Abstract
Agrobacterium-mediated plant galls are often misdiagnosed as nematode-mediated knots, even by experts, because the gall symptoms in both conditions are very similar. In the present study, we developed biosensor strains based on agrobacterial opine metabolism that easily and simply diagnoses Agrobacterium-induced root galls. Our biosensor consists of Agrobacterium mannitol (ABM) agar medium, X-gal, and a biosensor. The working principle of the biosensor is that exogenous nopaline produced by plant root galls binds to NocR, resulting in NocR/nopaline complexes that bind to the promoter of the nopaline oxidase gene (nox) operon and activate the transcription of noxB-lacZY, resulting in readily visualized blue pigmentation on ABM agar medium supplemented with X-gal (ABMX-gal). Similarly, exogenous octopine binds to OccR, resulting in OoxR/octopine complexes that bind to the promoter of the octopine oxidase gene (oox) operon and activate the transcription of ooxB-lacZY, resulting in blue pigmentation in the presence of X-gal. Our biosensor is successfully senses opines produced by Agrobacterium-infected plant galls, and can be applied to easily distinguish Agrobacterium crown gall disease from nematode disease.
Similar content being viewed by others
Introduction
Plant root galls are abnormal root tissue outgrowths that are caused by various parasites, including viruses, fungi, microscopic soil nematodes, and bacteria. Economic losses due to damage by nematodes and bacteria are significant; however, their galls are difficult to distinguish. Root-knot nematodes (Meloidogyne spp.) cause approximately 5% of global crop losses in over 2,000 susceptible plants1. Crown galls caused by Agrobacterium tumefaciens occur worldwide, causing great damage annually. This rod-shaped Gram-negative soil bacterium belongs to the class α-proteobacterium, and causes galls on the roots of susceptible plants2. The A. tumefaciens host range is often very broad; the bacterium causes crown galls that compromise the commercialization of plants in more than 60 families, including dicotylodonous plants, ornamental plants, brambles, and stone fruit and pome trees3,4,5,6.
The accurate and rapid diagnosis of crown gall is difficult but important for control and quarantine. There have been various attempts to diagnose and detect A. tumefaciens from plant tumors, including isolation using selective media, pathogenicity tests, biochemical and physiological tests, serological assays using antibodies, and polymerase chain reaction (PCR) techniques. In previous studies, the use of selective media for different biovars has allowed successful A. tumefaciens isolation7,8,9. However, this method of A. tumefaciens isolation is difficult and time-consuming. Several studies have applied serological techniques for A. tumefaciens detection, but this method has not been useful for the detection of pathogenic strains10,11. PCR techniques are the most frequently used methods for A. tumefaciens detection and diagnosis. Methods to diagnose crown galls with PCR have been developed in various ways12,13,14. Rhizobium and Agrobacterium are very similar in many respects, and it is difficult to distinguish these genera using PCR-based assays. Some studies have found no difference between Rhizobium and Agrobacterium in phylogenetic studies using 16S rRNA gene sequence. One method used to differentiate Agrobacterium from Rhizobium is to determine whether the bacteria induce pathogenic symptoms or root nodules; these symptoms are plasmid dependent. Thus, much effort has been made to avoid confusion between Rhizobium and Agrobacterium by designing the primer pairs used in PCR based on the gene located on the Ti (tumor-inducing) plasmid of A. tumefaciens. Another fundamental problem of the PCR method is that bacterial pathogens generally multiply into large populations in diseased lesions, but crown gall bacterial pathogens persist at significantly lower populations away from lesions and PCR-mediated diagnosis in plant samples requires a threshold population density. A major limitation of routine PCR application to diagnose plant disease is that successful PCR can be prevented by the frequent occurrence of polyphenolic inhibitors and thermostable DNA polymerase inhibitors in plant tissues12,15,16,17. To overcome these problems, a new approach to crown gall diagnosis is needed. Specifically, there is an urgent need for the development of new, simple, and rapid molecular-based diagnostic techniques. Therefore, in this study, we developed an easy and simple Agrobacterium biosensor based on opine catabolism.
