Abstract
Soil nitrification via ammonia oxidation is a key ecosystem process in terrestrial environments, but little is known of how increasing irrigation of farmland soils with saline waters effects these processes. We investigated the effects of long-term irrigation with saline water on the abundances and community structures of ammonia-oxidizing bacteria (AOB) and archaea (AOA). Irrigation with brackish or saline water increased soil salinity (EC1:5) and NH4-N compared to irrigation with freshwater, while NO3-N, potential nitrification rates (PNR) and amoA gene copy numbers of AOA and AOB decreased markedly under irrigation regimes with saline waters. Moreover, irrigation with brackish water lowered AOA/AOB ratios. PNR was positively correlated with AOA and AOB amoA gene copy numbers across treatments. Saline and brackish water irrigation significantly increased the diversity of AOA, as noted by Shannon index values, while saline water irrigation markedly reduced AOB diversity. In addition, irrigation with brackish or fresh waters resulted in higher proportions of unclassified taxa in the AOB communities. However, irrigation with saline water led to higher proportions of unclassified taxa in the AOA communities along with the Candidatus Nitrosocaldus genus, as compared to soils irrigated with freshwater. AOA community structures were closely associated with soil salinity, NO3−N, and pH, while AOB communities were only significantly associated with NO3−N and pH. These results suggest that salinity was the dominant factor affecting the growth of ammonia-oxidizing microorganisms and community structure. These results can provide a scientific basis for further exploring the response mechanism of ammonia-oxidizing microorganisms and their roles in nitrogen transformation in alluvial grey desert soils of arid areas.
Similar content being viewed by others
Introduction
Nitrification in soils is a central component of N biogeochemical cycling and involves the oxidation of ammonia \(({{\rm{NH}}}_{4}^{+})\) to nitrite \(({{\rm{NO}}}_{2}^{-})\), along with further oxidation to nitrate \(({{\rm{NO}}}_{3}^{-})\). The process has significant environmental and agricultural impacts on N nutrient availability for plants, NO3−N leaching, and greenhouse gases emissions1,2,3. Specifically, ammonia oxidation and nitrite oxidation are the two key steps of nitrification. Ammonia oxidation is the first and rate-limiting step of nitrification, plays a major role in sustaining the global N cycle, and is accomplished primarily by the participation of ammonia oxidizing Archaea (AOA) and Bacteria (AOB)4,5. Both AOA and AOB have similar ammonia-monooxygenase (Amo) enzymatic pathways, and the amoA gene can be used as a useful marker to evaluate the distribution of these guilds. The extensive development of molecular biology in recent decades has led to an increasing number of studies that have investigated the ecology of AOA and AOB via amoA gene surveys. For example, such studies have evaluated the effects of different environmental factors on AOA and AOB abundances6,7, in addition to influences on their community structures8,9, as well as the relative contributions of AOA and AOB to nitrification10,11.
Irrigation is a critical agricultural practice that ensures crop yields. However, freshwater scarcity and high water salinity have become threats to sustainable agricultural development in many regions. Consequently, a greater reliance has been placed on brackish or saline waters for agricultural irrigation. However, saline or brackish waters can cause salt accumulation in soils and alter other soil physicochemical and biological properties12,13. These changes can then influence soil nitrification processes and the microorganisms involved in nitrification. Previous studies have shown that the inhibition of nitrification increased with increasing soil salt levels14 and that the abundances of AOA and AOB are negatively correlated with soil salinity15. In contrast, other studies have shown that increases in the potential nitrification rates of soils and the abundances of AOA increased under moderate salinity levels (10–20 ppt), while the abundances of AOB were either negatively correlated, or not correlated at all with increased soil salinity16,17. Moreover, Cui et al.1 reported that salinity inhibited the activity of AOB and decreased the number of dominant AOBs, while not markedly impacting the composition of dominant AOB species. Furthermore, Beman and Francis18 observed that AOA community structure varied among environments with different salinities, while no correlations were observed between AOB community diversity and salinity gradients. In addition, He et al.19 observed that salinity was significantly associated with variance in AOA and AOB community structures in wetland soils. Lastly, Cao et al.20 observed that AOB diversity increased with increased soil salinity, while Dang et al.21 observed the opposite result. Thus, salinity is a key factor affecting soil nitrification and has received widespread attention in recent years, but the impact of soil salt concentrations on the abundances and community structures of AOA and AOB is still unclear.
Ammonia-oxidizing bacteria and archaea coexist in soils, yet there is very little data regarding AOB and AOA community compositions and their relative contributions to ammonia oxidation responses to irrigation of alluvial gray desert soils with saline waters. We hypothesized that long-term (10-year) irrigation with saline waters will negatively influence the abundances and community structures of AOB and AOA. Specifically, we leveraged field experiments to investigate the effects of saline water irrigation on (i) the abundances and community structures of AOB and AOA, (ii) the relationships between soil properties and the abundances and community structures of AOB and AOA.
Results
Soil physicochemical properties
Increased irrigation water salinity was associated with significant increases (P < 0.001) in the EC1:5, SWC, and NH4-N contents of soils, but significant decreases (P < 0.001) in soil pH, SOC, TN, and NO3-N contents (Table 1). Soil NO3-N content ranged from 31.96 to 46.19 mg/kg, with the highest value in the FW treatment. NO3-N content was 13.5% and 30.8% lower in the BW and SW treatment soils compared to the FW treatment soils, respectively. Soil NH4-N content ranged from 6.82 to 7.86 mg/kg, with the lowest value observed in the FW treatment. NH4-N contents in the BW and SW treatment soils were 10.4% and 15.2% higher than in those of the FW treatment.
amoA gene abundances and the potential nitrification rate
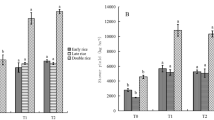
The BW and SW treatment soils had markedly lower amoA gene copy numbers belonging to AOA and AOB compared to the FW treatment (Fig. 1). Specifically, the amoA gene copy number of AOA in different treatments ranged from 2.2 × 106 and 3.6 × 106 copies/g dry soil (Fig. 1a), while those of AOB ranged between 1.9 × 105 and 3.2 × 105 copies/g dry soil (Fig. 1b). amoA gene copy numbers of AOA and AOB in soils of the BW and SW treatments were 28.4%/39.0% and 23.3%/38.4% lower than in those of the FW treatments, respectively.
