012).
Hoser, R. et al. Nucleocytoplasmic partitioning of tobacco N receptor is modulated by SGT1. New Phytol. 200(1), 158–71, https://doi.org/10.1111/nph.12347 (2013).
Inoue, H. et al. Blast resistance of CC-NB-LRR protein Pb1 is mediated by WRKY45 through protein–protein interaction. Proc. Natl. Acad. Sci. 110, 9577–82 (2013).
Qi, D., DeYoung, B. J. & Innes, R. W. Structure-function analysis of the coiled-coil and leucine-rich repeat domains of the RPS5 disease resistance protein. Plant Physiol. 158, 1819–32, https://doi.org/10.1104/pp.112.194035 (2012).
Marone, D., Russo, M. A., Laido, G., De Leonardis, A. M. & Mastrangelo, A. M. Plant nucleotide binding site–leucine-rich repeat (NBS-LRR) genes: active guardians in host defense responses. Int. J. Mol. Sci. 14, 7302–7326, https://doi.org/10.3390/ijms14047302 (2013).
Wang, G. F. & Balint-Kurti, P. J. Cytoplasmic and Nuclear Localizations Are Important for the Hypersensitive Response Conferred by Maize Autoactive Rp1-D21 Protein. Mol. Plant Microbe Interact. 28, 1023–31, https://doi.org/10.1094/MPMI-01-15-0014-R (2015).
Chou, K. C. & Shen, H. B. Large-scale plant protein subcellular location prediction. J. Cell Biochem. 100, 665–678, https://doi.org/10.1002/jcb.21096 (2007).
Meier, I. & Somers, D. E. Regulation of nucleocytoplasmic trafficking in plants. Curr. Opin. Plant Biol. 14, 538–546 (2011).
Rodriguez, M. S., Dargemont, C. & Stutz, F. Nuclear export of RNA. Biol. Cell 96, 639–55 (2004).
Marfori, M., Mynott, A., Ellis, J. J., Mehdi, A. M. & Saunders, N. F. Molecular basis for specificity of nuclear import and prediction of nuclear localization. Biochim. Biophys. Acta. 1813, 1562–77, https://doi.org/10.1016/j.bbamcr.2010.10.013. (2011).
Jia, Y. & Martin, R. Identification of a new locus, Ptr (t), required for rice blast resistance gene Pi-ta-mediated resistance. Mol. Plant Microbe Interact. 21(4), 396–403 (2008).
Bhattacharjee, S., Garner, C. M. & Gassmann, W. New clues in the nucleus: transcriptional reprogramming in effector-triggered immunity. Front. Plant Sci. https://doi.org/10.3389/fpls.2013.00364 (2013).
Slootweg, E. J. et al. Structural determinants at the interface of the ARC2 and leucine-rich repeat domains control the activation of the plant immune receptors Rx1 and Gpa2. Plant Physiol. https://doi.org/10.1104/pp.113.218842 (2013).
Tameling, W. I. et al. Ran GAP2 mediates nucleocytoplasmic partitioning of the NB-LRR immune receptor Rx in the Solanaceae, thereby dictating Rx function. Plant Cell. https://doi.org/10.1105/tpc.110.077461 (2010).
Wang, G. F. et al. Molecular and functional analyses of a maize autoactive NB-LRR protein identify precise structural requirements for activity. PLoS Pathog. 10, 11(4):e1004830, https://doi.org/10.1371/journal.ppat.1004830 (2015).
Luo, L., Zhou, W. Q. & Liu, P. LI, C. X., & Hou, S. W. The development of stomata and other epidermal cells on the rice leaves. Biologia Plantarum 56, 521–527 (2012).
Currie, H. A. & Perry, C. C. Silica in Plants: Biological, Biochemical and Chemical Studies. Ann. Bot. 100, 1383–1389, https://doi.org/10.1093/aob/mcm247 (2007).
Volk, R. J., Kahn, R. P. & Weintraub, R. L. Silicon content of the rice plant as a factor in influencing its resistance to infection by the rice blast fungus, Piricularia oryzae. Phytopathol. 48, 121–178 (1958).
Sudhakar, G. K., Singh, R. & Mishra, S. B. Susceptibility of rice varieties of different durations to rice leaf folder, Cnaphalocrocis medinalis Gueneee valuated under varied land situations. J. Entomol. Res. 152, 79–87 (1991).
Wang, Q. X., Xu, L. & Wu, J. C. Physical and biochemical mechanisms of resistance of different rice varieties to the rice leaf folder, Cnaphalocrocis medinalis (Lepidoptera:Pyralidae). Acta. Entomol. Sinica 51, 1265–1270 (2008).
