Abstract
There is evidence for an age-related decline in male reproductive functions, yet how the human testis may age is not understood. Human testicular peritubular cells (HTPCs) transport sperm, contribute to the spermatogonial stem cell (SSC) niche and immune surveillance, and can be isolated and studied in vitro. Consequences of replicative senescence of HTPCs were evaluated to gain partial insights into human testicular aging. To this end, early and advanced HTPC passages, in which replicative senescence was indicated by increased cell size, altered nuclear morphology, enhanced β-galactosidase activity, telomere attrition and reduced mitochondrial DNA (mtDNA), were compared. These alterations are typical for senescent cells, in general. To examine HTPC-specific changes, focused ion beam scanning electron microscopy (FIB/SEM) tomography was employed, which revealed a reduced mitochondrial network and an increased lysosome population. The results coincide with the data of a parallel proteomic analysis and indicate deranged proteostasis. The mRNA levels of typical contractility markers and growth factors, important for the SSC niche, were not significantly altered. A secretome analysis identified, however, elevated levels of macrophage migration inhibitory factor (MIF) and dipeptidyl peptidase 4 (DPP4), which may play a role in spermatogenesis. Testicular DPP4 may further represent a possible drug target.
Similar content being viewed by others
Introduction
Aging of any given organ is characterized by a decline of its function. This may apply also for the male gonad. It was postulated that with increasing age, an overall decline of male reproductive functions occurs, which involves reduced sperm counts and androgen production, paralleled by structural changes1,2. Several investigations indicate that structural and functional changes in the testis of men in advanced age involve the androgen-producing Leydig cells and the Sertoli cells, which are regarded as the nurse cells for male gem cells3,4,5,6,7. In addition, age-related alterations of the tubular wall compartment (also called tunica propria) in men were reported. These changes include progressive enlargement and sclerosis, accompanied by impaired spermatogenesis and (in some cases) complete tubular sclerosis2,5,6. In contrast, a study in men with proven fertility, who were examined later in life, suggested that aging of the human testis may not necessarily be associated with structural/functional changes, in general8. Hence, there is debate about the selection criteria of men in the mentioned studies, as patients with pre-existing infertility and accompanying structural changes may have been included2,5,6. In addition, all these investigations in elderly men have limitations with respect to statistically relevant amounts of samples. Conceivably, age-related alterations of testicular structure and function in men are likely a consequence of many factors acting together in the long-lived human species. Besides age itself, medical conditions, use of drugs and overall lifestyle can affect both, the regulation of testicular functions and/or the male gonads itself. These issues cannot be well separated in human and there is no established, long-lived animal model to adequately mirror this situation.
A possible approach to investigate testicular aging would be the examination of senescence of testicular cells, yet most human testicular cells cannot be propagated in vitro, with the exception of peritubular cells. Human testicular peritubular cells form several layers within the wall surrounding the germinal epithelium and morphological studies suggested that they may change during testicular aging2,5,6. Peritubular cells of the adult testis are contractile, smooth muscle-like cells and secrete extracellular matrix (ECM) proteins. Imbalances between secretion/maintenance of the ECM and cellular contractile abilities are documented in the testes of men with impaired spermatogenesis9,10. The resulting fibrosis of the tubular wall is furthermore accompanied by accumulation of immune cells, implying a sterile type of inflammation in the tubular wall of infertile men10,11,12. As mentioned, human testicular peritubular cells (HTPCs) can be isolated and cultured from human testicular tissue13. HTPCs produce important growth factor molecules, e.g. glial cell line derived factor (GDNF) and C-X-C motif chemokine ligand 12 (CXCL12) and thereby most likely contribute to the spermatogonial stem cell (SSC) niche of the testis and to life-long spermatogenesis14,15,16. Initially, Sertoli cells were known as sole producers of GDNF and an overall important part of the SSC niche. Yet the importance of complementary GDNF, derived from peritubular cells, was confirmed in a systemic animal model17. The GDNF-mutant mice lost their fertility with increasing age. This observation links aging to a function of peritubular cells and concurs with the concept that impairments of the SSC niche may be part of testicular aging18. In this context a recent study of the human testes by Pohl et al.19 documented the occurrence of diminished spermatogenic efficiency, increased amount of proliferating Adark spermatogonia and altered nuclear morphology of Sertoli cells with increasing age. This suggests alterations of Sertoli cell function. It does not rule out changes in peritubular cell functions, although obvious structural alterations of the tubular wall or peritubular cells with increasing age in men were not described.
