Abstract
In the Drosophila ovary, somatic escort cells (ECs) form a niche that promotes differentiation of germline stem cell (GSC) progeny. The piRNA (Piwi-interacting RNA) pathway, which represses transposable elements (TEs), is required in ECs to prevent the accumulation of undifferentiated germ cells (germline tumor phenotype). The soma-specific piRNA cluster flamenco (flam) produces a substantial part of somatic piRNAs. Here, we characterized the biological effects of somatic TE activation on germ cell differentiation in flam mutants. We revealed that the choice between normal and tumorous phenotypes of flam mutant ovaries depends on the number of persisting ECs, which is determined at the larval stage. Accordingly, we found much more frequent DNA breaks in somatic cells of flam larval ovaries than in adult ECs. The absence of Chk2 or ATM checkpoint kinases dramatically enhanced oogenesis defects of flam mutants, in contrast to the germline TE-induced defects that are known to be mostly suppressed by сhk2 mutation. These results demonstrate a crucial role of checkpoint kinases in protecting niche cells against deleterious TE activation and suggest substantial differences between DNA damage responses in ovarian somatic and germ cells.
Similar content being viewed by others
Introduction
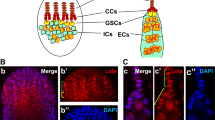
Many fundamental questions concerning the mechanisms of self-renewal and differentiation of stem cells are addressed using Drosophila oogenesis as a model1. Drosophila ovaries consist of ovarioles, chains of egg chambers connected to the germarium, which houses germline stem cells (GSCs). A microenvironment of somatic cells known as a niche regulates GSC state via different cell signaling pathways1,2,3. The ovarian niche includes terminal filament (TF) cells, cap cells (CCs), and escort cells (ECs). GSCs directly contact CCs and the most anterior ECs, which prevent GSC differentiation by secreting decapentaplegic (Dpp) and glass bottom boat (Gbb) protein ligands4,5,6,7. These ligands interact with GSC surface receptors and activate BMP signaling, which represses transcription of the bam gene required for GSC differentiation. After GSC division, one of the daughter cells retains its stem state, whereas the other one leaves the self-renewal niche and begins to differentiate into a cystoblast, which then divides and differentiates to form a cyst of germ cells surrounded by somatic follicle cells. A special marker of GSCs and cystoblasts is the spectrosome, a cytoplasmic body, which transforms into a branching structure called the fusome connecting the dividing germ cells. To initiate the differentiation of the cystoblast, BMP signaling must be decreased by different intrinsic and extrinsic mechanisms8. The majority of ECs limit the spreading of BMP ligands and therefore promote differentiation of the cystoblasts and dividing cysts9,10. Thus, the renewal somatic niche provides maintenance signals for GSCs, while a more posteriorly located differentiation niche, represented by ECs, is required for proper differentiation of GSC progeny.
The piRNA (Piwi-interacting RNA) pathway controls expression of transposable elements (TEs) in both somatic and germ cells of Drosophila ovaries. Piwi proteins guided by small piRNAs (24–30 nt) recognize complementary RNA molecules leading to their degradation or the repression of transcription with the help of other proteins (for review see11). The known molecular function of the piRNA pathway in the ovarian soma is the repression of a specific group of somatically active LTR retrotransposons12,13,14,15,16. The piRNA machinery in Drosophila ovarian somatic cells seems to be simpler than its counterpart in the germline. It operates via a single Piwi protein unlike the three proteins in germ cells and a substantial part of somatic piRNAs originates from a single source, the piRNA cluster flamenco (flam)14,15,17 that is an extended 180 kb region of X-chromosome heterochromatin, filled by TE copies and their fragments18,19,20. The flam locus is responsible for the repression of at least three somatically expressed retrotransposons: gypsy, ZAM and Idefix21,22,23,24. Cleavage of flam transcripts into small RNA molecules occurs in cytoplasmic Yb bodies. The cytoplasmic piRNA biogenesis machinery in somatic cells includes the nuclease Zucchini (Zuc), the RNA helicase Armitage (Armi), the TUDOR domain-containing proteins fs(1)Yb (Yb) and Vreteno (Vret), and other components16,25,26,27. In the course of flam transcript cleavage, piRNAs are loaded into Piwi and then move into the nucleus, where mature piRNA-Piwi complexes recognize complementary TE transcripts and repress their transcription with the help of adaptors, which recruit histone modification proteins, such as H3K9 methyltransferase Eggless (Egg) and H3K4 demethylase dLSD128,29,30,31,32.
piRNA pathway mutations cause upregulation of TEs and lead to different oogenesis defects and sterility. Initially, two key components of the piRNA system, Piwi and Yb, have been shown to be required in somatic cells to prevent GSC loss33,34. Later it was found that the lack of several components of the somatic piRNA pathway, including Piwi35,36,37, Vret27, flam23,38 and Egg38,39 lead to the accumulation of undifferentiated germ cells in germaria, known as a germline tumor phenotype. The germ cell differentiation defects observed in piRNA pathway mutants are thought to be related to the dysfunction of ECs36,37,39. Knockdowns of Piwi and Yb specifically in ECs induced large numbers of ectopic GSC-like cells36,37. However, the underlying mechanisms are contradictory. Several papers noted an increased rate of somatic cell death in ovaries due to TE activation27,38. Others have found that Piwi downregulates expression of the dpp gene in ECs36,37 and that TE activation decreases the expression of Wnt4 ligand, which ensures EC function in germ cell differentiation39. It has been shown also that piwi mutations disrupt the spatial position of gonadal intermingled cells (the EC progenitors) and germ cells in early development36.
Here we provide results indicating that the germ cell differentiation defects caused by somatic TE activation in flam mutants are due to a decrease of EC precursor population at the larval stage, whereas no EC death or additional decline of their production rate was observed in flam adult ovaries. We also found drastic oogenesis defects in flam mutants combined with mutations of genes encoding Chk2 (Checkpoint kinase 2) or ATM (ataxia telangiectasia-mutated) checkpoint kinases, contrary to known suppressor effect of chk2 mutation on ovarian development caused by TE derepression in the germline38,40,41,42,43. These results indicate that the somatic cells of ovaries are especially sensitive to TE upregulation upon loss of the Chk2 DNA damage response pathway.
Results
The occurrence of germ cell differentiation defects caused by somatic TE activation correlates with a reduced number of ECs
To extend previous observations27,37,38,39 that activation of TEs in ovarian somatic cells leads to germ cell differentiation defects, we estimated the spectrosome-containing cell number in ovaries lacking various components of the somatic piRNA pathway, some of which have not been tested in this regard before. For this and most subsequent experiments, we analyzed ovaries of 7-day-old females to allow tumor phenotype to develop to a pronounced degree. α-spectrin immunostaining revealed a drastic increase in the number of spectrosome-containing cells upon somatic depletion of Asterix (Arx) (also known as GTSF1) (Fig. 1a,b), a nuclear Piwi cofactor29,30. Depleting Armi, a cytoplasmic component of piRNA biogenesis machinery16,25, in all somatic cells of ovaries (Fig. 1a,b) or only in ECs (Fig. S1a) also caused germline tumors. Moreover, this phenotype was observed in ovaries lacking Zuc and Yb proteins (Fig. S1b). piwiNt mutation causing TE derepression due to cytoplasmic Piwi localization44 also led to the excess of spectrosome-containing germ cells (Fig. S1c), whereas in agreement with our previous report44 a GSC loss phenotype was rare in piwiNt ovaries in contrast to piwi null mutants (Fig. S1d). Thus, our results together with previous findings27,38,39, show that defects in germ cell differentiation are associated with the disruption of any component of the TE silencing pathway in somatic cells of ovaries.
