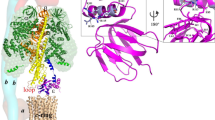
Abstract
The fatty acid synthase type II (FAS-II) multienzyme system builds the main chain of mycolic acids (MAs), important lipid pathogenicity factors of Mycobacterium tuberculosis (Mtb). Due to their original structure, the identification of the (3 R)-hydroxyacyl-ACP dehydratases, HadAB and HadBC, of Mtb FAS-II complex required in-depth work. Here, we report the discovery of a third dehydratase protein, HadDMtb (Rv0504c), whose gene is non-essential and sits upstream of cmaA2 encoding a cyclopropane synthase dedicated to keto- and methoxy-MAs. HadDMtb deletion triggered a marked change in Mtb keto-MA content and size distribution, deeply impacting the production of full-size molecules. Furthermore, abnormal MAs, likely generated from 3-hydroxylated intermediates, accumulated. These data strongly suggest that HadDMtb catalyzes the 3-hydroxyacyl dehydratation step of late FAS-II elongation cycles during keto-MA biosynthesis. Phenotyping of Mtb hadD deletion mutant revealed the influence of HadDMtb on the planktonic growth, colony morphology and biofilm structuration, as well as on low temperature tolerance. Importantly, HadDMtb has a strong impact on Mtb virulence in the mouse model of infection. The effects of the lack of HadDMtb observed both in vitro and in vivo designate this protein as a bona fide target for the development of novel anti-TB intervention strategies.
Similar content being viewed by others
Introduction
Tuberculosis (TB) is the top infectious killer worldwide1. The control of this disease has been challenged by the emergence of multidrug and extensively drug-resistant Mycobacterium tuberculosis (Mtb) strains. According to the WHO, the development of a new generation of drugs effective against these strains is urgently needed1. The very thick lipid-rich envelope of the tubercle bacillus provides a vital protective coat against the attacks of the infected host. In particular, the mycolic acid (MA)-containing lipids, which play a strategic role in the envelope architecture and permeability, are essential to the mycobacterial survival and constitute important pathogenicity factors. As a consequence, their biosynthesis pathway represents one of the Achilles’ heels of the tubercle bacillus. It is the primary target of several anti-TB antibiotics, including the first line drug isoniazid2,3. Furthermore, recently discovered small molecules, that are showing great promise as TB therapeutics, affect their metabolism4.
MAs, extremely long-chain α-alkylated β-hydroxylated fatty acids (FAs), are the major components of a highly efficient permeability barrier, the mycobacterial outer membrane (called mycomembrane)5, where they are covalently linked to the arabinogalactan layer or to polyol molecules such as trehalose6. Their production requires three distinct multienzyme systems, including the acyl carrier protein (ACP)-dependent fatty acid synthase type II (FAS-II) that is responsible for the synthesis of their main chain called ‘meromycolic chain’6. The latter carries various types of chemical function (cyclopropane ring, double bond, methyl branch and oxygenated groups) that differentiate the MA subclasses and modulate their biological activities. While typical FAS-II systems found in plants, bacteria, parasites, and mitochondria perform de novo biosynthesis7, the system from the Corynebacteriales order, which includes the Mycobacterium genus, elongates standard-size FAs (C16–C18)8. This unique property explains the success of the mycobacterial FAS-II system as a target for specific anti-TB drug therapy, illustrated by the modes of action of the drugs isoniazid, ethionamide and thiacetazone2. In mycobacteria, enzymes catalyzing the four main elongation steps have been characterized9,10,11,12,13. The last enzymes identified in Mtb FAS-II were two heterodimeric (3R)-hydroxyacyl-ACP dehydratases (HADs), HadAB and HadBC13. Belonging to the hydratase 2 family, they have an original structure for HADs of FAS-II system13,14,15, which are classically FabZ/FabA-type proteins.
In a recent survey, a combination of pull-downs, using the dehydratase HadAB of FAS-II as a bait, and proteomic analyses led to the discovery of a novel (R)-specific dehydratase, HadDMsm, of the FAS-II system from M. smegmatis, a non-tuberculous mycobacterium16. HadD displays highly specific interactions with HadAB. Bioinformatic analyses showed that, unlike HadAB, there is no HadDMsm ortholog in the genera of the Corynebacteriales order, such as Nocardia, Rhodococcus, and Gordonia, that produce medium-chain mycolic acids16. This suggested that HadDMsm might have a role during the late FAS-II elongation cycles. Consistent with this, the deletion of hadDMsm (MSMEG_0948) gene quasi-totally abolished the biosynthesis of the long chain α- and epoxy-MAs, and the resulting mutant strain produced only the medium-size α’-MAs. Thus, HadDMsm is most likely involved in building the third meromycolic segment leading to the synthesis of the full-size α- and epoxy-MAs. Importantly, hadDMsm inactivation induced an upheaval of both the bacterial cell surface and envelope properties of M. smegmatis, strongly altering the bacterial fitness and capacities to aggregate, assemble into colonies or biofilms and spread by sliding motility, critical for mycobacterial survival under stress and hostile conditions and for colonization16. It also conferred a hypersensitivity to the first line TB drug rifampicin.
Interestingly, a putative ortholog of HadDMsm was found in all of the sequenced mycobacterial genomes16, including those of the pathogen Mtb. Yet, Mtb produces a MA combination distinct from that of M. smegmatis, i.e. α-, keto- and methoxy-MAs, and has no α’- and epoxy-MAs2. In this context, we sought to elucidate the role of the putative ortholog of HadDMsm in Mtb, which in the reference strain H37Rv is represented by the yet uncharacterized protein Rv0504c. Furthermore, given the importance of HadDMsm for M. smegmatis physiology, we generated a Rv0504c knock-out Mtb mutant and examined the impact of Rv0504c inactivation on different physiological properties of Mtb in axenic cultures as well as on its virulence in the mouse model of infection.
