Abstract
The co-occurrence of Geobacter and Methanosarcinales is often used as a proxy for the manifestation of direct interspecies electron transfer (DIET) in the environment. Here we tested eleven new co-culture combinations between methanogens and electrogens. Previously, only the most electrogenic Geobacter paired by DIET with Methanosarcinales methanogens, namely G. metallireducens and G. hydrogenophilus. Here we provide additional support, and show that five additional Methanosarcinales paired with G. metallireducens, while a strict hydrogenotroph could not. We also show that G. hydrogenophilus, which is incapable to grow with a strict hydrogenotrophic methanogen, could pair with a strict non-hydrogenotrophic Methanosarcinales. Likewise, an electrogen outside the Geobacter cluster (Rhodoferrax ferrireducens) paired with Methanosarcinales but not with strict hydrogenotrophic methanogens. The ability to interact with electrogens appears to be conserved among Methanosarcinales, the only methanogens with c-type cytochromes, including multihemes (MHC). Nonetheless, MHC, which are often linked to extracellular electron transfer, were neither unique nor universal to Methanosarcinales and only two of seven Methanosarcinales tested had MHC. Of these two, one strain had an MHC-deletion knockout available, which we hereby show is still capable to retrieve extracellular electrons from G. metallireducens or an electrode suggesting an MHC-independent strategy for extracellular electron uptake.
Similar content being viewed by others
Introduction
Direct interspecies electron transfer (DIET) was discovered in an artificial co-culture of an ethanol-oxidizing Geobacter metallireducens with a fumarate-reducing Geobacter sulfurreducens where the possibility of hydrogen gas (H2) or formate transfer was invalidated through genetic studies1,2. Gene deletions rendering H2 and formate transfer impossible resulted in active co-cultures1,2, whereas deletion of genes for extracellular electron transfer proteins (EET) such as pili, and multiheme c-type cytochromes made the interspecies interaction impossible1,3. Remarkably, a deletion mutant lacking an outer membrane multiheme c-type cytochrome (OmcS) could be rescued by the addition of extracellular conductive particles, whereas a pili knock-out strain could not be rescued by conductive particles4. Additionally, previous studies have shown that G. metallireducens is also highly effective as an anode-respiring bacteria (ARB) generating some of the highest current densities of all Geobacter tested5. During DIET with G. sulfurreducens, G. metallireducens requires pili and certain multiheme c-type cytochromes3, which were also required during anode respiration6,7.
G. metallireducens is a strict respiratory microorganism unable to ferment its substrates to produce H2 for interspecies H2-transfer8,9. Consequently, G. metallireducens could not provide reducing equivalents for strict hydrogenotrophic methanogens (Methanospirilum hungatei and Methanobacterium formicicum)10,11. However, G. metallireducens did interact syntrophically with Methanosarcinales (Methanosarcina barkeri 800, Methanosarcina horonobensis, Methanothrix harundinacea)10,11,12 of which the last two are unable to consume H213,14. When incubated with Methanosarcinales, G. metallireducens upregulated EET-proteins, and if some of these EET proteins were deleted, co-cultures became inviable demonstrating the direct electron transfer nature of the interaction10,12.
For co-culture incubations, G. metallireducens has been typically provided with ethanol as electron donor, but without a soluble electron acceptor. In the absence of a soluble electron acceptor, G. metallireducens releases electrons extracellularly (reaction 1) and uses the methanogen as its extracellular terminal electron acceptor (reaction 2 & 3). If electrons released by G. metallireducens cannot reach a terminal electron acceptor, than ethanol oxidation would stop, because the electron transport chain would become ineffective, and NADH produced during ethanol oxidation could not get re-oxidized.
Reaction 1. Ethanol oxidation and electron release by G. metallireducens:
Reaction 2. Electron uptake coupled with CO2 reductive methanogenesis by Methanosarcinales:
Reaction 3. Acetoclastic methanogenesis:
Reaction 4. Total DIET reaction by a syntrophic association between Geobacter and Methanosarcinales:
There are indications for direct interspecies electron transfer occuring in methane producing environments such as anaerobic digesters15, rice paddy soils16, and aquatic sediments17,18, as well as in methane consuming environments such as hydrothermal vents19,20. In these environments, DIET is typically inferred either because conductive materials stimulate the syntrophic metabolism but also by the co-presence of DNA and/or RNA of phylotypes related to DIET-microorganisms. DIET-pairing of Geobacter with methanogens was only described in two Geobacter species (G. metallireducens, G. hydrogenophilus)5,10,11,12. On the other hand, a series of six Geobacter species were unable to interact syntrophically with Ms. barkeri 800. All these Geobacter were modest anode respiring bacteria and did not produce high current densities at the anode5. Species outside of the Geobacter-clade have never been shown to do DIET with methanogens.
In this study, we expand the list of syntrophic-DIET pairs and investigated whether electrogenic bacteria other than Geobacter, could interact syntrophically with methanogens. We determined whether DIET was widespread among methanogens or was a specific trait of Methanosarcinales because of their high c-type cytochrome content21. Multiheme c-type cytochromes (MHC) were previously implicated in EET in bacteria22 and an MHC of Methanosarcina acetivorans was required for anthraquinone-2, 6-disulfonate (AQDS) respiration23. Here we asked whether MHC in Methanosarcina are required for DIET and electron uptake from electrodes.
Materials and Methods
Microorganisms and cultivation conditions
Cultures were purchased from the Leibniz Institute DSMZ-German Collection of Microorganisms and Cell Cultures GmbH (DSMZ) and grown on the media advised by the collection until pre-adaption to co-cultivation media.
Electrogens
We used the following strains of bacteria: Rhodoferax ferrireducens (DSM 15236), Pelobacter carbinolicus (DSM 2380), Geobacter hydrogenophilus (DSM 13691) and Geobacter metallireducens GS-15 (DSM 7210) available at the University of Massachusetts. For experiments with G. metallireducens strain GS15 at the University of Southern Denmark GS-15 (DSM 7210) was purchased from the DSMZ culture collection. G. metallireducens, G. hydrogenophilus and R. ferrireducens were maintained on the typical co-culture freshwater media10 with 55 mM ferric citrate as electron acceptor. The two Geobacter species were maintained with 10–20 mM ethanol as electron donor11,12 whereas Rhodoferax was maintained with 5 mM glucose. P. carbinolicus which was used as H2-donating strain (DIET-negative control), was pre-cultivated under fermentative conditions on 10 mM acetoin as previously described2.