The effects of irrigation with saline water on ammonia oxidizing microbial communities. Panels show the effects of freshwater (FW), brackish water (BW), and saline water (SW) on amoA gene copy numbers of AOA (a), amoA gene copy numbers of AOB (b), the AOA/AOB ratio (c), and the potential nitrification rate (d). Mean data are shown while error bars show standard deviations, n = 3. FW, BW, and SW correspond to waters with electrical conductivity (EC) of 0.35, 4.61, and 8.04 dS m−1, respectively. Different lowercase letters indicate statistically significant differences among water salinity treatments (P < 0.05).
The ratio of AOA to AOB in the BW treatment soils was significantly lower than in those of the FW and SW treatments, while no significant difference was observed in the soils of the FW and SW treatments (Fig. 1c). These results imply that brackish water increased the relative proportion of AOB in those soils. In addition, BW and SW irrigation treatments resulted in significantly lower soil PNR values (Fig. 1d). The PNR of the FW treatment soils were 18.1% and 37.3% higher than those in the BW and SW treatments, respectively.
Relative contributions of AOA and AOB to PNR
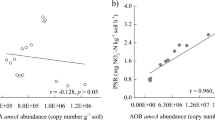
Regression analysis indicated that soil PNR was significantly and positively correlated with amoA gene copy numbers of AOA (R2 = 0.9228, P < 0.001) (Fig. 2). Similarly, PNR was significantly and positively correlated with amoA gene copy numbers of AOB (R2 = 0.9489, P < 0.001). Thus, variation in PNR was highly correlated with the abundances of both AOA and AOB.
Diversity of amoA genes within soils
The sequencing coverage of amoA gene libraries for AOA and AOB communities among all soil samples was greater than 99%, indicating that adequate sequencing depth was used to evaluate the native diversity in the soils (Table 2). The AOA and AOB sequences were clustered into 661–664 and 130–140 OTUs (defined at the 97% nucleotide similarity level), respectively. The number of AOB OTUs was significantly lower in the BW and SW irrigated soils relative to those of the FW treatment (P = 0.011), while no relationships for AOA OTUs among treatments was observed (P = 0.414). Richness indices (ACE and Chao1) were not significantly associated with salinity. However, the Shannon index of AOA diversity was significantly higher in the BW and SW treatment soils (P < 0.001) compared to those of the FW treatment, while the Simpson index showed a significant (P < 0.001) opposite relationship. In addition, the SW treatment soils had a significantly lower AOB Shannon index (P = 0.002) compared to the FW soils, but a significantly higher Simpson index value (P = 0.003). The BW treatment did not have a significant effect on AOB diversity index.
Association among AOA, AOB, and environmental parameters
The correlations among soil physicochemical properties with PNR, amoA gene copy number, and the diversity of AOA and AOB were evaluated (Table 3). Soil EC1:5, SWC, and NH4-N were negatively correlated with PNR, amoA gene copy numbers of AOA and AOB, and the Simpson index values of the AOA communities. However, soil EC1:5, SWC, and NH4-N were positively correlated with the Shannon index values of the AOA communities and the Simpson index of AOB communities. In addition, soil NO3-N, SOC, and TN were positively correlated with PNR, amoA gene copy numbers of AOA and AOB, the Simpson index values of AOA communities, and the Shannon index of AOB communities. However, soil NO3-N, SOC, and TN were negatively correlated with the Shannon index of AOA communities and the Simpson index of AOB communities. Soil pH was positively correlated with PNR, amoA gene copy numbers of AOA and AOB, and the Simpson index of AOA communities. However, soil pH was negatively correlated with the Shannon index values of AOA communities. Soil salinity was significantly and negatively correlated with the Shannon index values of AOB, while soil environmental factors were not significantly correlated with ACE and Chao1 indices.
Community structure
Irrigation with saline water also significantly affected the composition of AOA and AOB communities (Fig. 3). The Crenarchaeota,Proteobacteria, and Thaumarchaeota were the three most dominant phyla among the communities (Fig. 3a), with the Crenarchaeota comprising the largest portion of the communities. However, no significant differences were observed in the relative abundances of Crenarchaeota and Thaumarchaeota among the FW, BW, and SW treatments. The relative abundances of Proteobacteria were significantly greater in FW treatment soils than in BW and SW treatment soils (P < 0.001). The Candidatus Nitrosocaldus, Candidatus Nitrososphaera, Betaproteobacteria, and Marine archaeal group 1 lineage were the four most dominant classes, of which the relative abundances of Candidatus Nitrosocaldus were highest. The relative abundances of Candidatus Nitrosocaldus were significantly greater in the SW treatment soils than in FW and BW treatment soils (P < 0.001). In addition, the relative abundances of Betaproteobacteria were significantly lower in the BW and SW soils than in the FW soils (P = 0.003). No significant differences were observed in the relative abundances of Candidatus Nitrososphaera and Marine archaeal group 1 among the FW, BW, and the SW treatment soils (P > 0.05). The Nitrosomonadales and Nitrosopumilales were the two most dominant orders in the dataset. The relative abundances of Nitrosomonadales were significantly higher in the FW treatment soils than in the BW and SW treatment soils (P < 0.001). However, no significant differences were observed in the relative abundances of Nitrosopumilales among the FW, BW, and SW treatment soils. The Nitrosomonadaceae and Nitrosopumilaceae were the two most dominant families in the dataset. The relative abundances of Nitrosomonadaceae were significantly greater in the FW treatment soils than in those of the BW and SW treatments (P < 0.001). However, no significant differences were observed for the relative abundances of Nitrosopumilaceae among the FW, BW, and SW treatment soils. Nitrosopumilus and Nitrosospira were the two most dominant genera in the sequencing datasets. The relative abundances of Nitrosospira were significantly higher in the FW treatment soils than in those of the BW and SW treatments (P < 0.001). However, no significant differences were observed for the relative abundances of Nitrosopumilus among the FW, BW, and SW treatment soils. Lastly, irrigation with saline water was associated with higher abundances of unknown AOA taxa at all taxonomic levels compared to those soils irrigated with fresh water.
