Abstract
Sonic Hedgehog (SHH) medulloblastomas are brain tumours that arise in the posterior fossa. Cancer-propagating cells (CPCs) provide a reservoir of cells capable of tumour regeneration and relapse post-treatment. Understanding and targeting the mechanisms by which CPCs are maintained and expanded in SHH medulloblastoma could present novel therapeutic opportunities. We identified the aryl hydrocarbon receptor (AHR) pathway as a potent tumour suppressor in a SHH medulloblastoma mouse model. Ahr-deficient tumours and CPCs grown in vitro, showed elevated activation of the TGFβ mediator, SMAD3. Pharmacological inhibition of the TGFβ/SMAD3 signalling axis was sufficient to inhibit the proliferation and promote the differentiation of Ahr-deficient CPCs. Human SHH medulloblastomas with high expression of the AHR repressor (AHRR) exhibited a significantly worse prognosis compared to AHRRlow tumours in two independent patient cohorts. Together, these findings suggest that reduced AHR pathway activity promotes SHH medulloblastoma progression, consistent with a tumour suppressive role for AHR. We propose that TGFβ/SMAD3 inhibition may represent an actionable therapeutic approach for a subset of aggressive SHH medulloblastomas characterised by reduced AHR pathway activity.
Similar content being viewed by others
Introduction
Medulloblastoma represents one of the most common forms of paediatric, malignant brain tumours accounting for around 20% of all paediatric tumours of the CNS1. Typical treatment consists of a combination of chemotherapy, surgical resection and neuraxis irradiation, with a cure rate of approximately 70–75% in children ≥3 years of age2. However, survivors of medulloblastoma are left with a host of long-term adverse sequelae, including cognitive deficits, problems with neuroendocrine function and fertility3,4. These drawbacks to traditional treatment options necessitate more effective patient stratification strategies based on biomarkers that predict outcome and identify specific molecular medulloblastoma subtypes for personalized, targeted therapies.
Efforts to dissect the molecular underpinnings of medulloblastoma have identified four main subgroups - WNT, Sonic hedgehog (SHH), Group 3 and Group 4 - with distinct transcriptional, DNA methylation and mutational profiles, and different clinical characteristics and outcomes5,6,7. These subgroups are associated with different cells of origin8. SHH tumours originate from cerebellar granule cell progenitors (GCPs), and display excessive activation of the SHH signalling pathway9. The prognosis of SHH subgroup medulloblastomas is mixed, with some patients doing well and others not, suggesting significant heterogeneity within this subgroup7. Indeed, recent work has identified SHH medulloblastoma subtypes with specific molecular characteristics and associated clinical features5,6. Independent efforts showed that SHH MBs could be subdivided depending on age and molecular signature. Among these, infant (<3.0–4.0 years at diagnosis) and non-infant SHH subgroups are consistently observed, alongside a subgroup of non-infant tumours associated with a particularly poor prognosis, characterized by amplifications of MYCN and GLI2 and loss of function mutations in TP535,6. These molecular signatures have previously been linked to medulloblastomas with metastasis and high rates of post-treatment relapse6,10,11. Non-infant SHH tumours without these features are associated with better outcomes (>80% 5-year progression-free survival5).
The Ahr gene, encoding the aryl hydrocarbon receptor, has been studied extensively in the context of hepatocarcinoma, immune cell development and toxicology12,13,14, where this gene appears to have context-specific roles. In hepatocarcinoma, Ahr promotes cell proliferation and tumourigenesis12, while it has multiple roles in the immune system15. Intriguingly, activation of the AHR pathway by endogenous ligands has been shown to promote brain cancers by the anti-tumour immune response16, also suggestive of context-dependent, multi-faceted roles for this pathway in cancer biology. AHR is a bHLH (basic helix-loop-helix) transcription factor which acts as a receptor for endogenous tryptophan metabolites and xenobiotics such as TCDD17. Upon ligand binding, AHR translocates to the nucleus, where it forms a complex with formation of a heterodimer with ARNT (Aryl hydrocarbon Receptor Nuclear Translocator). AHR-ARNT heterodimers are recruited to Dioxin Response Elements (DRE) in the genome to regulate gene transcription18. An AHR Repressor (AHRR) protein can also dimerise with ARNT to competitively interfere with AHR-ARNT complex formation and inhibit AHR-regulated gene expression19.
Sox2+ cancer propagating cells (CPCs), capable of driving tumour initiation and exhibiting enhanced resistance to cytostatic therapy have been identified in SHH medulloblastoma mouse models20,21. Lineage tracing experiments of these cells demonstrated their capacity for tumour regeneration following anti-mitotic chemotherapy, suggesting these cells are responsible for tumour relapse20. However, the mechanisms governing CPC formation and maintenance in SHH medulloblastomas remain to be fully elucidated. AHR function has been linked to CPC and haematopoietic stem cell maintenance22,23. In a recent study, AHR was shown to regulate the balance between quiescence and proliferation in hematopoietic stem cells, with these stem cells becoming less quiescent and more proliferative in Ahr-deficient animals23.
Previous studies have implicated AHR in cerebellar development and medulloblastoma cell proliferation. Ahr deletion in primary cerebellar GCPs24 or AHR knock-down in a SHH-associated medulloblastoma cell line25 resulted in proliferative deficits. To determine if AHR has a direct role in SHH medulloblastoma in vivo, we conditionally deleted the Ahr gene in mouse cerebellar GCPs, either alone or in combination with medulloblastoma-initiating Ptch1 gene deletion. Our analyses of these mice revealed a striking tumour-suppressive role for AHR in mouse SHH medulloblastoma development. We identify a specific role for AHR in regulating the TGFβ-SMAD3 signalling axis in CPCs from these tumours and identify a new role for TGFβ-SMAD3 activity in medulloblastoma CPC differentiation. Examination of the expression of AHR pathway genes in human medulloblastoma cohorts support an important role for the AHR pathway in SHH medulloblastoma biology.
Results
AHR modulates primary mouse GCP proliferation and differentiation by repressing TGFβ/SMAD3 signalling
To investigate the role of the AHR pathway in neural progenitor fate in the developing cerebellum, we conditionally deleted the Ahr gene from Math1+ GCPs during cerebellar development. In agreement with a previous report24, we observed reduced GCP proliferation and enhanced cell cycle exit (as measured by cell Q fraction) (Fig. S1a) of GCPs in Ahr conditional knockout Math1cre; Ahrf/f (Ahr cKO) cerebella, compared to control Ahrf/f cerebella (Fig. S1b,c). The phenotype was particularly prominent in anterior lobules I/II, III, V and VI (Fig. S1d,e). This effect was not observed in the posterior lobules IX/X, which is attributable to lack of Cre activity within these lobules, as described previously26.
We confirmed that this proliferative deficit was retained in vitro. Ahr-deficient, primary GCPs isolated from P7 Ahr cKO mice proliferated less compared to control GCPs (Fig. 1a,b). Furthermore, we found that more Ahr-deficient GCPs commenced differentiation as evidenced by expression of the definitive differentiation marker Neurod1 after a 24 hour culture period, compared to controls (Fig. 1a,c). A substantial fraction of GCPs displayed positivity for both Ki67 and Neurod1 (Fig. 1a), which can be expected to occur at 24 hours as cells are transitioning from a proliferative to a terminally differentiated state. To determine whether Ahr-deficient cells matured faster, primary GCPs were cultured in the absence of exogenous Sonic hedgehog (SHH) for six days in vitro and neurite length measured as an indicator of granule cell differentiation27. Map2 was used as a marker for neurites due to its importance in stabilizing microtubule activity in mature neurons28. Ahr-deficient GCPs displayed >2-fold increase in neurite length on average compared to controls (Fig. 1d,e), confirming a role for Ahr in suppressing GCP differentiation and maturation.