Methanogens
The following strains of methanogens were tested: Methanoculleus marisnigri JR1 (DSM 1498), Methanospirillum hungatei JF1 (DSM 864), Methanosaeta harundinacea 8Ac (DSM 17206), Methanothrix soehngenii (DSMZ 2139), Methanosarcina mazei Gö1 (DSM 3647) and three strains of Methanosarcina barkeri MS (DSM 800), Fusaro (DSM 804), and 227 (DSM 1538).
Methanothrix soehengii, Methanosaeta harundinacea, Methanospirilum hungatei and Methanoculleus marisnigri were pre-grown on the typical co-culture freshwater media however without ethanol10 and instead containing their respective substrate 20–80 mM acetate for Methanosaeta/Methanothrix and H2:CO2 (80:20) for Methanoculleus and Methanospirillum.
Prior to co-cultivation, all Methanosarcina cultures were pre-grown on their respective substrates (20–30 mM acetate and/or 20 mM methanol) in a modified DSMZ 120c media in which tryptone was omitted and NaCl was reduced down to 1 g/L11,12. M. mazei (wild type and 633 k.o.) and M. barkeri 227 were pregrown with acetate and methanol, whereas M. barkeri 804 was pregrown with acetate.
A mutant strain of Methanosarcina mazei lacking the gene MM_0633 encoding for the multi-heme cytochrome c family protein (mutant name - M. mazei 633 k.o.) was kindly provided by Prof. Uwe Deppenmeier and Prof. Cornelia Welte. This mutant strain has been characterized in detail in Christian Krätzer’s Ph.D. thesis24. For the multiheme cytochrome mutant of M. mazei (633 k.o.) an antibiotic mix (ampicillin 100 µg/ml and puromycin 5 µg/ml) was added for selection and safe maintanance. This antibiotic mix was omitted in the transfer prior to co-culture inoculation and in the co-culture media.
Co-cultures
Co-cultures were set up without a methanogenic electron donor. As such, Methanosarcina co-cultures were prepared in modified DSMZ 120c media10,11,12, while co-cultures with Methanothrix/Methanosaeta or strict H2-utilizing methanogens (Methanospirillum and Methanoculleus) in modified freshwater media10. When the electron donating partner was G. metallireducens, G. hydrogenophilus or P. carbinolicus, 10–20 mM ethanol was provided as the sole substrate, whereas for co-cultures with R. ferrireducens, 5 mM glucose was provided as substrate.
Co-cultures with electrically conductive granular activated carbon (GAC)
For experiments with conductive materials, we added 25 g/L GAC (charcoal activated, Merck kGaA, Darmstadt, Germany) to the co-culture media and controls were run in parallel. No-food controls were set up to monitor for background product formation from carry-over substrates or the possible use of GAC as a food source. Single-species culture controls with and without the addition of GAC to assess the probability of ethanol oxidation using GAC as electron acceptor by a single species.
All co-cultures were incubated at 37 °C at a final volume of 10 ml and all co-cultures were set up in duplicates or more replicates.
Analytical measurements
Samples were withdrawn anaerobically using N2:CO2 (80:20) flushed hypodermic needles to verify the levels of methane, hydrogen, acetate, ethanol, and glucose. Determination of methane (CH4), hydrogen gas (H2), ethanol and volatile fatty acids (e.g. acetate, formate) was carried out as described before11,12. Briefly, for experiments done at the University of Southern Denmark, gases were measured with a Trace 1300 gas chromatograph (GC) (Thermo-Scientific) equiped with a TracePLOT™ TG-BOND Msieve 5A column and a thermal conductivity detector (TCD). The carrier gas was argon at a flow rate of 25 mL/min with the temperatures set for the injector, oven and detector at 150 °C, 70 °C and 200 °C, respectively. To monitor ethanol, the same GC system was used with a different column, TRACE™ TR-Wax and a flame ionization detector (FID). The filter-sterilised liquid sample (0.5 mL) was first heated to 60 °C for 5 mins in an air-tight exetainer after which the vaporised sample was collected for measurement. The carrier gas was nitrogen flowing at 1 mL/min, with the temperatures of injector, oven and detector set at 220 °C, 40 °C and 230 °C respectively. Acetate was analysed with a Dionex™ ICS-1500 Ion Chromatography system, using a Dionex™ IonPac™ AS15 IC Column. The eluent was a mixture of 1.4 mM NaHCO3 and 4.5 mM Na2CO3 detected on an electron capture detector (ECD) at 30 mA. For experiments done at the University of Massachusetts, gases were measured by a Shimadzu 8 A gas chromatograph with a 80/100 Hayasep Q column connected to an FID. The temperatures of injector, oven and detector were set at 200 °C, 120 °C and 200 °C respectively. Ethanol was measured on a Perkin Elmer GC equiped with an Elite 5 column with helium as the carrier gas following a gradient separation protocol as follows: 50 °C for 1 min, a increase of 12 °C per minute to reach 200 °C, and a final 1.5 min at 200 °C. The temperatures for injector and detector were set at 200 °C and 300 °C, respectively.
Acetate was measured on a high-pressure liquid chromatography (HPLC) via an Aminex NPX-87H column using 8 mM H2SO4 as the eluent with a UV detector set at 210 nm. Glucose was determined at the end of the incubation, as previously described25 by separating on an HPLC with an Aminex Ion-Exclusion organic acid analysis column HPX-87H (Bio-Rad) and detected with a SP8430 refractive-index detector (Spectra-Physics).
Electrochemical reactor setup
Incubations using a cathode as sole electron donor, were carried out as previously described11, with slight modifications. Briefly, a two-chambered H cell reactor (Adams and Chittenden, USA) with a total volume of 650 ml in each chamber was separated via a Nafion™ N117 proton exchange membrane (Ion power). The working and counter electrodes were made of graphite rods with dimensions of 2.5 × 7.5 × 1.2 cm and were connected to titanium rods. A leak-free Ag/AgCl reference electrode (3.4 M KCl) (CMA Microdialysis, Sweden). To ensure low carry over of substrates, cells were harvested in an anaerobic chamber at 4000 rpm for 10 mins, resuspended in fresh media (modified 120c – see above) prior to inoculation of the cell suspension at a final concentration of 20% in the reactor. In this set-up, the working and counter electrodes were connected via a resistor (250 Ω) and a potentiostat, however the resistor was unecessary since the cathodic potential was fully controlled by the potentiostat. The potential of working electrode was controlled with the MultiEmstat potentiostat (Palmsens, The Netherlands) set at −400 mV (vs. the standard hydrogen electrode).