The occurrence of germ cell differentiation defects caused by somatic TE activation correlates with a reduced number of ECs. (a) Examples of wild-type and tumorous germaria stained for α-spectrin (red) to detect spectrosomes and fusomes and for lamin (blue) to visualize cell nuclei. A wild-type germarium (upper panel) usually contains 2–3 GSCs and a few cystoblasts marked by round spectrosomes (s). GSC are located at the anterior end of the germarium in close proximity to somatic cap cells (cc). Dividing cysts carry branched fusome structures (f). Germaria of Arx (middle panel) and Armi (lower panel) knockdowns (KDs) driven by traffic jam Gal4 (tj-Gal4) in ovarian somatic cells carry an excess of spectrosome-containing cells and lack fusomes. (b) Quantification of spectrosome-containing cells in 7-day-old females with KDs of piRNA pathway components in ovarian somatic cells. Each dot corresponds to a single germarium. The central mark indicates the median, and the bottom and top lines indicate the 25th and 75th percentiles, respectively. All tested KD germaria contain significantly more spectrosomes than control (Mann–Whitney U-test; *p < 0.00001). Effects of Piwi and Vret KDs corroborate previously reported result27,35,36,37,38,39 (c). Quantification of spectrosome-containing cells in 7-day-old flam mutants (Mann–Whitney U-test; *p < 0.00001). (d) Quantification of ECs in flam germaria using PZ1444-lacZ line. EC number per germarium is indicated (Mann–Whitney U-test; *p < 0.00001). (e) Immunostaining of flamKG/Df and control germaria for α-spectrin (red), PZ1444-lacZ (green) and lamin (blue). (f) Increase of spectrosome-containing cell number in mutant germaria containing a reduced number of ECs. Percentage of flamKG/Df germaria with more than 10 spectrosome-containing cells in groups of germaria with different number of ECs is shown, based on three replicates (n = 181). Mean +/− s.d. are indicated. (Student’s t-test; **p < 0.05). Scale bars, 10 µm.
Mutations affecting protein-coding genes could exert pleiotropic effects, if corresponding proteins have additional specific functions in oogenesis, unrelated to TE repression, as has been reported for Piwi45,46. Therefore, to directly examine the influence of activated somatic TEs on germline differentiation, we focused on the studies of flam piRNA cluster mutants. In most experiments, we analyzed flamBG/Df and flamKG/Df mutants carrying P-element-induced mutations17,23 and an X chromosome deletion (Df) covering the whole flam locus. Both mutants exhibit the derepression of flam-regulated somatic TEs (Fig. S2b) and about one-half of mutant germaria show a prominent germline tumor phenotype (Figs. 1c and S3a) and some other defects (Fig. S3b). We found no or faint bam-GFP reporter4 expression in germ cells constituting tumors in flamKG/Df germaria (Fig. S3c), indicating an abnormally enhanced BMP-signaling, which may be caused by the failure of ECs to restrict Dpp spreading9,10. The number of ECs visualized by immunostaining for PZ1444 lacZ reporter expression47,48 was reduced about two-fold from an average of 29 ECs per germarium in the flam/+ control to 16 and 18 ECs in flamKG/Df and flamBG/Df mutants, respectively (Figs. 1d and S3d). ECs of flam mutants lacked cellular processes that wrap up differentiating germ cells in wild-type ovaries (Fig. S3e). The latter effect may be a consequence of defective germline differentiation according to literature. For example, it has been shown that bam mutation impedes the formation of EC processes9.
Although the decrease in the EC number was previously reported for piwi somatic knockdown37, it was not clear whether EC reduction directly affects the germ cell differentiation. Since both the spectrosome and EC numbers substantially varied among individuals carrying flam mutations, we wondered how these parameters would be related within a single genotype. Simultaneous immunostaining of mutant ovaries with antibodies against α-spectrin and β-galactosidase (PZ1444 reporter) (Fig. 1e) revealed that germline tumors were rarely detected in flam germaria containing more than 20 ECs, whereas germaria with a small number of ECs more often accumulated large numbers of spectrosome-containing cells (Fig. 1f). This result clearly shows a correlation between EC number reduction and germline tumor formation. Alternatively, the disruption of the differentiation niche may be caused by the loss of EC functional status, such as an abnormal or reduced production of signaling molecules. Impairment of different signaling pathways, including Wnt35,49,50,51, Rho9,52, EGFR53,54, Hh and Hpo/Yki52, as well as the enhancement of BMP signaling in ECs may lead to the germline tumor phenotype. Specifically, loss of piRNA pathway in ECs has been shown to be associated with enhancement of Dpp expression36,37 and a decrease in Wnt signaling39. However, we observed no significant changes in the Wnt2 ligand mRNA expression, a two-fold decrease of the Wnt4 ligand and Frizzled3 (Fz3, target of Wnt pathway) mRNAs and a slight upregulation of Dpp in both flamKG/Df and flamBG/Df germaria compared to control siblings (Fig. S4a). The two-fold decrease of wnt4 expression is likely explained by the observed two-fold reduction of EC number in flam mutants (Fig. 1d), given that Wnt4 (but not Wnt2) is expressed only in ECs and is not detected in other cell types in the germarium49. The observed Dpp upregulation in flam germaria (Fig. S4a) can also be interpreted as a consequence of EC number reduction, because the antagonism between Wnt and BMP pathways in the ovarian somatic cells has been established50,51. The Wnt4 target tkv-lacZ expression49 was similar in ECs of flam and control ovaries indicating active Wnt signaling (Fig. S4b). Secreted Wnt ligands are known to act in ECs in an autocrine manner49,51 resulting in stabilization of a downstream effector protein β-catenin/Armadillo (Arm)55. We failed to find any alteration of Arm protein level in flam germaria by Western blot (Fig. S4c). Overexpression of Arm in flam mutants did not cause a decrease in spectrosome-containing germ cell number (Fig. S4d). Similarly, expression of the constitutive Arm form (UAS-Arm-S10) driven by c587-Gal4 in ECs did not rescue the germline tumor phenotype of piwi mutants (Fig. S4e). As a whole, these results suggest that the germ cell differentiation defect in flam mutants is mediated rather by a decrease in the number of ECs, than by dysfunction of remaining ECs.
EC number and the formation of germline tumor phenotype in flam mutants are determined at the larval stage
ECs are initially produced from the intermingled cells during larval and pupal development56,57,58. Then, in adult ovaries ECs exhibit slow turnover rates, though a fraction of ECs is renewed. Escort stem cells59 or self-duplications of ECs9,10 were previously suggested as a source of new ECs in the adult gonads. A recent study revealed that new ECs in the imago are produced by divisions of follicle stem cells60. The reduction of EC number in flam germaria could be attributed to an increased rate of EC death, to defects of their renewal in adult ovaries or to a decline of EC production during earlier development. The TUNEL assay revealed less than 10% of flam/+ germaria containing at least one apoptotic EC. Unexpectedly, in flamKG/Df germaria the apoptotic ECs were even less frequently detected (Fig. S5a–c). To examine the formation of new ECs in flam ovaries, we carried out immunostaining for phosphohistone H3 Ser10 (PH3) mitotic marker and EdU incorporation assay. Both methods failed to detect a significant number of newly formed ECs in the flamKG/Df and control flies (Fig. S5d,e). Furthermore, most ovaries of females fed on EdU-containing food for three days did not contain EdU-positive ECs (Fig. 2a), suggesting that ECs in tested lines are mainly produced at earlier developmental stages. Importantly, we found about the same number of ECs in germaria of one-, four- or seven-day-old flam adults (Fig. 2b). Thus, the decrease of EC number is observed already in one-day-old flam mutants (Fig. 2b) and, therefore, is determined prior to the imago stage.