Ahr regulates the balance between proliferation and differentiation in GCPs. (a) Isolated P7 GCPs were cultured for 24 hours in the presence of exogenous SHH. Each cerebellum was processed independently (without pooling) from 3 control and 3 cKO littermates and the experiment repeated three times on separate occasions with independent litters. The data shown is the average of the three experiments with 9 WT and 9 cKO cerebella in total. Panels show immunostaining for Ki67 (green), Neurod1 (red) and Hoechst counterstained nuclei (blue). Merged panels show composite images. White arrows indicate Ki67+ cells. (b) Quantification of %Ki67 positive cells from a. (c) Quantification of %Neurod1 positive cells from a. (d) Isolated P7 GCPs were seeded in 6 well plates and allowed to differentiate in the absence of SHH, before immunocytochemical staining was performed for Map2 (green), followed by neurite tracing and quantification after 6 days in vitro (6 DIV). White arrows indicate neurites extending from granule neuron soma. (e) Quantification of neurite length (um). Data in panels b, c and e were analyzed by Student’s t test (p < 0.001 (***), p < 0.01 (**), p < 0.05 (*)). Bars represent mean values +/− SEM. Scale bars: 10 um (a,d).
To identify molecular pathways that were altered in Ahr-deficient cells, we performed immunoblots of lysates from purified GCPs to detect activated signalling mediators. For instance, as AHR has been shown to regulate the TGFβ-SMAD signalling pathway in several other contexts29, including brain tumours30, we assayed for the activated, phosphorylated (S423 and S425) form of SMAD3, an intracellular mediator of TGFβ receptor activation31. P-SMAD3 levels were elevated several-fold in Ahr cKO GCPs compared to control GCPs (Fig. 2a,b). In comparison, neither P-SMAD2 levels, nor the total amount of SMAD2 and SMAD3 proteins were altered in these cells (Fig. 2a). To determine whether SMAD3 hyperactivation was responsible for the altered phenotype of Ahr-deficient GCPs, primary GCPs from control and Ahr conditional mutant mice were cultured in vitro in the presence of exogenous SHH, with or without the selective SMAD3 inhibitor SIS332. After 24 hours in culture, the fraction of proliferating and differentiating cells was quantified after immunostaining with antibodies to Ki67 and Neurod1, respectively. SIS3 treatment had no effect on the proliferation or differentiation of control GCPs in culture, suggesting that TGFβ-SMAD3 signalling is not an essential regulator of the proliferation or differentiation of these cells. However, the proliferative deficit of Ahr-deficient cells was fully rescued by SIS3 treatment, as was the tendency of these cells to differentiate (Fig. 2c,d). SIS3 treatment did not impact total GCP cell numbers over the course of 24 hours (Fig. S4a).
Loss of Ahr promotes GCP cycle arrest through enhanced activation of the TGF-β/SMAD3 axis. (a) Western blots of total cell lysates of isolated P7 GCPs from control and Ahr cKO cerebella with antibodies specific to phosphorylated S423/S425 residues of SMAD3 and phosphorylated S465/467 residues of SMAD2 proteins as well as antibodies against total SMAD2/3 and GAPDH (loading controls). Molecular weight markers are indicated on the left. (b) Quantification of band optic density for phosphorylated SMAD3, relative to total SMAD2/3, normalized to GAPDH levels. (c) Quantification of %Ki67 positive cells in non-treated (−) and 1 uM SIS3 (+SIS3) treated cultures. (d) Quantification of %Neurod1 positive cells. Data shown in (c,d) is representative of GCPs (on average 100 cells counted from 4 different fields of view from triplicate wells) isolated from 3 animals of each genotype, cultured for 24 hours in triplicate. Data was analyzed by Student’s t test (p < 0.001 (***), p < 0.01 (**), p < 0.05 (*)). Bars represent mean values +/− SEM.
Together, these findings identify a role for AHR in keeping SMAD3 activation in check during normal GCP development. We conclude that SMAD3 hyperactivation is at least in part responsible for the proliferative deficit and enhanced differentiation of Ahr-deficient GCPs.
AHR supresses tumour progression in a SHH medulloblastoma mouse model
Cerebellar GCPs have been identified as the cell of origin for SHH medulloblastoma33,34. In mice, the conditional deletion of Ptch1, encoding the SHH receptor Patched1, which functions as an inhibitor of the SHH pathway, results in hyper-proliferation of GCPs, their rapid transformation and lethal medulloblastoma development in 100% of animals35. To determine whether Ahr has a role in SHH medulloblastoma, we deleted Ahr in these cells together with Ptch1. As previously reported35, 100% of Math1cre; Ptch1f/f (Ptch1 cKO) animals succumbed to medulloblastoma within 3 months of age, with a median survival of 63.5 days (Fig. 3a). By comparison, all Math1cre; Ptch1f/f; Ahrf/f (Ptch1 cKO Ahr cKO) animals died within 40 days of age, with a median survival of 33 days (Fig. 3a). The shorter survival time of animals with Ahr-deficient GCPs was highly significant (p < 0.0001, log-rank test). All animals presented with large medulloblastoma tumours with characteristic classic histology (Fig. 3b).
Ahr deletion in GCPs exacerbates tumorigenesis in a mouse model of SHH medulloblastoma. (a) Kaplan-Meier survival curves for Math1cre; Ptch1f/f (Ptch1 cKO; n = 19) and Math1cre; Ahrf/f; Ptch1f/f (Ptch1 cKO; Ahr cKO; n = 15) mice. (b) Representative haematoxylin/eosin (H&E) staining showing classic histology in both Ptch1 cKO and Ptch1 cKO; Ahr cKO medulloblastomas. (c) Immunohistochemical staining for mitotic marker pH3B (phosphorylated histone H3B) in brown. (d) Quantification of pH3B+ cells/mm2 of tumour (4 non-adjacent sections from 3 tumours of each genotype). (e) Immunostaining for neural differentiation markers synaptophysin, Map2 and NeuN (brown). Statistical comparison of survival curves in panel (a) was performed by log-rank test (p < 0.001 (***)). Data shown is of a single tumor assessed from each animal (n = 3 WT and 3 cKOs), as each animal presented with a single large tumour. Proliferation data in panel (d) was analyzed by Student’s t test (p < 0.05 (*)). Bars represent mean values +/− SEM. Scale bars: 50 um.
Ahr-deficient medulloblastoma tumours have an undifferentiated phenotype
To characterize the salient features that distinguish Ahr-deficient from control tumours, we compared cell proliferation and differentiation. Overall, cellular proliferation was slightly, but significantly increased in Ahr-deficient tumours compared to control tumours (Fig. 3c,d). A comparison of control and Ahr-deficient tumours for steady-state levels of neuronal differentiation markers did not reveal any obvious difference in the levels of differentiation in Ahr-deficient tumours (Fig. 3e).
Ahr controls cancer-propagating cell proliferation and differentiation via TGFβ-SMAD3 inhibition
Next, we asked whether AHR also inhibited SMAD3 activation in SHH medulloblastoma. Immunoblot analysis of tumour lysates found elevated levels of P-SMAD3 in Ahr-deficient samples, compared to controls (Fig. 4a,b). Levels of total SMAD2/3 protein were also increased in Ahr cKO tumours (Fig. S3c,d). Immunostaining of medulloblastoma tissue revealed a salt and pepper distribution of P-SMAD3 positive cells in the tissue, suggesting that only a subset of cells in Ahr cKO tumours responded to TGFβ signals at a given time point (Fig. 4c). When comparing P-SMAD3 immunostaining between control and Ahr cKO medulloblastomas, we found that the number of cells with detectable P-SMAD3, as well as the intensity of P-SMAD3 staining in these cells, were increased in Ahr-deficient tumour tissue (Fig. 4d).