Results and Discussion
Electrogens establish DIET syntrophy with Methanosarcinales methanogens
To examine the ability for direct interspecies electron uptake in new strains of methanogens, G. metallireducens was used as the default DIET-partner for co-culture experiments due to its high electrogenic capability5, correlated to a syntrophic ability10,12, which was independent of H2-transfer, as this Geobacter was incapable to produce H29,26. We selected seven methanogenic strains as representatives of two groups: strict hydrogenotrophic methanogens (H2-consuming) and Methanosarcinales-methanogens, including H2-utilizing and strict non-hydrogenotrophic strains.
G. metallireducens paired syntrophically with five new Methanosarcinales, independent of the methanogen’s ability to consume H2
The electrogen Geobacter metallireducens was previously shown to establish DIET interactions with three species of the order Methanosarcinales of which two were non-hydrogenotrophic methanogens (Methanosarcina horonobensis and Methanosaeta harundinacea) and one was a H2-utilizing methanogen (Methanosarcina barkeri strain MS)10,11,12. Nevertheless, we recently reported that of two Methanosarcina species which are capable of DIET with G. metallireducens only one could use a cathode at −400 mV (vs. SHE)11 as sole electron donor. This variability in the Methanosarcinales capability to carry out direct electron uptake prompted us to verify whether DIET is conserved in other Methanosarcinales.
In this study, we evaluated five aditional strains of Methanosarcina, two of M. barkeri (M. barkeri 227 and M. barkeri strain Fusaro/804) two of M. mazei (Go1 and 633 k.o.) and one of the strict non-hydrogenotrophic methanogen - Methanothrix soehngenii.
All co-cultures were provided with ethanol as electron donor in the absence of any electron acceptor other than carbon dioxide. If successful, co-cultures were anticipated to reach mid exponential growth after circa two months, in agreement to preliminary tests and previous reports on DIET-consortia10,11,12. After 75 days, all co-cultures of G. metallireducens with the five new Methanosarcinales effectively converted their substrate (ethanol) to products (methane) (Fig. 1). The respiratory metabolism of G. metallireducens (see reaction 1) resulted in extracellular transfer of electrons and transient formation of acetate (Fig. 2). The products of ethanol oxidation were then converted into methane (reaction 2 & 3; Fig. 2). Alone, none of the five Methanosarcinales converted ethanol to methane (Fig. 2).
Overview of co-culture tests with 14 species of methanogens and various electroactive bacteria. As electron donors we used 5 mM glucose for tests with Rhodoferax; 20 mM ethanol for the rest of the co-cultures; except for tests with M. mazei strains, M. horonobensis and M. barkeri 227, which were provided with 10 mM ethanol.
G. metallireducens co-cultivated on 10 mM ethanol together with (a) M. barkeri 227 and (b) M. mazei with or without electrically conductive particles of granular activated carbon (GAC). Control incubations of M. barkeri 227 (c) and M. mazei (d) on 10 mM ethanol as sole electron donor, with or without GAC. Changes in ethanol, acetate and methane concentrations are shown for triplicate incubations (n = 3). Whiskers represent standard deviations of the replicate incubations, and if invisible they are smaller than the symbol.
Previously, it was shown that interactions between G. metallireducens and M. barkeri strain MS (DSM 800) could be initiated much faster and continue at a faster pace, when amended with electrically conductive materials such as granular activated carbon (GAC)12,27 whereas non-conductive materials such as cotton cloth had no effect28. GAC was shown to promote the respiratory metabolism of G. metallireducens, which oxidizes ethanol and releases electrons extracellularly onto this conductor until it reaches charge-saturation29,30. Afterwards, the presence of a methanogen reclaiming electrons keeps the process from coming to a halt11.
In this study, we subjected three additional co-cultures of G. metallireducens and Methanosarcina (two strains of M. mazei and one M. barkeri 227) to 25 g/L GAC to verify whether GAC can accelerate their growth. All co-cultures amended with this electrically conductive material exhibited shorter lag-phases, reached mid-exponential growth much quicker, and at a minimum tripled their methanogenesis rates (Fig. 2). On the other hand, the addition of electrically conductive particles to the three Methanosarcina-strains in pure culture had no impact on methane production or ethanol utilization (Fig. 2).
We have now expanded the repertoire of strains of methanogens capable of DIET with G. metallireducens by an additional five strains adding to the original three strains described in the past. Of these eight strains, three were not hydrogenotrophic. We could conclude that the capacity to carry out DIET with G. metallireducens was conserved among all tested Methanosarcinales, independent of their ability to use H2 or not.
G. metallireducens was unsuccessful to pair syntrophically with a strict hydrogenotroph - Methanoculleus marisnigri
Although G. metallireducens does not have the genetic possibility to form H226, and could not evolve H2 when grown in pure culture on its own substrate9, we nevertheless previously tested whether H2-utilizing methanogens may have discovered a strategy to interact with this electrogenic Geobacter. In these former investigations we have shown that G. metallireducens was unsuccessful in developing ethanol-utilizing consortia in co-culture with two strict hydrogen-utilizing methanogens Methanospirillum hungatei (order Methanomicrobiales) and Methanobacterium formicicum (order Methanobacteriales)10, even in the presence of electrically conductive materials11.
In this study, we evaluated DIET between G. metallireducens with a third strict hydrogenotrophic species: Methanoculleus marisnigri (order Methanomicrobiales). Order Methanomicrobiales comprises the families Methanomicrobiaceae and Methanospirillaceae. Members of the family Methanomicrobiaceae (includins Methanoculleus marisnigri), were high in transcript abundance (13%) in rice paddies co-dominated by transcripts of Geobacter16, whereas Methanospirillaceae were not. This was hinting at the possibility of an interaction between Geobacter and Methanomicrobiaceae. We incubated G. metallireducens with Mcl. marisnigri for ca. 5 months, but did not observe methane production from ethanol during this timeframe (Fig. 1).
DIET evaluation by co-cultivation with G. metallireducens has now been carried out for three strict hydrogenotrophs from two major orders: Methanomicrobiales (M. hungatei, M. marisnigri) and Methanobacteriales (M. formicicum), showcasing the inability of strict hydrogenotrophic methanogens to pair with G. metallireducens (Fig. 1).