Results
M. tuberculosis holds a putative HadD ortholog that is not essential for survival
Protein–protein BLAST searches performed against Mtb H37Rv genome17, using the MSMEG_0948 (HadDMsm) protein sequence as a probe, showed the presence of a potential ortholog of HadDMsm with a sequence identity rate of 68% in Mtb, Rv0504c, which we named HadDMtb (Fig. 1A). The latter, annotated as “conserved protein” and having a theoretical monomeric mass of 18.4 kDa17, bears similarly to HadDMsm a degenerate hydratase 2 motif ‘F-x(2)-a-x(2)-D-x(2)-P-x-H-x(5)-A’ containing the putative catalytic dyad Asp (D37) and His (H42) (Fig. 1A). The chromosomal region of hadD gene is partially conserved between Mtb and M. smegmatis (Fig. 1B). Interestingly, cmaA2 (Rv0503c) gene sits downstream of hadDMtb (Rv0504c) on Mtb chromosome and is transcribed in the same direction. It encodes the mycolic acid methyltransferase (MA-MT) CmaA2 that has a function of cyclopropane synthase and introduces a trans or cis cyclopropane at the proximal position of both keto- and methoxy-MAs18. The lack of a cmaA2 ortholog in M. smegmatis (Fig. 1B) is in agreement with the absence of these MA classes in this species2.
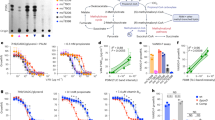
Analysis of HadDMtb sequence, hadD chromosomic region and Mtb ∆hadD mutant. (A) Sequence alignment of HadDMtb (Rv0504c) with HadDMsm (MSMEG_0948) and HadBMtb (Rv0636) proteins. Black and gray shadings indicate strictly conserved and similar residues, respectively. HadDMtb shares a sequence identity of 63% with HadDMsm (68% using BlastP alignment) and only 19% with HadBMtb (Clustal Omega scores). HadDMtb bears a degenerate hydratase 2 motif ‘F-x(2)-a-x(2)-D-x(2)-P-x-H-x(5)-A’ (uppercase: strictly conserved; lowercase: similar residue) indicated by blue stars; the putative catalytic Asp and His residues are labeled by red stars. The hydratase 2 motif ‘[YF]-x(1,2)-[LIVG]-[STGC]-G-D-x-N-P-[LIV]-H-x(5)-[AS]’ of HadBMtb13 is indicated by black stars. Alignment was performed by using Clustal Omega program, and the figure was shaped with Box shade. Database accession numbers: HadDMtb, P9WFK3 (166 aa); HadDMsm, A0QR13 (177 aa); HadBMtb, I6WYY7 (142 aa). (B) Genomic organization of hadD gene region in M. smegmatis, Mtb H37Rv and Mtb ΔhadD strains. Matching genes are drawn with identical colors. Mtb ΔhadD mutant strain was produced by an in-frame deletion of a 308 bp internal fragment (dashed lines) of hadDMtb gene (501 bp). In Mtb, Rv0502 (1,077 bp), cmaA2 (909 bp), serB1 (1,122 bp) and mmpS2 (444 bp) are annotated as encoding a conserved protein, the mycolic acid cyclopropane synthetase CmaA2, a possible phosphoserine phosphatase SerB1 and a probable conserved membrane protein MmpS2, respectively. The distinct genes in M. smegmatis, MSMEG_0946, MSMEG_0950 and MSMEG_0951, are annotated as encoding a NAD-dependent epimerase/dehydratase family protein, a hypothetical protein and the glutaredoxin 2, respectively. (C) Verification of hadDMtb gene deletion by PCR analysis. The primers (x and y; symbolized by black arrows in panel B) used for the PCR are located outside the hadDMtb gene. The genomic DNA of each strain was used as a template. HadDMtb gene length: 501 bp; ΔhadDMtb gene length: 193 bp. L: DNA ladder.
The essentiality of hadDMtb gene was examined by generating an in-frame unmarked deletion (Fig. 1B) using a two-step homologous recombination method19, so that it does not cause any polar effect on cmaA2 expression. The gene deletion was verified by PCR analysis (Fig. 1C). The viability of the resulting mutant strain showed that hadDMtb is not essential for the survival of Mtb in axenic culture, similar to the situation seen for hadDMsm in M. smegmatis16. This is consistent with the prediction of non-essentiality of hadDMtb made from a microarray-based study20, but in discrepancy with a more recent survey using high-resolution phenotypic profiling where hadDMtb was predicted to be essential21.
HadD Mtb deletion alters Mtb physiology and virulence
In the aim of evaluating the importance of HadDMtb function in the physiology of Mtb, phenotypic assays were realized. Although Mtb ΔhadD is viable, this mutant strain showed difficulties to grow in planktonic culture as compared to the wild type (wt) strain (Fig. 2A). HadDMtb inactivation also resulted in a strong alteration of the structuration of biofilms at the air-liquid interface, which appeared thinner with large clumps (Fig. 2B). There was a deep change of the colony morphology as well, with much larger, flat and spread colonies (Fig. 2C). The wt phenotype was restored in the mutant upon complementation meaning that these phenomena were linked to hadDMtb deletion. The lack of HadDMtb also conferred a high sensitivity to low temperature (Fig. 2D). In addition, Mtb ΔhadD was slightly more susceptible than Mtb wt to rifampicin, a first line antituberculous drug targeting the DNA-dependent RNA synthesis. In contrast, there was no significant difference in the sensitivity of both strains to isoniazid and ethambutol, two other firstline TB drugs, and to ciprofloxacin, a broad spectrum fluoroquinolone, as well as to the SDS detergent (Supplementary Table S1).
Inactivation of hadDMtb alters the bacterial fitness, biofilm and colony formation, and tolerance to low temperature. Comparison of Mtb H37Rv wt, ΔhadD and the complemented ΔhadD::hadDMtb strains in different phenotyping assays. All of the data are representative of at least three independent experiments. (A) Planktonic growth. Cultures were performed under shaking (120 rpm) at 37 °C in 7H9-based medium supplemented with 0.05% (w/v) Tween-80. Data are means and average deviations of three independent experiments. Some deviation bars are too small to be visible. (B) Biofilm growth at the air-liquid interface. The growth was followed for three weeks at 37 °C on 7H9-based medium. Photographs were taken after three weeks. Scale bars represent 1 cm. (C) Colony morphotype. Five μl culture aliquots were spotted on 7H11-based medium and grown for three weeks at 37 °C. Scale bars represent 0.5 cm. (D) Sensitivity to low temperature. Liquid precultures were adjusted to the same OD then serially diluted, spotted onto 7H11-based medium and incubated for four weeks at 37 °C or 30 °C.
