Abstract
We investigated the relationship between germline single nucleotide polymorphisms (SNPs) in Von Hippel-Lindau (VHL) and Hypoxia-inducible factor 1-alpha (HIF1A), and their gene-environment and gene-gene interactions, and clear-cell RCC (ccRCC) risk. Furthermore, we assessed the relationship between VHL SNPs and VHL promoter methylation. Three VHL polymorphisms and one HIF1A polymorphism were genotyped in the Netherlands Cohort Study. In 1986, 120,852 participants aged 55–69 completed a self-administered questionnaire on diet and lifestyle and toenail clippings were collected. Toenail DNA was genotyped using the Sequenom MassARRAY platform. After 20.3 years, 3004 subcohort members and 406 RCC cases, of which 263 ccRCC cases, were eligible for multivariate case-cohort analyses. VHL_rs779805 was associated with RCC (Hazard Ratio (HR) 1.53; 95% Confidence Interval (CI) 1.07–2.17) and ccRCC risk (HR 1.88; 95% CI 1.25–2.81). No associations were found for other SNPs. Potential gene-environment interactions were found between alcohol consumption and selected SNPs. However, none remained statistically significant after multiple comparison correction. No gene-gene interactions were observed between VHL and HIF1A. VHL promoter methylation was not associated with VHL SNPs. VHL SNPs may increase (cc)RCC susceptibility. No associations were found between gene-environment and gene-gene interactions and (cc)RCC risk and between VHL promoter methylation and VHL SNPs.
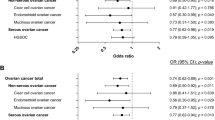
Similar content being viewed by others
Introduction
Genetic and epigenetic alterations in the Von Hippel-Lindau (VHL) gene are important drivers of carcinogenesis in clear-cell renal cell carcinoma (ccRCC)1. For sporadic ccRCC, biallelic inactivation of VHL because of rare, but highly penetrant, somatic mutations is relatively common2,3. Previous studies have estimated that 50–82% of patients with sporadic ccRCC have a mutation in the VHL gene4,5,6,7,8. The VHL gene encodes the VHL tumor suppressor protein (pVHL). Inactivation of pVHL leads to the unchecked accumulation of hypoxia-inducible factor 1 alpha (HIF1A), which facilitates oxygen delivery, adaptation to oxygen deprivation and angiogenesis1,9. Therefore, genetic or epigenetic alterations in VHL and HIF1A may lead to enhanced cell survival and carcinogenesis.
In contrast to the rare, but highly penetrant, sequence alterations leading to VHL loss, some germline Single Nucleotide Polymorphisms (SNPs) are highly frequent, but have a low penetrance. In general, SNPs account for many different phenotypes as they may alter disease susceptibility by affecting the gene’s function10. Genome-wide association studies (GWAS) have not found an association with VHL and HIF1A loci11,12,13,14,15,16,17,18. However, candidate gene studies have found conflicting evidence on the relationship between VHL SNPs and (cc)RCC risk, with some studies indicating a positive association19,20, while others indicate no association21. In previous studies, HIF1A SNPs have been associated with RCC prognosis, but not with (cc)RCC development21,22.
Previous studies have indicated the importance of assessing the interplay between genetic, epigenetic and environmental triggers when assessing ccRCC risk. Moore et al. found increased promoter hypermethylation in sporadic ccRCC when certain VHL polymorphisms were present23. In addition, multiple studies have indicated potential gene-environment interactions between germline SNPs and environmental factors in RCC24,25,26,27,28. To our knowledge, the relationship between established environmental risk factors associated with RCC risk, namely smoking, hypertension, obesity and alcohol consumption29, and VHL and HIF1A SNPs remains unstudied.
Therefore, we investigated the relationship between three selected germline VHL SNPs and one HIF1A SNP and (cc)RCC risk in the Netherlands Cohort Study on diet and cancer (NLCS). In addition, interactions between VHL and HIF1A SNPs and smoking, hypertension, body mass index (BMI) and alcohol consumption were studied. Lastly, we investigated the association between VHL promoter methylation and VHL SNPs.
Results
After excluding participants with missing values for predefined confounders 3004 subcohort members and 406 RCC cases, of which 263 ccRCC cases, were included in the analyses. The proportion of men was higher in both RCC and ccRCC cases when compared to the subcohort (Table 1). In addition, cases were more often smokers and were more often diagnosed with hypertension when compared to the subcohort.