G. hydrogenophilus paired syntrophically with a second Methanosarcinales
Previously, we demonstrated that a second Geobacter species (G. hydrogenophilus) paired syntrophically with M. barkeri strain MS (DSM800). This was the only other Geobacter out of seven strains tested which exhibited the highest current density on the anode, similar to G. metallireducens5. Unlike G. metallireducens, G. hydrogenophilus does produce H2 during respiration9, but could not pair with the strict H2-utilizing methanogen – Methanospirillum hungatei5.
G. hydrogenophilus was only tested with strains that did have the possibility to use H2 and it was never tested whether it can pair syntrophically with strict non-hydrogenotrophic methanogens. In this study, we tested whether G. hydrogenophilus can pair syntrophically with the non-hydrogenotrophic methanogen Methanosaeta harundinacea (order Methanosarcinales). Methane production and acetate accumulation from ethanol were used as a proxy for the efficiency of the interaction. Previously, we have shown that acetate and methane can only accumulate in successfully paired co-cultures. For example, control tests with Methanosaeta could not sustain methane production and acetate accumulation from ethanol in the absence of a functional ethanol-oxidizing Geobacter10. In this study, we show that over the course of 117 days, co-cultures of G. hydrogenophilus and Methanosaeta harundinacea accumulated 5-times more methane and 4-times more acetate than a control culture of Methanosaeta on ethanol run in paralel (Fig. 3). However, a small amount of methane (ca. 1 mM) was produced by Methanosaeta in the control culture likely due to cells being transferred together with traces of acetate (an effective substrate for this methanogen). This was consistent with previous observations10.
G. hydrogenophilus in co-culture with the non-hydrogenotrophic methanogen Methanosaeta harundinacea (a) produced acetate and methane when provided with 20 mM ethanol (n = 2). Electron recoveries from day 32 to day 177, showed circa half recovery of the electrons from ethanol as acetate and the other half as methane. 1 mM methane and 1 mM acetate contain 8 mM electron equivalents (eeq) according to the stoichiometry of the following reactions: CH4 + 2H2O → CO2 + [8e− + 8H+]; CH3COOH + 2H2O → 2CO2 + [8e− + 8H+]. (b) Methane and acetate in a parallel control experiment with Methanosaeta harundinacea provided with 20 mM ethanol (n = 1). Carry over acetate of ca. 1.5 mM is typical for Methanosaeta. Here we verified whether there was an effective increase in acetate due to ethanol fermentation, above the typical methane production from acetate carry over. Whiskers represent standard deviations of the replicate incubations, and if invisible they are smaller than the symbol.
Moreover, the co-culture of G. hydrogenophilus and Methanosaeta harundinacea produced 9-times more methane than co-cultures with Methanospirillum run in parallel (Fig. 1, p = 0.03). Methanospirrilum was previously shown not to pair successfully with Geobacter hydrogenophilus5. Here we have shown additionally that that G. hydrogenophilus favors pairing with a strict non-hydrogenotrophic methanogen (Methanosaeta harundinacea) over a strict hydrogenotroph (Methanospirillum hungatei). This supports the notion that while G. hydrogenophilus can produce H29 it is an ineffective H2 donor, unlike the H2-donating syntroph - P. carbinolicus12 (Fig. 1).
Rhodoferrax ferrireducens paired syntrophically with Methanosarcinales but not with strict hydrogenotrophic methanogens
With only two out of seven Geobacter species showing aptitude for DIET with Methanosarcinales, we verified whether other highly effective electrogens outside the Geobacter clade could pair syntrophically via DIET. We tested this possibility for an effective anode respiring Betaproteobacteria - Rhodoferax ferrireducens25. Interestingly, Rhodoferax was predicted to outcompete Geobacter in a subsurface environment with low substrate flux and relatively high ammonia31, where non-hydrogenotrophic Methanosarcina species co-exists32. The co-existence of Rhodoferax, Geobacter and Methanosarcina was also noted in coastal Baltic Sea sediments32. It is possible Methanosarcina species could receive DIET-electrons from Rhodoferax as well as Geobacter. Here we examined whether R. ferrireducens could establish interspecies electron transfer in co-cultures with 2 DIET methanogens (M. harundinacea, M. barkeri) or with 2 strict hydrogenotrophic methanogens (M. hungatei and M. formicicum). We expected that this efficient anode-respiring bacterium25 would prefer DIET syntrophic partners to H2-utilizing partners. R. ferrireducens cannot utilize ethanol, therefore these co-cultures were provided with glucose (5 mM) as sole electron donor33. On glucose, Rhodoferax acts a respiratory organism and could not oxidize this substrate in the absence of an electron acceptor34. In these co-cultures, methane was used as a proxy for syntrophic metabolism. In order to estimate electron recovery from glucose, volatile fatty acid accumulation was determined during stationary phase. All co-cultures consumed the 5 mM glucose added (<4 µM detected after 270 days). Product recoveries varied significantly in Rhodoferax co-cultures with DIET-methanogens versus co-cultures with hydrogenotrophic methanogens (Fig. 4 and 4-inset). By comparing methane production in co-cultures with Rhodoferax, it was evident that M. harundinacea was the most effective at accumulating methane followed by Ms. barkeri and then strict hydrogenotrophs (Fig. 4). Rhodoferax in co-culture with Methanothrix had the highest total electron recovery (45%) with all electrons being recovered as methane, and none as acetate (Fig. 4-inset). The electrons not accounted for in products, are likely assimilated into biomass, typical of methanogenic metabolisms35. The Rhodoferax co-cultured with Methanosarcina was 3-fold less effective at recovering electrons into products (14%; Fig. 4-inset), yet the majority of the electrons were recovered as methane (12%) and only traces as acetate (2%). On the other hand, both co-cultures of Rhodoferax with strict hydrogenotrophs resulted in low electron recoveries as methane (ca. 5%) and more as acetate (18%) indicating hydrogenotrophic methanogens do not thrive in partnership with Rhodoferax. These results indicate that Rhodoferax favors interactions with DIET-methanogens rather than strict hydrogenotrophic methanogens. However, we do not know how Rhodoferax releases electrons to DIET methanogenic partners, although hints about its EET metabolism have been projected from genome screening34,36. The genome of R. ferrireducens contains 45 putative c-type cytochromes34 and the entire Mtr-pathway suggesting R. ferrireducens may be doing EET similar to Shewanella36,37. It remains to be tested whether this pathway is also used for DIET syntrophy with methanogens.