To assess the influence of hadDMtb on the virulence level of Mtb, infection trials were performed with severe combined immunodeficiency (SCID) mice. This murine model provides a rapid and sensitive method for evaluating in vivo growth characteristics of mycobacterial strains during the acute phase of infection22,23,24,25. Although overexpressing hadDMtb gene had no significant impact on the virulence level, we observed marked and reproducible reductions of the bacterial loads in lungs (2 logs) and spleens of mice infected with the deletion mutant after 28 days of infection relative to controls (Fig. 3A). This effect correlated with a decrease of the spleen size (Fig. 3B). This phenomenon likely reflected a specific in vivo phenotype since the bacterial density of Mtb ∆hadD reached that of the wt strain after 11 days of growth in axenic cultures (Fig. 2A). The complementation observed upon transformation of the mutant strain with a wt hadDMtb copy provides explicit evidence for a link between HadDMtb function and Mtb virulence (Fig. 3A).
The deletion of hadD strongly affects Mtb virulence. The Mtb ∆hadD mutant had a reduced capacity to grow in the lungs and spleen of SCID mice. Mice were infected intravenously with 1–2 × 105 CFU of the different Mtb wt and mutant strains. (A) Bacterial loads in lungs (top panel) and spleens (bottom panel) determined after 1 (white bars) and 28 days (black bars). Data reported are mean and standard deviation of values obtained in a representative experiment of three performed each with four mice per group. (B) Photographs of the spleens from infected mice at 28 days post-infection.
In conclusion, HadDMtb plays a key role in the faculty of the tubercle bacillus to invade and multiply within the infected host. Furthermore, it is also important for the fitness and the capacities of Mtb bacilli to assemble into colonies or biofilms, and for their tolerance to low temperature. The deep changes observed in hadDMtb-deficient strain likely are the consequences of an alteration of the cell envelope composition and architecture.
HadDMtb influences the mycolic acid profile of Mtb
Mtb produces three main MA classes, α-, methoxy- and keto-MAs (Fig. 4A). To examine the potential involvement of HadDMtb protein in their biosynthesis, the MAs were extracted from Mtb ∆hadDMtb strain and analyzed. The MA content, expressed as the ratio ‘MA dry weight/delipidated bacterial residue dry weight’, was similar between the wt (10.2 ± 1.1%) and the mutant (10.4 ± 1.7%) strains. Yet, the MA distribution was changed in the deletion mutant, as shown by HPTLC analysis of the MA methyl esters (MAMEs) (Fig. 4B,C). The abundance of keto-MAs was reduced by 63%. This was compensated by an increase in the α-MA content. The recovery of the wt profile upon complementation with a wt hadDMtb copy suggested that this gene is directly involved in keto-MA biosynthesis26. The overexpression of hadDMtb in Mtb wt strain supported this conclusion, since it induced a strong increase (of 87%) in the keto-MA relative content (Fig. 4C). This had repercussions on the relative abundance of the biosynthetically affiliated methoxy-MAs, which increased slightly (Fig. 4C).
HadDMtb protein is implicated in keto-mycolic acid biosynthesis pathway. (A) Structures of the main MA classes produced by M. tuberculosis. One of the main molecules was drawn for each class, as an example. The three segments of the meromycolic chain, delimited by the distal (D) and the proximal (P) positions of the chemical functions, are indicated. (B) Mycolic acid methyl ester (MAME) profiles in HPTLC. Five µg of MAMEs from each strain were loaded onto a HPTLC plate developed five times in petroleum ether/diethyl ether, 9:1 (v/v) and stained by immersion in CuSO4 and heating. The figure is representative of six independent experiments. The hydroxy-MAs are minority MAs bearing an hydroxyl group at the distal position, and precursors of the keto- and methoxy-MAs28. (C) MA distribution in the different strains deduced from the quantification of the HPTLC band intensities as in panel B. Data are means ± average deviations of at least six independent experiments.
These results altogether clearly demonstrate that HadDMtb plays an important role in the biosynthesis of the keto-MAs. It is noteworthy that no major change in the profile of other lipids from Mtb was detected (Supplementary Fig. S1) suggesting that hadDMtb is not involved in another lipid biosynthesis pathways. Therefore, the significant alteration of MA distribution in Mtb ∆hadD is most likely responsible for its loss of virulence (Fig. 3A).
Production of full-size keto-mycolic acids requires an active HadDMtb protein
To investigate further the function of HadDMtb, the fine structures of MAs were analyzed using MALDI-TOF mass spectrometry (MS) and 1H-NMR spectroscopy. The mass spectrum of the total MA mixture from Mtb ∆hadD displayed an increase of the relative intensities of α-MAMEs signals with respect to the wt strain (Fig. 5). More importantly, the size distribution of the keto-MAs changed significantly. The signal intensities of the long chain molecules (C82-C88) markedly decreased, while those of the short-chain keto-MAs (C78, C80) raised (Fig. 5). The same observations were made on the spectrum of keto-MAs purified from the deletion mutant (Supplementary Fig. S2). The wt profiles were partially recovered in the complemented strain Mtb ΔhadD::hadDMtb (Fig. 5, Supplementary Fig. S2), confirming the role played by hadDMtb deletion in this phenotype. It is noteworthy that a reduction in the proportion of the longest molecules was also detected for the methoxy-MAs in Mtb ∆hadD (Fig. 5). Moreover, when hadDMtb was overexpressed in Mtb wt, the relative content of the long-chain keto-MAs (C82-C86) strongly rised (Fig. 5).
HadDMtb is involved in the late steps of keto-MA biosynthesis. MALDI-TOF MS spectra of the MAME mixtures from the different Mtb strains. Ion peaks are labeled with the matching MA type and the total carbon number (of the free acid form). They correspond to monosodium adducts. The spectra are representative of three independent experiments. k, keto-MA; m, methoxy-MA.