SMAD3 phosphorylation is increased in Ahr-deficient SHH medulloblastomas. (a) Western blots of total cell lysates of isolated end stage medulloblastoma tissue with antibodies specific to phosphorylated S423/S425 residues of SMAD3 (P-SMAD3), total SMAD2/3 and GAPDH proteins. (b) Quantification of band optic density for phosphorylated SMAD3 relative to SMAD2/3 levels, normalized to GAPDH levels. Data is representative of 3 animals/genotype. (c) Immunohistochemical staining of end stage tumour sections with P-SMAD3 antibody (in brown) with methyl green counterstaining nuclei. (d) Quantification of P-SMAD3+ cells/mm2 of tumour tissue (4 non-adjacent sections from 3 tumours of each genotype). Data was analyzed by Student’s t test (p < 0.001 (***)). Bars represent mean values +/− SEM. Scale bars: 100 um.
Given previous studies linking Ahr function with maintenance of the cancer-propagating cell (CPC) compartment36,37 and the evidence supporting a role for Sox2+ CPCs in promoting SHH medulloblastoma aggressiveness20,21, we decided to investigate whether the elevated SMAD3 activity in Ahr-deficient medulloblastoma modulated important CPC properties. To achieve this, we established primary cultures of CPCs from end stage medulloblastomas and maintained these cells in serum-free stem cell medium supplemented with growth factors, as described previously38 (Fig. 5a). As expected, the majority (70–80%) of medulloblastoma CPCs were positive for the neural stem cell marker Sox2 and TGFβ inhibitor treatments had no effect on the proportion of Sox2 expressing cells (Fig. S2a), indicating that Sox2+ cell identity or Sox2 expression were not dependent on TGFβ signalling. Immunostaining of these cells revealed that all Ahr-deficient medulloblastoma CPCs were strongly positive for p-SMAD3, compared to control culture that did not display SMAD3 activation (Fig. 5b). This finding suggested that the AHR pathway suppressed TGFβ-SMAD3 signalling in Sox2+ cells.
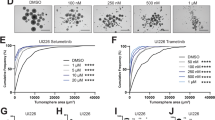
Ahr suppresses SHH MB CPC proliferation by repressing SMAD3 activation. (a) Procedure involved in isolation and maintenance of MB CPCs38. (b) Immunocytochemical staining of proliferating MB CPC cultures for phosphorylated SMAD3 (green) with Hoechst counterstained nuclei (blue). (c) The same cultures were treated for 24 hours with or without TGFβ pathway inhibitors and stained for Ki67 (green), Sox2 (red) and Hoechst (blue). (d) Quantification of %Ki67+ cells in non-treated (−), 500 nM SB43 (+SB43) and 1uM SIS3 (+SIS3) treated cells. (e) Quantification of %Ki67+; Sox2+ (double positive) cells in each culture condition. (f) Quantification of total cell number/field of view based on Hoechst staining. On average 100 cells were counted from 4 different fields of view from triplicate wells of each condition. Data shown is representative of CPCs isolated from 2 animals/genotype with experiments performed in triplicate for each. Data was analyzed by Student’s t test (p < 0.01 (**), p < 0.05 (*)). Bars represent mean values +/− SEM. Scale bars: 10 um (b,c).
In agreement with observations in tumour sections (see Fig. 3d), Ahr-deficient cultures contained significantly higher numbers of proliferating cells compared to controls (Fig. 5c,d). Inclusion of the SMAD3 phosphorylation inhibitor SIS332 in these proliferating cultures reduced levels of cycling to near control levels (Fig. 5c,d). Quantification of the proportion of cycling (Ki67+) Sox2+ CPCs revealed the same trend (Fig. 5e). These data implicating TFGβ/SMAD3 signalling in medulloblastoma CPC proliferation were corroborated by treating the cells with the selective TGF-β receptor I inhibitor SB-431542 (SB43)39 (Fig. 5d,e). Treatment of Ahr-deficient CPCs with both SIS3 and SB43 inhibitors led to a significant reduction in cell numbers (~25% with SIS3 and ~35% with SB43), suggesting that TFGβ/SMAD3 signalling may function by promoting CPC survival. Together, these findings implicated the TGF-β/SMAD3 pathway in mediating the effect of AHR signalling on proliferation and cell survival of medulloblastoma CPCs.
To ask whether Ahr deletion prevented medulloblastoma CPC differentiation via TGFβ-SMAD3 signalling, cells were transferred to a culture medium that promotes the differentiation of these cells38. Under these differentiation conditions, control cells completely lost expression of Sox2 with 7 days, while approximately 60% of Ahr-deficient medulloblastoma CPCs retained high levels of Sox2 expression, indicative of a retention of an undifferentiated phenotype (Fig. 6b,d). Control Ptch1 cKO CSC cultures maintained under these differentiation conditions were still characterised by low levels of SMAD3 activity, compared to Ptch1 cKO; Ahr cKO cells that had high levels of nuclear P-SMAD3 (Fig. 6a).
Ahr-deficient MB CPCs resist differentiation partly through SMAD3 activation. (a) Immunocytochemical staining of 7 DIV (day in vitro) differentiation cultures for P-SMAD3 (green) and Hoechst (blue). Note increased P-SMAD3 signal intensity in Ahr cKO cultures. (b) Staining of Ptch1 cKO and Ptch1 cKO; Ahr cKO cultures with/without 1uM SIS3 for Sox2 (red). (c) Staining of the same cultures for TuJ1/βIII tubulin (green). (d) Quantification of %Sox2+ cells in all culture conditions. (e) Quantification of %TuJ1+ cells in all culture conditions. On average 100 cells were counted from 4 different fields of view from triplicate wells of each condition. Note increased number of Sox2+ and reduced number of TuJ1+ cells in Ahr cKO CPCs and partial rescue with SIS3 treatment. Data shown is representative of CPCs isolated from 2 animals/genotype performed in triplicates. Data was statistically analyzed by Student’s t test (p < 0.001 (***), p < 0.01 (**), p < 0.05 (*)). Bars represent mean values +/− SEM. Scale bars: 10 um (a–c).
Treatment of Ahr cKO cultures with either TGFβ/SMAD inhibitors (SB43 and SIS3) over 7 days significantly reduced the proportion of Sox2+ cells by 30–40% (Fig. 6d). Inhibitor treatments had no effect on Sox2+ cells in control cultures. To confirm and compare levels of differentiation, both control and Ahr cKO CPC cultures were stained for TuJ1/βIII tubulin, a marker of immature neuron differentiation38. After 7 days of differentiation, nearly 80% of control Ptch1 cKO CPCs were positive for TuJ1, while only 5–10% of Ptch1 cKO; Ahr cKO CPCs were positive for this marker (Fig. 6c,e). TGFβ inhibition increased TuJ1 expression in Ahr cKO cultures, while having no effect on control CPC differentiation (Fig. 6c,e). The rescue effect on Ahr cKO cultures was partial, suggesting involvement of other pathways beside TGFβ/SMAD3 in promoting resistance to differentiation in Ahr cKO CPCs.
Taken together, these studies suggested that Ahr deletion in SHH medulloblastoma promoted CPC fate via induction of TGFβ/SMAD3 signalling. Correspondingly, TGFβ/SMAD3 inhibition promoted CPC differentiation.
SHH medulloblastoma patients with high levels of AHRR expression show reduced survival
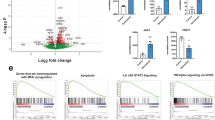
To determine if our findings of a tumour-suppressive role for the AHR pathway may have direct clinical relevance, we examined the expression of AHR in a cohort of human medulloblastomas, profiled by the Clifford group, with the following subgroup distribution: WNT (n = 28), SHH (n = 58), Grp 3 (n = 59), Grp 4 (n = 95). AHR gene expression was significantly higher in the WNT subgroup, with no difference between other subgroups (Fig. 7a). Furthermore, AHR expression levels did not correlate with patient survival in any subgroup (data not shown). Intriguingly, we found that the expression of the AHRR gene, which encodes an AHR Repressor protein, was elevated specifically in the SHH subgroup (Fig. 7b). When examining the association between AHRR expression in SHH tumours and patient survival, we found a statistically significant reduction in patient survival in medulloblastomas with high (>median) AHRR expression (Fig. 7c). This relationship between AHRR expression and survival was specific to the SHH subgroup with no associations found in other subgroups (data not shown). These findings are consistent with a model whereby a reduction of the AHR pathway, either via reducing Ahr expression (as in our mouse model), or increased AHRR expression in human tumours, is associated with more aggressive SHH medulloblastoma tumours and reduced patient survival.