Co-cultures of four species of methanogens with Rhodoferax ferrireducens. Methane profiles in co-cultures provided with 5 mM glucose as sole electron donor. (inset) Estimated electron recoveries into products (acetate and methane). Incubations were carried out in triplicate (n = 3). Whiskers represent standard deviations of the replicate incubations, and if invisible, they are smaller than the symbol.
Multiheme c-type cytochromes were not required for DIET or cathodic EET
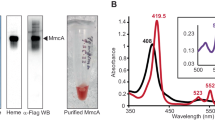
Many studies have shown that multi-heme c-type cytochromes (MHC) are important for extracellular electron transfer38 including EET during DIET1,3,12,19,39. However, MHCs are neither ubiquitous nor restricted to DIET-methanogens (Fig. 5). Of the hydrogenotrophic methanogens tested, M. hungatei and M. marisnigri contained potential multiheme cytochrome proteins (Fig. 5), apparently localized in the cytoplasm (Fig. 5). Of the DIET-methanogens, 5 out of 7 species did not contain MHCs including all of the Methanosarcina barkeri strains and both Methanosaetaceae species (M. harundinacea and M. soehngenii). M. horonobensis and M. mazei were the only 2 species with MHC apparently localized on the membrane or secreted (Fig. 5). M. mazei’s predicted MHC was detected within the surface/membrane-bound fraction by biochemical testing and predicted to be secreted extracellularly through leaderless secretion40 where it is suggested to join a membrane-bound complex containing flavoproteins and iron-sulfur flavoproteins41.
Predicted c-type cytochromes in methanogens tested for DIET. c-type cytochrome heme biding sites in 10 species of methanogens as predicted by CxxCH motif and/or annotated, this includes multiheme cytochromes; the predicted localization of the multiheme c-type cytochromes according to various bioinformatics tools. PSORTb - subcellular localization prediction tool58; TMHMM - prediction of transmembrane helices in proteins59; SOSUI- classification and secondary structure prediction system for membrane proteins60; SignalP - location of signal peptide cleavage sites, which is Archaea specific61; Pred-Signal - prediction of signal peptides in Archaea62; TatP - prediction of Twin-arginine signal peptide cleavage sites63; SecretomeP - prediction of non-classical protein secretion64.
Our hypothesis was that if M. mazei required its MHC for extracellular electron transfer, cells without it would be unable to interact with a DIET syntroph or with a poised electrode. This approach was previously used to determine Geobacter’s necessity for cell surface MHCs during EET to electrodes42, iron-oxide minerals22 and DIET-partners1,3,12.
To test this hypothesis, we used a knock-out mutant of M. mazei (633k.o.) in which the gene (MM_0633) encoding for the putative multiheme c-type cytochrome was deleted24. The deletion mutant showed no phenotypic variability to the wild type when growing on its typical substrates (methanol and acetate)24. To determine whether this MHC was required to receive DIET electrons from Geobacter, M. mazei 633k.o. was incubated with G. metallireducens in syntrophic media with ethanol (Fig. 6). Methane production and ethanol oxidation progressed similar to wild type control incubations (Fig. 6) demonstrating that this MHC is not required for DIET.
A genetically manipulated Methanosarcina mazei (strain 633k.o.), lacking the gene encoding a multiheme c-type cytochrome (MM_633), was tested for ethanol metabolism as single species (a) and in co-culture (b) with G. metallireducens. Incubations were carried out with or without the addition of electrically conductive GAC. The whiskers represent standard deviations of triplicate incubations (n = 3) and if invisible, they are smaller than the symbol.
Recently, it was reported that M. mazei was not electroactive and incapable to retrieve electrons from a poised cathode at −700 mV (vs. SHE)43. However, these experiments were carried out under conditional typically associated with electrochemical H2-generation44, and for a time frame of only 3 days, which is too short to test for the formation of electrical contacts for direct electromethanogenesis11,45. In fact, the authors observed only the growth of strict hydrogenotrophic methanogens on cathodes poised at −700 mV43. Strict hydrogenotrophic methanogens have low H2-thresholds (ca. 6 nM)46, unlike M. mazei, which grows poorly on H2, and has a high H2-threshold (ca. 300 nM) like all other Methanosarcinales46. Unlike the study by Meyer et al.43, our experiements were run at −400 mV vs. SHE, condition that does not allow for abiotic electrochemical H2 accumulation even after several months11. In a recent study, we showed that a cathode at −400 mV could not be used as sole electron donor by H2-utilizing methanogens like Methanobacterium formicicum, but it was used by a strain capable of direct electron uptake (via DIET) - Methanosarcina barkeri strain MS11.
Here we tested whether another Methanosarcina, M. mazei could carry out direct electron uptake from a cathode at −400 mV. We compared a wild type M. mazei and an MHC deletion mutant of M. mazei (633k.o.) to see whether the absence of its one and only MHC impacts EET from a cathode. Both M. mazei strains with and without the multiheme cytochrome were incubated with a cathode poised at a voltage of −400 mV (vs. SHE), unfavorable for the H2-evolution reaction11. Control experiments were run alongside, without applying a voltage at the cathode, to verify whether methanogenesis can be induced by carry-over substrates. Only in experiments with a poised cathode, methane production proceeded effectively for 633k.o. and wild type M. mazei (Fig. 7), showing that the MHC is not required for electron uptake from a cathode. Previously, we observed that M. horonobensis which contains the highest number of MHCs among DIET-methanogens was also unable to use a cathode as electron donor11. Combined, these results disprove the hypothesis that Methanosarcina species require a multiheme c-type cytochrome for extracellular electron uptake.
For Methanosarcinales involved in EET/DIET, the first barrier for electrons to enter a cell is the cell envelope. Thus, for DIET to take place, the cell surface of Methanosarcinales is anticipated to harbor charge transferring molecules. Methanogens exhibit very different cell envelopes and among the methanogens examined in this paper, as many as five types of distinct cellular surface composition have been observed (Fig. 8).
Representative cell envelope structures of DIET-methanogens (order Methanosarcinales) in contrast with hydrogen transfer (HT)-methanogens (Methanobacteriales; Methanomicrobiales). Abbreviations: DIET (direct interspecies electron transfer); HT (interspecies hydrogen transfer); MC (methanochondroitin); SL (S-layer); CM (cell membrane); protein sheath (PS); protein sheath amyloid type (PSa); protein sheet S-layer type (PSb); GP (glycosylated protein); PM (pseudomurein).