Genotype and allele frequencies for the four selected SNPs in subcohort members of the NLCS are presented in Supplementary Table 1. All selected SNPs adhered to the Hardy-Weinberg Equilibrium. Only VHL_rs779805 had a minor allele frequency (MAF) above 25% and is, therefore, assessed primarily using additive models.
Main SNP effects
In both age- and sex-adjusted analyses and multivariable-adjusted analyses, an association with (cc)RCC risk was observed for SNPs in VHL_rs779805, but not for SNPs in VHL_rs1642739, VHL_rs265318 and HIF1A_rs2301111 (Table 2). In multivariable-adjusted analyses individuals carrying the AG (vs. AA) genotype of VHL_rs779805 had a statistically significantly increased RCC risk (HR 1.32, 95%CI 1.06–1.66), and the GG (vs. AA) genotype was associated with a statistically significantly increased RCC risk (HR 1.53, 95%CI 1.07–2.17). In addition, a statistically significant per-allele p for trend was observed (p = 0.004). In multivariable-analyses for ccRCC risk, the AG (vs. AA) genotype for VHL_rs779805 was associated with a statistically significantly increased ccRCC risk (HR 1.35, 95% CI 1.02–1.78), as was the GG (vs. AA) genotype of VHL_rs779805 (HR 1.88, 95%CI 1.25–2.81).
Gene-environment interactions
In multivariable-adjusted models for RCC risk, potential gene-environment interactions were observed between VHL_rs1642739, VHL_rs779805 and HIF1A_rs2301111 SNPs and alcohol consumption (Table 3). A weak inverse association between alcohol consumption (per 5 g/day) and RCC risk was observed in participants carrying the rare genotype for VHL_rs1642739 and VHL_rs779805, but not in participants carrying the wild-type genotype. For carriers of the wild-type HIF1A_rs2301111 genotype a weak inverse association was observed between alcohol consumption and RCC risk, but not for individuals carrying the rare genotype. No interaction was observed between either of the selected SNPs and self-reported hypertension (yes, no), smoking status (never, former, current) and BMI (per kg/m2) for RCC risk. For ccRCC, a potential interaction between VHL_rs779805 SNPs and alcohol consumption was observed. However, after correction for multiple comparisons using the adaptive Benjamini-Hochberg method none of the potential gene-environment interactions maintained statistical significance30.
In sensitivity analyses, a potential gene-environment interaction was apparent between categorized alcohol consumption (0 g/d, 0.1–4 g/d, 5–14 g/d, 15–29 g/d and 30+ g/d) and VHL_rs164273 status for ccRCC risk (p = 0.009; Supplementary Table 2). The direction of associations for VHL_rs779805 was similar to main analyses using alcohol consumption (per 5 g/day). Sensitivity analyses between smoking status (ever/never), hypertension (no self-reported hypertension or no self-reported antihypertensive medication, hypertension with self-reported hypertensive medication) and BMI (<20 kg/m2, 20–<25 kg/m2, 25–<30 kg/m2 and 30+ kg/m2) and SNP status showed similar associations compared to main gene-environment analyses (Supplementary Table 2). Similar to main analyses, no sensitivity analysis remained statistically significant after multiple comparison correction.
Gene-gene interactions
No gene-gene interactions, as tested with the Wald χ2-test, were found between the three selected VHL SNPs and HIF1A_rs2301111 for both RCC (p = 0.310, p = 0.321 and p = 0.514 for VHL_rs1642739, VHL_rs779805 and VHL_rs265318, respectively) and ccRCC (p = 0.762, p = 0.442 and p = 0.978 for VHL_rs1642739, VHL_rs779805 and VHL_rs265318, respectively).
Association between SNPs and VHL promoter methylation status
In total, information on VHL promoter methylation was available from 253 ccRCC cases. Among ccRCC cases, 19 (7.5%) participants had a methylated CpG island in the VHL promoter region of which 13 had at least one mutant allele for the selected VHL SNPs (Supplementary Table 3). VHL promoter methylation was apparent in three, twelve and two participants for the rare genotype of VHL_rs1642739 (GG vs. GT + TT), VHL_rs779805 (AA vs. AG + GG) and VHL_rs264318 (AA vs. AC + CC), respectively. In multivariable-adjusted analyses a non-significant inverse association was observed between both VHL_rs1642739 (HR 0.45, 95%CI 0.12–1.69) and VHL_rs265318 (HR 0.38, 95%CI 0.07–2.00) and VHL promoter methylation in ccRCC cases. No association was observed for the VHL_rs779805 SNP (HR 0.99, 95%CI 0.37–2.69).