To diagnose Agrobacterium crown galls, it is important to understand their complex opine biology. A. tumefaciens pathogenicity is initiated by transferring a segment comprising roughly 20% of the Ti plasmid, called the T-DNA (~40 kb) into plant cells during infection3,18. Genes in the transferred DNA are expressed in the plant nucleus, and are responsible for inducing tumorous growth of the transformed cells and for synthesizing opines, which serve as nutrient for A. tumefaciens that colonize the infected tissue19. Two very common opines are octopine and nopaline, which are produced in plant cells transformed with A. tumefaciens which harbor octopine- and nopaline-type Ti plasmids, respectively20. Opine biosynthetic genes on the T-DNA are distinct from their catabolic genes. Opine produced by transformed plant cells stimulate the expression of catabolic genes that are carried in the non-transferred portion of the A. tumefaciens Ti plasmid21. Nopaline tumors are caused by T-DNA transfer from nopaline-utilizing strains, and octopine tumors are caused by T-DNA transfer from strains that metabolize octopine22. In contrast, one group of strains can utilize both nopaline and octopine, although their tumors synthesize only nopaline, and another group utilizes nopaline, but their tumors produce either nopaline or octopine23. Additionally, some strains can utilize both types of opines, but their tumors produce neither nopaline nor octopine21,22,23,24. The nox or oox regions of the pTiC58 (nopaline-type) or pTi15955 (octopine-type) Ti plasmids are responsible for the catabolic utilization of nopaline or octopine in the A. tumefaciens strains C58 or 15955, respectively20,24. Catabolic functions are activated in the presence of exogenous nopaline or octopine, and regulatory controls are mediated by the LysR-type transcriptional regulatory proteins NocR or OccR; the genes encoding these proteins are located in the opine transporter regions (noc and occ) of the C58 and 15955 strains, respectively24,25.
The aims of the present study were to (1) develop a method to diagnose crown gall using opine detection and (2) evaluate the sensitivity of this method in practical application. Conventional density-based diagnostic methods for other bacterial plant diseases are not suitable for crown gall diagnosis. Therefore, a new diagnostic method was developed using opine metabolic genes on the Ti plasmid of Agrobacterium. Our diagnosis method is based on sensing the presence of external opines, and allows visualization of the expression of opine catabolism genes. The engineered opine-responsive, regulator based bacterial biosensors will be helpful for researchers and plant growers to detect plant diseases caused by A. tumefaciens, and will reduce detection time and cost. These biosensors can also be applied to evaluate and quantify opines produced by plant gall tissues.
Results and Discussion
Two plant samples showing root gall symptoms were delivered to our laboratory by farmers in 2018, one from squash and the other from raspberry. The farmers questioned whether their crops were infested with nematodes or bacteria. Nematode-mediated knots are often mistaken for Agrobactgerium-mediated galls, because the symptoms of both infections are difficult to distinguish, even for experts. General identification and detection of plant pathogens from plant materials typically requires 1–2 weeks, followed by an additional 3–4 weeks of pathogenicity tests. An easy and fast detection method is needed for local extension services. Although current PCR techniques have opened a new era of pathogen diagnosis, their application and interpretation can be complex. PCR techniques require PCR machinery, polymerases, buffers, and oligonucleotide primers; visualization of the results can also require UV lamps, ethidium bromide, and electrophoresis. Thus, we developed a simple molecular Agrobacterium biosensor detection method, based on two engineered, opine-responsive A. tumefaciens derivatives. As shown in Fig. 1, exogenous nopaline binds to NocR, a LysR-type transcriptional activator. The resulting NocR/nopaline complex activates the transcription of noxB-lacZY, resulting in β-galactosidase expression (Fig. 1a). Similarly, exogenous octopine binds to OccR, a LysR-type transcriptional activator, and the resulting OccR/octopine complex activates ooxB-lacZY transcription, resulting in β-galactosidase expression (Fig. 1b)23,25.
Working model of the opine-based biosensor strains. (a) Working model of the nopaline-based biosensor strain. Exogenous nopaline binds to NocR, a LysR-type transcriptional activator; the NocR/nopaline complex activates noxB-lacZY transcription, resulting in β-galactosidase expression. (b) Working model of the octopine-based biosensor strain. Exogenous octopine binds to OccR, a LysR-type transcriptional activator; the OccR/octopine complex activates ooxB-lacZY transcription, resulting in β-galactosidase expression.