Effects of irrigating soils with varying salinities on the community structure of ammonia-oxidizing archaea (a) and ammonia-oxidizing bacteria (b). FW, BW, and SW treatments are defined as in Fig. 1.
The dominant AOB phyla, class, order, and family were the Proteobacteria, Betaproteobacteria, Nitrosomonadales,and Nitrosomonadaceae, respectively (Fig. 3b). The relative abundances of Proteobacteria, Betaproteobacteria, Nitrosomonadales, and Nitrosomonadaceae were significantly greater in the SW treatment soils than in the BW soils (P < 0.001). However, no significant differences were observed in the relative abundances of Proteobacteria, Betaproteobacteria, Nitrosomonadales, and Nitrosomonadaceae between the SW and FW treatment soils. The Nitrosospira, Nitrosomonas, and Nitrosovibrio were the three most dominant AOB genera, of which the Nitrosospira (53.96–61.15% relative abundances) were dominant. The relative abundance of Nitrosospira in the BW treatment soils was significantly lower than in the FW and SW soils (P < 0.05). The relative abundances of Nitrosomonas decreased with increasing irrigation water salinity, while no Nitrosomonas were detected in the SW soils. Moreover, there were no significant differences in the relative abundances of Nitrosovibrio among the FW, BW, and SW treatment soils (P > 0.05). Lastly, brackish water irrigation was associated with higher abundances of unknown taxonomic classes compared to AOB communities from soils irrigated with saline waters.
RDA analysis
The associations between environmental factors and AOA and AOB community structures were also analyzed using RDA (Fig. 4). Axes 1 and 2 explained 54.7% and 27.5% of the total variation in AOA community variation (Fig. 4a). AOA community structure was significantly associated with NO3-N contents (interpretation degree of 59.1%, P = 0.00), pH (interpretation degree of 23.2%, P = 0.032), and soil salinity (interpretation degree of 10.4%, P = 0.042). In the AOB analysis, axes 1 and 2 explained 57.5% and 31.2% of the total variation in community structure (Fig. 4b). AOB community structure was also significantly associated with NO3-N contents (interpretation degree of 33.3%, P = 0.04) and pH (interpretation degree of 47.7%, P = 0.012).
RDA analysis of the correlation among ammonia-oxidizing archaea (a) and ammonia-oxidizing bacteria (b) community structures with environmental factors. FW, BW, and SW treatments are defined as in Fig. 1. Arrows show statistically significant environmental parameters, wherein the direction of the arrow indicates the direction of highest correlation and the length of the arrow indicates the magnitude of the correlation. The percent of the variation explained by each axis is indicated in each axis title.
Discussion
Freshwater shortage is an important factor that limits the sustainable development of agriculture. Consequently, the rational use of saline water irrigation is crucial for alleviating freshwater shortages in arid areas22,23. However, the long-term use of saline or brackish water irrigation may cause salt accumulation in root zones and increase the risk of soil salinization, thereby altering soil physicochemical properties. These changes in turn affect nutrient cycling and transformations, and especially the critical ecosystem processes of nitrogen transformations24,25. Here, we observed that soil NH4-N content significantly increases with increased irrigation water salinity, while NO3-N content exhibited an opposite trend that might be attributed to inhibited soil nitrification due to increased soil salinity26. Moreover, long-term irrigation with brackish or saline waters significantly inhibited soil PNR. These results are consistent with those of He et al.27 that indicated that PNR markedly decreased with increased soil salinity. Together, these results suggest that increased salinity inhibits the critical microbial ecosystem processes of nitrification. It should be noted that other studies have shown that moderate salinity can increase soil PNR, while high salinity levels inhibit soil PNR28. These results could be explained by the specific salinity tolerances of microorganisms involved in nitrification, with the potential promotion of certain microorganisms involved in nitrification and consequent increases in soil PNR over certain salinity ranges29.
The first and rate limiting step of nitrification is ammonia oxidation that is primarily conducted by ammonia-oxidizing bacteria (AOB) and archaea (AOA)30. Higher AOA amoA gene copy numbers relative to those of AOB have been observed in a range of soil environments31,32,33,34,35. Indeed, AOA are considered to be the primary drivers of nitrification in many soil ecosystems36,37. Likewise, AOA amoA gene copy numbers were greater than those of AOB in this study. These results are in accordance with those of Bernhard et al.16 who observed that AOA amoA gene copy numbers were consistently higher than those of AOB along salinity gradients. Zhang et al.17 also observed that AOA exhibited higher growth and ammonia-oxidizing activity than AOB in moderate and high salinity environments. In contrast to the above studies, Magalhaes et al.38 observed that amoA gene copy numbers of AOB were always higher than those of AOA in sediments. The higher AOA amoA gene copy numbers observed in the soils evaluated here suggest that AOA may be the dominant ammonia-oxidizing guild in these environments. Furthermore, amoA gene copy numbers of AOA and AOB also significantly decreased with increasing irrigation water salinity. These results are consistent with those indicating that high salinity inhibits AOB growth39.