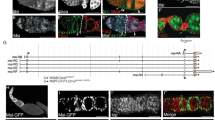
flam mutation leads to a reduction in EC number in larval development but not in adults. (a) flamKG/+ germarium of an adult female after feeding EdU for three days, stained for EdU (purple) and lamin (red). (a’) The same germarium with PZ1444 immunostaining (green). ECs (indicated by green arrows) and CCs are EdU-negative. (b) Quantification of ECs in flamKG/+ (red dots) and flamKG/Df (green dots) flies at the age of 1, 4 and 7 days. The differences between samples of different ages of the same genotypes are not significant (n.s.) (Mann–Whitney U-test; p > 0.1). (c,d) Germaria of females, obtained from larvae reared on EdU-containing food. White and green arrows indicate EdU-positive and EdU-negative ECs, respectively. (e,e’) An example of flamKG/Df germarium with large number of ECs, most of which are EdU-negative after larval EdU incorporation. Full Z-series projections are shown. (f,f’) flamKG/+ germarium. EdU-positive ECs are located at the more anterior region of the germarium compared to EdU-negative ECs. (g) Quantification of EdU-positive and EdU-negative ECs in flamKG/+ and flamKG/Df germaria after larval EdU feeding. Mean +/− s.d. are indicated, based on three replicates (Student’s t test; *p = 0.01; n.s. = not significant). Scale bars, 10 µm.
To find out the developmental stage when ECs are lost, we reared larvae on EdU-containing food, then placed eclosed flies on standard food and analyzed 3-day-old fly ovaries. In this case, all dividing larval cells will contain EdU signals, which then will be diluted with each round of replication in pupae and adults. Expectedly, the follicle cells and most of the germ cells were EdU-negative. Conversely, strong EdU immunostaining was observed in CCs and TF cells (Fig. 2c–f), which are known to be formed in larvae and then do not divide or renew56,57,58,61. About 70% of ECs were also labeled by EdU in flamKG/+ germaria. Apparently, the EdU-positive ECs were formed as a result of a few divisions of parental cells marked by EdU incorporation at the larval stage, while the EdU-negative ECs were likely produced later in development or originated from more actively proliferating cells. Interestingly, EdU-positive ECs were usually located more anteriorly than EdU-negative ECs (Fig. 2c,f), which is consistent with the possible origin of the latter from follicle stem cells60. In flamKG/Df germaria we observed a significant decrease of EdU-positive EC number compared to flamKG/+ sisters (Fig. 2d,e,g), which demonstrates decreased EC precursors formation in flam larvae. However, the number of EdU-negative ECs in flam mutant showed a large scatter of values (Fig. 2g). In some flam germaria the number of the EdU-negative ECs was even increased compared to control (as exemplified in Fig. 2e), suggesting that new ECs can be actively produced after the larval stage to compensate for the lack of EC precursors in earlier development.
Primordial germ cells (PGCs) starting from mid-larval third instar stage are associated with intermingled cells that are EC progenitors. At this stage, all germ cells of the developing ovary are grouped together. Germaria formation occurs later in pupae, when TFs, CCs and their attached PGCs are separated into individual germaria units56,62,63. If EC number and germline differentiation defects are determined during larval development of flam gonads, a correlation can be expected between phenotypes of germaria within the same ovary. Indeed, we found that the numbers of both ECs and spectrosomes were quite similar in flam germaria belonging to the same ovary but varied substantially between individual ovaries (Fig. 3). Thus, developmental events prior to the pupal stage predetermine the germ cell differentiation defects in flam mutants.
flam germaria belonging to the same ovary exhibit similar phenotypes. Quantification of spectrosome-containing cells (a) and ECs (b) per germarium in the same ovaries of flamKG/Df mutants. Germaria in one оvary are shown by grouped dots of the same color. Spectrosome numbers in germaria from tumorous ovaries (A and B) are significantly higher than their numbers in the non-tumorous ovaries (C-G) (Mann–Whitney U-test; *p < 0.05 for A and B vs C-G).
flam mutation induces DNA breaks in somatic cells of larval ovaries
The observed decline of EC precursor production in larval ovaries may be caused by the appearance of TE-induced DNA lesions in their genomes. To check this, we examined the presence of phosphorylated H2Av (γ-H2Av) histone, a commonly used DNA break marker64, in larval somatic intermingled cells marked by Traffic jam (Tj) immunostaining56,58. γ-H2Av dots were observed in 10–20% of Tj-positive cells in wild-type (Batumi) and flamKG/+ (Fig. 4a) third instar larval (L3) ovaries. In flamKG/Df L3 ovaries about 80% of Tj-positive cells contained γ-H2Av signals (Fig. 4b,c). γ-H2Av foci were also detected in Tj-negative somatic cells, including TF cells, as well as somatic apical (AP) and basal (BS) cells (Fig. 4b), which are known to be not incorporated into germaria57. However, most PGCs surrounded by intermingled cells did not contain γ-H2Av foci in mutant ovaries (Fig. 4b). Thus, the flam mutation leads to DNA breaks in somatic, but not germline cells of the larval ovaries. Immunostaining with activated Caspase3 antibodies, as well as TUNEL assay detected an increase of somatic cell death in flamKG/Df larval ovaries (Fig. S6). However, we cannot exclude that a reduction of EC number is partially caused by a decrease of division rate of EC precursors.
Intermingled cells in flam larval ovaries more often contain DNA breaks than ECs in adult ovaries. (a,a’) The flamKG/+ ovary of third instar larval stage stained for lamin (green), γ-H2Av DNA break marker (red) and Traffic jam (Tj, blue) showing intermingled cells (IC). Tj-negative cells include somatic apical (AP) and basal (BC) cells, TF, and Primordial germ cells (PGC). (b,b’) flamKG/Df L3 ovaries accumulate γ-H2Av in most ICs and other somatic cells, including TF and AP, but not in PGCs. (c) Quantification of γ-H2Av-positive among Tj-positive cells in flamKG/+ and flamKG/Df larval ovaries. Mean + /− s.d. are indicated (Student’s t test; *p < 1e-26). (d) flamKG/+ germarium of adult ovary stained for lamin (blue), γ-H2Av (red) and PZ1444 EC marker (green). γ-H2Av signals are observed mainly in germ cells (GC, indicated by yellow arrows). (e,e’) In flamKG/Df germaria γ-H2Av foci appear in most follicle cells (FC, white arrows) and only in some PZ1444-marked ECs (green arrows). (f) Quantification of γ-H2Av signals in FCs and ECs in ovaries of adults. Mean + /− s.d. are indicated (Student’s t test; *p < 1e-9). Scale bars, 10 µm.
Then we monitored γ-H2Av presence in the somatic cells of adult flam ovaries. In flam/+ germaria, as in wild-type, γ-H2Av signals were absent in ECs and CCs, but were detected in the meiotic germ cells and endocycling nurse cells (Fig. 4d) where DNA breaks are generated during normal development65,66,67,68,69. In the flamKG/Df germaria only about 20% of ECs contained γ-H2Av foci, whereas follicle cells were mostly γ-H2Av-positive (Fig. 4e,f). These observations indicate that DNA damage events occur in mature flam ECs less often than in their precursors, intermingled cells, at the larval stage and/or mature ECs have an enhanced capacity to repair DNA lesions.