Construction of opine biosensor strains
Nopaline and octopine catabolism operons carry genes responsible for transport and catabolism. To allow the entry of external opines, genes responsible for opine transport must remain intact and be active. However, disruption of the first cytoplasmic step of opine catabolism does not prevent transport of the opine into the cell, and opine-responsive gene regulation would be maintained. Thus, opine catabolism genes encoding opine oxidase were targeted for lacZY reporter fusions, and noxB of A. tumefaciens C58 and ooxB of A. tumefaciens 15955 simultaneously disrupted and fused to the lacZY reporter via Campbell integration as described previously26. The internal fragment of the target gene was inserted into pVIK112. The plasmids were then transferred from S17-1/λpir into the A. tumefaciens strain C58 or 15955 by conjugation, selecting for kanamycin-resistant target–lacZY transcriptional fusion. The expression levels of noxB-lacZY and ooxB-lacZY were visualized using X-gal or ONPG when nopaline and octopine were provided exogenously.
Responses of opine biosensor strains to synthetic opines
Synthetic opines were used to determine whether the opine biosensor strains were functional as predicted. A blue ring was observed around a paper disc containing nopaline and the C58 noxB-lacZY biosensor strain embedded in ABMX-gal, indicating the presence of nopaline and β-galactosidase expression (Fig. 2a). Similarly, a blue ring was observed around a paper disc containing octopine and the 15955 ooxB-lacZY biosensor strain embedded in ABMX-gal, indicating the presence of octopine and β-galactosidase expression (Fig. 2b). Neither biosensor responded detectably to the non-cognate opine (octopine for noxB-lacZY and nopaline for ooxB-lacZY).
Responses of the opine biosensor strains to synthetic opines, potato and tomato tumor tissues induced by the A. tumefaciens strain C58 or 15955. (a) Responses of the nopaline-based biosensor (C58 noxB-lacZY) to synthetic opines, potato and tomato tumor tissues induced by the A. tumefaciens strain C58. (b) Responses of the octopine-based biosensor (15955 ooxB-lacZY) to synthetic opines, potato and tomato tumor tissues induced by the A. tumefaciens strain 15955. Blue color around paper discs containing opines (10 nM) and a biosensor strain embedded in ABMX-gal plates indicates the presence of opines and β-galactosidase expression. w, Water control; n, nopaline; and o, octopine. Blue color around plant tumor tissues (right side on each plate) on a biosensor embedded in ABMX-gal indicates the presence of opines and β-galactosidase expression. Healthy plant tissues exhibited no color (left side on each plate). The biosensor functioned well as an opine sensor.
Responses of opine biosensor strains to plant tumor tissues
To determine whether the opine-based biosensors responded to plant opines, tumor tissue was induced in potato and tomato and used for plant assays. Plant tumor tissue samples induced with A. tumefaciens strains C58 or 15955 and healthy plant tissues (negative control) were placed on biosensor-imbedded ABMX-gal or on ABMX-gal agar plates close to streaks of the biosensor strain. The nopaline-based biosensor (C58 noxB-lacZY) exhibited a blue color in response to plant tumor tissues induced by A. tumefaciens C58 (Fig. 2a). The octopine-based biosensor (15955 ooxB-lacZY) exhibited a blue color in response to plant tumor tissues induced by A. tumefaciens 15955 (Fig. 2b). Again, each biosensor was specific; the nonB-lacZY biosensor only responded to tissue infected by the nopaline strain C58, not tissue infected with 15955, and the ooxB-lacZY only responded to the tissue infected with 15955. The uninfected plant tissue controls also resulted in no color changes.
Thin layer chromatography assay
To determine the type of opines present with Agrobacterium strains, plant opines were historically subjected to paper electrophoresis; the devices used in this type of electrophoresis are not currently used or readily available. We therefore attempted to perform opine analysis using TLC plates, which were readily obtained. Crude extracts from tomato tumor tissue samples or synthetic nopaline and octopine were loaded onto a reverse-phase TLC system. Once developed these plates were dried and overlaid with a suspension containing both the nopaline and the octopine biosensor strains in ABMX-gal agar. After incubation, inverted treardrop-shaped regions of light blue pigmentation were observed for all of the opine-containing samples. The retardation factor (Rf) of nopaline (0.35) and octopine (0.4) were comparable. We also confirmed nopaline and octopine could be detected in extracts from infected plants, and appeared identical to the synthetic opines by TLC (Fig. 3). This study is the first to perform TLC-based opine analysis, which will be helpful to researchers performing similar studies.