Discussion
In this study, a statistically significantly increased RCC risk was found for individuals that carry genotypes with at least one variant allele for the VHL_rs779805 SNP. This association was especially pronounced for ccRCC risk. No association was found for VHL_rs164239, VHL_rs265318 and HIF1A_rs2301111. After adjustment for multiple comparisons, no statistically significant gene-environment interactions were found between the selected SNPs and smoking, hypertension, BMI and alcohol for both RCC and ccRCC cases. No gene-gene interactions were found between selected VHL SNPs and the HIF1A SNP.
Several studies have assessed the relationship between the VHL_rs779805 SNP and sporadic RCC19,20,21. Lv et al. found an association between the germline SNP VHL_rs779805 and RCC risk. Similarly, we found a statistically significant positive trend for the G allele and a positive association between the GG genotype for VHL_rs779805 and RCC risk20. The aforementioned studies did not report associations between VHL SNPs and ccRCC risk. In our study, rare VHL_rs779805 genotypes had a stronger association with ccRCC risk than with RCC risk. This might indicate that VHL polymorphisms lead to an increased susceptibility for ccRCC in particular. To our knowledge, no other study has investigated the relationship between VHL_rs1642739, VHL_rs265318 and HIF1A_rs2301111 and (cc)RCC risk. In this study, no association was found between (cc)RCC risk and VHL_rs1642739, VHL_rs265318 or HIF1A_rs2301111.
Multiple studies have assessed gene-environment interactions in RCC and ccRCC. RCC risk has been found to be associated with interactions between alcohol consumption and ADH726; sodium and hypertension and AGTR, AGT and ACE31; calcium and vitamin D intake and RXRA28; tobacco smoking and NAT2, CYP1A1 and GSTM125; and meat-cooking mutagens and ITPR2 and EPAS127. To our knowledge, we are the first to study gene-environment interactions between the selected VHL and HIF1A SNPs and smoking, hypertension, BMI and alcohol consumption. Solely the interaction between VHL_rs779805 and alcohol consumption was associated with both RCC and ccRCC risk. However, this association did not maintain statistical significance after correction for multiple comparisons with the adaptive Benjamini-Hochberg method. Dominant models were used for all gene-environment analyses because of the low MAF of most included SNPs. However, SNPs may not have adhered to a dominant model, as there may be differences in disease susceptibility between heterozygous and homozygous rare genotypes, as was found for VHL_rs779805 (Table 2). This exemplifies that our gene-environment analyses may have been hampered by the inability to assess interactions per genotype. Further research is needed to ascertain the interaction between alcohol and VHL SNP status on (cc)RCC risk.
Disruptions in the VHL tumor suppressor gene are thought to play a role in the constitutive activation of hypoxia-inducible factors, as regulated in part by HIF1A, which may lead to carcinogenesis1. Therefore, it is plausible for gene-gene interactions to occur. However, in this study, we did not find gene-gene interactions between selected VHL and HIF1A SNPs on the risk of developing (cc)RCC.
Previous studies have found a relationship between VHL promoter hypermethylation and SNPs in VHL_rs779805 in sporadic ccRCC cases6,23. Moore et al. also reported a positive association between promoter hypermethylation and VHL_rs265318 and VHL_rs1642739. In contrast, we found no association between promoter methylation status and VHL_rs779805 in ccRCC cases. VHL_rs1642739 and VHL_rs265318 seemed inversely associated with VHL promoter methylation in ccRCC cases. However, this association was based on a limited sample size. While the number of cases with known promoter methylation status was similar in size to the study of Moore et al., our study had a smaller proportion of cases with VHL promoter methylation (7.5% vs. 9.8%)23. Banks et al. reported an even higher proportion of sporadic ccRCC cases with a methylated VHL promoter (20.4%), but had a smaller study population6. In general, there are large differences in the proportion of methylated VHL promoters per SNP between studies, which may explain these unstable point estimates23. Therefore, more research with a larger number of sporadic ccRCC cases is needed to elucidate the relationship between VHL promoter methylation and VHL SNPs.