The absence of Chk2 or ATM checkpoint kinases enhanced oogenesis defects of flam mutants
DNA damage is known to block cell proliferation through the activation of checkpoint kinases, which induce cell cycle arrest followed by apoptosis or DNA repair (for review see70). Drosophila Chk2 encoded by the Mnk/Loki gene together with other checkpoint kinases is required for cell cycle arrest in response to DNA breaks in both somatic and germ cells71,72,73,74. Another function of Chk2 is p53 phosphorylation that activates transcription of genes involved in DNA repair and/or apoptosis pathways75,76. To examine whether the flam mutant phenotype is mediated by the checkpoint response to TE-induced DNA breaks, we crossed the mnkp6 mutation (the well-characterized loss of function allele42,43,71,77) into a flam mutant background. Although the chk2 mutation was shown to partially rescue the germline differentiation defects induced by TE activation in germ cells38,42,43, we unexpectedly observed its opposite effect in flam mutants. The flamKG/Df; mnkp6/mnkp6 double mutants had drastically more defective ovaries than flamKG/Df; mnkp6/+ individuals, whereas flamKG/+; mnkp6/mnkp6 ovaries displayed no visible morphological defects (Fig. 5a–d). The formation of germaria was abolished in most flamKG/Df; mnkp6/mnkp6 ovaries (Fig. 5c,d) and in some of them the number of Tj-positive ovarian somatic cells was highly reduced (Fig. 5d). Severe oogenesis defects were also observed when we combined flam with two different mutations in the tefu gene (Fig. S7), encoding a Drosophila homolog of ATM kinase that is directly recruited and activated by DNA double-strand breaks, acting upstream of Chk270,78. We suggested that the observed catastrophic ovarian phenotypes can be induced by the death or dysfunction of ovarian somatic cells, including ECs due to their inability to repair TE-induced DNA lesions. Then, we checked whether DNA breaks are accumulated in ECs of flam mutants lacking checkpoint response. We found that somatic cell nuclei in rarely observed germaria-like structures of flamKG/Df; mnkp6/mnkp6 ovaries were dramatically enriched in γ-H2Av signals compared to both flamKG/Df; mnkp6/+ and flamKG/+; mnkp6/mnkp6 germaria (Fig. 5e–g). Somatic depletion of Mnk in the flamKG/Df, but not flamKG/+ background also caused accumulation of γ-H2Av in nearly all ECs (Fig. 5h,i) in contrast to about 20% of γ-H2Av-positive ECs observed in flamKG/Df ovaries (Fig. 4f). In addition, γ-H2Av foci were accumulated in ECs upon depletion of another checkpoint kinase, mei-41 (Drosophila homolog of ATR) (Fig. 5h,j), which, however, did not enhance oogenesis defects. Thus, Chk2, ATM and ATR kinases are involved in cellular response upon TE activation in ECs or their progenitors, but their specific molecular functions in this process warrant further examination.
chk2 mutation enhances ovarian defects in flam mutants. (a) flamKG/Df; mnkp6/+ germarium stained for lamin (blue), α-spectrin (red) and Tj (green) showing nuclei of CCs, ECs and FCs. (b) flamKG/+; mnkp6/mnkp6 germarium with no morphological defects. (c,d) Fragments of flamKG/Df; mnkp6/mnkp6 ovaries showing an impaired formation of germaria and ovarioles. Abnormal germaria-like structures (G) are filled with spectrosome-containing cells and lack Tj-positive somatic cells or lack both germ cells and ECs. Separate accumulations of Tj-positive somatic cells (S) are indicated. (e) Immunostaining of flamKG/+; mnkp6/mnkp6 germarium with Tj (green), γ-H2Av (red) and lamin (blue). γ-H2Av signals are observed in meiotic germ cells and in a few somatic cells. (f) flamKG/Df; mnkp6/+ germarium containing γ-H2Av dots in follicle cells and in some ECs. (g) flamKG/Df; mnkp6/mnkp6 germarium showing increased intensity of γ-H2Av signals in Tj-positive somatic cells. (h) Quantification of γ-H2Av-positive ECs in flamKG/Df; tj-Gal4/Cy (no KD, control), flamKG/Df; tj-Gal4 > mnk KD and flamKG/Df; tj-Gal4 > mei-41 KD ovaries. Mean +/− s.d. are indicated (Student’s t test; *p < 0.001). (i) Immunostaining of flamKG/Df; tj-Gal4 >mnk KD germarium with γ-H2Av (red) and lamin (blue) and Tj-marked somatic cells (green) (i’). (j,j’) Immunostaining of flamKG/Df; tj-Gal4 >mei-41 KD germarium with the same antibodies. (k) A working model of the occurrence of germ cell differentiation defects due to TE activation in ovarian somatic cells. Scale bars, 10 µm.
Discussion
Activation of TEs in ovarian somatic cells is known to compromise differentiation of germ cells27,38,39. Here, we found that the accumulation of GSC-like cells caused by mutations in the somatic piRNA cluster, flam, is determined by an insufficient number of somatic ECs. We demonstrated that the decrease of EC production in flam mutants, as well as the formation of germline tumor phenotype, depend on the events which occur in larvae and possibly at earlier stages of development, but not in the adult ovaries (Fig. 5k). These observations are consistent with previous report showing that Piwi expression in intermingled cells during the larval L3 stage is required to restrict GSC number in adults36. Of note, somatic piRNAs against TEs were found to be produced de novo in large amounts between embryogenesis and the L3 stage79 that shows the piRNA pathway activity in somatic cells of larval ovaries. Our finding of abundant DNA breaks in intermingled cells of flam mutant (Fig. 4b,c) indicates that activated TEs affect the genome of EC precursors at the larval stage. Interestingly, the intermingled cells accumulated more DNA breaks than mature ECs (Fig. 4). The reason for this vulnerability of somatic niche to TE activation during larval development remains unclear.
Mobilization of TEs in the germ cells was shown to initiate the checkpoint response. In developing oocytes, TE-induced DNA breaks trigger Chk2-dependent oocyte polarization abnormalities40,41. Consistent with this, chk2 mutation suppresses polarization defects in the oocytes of piRNA pathway mutants40,41. Derepression of TEs in GSCs leads to Chk2-mediated arrest of cell cycle42,43,80 and induction of p53 activity81, which launches DNA repair or apoptosis. In particular, mutation of aub gene encoding a germline-specific piRNA-binding protein is phenotypically manifested as a decrease of GSC number and a delayed differentiation of cystoblasts42,43. The chk2 mutation partially rescues these defects42,43. Transpositions of P element during hybrid dysgenesis also induce Chk2-dependent arrest of germ cell differentiation and selective apoptosis of some GSCs, whereas mutating Chk2 restores GSC self-renewal and normal looking germaria38,80. However, in this case chk2 mutants show strong γ-H2Av signals and death of some cells at all oogenesis stages and never restore fertility80. Interestingly, GSCs in dysgenic females are able, over time, to acquire resistance to P element due to the piRNA amplification by ping-pong mechanism, whereas this adaptation does not occur in chk2 mutants80. As a result, the ovarian defects in older dysgenic females are enhanced by chk2 mutation80. Here, we for the first time examined the role of checkpoint response upon genomic stress caused by TE activation in somatic cells of ovaries, which lack ping-pong piRNA amplification11,14,15. We found that the absence of Chk2 or ATM kinases in the flam mutant background leads to dramatically more severe oogenesis defects compared to those induced by the flam mutation alone (Figs. 5 and S7). Thus, in contrast to germ cells, the Chk2-dependent response to TE activation in somatic ovarian cells is critical for the preservation of normal ovarian structure. The observed phenotypes of flam mnk double mutants indicate the loss of somatic cells due to the accumulation of unrepaired DNA lesions (Fig. 5). Our results suggest that the primary function of the Chk2-mediated response in ovarian somatic cells is the induction of DNA repair (Fig. 5k). The canonical activation of DNA repair/apoptosis pathways following DNA damage requires Chk2-mediated phosphorylation of p5375,76. However, p53 activity in Drosophila ovaries was shown to be restricted to GSCs and cystoblasts81, suggesting that in ovarian somatic cells Chk2 induces DNA repair by an unknown mechanism, which is of interest for further research.