Thin-layer chromatography (TLC) analysis of opines using the biosensor strains. Reverse-phase TLC plates (12 × 10 cm) were developed using chloroform:acetic acid:water (2.8:3.5:0.7, v:v:v) and overlaid with ABMX-gal containing the nopaline- and octopine-based biosensor strains (C58 noxB-lacZY and 15955 ooxB-lacZY, respectively). n, Synthetic nopaline; 1, extract of tomato tumor induced by C58; 2, extract of tomato tumor induced by 15955; o, synthetic octopine; and c–, extract of healthy tomato tissue (negative control). Rf, retardation factor.
Quantification of opines based on β-galactosidase activity assay
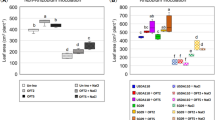
We then evaluated the sensitivity of opine-based biosensors by measuring β-galactosidase enzyme activity using ONPG as a substrate. The linear dose response was calculated using synthetic nopaline and octopine. The results of the nopaline-based biosensor (C58 noxB-lacZY) exhibited a good linear relationship in the ranges of 0–100 and 100–500 nM, with a greater slope for the lower concentration range (0–100 nM) than for the higher concentration range (100–500 nM) (Fig. 4a). The octopine-based biosensor (15955 ooxB-lacZY) showed even better sensitivity for octopine detection (Fig. 4b) than the nopaline biosensor for nopaline, with a linear relationship in the ranges of 0–50 and 50–200 nM, with slopes following the same pattern exhibited by C58 noxB-lacZY. The octopine-based biosensor showed excellent sensitivity to octopine detection, and higher sensitivity than the nopaline-based biosensor.
Plots of the influence coefficient of β-galactosidase activity versus the opine concentration of the proposed biosensors. β-galactosidase activity responses to (a) nopaline and (b) octopine. The results of the nopaline-based biosensor (C58 noxB-lacZY) exhibited a good linear relationship in the ranges of 0–100 (blue) and 100–500 nM (red) with different slops. The octopine-based biosensor (15955 ooxB-lacZY) showed even better sensitivity for octopine detection than the nopaline biosensor for nopaline, with a linear relationship in the ranges of 0–50 nM (blue). Error bars represent the standard deviations of three independent measurements. The distinct yellow color of o-nitrophenol from the ONPG substrate in microplates indicates the presence of opines.
Application of biosensor strains for gall diagnosis
The gall symptoms of the raspberry and squash samples that prompted this study were very similar (Figs. 5a and 6a). Both samples were diagnosed using the biosensor strains developed in this study. The raspberry gall samples exhibited a blue color in the nopaline-based biosensor, but no reaction in the octopine-based biosensor (Fig. 5b). The raspberry gall was ultimately confirmed to be a crown gall and diagnosed as a nopaline-type A. tumefaciens infection. Although the diagnosis was completed using the biosensor developed in this study, we isolated the causative bacterium from the root gall tissues to perform a PCR test using the bacterial DNA and the RBF (5′-TGACAGGATATATTGGCGGGTAA-3′) and RBR (5′-TGCTCCGTCGTCAGGCTTTCCGA-3′) primer set designed based on the nopaline type T-DNA right border14. The A. tumefaciens strains from the raspberry crown gall, which included C58 and R18 (four colonies, 1–4), exhibited the anticipated bands. The octopine type strain 15955 resulted in no bands (Fig. 5c).
Responses of the biosensor strains to raspberry galls. (a) Photograph of natural raspberry root gall symptoms. (b) Plate-based biosensor assays. Blue color around raspberry gall tissues (right side of each plate) indicates the presence of nopaline and β-galactosidase expression. Healthy raspberry tissues resulted in no color change (left side of each plate) in the octopine-based biosensor. (c) Nopaline type T-DNA fragment amplification of isolate R18 (1–4 colonies) from raspberry gall using RBF and RBR primers. M, 100-bp ladder marker; C58, A. tumefaciens strain; 15955, A. tumefaciens strain. The raspberry root gall was diagnosed as being caused by nopaline-type A. tumefaciens.