At present, genome-wide association studies (GWAS) have identified multiple novel risk loci that may contribute to RCC susceptibility. Interestingly, SNPs in the VHL and HIF1A genes have not (yet) been identified as potential risk variants, while there is a biological plausibility for the involvement of these genes based on current evidence on the development of RCC2,9. For example, risk loci have been identified in EPAS111,13,17,18, which is known to be involved in the VHL-HIF-1 pathway32. While we found no evidence for an association between three of our selected SNPs, VHL_rs779805 was associated with an increased risk of RCC. This finding was in line with two prior published studies, in which a potential association between VHL_rs779805 and RCC risk was found19,20. While this particular SNP is present on commonly used SNP arrays, this SNP remains unidentified in large-scale GWAS studies11,12,13,14,15,16,17,18. It is estimated that the currently available risk loci for RCC account for approximately 10% of the familial risk for RCC11. Therefore, it may well be possible for minor susceptibility loci to remain unidentified in GWAS studies, due to their tendency to convey small-to-moderate changes in risk, while major susceptibility loci are detectable in the stringent false discovery rate correction criteria of GWAS studies. This could be a reason why SNPs like VHL_rs779805 may remain unidentified, unless alternative methodologies are employed11. As a result, there is ample opportunity to discover new, rarer, RCC risk variants in future research. Additional evidence on risk loci from GWAS studies, combined with extensive information on direct effects, environmental factors and other potential modulators of disease etiology from candidate SNP studies, should lead to new insights into the biology of RCC to further the potential for new prevention, early detection and intervention strategies to be employed11.
This study also has several strengths. Strengths of this study were the detailed questionnaire information, the long duration and the histological revision of RCC cases by two experienced pathologists. Furthermore, cases in our study were obtained prospectively from a population of 120,852 men and women from 204 Dutch municipalities. Combined with the completeness of follow-up, we assume that these cases are a representative of kidney cancer cases in the Netherlands at the time.
In conclusion, this study confirmed the association between germline SNP VHL_rs779805 with RCC risk. In addition, a slightly stronger association for ccRCC was found compared to RCC. Potential gene-environment interactions were found between alcohol and VHL SNPs. However, results did not remain statistically significant after correction for multiple comparisons. No gene-gene interactions were observed between the VHL and HIF1A SNPs. Lastly, tumor promoter methylation was not significantly associated with VHL SNPs.
Methods
Study design
The NLCS is a nation-wide prospective cohort study initiated in September 1986 with the inclusion of 120,852 participants aged 55–69 years to study the relationship between diet and cancer. The study design has been described in detail elsewhere33. In short, a case-cohort design was used for efficiency in data processing and follow-up for vital status. Cases were derived from the entire cohort, whereas a subcohort of 5000 participants, consisting of 2411 men and 2589 women, was randomly sampled at baseline to estimate person years at risk for the entire cohort. The subcohort was followed up biennially for migration and vital status information by contacting participants and using computerized municipalities registries. Using the subcohort, person-years at risk were calculated from baseline until registration of RCC, or until date of censoring by death, emigration, loss to follow-up or end of follow-up, whichever occurred first. Cancer follow-up for the full cohort was conducted by computerized record linkage with the Netherlands Cancer Registry (NCR), the Netherlands Pathology Registry (PALGA), and causes of death registry maintained by Statistics Netherlands (CBS)34. Follow-up for vital status of the subcohort was nearly 100% complete after 20.3 years. The completeness of cancer follow-up is estimated to be over 96%35.
Individuals with prevalent cancer, excluding skin cancer, at baseline were excluded. After 20.3 years of follow-up, 608 RCC cases were identified (International Classification of Diseases for Oncology 3 (ICD-O-3):C64). Histologically confirmed epithelial RCC cases were eligible for the collection of formalin-fixed paraffin-embedded (FFPE) tumor tissue. Tumor blocks were collected for 454 out of 568 eligible cases (80%). Two experienced pathologists revised the tumor histology according to the WHO-classification of RCC tumors36. Based on this revision 366 (81%) of the cases with available tumor blocks were classified as ccRCC cases, 60 (13%) as papillary RCC cases, 15 (3.3%) chromophobe RCC cases, and 13 (2.9%) other or undefined RCC cases.
Ethics statement
Individuals invited to participate in the NLCS received an invitation letter with details on the study and the use of their data. In addition, they received the baseline questionnaire, which included an envelope for returning toenail clippings. By completing and returning the baseline questionnaire, individuals consented to participate in the NLCS (response rate 35.5%). Individuals were informed about the possibility to end their participation at any time, at which point all their data would be removed. All methods were performed in accordance with the relevant guidelines and regulations that were applicable at that time (1986). The institutional review boards of Maastricht University (Maastricht) and the Netherlands Organization for Applied Scientific Research TNO (Zeist) approved the NLCS (February 2, 1985 and January 6, 1986, respectively). The institutional review board of Maastricht University (Maastricht) later re-evaluated the original approval of the study protocol and procedures (2010). Based on the re-evaluation the institutional review board amended the original approval to include the genotyping of SNPs (April 12, 2010). Participants did not provide written informed consent to the sharing of data.