Methods
Drosophila stocks
Drosophila melanogaster stocks were maintained under standard conditions at 25 °C. For analysis of flam mutations the following stocks were obtained from the Bloomington Drosophila Stock Center: w1118 P{GT1}1, flamBG02658 (#13912, flamBG), y1 P{SUPor-P}flamKG00476 (BDSC #16453, flamKG) and Df(1)Exel6255, w1118 P{XP-U}Exel6255/FM7c (BDSC #7723, flamDf). To distinguish between flamKG/Df and flamKG/+ larvae we used the y1 w67c23 Alr1/FM7i, P{w[ + mC] = ActGFP}JMR3 (BDSC #25048) balancer and manually selected GFP-positive and GFP-negative larvae. To visualize ECs flam mutations were combined with the PZ1444 lacZ enhancer trap line47,48. For piwi, we used piwi2 and piwi3 null mutations34 and piwiNt mutation with disrupted Piwi nuclear localization44. To analyze piwiNt/piwiNt females we used a strain with a higher survival rate of homozygous flies due to a change of genetic background. The following UAS-RNAi stocks were obtained from Vienna Drosophila Resource Center (VDRC): piwi-RNAi (#101658), vret-RNAi (#34897, #101134), armi-RNAi (#103589), arx-RNAi (#40480, #40479), mei-41-RNAi (#11251), Chk2-RNAi (#110342). RNAi depletion or expression of proteins was induced by UAS-Dicer tj-Gal4 driver active in most somatic ovarian cells16,82 or c587-Gal4 driver active in ECs and early follicle progenitors9,37. Other fly stocks were the following: Df(2 L)Prl and zucHM27 (from T. Schüpbach lab), mnkp6 (lokp6)71 (from M. Simonelig lab), tefu1, tefured31 83, fs(1)Yb1 (Yb1), fs(1)Yb72 (Yb72)33, bamGFP4, Batumi, P{w(+mC) = UAS-arm.S10}C, y(1) w(1118) (BDSC #4782, UAS-arm.S10), y(1) w(1118); P{w(+mC) = UAS-arm.Exel}2 (BDSC #8369, UAS-arm), y(1) w(67c23); P{w(+mC) = lacW}tkv(k16713)/CyO (BDSC #11191, tkv-lacZ), y(1) w(*); P{w(+mC) = UAS-mCD8::GFP.L}LL5 (BDSC #5137, UAS-mCD8::GFP).
Immunostaining
For spectrosome analysis ovaries from 7-day-old females were used, and for other purposes - as indicated in the text. We revealed that germline differentiation defects in flam and piwi mutants are more pronounced in the progeny of older parents and to standardize further analysis we used offspring from the parents less than three weeks old. Immunostaining was basically performed as described previously84 with some modifications. Ovaries were manually isolated in PBT (PBS containing 0.01% Tween-20) at 4 °C, rinsed in PBS and fixed in 4% formaldehyde (in PBT) for 25 min at room temperature. Fixation was stopped by incubation with 0.25 M glycine (Sigma-Aldrich) for 5 min. Then ovaries were washed in PBS three times for 10 min at room temperature, permeabilized with PBTX (PBS with 0.1% Tween-20, 0.3% Triton X-100) for 10 min, blocked with PBTX containing 3% normal goat serum (NGS, Invitrogen) for 3 h, incubated with primary antibody in PBTX containing 3% NGS for 7 h at room temperature, or overnight at 4 °C, washed in PBTX three times for 10 min, incubated with secondary antibodies (1:1000) in PBTX containing 3% NGS for 7 h or overnight in a dark chamber, and then washed in PBTX three times for 10 min. Coverslips were mounted with a drop of SlowFade Gold Antifade reagent (Invitrogen) containing DAPI. The following primary antibodies were used: rabbit anti-lamin Dm0 (1:500, provided by P. Fisher85), chicken anti-β-galactosidase (1:500, Abcam, ab9361), mouse anti-β-galactosidase (1:200, DSHB #40-1а), rabbit anti-pS10H3 (1:200, Millipore #MC463), rabbit anti-GFP (1:500, Abcam, ab290), mouse anti-α-spectrin (1:200, DSHB, 3А9), rabbit anti-γ-H2av (1:100, Rockland, anti-H2AvD pS137), rat anti-Vasa (1:100; DSHB), guinea pig anti-Tj (1:5000, a gift from Dorothea Godt), rabbit anti-Caspase-3 antibody (1:200; Abcam, ab13847). The following secondary antibodies (Invitrogen, Thermo Fisher Scientific) were used: anti-rat IgG Alexa Fluor 546; anti-rabbit IgG Alexa Fluor 488; anti-rabbit IgG Alexa Fluor 546; anti-rabbit IgG Alexa Fluor 633; anti-mouse IgG Alexa Fluor 488; anti-mouse IgG Alexa Fluor 633; anti-chicken IgG Alexa Fluor 633; anti-guinea pig IgG Alexa Fluor 488; anti-guinea pig IgG Alexa Fluor 633. Confocal microscopy was done using LSM 510 META system (Zeiss).
TUNEL assay
TUNEL staining was performed using Click-iT™ Plus TUNEL Assay for In Situ Apoptosis Detection, Alexa Fluor™ 647 dye kit (#C10619, Invitrigen, Thermo Fisher Scientific) according to the manufacturer’s instructions.
EdU incorporation assays
For the two-hour EdU labeling, the ovaries were incubated in Grace’s medium containing 10 µM EdU for 2 hours at 25 °C. For the EdU in vivo incorporation assay, females were fed on food with yeast paste containing EdU (0.5 mM) for three days. For larval EdU assay, parental flies were placed on EdU-containing food (0.5 mM), where larvae developed. Then newly eclosed flies were placed on food without EdU and after 3 days the ovaries were dissected and analyzed.
The ovaries from all these types of assays were fixed, permeabilized as described above, and processed for EdU label detection using the Click-iT™ reaction according to the manufacturer’s instructions. Click-iT reaction was carried out in a cocktail containing Alexa Fluor 647 azide, triethylammonium salt (#A10277, Invitrogen) and Reaction Buffer Kit (#C10269, Invitrogen) 30 min in the dark at room temperature. Then ovaries were washed in PBTX and processed for immunostaining.
Western blot
Ovarian lysates were fractionated by SDS-PAGE (10% acrylamide gel) and transferred to a PVDF membrane (Immobilon-P, Millipore). Blots were developed using alkaline phosphatase-conjugated secondary antibody (Sigma) and the Immun-Star AP detection system (Bio-Rad). The following primary antibodies were used: mouse anti-Arm (1:500, DSHB), and mouse anti-β-Actin (1:3000; Abcam, ab8224).
RT-qPCR analysis
Total RNA was isolated from manually dissected ovaries using Trizol reagent (Invitrogen, Thermo Fisher Scientific) and cleared of genomic DNA by DNA-free kit (Ambion).