HTPCs also secrete cytokines, express Toll-like receptors, purinergic receptors and produce reactive oxygen species (ROS)20. Previous work implies that peritubular cells are involved in sterile inflammation, associated with some cases of male infertility10,21,22. Inflammation may also be part of aging. Indeed the term “inflammaging”23 was coined. A possible functional involvement of peritubular cells in aging of the human testis was not examined yet. We reasoned that replicative senescence, triggered by serial passaging of cultured HTPCs, may allow the investigation of their contribution to testicular aging. Replicative senescence is an established cell culture model for research in aging and senescence24,25. It describes a cellular response, which limits proliferation of aged or damaged cells, in association with restricted cellular function26 and a characteristic secretory phenotype, i.e. senescence-associated secretory phenotype (SASP)27. The consequences of replicative senescence in early and advanced passages of cultured HTPCs were investigated employing complementary approaches, including proteomics, qPCR studies and light microscopy (LM) combined with focused ion beam scanning electron microscopy (FIB/SEM) tomography, which allowed 3D reconstructions of various cellular compartments.
Results
Replicative senescence in HTPCs
HTPCs divide in culture with a doubling time of 2–3 days and can be propagated until cell division ceases after approximately one year. Cells from the same donors without (low passage number) and with signs of replicative senescence (advanced passage number) were compared. In addition to growth arrest, the following criteria were used to define replicative senescence of HTPCs: a change of the spindle-like morphology to an irregularly shaped and flattened morphology (Fig. 1a), increased β-galactosidase activity (Fig. 1b), increased cell size (Fig. 1c) and reduced telomere length (Fig. 1d). Cells fulfulling these criteria were further examined. Furthermore, we found that mtDNA was reduced in advanced passges, compared to the corresponding early passages (Fig. 1e).
Changes associated with replicative senescence in HTPCs. Schematic presentation of HTPCs in transition from early to advanced passages (a). Light micrographs of senescence associated β-galactosidase staining of HTPCs: in contrast to HTPCs from early passage (passage = P; P5, left), β-galactosidase activity is prominent in advanced passage (P15, right) (b). Measurement of cell sizes revealed a significant increase in diameter in advanced passages from the same donor (n = 7), statistics were calculated with a paired two-sample t-test (c). Quantification of the relative telomere length from isolated gDNA from advanced passages of HTPCs from the same donor (n = 5) with significant telomere attrition, normalized to early passages (d). Quantification of mtDNA copy number by qPCR indicates significantly decreased amount of mtDNA in advanced HTPC passages (e). Statistical analysis was done with an one-sample t-test (d,e), mean ± SEM are given, asterisks indicate statistical significance, *p < 0.05, **p < 0.01, ***p < 0.001.
FIB/SEM tomography of early vs. advanced passages of HTPCs
Ultrathin embedding of HTPCs allowed identification of target cells in SEM due to the coordinate system engraved on the slides and the specific cell topography. Superposition of LM (phase contrast) and SEM images allowed rapid correlation of fine structural details in the nanometer range. HTPCs of early passages are elongate, spindle-shaped cells, with a prominent nucleus and several nucleoli (Figs 1a, 2a,b). In phase contrast, granular substructures dispersed within the cytoplasm were observed in all cells. At one side of the nucleus the cytoplasm appeared to be devoid of larger substructures over a distance of 10–20 µm (Fig. 2a). In contrast, cells of advanced passages exhibited an abounding number of granules, either bright or dark in phase contrast, evenly distributed within the cytoplasm (Fig. 2a’). For better insight into the distribution of cellular substructures in relation to the nucleus, longitudinal sections were chosen for FIB/SEM analysis of both passages (Fig. 2c,c’,d,d’).