Significantly reduced survival of patients with SHH MB expressing high levels of AHR repressor (AHRR). (a) AHR gene expression comparison across different medulloblastoma subgroups (WNT (n = 28), SHH (n = 58), Grp 3 (n = 59), Grp 4 (n = 95)) from the Clifford cohort. Note significant difference in WNT subgroup. (b) AHRR gene expression comparison across different medulloblastoma subgroups. Note significant difference in SHH subgroup. (c) Kaplan-Meier survival curves from Clifford group patient dataset showing survival rate comparison between SHH MB patients with high and low expression of AHRR for the 56 SHH patients for whom follow-up data was available. (d) AHRR gene expression comparison across different medulloblastoma subgroups (WNT (n = 70), SHH (n = 223), Grp 3 (n = 144), Grp 4 (n = 326)) from the SickKids cohort6. Note significant difference in SHH subgroup. (e) Kaplan-Meier survival curves from Taylor group patient dataset showing survival rate comparison between SHH MB patients with AHRR expression below or above median levels. Note similar trend with Clifford dataset of reduced survival probability with heightened AHRR expression. (f) AHRR gene expression comparison across different medulloblastoma subgroups (WNT (n = 53), SHH (n = 112), Grp 3 (n = 94), Grp 4 (n = 164)) from the Kool cohort. Note significant difference in SHH subgroup. (g) Kaplan-Meier survival curves from Kool group patient dataset showing survival rate comparison between SHH MB patients with AHRR expression below or above median levels. Data in a,b and d were statistically analyzed by an Anova test and data in c,e and g by log rank test.
Finally, we asked if we could replicate our human medulloblastoma findings in an independent, larger patient cohort. An analysis of 172 SHH tumours from the Taylor group in Toronto, confirmed both the increased expression of AHRR in SHH tumours, compared to other subgroups (Fig. 7d), as well as reduced survival of patients with high AHRR expression (Fig. 7e). The same AHRR expression profile was observed across medulloblastoma subgroups in a third, independent cohort profiled by the Kool group in Heidelberg (Fig. 7f), however no correlation between AHRR expression levels and survival was observed in this particular cohort of SHH medulloblastomas (Fig. 7g).
Discussion
Here we identified a critical role for Ahr in preventing activation of the TGFβ mediator SMAD3, both in primary cerebellar GCPs and GCP-derived SHH meduloblastoma. We further found that CPCs derived from Ahr-deficient tumours exhibited very high levels of P-SMAD3 compared to tumours with intact Ahr. Ahr deletion in GCPs together with the cancer-initiating Ptch1 gene deletions dramatically reduced survival, identifying a potent tumour-suppressive role for Ahr. CPCs from these Ahr-deficient tumours were refractory to differentiation in vitro. Most importantly, pharmacological inhibition of the TGFβ-SMAD3 pathway was sufficient to drive Ahr-deficient CPCs towards differentiation, identifying this pathway as a potentially viable therapeutic target for aggressive medulloblastoma subtypes with reduced AHR pathway activity and elevated TGFβ-SMAD3 signalling. Transcriptomic analyses of human SHH medulloblastomas indeed identified a substantial subset of SHH primary tumours with high AHRR expression and poor prognosis in two independent patient cohorts. As these aggressive SHH medulloblastoma subtypes are highly resistant to conventional therapies, future studies to fully characterise these AHRRhigh tumours, establish to what extent they resemble Ahr-deficient SHH tumours in the mouse, and explore the potential of TGFβ-SMAD3 pathway inhibition will be important.
Several mechanistic questions remain to be answered. Exactly how Ahr deletion leads to specific activation of SMAD3, and not SMAD2 is not known. This apparently exquisite specificity argues against a general induction of TGFβ ligands or membrane receptors, which would be expected to activate both SMAD2 and SMAD3. The increase in total SMAD2/3 protein levels suggests Ahr might also function by either suppressing SMAD2/3 transcription in the tumour context or promoting SMAD2/3 proteolytic turnover.
Our findings further support the idea that the role of the AHR pathway is highly context-specific. In primary GCPs, Ahr deletion leads to reduced proliferation and enhanced differentiation, while in SHH medulloblastomas derived from these cells, Ahr deletion has the opposite effect. These observations are particularly important in the light of a previous study showing that Ahr knockdown in the SHH-like medulloblastoma cell line DAOY resulted in reduced cell proliferation25.
It should be noted that other genes in the AHR pathway have been implicated in medulloblastoma. In particular, the Arnt (Aryl hydrocarbon receptor nuclear translocator) gene, which encodes an AHR interacting protein necessary for its function, can promote leptomeningeal metastatic dissemination when overexpressed in SHH medulloblastomas in the mouse40. Whether these effects are as a result of modulation of the AHR pathway remains to be determined.
It is intriguing to consider our observations in SHH medulloblastoma CPCs in the context of other brain tumours. Gramatzki et al. have shown that AHR inhibition in glioma cells also resulted in upregulation of the TGFβ-SMAD pathway30. Elevated TFGβ-SMAD signalling promotes glioma stem cell self-renewal and is associated with more aggressive gliomas and poorer survival41,42. Together, these findings imply some conservation in the role of AHR in repressing the TGFβ-SMAD pathway in brain tumour stem cells and suggest the possibility that inactivation or repression of this pathway may represent a mechanism whereby tumour cells retain a stem-like character and become resistant to differentiation. This study did not assess the effects of TGFβ-SMAD pathway modulation on apoptosis, however a significant decrease in Ahr cKO CPC number observed with SB43 and SIS3 treatment suggests that enhanced CPC survival may be an important mechanism of increased tumorigenicity in Ahr-deficient SHH medulloblastoma.
The role of the TGFβ-SMAD pathway specifically in medulloblastoma remains unclear. The present study links hyper-activation of this pathway in SHH medulloblastoma with resistance to differentiation and poor prognosis. This finding is in disagreement with Aref et al. who suggested that nuclear SMAD3 localisation as a proxy read-out of SMAD3 activation in SHH medulloblastoma samples correlated with good prognosis43. This study only assessed SHH medulloblastomas from 35 patients and using nuclear localisation of SMAD3 as a read-out of pathway activation may not be ideal. Clearly, a larger study to assess P-SMAD3 immunoreactivity will be important. Interestingly, elevated TGFβ signalling has also been implicated as a driver of Group 3 medulloblastoma44, suggesting that this pathway may also play important roles in other medulloblastoma subtypes.
Our human transcriptomic analyses identified AHRR expression as a novel biomarker for aggressive SHH medulloblastoma in two of three independent cohorts investigated. Demographic differences and the mixed therapies deployed within these retrospective cohorts may explain the lack of consistent observations in all three cohorts, and these encouraging initial findings now require prospective validation in clinical trials-based cohorts. Determining to what extent high AHRR expression in human SHH medulloblastoma is associated with elevated TGFβ-SMAD3 signalling will be an important next step to identify patients that may benefit from TGFβ-SMAD3 inhibition and/or AHR agonist therapies.
Materials and Methods
Animals
Math1cre45, Ptch1 flox46, and Ahr flox47 mouse lines have been described and were genotyped by PCR using tail or ear DNA extracted using proteinase K digestion or the HotSHOT method48. PCR primers used are indicated in the table below. Ptch1 cKO mice were generated by crossing Math1cre and Ptch1f/f mice, followed by crosses of the resultant Math1cre; Ptch1f/+ with Ptch1f/f animals. Ahr cKO medulloblastoma mice were generated by crossing Math1cre and Ahrf/f; Ptch1f/f mice followed by crosses of the resulting Math1cre; Ahrf/+; Ptch1f/+ mice with Ahrf/f; Ptch1f/f animals. Mice were bred and maintained according to Home Office regulations in New Hunt’s House, Biological Services Unit, King’s College, London. For postnatal stages the day of birth was designated postnatal day 0 (P0). All mice on tumour watch were monitored closely for symptoms of medulloblastoma and were sacrificed upon discovery of symptoms such as hydrocephalus, weight loss and ataxic gait49. The institutional Local Ethical Review Panel and the UK Home Office approved all experimental procedures (Project licence numbers: 70/7506 and P8DC5B496). All procedures were carried out by a personal licence holder.