Consistent with HPTLC analyses (Fig. 4B,C), these data altogether confirm the involvement of HadDMtb in the keto-MA pathway. Furthermore, they show that HadDMtb is implicated in the late elongation cycles during the biosynthesis of the longest chain keto-meromycolic chains. This is in agreement with the lack of HadD ortholog in other genera of the Corynebacteriales order, such as Nocardia, Rhodococcus, and Gordonia, which produce medium-chain MAs16. Despite a slight difference in the cis/trans cyclopropane distribution in the keto-MAs, there was no change in the type of unsaturations carried by the MAs in Mtb ∆hadD, which remained cyclopropanes (Supplementary Table S2), whereas inactivation of cmaA2 leads to accumulation of ethylenic oxygenated MAs in Mtb18. Thus, as expected, the in-frame unmarked hadDMtb deletion performed had no visible polar effect on the expression of the adjacent cmaA2 gene (Fig. 1B) encoding CmaA2 cyclopropane synthase.
HadDMtb and HadDMsm have distinct substrate specificities
Interestingly, HPTLC analysis showed that hadDMsm gene from M. smegmatis expressed in Mtb ∆hadD could restore a MA distribution similar to that of Mtb wt strain (Fig. 4B). Furthermore, the fine structures of the MAs in Mtb ΔhadD::hadDMsm were also largely recovered, as observed by MALDI-TOF MS (Fig. 5) and NMR spectroscopy (Supplementary Table S2). This suggested that the functions of HadDMtb and HadDMsm proteins are similar. Indeed, the present work and previous data16 show that they are both involved in dehydratation steps during late FAS-II elongation cycles. Yet, unlike hadDMtb, the hadDMsm gene was not able to restore the virulence level of Mtb wt in Mtb ΔhadD strain (Fig. 3A,B). Furthermore, the reverse cross-complementation, i.e. the expression of hadDMtb in M. smegmatis ∆hadD, was far to be as successful in terms of MA profile and physiology. The MA distribution was much closer to that of M. smegmatis ∆hadD than that of M. smegmatis ∆hadD::hadDMsm (Supplementary Fig. S3A,B). The same conclusions were drawn from the colony morphology observation (Supplementary Fig. S3C) as well as from the sensitivity assays to rifampicin (Supplementary Fig. S3D), towards which M. smegmatis ∆hadD exhibits a hypersensitivity16. These data show that the functions of HadDMtb and HadDMsm are not completely superimposable. This is in perfect agreement with the specificities of HadDMsm for the α- and epoxy-MA biosynthesis16 and of HadDMtb for the keto-MA pathway. The inactivation of hadDMtb did not induce a reduction of α-MA content in Mtb as in M. smegmatis16. Therefore, although HadDMtb and HadDMsm possess similar functions, they are not strict orthologs, and the substrate specificity of HadDMtb seems important for its role in virulence.
Accumulation of dehydratation substrates triggered by hadD Mtb inactivation
It is noteworthy that Mtb ∆hadD accumulated a polar compound ‘Z’, undetectable in Mtb wt and hardly visible in Mtb ∆hadD::hadDMtb by HPTLC (Fig. 4B). This compound, appearing as a double band, likely corresponded to a heterogeneous mixture of molecules. After purification, the MALDI-TOF mass spectrum of compound Z displayed peak envelopes within a mass range from 1148 to 1362 Da (Supplementary Fig. S4A) reminiscent of the classical MAME mixture (Fig. 5), but with an addition of 16 mass units. Consistent with this, the mass increment of 84 Da (corresponding to two acetyl groups) observed after per-O-acetylation of compound Z indicated the presence of two hydroxyl groups in the intact compound (Supplementary Fig. S4A), which explained its low Rf in HPTLC (Fig. 4B). This was confirmed by 1H-NMR spectroscopy of intact compound Z generating two signals at 3.85 ppm and 3.98 ppm assigned to methines bearing both hydroxyl groups (-CHOH-). After peracetylation of compound Z, these signals shifted at 4.92 ppm and 5.13 ppm, corresponding to the chemical shifts of methines bearing O-acetyl groups (Supplementary Fig. S4B). The spin system in the 1H-1H 2D NMR COSY spectrum of per-O-acetylated compound Z revealed that, in the intact compound, one hydroxyl group is carried by the C-3 like in the classical MAs, whereas the additional hydroxyl group is located on the C-5 (Supplementary Fig. S4C,D). These data altogether clearly showed that compound Z is a mixture of 5-hydroxylated MAs. These molecules are mostly cis-cyclopropanated like regular Mtb MAs (Supplementary Table S2).
The additional hydroxyl group on the C-5 in compound Z was located on the C-3 in the precursor meromycolic chains, before the mycolic condensation step (Supplementary Fig. S5). The 3-hydroxylated meromycolic acids were taken over by the mycolic condensation system leading to the synthesis of abnormal 5-hydroxylated MAs. The accumulation of 3-hydroxylated intermediates corresponding to dehydratase substrates strongly suggests that, in the absence of HadDMtb protein, the dehydratation step of the late FAS-II elongation cycles is partially blocked, preventing the formation of mature meromycolic chains (Supplementary Fig. S5). The 3-hydroxylated α-meromycolic chains might come from the group of diethylenic precursors of oxygenated MAs (Fig. 6), which, accumulating abnormally, would later be cyclopropanated by default like the regular α-meromycolic chains.
Proposed function of HadDMtb in the mycolic acid biosynthetic pathway of M. tuberculosis. FAS-II first catalyzes several series of early elongation cycles involving HadAB dehydratase, and punctuated by the introduction of two double bonds potentially leading to the biosynthesis of diethylenic precursors. These double bonds are modified into the appropriate chemical functions (cis/trans cyclopropane, methoxy, keto) by dedicated MA-methyltransferases that differentiate the pathways of the three main MA classes. The Claisen-type condensation of the resulting dicyclopropyl meromycoloyl chains with a carboxy-fatty acyl-CoA by the mycolic condensation system generates the trehalose α-MA esters. The formation of the main methoxy- and keto-MAs, whose meromycolic chain is longer, requires an additional set of FAS-II late elongation cycles, which specifically involve HadBC and HadDMtb dehydratases during the methoxy-MA and keto-MA biosyntheses, respectively. HadBC and HadDMtb functions would partially overlap. The corresponding mature MAs are then generated by the mycolic condensation system. The total carbon number of the meromycolic and MA chains are indicated. AcpM, mycobacterial FAS-II acyl carrier protein.