Gene and SNP selection
Genes and SNPs related to RCC risk were selected through literature search. Priority was given to SNPs with a MAF ≥ 20% in Caucasians and primers had to be compatible with RAAS-pathway SNPs present on the multiplex assay31. Consequently, three VHL SNPs (rs779805, rs265318 and rs1642739) and one HIF1A SNP (rs2301111) were selected. All included VHL SNPs were selected based on their association with VHL promoter methylation in previous research23. The included HIF1A tag-SNP had the highest MAF of the HIF1A SNPs compatible with the assay.
Tissue collection and DNA isolation
Approximately 90,000 participants provided toenail clippings at baseline, which have been shown to be a valid source of DNA for the genotyping of germline genetic variants37. DNA was isolated according to the DNA isolation protocol by Cline et al.38. To increase the number of cases with available DNA, DNA was isolated from FFPE healthy tissue, as described by van Houwelingen et al.5, for 67 RCC cases without toenail clippings. There were no substantial quality differences between DNA samples from toenail and FFPE healthy tissue31. In total, 3582 (75%) subcohort members and 502 (83%) RCC cases were genotyped.
SNP genotyping was performed on the Sequenom MassARRAY platform using the iPLEX assay (Sequenom Inc., Hamburg, Germany), as described previously31. This method provides suitable SNP call rates and reproducibility using toenail DNA37.
DNA methylation of the CpG island of the VHL gene promoter region, of which methylation has been associated with inhibition of VHL gene expression39, in RCC tumor blocks was determined by chemical modification of genomic DNA with sodium bisulfite and subsequent methylation-specific PCR analysis (MSP) as previously described elsewhere40,41,42. MSP primer design was based on the MBD-affinity massive parallel sequencing data. Detailed information on primer sequences and MSP conditions are available elsewhere24.
Questionnaire information
All participants completed a mailed, self-administered, questionnaire on diet and other cancer risk factors for cancer at baseline (1986)43. Information on dietary habits was obtained through a 150-item, semi-quantitative food frequency questionnaire (FFQ) focusing on habitual consumption of food and beverages during the year preceding baseline.
Cigarette smoking status, frequency and duration were based on self-reported information. Participants reported hypertension as diagnosed by a physician, preceding baseline. Participants were asked to report the use of any drugs that they used longer than 6 months. From this information, the use of antihypertensive medication was extracted. BMI was calculated using self-reported height and weight from the baseline questionnaire. Questions on beer, red wine, white wine, sherry, fortified wines, liqueur, and liquor were used to assess the consumption of alcohol. Participants who consumed alcoholic beverages less than once a month were considered non-users. Standard glass sizes were defined as 200 ml for beer, 105 ml for wine, 80 ml for sherry, and 45 ml for both liqueur and liquor44. These values corresponded to 8, 10, 11, 7 and 13 grams of alcohol, respectively. Mean daily alcohol consumption was calculated by multiplying the consumption frequency and the standardized item unit.
Statistical analyses
Cox proportional hazards models were used to estimate age- and sex-adjusted and multivariable-adjusted hazard ratios (HR) and 95% confidence intervals (CIs). A priori selected covariables in the multivariable-adjusted model were BMI (kg/m2, continuous), hypertension (yes,no), cigarette smoking status (never, former, current), intensity (cig/d, centered; continuous), duration (years, centered; continuous) and alcohol consumption (g/d, continuous).
The most common allele was used as the reference allele. Associations between genotypes and RCC and ccRCC risk were assessed using additive and dominant models. Results of SNPs with a MAF < 0.25 were interpreted using a dominant model for power reasons. SNP allele frequencies in the subcohort were tested against departure from the Hardy-Weinberg Equilibrium using the Pearson χ2-test, as calculated with the Stata program ‘hwsnp’45. Gene-environment interactions were tested with the Wald χ2-test. Gene-environment analyses were adjusted for multiple comparisons with the adaptive Benjamini-Hochberg false discovery rate (FDR) procedure with a q-value threshold of 10%30. Sensitivity analyses were performed to explore the impact of using alternative categorizations for BMI (<20 kg/m2, 20–<25 kg/m2, 25–<30 kg/m2 and 30 + kg/m2), smoking status (never, ever), hypertension (no self-reported hypertension or no self-reported antihypertensive medication, hypertension with self-reported hypertensive medication) and alcohol consumption (0 g/d, 0.1–4 g/d, 5–14 g/d, 15–29 g/d and 30+ g/d) when assessing gene-environment interactions. Gene-gene interactions between VHL SNPs and the selected HIF1A SNP were tested using the Wald χ2-test. In a case-only analysis, the association between VHL SNPs and VHL tumor promoter methylation status (methylated, unmethylated) was assessed using multiple logistic regression for both RCC and ccRCC.