However, some studies have indicated that salinity is not significantly related to AOA amoA gene copy numbers40. Further, moderate salinity (10–30 ppt) may even stimulate AOA growth and AOB amoA copy numbers, as observed by insensitivity to soil salinity gradients17. Here, decreases in amoA abundances could indicate lower potentials for soil nitrification. Tao et al.11 found that soil PNR were positively correlated with AOB abundance but not AOA abundance, which suggest that AOB contribute more than AOA to nitrification in this fertilized calcareous field soil. Indeed, AOA and AOB amoA gene copy numbers were highly correlated with PNR. Thus, AOA and AOB dynamics likely explained the variation in PNR in the soils analyzed here. Moreover, the AOA/AOB ratio was significantly lower in soils with brackish water irrigation than in those with freshwater irrigation. The shift in ammonia oxidizing soil communities from low to high AOA/AOB indicates that the ammonia oxidizing community in these soils underwent selective pressure against AOB following saline water irrigation. Nevertheless, the AOA/AOB ratio alone is not sufficient to determine which of the two guilds are functionally dominant in ammonia oxidation activities10,41,42.
In salt-affected soils, salinity is the major factor that controls the diversity and distribution of AOB and AOA43. In this study, strong associations were observed between irrigation water salinity levels with AOA and AOB community structures. AOA alpha-diversity (i.e., the Shannon index) significantly increased with increased irrigation water salinity. However, the Shannon index of AOB communities was higher when fresh and brackish water was used for irrigation compared to irrigation with saline waters. These results suggest that AOB communities were more sensitive to changes in salinity than were AOA communities. Gao et al.44 also found that AOA community diversity increased as soil salinity increased, while AOB community diversity was highest in moderate or lower salinity soils. Alternatively, AOA communities can be adapted to low pH and low nutrient soil conditions34,45,46,47, while AOB are thought to preferentially inhabit soil with neutral pH and high-N agricultural soils10,44. Following these observations, Schauss et al.48 speculated that AOB mainly dominate “nutrient-rich” environments, while AOA are better adapted to “nutrient-poor” environments. In this study, saline water and brackish water irrigation changed the community structure of AOA and AOB. Fresh water irrigation was associated with higher abundances of the AOA taxa comprising the Proteobacteria phylum, the Betaproteobacteria class, the Nitrosomonadales order,the Nitrosomonadaceae family, and the Nitrosospira genus compared to soils irrigated with saline or brackish water. Among AOA classes, Candidatus Nitrosocaldus was the main taxonomic group associated with saline water irrigation, with significant increases in their relative abundances in the high salinity treatments. In addition, saline water irrigation led to higher abundances of AOB taxa comprising the Proteobacteria phylum, Betaproteobacteria class, Nitrosomonadales order, and Nitrosomonadaceae family, as compared with soils irrigated with brackish waters. In contrast, soils with brackish water irrigation contained higher levels of unclassified taxa compared to soils with saline water irrigation. Nitrosospira was the dominant AOB genus, with relative abundances higher in soils irrigated with saline water compared to those irrigated with brackish water. In contrast, the relative abundances of Nitrosomonas significantly decreased in soils irrigated with increasingly saline water. These results were consistent with those of Sahan and Muyzer43 wherein Nitrosospira were enriched in a high salinity environment, while Nitrosomonas was enriched in low or moderate salinity environments. Kowalchuk and Stephen49 also observed that Nitrosospira were more dominant components of AOB communities in terrestrial ecosystems than Nitrosomonas.
The contribution of AOB and AOA to potential nitrification in soils is highly dependent on the initial environments of soils11. After 10 years of irrigation with saline waters, the physical and chemical properties of the soils analyzed here significantly changed, thereby affecting the ammonia-oxidizing microbial communities. Irrigation with saline or brackish waters inhibited nitrification and significantly decreased soil mineral N concentrations (NH4+, NO3−). The significantly lower amoA copy numbers of AOA and AOB in the brackish and saline water treatments compared to fresh water treatments indicated that N inputs could be another key factor influencing AOA and AOB abundances. This supposition was further supported by the RDA analysis indicating that NO3-N accounted for 59.1% and 33.3% of the total variation in AOA (P = 0.002) and AOB community (P = 0.04) structures, respectively. Thus, NO3-N variation was one of the main factors affecting AOA and AOB community structures when irrigated with saline or brackish waters. NO3-N is closely related to AOA and AOB activities, since it is a product of nitrification50. amoA gene copy numbers were positively correlated with NO3-N concentrations, indicating that both AOA and AOB contributed to the whole ammonia oxidation process. In contrast, Tago et al.51 observed that soil NO3-N content was only significantly correlated with changes in AOB community structures, which could be due to different soil nutrient conditions compared to soils analyzed here. In addition, AOA actively grow in harsh environments (e.g., low nitrogen availability, strong acidity, and high temperatures), which in turn, leads to robust functional activities52. Soil pH is thought to be one of the most important factors affecting AOA and AOB abundances53. pH only differed by 0.2 units between fresh and saline water irrigation treatments, but was nevertheless associated with decreases in AOA and AOB abundances by 36.22% and 38.38%, respectively. These results agree with Hu et al.54 wherein the abundances of AOA and AOB were positively correlated with pH. In contrast, Nicol et al.34 found that AOB abundances significantly decreased with decreasing pH, while AOA exhibited an opposite relationship. Further, Xi et al.55 observed that a decrease in soil pH resulted in a significant increase in AOA abundances without affecting AOB abundances. The lack of major change in AOA or AOB abundances observed here could be due to the relatively small range of pH change in this study, with salinity having a more important effect. Nevertheless, RDA indicated that variation in soil pH alone explained a large component (interpretation degree of 47.7%, P = 0.012) in AOB community structure as well as AOA community structure (interpretation degree of 23.2%, P = 0.012). These results are consistent with those of Ying et al.56 wherein AOB community structure variation was associated with the combined effects of numerous environmental factors including soil pH and N availability, even when gradients were minimal. Together, these findings indicate that pH was likely an important factor associated with changes in the AOA and AOB structure in these alluvial gray desert soils.