In conclusion, HadDMtb most likely catalyzes the 3-hydroxyacyl-ACP dehydratation step of the late elongation cycles during the biosynthesis of the oxygenated meromycolic chains.
Discussion
The present study reports the existence in Mtb of a putative ortholog, which we named HadDMtb, of the recently discovered (3R)-hydroxyacyl-ACP dehydratase HadDMsm of the FAS-II system from M. smegmatis16. The in-depth analysis of the distribution and fine structure of the MAs produced by Mtb hadD deletion mutant revealed that HadDMtb is involved in the oxygenated MA biosynthesis, and more particularly dedicated to the keto-MA pathway. Indeed, in the mutant, the overall keto-MA content was strongly reduced. The fact that the relative abundance of the longest keto-MAs decreased in Mtb ∆hadDMtb whereas it increased in Mtb wt::hadDMtb overexpression strain shows that HadDMtb is involved in late FAS-II elongation cycles during the biosynthesis of the keto-meromycolic chains. This conclusion is supported by the lack of HadD ortholog in the genera of the Corynebacteriales order that produce only intermediate size and not full size mycolic acids as found in mycobacteria16. The abnormal 5-hydroxylated MAs observed in HadDMtb deficient strain result from the condensation of 3-hydroxy-meromycoloyl chains with a carboxyacyl chain (Supplementary Fig. S5). Their accumulation together with the belonging of HadDMtb to the hydratase 2 protein family and its high sequence similarity with HadDMsm leads to the conclusion that HadDMtb most likely catalyzes the 3-hydroxyacyl-ACP dehydratation step of these FAS-II elongation cycles (Fig. 6). We had previously discovered two (3 R)-hydroxyacyl-ACP dehydratases of the FAS-II system from M. tuberculosis, HadAB and HadBC13. Thus, HadDMtb constitutes a third dehydratase of this system. It has been shown that HadAB is involved in the early elongation cycles common to both α and oxygenated MA pathways, while HadBC is required for the late cycles leading to the biosynthesis of the sole oxygenated MAs, which are 4–6 carbon longer than α-MAs13,27. Since the keto- and methoxy-MAs are biosynthetically affiliated28, the MA distribution in Mtb ∆hadC strain was interpreted in terms of α/oxygenated ratio27. Yet, it is noteworthy that, in this mutant, the keto-MA content remains stable whereas that of the methoxy-MAs drops dramatically27. In the light of our new findings, this strongly suggests that HadBC is preferentially dedicated to the methoxy-MA biosynthesis and HadDMtb to the keto-MA pathway. However, the functions of both enzymes are likely partially redundant since their individual inactivation does not totally inhibit the production of methoxy- or keto-MAs. This is supported by the variations in methoxy-MA content and size distribution observed in Mtb wt::hadDMtb and in Mtb ∆hadDMtb, respectively (Figs. 4 and 5), and in keto-MA fine structure in Mtb ΔhadC strain27, which may also be partly due to a tight regulation between both biosynthesis pathways. These surveys allow us to draw a biosynthesis scheme for the three MA classes from Mtb, which details the specific roles of the three FAS-II dehydratases (Fig. 6).
The functions of HadDMtb and HadDMsm in M. smegmatis appear closely related since they both catalyze the dehydratation step during late elongation cycles16. Yet, the protein from Mtb has a substrate specificity distinct from that of M. smegmatis, where it is required for the biosynthesis of α- and epoxy-MAs16. The phenotypic analyses of the cross-complemented M. smegmatis ∆hadD mutant in terms of MA profile, colony morphology and rifampicin sensitivity, and of the cross-complemented Mtb ∆hadD mutant in terms of virulence level confirmed this conclusion and definitely showed that hadDMtb and hadDMsm are not strict orthologs, despite the partial genomic conservation of their chromosomal regions. This suggests that there has been a functional divergence of both genes after the speciation event. Indeed, during its adaption to a distinct environment, the ortholog in a new species may undergo neofunctionalization, resulting in a species-specific function for this gene29.
We showed that Rv0504c gene encoding HadDMtb is not essential for the survival of the tubercle bacillus in axenic culture. However, we observed that it has an important impact on the fitness of bacteria, their organization into biofilms and colonies, as well as their tolerance to low temperature (30 °C). This is certainly linked to the determinant role of MAs in the architecture and the fluidity of the envelope due to their strategic location within the mycomembrane2,30. In agreement with our data, the growth of an Mtb strain overproducting mmaA3 MA-methyltransferase gene and lacking keto-MAs was impaired at reduced temperature (32 °C)31. MAs are also known to be important for the formation of mycobacterial biofilms, where the extracellular matrix contains large amounts of free MAs32. In particular, the production of keto-MAs in which HadDMtb is involved is essential for biofilm growth33. MAs represent also key Mtb pathogenicity factors, their fine structures strongly potentiating the immune response to infection2. Here, we show that HadDMtb function greatly influences the virulence level of Mtb in the mouse model of infection. This is consistent with previous findings showing that a Mtb hma (mmaA4) mutant devoid of oxygenated MAs is attenuated in mice34 and that keto-MAs are critical for Mtb growth within the natural host cells31. Additionally, oxygenated MAs play a role in the selective repression of macrophage IL-12p40 cytokine production aimed to evade elimination by the host immune system35, and the hma gene required for their biosynthesis is actively expressed during human pulmonary tuberculosis36. Importantly, the recently approved drug delamanid, active against MDR-TB, kills Mtb by blocking the oxygenated MA production37. Altogether, these data indicate that targeting this metabolism would constitute a relevant strategy for the development of new therapeutics against Mtb. Because of its involvement in the keto-MA biosynthesis pathway, its role in Mtb virulence and its specificity to mycobacterial cells16, HadD protein represents a promising pharmaceutical target. Besides, targeting non-essential enzymes of the tubercle bacillus should limit the occurrence of antibiotic resistance mechanisms.
Methods
Bio-containment measures
All of the experiments using Mtb living strains were performed either in a regular Biosafety Level-3 (BSL-3) laboratory or in an Animal Biosafety Level-3 (ABSL-3) laboratory in the case of mouse infection experiments.