All analyses were performed using Stata Statistical Software: Release 15 (StataCorp., 2017, College Station, TX). The proportional hazards assumption was tested using scaled Schoenfeld residuals46. A violation of the assumption was apparent for age. Therefore, all models were adjusted for age as a time-dependent covariable. With the exception of FDR-corrected analyses, a p-value < 0.05 was considered statistically significant.
Data availability
The datasets generated during and/or analyzed during the current study are not publicly available because the participants did not provide written informed consent to the sharing of data.
References
Hsieh, J. J. et al. Renal cell carcinoma. Nat. Rev. Dis. Primers 3, 17009, https://doi.org/10.1038/nrdp.2017.9 (2017).
Kaelin, W. G. Von Hippel-Lindau disease. Annu. Rev. Pathol. 2, 145–173, https://doi.org/10.1146/annurev.pathol.2.010506.092049 (2007).
Kim, W. Y. & Kaelin, W. G. Role of VHL Gene Mutation in Human Cancer. Journal of Clinical Oncology 22, 4991–5004, https://doi.org/10.1200/jco.2004.05.061 (2004).
Nickerson, M. L. et al. Improved identification of von Hippel-Lindau gene alterations in clear cell renal tumors. Clin Cancer Res 14, 4726–4734, https://doi.org/10.1158/1078-0432.ccr-07-4921 (2008).
van Houwelingen, K. P. et al. Prevalence of von Hippel-Lindau gene mutations in sporadic renal cell carcinoma: results from The Netherlands cohort study. BMC Cancer 5, 57, https://doi.org/10.1186/1471-2407-5-57 (2005).
Banks, R. E. et al. Genetic and epigenetic analysis of von Hippel-Lindau (VHL) gene alterations and relationship with clinical variables in sporadic renal cancer. Cancer Res 66, 2000–2011, https://doi.org/10.1158/0008-5472.can-05-3074 (2006).
Gnarra, J. R. et al. Mutations of the VHL tumour suppressor gene in renal carcinoma. Nat Genet 7, 85–90, https://doi.org/10.1038/ng0594-85 (1994).
Cancer Genome Atlas Research, N. Comprehensive molecular characterization of clear cell renal cell carcinoma. Nature 499, 43–49, https://doi.org/10.1038/nature12222 (2013).
Schodel, J. et al. Hypoxia, Hypoxia-inducible Transcription Factors, and Renal Cancer. Eur. Urol 69, 646–657, https://doi.org/10.1016/j.eururo.2015.08.007 (2016).
Hunter, D. J. Gene-environment interactions in human diseases. Nat. Rev. Genet 6, 287–298, https://doi.org/10.1038/nrg1578 (2005).
Scelo, G. et al. Genome-wide association study identifies multiple risk loci for renal cell carcinoma. Nat. Commun. 8, 15724, https://doi.org/10.1038/ncomms15724 (2017).
Henrion, M. et al. Common variation at 2q22.3 (ZEB2) influences the risk of renal cancer. Hum Mol Genet 22, 825–831, https://doi.org/10.1093/hmg/dds489 (2013).
Purdue, M. P. et al. Genome-wide association study of renal cell carcinoma identifies two susceptibility loci on 2p21 and 11q13.3. Nat Genet 43, 60–65, https://doi.org/10.1038/ng.723 (2011).
Wu, X. et al. A genome-wide association study identifies a novel susceptibility locus for renal cell carcinoma on 12p11.23. Hum Mol Genet 21, 456–462, https://doi.org/10.1093/hmg/ddr479 (2012).
Gudmundsson, J. et al. A common variant at 8q24.21 is associated with renal cell cancer. Nat. Commun. 4, 2776, https://doi.org/10.1038/ncomms3776 (2013).
Purdue, M. P. et al. A genome-wide association study of renal cell carcinoma among African Americans. Cancer Epidemiol Biomarkers Prev. 23, 209–214, https://doi.org/10.1158/1055-9965.epi-13-0818 (2014).