Responses of the biosensors to squash root galls. (a) Photograph of natural squash root gall symptoms. (b) No response around squash root gall tissues (left side of each plate) on both nopaline- and octopine-based biosensors indicates the absence of opines. Positive control potato disc tumors (C+) artificially induced by Agrobacterium strains C58 (upper) or 15955 (lower) exhibited blue color. (c) Micrographs of root-knot nematode isolated from the squash samples. The arrow indicates the stylet. The squash root gall was suspected to have been caused by root knot nematodes.
The squash gall samples were examined in a similar manner; they resulted in no color changes in either the nopaline- or octopine-based biosensor (Fig. 6b). Only positive controls caused blue color to form. Microscopic examination of the squash root gall tissues revealed root knot nematodes possessing a stylet (Fig. 6c).
A previous study compared the expression of the pathogenic A. tumefaciens gene pinF using enzyme activity assays at different concentrations of nopaline and octopine21. The opine concentrations and enzyme activity levels (Miller unit) were different in the present study from those reported previously21, likely due to differences in the expression levels of each gene. Millimolar-level amounts (mM) of opines are required for pinF expression levels in the hundreds of units; however, nanomolar-level amounts (nM) of opines are detectable for the biosensors developed in this study. Our results indicate that noxB and ooxB gene expression levels were higher than that of pinF. These biosensors may also have enhanced sensitivity because they are unable to degrade the opines, due to disruption in the oxidase genes.
We also quantified the plant opines found in tumor samples using standard curves prepared using the slopes in the lower concentration ranges (0–100 and 0–50 nM for nopaline and octopine, respectively). The response of the biosensors suggested that the amount of nopaline and octopine were 10–20 and 70–74 mg/g dry weight of potato and tomato tumor tissue, respectively. The biosensors were sufficiently sensitive to detect very small amounts of nopaline and octopine indicating that these opine-based biosensors will be useful for diagnosing small opine amounts in plant gall samples.
The grapevine crown gall caused by A. vitis is still a major problem in nurseries, and pathogenic strains produce opine vitopine27. In addition, the infestation of grapevine with root-knot nematodes is spreading all over the world28. Differentiation between these two seems more relevant. Since vitopine catabolism is not induced by nopaline or octopine, our biosensors may not applicable to the diagnosis of grapevine crown gall, but if the same strategy is applied to the vitopine catabolism genes, vitopine biosensors could be developed as well.
The worldwide economic damage caused by nematodes is significant, and quarantines between countries are strengthened to prevent nematode infection. However, no diagnostic kits or equipment are currently available for the diagnosis of nematode disease. Our opine-based biosensor can be applied to easily distinguish Agrobacterium crown gall disease from nematode disease.
Methods
Bacterial strains, plasmids, growth conditions, and chemicals
The bacterial strains and plasmids used in this study are listed in Table 1. E. coli strains were cultured at 37 °C in Luria-Bertani (LB) medium with or without 1.5% agar. A. tumefaciens strains were grown at 28 °C in Agrobacterium minimal medium with 0.25% mannitol (ABM) as a carbon source. Chemicals and antibiotics were purchased from Sigma-Aldrich and Fisher Scientific (St. Louis, MO, USA). When required, appropriate antibiotics were added to media: for E. coli, ampicillin (50 μg/mL) and kanamycin (50 μg/mL), and for A. tumefaciens, kanamycin (100 μg/mL). 5-bromo-4-chloro-indolyl-β-d-galactopyranoside (X-gal) was added to 40 μg/mL. Octopine was purchased from Sigma-Aldrich and Fisher Scientific and nopaline was synthesized (Basnet A, Department of Chemistry, Indiana University Bloomington) according to previously reported protocols29,30,31,32,33.
General DNA manipulation
DNA manipulation, cloning, restriction enzyme digestion and agarose gel electrophoresis were performed using standard techniques. For most techniques, the procedures described by Sambrook and Russell34 were used. DNA sequencing was performed using an ABI3730 sequencer. Restriction enzymes and DNA-depleting enzymes (New England Biolabs, MA, USA and TaKaRa, Otsu, Japan) were used according to the manufacturers’ recommendations. DNA fragments were purified from agarose gel using a QIAEX II gel extraction kit (Qiagen, Hilden, Germany) according to the manufacturer’s instructions. DNA amplification by PCR was performed using a T100 thermal cycler (Bio-Rad, Hercules, CA, USA).