DIET-associations have now been demonstrated strictly with methanogens of the order Methanosarcinales including members of the families Methanosarcinaceae (Methanosarcina-genus) and Methanosaetaceae (Methanothrix and Methanosaeta). What distinguishes the Methanosarcina genus from all other methanogens is a thick methanochondroitin sulfate (MS) layer (Fig. 8), which is steadily represented on cell surfaces grown at low osmolarity47. Until now, DIET has been only demonstrated under freshwater conditions when Methanosarcina cells would be coated by methanochodroitin sulfate (MCS). MCS is an exopolysacharide resembling chondroitin sulfate in eukaryotes48 where it confers conduction via axonal length49. It is typical of exopolysaccharides like MCS to absorb metals50 or even trap redox cofactors and c-type cytochromes51, thus the embedded redox centers within the surface matrix may confer very different electric properties. What distinguishes the cell surface of Methanosaeta/Methanothrix genus from other methanogens is that only a protein sheet is delineating the cell surface from the environment52 (Fig. 8). The protein sheet was recently described in Methanosaeta thermophila to be composed of amyloid proteins53. Amyloid proteins are known to cluster together, while binding peptides54, and concentrating metal ions55. For example, the protein sheet of Methanothrix shoeghenii was described to concentrate metal ions like iron, copper, nickel and zinc56.
The surface structures of DIET methanogens of the order Methanosarcinales, although different in structure, have a shared attribute in binding/traping metal-ions to the cell surface, which we hypothesize to play a role in extracellular electron uptake by these Archaea.
Of the strict H2-utilizing methanogens, we observed that members of Methanomicrobiales and one Methanobacteriales were incapable to interact by DIET with electrogens. Typically, the membrane of these two groups are delineated from the environment by one single layer made either of pseudomurein (Methanobacteriales) or glycosylated S-layer proteins (Methanomicrobiales)57. Methanospirillum which is coated by an S-layer protein sheath was often compared to the protein sheet of Methanothrix (Fig. 8). However, the S-layer protein sheath of Methanospirillum traps less metal ions (2–5 fold less) than that of Methanothrix56.
These differences in surface biology may provide DIET-methanogens with a specific advantage to retrieve electrons from the extracellular environment and consequently an ecological niche where they outcompete H2-utilizers (e.g. mineral rich environments).
Conclusion
The incidence of Geobacter in methanogenic environments is often used as signature for direct interspecies electron transfer. However, only two (G. metallireducens, G. hydrogenophilus), which were the most electroactive species out of seven Geobacter species tested were previously shown to pair succesfully with Methanosarcinales. On the other hand these two Geobacter were previously shown to be unable to pair with methanogens of the orders Methanobacteriales (Methanobacterium formicicum) and a Methanomicrobiales (Methanospirillum hungatei). Here, we have expanded the list of DIET partnerships between Geobacter and Methanosarcinales by six additional combinations; and confirm using another Methanomicrobiales (Methanoculleus marisnigri) that these electrogenic Geobacter cannot pair with a strict hydrogenotrophic methanogens. Additionally, we show that DIET may favor another effective electrogen outside the Geobacter-cluster - Rhodoferrax ferrireducens. We observed that similar to the two electrogenic Geobacter, Rhodoferax ferrireducens promoted methanogenesis with Methanothrix and Methanosarcina and not with two strict hydrogenotrophic methanogens of the orders Methanobacteriales and Methanomicrobiales.
All Methanosarcinales tested formed metabolically active DIET consortia with G. metallireducens. Similar to other microorganisms involved in extracellular electron transfer, Methanosarcinales were anticipated to require MHCs. Here we documented that MHCs are infrequent within DIET-capable Methanosarcinales and unrestricted to DIET-methanogens. Moreover, a deletion of the MHC in one of the two MHC-containing Methanosarcinales (M. mazei) did not impact its ability to retrieve extracellular electrons from a DIET-partner or from an electrode. These data confirmed that extracellular electron uptake in Methanosarcina did not require MHCs, prompting a quest for cell surface constituents required for EET.
References
Summers, Z. M. et al. Direct exchange of electrons within aggregates of an evolved syntrophic coculture of anaerobic bacteria. Science 330, 1413–1415 (2010).
Rotaru, A.-E. et al. Interspecies electron transfer via hydrogen and formate rather than direct electrical connections in cocultures of Pelobacter carbinolicus and Geobacter sulfurreducens. Appl. Environ. Microbiol. 78, 7645–51 (2012).
Shrestha, P. M. et al. Transcriptomic and genetic analysis of direct interspecies electron transfer. Appl. Environ. Microbiol. 79, 2397–404 (2013).
Liu, F. et al. Magnetite compensates for the lack of a pilin-associated c-type cytochrome in extracellular electron exchange. Environ. Microbiol. 17, 648–55 (2015).
Rotaru, A.-E., Woodard, T. L., Nevin, K. P. & Lovley, D. R. Link between capacity for current production and syntrophic growth in Geobacter species. Front. Microbiol. 6, 744 (2015).
Tremblay, P.-L. L., Aklujkar, M., Leang, C., Nevin, K. P. & Lovley, D. A genetic system for Geobacter metallireducens: role of the flagellin and pilin in the reduction of Fe(III) oxide. Environ. Microbiol. Rep. 4, 82–88 (2012).
Gregory, K. B., Bond, D. R. & Lovley, D. R. Graphite electrodes as electron donors for anaerobic respiration. Environ. Microbiol. 6, 596–604 (2004).
Aklujkar, M. et al. The genome sequence of Geobacter metallireducens: features of metabolism, physiology and regulation common and dissimilar to Geobacter sulfurreducens. BMC Microbiol. 9, 1–22 (2009).
Cord-Ruwisch, R., Lovley, D. R. & Schink, B. Growth of Geobacter sulfurreducens with acetate in syntrophic cooperation with hydrogen-oxidizing anaerobic partners. Appl. Environ. Microbiol 64, 2232–2236 (1998).
Rotaru, A. E. et al. A new model for electron flow during anaerobic digestion: direct interspecies electron transfer to Methanosaeta for the reduction of carbon dioxide to methane. Energy Environ. Sci. 7, 408–415 (2014).