Henrion, M. Y. et al. Common variation at 1q24.1 (ALDH9A1) is a potential risk factor for renal cancer. PLoS One 10, e0122589, https://doi.org/10.1371/journal.pone.0122589 (2015).
Laskar, R. S. et al. Sex specific associations in genome wide association analysis of renal cell carcinoma. Eur. J. Hum Genet 27, 1589–1598, https://doi.org/10.1038/s41431-019-0455-9 (2019).
Wang, W. C., Tsou, M. H., Chen, H. J., Hsu, W. F. & Lai, Y. C. Two single nucleotide polymorphisms in the von Hippel-Lindau tumor suppressor gene in Taiwanese with renal cell carcinoma. BMC Res. Notes 7, 638, https://doi.org/10.1186/1756-0500-7-638 (2014).
Lv, C., Bai, Z., Liu, Z., Luo, P. & Zhang, J. Renal cell carcinoma risk is associated with the interactions of APOE, VHL and MTHFR gene polymorphisms. Int. J. Clin. Exp. Pathol. 8, 5781–5786 (2015).
Qin, C. et al. The polymorphisms in the VHL and HIF1A genes are associated with the prognosis but not the development of renal cell carcinoma. Ann Oncol 23, 981–989, https://doi.org/10.1093/annonc/mdr325 (2012).
Qin, C. et al. Variants in angiogenesis-related genes and the risk of clear cell renal cell carcinoma. Mutagenesis 29, 419–425, https://doi.org/10.1093/mutage/geu046 (2014).
Moore, L. E. et al. Von Hippel-Lindau (VHL) inactivation in sporadic clear cell renal cancer: associations with germline VHL polymorphisms and etiologic risk factors. PLoS Genet 7, e1002312, https://doi.org/10.1371/journal.pgen.1002312 (2011).
Schouten, L. J., Deckers, I. A., van den Brandt, P. A., Baldewijns, M. M. & van Engeland, M. Alcohol and Dietary Folate Intake and Promoter CpG Island Methylation in Clear-Cell Renal Cell Cancer. Nutr. Cancer 68, 1097–1107, https://doi.org/10.1080/01635581.2016.1187283 (2016).
Smits, K. M. et al. Polymorphisms in genes related to activation or detoxification of carcinogens might interact with smoking to increase renal cancer risk: results from The Netherlands Cohort Study on diet and cancer. World J. Urol 26, 103–110, https://doi.org/10.1007/s00345-007-0220-5 (2008).
Antwi, S. O. et al. Alcohol consumption, variability in alcohol dehydrogenase genes and Risk of Renal Cell Carcinoma. Int J Cancer, https://doi.org/10.1002/ijc.31103 (2017).
Melkonian, S. C. et al. Gene-environment interaction of genome-wide association study-identified susceptibility loci and meat-cooking mutagens in the etiology of renal cell carcinoma. Cancer 122, 108–115, https://doi.org/10.1002/cncr.29543 (2016).
Karami, S. et al. Vitamin d pathway genes, diet, and risk of renal cell carcinoma. Int. J. Endocrinol 2010, 879362, https://doi.org/10.1155/2010/879362 (2010).
Chow, W. H., Dong, L. M. & Devesa, S. S. Epidemiology and risk factors for kidney cancer. Nat. Rev. Urol 7, 245–257, https://doi.org/10.1038/nrurol.2010.46 (2010).
Benjamini, Y. & Hochberg, Y. On the Adaptive Control of the False Discovery Rate in Multiple Testing with Independent Statistics. Journal of Educational and Behavioral Statistics 25, 60–83, https://doi.org/10.2307/1165312 (2000).
Deckers, I. A. et al. Polymorphisms in genes of the renin-angiotensin-aldosterone system and renal cell cancer risk: interplay with hypertension and intakes of sodium, potassium and fluid. Int. J. Cancer 136, 1104–1116, https://doi.org/10.1002/ijc.29060 (2015).
Schödel, J. et al. Common genetic variants at the 11q13.3 renal cancer susceptibility locus influence binding of HIF to an enhancer of cyclin D1 expression. Nat. Genet 44, 420–S422, https://doi.org/10.1038/ng.2204 (2012).
van den Brandt, P. A. et al. A large-scale prospective cohort study on diet and cancer in The Netherlands. J. Clin. Epidemiol 43, 285–295 (1990).
Van den Brandt, P. A., Schouten, L. J., Goldbohm, R. A., Dorant, E. & Hunen, P. M. Development of a record linkage protocol for use in the Dutch Cancer Registry for Epidemiological Research. International Journal of Epidemiology 19, 553–558 (1990).