Protein sequence analyses
Sequence alignments were performed using BLAST and Clustal Omega softwares38,39, with default parameters. The presence of a putative HadDMsm ortholog in Mtb H37Rv was analyzed by BlastP searches against the fully sequenced genome available at NCBI website (www.ncbi.nlm.nih.gov/genomes/lproks.cgi), using MSMEG_0948 protein sequence (database accession number: A0QR13) as a probe. Genomic organization of hadDMsm and hadDMtb gene regions in M. smegmatis mc2155 and Mtb H37Rv, respectively, was visualized at Mycobrowser website (https://mycobrowser.epfl.ch/)17.
Construction of the deletion mutant and complemented strains
The unmarked deletion of Rv0504c (hadDMtb) gene in Mtb H37Rv (ATCC 27294) strain was done using a previously described method19. Briefly, the Rv0504c deletion delivery vector was constructed by amplifying the upstream (0.95 kb) and downstream (1.1 kb) regions of the gene using the primer pairs EUF24 (5′-GCTGCAGCTTCTTCGACGTGGACAACA-3′) and EUR24 (5′- CAAGCTTGCCGATCAGTGTCTGGGCTTCT-3′) and EUF25 (5′-CAAGCTTGCCGAGATCCGAAGCGAAGTTA-3′) and EUR25 (5′- CGGTACCTTTCGGCTGACCCTTATTG-3′), respectively. PCR-amplified fragments were cloned into p2NIL19 using the restriction sites in the primers (PstI, HindIII and Kpn I, underlined in the above sequences) and the sequence was verified. The PacI cassette containing lacZ, sacB, and hyg from pGOAL1919 was introduced to construct the final vector pTACK0504G. The plasmid was electroporated into Mtb and single crossovers were isolated. Double crossovers were isolated from the single crossover strain as previously described19. Colonies were screened for the presence of the wild-type (wt) or deletion alleles by PCR using primers 0504D1 (5′-AGCCTCTAGACGCCAATCAC-3′) and 0504D2 (5′-GGCTCAAGGTTCAGCTTGTC-3′). The deletion was checked by PCR and sequencing. For complementation of Mtb ∆hadD strain, the wt copy of Rv0504c gene, containing its natural promoter (186 bp before the start codon), was amplified with the primer pairs 0504C1 (5′-AGCCTCTAGACGCCAATCAC-3′) and 0504C2 (5′-CAAGCTTCGTCATTGAACGGACCCTAC-3′) that incorporates a HindIII restriction site and cloned into pSC-A (Agilent). The HindIII fragment from pUC-GM-Int containing the mycobacteriophage L5 integrase, att site, and Gm resistance40 was inserted to make the complementing vector. The resulting plasmid pUC-hadDMtb and the empty plasmid pUC (as a control) were used to transform both Mtb wt and Mtb ∆hadD strains (Supplementary Table S3). For cross-complementation experiments, pUC-hadDMsm plasmid carrying a wt copy of MSMEG_0948 (hadDMsm) gene16 was used to transform Mtb ∆hadD and pUC-hadDMtb plasmid was used to transform M. smegmatis ΔhadD strain previously constructed16 (Supplementary Table S3). Data were compared to those obtained for M. smegmatis mc2155 (wt) strain as well as M. smegmatis ΔhadD complemented by pUC-hadDMsm and previously described16.
Culture conditions and phenotyping assays
For biofilm growth and lipid analyses, Mtb strains were cultured as pellicles in glass bottles for 4–5 weeks at 37 °C in 7H9 broth (Difco) containing 0.2% glycerol, 10% Middlebrook ADC (Difco) and 10 µg/ml gentamycin. Cultures were inoculated with identical volumes of precultures in exponential growth phase grown in the same medium then adjusted at OD600 ~ 1. For the growth curves, planktonic cultures were realized under shaking (120 rpm) in the above medium supplemented with 0.05% (w/v) Tween-80, and OD600 was measured at different time points. For colony morphology and susceptibility to temperature assays on Mtb strains, liquid precultures were done in the latter medium (with Tween-80) then adjusted to the same OD and serially diluted. Five µl aliquots of each dilution were spotted on Middlebrook 7H11 medium (Sigma) supplemented with 0.5% glycerol, 10% Middlebrook OADC (Difco) and 10 µg/ml gentamycin. Cultures were incubated for at least 3 weeks at 37 °C or at 30 °C (for temperature testing). The minimum inhibitory concentrations for several antibiotics and SDS were determined in 7H9 broth supplemented with glycerol by using a colorimetric microassay based on the reduction of 3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (MTT, Sigma-Aldrich) into formazan by metabolically active cells, as described41. Colony morphology and rifampicin susceptibility assays on M. smegmatis strains were performed as previously described16.
Lipid extractions, (HP)TLC analyses and purification
The MAs and the total extractable lipids were extracted and analyzed as described previously16, except that the (HP)TLC plates for MAME separation were developed in ether/diethyl ether (9:1, v/v, five runs). Purification of MAMEs and compound Z was performed by preparative TLC using silica gel 60 plates (Merck) developed in the above eluent; the compounds were then scraped off and extracted from silica gel three times with diethyl ether.
Lipid structural analyses
Peracetylation of compound Z was performed in pyridine:acetic anhydride 1:1 for 1 h at 100 °C. After drying, three extractions with H2O:diethyl ether (1:1) were done; the ether phases were collected, washed three times with water and dried. MALDI-TOF MS analyses were performed in the positive ionization and reflectron mode, using the 5800 MALDI-TOF/TOF Analyzer (Applied Biosystems/ABsciex) equipped with a Nd:YAG laser (349 nm wavelength) as described previously16. 1H-NMR and 1H-1H-NMR COSY spectra were recorded in CDCl3 at 298° K using a 600-MHz Bruker Avance III spectrometer (Bruker Biospin) equipped with a TCI cryoprobe. Chemical shift values were referenced to CHCl3 resonance (δH 7.26 ppm). The quantification of MAME unsaturations was performed with TopSpin 3.5 pl7 software.