Focused ion beam scanning electron microscopy (FIB/SEM) tomography of early vs. advanced passages of HTPCs. Correlative LM and FIB/SEM tomography of ultrastructural changes of HTPCs (early passages: a–g; advanced passages: a’-g’). In early passages the cells are spindle-shaped with a rather homogeneous region of cytoplasm proximal to the nucleus (a: A) and a distal, granular region (a: B). Senescent cells flatten out and their cytoplasm becomes packed with granules (a’). Selected cells from LM were re-localized with SEM (b and b’) and sectioned with FIB longitudinally (c,c’; d,d’). In early passages the homogeneous part of the cytoplasm is characterized by large, parallel sheets of rough endoplasmic reticulum (ER) and granules, which are classified as lysosomes (Ly) (c–e), moderate electron dense after osmium fixation (c,c’) and very electron dense after rOTO-staining (d,d’). In senescent cells lysosomes accumulate in enormous numbers (c’,d’,e’). Mitochondria are elongated and form branched larger networks, independent of the passage (f,f’; individual mitochondria are differently colored). However, in senescent cells the portion of mitochondria related to the cytoplasm is significantly reduced (f,f’). The ER is densely packed with ribosomes during all stages and is associated with both, mitochondria (g,g’) and lysosomes (g’). M = mitochondrion; N = nucleus
Cells in early passages were rather flat with a height in the range of 3 µm (Fig. 2c). The nucleus was lens shaped with parallel envelope membranes. Nuclear pores could be visualized by volume rendering of the 3D-data set. Several strands of ER were typically connected to the outer nuclear membrane. The ER formed: i) large, parallel sheets densely packed with ribosomes, ii) smaller sheets of fenestrated rough ER, randomly orientated and iii) bulb shaped dilatations with either electron translucent or fine granular lumen (Fig. 2c,d). Mitochondria were up to 15 µm long and very variable in their diameter, ranging from 0.3 µm to 0.1 µm, without any cristae. Only in the 3D context could they be recognized as mitochondria (Fig. 2c,d,f). Most mitochondria were branched and formed a complex network including numerous loops. Characteristic for all mitochondrial networks were multiple intimate contact sites with sheets of rough ER (Fig. 2g). The mitochondrial tubes were locally tightly enwrapped so that the membranes of ER and the outer mitochondrial membrane could not be discriminated, even at the resolution limit of FIB/SEM (2 nm isovoxel). At the site of contact, the ER was depleted of ribosomes. There was a population of roughly globular organelles composed of two components: a very electron dense matrix of packed/and or fused membranes and electron translucent areas resembling vacuoles, ranging from approximately 200 nm to 2 µm in diameter (Fig. 2c). When present in higher densities, they were interconnected. They were regarded as lysosomes, rather than autophagosomes or autophagolysosomes, as there was no indication for the presence of membranous phagophores enwrapping cell organelles28, e.g. mitochondria. When examining lysosomes in a 3D context, it became evident that the ER lumen was in continuity to both, the vacuolar, and the membrane part of the lysosomes. In early passages lysosomes were not distributed randomly within the cell but rather accumulated in one half of the cell, typically 10–15 µm distal from the nucleus. This concurs with the granular structures observed in the phase contrast images (Fig. 2a, a’).