For analysis of signaling pathway genes in germaria, RNA was isolated from 0-1-day ovaries containing no late stage egg chambers. 1 μg of total RNA was used for the reverse transcription reaction with oligo(dT) primer and Superscript II reverse transcriptase (Invitrogen). The resulting cDNAs in at least three biological replicates were analyzed by RT-qPCR performed in MJ Mini thermal cycler (Bio-Rad) using SYBR Green chemistry (Applied Biosystems). The following primers were used for PCR:
Gypsy for CTTCACGTTCTGCGAGCGGTCT,
Gypsy rev CGCTCGAAGGTTACCAGGTAGGTTC,
Zam for3 TCACATCCTTCCAGCAATCTTCAA,
Zam rev3 TATTACAGTTTCTGACATTATTTCTTCGTG,
MDG1 dir AACAGAAACGCCAGCAACAGC,
MDG1 rev CGTTCCCATGTCCGTTGTGAT,
Idefix for AACAAAATCGTGGCAGGAAG,
Idefix rev TCCATTTTTCGCGTTTACTG,
dpp for2 GGCTTCTACTCCTCGCAGTG,
dpp rev2 TGCTTTTGCTAATGCTGTGC,
wnt4 for5 ATGATCCTCACCCACCTGAG,
wnt4 rev5 ACCTGACCAGCATTGTTTCC,
wnt2 for CAATAACCGAGCAGGGAGAAC,
wnt2 rev CATGAGTCTATCGCCAACCAG,
fz3 for TCTGCTTCGTCCTGACACTG,
fz3 rev CCTTGCTTGATTGTGGAACAC,
Rp49_up ATGACCATCCGCCCAGCATAC,
Rp49_rev2 GCTTAGCATATCGATCCGACTGG.
Data availability
All data generated or analyzed during this study are included in this published article (and its Supplementary Information Files).
Change history
20 December 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41598-021-03387-5
References
Spradling, A., Fuller, M. T., Braun, R. E. & Yoshida, S. Germline stem cells. Cold Spring Harbor perspectives in biology 3, a002642, https://doi.org/10.1101/cshperspect.a002642 (2011).
Chen, S., Wang, S. & Xie, T. Restricting self-renewal signals within the stem cell niche: multiple levels of control. Current opinion in genetics & development 21, 684–689, https://doi.org/10.1016/j.gde.2011.07.008 (2011).
Losick, V. P., Morris, L. X., Fox, D. T. & Spradling, A. Drosophila stem cell niches: a decade of discovery suggests a unified view of stem cell regulation. Developmental cell 21, 159–171, https://doi.org/10.1016/j.devcel.2011.06.018 (2011).
Chen, D. & McKearin, D. Dpp signaling silences bam transcription directly to establish asymmetric divisions of germline stem cells. Current biology: CB 13, 1786–1791 (2003).
Liu, Z. et al. Coordinated niche-associated signals promote germline homeostasis in the Drosophila ovary. The Journal of cell biology 211, 469–484, https://doi.org/10.1083/jcb.201503033 (2015).
Song, X. et al. Bmp signals from niche cells directly repress transcription of a differentiation-promoting gene, bag of marbles, in germline stem cells in the Drosophila ovary. Development 131, 1353–1364, https://doi.org/10.1242/dev.01026 (2004).
Xie, T. & Spradling, A. C. decapentaplegic is essential for the maintenance and division of germline stem cells in the Drosophila ovary. Cell 94, 251–260 (1998).
Xie, T. Control of germline stem cell self-renewal and differentiation in the Drosophila ovary: concerted actions of niche signals and intrinsic factors. Wiley interdisciplinary reviews. Developmental biology 2, 261–273, https://doi.org/10.1002/wdev.60 (2013).
Kirilly, D., Wang, S. & Xie, T. Self-maintained escort cells form a germline stem cell differentiation niche. Development 138, 5087–5097, https://doi.org/10.1242/dev.067850 (2011).
Morris, L. X. & Spradling, A. C. Long-term live imaging provides new insight into stem cell regulation and germline-soma coordination in the Drosophila ovary. Development 138, 2207–2215, https://doi.org/10.1242/dev.065508 (2011).
Czech, B. & Hannon, G. J. One Loop to Rule Them All: The Ping-Pong Cycle and piRNA-Guided Silencing. Trends in biochemical sciences 41, 324–337, https://doi.org/10.1016/j.tibs.2015.12.008 (2016).
Sarot, E., Payen-Groschene, G., Bucheton, A. & Pelisson, A. Evidence for a piwi-dependent RNA silencing of the gypsy endogenous retrovirus by the Drosophila melanogaster flamenco gene. Genetics 166, 1313–1321 (2004).
Pelisson, A., Sarot, E., Payen-Groschene, G. & Bucheton, A. A novel repeat-associated small interfering RNA-mediated silencing pathway downregulates complementary sense gypsy transcripts in somatic cells of the Drosophila ovary. Journal of virology 81, 1951–1960, https://doi.org/10.1128/JVI.01980-06 (2007).
Malone, C. D. et al. Specialized piRNA pathways act in germline and somatic tissues of the Drosophila ovary. Cell 137, 522–535, https://doi.org/10.1016/j.cell.2009.03.040 (2009).
Li, C. et al. Collapse of germline piRNAs in the absence of Argonaute3 reveals somatic piRNAs in flies. Cell 137, 509–521, https://doi.org/10.1016/j.cell.2009.04.027 (2009).
Olivieri, D., Sykora, M. M., Sachidanandam, R., Mechtler, K. & Brennecke, J. An in vivo RNAi assay identifies major genetic and cellular requirements for primary piRNA biogenesis in Drosophila. The EMBO journal 29, 3301–3317, https://doi.org/10.1038/emboj.2010.212 (2010).
Brennecke, J. et al. Discrete small RNA-generating loci as master regulators of transposon activity in Drosophila. Cell 128, 1089–1103, https://doi.org/10.1016/j.cell.2007.01.043 (2007).
Robert, V., Prud’homme, N., Kim, A., Bucheton, A. & Pelisson, A. Characterization of the flamenco region of the Drosophila melanogaster genome. Genetics 158, 701–713 (2001).
Zanni, V. et al. Distribution, evolution, and diversity of retrotransposons at the flamenco locus reflect the regulatory properties of piRNA clusters. Proceedings of the National Academy of Sciences of the United States of America 110, 19842–19847, https://doi.org/10.1073/pnas.1313677110 (2013).
Goriaux, C., Theron, E., Brasset, E. & Vaury, C. History of the discovery of a master locus producing piRNAs: the flamenco/COM locus in Drosophila melanogaster. Frontiers in genetics 5, 257, https://doi.org/10.3389/fgene.2014.00257 (2014).
Prud’homme, N., Gans, M., Masson, M., Terzian, C. & Bucheton, A. Flamenco, a gene controlling the gypsy retrovirus of Drosophila melanogaster. Genetics 139, 697–711 (1995).
Pelisson, A. et al. Gypsy transposition correlates with the production of a retroviral envelope-like protein under the tissue-specific control of the Drosophila flamenco gene. The EMBO journal 13, 4401–4411 (1994).
Mevel-Ninio, M., Pelisson, A., Kinder, J., Campos, A. R. & Bucheton, A. The flamenco locus controls the gypsy and ZAM retroviruses and is required for Drosophila oogenesis. Genetics 175, 1615–1624, https://doi.org/10.1534/genetics.106.068106 (2007).
Desset, S., Buchon, N., Meignin, C., Coiffet, M. & Vaury, C. In Drosophila melanogaster the COM locus directs the somatic silencing of two retrotransposons through both Piwi-dependent and -independent pathways. PloS one 3, e1526, https://doi.org/10.1371/journal.pone.0001526 (2008).
Saito, K. et al. Roles for the Yb body components Armitage and Yb in primary piRNA biogenesis in Drosophila. Genes & development 24, 2493–2498, https://doi.org/10.1101/gad.1989510 (2010).
Nishimasu, H. et al. Structure and function of Zucchini endoribonuclease in piRNA biogenesis. Nature 491, 284–287, https://doi.org/10.1038/nature11509 (2012).