Conclusion
These results indicate that irrigation with saline or brackish waters significantly reduced soil NO3-N contents and PNR, while soil PNR was significantly and positively correlated with amoA gene copy numbers of AOB and AOA. Long-term (10 years) irrigation with saline water significantly decreased the abundances of AOB and AOA and also altered the community composition of both AOB and AOA. Fresh water irrigation led to higher abundances of the AOA phylum Proteobacteria, the class Betaproteobacteria, the order Nitrosomonadales, the family Nitrosomonadaceae, and the genus Nitrosospira, as compared with soils irrigated with saline or brackish waters. However, irrigation with saline water led to higher abundances of unclassified archaeal taxa compared to irrigation with fresh water. For AOB, irrigation with brackish water led to higher abundances of unknown bacterial taxa compared to irrigation with saline waters. Irrigation with brackish and saline water significantly increased the Shannon diversity index of AOA communities, while saline water treatment significantly decreased the Shannon diversity index of AOB communities. Soil properties, including NO3-N, pH, and salinity, were significantly associated with variation in AOA community structure, while AOB community structure was only significantly correlated with NO3-N and pH. This information will aid in understanding and managing N transformations in agriculturally-productive alluvial grey desert soils.
Materials and Methods
Experiment site and experimental design
The long-term field experiment site was located at the Shihezi University Agricultural Experimental Station (N 44°18′, E 86°02′). The region is classified as a temperate arid zone with an average temperature of 7.8 °C and 168–171 frost-free days each year. There is annual average sunshine of 2,721–2,818 h, annual average evaporation of 1,660 mm, and annual average precipitation between 125–200 mm, without significant annual variation. The soil of the field site is an alluvial gray desert type (Calcaric Fluvisol in the FAO system). Physicochemical properties of soils at 0–30 cm depth prior to the experiment were: 0.13 dS/m EC1:5, pH1:2.5 7.90, 16.8 g/kg soil organic matter, 1.08 g/kg total N, 25.9 mg/kg available P, and 253 mg/kg available K.
The experiment was conducted from 2009 to 2018 using a completely randomized block design with three replicates of three irrigation treatments. The three treatments comprised (1) fresh water (FW) with irrigation water electrical conductivities (ECw) of 0.35 dS/m; (2) brackish water (BW) with ECw of 4.61 dS/m; and (3) saline water (SW) with ECw of 8.04 dS/m. FW was obtained from a well at the site, while saline water for the BW and SW treatments was prepared by adding equal amounts of NaCl and CaCl2 (1:1 mass ratio) to the well waters. The chemical compositions of the irrigation waters are shown in Table 4.
A traditionally farmed cotton cultivar (Gossypium hirsutum L. cv Xinluzao 52) was planted in the experiment soils. The plots were drip-irrigated and mulched with transparent polyethylene plastic films. One sheet of plastic film was laid side by side in each plot that was 18 m × 1.2 m. Each transparent polyethylene plastic film had four rows of cotton with two irrigation drip lines that were installed below the transparent polyethylene plastic film. A 30 cm strip of bare soil was maintained between each plot. Cotton plants were sown at 10 cm intervals within each row, with 30 cm spacing between the first and second rows, second and third rows, and third and fourth rows. Cotton was sown in mid-late April with a planting density of 222,000 plants/ha each year. The growing season of cotton ranged from April to September. Each plot was irrigated nine times at irrigation intervals of 7 to 10 d between June and August. A total 450 mm of water was used to irrigate during the cotton growing season each year. Before sowing, each plot was fertilized with 105 kg P2O5/ha and 60 kg K2O/ha. A total of 360 kg N/ha was applied via drip in five applications into each plot between June and August. The same cultivation techniques were used in each of the years.
Soil sampling
Soil samples were collected from the 0–20 cm layer from three random locations in each plot on July 28, 2018 during the tenth year of the long-term study. The soil samples were packed with ice packs and brought back to the laboratory. Soil samples were passed through a 2 mm sieve and then divided into two portions. One was used to determine the soil physico-chemical properties and potential nitrification rates (PNR), while the other was stored at −80 °C and used to analyze the abundances and diversity of ammonia-oxidizing microorganisms via molecular analyses.
Soil physico-chemical analyses
Soil water content (SWC) was determined by oven-drying the soil at 105 °C for 1 d. Soil salinity and pH were determined with an MP521 Lab pH/conductivity meter. The soil-to-water ratios used to measure electrical conductivity and pH were 1:5, and 1:2.5, respectiely. NH4-N and NO3-N contents were determined with a SmartChem140 discrete Analyzer (Westco Scientific, Danbury, Connecticut, USA), after extraction with 2 M KCl. Soil organic carbon (SOC) and total nitrogen (TN) were determined using the K2Cr2O7-H2SO4 oxidation-reduction titration method and the semi-micro Kjeldahl method, respectively.
Potential nitrification rates (PNR)
Soil PNRs were determined using methods described by Kurola et al.57. Briefly, 5 g of fresh soil was placed into 50 mL centrifuge tubes containing 20 mL of phosphate buffer saline solution with 1 mM (NH4)2SO4. To inhibit nitrite oxidation, potassium chlorate was placed in the centrifuge tubes at a final concentration of 10 mmol/l. After incubation for 24 h at room temperature in the dark, nitrite (NO2-N) was extracted with 5 mL of 2 M KCl and determined spectrophotometrically at 545 nm with N-(1-naphthyl) ethylenediamine dihydrochloride.