Gene | Forward Primer | Reverse Primer |
Cre | 5′-CCTGGAAAATGCTTCTGTCCG-3′ | 5′-CAGGGTGTTATAAGCAATCCC-3′ |
Ahrflox | 5′-CAGTGGGAATAAGGCAAGAGTGA-3′ | 5′-GGTACAAGTGCACATGCCTGC-3′ |
Ptch1flox | 5′-CCACCAGTGATTTCTGCTCA-3′ | 5′-AGTACGAGGCATGCAAGACC-3′ |
Thermal cycles for Cre and Ptch1flox genotyping were as follows: 94 °C, 10 minutes; 40× (94 °C, 45 sec; 57 °C, 45 sec; 72 °C, 60 sec); 72 °C, 7 minutes. Thermal cycles for Ahrflox genotyping were: 94 °C, 3 minutes; 35 × (94 °C, 30 sec; 69 °C, 60 sec; 72 °C, 60 sec); 72 °C, 2 minutes.
Tissue processing and histology
Brains were dissected in ice cold PBS and fixed in 4% paraformaldehyde (PFA) at 4 °C. Samples were dehydrated, cleared and infiltrated with paraffin wax using a Leica ASP300 Tissue Processor, followed by sagittal embedding in paraffin. Paraffin tissue blocks were sectioned using a microtome (Leica RM2145). Serial sections were cut at 4–10 μm thickness and mounted onto glass slides (SuperfrostPlus®, VWRTM).
Immunohistochemistry
3, 3′-diaminobenzidine (DAB) immunohistochemistry
Sections were deparaffinised and rehydrated through a series of graded ethanols to PBS. Endogenous peroxidases were blocked in a solution containing 3% H2O2 (stock 30%) and 10% methanol in PBS for 15 minutes and were subsequently rinsed with dH2O. Sections were then heated in the microwave at full power in a 10 mM sodium citrate solution (pH 6.0) (4 × 5 mins) to break methylene bridges associated with the fixation process and expose antigenic sites. After cooling for 20 minutes at room temperature, cells were then permeabilised using 0.2% PBSTx (Triton®X-100 in PBS) (1 × 10 minutes) and non-specific antibody binding was blocked by incubating slides in 10% goat serum in PBSTx for 1 hour at room temperature. Sections were then incubated with the primary antibody diluted in 5% goat serum in PBSTx overnight at 4 oC. The following day unbound antibody was removed using three 10 minute 0.1% PBSTx washes and sections were then incubated in the appropriate biotinylated secondary antibody (1/200) diluted in 5% goat serum in 0.1% PBSTx. After 1 hour at room temperature unbound secondary antibody was washed off using PBS (3 × 10 minutes). The signal was then amplified using the VECTASTAIN Avidin Biotin Complex (ABC) kit by incubating sections in a solution containing 1/200 dilutions of A and B in PBS for 1 hour at room temperature. Sections were then washed in PBS (3 × 10 minutes) and visualised with 0.03% 3, 3′- diaminobenzidine (DAB) substrate (stock 30% in Tris-HCl)/ 0.0003% hydrogen peroxide (stock 30%). After visualization sections were then counterstained with Methyl Green (1 × 5 minutes), briefly washed under running water and then dehydrated in a series of ascending ethanols, cleared in xylene, mounted using DPX and imaged with a Nikon Eclipse 80i microscope.
Immunofluorescence
Paraffin sections
Sections were deparaffinised and rehydrated as above and washed with PBS (2 × 5 mins). Antigen retrieval was performed by heating sections in a solution of 10 mM sodium citrate (pH 6.0) for 4 × 5 minutes at full power in the microwave. Sections were then left to cool to room temperature for 20 minutes. Tissue was permeabilized in 0.2% PBSTx and blocked in 10% heat inactivated goat serum in PBSTx for 1 hour before incubating in primary antibody in 5% goat serum in PBSTx overnight at 4 oC. The following day, unbound antibody was removed using 3 × 10 minute PBS washes. Slides were incubated with Alexa-Fluor-labelled secondary antibodies (1/200, Life technologies) in 5% goat serum in PBSTx. Unbound secondary antibody was then washed off with 3 × 10 minute PBS washes. The nuclear counterstain, 4′-6-diamidino-2-phenylindole (Dapi) (1:5000, Invitrogen) was added to the final wash and slides were mounted using Citifluor (www.citifluor.com). Tumour histology for differentiation markers Synaptophysin, MAP2 and NeuN were performed on a Ventana Medical System Benchmark automated immunostainer as described50.
Fixed cells
Cells fixed on coverslips with 4% PFA were initially washed with 2 × 10 minute PBS washes. They were then permeabilized in 0.2% PBSTx and blocked in 10% heat inactivated goat serum in PBSTx for 1 hour before incubating in primary antibody in 5% goat serum in PBSTx overnight at 4oC. The following day, unbound antibody was removed using 3 × 10 minute PBS washes. Slides were incubated with Alexa-Fluor-labelled secondary antibodies (1/200, Life technologies) in 5% goat serum in PBSTx. Unbound secondary antibody was then washed off with 3 × 10 minute PBS washes. The nuclear counterstain, 4′-6-diamidino-2-phenylindole (Dapi) (1:5000, Invitrogen) was added to the final wash and slides were mounted using Citifluor (www.citifluor.com). Fluorescent images were captured using Nikon Eclipse 80i with Nikon Y-QT Hamamatsu C4742-95 camera.
Antibodies
Primary
Primary antibodies used were as follows: rabbit anti-GAPDH (1/2500, Abcam, ab9485), rabbit anti-phospho histone H3B (1/200, NEB, 9701S), rat anti-BrdU (1/50, Abcam, ab6326), mouse anti-BrdU (1/100, BD, 347580), mouse anti-NeuN (1/2000, Chemicon), mouse anti-Map2 (1/500, Chemicon), rabbit anti-Ki67 (1/1000, Abcam, ab15580), rabbit anti-P-SMAD2/3 (1/1000, NEB, D27F4), rabbit anti-SMAD2/3 (1/1000, NEB, 3102S), mouse anti-Neurod1 (1/100, Abcam, ab60704), rabbit anti-synaptophysin (Pe-dilued, Zymed, 080130), rabbit anti-P-SMAD3 (1/100, Abcam, ab52903), rabbit anti-TuJ1 (1/500, Abcam, ab18207) and mouse anti-Sox2 (1/500, Abcam, ab79351).
Secondary
For western blots primary antibodies were detected using polyclonal goat anti-rabbit IgG horse radish peroxidase (HRP) conjugated secondary antibody (1/2000, Thermofisher, 65-6120). For DAB immunohistochemistry primary antibodies were detected using polyclonal goat anti-mouse IgG biotinylated secondary antibody (1/200, Dako, E0433), polyclonal goat anti- rabbit IgG biotinylated secondary antibody (1/200, Dako, E0432). For immunofluorescence primary antibodies were detected using Alexafluor-488 goat anti-rabbit IgG (Invitrogen, 1/200, A11034), Alexafluor-568 goat anti-mouse IgG (Invitrogen, 1/200, A21124), Alexafluor-488 goat anti-mouse IgG (Invitrogen, 1/200, A11001), Alexafluor-568 goat anti-rabbit IgG (Invitrogen, 1/200, A11011) secondary antibodies.