Construction of lacZY transcriptional integration
The plasmid pBY34, carrying the internal fragment of the noxB gene was constructed using C58 genomic DNA as the PCR template and NoxBE (5′-GAATTCGCAATTGGATACGGGTTA-3′) and NoxBK (5′-GGTACCGCGATAGTCAGGATGAAT-3′) as primers. The amplified region (318 bp) was purified from agarose gel and ligated into the pGEM-T Easy Vector System (Promega, Mannheim, Germany) to generate pBY33, and the correct sequence was confirmed by sequencing. EcoRI-KpnI fragments of pBY33 digested with appropriate restriction enzymes (TaKaRa) were purified after electrophoresis from an agarose gel and inserted into the suicide vector pVIK11226, generating pBY34. The resulting construct pBY34 was transferred into E. coli S17-1 λpir, introduced into A. tumefaciens C58 by conjugation, generating noxB-lacZY (Fig. 1a). The lacZY reporter gene fusion insertion mutants were selected based on the kanamycin-resistance phenotype and confirmed using PCR analysis with primers that annealed upstream of the truncated fragments of noxB, NoxPro (5′-TTCGAGACAGCCATTGTT-3′) and LacFuse (5′-GGGGATGTGCTGCAAGGCG-3′).
The plasmid pBY36, carrying the internal fragment of the ooxB gene was constructed using 15955 genomic DNA as the PCR template and OoxBE (5′-GAATTCCAGCAAGACGGAGCATTT-3′) and OoxBK (5′-GGTACCGCGTGACAGGATAGAAAA-3′) as primers. The amplified region (345 bp) was purified from agarose gel and ligated into the pGEM-T Easy Vector System (Promega) to generate pBY35, and the correct sequence was confirmed by sequencing. EcoRI-KpnI fragments of pBY35 digested with appropriate restriction enzymes (TaKaRa) were purified after electrophoresis from an agarose gel and inserted into the suicide vector pVIK112, generating pBY36. The resulting construct pBY36 was transferred into E. coli S17-1 λpir, introduced into A. tumefaciens 15955 by conjugation, generating ooxB-lacZY (Fig. 1b). Reporter insertion mutants were selected based on the kanamycin-resistance phenotype and confirmed using PCR analysis with primers that annealed upstream of the truncated fragments of ooxB, OoxBPro (5′-ATGGCAAACACCCTGCTG-3′) and LacFuse.
Plant tumor induction
Potato tumors were artificially induced by inoculation with A. tumefaciens C58 and 15955 as previously reported35. Briefly, five 1-cm-diameter potato tissue discs were placed on water agar, and each was inoculated with 100 μL of suspension of the Agrobacterium strains C58 and 15955. The inoculated discs were incubated at room temperature for 3–4 weeks. To induce tumors in healthy tomato plants, a V-shaped wound (10–60 mm in length) was made in the middle of a young internode using a razor blade. The wound was inoculated with concentrated bacterial suspensions of A. tumefaciens strains C58 and 15955. The wounds were wrapped with wet gauze, sealed with parafilm, and wet treated for 1 week. The inoculated plants were incubated at room temperature for 5–6 weeks.
Crude opine extraction from plant tumors
Crude opines were extracted from tumors of potato, tomato and raspberry as previously described36 but with modifications. The tumor samples were dried at room temperature for 2 days. The dried tumor samples were ground and dissolved in sterile distilled water (0.01 g tumor/mL). After 1 h, the supernatant was centrifuged and filtered using a 0.45 μm filter (Sartorius, Gottingen, Germany).
Bacterial culture assay
The responses of opine biosensor strains to synthetic nopaline and octopine and plant gall tissues were analyzed using the imbedding and streaking method. In the first method, tumor tissues or paper discs containing chemical opines were placed onto a biosensor strain imbedded in an ABM agar plate containing X-gal (ABMX-gal). In the second method, a biosensor strain was streaked close to the tumor or chemical samples on an ABMX-gal.
Thin layer chromatography (TLC) assay
Prior to analytical TLC, samples (1–5 μL) were applied to reverse-phase TLC plates (Merck, Darmstadt, Germany), and chromatograms were developed using chloroform/acetic acid/water (2.8:3.5:0.7, v:v:v). After development, the solvent was evaporated, and the dried plates were overlaid with ABMX-gal agar media containing both biosensor strains (1/100 volume of medium), which were overnight cultured in ABM broth separately, plus 100 μg/mL kanamycin and incubated at 28 °C.