DNA extraction, qPCR assays, and high-throughput sequencing
Soil microbial DNA was extracted from 0.3 g of fresh soil samples using the Power Soil DNA Isolation Kit (Mo Bio Laboratories Inc., USA) according to manufacturer’s instructions. The concentration and purification of DNA were determined by UV-vis spectrophotometry, while the quality of extracted DNA was evaluated by 1% agarose gel electrophoresis. Extracted DNA was stored at −20 °C prior to further analysis.
AOA and AOB abundances were determined by real-time quantitative PCR assays using the LightCycler 480 SYBR Green I Master (Roche) by LightCycler 480 Real-Time PCR System (Roche). AOA amoA genes were amplified with the primers Arch-amoAF (5′-STAATGGTCTGGCTTAGACG-3′), and Arch-amoAR (5′-GCGGCCATCCATCTGTATGT-3′)58. AOB amoA genes were amplified with the primers amoA-1F (5′-GGGGTTTCTACTGGTGGT-3′), and amoA-2R (5′- CCCCTCKGSAAAGCCTTCTTC -3′)59. Each reaction mixture was performed in triplicate 20 μL volume reactions containing 10 μL of SYBR Green I Master Mix, 2 μL of DNA template, 1 μL of each primer and 6 μL ddH2O. The PCR reactions were conducted using the following program: 95 °C for 5 min; 40 cycles of 10 s at 95 °C, 55 °C for 20 s, and 72 °C for 30 s. amoA gene fragments were cloned into PMD-18 plasmids (Takara), followed by validation of correctly inserted gene fragments. Standard curves were generated over the range of 10−1–10−6 gene copies mL−1.
amoA gene amplicons were also analyzed using high-throughput sequencing at Beijing Biomarker Technology Co., Ltd. (Beijing, China). The PCR primers used for sequence analyses were identical to those of the qPCR assays. PCR amplifications were conducted in 25 µL reactions including 2 μL of DNA template, 1 μL of primers (10 µM stock concentrations) before and after amplification, 5 μL of 5X PCR buffer, 2 μL (2.5 mM stock) of dNTPs, 5 μL of 5X Q5High GC enhancer buffer, 0.25 μL (0.02 U/µL) of Q5 High-Fidelity DNA polymerase (NEB), and 8.75 μL of ddH2O. The thermal cycle reaction program for amoA amplification comprised: 5 min of initial denaturation at 98 °C, 35 cycles of 98 °C for 30 s, annealing at 55 °C for 30 s, and elongation at 72 °C for 45 s, followed by a final extension at 72 °C for 5 min. PCR amplicons were purified using Agencourt AMPure Beads (Beckman Coulter, Indianapolis, IN, USA), and quantified using the QuantiFluor-ST system (Promega, USA) according to manufacturer instructions. Equimolar amounts of samples were pooled before high-throughput sequencing on the Illumina MiSeq platform.
Data analysis
One-way analysis of variance (ANOVA) tests and Pearson correlation analyses were conducted using the SPSS statistical program (version SPSS 21.0). Tukey’s tests were used to evaluate differences in values among groups at an alpha level of P < 0.05. The sequence data were analyzed using QIIME, as described previously60. The diversity and richness indices were calculated using the Mothur software program (version 1.30.1). Visualization of taxonomic classifications and abundances was conducted in MEGAN. Redundancy analysis (RDA) was used to elucidate the soil properties that explained variation in community structures, as implemented in Canoco version 4.5.
Data availability
All data generated or analysed during this study are included in this published article (and its Supplementary Information files).
References
Cui, Y. W., Zhang, H. Y., Ding, J. R. & Peng, Y. Z. The effects of salinity on nitrification using halophilic nitrifiers in a Sequencing Batch Reactor treating hypersaline wastewater. Scientific reports 6, 24825 (2016).
Li, Y., Watanabe, T., Murase, J., Asakawa, S. & Kimura, M. Abundance and composition of ammonia oxidizers in response to degradation of root cap cells of rice in soil microcosms. Journal of soils and sediments 14(9), 1587–1598 (2014).
Zhalnina, K., de, Quadros, P. D., Camargo, F. A. & Triplett, E. W. Drivers of archaeal ammonia-oxidizing communities in soil. Frontiers in Microbiology 3, 210 (2012).
Martens-Habbena, W. et al. Ammonia oxidation kinetics determine niche separation of nitrifying archaea and bacteria. Nature 461(7266), 976–981 (2009).
Morimoto, S. et al. Quantitative analyses of ammonia-oxidizing archaea (AOA) and ammonia-oxidizing bacteria (AOB) in fields with different soil types. Microbes and environments 1105130304-1105130304. (2009).
Limpiyakorn, T., Sonthiphand, P., Rongsayamanont, C. & Polprasert, C. Abundance of amoA genes of ammonia-oxidizing archaea and bacteria in activated sludge of full-scale wastewater treatment plants. Bioresource technology 102(4), 3694–3701 (2011).
Szukics, U. et al. Management versus site effects on the abundance of nitrifiers and denitrifiers in European mountain grasslands. Science of the total environment 648, 745–753 (2019).
Francis, C. A., O’Mullan, G. D. & Ward, B. B. Diversity of ammonia monooxygenase (amoA) genes across environmental gradients in Chesapeake Bay sediments. Geobiology 1(2), 129–140 (2003).
Wang, M. et al. Newly designed primer pair revealed dominant and diverse comammox amoA gene in full-scale wastewater treatment plants. Bioresource technology 270, 580–587 (2018).
Di, H. J. et al. Nitrification driven by bacteria and not archaea in nitrogen-rich grassland soils. Nature Geoscience 2(9), 621 (2009).