Western blots
Purified GCPs were isolated from P7 cerebella and either whole cell protein or subcellular fractions were prepared by lysing in N-PER lysis buffer (Thermofisher) or subcellular fractionation buffers (Thermofisher) respectively, following the manufacturer’s instructions. All buffers contained protease inhibitors (PMSF, Pepstatin A, Leupeptin, Aprotinin; Roche) and a phosphatase inhibitor cocktail (Sigma). Protein loading samples were made by diluting samples in Laemmli buffer containing 10% β-mercaptoethanol, followed by boiling at 95 °C for 5 minutes. Samples were loaded (10 µg total protein per lane) onto a Mini-PROTEAN pre-cast gel (Bio-Rad) and resolved using gel electrophoresis. Protein was transferred to a nitrocellulose membrane (Bio-Rad) which was then blocked in 5% non-fat milk powder (Bio-Rad) or 3% bovine serum albumin (BSA, Sigma) in TBS with 0.1% Tween-20 (TBST) for one hour at room temperature, followed by incubation with primary antibodies diluted in 3% BSA/TBST overnight at 4 °C. The next day the membranes were washed 3 × 10 mins in TBST, followed by incubation in secondary antibodies diluted in 5% non-fat milk powder in TBST for one hour at room temperature. Membranes were subsequently washed again 3 × 10 minutes in TBST and HRP was detected with Clarity ECL reagent (Bio-Rad) and the membranes imaged using a Bio-Rad ChemiDoc system. Relative protein quantity was calculated using Bio-Rad ImageLab software.
Cerebellar GCP isolation
GCPs were isolated as described previously51. The cerebella from P7 control and Math1cre; Ahrf/f pups were dissected in ice-cold DPBS. The lobules that retain Ahr expression (IX + X) along with the flocculus and paraflocculus were removed. Each cerebellum was processed separately. Papain-I (100 U in 10 ml DPBS) was dissolved in 10 ml of DPBS at 37 °C. Once dissolved, 200 μl of DNaseI (12,500 U/ml, Sigma) was added and the Papain-I enzyme was activated by adding L-Cysteine (2 mg/10 ml). After adjusting the pH to using 2 N NaOH, cells were dissociated by incubating the cerebella in the Papain-I containing solution for 30 minutes at 37 °C followed by trituration in ovo solution (2 mg/ml ovomucoid, 125 U/ml DNaseI in DPBS). Dissociated cells were then centrifuged for 10 minutes at 1000 rpm. The supernatant was then aspirated and the pellet suspended in DPBS-BSA (DPBS containing 1% BSA). The suspension was then passed through a cell strainer before underlaying the solution containing the dissociated cells with 35% Percoll followed by 65% Percoll. Cells were then separated according to size by centrifugation (12 minutes, 2500 rpm). The layer constituting the interphase between the 35% and 65% Percoll layers, containing GCPs, was then removed and placed in a separate falcon tube containing 14 ml of DPBS-BSA. GCPs were counted using a haemocytometer, the solution containing GCPs was then centrifuged (10 minutes, 1400 rpm) and the supernatant aspirated.
Cell culture
Purified GCPs were cultured in Neurobasal media (Thermofisher) supplemented with B27 (Thermofisher), exogenous SHH, glutamine (Thermofisher) and penicillin/streptomycin (Sigma). The SHH came from supernatant obtained from conditioned media from HEK293T cells transfected with pcDNA3.1 ShhN plasmid. pcDNA3.1 ShhN was a kind gift from Philip Beachy (Addgene plasmid # 37680). Culture vessels were pre-coated with poly-D-lysine (Sigma) before cell seeding. Media was changed every other day to maintain growth conditions. Omission of exogenous SHH promoted differentiation. CPCs were isolated from end-stage tumours as described in Fig. 5a. CPCs were cultured as described in38. Briefly, enriched CPCs were cultured in Neurobasal media supplemented with B27 (Thermofisher), N2 (Thermofisher), bFGF (Peprotech), EGF (Peprotech), glutamine (Thermofisher) and penicillin/streptomycin (Sigma) to maintain growth conditions. Media was changed every other day in expanding cultures. Culture vessels were pre-coated with 0.1% gelatin (Sigma) before cell seeding. Upon switching the medium to 10% serum in DMEM (Thermofisher) the cells underwent differentiation. In experimental cultures SB-431542 (SB43) (Sigma) or SIS3 (Sigma) were added to some culture wells as part of a daily change of medium. SB43 functions as an inhibitor of the transforming growth factor-beta superfamily type I (TGFβRI) receptor39 while SIS3 is a specific inhibitor of SMAD3 phosphorylation and its interaction with SMAD432.
Quantitative analysis
Ki67+ and Neurod1+ cells quantified in Figs. 1b,c and 2c,d were counted from four different fields of view/quadrants (top to bottom, left to right) from triplicate wells (of 24 well plates) of each condition. Hoechst counterstained cells were counted alongside and the final % positivity calculated and averaged for each condition. The Hoechst staining quantification is from the same fields with filter change. Neurites were traced using the Simple Neurite Tracer plugin in ImageJ and the path lengths of traced neurites from each cell in the field of view were converted to a micron scale before calculating the mean length. The starting point of the neurite was taken as the point of incidence from the cell soma. The data was assessed by a single, blinded researcher, and the experiment was performed in three independent instances from three separate control and cKO cerebella. Proliferation data in Fig. 3d was quantified by counting the number of pH3B+ cells in 100um x 100um squares from every 30th section (10um thick sections) of three tumours of each genotype (each from a separate animal), followed by averaging. The same method was used to obtain data in Fig. 4d. Ki67+, Sox2+ and TuJ1+ CPCs quantified in Figs. 5d,e, 6d,e were counted in the same manner as the GCP data described above. Fluorescent images were captured using Nikon Eclipse 80i with Nikon Y-QT Hamamatsu C4742-95 camera. Acquired fluorescent images from cell culture experiments were subjected to an identical intensity threshold, as judged by the investigator, in ImageJ (separately for each antigen) before counting positive cells. The experimenter was blinded for the IHC/ICC quantification in (Figs. 1, 3 and 4) and data quantification in the cell culture experiments (Figs. 2 and 5) was done and repeated by another researcher in double-blind fashion where neither of the researchers was aware of the culture conditions or group identity.
Image processing
Images were processed using Adobe Photoshop CC 2017 and figures assembled in Adobe Illustrator CC 2017.
Statistics
Statistical analysis was carried out and graphs generated using GraphPad Prism 5®. Most data were analysed using a Student’s t-test with the exception of the Kaplan-Meier survival analysis which used the log-rank test or Cox regression analysis. P < 0.05 was considered significant.
Transcriptomic analyses from human data
Read counts for AHR and AHRR expression were produced by aligning paired end RNA-seq (~90 M read/sample Illumina HiSeq 2500) reads to HG19 genome using STAR-align52. Read counts were produced using HT-SEQ-count. DESeq 2 (R/Bioconductor) was used to normalise reads to library size and variance stabilised data (VSD) was generated using the vsd function. Statistical testing for differential expression across groups was performed using an ANOVA test. Affymetrix expression data were obtained from previously reported series through GEO accession numbers GSE103278, GSE1299253, GSE3741854, GSE4924355 and published previously56. All data were MAS5.0 normalized and analysed using the genomics analysis and visualization platform R2 (http://r2.amc.nl). For survival analyses the Kaplan scanning tool in R2 was used that identifies the optimal cut off in expression in a dataset that results in the lowest log rank p value in overall survival analyses between the subset with high expression and the subset with low expression of the gene of interest. Log rank p-value is corrected for multiple testing using the Bon Ferroni method.