Zamparini, A. L. et al. Vreteno, a gonad-specific protein, is essential for germline development and primary piRNA biogenesis in Drosophila. Development 138, 4039–4050, https://doi.org/10.1242/dev.069187 (2011).
Sienski, G., Donertas, D. & Brennecke, J. Transcriptional silencing of transposons by Piwi and maelstrom and its impact on chromatin state and gene expression. Cell 151, 964–980, https://doi.org/10.1016/j.cell.2012.10.040 (2012).
Donertas, D., Sienski, G. & Brennecke, J. Drosophila Gtsf1 is an essential component of the Piwi-mediated transcriptional silencing complex. Genes & development 27, 1693–1705, https://doi.org/10.1101/gad.221150.113 (2013).
Ohtani, H. et al. DmGTSF1 is necessary for Piwi-piRISC-mediated transcriptional transposon silencing in the Drosophila ovary. Genes & development 27, 1656–1661, https://doi.org/10.1101/gad.221515.113 (2013).
Sienski, G. et al. Silencio/CG9754 connects the Piwi-piRNA complex to the cellular heterochromatin machinery. Genes & development 29, 2258–2271, https://doi.org/10.1101/gad.271908.115 (2015).
Yu, Y. et al. Panoramix enforces piRNA-dependent cotranscriptional silencing. Science 350, 339–342, https://doi.org/10.1126/science.aab0700 (2015).
King, F. J., Szakmary, A., Cox, D. N. & Lin, H. Yb modulates the divisions of both germline and somatic stem cells through piwi- and hh-mediated mechanisms in the Drosophila ovary. Molecular cell 7, 497–508 (2001).
Lin, H. & Spradling, A. C. A novel group of pumilio mutations affects the asymmetric division of germline stem cells in the Drosophila ovary. Development 124, 2463–2476 (1997).
Hamada-Kawaguchi, N., Nore, B. F., Kuwada, Y., Smith, C. I. & Yamamoto, D. Btk29A promotes Wnt4 signaling in the niche to terminate germ cell proliferation in Drosophila. Science 343, 294–297, https://doi.org/10.1126/science.1244512 (2014).
Jin, Z., Flynt, A. S. & Lai, E. C. Drosophila piwi mutants exhibit germline stem cell tumors that are sustained by elevated Dpp signaling. Current biology: CB 23, 1442–1448, https://doi.org/10.1016/j.cub.2013.06.021 (2013).
Ma, X. et al. Piwi is required in multiple cell types to control germline stem cell lineage development in the Drosophila ovary. PloS one 9, e90267, https://doi.org/10.1371/journal.pone.0090267 (2014).
Rangan, P. et al. piRNA production requires heterochromatin formation in Drosophila. Current biology: CB 21, 1373–1379, https://doi.org/10.1016/j.cub.2011.06.057 (2011).
Upadhyay, M. et al. Transposon Dysregulation Modulates dWnt4 Signaling to Control Germline Stem Cell Differentiation in Drosophila. PLoS genetics 12, e1005918, https://doi.org/10.1371/journal.pgen.1005918 (2016).
Chen, Y., Pane, A. & Schupbach, T. Cutoff and aubergine mutations result in retrotransposon upregulation and checkpoint activation in Drosophila. Current biology: CB 17, 637–642, https://doi.org/10.1016/j.cub.2007.02.027 (2007).
Klattenhoff, C. et al. Drosophila rasiRNA pathway mutations disrupt embryonic axis specification through activation of an ATR/Chk2 DNA damage response. Developmental cell 12, 45–55, https://doi.org/10.1016/j.devcel.2006.12.001 (2007).
Ma, X. et al. Aubergine Controls Germline Stem Cell Self-Renewal and Progeny Differentiation via Distinct Mechanisms. Developmental cell 41, 157–169 e155, https://doi.org/10.1016/j.devcel.2017.03.023 (2017).
Rojas-Rios, P., Chartier, A., Pierson, S. & Simonelig, M. Aubergine and piRNAs promote germline stem cell self-renewal by repressing the proto-oncogene Cbl. The EMBO journal 36, 3194–3211, https://doi.org/10.15252/embj.201797259 (2017).
Klenov, M. S. et al. Separation of stem cell maintenance and transposon silencing functions of Piwi protein. Proceedings of the National Academy of Sciences of the United States of America 108, 18760–18765, https://doi.org/10.1073/pnas.1106676108 (2011).
Klein, J. D. et al. c-Fos Repression by Piwi Regulates Drosophila Ovarian Germline Formation and Tissue Morphogenesis. PLoS genetics 12, e1006281, https://doi.org/10.1371/journal.pgen.1006281 (2016).
Peng, J. C., Valouev, A., Liu, N. & Lin, H. Piwi maintains germline stem cells and oogenesis in Drosophila through negative regulation of Polycomb group proteins. Nature genetics 48, 283–291, https://doi.org/10.1038/ng.3486 (2016).
Margolis, J. & Spradling, A. Identification and behavior of epithelial stem cells in the Drosophila ovary. Development 121, 3797–3807 (1995).
Xie, T. & Spradling, A. C. A niche maintaining germ line stem cells in the Drosophila ovary. Science 290, 328–330 (2000).
Luo, L., Wang, H., Fan, C., Liu, S. & Cai, Y. Wnt ligands regulate Tkv expression to constrain Dpp activity in the Drosophila ovarian stem cell niche. The Journal of cell biology 209, 595–608, https://doi.org/10.1083/jcb.201409142 (2015).
Mottier-Pavie, V. I., Palacios, V., Eliazer, S., Scoggin, S. & Buszczak, M. The Wnt pathway limits BMP signaling outside of the germline stem cell niche in Drosophila ovaries. Developmental biology 417, 50–62, https://doi.org/10.1016/j.ydbio.2016.06.038 (2016).
Wang, S. et al. Wnt signaling-mediated redox regulation maintains the germ line stem cell differentiation niche. eLife 4, e08174, https://doi.org/10.7554/eLife.08174 (2015).
Huang, J., Reilein, A. & Kalderon, D. Yorkie and Hedgehog independently restrict BMP production in escort cells to permit germline differentiation in the Drosophila ovary. Development 144, 2584–2594, https://doi.org/10.1242/dev.147702 (2017).
Banisch, T. U., Maimon, I., Dadosh, T. & Gilboa, L. Escort cells generate a dynamic compartment for germline stem cell differentiation via combined Stat and Erk signalling. Development 144, 1937–1947, https://doi.org/10.1242/dev.143727 (2017).
Liu, M., Lim, T. M. & Cai, Y. The Drosophila female germline stem cell lineage acts to spatially restrict DPP function within the niche. Science signaling 3, ra57, https://doi.org/10.1126/scisignal.2000740 (2010).
Clevers, H., Loh, K. M. & Nusse, R. Stem cell signaling. An integral program for tissue renewal and regeneration: Wnt signaling and stem cell control. Science 346, 1248012, https://doi.org/10.1126/science.1248012 (2014).
Gancz, D. & Gilboa, L. Insulin and Target of rapamycin signaling orchestrate the development of ovarian niche-stem cell units in Drosophila. Development 140, 4145–4154, https://doi.org/10.1242/dev.093773 (2013).
Lai, C. M. et al. Hedgehog signaling establishes precursors for germline stem cell niches by regulating cell adhesion. The Journal of cell biology 216, 1439–1453, https://doi.org/10.1083/jcb.201610063 (2017).
Panchal, T. et al. Specification and spatial arrangement of cells in the germline stem cell niche of the Drosophila ovary depend on the Maf transcription factor Traffic jam. PLoS genetics 13, e1006790, https://doi.org/10.1371/journal.pgen.1006790 (2017).