HTPCs in advanced passages lost their original spindle-like shape and flatten out to irregular shapes (Fig. 2a’, b’, c’). Although the nucleus still appeared to be roughly lens shaped in LM, deduced from DAPI images (Fig. 2a’), the 3D analysis revealed numerous cytoplasmic invaginations (Fig. 2c’). Nuclear pores were still visible with volume rendering of the 3D data set. The mitochondrial networks were pertained (Fig. 2f’), however, their number decreased in relation to the corresponding cytoplasmic volume (normalized to 100 µm3) (Table 1). Deducing from statistical measurements, the mitochondria (network) became smaller in diameter (0.22 µm vs. 0.18 µm) but extended in length (5.8 µm vs. 8.8 µm) (Fig. 2f’). However, the total mitochondrial surface in a given volume of 100 µm3 was reduced to approximately 25% compared to cells of early passages. The characteristic contact sites between mitochondria and ER were preserved, however, the shape of ER changed from large parallel sheets to smaller slabs with random orientations (Fig. 2g’). There was indication for at least partial degradation of mitochondria or mitochondrial segments, which became visible upon large volume serial block face sectioning: the mitochondrial matrix became locally electron translucent, formed blebs, with electron dense membrane fragments, likely residual cristae (Fig. 3a,b). The 3D reconstruction revealed that several segments of the mitochondrial network can be interconnected by vacuolated blebs (Fig. 3d’; Supplementary Fig. 1). Endosomes and small vesicular structures were in close vicinity to both, ER and mitochondria (Fig. 3a,c,d). Within the sheets of rough ER, small lens-shaped vacuoles were frequently observed (Fig. 3a’) and several vacuoles of different sizes can be interconnected via rough ER (Fig. 3c’). The most striking structural alteration was an enormous accumulation of lysosomes (Fig. 2d,d’; e,e’, 3a’). Their volume portion increased almost tenfold (from 7% to 60%). 3D reconstructions revealed that most lysosomes, are interconnected, either directly by their electron dense and electron translucent components or by short strands of ER (Fig. 2e’, 3a’). The ER was in luminal connection to the lysosomes (Fig. 3a’; Supplementary Fig. 1). The intimate contact of the mitochondria with sheets of rough ER was pertained, as shown by 3D reconstructions (Fig. 3c,c’, d).
Selected FIB/SEM micrographs of a series of HTPCs from early (a–d) and advanced passages (a’–d’). The numbers (upper right) indicate the selected micrographs from the series. In the early passage the HTPC shows contact sites of ER with a branched mitochondrion (M) with several cross sections within one micrograph (4 nm isovoxel). Vacuoles (V) form within the mitochondrial matrix, leading to swellings (b; circle/transparent magenta) with some cristae still visible (a, arrows). Sheets of ER are attached to the mitochondrial network, noticeable after 3D reconstruction (c,d). The advanced passage exhibits numerous lens shaped vacuoles (a’; asterisks; voxel size: 7.5 × 7.5 × 14 nm). Strands of rough ER are connected at both ends to the vacuoles (a’, circles). Electron dense inclusions within the vacuole were identified as lysosomes (a’, arrow), when following the FIB/SEM series. Lysosomes (Ly) are connected to strands of ER at multiple sites (a’; squares). 3Dreconstruction of vacuoles (V) and lysosomes (Ly) show that they form clusters connected to each other (b’; arrow). The ER forms large sheets, best visible in top view (c’) compared to the FIB/SEM block face micrographs (a’). The ER membrane is in continuity with the vacuole membrane (c’; rectangle). The mitochondrion (M) shown is also locally fused with smaller and larger vacuoles (d’; circles).