β-Galactosidase activity assay
A liquid-culture based β-galactosidase activity assay using ortho-nitrophenyl-β-galactoside (ONPG) as a colorimetric substrate was performed as previously described37. Mid-log phase biosensor strain cultures were diluted 1:100 with ABM broth to an OD600 of 0.01, mixed with different concentrations of nopaline or octopine (1–500 nM) in 2-mL of ABM broth, and incubated at 28 °C to an OD600 of ~0.6. Mid-log phase cultures were used for subsequent assays.
References
Moens, M., Roland, N. P. & James, L. S. Meloidogyne Species: a diverse group of novel and important plant parasites in Root-knot Nematodes (eds Roland, N. P., Maurice, M. & James, L. S.) 1–17 (CABI Publishing, 2009).
Smith, E. F. & Townsend, C. O. A plant-tumor of bacterial origin. Science 25, 671–673 (1907).
De Cleene, M. & De Ley, J. The host range of crown gall. Bot Rev 42, 389–466 (1976).
Alconero, R. Crown gall of peaches from Maryland, South Carolina, and Tennessee and problems with biological control. Plant Dis 64, 835–838 (1980).
Kennedy, B. W. & Alcorn, S. M. Estimates of U.S. crop losses to prokaryote plant pathogens. Plant Dis 64, 674–676 (1980).
Bazzi, C. & Rosciglione, B. Agrobacterium tumefaciens biotype 3, causal agent of crown gall of Chrysanthemum in Italy. Phytopathology 103, 280–284 (1982).
Schroth, M. N., Thompson, J. P. & Hidebrand, D. C. Isolation of the Agrobacterium tumefaciens - A. radiobacter group from soil. Phytopathology 55, 645–647 (1965).
New, P. B. & Kerr, A. A selective medium for Agrobacterium radiobacter Biotype 2. J Appl Bacteriol 34, 233–236 (1971).
Roy, M. A. & Sasser, M. A medium selective for Agrobacterium tumefaciens biotype 3. (Abstr.). Phytopathology 73, 810 (1983).
Alarcón, B., López, M. M., Cambra, M. & Ortiz, J. Comparative study of Agrobacterium biotypes 1, 2 and 3 by electrophoresis and serological methods. J App Microbiol 62, 295–308 (1987).
Goodman, R. N., Burton, D. & Tarbah, F. The occurrence of Agrobacterium tumefacines in grapevine-propagative material and simplified indexing system. Am J Enol Vitic 38, 189–193 (1987).
Cubero, J., Manjavacas, M. C., Llop, P. & López, M. M. A simple and efficient PCR method for the detection of Agrobacterium tumefaciens in plant tumours. J Appl Microbiol 86, 591–602 (1999).
Cubero, J., Van Der Wolf, J., van Beckhoven, J. R. C. M. & López, M. M. An internal control for the diagnosis of crown gall by PCR. J Microbiol Methods 51, 387–392 (2002).
Tan, B. S., Yabuki, J., Matsumoto, S., Kageyama, K. & Fukui, H. PCR primers for identification of opine types of Agrobacterium tumefaciens in Japan. J Gen Plant Pathol 69, 258–266 (2003).
John, M. E. An efficient method for isolation of RNA and DNA from plants containing polyphenolics. Nucleic Acids Res 20, 2381 (1992).
Pitch, U. & Schubert, I. Midiprep method for isolation of DNA from plants with a high content of polyphenolics. Nucleic Acids Res 21, 3328 (1993).
De Boër, S. H., Ward, L. J., Li, X. & Chittaranjan, S. Attenuation of PCR inhibition in the presence of plant compounds by addition of BLOTTO. Nucleic Acids Res 23, 2567–2568 (1995).
Escobar, M. A. & Dandekar, A. M. Agrobacterium tumefaciens as an agent of disease. Trends Plant Sci 8, 380–386 (2003).
Weiler, E. W. & Schroder, J. Hormone genes and crown gall disease. Trends Biochem Sci 12, 271–275 (1987).
Zanker, H. et al. Octopine and nopaline oxidases from Ti plasmids of Agrobacterium tumefaciens: molecular analysis, relationship, and functional characterization. J Bacteriol 176, 4511–4517 (1994).
Veluthambi, K., Krishnan, M., Gould, J. H., Smith, R. H. & Gelvin, S. B. Opines stimulate induction of the vir genes of the Agrobacterium tumefaciens Ti plasmid. J Bacteriol 171, 3696–3703 (1989).