Mouse infection experiments
Virulence studies were performed in SCID mice. Briefly, groups of 6 week old C.B-17/Icr SCID mice (Charles River) were intravenously infected with a bacterial suspension containing 1–2 × 105 CFU/mouse. One day and 28 days after infection, mice were euthanized, and spleen and lungs were homogenized in 2 ml tubes containing 500 µl Sauton medium and 2.5 mm diameter glass beads using an MM300 apparatus (Qiagen). CFU numbers in target organs were determined by plating 5- or 10-fold serial dilutions of organ homogenates on solid medium and incubation at 37 °C. These studies were approved by the Institut Pasteur Safety Committee (Protocol 11.245; experimentation authorization number 75–1469), in accordance with European and French guidelines (Directive 86/609/CEE and Decree 87–848 of 19 October 1987), and implicating approval from local ethical committees (CETEA 2013–0036 and CETEA dab180023).
Data availability
All data generated or analysed during this study are included in this published article (and its Supplementary Information files).
References
WHO. Global tuberculosis report 2019. WHO Press edn, (World Health Organization, 2019).
Daffé, M., Quémard, A. & Marrakchi, H. Mycolic acids: from Chemistry to Biology in Biogenesis of Fatty Acids, Lipids and Membranes (ed Geiger, O.) 181–216, Handbook of Hydrocarbon and Lipid Microbiology series (Springer, Cham, 2019).
Quémard, A. et al. Binding of catalase-peroxidase-activated isoniazid to wild-type and mutant Mycobacterium tuberculosis enoyl-ACP reductases. J. Am. Chem. Soc. 118, 1561–1562 (1996).
North, E. J., Jackson, M. & Lee, R. E. New Approaches to Target the Mycolic Acid Biosynthesis Pathway for the Development of Tuberculosis Therapeutics. Curr. Pharm. Des 20, 4357–4378 (2014).
Zuber, B. et al. Direct visualization of the outer membrane of mycobacteria and corynebacteria in their native state. J. Bacteriol. 190, 5672–5680 (2008).
Quémard, A. New Insights into the Mycolate-Containing Compound Biosynthesis and Transport in Mycobacteria. Trends Microbiol. 24, 725–738, https://doi.org/10.1016/j.tim.2016.04.009 (2016).
Rock, C. O. & Cronan, J. E. Escherichia coli as a model for the regulation of dissociable (type II) fatty acid biosynthesis. Biochim. Biophys. Acta 1302, 1–16 (1996).
Bloch, K. Control mechanisms for fatty acid synthesis in Mycobacterium smegmatis. Adv. Enzymol. Relat. Areas Mol. Biol 45, 1–84 (1977).
Schaeffer, M. L. et al. Purification and biochemical characterization of the Mycobacterium tuberculosis beta-ketoacyl-acyl carrier protein synthases KasA and KasB. J. Biol. Chem. 276, 47029–47037 (2001).
Kremer, L. et al. Mycolic acid biosynthesis and enzymic characterization of the beta-ketoacyl-ACP synthase A-condensing enzyme from Mycobacterium tuberculosis. Biochem. J 364, 423–430 (2002).
Marrakchi, H. et al. MabA (FabG1), a Mycobacterium tuberculosis protein involved in the long-chain fatty acid elongation system FAS-II. Microbiology 148, 951–960 (2002).
Quémard, A. et al. Enzymatic characterization of the target for isoniazid in Mycobacterium tuberculosis. Biochemistry 34, 8235–8241 (1995).
Sacco, E. et al. The missing piece of the type II fatty acid synthase system from Mycobacterium tuberculosis. Proc. Natl. Acad. Sci. USA 104, 14628–14633, https://doi.org/10.1073/pnas.0704132104 (2007).
Biswas, R. et al. Crystal structure of dehydratase component HadAB complex of mycobacterial FAS-II pathway. Biochem. Biophys. Res. Commun. 458, 369–374, https://doi.org/10.1016/j.bbrc.2015.01.119 (2015).
Dong, Y. et al. Molecular basis for the inhibition of beta-hydroxyacyl-ACP dehydratase HadAB complex from Mycobacterium tuberculosis by flavonoid inhibitors. Protein Cell 6, 504–517, https://doi.org/10.1007/s13238-015-0181-1 (2015).
Lefebvre, C. et al. HadD, a novel fatty acid synthase type II protein, is essential for alpha- and epoxy-mycolic acid biosynthesis and mycobacterial fitness. Sci. Rep 8, 6034, https://doi.org/10.1038/s41598-018-24380-5 (2018).
Kapopoulou, A., Lew, J. M. & Cole, S. T. The MycoBrowser portal: a comprehensive and manually annotated resource for mycobacterial genomes. Tuberculosis 91, 8–13, https://doi.org/10.1016/j.tube.2010.09.006 (2011).
Barkan, D., Rao, V., Sukenick, G. D. & Glickman, M. S. Redundant function of cmaA2 and mmaA2 in Mycobacterium tuberculosis cis cyclopropanation of oxygenated mycolates. J. Bacteriol. 192, 3661–3668, https://doi.org/10.1128/JB.00312-10 (2010).
Parish, T. & Stoker, N. G. Use of a flexible cassette method to generate a double unmarked Mycobacterium tuberculosis tlyA plcABC mutant by gene replacement. Microbiology 146(Pt 8), 1969–1975, https://doi.org/10.1099/00221287-146-8-1969 (2000).
Sassetti, C. M., Boyd, D. H. & Rubin, E. J. Genes required for mycobacterial growth defined by high density mutagenesis. Mol. Microbiol 48, 77–84 (2003).
Griffin, J. E. et al. High-resolution phenotypic profiling defines genes essential for mycobacterial growth and cholesterol catabolism. PLoS Pathog. 7, e1002251, https://doi.org/10.1371/journal.ppat.1002251 (2011).
Pym, A. S., Brodin, P., Brosch, R., Huerre, M. & Cole, S. T. Loss of RD1 contributed to the attenuation of the live tuberculosis vaccines Mycobacterium bovis BCG and Mycobacterium microti. Mol. Microbiol 46, 709–717 (2002).
Bottai, D. et al. ESAT-6 secretion-independent impact of ESX-1 genes espF and espG1 on virulence of Mycobacterium tuberculosis. J. Infect. Dis 203, 1155–1164, https://doi.org/10.1093/infdis/jiq089 (2011).
Hsu, T. et al. The primary mechanism of attenuation of bacillus Calmette-Guerin is a loss of secreted lytic function required for invasion of lung interstitial tissue. Proc. Natl. Acad. Sci. USA 100, 12420–12425, https://doi.org/10.1073/pnas.1635213100 (2003).