Tao, R., Wakelin, S. A., Liang, Y. & Chu, G. Response of ammonia-oxidizing archaea and bacteria in calcareous soil to mineral and organic fertilizer application and their relative contribution to nitrification. Soil Biology and Biochemistry 114, 20–30 (2017).
Singh, K. Microbial and enzyme activities of saline and sodic soils. Land Degradation and Development 27(3), 706–718 (2016).
Min., W. et al. Irrigation water salinity and N fertilization: Effects on ammonia oxidizer abundance, enzyme activity and cotton growth in a drip irrigated cotton field. Journal of integrative agriculture 15(5), 1121–1131 (2016).
Akhtar, M. et al. Influence of Salinity on Nitrogen Transformations in Soil. Communications in soil science and plant analysis 43(12), 1674–1683 (2012).
Li, X. R. et al. Abundance and composition of ammonia-oxidizing bacteria and archaea in different types of soil in the Yangtze River estuary. Journal of Zhejiang University SCIENCE B 13(10), 769–782 (2012).
Bernhard, A. E. et al. Abundance of ammonia-oxidizing archaea and bacteria along an estuarine salinity gradient in relation to potential nitrification rates. Applied and Environmental Microbiology 76(4), 1285–1289 (2010).
Zhang, Y. et al. The influence of salinity on the abundance, transcriptional activity, and diversity of AOA and AOB in an estuarine sediment: a microcosm study. Applied microbiology and biotechnology 99(22), 9825–9833 (2015).
Beman, J. M. & Francis, C. A. Diversity of ammonia-oxidizing archaea and bacteria in the sediments of a hypernutrified subtropical estuary: Bahia del Tobari, Mexico. Applied and Environmental Microbiology 72(12), 7767–7777 (2006).
He, Y. et al. Abundance and diversity of ammonia-oxidizing archaea and bacteria in the rhizosphere soil of three plants in the Ebinur Lake wetland. Canadian journal of microbiology 63(7), 573–582 (2017).
Cao, H., Hong, Y., Li, M. & Gu, J. D. Community shift of ammonia-oxidizing bacteria along an anthropogenic pollution gradient from the Pearl River Delta to the South China Sea. Applied microbiology and biotechnology 94(1), 247–259 (2012).
Dang, H. et al. Diversity, abundance, and spatial distribution of sediment ammonia-oxidizing Betaproteobacteria in response to environmental gradients and coastal eutrophication in Jiaozhou Bay, China. Applied and Environmental Microbiology 76(14), 4691–4702 (2010).
Wang, Q. et al. Impact of saline water irrigation on water use efficiency and soil salt accumulation for spring maize in arid regions of China. Agricultural Water Management 163, 125–138 (2016).
Yuan, C., Feng, S., Huo, Z. & Ji, Q. Effects of deficit irrigation with saline water on soil water-salt distribution and water use efficiency of maize for seed production in arid Northwest China. Agricultural water management 212, 424–432 (2019).
Pereira, L. S., Oweis, T. & Zairi, A. Irrigation management under water scarcity. Agricultural Water Management 57, 175–206 (2016).
Ma, T. et al. Effects of water, salt and nitrogen stress on sunflower (Helianthus annuus L.) at different growth stages. Journal of soil science and plant nutrition 16(4), 1024–1037 (2016).
Bernhard, A. E. & Bollmann, A. Estuarine nitrifiers: new players, patterns and processes. Estuarine, coastal and shelf science 88(1), 1–11 (2010).
He, H. et al. Ammonia-oxidizing Archaea and Bacteria differentially contribute to ammonia oxidation in sediments from adjacent waters of Rushan Bay, China. Frontiers in Microbiology 9, 116 (2018).
Bernhard, A. E. et al. Functionally distinct communities of ammonia-oxidizing bacteria along an estuarine salinity gradient. Environmental Microbiology 9(6), 1439–1447 (2007).
Duan, M. et al. Contrasting responses of gross and net nitrogen transformations to salinity in a reclaimed boreal forest soil. Biology and Fertility of Soils 54(3), 385–395 (2018).
Gleeson, D. B. et al. Response of ammonia oxidizing archaea and bacteria to changing water filled pore space. Soil Biology and Biochemistry 42, 1888–1891 (2010).
Adair, K. L. & Schwartz, E. Evidence that ammonia-oxidizing archaea are more abundant than ammonia-oxidizing bacteria in semiarid soils of northern Arizona, USA. Microbial Ecology 56(3), 420–426 (2008).
Fan, F. L. et al. Impacts of organic and inorganic fertilizers on nitrification in a cold climate soil are linked to the bacterial ammonia oxidizer community. Microbial Ecology 62, 982e990 (2011).
He, J. Z. et al. Quantitative analyses of the abundance and composition of ammonia-oxidizing bacteria and ammonia-oxidizing archaea of a Chinese upland red soil under long-term fertilization practices. Environmental Microbiology 9, 2364e2374 (2006).
Nicol, G. W., Leininger, S., Schleper, C. & Prosser, J. I. The influence of soil pH on the diversity, abundance and transcriptional activity of ammonia oxidizing archaea and bacteria. Environmental microbiology 10(11), 2966–2978 (2008).
Trias, R. et al. Emergent macrophytes act selectively on ammonia-oxidizing bacteria and archaea. Applied and environmental microbiology 78(17), 6352–6356 (2012).
Leininger, S. et al. Archaea predominate among ammoniaoxidizing prokaryotes in soils. Nature 442, 806e809 (2006).
O’Sullivan, C. A., Wakelin, S. A., Fillery, I. R. P. & Roper, M. M. Factors affecting ammonia-oxidising microorganisms and potential nitrification rates in southern Australian agricultural soils. Soil Research 51, 240e252 (2013).
Magalhães, C. M., Machado, A. & Bordalo, A. A. Temporal variability in the abundance of ammonia-oxidizing bacteria vs. archaea in sandy sediments of the Douro River estuary, Portugal. Aquatic Microbial Ecology 56(1), 13–23 (2009).