Decotto, E. & Spradling, A. C. The Drosophila ovarian and testis stem cell niches: similar somatic stem cells and signals. Developmental cell 9, 501–510, https://doi.org/10.1016/j.devcel.2005.08.012 (2005).
Reilein, A. et al. Alternative direct stem cell derivatives defined by stem cell location and graded Wnt signalling. Nature cell biology 19, 433–444, https://doi.org/10.1038/ncb3505 (2017).
Sahut-Barnola, I., Godt, D., Laski, F. A. & Couderc, J. L. Drosophila ovary morphogenesis: analysis of terminal filament formation and identification of a gene required for this process. Developmental biology 170, 127–135, https://doi.org/10.1006/dbio.1995.1201 (1995).
Zhu, C. H. & Xie, T. Clonal expansion of ovarian germline stem cells during niche formation in Drosophila. Development 130, 2579–2588 (2003).
Sato, T., Ogata, J. & Niki, Y. BMP and Hh signaling affects primordial germ cell division in Drosophila. Zoological science 27, 804–810, https://doi.org/10.2108/zsj.27.804 (2010).
Madigan, J. P., Chotkowski, H. L. & Glaser, R. L. DNA double-strand break-induced phosphorylation of Drosophila histone variant H2Av helps prevent radiation-induced apoptosis. Nucleic acids research 30, 3698–3705 (2002).
Jang, J. K., Sherizen, D. E., Bhagat, R., Manheim, E. A. & McKim, K. S. Relationship of DNA double-strand breaks to synapsis in Drosophila. Journal of cell science 116, 3069–3077, https://doi.org/10.1242/jcs.00614 (2003).
Joyce, E. F. et al. Drosophila ATM and ATR have distinct activities in the regulation of meiotic DNA damage and repair. The Journal of cell biology 195, 359–367, https://doi.org/10.1083/jcb.201104121 (2011).
Mehrotra, S., Maqbool, S. B., Kolpakas, A., Murnen, K. & Calvi, B. R. Endocycling cells do not apoptose in response to DNA rereplication genotoxic stress. Genes & development 22, 3158–3171, https://doi.org/10.1101/gad.1710208 (2008).
Mehrotra, S. & McKim, K. S. Temporal analysis of meiotic DNA double-strand break formation and repair in Drosophila females. PLoS genetics 2, e200, https://doi.org/10.1371/journal.pgen.0020200 (2006).
Narbonne-Reveau, K. & Lilly, M. The Cyclin-dependent kinase inhibitor Dacapo promotes genomic stability during premeiotic S phase. Molecular biology of the cell 20, 1960–1969, https://doi.org/10.1091/mbc.E08-09-0916 (2009).
Song, Y. H. Drosophila melanogaster: a model for the study of DNA damage checkpoint response. Molecules and cells 19, 167–179 (2005).
Masrouha, N., Yang, L., Hijal, S., Larochelle, S. & Suter, B. The Drosophila chk2 gene loki is essential for embryonic DNA double-strand-break checkpoints induced in S phase or G2. Genetics 163, 973–982 (2003).
Shim, H. J., Lee, E. M., Nguyen, L. D., Shim, J. & Song, Y. H. High-dose irradiation induces cell cycle arrest, apoptosis, and developmental defects during Drosophila oogenesis. PloS one 9, e89009, https://doi.org/10.1371/journal.pone.0089009 (2014).
Xu, J. & Du, W. Drosophila chk2 plays an important role in a mitotic checkpoint in syncytial embryos. FEBS letters 545, 209–212 (2003).
Xu, J., Xin, S. & Du, W. Drosophila Chk2 is required for DNA damage-mediated cell cycle arrest and apoptosis. FEBS letters 508, 394–398 (2001).
Brodsky, M. H. et al. Drosophila melanogaster MNK/Chk2 and p53 regulate multiple DNA repair and apoptotic pathways following DNA damage. Molecular and cellular biology 24, 1219–1231 (2004).
Peters, M. et al. Chk2 regulates irradiation-induced, p53-mediated apoptosis in Drosophila. Proceedings of the National Academy of Sciences of the United States of America 99, 11305–11310, https://doi.org/10.1073/pnas.172382899 (2002).
Ma, X. et al. DNA damage-induced Lok/CHK2 activation compromises germline stem cell self-renewal and lineage differentiation. Development 143, 4312–4323, https://doi.org/10.1242/dev.141069 (2016).
Smith, J., Tho, L. M., Xu, N. & Gillespie, D. A. The ATM-Chk2 and ATR-Chk1 pathways in DNA damage signaling and cancer. Advances in cancer research 108, 73–112, https://doi.org/10.1016/B978-0-12-380888-2.00003-0 (2010).
Marie, P. P., Ronsseray, S. & Boivin, A. From Embryo to Adult: piRNA-Mediated Silencing throughout Germline Development in Drosophila. G3 7, 505–516, https://doi.org/10.1534/g3.116.037291 (2017).
Moon, S. et al. A Robust Transposon-Endogenizing Response from Germline Stem Cells. Developmental cell 47, 660–671 e663, https://doi.org/10.1016/j.devcel.2018.10.011 (2018).
Wylie, A., Lu, W. J., D’Brot, A., Buszczak, M. & Abrams, J. M. p53 activity is selectively licensed in the Drosophila stem cell compartment. eLife 3, e01530, https://doi.org/10.7554/eLife.01530 (2014).
Tanentzapf, G., Devenport, D., Godt, D. & Brown, N. H. Integrin-dependent anchoring of a stem-cell niche. Nature cell biology 9, 1413–1418, https://doi.org/10.1038/ncb1660 (2007).
Oikemus, S. R. et al. Drosophila atm/telomere fusion is required for telomeric localization of HP1 and telomere position effect. Genes & development 18, 1850–1861, https://doi.org/10.1101/gad.1202504 (2004).
Ilyin, A. A. et al. Piwi interacts with chromatin at nuclear pores and promiscuously binds nuclear transcripts in Drosophila ovarian somatic cells. Nucleic acids research 45, 7666–7680, https://doi.org/10.1093/nar/gkx355 (2017).
Osouda, S. et al. Null mutants of Drosophila B-type lamin Dm(0) show aberrant tissue differentiation rather than obvious nuclear shape distortion or specific defects during cell proliferation. Developmental biology 284, 219–232, https://doi.org/10.1016/j.ydbio.2005.05.022 (2005).
Acknowledgements
We thank T. Schüpbach, M. Simonelig and the Bloomington Drosophila Stock Center for the fly stocks, M. Siomi and P. Fisher for antibodies, and Y. Shevelyov for helpful discussion on the manuscript. The work was carried out with the use of the equipment of the common use center «Center of Cell and Gene Technology», Institute of Molecular Genetics, RAS. This work was supported by Russian Science Foundation (RSF) [grant number 19-14-00382 to M.S.K.].
Author information
Authors and Affiliations
Contributions
M.K., O.S., V.G. and E.M. designed experiments; O.S., E.M., S.K., Y.A. and M.K. performed experiments; M.K. wrote the manuscript with support from V.G.; M.K. coordinated the project.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this Article was revised: The original version of this Article contained an error in the Acknowledgments section. “The work was supported by the grant from Russian Foundation for Basic Research [16-04-01524 for M.K.] and by the Presidium of the Russian Academy of Sciences program Molecular and Cell Biology (for V. G.).” Now reads: “This work was supported by the Russian Science Foundation (RSF) [grant number 19-14-00382 to M.S.K.].”
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sokolova, O.A., Mikhaleva, E.A., Kharitonov, S.L. et al. Special vulnerability of somatic niche cells to transposable element activation in Drosophila larval ovaries. Sci Rep 10, 1076 (2020). https://doi.org/10.1038/s41598-020-57901-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-020-57901-2
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.