Ethical approval
All animal experiments were approved by the King’s College London Animal Welfare and Ethical Review Board (AWERB) and the UK Home Office (Project licence P8DC5B496), in accordance with the relevant guidelines and practises. For studies using human tissue and clinical information, all methods were carried out in accordance with relevant guidelines and regulations, all experimental protocols were approved by a named institutional and/or licensing committee and informed consent was obtained from all participants or their legal guardians. For the Newcastle cohort, tumour samples were provided by the UK CCLG as part of CCLG-approved biological study BS-2007–04. Tumour investigations were done with approval from Newcastle North Tyneside Research Ethics Committee (study reference 07/Q0905/71). For the Toronto cohort, medulloblastoma samples were collected at diagnosis after obtaining informed consent from subjects as part of the Medulloblastoma Advanced Genomics International Consortium. Approval was obtained from institutional research ethics boards at all contributing institutions as outlined by Cavalli et al.6. For the Heidelberg cohort, all tumors were collected after receiving informed consent according to ICGC guidelines and approved by the institutional review board of contributing centers.
References
Gajjar, A. J. & Robinson, G. W. Medulloblastoma-translating discoveries from the bench to the bedside. Nat Rev Clin Oncol 11, 714–722, https://doi.org/10.1038/nrclinonc.2014.181 (2014).
Gajjar, A. et al. Risk-adapted craniospinal radiotherapy followed by high-dose chemotherapy and stem-cell rescue in children with newly diagnosed medulloblastoma (St Jude Medulloblastoma-96): long-term results from a prospective, multicentre trial. The Lancet. Oncology 7, 813–820, https://doi.org/10.1016/S1470-2045(06)70867-1 (2006).
Mulhern, R. K., Merchant, T. E., Gajjar, A., Reddick, W. E. & Kun, L. E. Late neurocognitive sequelae in survivors of brain tumours in childhood. The Lancet. Oncology 5, 399–408, https://doi.org/10.1016/S1470-2045(04)01507-4 (2004).
Hoppe-Hirsch, E. et al. Medulloblastoma in childhood: progressive intellectual deterioration. Child’s nervous system: ChNS: official journal of the International Society for Pediatric Neurosurgery 6, 60–65 (1990).
Schwalbe, E. C. et al. Novel molecular subgroups for clinical classification and outcome prediction in childhood medulloblastoma: a cohort study. Lancet Oncol 18, 958–971, https://doi.org/10.1016/S1470-2045(17)30243-7 (2017).
Cavalli, F. M. G. et al. Intertumoral Heterogeneity within Medulloblastoma Subgroups. Cancer Cell 31, 737–754 e736, https://doi.org/10.1016/j.ccell.2017.05.005 (2017).
Taylor, M. D. et al. Molecular subgroups of medulloblastoma: the current consensus. Acta Neuropathol 123, 465–472, https://doi.org/10.1007/s00401-011-0922-z (2012).
Kool, M. et al. Integrated genomics identifies five medulloblastoma subtypes with distinct genetic profiles, pathway signatures and clinicopathological features. PLoS One 3, e3088, https://doi.org/10.1371/journal.pone.0003088 (2008).
Ramaswamy, V. et al. Risk stratification of childhood medulloblastoma in the molecular era: the current consensus. Acta neuropathologica 131, 821–831, https://doi.org/10.1007/s00401-016-1569-6 (2016).
Zhukova, N. et al. Subgroup-specific prognostic implications of TP53 mutation in medulloblastoma. J Clin Oncol 31, 2927–2935, https://doi.org/10.1200/JCO.2012.48.5052 (2013).
Korshunov, A. et al. Biological and clinical heterogeneity of MYCN-amplified medulloblastoma. Acta Neuropathol 123, 515–527, https://doi.org/10.1007/s00401-011-0918-8 (2012).
Hsu, S. H. et al. Aryl hydrocarbon receptor promotes hepatocellular carcinoma tumorigenesis by targeting intestine-specific homeobox expression. Molecular carcinogenesis 56, 2167–2177, https://doi.org/10.1002/mc.22658 (2017).
Schiering, C. et al. Feedback control of AHR signalling regulates intestinal immunity. Nature 542, 242–245, https://doi.org/10.1038/nature21080 (2017).
Bradshaw, T. D. & Bell, D. R. Relevance of the aryl hydrocarbon receptor (AhR) for clinical toxicology. Clinical toxicology 47, 632–642, https://doi.org/10.1080/15563650903140423 (2009).
Stockinger, B., Di Meglio, P., Gialitakis, M. & Duarte, J. H. The aryl hydrocarbon receptor: multitasking in the immune system. Annu Rev Immunol 32, 403–432, https://doi.org/10.1146/annurev-immunol-032713-120245 (2014).
Opitz, C. A. et al. An endogenous tumour-promoting ligand of the human aryl hydrocarbon receptor. Nature 478, 197–203, https://doi.org/10.1038/nature10491 (2011).
Murray, I. A., Patterson, A. D. & Perdew, G. H. Aryl hydrocarbon receptor ligands in cancer: friend and foe. Nat Rev Cancer 14, 801–814, https://doi.org/10.1038/nrc3846 (2014).
Dere, E., Lo, R., Celius, T., Matthews, J. & Zacharewski, T. R. Integration of genome-wide computation DRE search, AhR ChIP-chip and gene expression analyses of TCDD-elicited responses in the mouse liver. BMC Genomics 12, 365, https://doi.org/10.1186/1471-2164-12-365 (2011).
Sakurai, S., Shimizu, T. & Ohto, U. The crystal structure of the AhRR-ARNT heterodimer reveals the structural basis of the repression of AhR-mediated transcription. J Biol Chem 292, 17609–17616, https://doi.org/10.1074/jbc.M117.812974 (2017).
Vanner, R. J. et al. Quiescent sox2(+) cells drive hierarchical growth and relapse in sonic hedgehog subgroup medulloblastoma. Cancer Cell 26, 33–47, https://doi.org/10.1016/j.ccr.2014.05.005 (2014).
Ahlfeld, J. et al. Sox2 requirement in sonic hedgehog-associated medulloblastoma. Cancer research 73, 3796–3807, https://doi.org/10.1158/0008-5472.CAN-13-0238 (2013).
Al-Dhfyan, A., Alhoshani, A. & Korashy, H. M. Aryl hydrocarbon receptor/cytochrome P450 1A1 pathway mediates breast cancer stem cells expansion through PTEN inhibition and beta-Catenin and Akt activation. Mol Cancer 16, 14, https://doi.org/10.1186/s12943-016-0570-y (2017).
Rentas, S. et al. Musashi-2 attenuates AHR signalling to expand human haematopoietic stem cells. Nature 532, 508–511, https://doi.org/10.1038/nature17665 (2016).
Dever, D. P. et al. Aryl hydrocarbon receptor deletion in cerebellar granule neuron precursors impairs neurogenesis. Dev Neurobiol 76, 533–550, https://doi.org/10.1002/dneu.22330 (2016).
Dever, D. P. & Opanashuk, L. A. The aryl hydrocarbon receptor contributes to the proliferation of human medulloblastoma cells. Mol Pharmacol 81, 669–678, https://doi.org/10.1124/mol.111.077305 (2012).
Whittaker, D. E. et al. The chromatin remodeling factor CHD7 controls cerebellar development by regulating reelin expression. The Journal of clinical investigation 127, 874–887, https://doi.org/10.1172/JCI83408 (2017).
Leto, K. et al. Consensus Paper: Cerebellar Development. Cerebellum 15, 789–828, https://doi.org/10.1007/s12311-015-0724-2 (2016).
Dehmelt, L. & Halpain, S. The MAP2/Tau family of microtubule-associated proteins. Genome Biol. 6, 204, https://doi.org/10.1186/gb-2004-6-1-204 (2005).
Chang, X. et al. Ligand-independent regulation of transforming growth factor beta1 expression and cell cycle progression by the aryl hydrocarbon receptor. Mol. Cell. Biol. 27, 6127–6139, https://doi.org/10.1128/MCB.00323-07 (2007).
Gramatzki, D. et al. Aryl hydrocarbon receptor inhibition downregulates the TGF-beta/Smad pathway in human glioblastoma cells. Oncogene 28, 2593–2605, https://doi.org/10.1038/onc.2009.104 (2009).