Jin, T. et al. Diversity and quantity of ammonia-oxidizing Archaea and Bacteria in sediment of the Pearl River Estuary, China. Applied microbiology and biotechnology 90(3), 1137–1145 (2011).
Wang, H., Gilbert, J. A., Zhu, Y. & Yang, X. Salinity is a key factor driving the nitrogen cycling in the mangrove sediment. Science of The Total Environment 631, 1342–1349 (2018).
Jia, Z. J. & Conrad, R. Bacteria rather than archaea dominate microbial ammonia oxidation in an agricultural soil. Environmental Microbiology 11, 1658–1671 (2009).
Wakelin, S. et al. Predicting the efficacy of the nitrification inhibitor dicyandiamide in pastoral soils. Plant and Soil 381, 35–43 (2014).
Sahan, E. & Muyzer, G. Diversity and spatio-temporal distribution of ammonia-oxidizing Archaea and Bacteria in sediments of the Westerschelde estuary. FEMS microbiology ecology 64(2), 175–186 (2008).
Gao, J. et al. Shifts in the Community Dynamics and Activity of Ammonia-Oxidizing Prokaryotes Along the Yangtze Estuarine Salinity Gradient. Journal of Geophysical Research: Biogeosciences 123(11), 3458–3469 (2018).
Di, H. J. et al. Ammonia-oxidizing bacteria and archaea grow under contrasting soil nitrogen conditions. FEMS Microbiology Ecology 72, 386–394 (2010).
Auguet, J. C. et al. Vertical segregation and phylogenetic characterization of ammoniaoxidizing Archaea ina deep oligotrophic lake. The ISME Journal 6, 1786–1797 (2012).
Lu, L. & Jia, Z. Urease gene-containing Archaea dominate autotrophic ammonia oxidation in two acid soils. Environmental Microbiology 15, 1795–1809 (2013).
Schauss, K. et al. Dynamics and functional relevance of ammonia-oxidizing archaea in two agricultural soils. Environmental Microbiology 11, 446–456 (2009).
Kowalchuk, G. A. & Stephen, J. R. Ammonia-oxidizing bacteria: a model for molecular microbial ecology. Annual Reviews in Microbiology 55(1), 485–529 (2001).
Yin, X. et al. Microbial community of nitrogen cycle-related genes in aquatic plant rhizospheres of Lake Liangzi in winter. Journal of basic microbiology 58(11), 998–1006 (2018).
Tago, K. et al. Environmental factors shaping the community structure of ammonia-oxidizing bacteria and archaea in sugarcane field soil. Microbes and environments ME14137 (2014).
Schleper, C. Ammonia oxidation: different niches for bacteria and archaea? The ISME journal 4(9), 1092 (2010).
Gubry-Rangin, C. et al. Niche specialization of terrestrial archaeal ammonia oxidizers. Proceedings of the National Academy of Sciences of the United States of America 108, 21206–21211 (2011).
Hu, H. et al. The large-scale distribution of ammonia oxidizers in paddy soils is driven by soil pH, geographic distance, and climatic factors. Frontiers in microbiology 6, 938 (2015).
Xi, R., Long, X. E., Huang, S. & Yao, H. pH rather than nitrification and urease inhibitors determines the community of ammonia oxidizers in a vegetable soil. AMB Express 7(1), 129 (2017).
Ying, J. et al. Contrasting effects of nitrogen forms and soil pH on ammonia oxidizing microorganisms and their responses to long-term nitrogen fertilization in a typical steppe ecosystem. Soil Biology and Biochemistry 107, 10–18 (2017).
Kurola., J. et al. Activity, diversity and population size of ammonia-oxidising bacteria in oil-contaminated landfarming soil. FEMS microbiology letters 250(1), 33–38 (2005).
Francis, C. A. et al. Ubiquity and diversity of ammonia-oxidizing archaea in water columns and sediments of the ocean. Proceedings of the National Academy of Sciences of the United States of America 102(41), 14683–14688 (2005).
Park, S. J., Park, B. J. & Rhee, S. K. Comparative analysis of archaeal 16SrRNA and amoA genes to estimate the abundance and diversity of ammonia-oxidizing archaea in marine sediments. Extremophiles 12(4), 605–615 (2008).
Caporaso, J. G. et al. QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7(5), 335–336 (2010).
Acknowledgements
This work was jointly funded by The National Natural Science Foundation of China [41661055] and the Youth Innovation Talent Cultivation Program of Shihezi University [CXRC201706].
Author information
Authors and Affiliations
Contributions
Huijuan Guo wrote the main manuscript. Huijuan Guo and Lijuan Ma performed the statistical analysis. Zhenan Hou and Wei Min conceived of this study. Zhenan Hou, Yongchao Liang and Wei Min participated in the design and helped to draft the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Guo, H., Ma, L., Liang, Y. et al. Response of ammonia-oxidizing Bacteria and Archaea to long-term saline water irrigation in alluvial grey desert soils. Sci Rep 10, 489 (2020). https://doi.org/10.1038/s41598-019-57402-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-57402-x
This article is cited by
-
Microbes in a neutral-alkaline paddy soil react differentially to intact and acid washed biochar
Journal of Soils and Sediments (2022)
-
Ammonia Monooxygenase Activity Connects Nitrification Rate with Dominant Edaphic Properties Under Salinity Stress in Coastal Fluvo-aquic Soil
Journal of Soil Science and Plant Nutrition (2022)
-
Saline and alkaline stresses alter soil properties and composition and structure of gene-based nitrifier and denitrifier communities in a calcareous desert soil
BMC Microbiology (2021)
-
Impact of irrigation with fish-processing effluents on nitrification and ammonia-oxidizer abundances in Patagonian arid soils
Archives of Microbiology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.