Goldman, A., Tempe, J. & Morel, G. Quelques particularities de diverses souches d’Agrobacterium tumefaciens. C R Acad Sci 162, 630–631 (1968).
Montoya, A. L., Chilton, M. D., Gordon, M. P., Sciaky, D. & Nester, E. W. Octopine and nopaline metabolism in Agrobacterium tumefaciens and crown gall tumor cells: Role of plasmid genes. J Bacteriol 129, 101–107 (1977).
Kreusch, D., von Lintig, J. & Schróder, J. Ti plasmid-encoded octopine and nopaline catabolism in Agrobacterium: specificities of the LysR-type regulators OccR and NocR, and protein-induced DNA bending. Mol Gen Genet 249, 102–110 (1995).
Habeeb, L. F., Wang, L. & Winans, S. C. Transcription of the octopine catabolism operon of the Agrobacterium tumorinducing plasmid pTiA6 is activated by a LysR-type regulatory protein. Mol Plant-Microbe Interact 4, 379–385 (1991).
Kalogeraki, V. S. & Winans, S. C. Suicide plasmids containing promoterless reporter genes can simultaneously disrupt and create fusions to target genes of diverse bacteria. Gene 188, 69–75 (1997).
Chilton, W. S., Petit, A., Chilton, M. D. & Dessaux, Y. Structure and characterization of the crown gall opines heliopine, vitopine and ridéopine. Phytochemistry 58, 137–142 (2001).
Howland, A. D. et al. Impact of grapevine (Vitis vinifera) varieties on reproduction of the northern root-knot nematode (Meloidogyne hapla). J Nematol 47, 141–147 (2015).
Jensen, R. E., Zdybak, W. T., Yasuda, K. & Chilton, W. S. A useful synthesis of nopaline, a crown gall tumor metabolite. Biochem. Biophys. Res Commun 75, 1066–1070 (1977).
Chilton, W. S. & Chilton, M. D. Mannityl opine analogs allow isolation of catabolic pathway regulatory mutants. J Bacteriol 158, 650–658 (1984).
Chilton, W. S., Rinehart, K. L. & Chilton, M. D. Structure and stereochemistry of succinamopine. Biochemistry 23, 3290–3297 (1984).
Chilton, W. S., Hood, E. & Chilton, M. D. Absolute stereochemistry of Leucinopine, a crown gall opine. Phytochemistry 24, 221–224 (1985).
Isogai, A. et al. Structure of a new opine, mikimopin, in hairy root induced by Agrobacterium rhizogenes. Agric Biol Chem 52, 3235–3237 (1988).
Sambrook, J. & Russell, D. W. Molecular Cloning: A Laboratory Manual (Cold Spring Harbor Laboratory Press, 2001).
Kim, J., Heindl, J. E. & Fuqua, C. Coordination of division and development influences complex multicellular behavior in Agrobacterium tumefaciens. PLoS One 8, e56682 (2013).
Ellis, J. G. & Murphy, P. J. Four new opines from crown gall tumours-their detection and properties. Mol Gen Genet 181, 36–43 (1981).
Miller, J. H. Experiments in molecular genetics (Cold Spring Harbor Laboratory Press, 1972).
Simon, R. U. P. A. P., Priefer, U. & Pühler, A. A broad host range mobilization system for in vivo genetic engineering: transposon mutagenesis in gram negative bacteria. Nat Biotechnol 1, 784–791 (1983).
Acknowledgements
This work was supported by the Gyeongsang National University Fund for Professors on Sabbatical Leave, 2017. Additional funds were provided by Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education (2015R1A6A1A03031413). We thank A. Basnet of the IU Department of Chemistry for synthesizing the purified nopaline used in this study.
Author information
Authors and Affiliations
Contributions
C.F. and J.K. conceived and designed research. O.C. and J.B. conducted experiments. B.K., Y.L. and S.K. analyzed data. O.C., J.B. and J.K. wrote the manuscript. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Choi, O., Bae, J., Kang, B. et al. Simple and economical biosensors for distinguishing Agrobacterium-mediated plant galls from nematode-mediated root knots. Sci Rep 9, 17961 (2019). https://doi.org/10.1038/s41598-019-54568-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-54568-2
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.