Yee, M. O., Snoeyenbos-West, O. L., Thamdrup, B., Ottosen, L. D. M. & Rotaru, A.-E. Extracellular electron uptake by two Methanosarcina species. Front. Energy Res 7, 458091 (2019).
Rotaru, A. E. et al. Direct interspecies electron transfer between Geobacter metallireducens and Methanosarcina barkeri. Appl. Environ. Microbiol. 80, 4599–4605 (2014).
Shimizu, S., Upadhye, R., Ishijima, Y. & Naganuma, T. Methanosarcina horonobensis sp. nov., a methanogenic archaeon isolated from a deep subsurface miocene formation. Int. J. Syst. Evol. Microbiol. 61, 2503–2507 (2011).
Ma, K., Liu, X. & Dong, X. Methanosaeta harundinacea sp. nov., a novel acetate-scavenging methanogen isolated from a UASB reactor. Int. J. Syst. Evol. Microbiol. 56, 127–131 (2006).
Dubé, C. D. & Guiot, S. R. Direct interspecies electron transfer in anaerobic digestion: a review. Biogas Sci. Technol. 151, 101–115 (2015).
Holmes, D. E. et al. Metatranscriptomic evidence for direct interspecies electron transfer between Geobacter and Methanothrix species in methanogenic rice paddy soils. Appl. Environ. Microbiol. 83, AEM.00223–17 (2017).
Rotaru, A.-E. et al. Conductive particles enable syntrophic acetate oxidation between Geobacter and Methanosarcina from coastal sediments. mBio 49, 1–14 (2018).
He, S., Lau, M. P., Linz, A. M., Roden, E. E. & McMahon, K. D. Extracellular electron transfer may be an overlooked contribution to pelagic respiration in humic-rich freshwater lakes. mSphere 4, 1–8 (2019).
McGlynn, S. E., Chadwick, G. L., Kempes, C. P. & Orphan, V. J. Single cell activity reveals direct electron transfer in methanotrophic consortia. Nature 526, 531–535 (2015).
Wegener, G., Krukenberg, V., Riedel, D., Tegetmeyer, H. E. & Boetius, A. Intercellular wiring enables electron transfer between methanotrophic archaea and bacteria. Nature 526, 587–590 (2015).
Thauer, R. K., Kaster, A.-K., Seedorf, H., Buckel, W. & Hedderich, R. Methanogenic archaea: ecologically relevant differences in energy conservation. Nat. Rev. Microbiol. 6, 579–91 (2008).
Smith, J. A., Lovley, D. R. & Tremblay, P. L. Outer cell surface components essential for Fe(III) oxide reduction by Geobacter metallireducens. Appl. Environ. Microbiol. 79, 901–907 (2013).
Holmes, D. E. et al. A Membrane-bound cytochrome enables Methanosarcina acetivorans to conserve energy from extracellular electron transfer. mBio 10, 1–12 (2019).
Kratzer, C. Substratumsetzung und Schutz vor Sauerstoffradikalen in Methanosarcina mazei. Hss.Ulb.Uni-Bonn.De (2011).
Chaudhuri, S. K. & Lovley, D. R. Electricity generation by direct oxidation of glucose in mediatorless microbial fuel cells. Nat. Biotechnol. 21, 1229–1232 (2003).
Shrestha, P. M. et al. Syntrophic growth with direct interspecies electron transfer as the primary mechanism for energy exchange. Environ. Microbiol. Rep 5, 904–10 (2013).
Liu, F. et al. Promoting direct interspecies electron transfer with activated carbon. Energy Environ. Sci. 5, 8982 (2012).
Chen, S. et al. Carbon cloth stimulates direct interspecies electron transfer in syntrophic co-cultures. Bioresour. Technol 173, 82–6 (2014).
Van Der Zee, F. P., Bisschops, I. A. E., Lettinga, G. & Field, J. A. Activated carbon as an electron acceptor and redox mediator during the anaerobic biotransformation of azo dyes. Environ. Sci. Technol. 37, 402–408 (2003).
Zhang, Y. et al. Progress of electrochemical capacitor electrode materials: A review. Int. J. Hydrogen Energy 34, 4889–4899 (2009).
Zhuang, K. et al. Genome-scale dynamic modeling of the competition between Rhodoferax and Geobacter in anoxic subsurface environments. ISME J. 5, 305–316 (2011).
Holmes, D. E. et al. Potential for Methanosarcina to contribute to uranium reduction during acetate-promoted groundwater bioremediation. Front. Microbiol. 660–667, https://doi.org/10.1007/s00248-018-1165-5 (2018).
Finneran, K. T., Johnsen, C. V. & Lovley, D. R. Rhodoferax ferrireducens sp. nov., a psychrotolerant, facultatively anaerobic bacterium that oxidizes acetate with the reduction of Fe(III). Int. J. Syst. Evol. Microbiol. 53, 669–673 (2003).
Risso, C. et al. Genome-scale comparison and constraint-based metabolic reconstruction of the facultative anaerobic Fe(III)-reducer Rhodoferax ferrireducens. BMC Genomics 10, 447 (2009).
Buan, N. R. Methanogens: pushing the boundaries of biology. Emerg. Top. Life Sci 2, 629–646 (2018).
Shi, L., Rosso, K. M., Zachara, J. M. & Fredrickson, J. K. Mtr extracellular electron-transfer pathways in Fe(III)-reducing or Fe(II)-oxidizing bacteria: a genomic perspective. Biochem. Soc. Trans 40, 1261–1267 (2012).
Coursolle, D., Baron, D. B., Bond, D. R. & Gralnick, J. A. The Mtr respiratory pathway is essential for reducing flavins and electrodes in Shewanella oneidensis. J. Bacteriol. 192, 467–474 (2010).
Shi, L. et al. Extracellular electron transfer mechanisms between microorganisms and minerals. Nat. Publ. Gr 14, 651–662 (2016).
Skennerton, C. T. et al. Methane-fueled syntrophy through extracellular electron transfer: Uncovering the genomic traits conserved within diverse bacterial partners of anaerobic methanotrophic archaea. MBio 8 (2017).
Leon, D. R. et al. Mining proteomic data to expose protein modifications in Methanosarcina mazei strain Gö1. Front. Microbiol. 6, 149 (2015).
Hovey, R. et al. DNA microarray analysis of Methanosarcina mazei Go1 reveals adaptation to different methanogenic substrates. Mol. Genet. Genomics 273, 225–239 (2005).