Derynck, R. & Zhang, Y. E. Smad-dependent and Smad-independent pathways in TGF-beta family signalling. Nature 425, 577–584, https://doi.org/10.1038/nature02006 (2003).
Jinnin, M., Ihn, H. & Tamaki, K. Characterization of SIS3, a novel specific inhibitor of Smad3, and its effect on transforming growth factor-beta1-induced extracellular matrix expression. Mol. Pharmacol. 69, 597–607, https://doi.org/10.1124/mol.105.017483 (2006).
Schuller, U. et al. Acquisition of granule neuron precursor identity is a critical determinant of progenitor cell competence to form Shh-induced medulloblastoma. Cancer Cell 14, 123–134, https://doi.org/10.1016/j.ccr.2008.07.005 (2008).
Oliver, T. G. et al. Loss of patched and disruption of granule cell development in a pre-neoplastic stage of medulloblastoma. Development 132, 2425–2439, https://doi.org/10.1242/dev.01793 (2005).
Yang, Z. J. et al. Medulloblastoma can be initiated by deletion of Patched in lineage-restricted progenitors or stem cells. Cancer Cell 14, 135–145, https://doi.org/10.1016/j.ccr.2008.07.003 (2008).
Stanford, E. A. et al. The role of the aryl hydrocarbon receptor in the development of cells with the molecular and functional characteristics of cancer stem-like cells. BMC biology 14, 20, https://doi.org/10.1186/s12915-016-0240-y (2016).
Prud’homme, G. J. et al. Breast cancer stem-like cells are inhibited by a non-toxic aryl hydrocarbon receptor agonist. PloS one 5, e13831, https://doi.org/10.1371/journal.pone.0013831 (2010).
Huang, X., Ketova, T., Litingtung, Y. & Chiang, C. Isolation, enrichment, and maintenance of medulloblastoma stem cells. J. Vis. Exp., https://doi.org/10.3791/2086 (2010).
Inman, G. J. et al. SB-431542 is a potent and specific inhibitor of transforming growth factor-beta superfamily type I activin receptor-like kinase (ALK) receptors ALK4, ALK5, and ALK7. Molecular pharmacology 62, 65–74 (2002).
Jenkins, N. C. et al. Genetic drivers of metastatic dissemination in sonic hedgehog medulloblastoma. Acta Neuropathol. Commun. 2, 85, https://doi.org/10.1186/s40478-014-0085-y 10.1186/PREACCEPT-1860372034135162 (2014).
Bruna, A. et al. High TGFbeta-Smad activity confers poor prognosis in glioma patients and promotes cell proliferation depending on the methylation of the PDGF-B gene. Cancer Cell 11, 147–160, https://doi.org/10.1016/j.ccr.2006.11.023 (2007).
Penuelas, S. et al. TGF-beta increases glioma-initiating cell self-renewal through the induction of LIF in human glioblastoma. Cancer Cell 15, 315–327, https://doi.org/10.1016/j.ccr.2009.02.011 (2009).
Aref, D. et al. Canonical TGF-beta pathway activity is a predictor of SHH-driven medulloblastoma survival and delineates putative precursors in cerebellar development. Brain Pathol. 23, 178–191, https://doi.org/10.1111/j.1750-3639.2012.00631.x (2013).
Northcott, P. A. et al. Subgroup-specific structural variation across 1,000 medulloblastoma genomes. Nature 488, 49–56, https://doi.org/10.1038/nature11327 (2012).
Matei, V. et al. Smaller inner ear sensory epithelia in Neurog 1 null mice are related to earlier hair cell cycle exit. Dev Dyn 234, 633–650, https://doi.org/10.1002/dvdy.20551 (2005).
Ellis, T. et al. Patched 1 conditional null allele in mice. Genesis 36, 158–161, https://doi.org/10.1002/gene.10208 (2003).
Walisser, J. A., Glover, E., Pande, K., Liss, A. L. & Bradfield, C. A. Aryl hydrocarbon receptor-dependent liver development and hepatotoxicity are mediated by different cell types. Proceedings of the National Academy of Sciences of the United States of America 102, 17858–17863, https://doi.org/10.1073/pnas.0504757102 (2005).
Truett, G. E. et al. Preparation of PCR-quality mouse genomic DNA with hot sodium hydroxide and tris (HotSHOT). BioTechniques 29(52), 54 (2000).
Huang, X. et al. Intracranial orthotopic allografting of medulloblastoma cells in immunocompromised mice. Journal of visualized experiments: JoVE, https://doi.org/10.3791/2153 (2010).
Sutter, R. et al. Cerebellar stem cells act as medulloblastoma-initiating cells in a mouse model and a neural stem cell signature characterizes a subset of human medulloblastomas. Oncogene 29, 1845–1856, https://doi.org/10.1038/onc.2009.472 (2010).
Yu, T., Yaguchi, Y., Echevarria, D., Martinez, S. & Basson, M. A. Sprouty genes prevent excessive FGF signalling in multiple cell types throughout development of the cerebellum. Development 138, 2957–2968, https://doi.org/10.1242/dev.063784 (2011).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21, https://doi.org/10.1093/bioinformatics/bts635 (2013).
Fattet, S. et al. Beta-catenin status in paediatric medulloblastomas: correlation of immunohistochemical expression with mutational status, genetic profiles, and clinical characteristics. The Journal of pathology 218, 86–94, https://doi.org/10.1002/path.2514 (2009).
Robinson, G. et al. Novel mutations target distinct subgroups of medulloblastoma. Nature 488, 43–48, https://doi.org/10.1038/nature11213 (2012).
Kool, M. et al. Genome sequencing of SHH medulloblastoma predicts genotype-related response to smoothened inhibition. Cancer cell 25, 393–405, https://doi.org/10.1016/j.ccr.2014.02.004 (2014).
Northcott, P. A. et al. The whole-genome landscape of medulloblastoma subtypes. Nature 547, 311–317, https://doi.org/10.1038/nature22973 (2017).
Acknowledgements
This work was supported by grants from the Wellcome Trust (091475) to MAB, the Medical Research Council (MR/N000528/1) to SM and MAB, Cancer Research UK (C8464/A13457 and C8464/A23391) to DW and SCC, and LoveOliver to DW and SCC. The INSTINCT network (SCC and DW) is co-funded by The Brain Tumour Charity, Great Ormond Street Children’s Charity, and Children with Cancer UK. NS was supported by a King’s College London International Graduate School studentship. We thank David Rowitch for the Math1cre mouse line and Brandon Wainwright for the Ptch1 line. We thank Stephen Crosier for assistance with neuropathological assessments of tumours, John Whittingham for technical assistance, Caroline Hill for advice on TGFβ-SMAD reagents and our laboratory colleagues for comments on the manuscript. Human tumour investigations were conducted as part of a Children’s Cancer and Leukaemia Group-approved biological study, and with approval from Newcastle and North Tyneside Research Ethics Committee (Study reference 07/Q0905/71). VR is supported by operating grants from the Canadian Institutes of Health Research, the American Brain Tumor Association, Brain Tumour Foundation of Canada, Nelina’s Hope and the CR Younger Foundation. MDT is supported by operating funds from the National Institutes of Health (R01CA148699 and R01CA159859) and the Pediatric Brain Tumor Foundation.
Author information
Authors and Affiliations
Contributions
N.S. and M.A.B. conceived the study and designed animal and in vitro experiments. N.S. performed the animal and in vitro experiments and analysed the data. M.S., V.R., M.K., D.W., M.T. and S.C.C. provided human datasets and performed bioinformatic analyses. B.S. and C.H. provided expertise, reagents and supervision related to AHR biology and SM provided expertise and analysed immunohistochemistry together with N.S. N.S. and M.A.B. wrote the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Sarić, N., Selby, M., Ramaswamy, V. et al. The AHR pathway represses TGFβ-SMAD3 signalling and has a potent tumour suppressive role in SHH medulloblastoma. Sci Rep 10, 148 (2020). https://doi.org/10.1038/s41598-019-56876-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-56876-z
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.