Goldbohm, R. A., Van den Brandt, P. A. & Dorant, E. Estimation of the coverage of Dutch municipalities by cancer registries and PALGA based on hospital discharge data. Tijdschr Soc Gezondheidsz 72, 80–84 (1994).
Eble, J., Sauter, G., Epstein, J. & Sesterhenn, I. World Health Organization Classification of Tumours. Pathology and Genetics. Tumours of the Urinary System and Male Genital Organs. (IARC Press, 2004).
Hogervorst, J. G. et al. DNA from nails for genetic analyses in large-scale epidemiologic studies. Cancer Epidemiol Biomarkers Prev 23, 2703–2712, https://doi.org/10.1158/1055-9965.EPI-14-0552 (2014).
Cline, R. E., Laurent, N. M. & Foran, D. R. The fingernails of Mary Sullivan: developing reliable methods for selectively isolating endogenous and exogenous DNA from evidence. J Forensic Sci 48, 328–333 (2003).
Schouten, L. J. et al. Alcohol consumption and mutations or promoter hypermethylation of the von Hippel-Lindau gene in renal cell carcinoma. Cancer Epidemiol Biomarkers Prev 17, 3543–3550, https://doi.org/10.1158/1055-9965.epi-08-0321 (2008).
Derks, S. et al. Methylation-specific PCR unraveled. Cell Oncol 26, 291–299 (2004).
Herman, J. G., Graff, J. R., Myohanen, S., Nelkin, B. D. & Baylin, S. B. Methylation-specific PCR: a novel PCR assay for methylation status of CpG islands. Proc Natl Acad Sci USA 93, 9821–9826, https://doi.org/10.1073/pnas.93.18.9821 (1996).
van Engeland, M. et al. K-ras mutations and RASSF1A promoter methylation in colorectal cancer. Oncogene 21, 3792–3795, https://doi.org/10.1038/sj.onc.1205466 (2002).
van den Brandt, P. A. & Goldbohm, R. A. A prospective cohort study on diet and cancer in the Netherlands: design, conduct, analysis and first results after 3.3 years of follow-up, Maastricht University (1993).
Goldbohm, R. A. et al. Prospective study on alcohol consumption and the risk of cancer of the colon and rectum in the Netherlands. Cancer Causes Control 5, 95–104 (1994).
Cleves, M. A. Exploratory analysis of single nucleotide polymorphism (SNP) for quantitative traits. Stata Journal 5, 141–153 (2005).
Schoenfeld, D. Partial Residuals for The Proportional Hazards Regression Model. Biometrika 69, 239–241, https://doi.org/10.2307/2335876 (1982).
Acknowledgements
The authors are indebted to all the participants of the Netherlands Cohort study on diet and cancer, and further wish to thank the Netherlands Cancer Registry, the Dutch Pathology Registry (PALGA) and the BioBank Maastricht UMC+. The authors are grateful to I. Deckers for the preparatory work and sample selection, S. van Breda for the pilot work to isolate DNA from toenails, the laboratory technicians L. Maas, J. Goessens, L. Jonkers, K. Lemmens and S. Lumeij for DNA isolation and sample preparation and C. Hulsbergen-van de Kaa and M. Baldewijns for revising the tumor histology. Finally, the authors thank C. Simons, S. van de Crommert, J. Nelissen, H. Brants, C. de Zwart, M. Moll and A. Pisters, for their assistance with data entry or data management; and H. van Montfort, T. van Moergastel, E. Dutman, and R. Schmeitz for programming assistance. This research received no specific grant from any funding agency in the public, commercial or not-for-profit sectors.
Author information
Authors and Affiliations
Contributions
L.S. conceived the study. All authors participated in the analysis and interpretation of data. J.v.d.P. carried out the statistical analyses and drafted the manuscript. P.v.d.B., M.v.E., R.G., F.S., J.H. and L.S. critically revised the manuscript for important intellectual content. All authors approved the final manuscript as submitted.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
van de Pol, J.A.A., van den Brandt, P.A., van Engeland, M. et al. Germline polymorphisms in the Von Hippel-Lindau and Hypoxia-inducible factor 1-alpha genes, gene-environment and gene-gene interactions and renal cell cancer. Sci Rep 10, 137 (2020). https://doi.org/10.1038/s41598-019-56980-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-56980-0
This article is cited by
-
Frequency Analysis of Flood Flow in Markanda Basin of Ghaggar River System in North Western India
Journal of the Geological Society of India (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.