Holmes, D. E. et al. Microarray and genetic analysis of electron transfer to electrodes in Geobacter sulfurreducens. Environ. Microbiol. 8, 1805–1815 (2006).
Mayer, F., Enzmann, F., Lopez, A. M. & Holtmann, D. Performance of different methanogenic species for the microbial electrosynthesis of methane from carbon dioxide. Bioresour. Technol 289, 121706 (2019).
Yee, M. O., Snoeyenbos-West, O., Thamdrup, B., Ottosen, L. D. M. & Rotaru, A.-E. Extracellular electron uptake by two Methanosarcina species. Front. Energy Res. 7, 1–10 (2019).
Beese-Vasbender, P. F., Grote, J.-P., Garrelfs, J., Stratmann, M. & Mayrhofer, K. J. J. Selective microbial electrosynthesis of methane by a pure culture of a marine lithoautotrophic archaeon. Bioelectrochemistry 102, 50–55 (2015).
Lovley, D. R. Minimum threshold for hydrogen metabolism in methanogenic bacteria. Appl. Environ. Microbiol 49, 1530–1 (1985).
Sowers, K. R., Boone, J. E. & Gunsalus, R. P. Disaggregation of Methanosarcina spp. and growth as single cells at elevated osmolarity. Appl. Environ. Microbiol. 59, 3832–3839 (1993).
Kreisl, P. & Kandler, O. Chemical structure of the cell wall polymer of Methanosarcina. Syst. Appl. Microbiol. 7, 293–299 (1986).
Hunanyan, A. S. et al. Role of chondroitin sulfate proteoglycans in axonal conduction in mammalian spinal cord. J. Neurosci. 30, 7761–9 (2010).
van Hullebusch, E. D., Zandvoort, M. H. & Lens, P. N. L. Metal immobilisation by biofilms: Mechanisms and analytical tools. Rev. Environ. Sci. Bio/Technology 2, 9–33 (2003).
Rollefson, J. B., Stephen, C. S., Tien, M. & Bond, D. R. Identification of an extracellular polysaccharide network essential for cytochrome anchoring and biofilm formation in Geobacter sulfurreducens. J. Bacteriol. 193, 1023–33 (2011).
Claus, H. & Konig, H. Prokaryotic Cell Wall Compounds: Structure and Biochemistry, https://doi.org/10.1007/978-3-642-05062-6 (Springer Berlin Heidelberg, 2010).
Dueholm, M. S. et al. The tubular sheaths encasing Methanosaeta thermophila filaments are functional amyloids. J. Biol. Chem. 290, 20590–20600 (2015).
Maji, S. K. et al. Functional amyloids as natural storage of peptide hormones in pituitary secretory granules. Science 325, 328–332 (2009).
Viles, J. H. Metal ions and amyloid fiber formation in neurodegenerative diseases. Copper, zinc and iron in Alzheimer’s, Parkinson’s and prion diseases. Coord. Chem. Rev. 256, 2271–2284 (2012).
Patel, G. B., Sprott, G. D., Humphrey, R. W. & Beveridge, T. J. Comparative analyses of the sheath structures of Methanothrix concilii GP6 and Methanospirillum hungatei strains GP1 and JF1. Can. J. Microbiol. 32, 623–631 (2010).
Meyer, B. H. & Albers, S.-V. Archaeal cell walls. eLS 1–13, https://doi.org/10.1002/9780470015902.a0000384.pub2 (2014).
Yu, N. Y. et al. PSORTb 3.0: Improved protein subcellular localization prediction with refined localization subcategories and predictive capabilities for all prokaryotes. Bioinformatics 26, 1608–1615 (2010).
Krogh, A., Larsson, B., Von Heijne, G. & Sonnhammer, E. L. L. Predicting transmembrane protein topology with a hidden Markov model: Application to complete genomes. J. Mol. Biol. 305, 567–580 (2001).
Hirokawa, T., Boon-Chieng, S. & Mitaku, S. SOSUI: classification and secondary structure prediction system for membrane proteins. Bioinformatics 14, 378–379 (1998).
Nielsen, H. Predicting secretory proteins with SignalP. Protein Function Prediction in Methods in Molecular Biology . 59–73 (Humana Press, New York, 2017).
Bagos, P. G., Nikolaou, E. P., Liakopoulos, T. D. & Tsirigos, K. D. Combined prediction of Tat and Sec signal peptides with hidden Markov models. Bioinformatics 26, 2811–2817 (2010).
Bendtsen, J. D., Nielsen, H., Widdick, D., Palmer, T. & Brunak, S. Prediction of twin-arginine signal peptides. BMC Bioinformatics 6, 1–9 (2005).
Bendtsen, J. D., Kiemer, L., Fausbøll, A. & Brunak, S. Non-classical protein secretion in bacteria. BMC Microbiol. 5, 1–13 (2005).
Acknowledgements
This work was a contribution to a grant from the Innovationsfonden Denmark awarded to Lars Ottosen at Aarhus University (4106–00017). We would like to thank Lasse Ørum-Smidt and Trevor Woodard for lab assistance, and Prof. Lars Ottosen for support. We would like to thank Prof. Derek Lovley for reading and commenting on the manuscript.
Author information
Authors and Affiliations
Contributions
A.E.R. and M.O.Y. designed the experiments, A.E.R. carried out some of the co-culture incubations including analytical experiments at the University of Massachusetts, M.O.Y. carried out additional co-culture incubations and analytical experiments at the University of Southern Denmark, did the bioinformatic analyses and tested M. mazei strains in co-cultures and in a bioelectrochemical set-up. Both authors wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Yee, M.O., Rotaru, AE. Extracellular electron uptake in Methanosarcinales is independent of multiheme c-type cytochromes. Sci Rep 10, 372 (2020). https://doi.org/10.1038/s41598-019-57206-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-57206-z
This article is cited by
-
MmcA is an electron conduit that facilitates both intracellular and extracellular electron transport in Methanosarcina acetivorans
Nature Communications (2024)
-
Direct interspecies electron transfer for environmental treatment and chemical electrosynthesis: A review
Environmental Chemistry Letters (2024)
-
Mechanisms of microbial co-aggregation in mixed anaerobic cultures
Applied Microbiology and Biotechnology (2024)
-
Viruses interact with hosts that span distantly related microbial domains in dense hydrothermal mats
Nature Microbiology (2023)
-
Humic acid-dependent respiratory growth of Methanosarcina acetivorans involves pyrroloquinoline quinone
The ISME Journal (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.