Xu, J., Wang, Q. X. & Wu, J. C. Resistance of cultivated rice varieties to Cnaphalocrocis medinalis (Lepidoptera:Pyralidae). J. Econ. Entomol. 103, 1166–1171 (2010).
Cai, K. et al. Physiological and cytological mechanisms of silicon-induced resistance in rice against blast disease. Physiol. Plant. 134, 324–33, https://doi.org/10.1111/j.1399-3054.2008.01140.x. (2008).
Allegre, M. et al. Stomatal deregulation in Plasmopara viticola-infected grapevine leaves. New Phytol. 173, 832–40 (2007).
Verchot-Lubicz, J. Plasmodesmata transport of GFP and GFP fusions requires little energy and transitions during leaf expansion. Plant Signal. Behav. 3, 902–905 (2008).
Rodrigues, F. A., Benhamou, N., Datnoff, L. E., Jones, J. B. & Belanger, R. R. Ultrastructural and cytochemical aspects of silicon-mediated rice blast resistance. Phytopathol. 93, 535–546 (2003).
Ji, H. et al. Map-based Cloning and Characterization of the BPH18 Gene from Wild Rice Conferring Resistance to Brown Plant hopper (BPH) Insect Pest. Sci. Rep. https://doi.org/10.1038/srep34376 (2016).
Psort, I. I. PSORT: a program for detecting sorting signals in proteins and predicting their subcellular localization. J. Mol. Biol. 266, 594–600 (1997).
Chou, K. C. & Shen, H. B. Cell-PLoc: a package of web servers for predicting subcellular localization of proteins in various organisms. Nat. Protoc. 3, 153–162, https://doi.org/10.1038/nprot.2007.494 (2008).
Chou, K. C. & Shen, H.B. Plant-mPLoc: a top-down strategy to augment the power for predicting plant protein subcellular localization. Plos One https://doi.org/10.1371/journal.pone.0011335 (2010).
Pierleoni, A., Martelli, P. L., Fariselli, P. & Casadio, R. BaCelLo: a balanced subcellular localization predictor. Bioinformatics 22, 408–416, https://doi.org/10.1093/bioinformatics/btl222 (2006).
Sanford, J. C., Klein, T. M., Wolf, E. D. & Allen, N. Delivery of substances into cells and tissues using a particle bombardment process. J. Particulate Sci. Technol. 5, 27–37 (1987).
Sambrook J., Fritsch E. F., Maniatis T. A laboratory manual. Second edition. 2, 1120 (1989).
Sharma, T. R. et al. RAPD and Pathotype Analyses of Magnaporthe grisea Populations from the north-western Himalayan Region of India. J. Phytopathol. 150, 649–656 (2002).
Bonman, J. M., Vergel, T. I., De, D. & Khin, M. M. Physiological specialization of Pyricularia oryzae in the Philippines. Plant Dis 70, 767–769 (1986).
Kauffman, H. E., Reddy, A. P. K., Hsieh, S. P. Y. & Merca, S. D. An improved technique for evaluating resistance to rice varieties of Xanthomonas oryzae pv. oryzae. Plant Dis. Rep. 57, 537–541 (1973).
Richa, K. et al. Novel Chitinase Gene LOC_Os11g47510 from Indica Rice Tetep Provides Enhanced Resistance against Sheath Blight Pathogen Rhizoctonia solani in Rice. Front. Plant Sci. https://doi.org/10.3389/fpls.2017.00596 (2009).
Hayashi, K. & Yoshida, H. Refunctionalization of the ancient rice blast disease resistance gene Pit by the recruitment of a retrotransposon as a promoter. Plant. J57, 413–425 (2017).
Takahashi, A., Hayashi, N., Miyao, A. & Hirochika, H. Unique features of the rice blast resistance Pish locus revealed by large scale retrotransposon-tagging. BMC Plant Biol. 10, 175 (2010).
Wang, Z. X. et al. The Pib gene for rice blast resistance belongs to the nucleotide binding and leucine-rich repeat class of plant disease resistance genes. Plant J. 19, 55–64 (1999).
Qu, S. et al. The broad-spectrum blast resistance gene Pi9 encodes a nucleotide-binding site-leucine-rich repeat protein and is a member of a multigene family in rice. Genetics. 172, 1901–14 (2006).