Frigui, W. et al. Control of M. tuberculosis ESAT-6 secretion and specific T cell recognition by PhoP. PLoS Pathog. 4, e33, https://doi.org/10.1371/journal.ppat.0040033 (2008).
Glickman, M. S., Cahill, S. M. & Jacobs, W. R. Jr. The Mycobacterium tuberculosis cmaA2 gene encodes a mycolic acid trans-cyclopropane synthetase. J. Biol. Chem. 276, 2228–2233, https://doi.org/10.1074/jbc.C000652200 (2001).
Slama, N. et al. The changes in mycolic acid structures caused by hadC mutation have a dramatic effect on the virulence of Mycobacterium tuberculosis. Mol. Microbiol. 99, 794–807, https://doi.org/10.1111/mmi.13266 (2016).
Dubnau, E. et al. Mycobacterium bovis BCG genes involved in the biosynthesis of cyclopropyl keto- and hydroxy-mycolic acids. Mol. Microbiol. 23, 313–322 (1997).
Das, M. et al. Expression pattern similarities support the prediction of orthologs retaining common functions after gene duplication events. Plant Physiol 171, 2343–2357, https://doi.org/10.1104/pp.15.01207 (2016).
Liu, J., Barry, C. E. III, Besra, G. S. & Nikaido, H. Mycolic acid structure determines the fluidity of the mycobacterial cell wall. J. Biol. Chem. 271, 29545–29551 (1996).
Yuan, Y., Zhu, Y., Crane, D. D. & Barry, C. E. III The effect of oxygenated mycolic acid composition on cell wall function and macrophage growth in Mycobacterium tuberculosis. Mol. Microbiol. 29, 1449–1458 (1998).
Ojha, A. K. et al. Growth of Mycobacterium tuberculosis biofilms containing free mycolic acids and harbouring drug-tolerant bacteria. Mol. Microbiol 69, 164–174, https://doi.org/10.1111/j.1365-2958.2008.06274.x (2008).
Sambandan, D. et al. Keto-mycolic acid-dependent pellicle formation confers tolerance to drug-sensitive Mycobacterium tuberculosis. Mbio. 4, e00222–00213, https://doi.org/10.1128/mBio.00222-13 (2013).
Dubnau, E. et al. Oxygenated mycolic acids are necessary for virulence of Mycobacterium tuberculosis in mice. Mol. Microbiol 36, 630–637 (2000).
Dao, D. N. et al. Mycolic acid modification by the mmaA4 gene of M. tuberculosis modulates IL-12 production. PLoS Pathog. 4, e1000081, https://doi.org/10.1371/journal.ppat.1000081 (2008).
Rachman, H. et al. Unique transcriptome signature of Mycobacterium tuberculosis in pulmonary tuberculosis. Infect. Immun. 74, 1233–1242, https://doi.org/10.1128/IAI.74.2.1233-1242.2006 (2006).
Liu, Y. et al. Delamanid: From discovery to its use for pulmonary multidrug-resistant tuberculosis (MDR-TB). Tuberculosis 111, 20–30, https://doi.org/10.1016/j.tube.2018.04.008 (2018).
Sievers, F. et al. Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol. Syst. Biol. 7, 539, https://doi.org/10.1038/msb.2011.75 (2011).
Altschul, S. F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic. Acids. Res 25, 3389–3402 (1997).
Mahenthiralingam, E. et al. Site-directed mutagenesis of the 19-kilodalton lipoprotein antigen reveals no essential role for the protein in the growth and virulence of Mycobacterium intracellulare. Infect. Immun 66, 3626–3634 (1998).
Vaubourgeix, J. et al. S-adenosyl-N-decyl-aminoethyl, a potent bisubstrate inhibitor of Mycobacterium tuberculosis mycolic acid methyltransferases. J. Biol. Chem. 284, 19321–19330, https://doi.org/10.1074/jbc.M809599200 (2009).
Acknowledgements
We are grateful to L. Spina (IPBS, Toulouse) for technical help and to F. Bardou (IPBS, Toulouse) for fruitful discussions. The NMR data were collected on the PICT-IbiSA facility; NMR experiments were performed on the PICT-Genotoul platform of Toulouse funded by CNRS, Université de Toulouse-UPS, Ibisa, European structural funds and the Midi-Pyrénées region. We acknowledge funding from the Agence Nationale de la Recherche (FASMY, grant ANR-14-CE16-0012, as well as ANR-10-LABX-62-IBEID, ANR-16-CE35-0009, and ANR-16-CE15-0003) and MSDAVENIR (FIGHT-TB: Unconventional Strategies for Tuberculosis Treatment).
Author information
Authors and Affiliations
Contributions
C.L., N.S., F.L.-V., M.Da and A.Q. designed and analyzed data of lipid structural analyses; W.F., R.B. and A.Q. designed and analyzed data of mouse infection experiments; C.L., P.C., M.Da and A.Q. designed and analyzed data of the other experiments. C.L., N.S., F.L.-V. and P.C. performed lipid extractions and structural analyses; W.F. performed mouse infection experiments; A.Q. performed protein sequence analyses; C.L. and P.C. performed the other experiments. F.L.-V., P.C., A.L., T.P. and N.E. contributed to experimental or analytical tools. A.Q. wrote the manuscript main text. C.L., R.B. and A.Q. wrote the Methods. W.F. and A.Q. prepared Fig. 3, and C.L. and A.Q. prepared the other figures and tables.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Lefebvre, C., Frigui, W., Slama, N. et al. Discovery of a novel dehydratase of the fatty acid synthase type II critical for ketomycolic acid biosynthesis and virulence of Mycobacterium tuberculosis. Sci Rep 10, 2112 (2020). https://doi.org/10.1038/s41598-020-58967-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-58967-8
This article is cited by
-
Lsr2 acts as a cyclic di-GMP receptor that promotes keto-mycolic acid synthesis and biofilm formation in mycobacteria
Nature Communications (2024)
-
Transcriptional portrait of M. bovis BCG during biofilm production shows genes differentially expressed during intercellular aggregation and substrate attachment
Scientific Reports (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.