Zhou, B. et al. The eight amino-acid differences within three leucine-rich repeats between Pi2 and Piz-t resistance proteins determine the resistance specificity to Magnaporthe grisea. Mol. Plant-Microbe Interact. 19, 1216–28 (2006).
Chen, J. et al. A “Pid3” allele from rice cultivar Gumei 2 confers resistance to “Magnaporthe oryzae”. J. Genet. Genom. 38, 209–16 (2011).
Liu, X., Lin, F., Wang, L. & Pan, Q. The in Silico Map-Based Cloning of Pi36, a Rice Coiled-Coil–Nucleotide-Binding Site–Leucine-Rich Repeat Gene That Confers Race-Specific Resistance to the Blast Fungus. Genetics. 176(4), 2541–2549, https://doi.org/10.1534/genetics.107.075465 (2007).
Lee, S. K. et al. Rice Pi5-mediated resistance to Magnaporthe oryzae requires the presence of two coiled-coil-nucleotide-binding-leucine-rich repeat genes. Genetics 181, 1627–1638 (2009).
Hua, L. et al. The isolation of Pi1, an allele at the Pik locus which confers broad spectrum resistance to rice blast. Theor. Appl. Genet. 125(5), 1047–55, https://doi.org/10.1007/s00122-012-1894-7 (2012).
Zhai, C. et al. The isolation and characterization of Pik, a rice blast resistance gene which emerged after rice domestication. New Phytolog. 189, 321–34 (2010).
Ashikawa, I. et al. Two adjacent nucleotide-binding site-leucine-rich repeat class genes are required to confer Pikm-specific rice blast resistance. Genetics. 180, 2267–76 (2008).
Yuan, B. et al. The Pik-p resistance to Magnaporthe oryzae in rice is mediated by a pair of closely linked CC-NBS-LRR genes. Theor. Appl. Genet. 122, 1017–28, https://doi.org/10.1007/s00122-010-1506-3 (2010).
Okuyama, Y. et al. A multifaceted genomics approach allows the isolation of the rice Pia-blast resistance gene consisting of two adjacent NBS-LRR protein genes. Plant J. 66(3), 467–79, https://doi.org/10.1111/j.1365-313X.2011.04502.x. (2011).
Tang, J. et al. Semi-dominant mutations in the CC-NB-LRR-type R gene, NLS1, lead to constitutive activation of defense responses in rice. Plant J. 66, 996–1007 (2011).
Su, J. et al. Functional divergence of duplicated genes results in a novel blast resistance gene Pi50 at the Pi2/9 locus. Theor. Appl. Genet. 128(11), 2213–25, https://doi.org/10.1007/s00122-015-2579-9 (2015).
Acknowledgements
TR Sharma is grateful to Department of Biotechnology Govt. of India for financial help and DST for JC Bose National Fellowship. We are thankful to Dr. Rithu Rai for sharing X. oryzae strains used in this study. Authors are thankful to Dr. Praveen Verma, Scientist NIPGR, New Delhi for their valuable suggestions in FM 4–64 staining. We also thank the in-charge of National Phytotron Facility IARI, New Delhi for providing facilities to maintain transgenic rice lines and conducting phenotyping experiments.
Author information
Authors and Affiliations
Contributions
Experiments were conceived and designed: T.R.S. Performed the experiments: J.S. Confocal Microscopy observation: S.K.S. and J.S., Analyzed the data: J.S., S.K.G. and B.D. Wrote the paper: T.R.S., J.S., B.D., S.K.G. and A.U.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Singh, J., Gupta, S.K., Devanna, B.N. et al. Blast resistance gene Pi54 over-expressed in rice to understand its cellular and sub-cellular localization and response to different pathogens. Sci Rep 10, 5243 (2020). https://doi.org/10.1038/s41598-020-59027-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-59027-x
This article is cited by
-
Low level of DNA variation in blast resistance Pi-ta gene of wild, weedy rice and Thai jasmine rice Khao Dawk Mali 105
Genetic Resources and Crop Evolution (2023)
-
Closer vein spacing by ectopic expression of nucleotide-binding and leucine-rich repeat proteins in rice leaves
Plant Cell Reports (2022)
-
Interactive effects of tropospheric ozone and blast disease (Magnaporthe oryzae) on different rice genotypes
Environmental Science and Pollution Research (2022)
-
Genome editing interventions to combat rice blast disease
Plant Biotechnology Reports (2022)
-
Deciphering the role of microRNAs during Pi54 gene mediated Magnaporthe oryzae resistance response in rice
Physiology and Molecular Biology of Plants (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.