Proteomic approach for comparison of early vs. advanced HTPC passages
A quantitative proteome analysis of early and advanced passages of cultured HTPCs (n = 6) and their conditioned media (CM) was performed to explore potential molecular alterations of replicative senescence of the HTPC proteome and secretome. A total of 3692 proteins were identified in HTPC-lysates at a false discovery rate (FDR) < 0.01. All identified proteins in early and advanced HTPC passages are listed in the Supplementary Table 1. Proteome analysis of CM identified 860 proteins, including 569 proteins with annotated extracellular localization, defining the secretome dataset. The identified proteins in CM of early and advanced HTPC passages are listed in the Supplementary Table 2. Hierarchical clustering and principal component analyses demonstrated passage-associated segregation of cellular proteomes and secretomes from early and advanced passages (Supplementary Fig. 2a,b). Label-free quantification (LFQ) revealed that 208 proteins were significantly different in quantity (t-test, FDR < 0.05) in cellular proteomes (Fig. 4c, Supplementary Table 3) and 131 proteins in the secretomes (Fig. 4d, Supplementary Table 3), between HTPCs from early and advanced passages, respectively. A large number of nuclear and mitochondrial proteins were decreased in contrast to increasing lysosomal proteins (Fig. 4a,b).
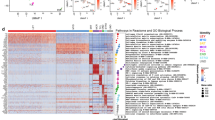
Proteomic analysis of advanced compared to early passages of HTPCs. Gene set enrichment analysis (GSEA) revealed significantly enriched gene sets (FDR q-value ≤ 0.05) and were summarized using REVIGO by clustering semantically similar GO terms. Each of the 20 characteristic gene sets enriched in early (a) and advanced passages (b) of HTPCs are shown. Color-coding refers to the corresponding highest GO hierarchy level. The x-axis shows the enrichment significance resulting from the GSEA and is depicted as –log10 (FDR q-value). The number of quantified proteins per gene set is shown in brackets. Volcano plots of intracellular and extracellular proteins, which are more abundant in passaged HTPC cellular proteomes (c) and secretomes (d) are depicted as red dots and proteins less abundant are shown as blue dots, respectively. Selected proteins with significant difference in abundance are labeled. P-values were calculated by a paired two-sample t-test. DPP4 expression in testicular peritubular cells (e). Light micrographs of immunohistochemical staining of human testicular sections. DPP4 is detected in several peritubular cells and cells of the interstitial space. Right micrograph: detail of the DPP4 staining (framed area). The negative control is without staining.
The secretome analysis of advanced passages showed, among others, a significant increase of the dipeptidyl peptidase 4 (DPP4) (Fig. 4d). As DPP4 expression has not been identified before in testicular peritubular cells, we explored its presence in situ. DPP4 staining was carried out with human testicular sections from middle-aged men (48–50 years). DPP4 was absent in many of the peritubular cells of the seminiferous tubular wall. However, in half of the samples examined (3 out of 6 examined) immuno-reactive peritubular cells were found in situ. DPP4 was also found in cells of the interstitial space (Fig. 4e). The gene set enrichment analysis (GSEA) showed substantial alterations between early and advanced passages in cell lysates and CM. The gene sets enriched in early passages were primarily related to mitochondrial and chromosomal functions, and more specifically related to RNA splicing, chromatin organization and DNA repair (Fig. 4a), while secretory, lysosomal, and metabolic processes were enriched in advanced passages (Fig. 4b). In the HTPC CM, gene sets related to extracellular matrix were enriched in early passages, while gene sets related to different binding and signaling processes were enriched in advanced passages. The detailed results of the GSEA are listed in Supplementary Tables 5, 6 and are visualized as functional networks in Supplementary Fig. 3.
qPCR study of characteristic HTPC marker genes
For complementation of the study, possible alterations between early and advanced passages were examined by qPCR. Transcription levels of AR, ACTA2, CNN1, StAR and PTGS1 remained stable (Fig. 5a). Inflammatory genes, namely IL6, IL8, PTGS2 and PTX3 were also not affected by passaging except for CCL2, which was increased about 2-fold (Fig. 5b). However, ELISA-measurements did not confirm such a change at the protein level (Supplementary Fig. 5). Expression levels of the growth factors CXCL12 and GDNF varied between the different donors, although without detectable tendency (Fig. 5c). ELISA-measurements of CXCL12 did not indicate a significant age-associated change, but rather inter-individual alterations (Supplementary Fig. 4).
qPCR study of typical genes expressed in peritubular cells. mRNA levels of characteristic HTPC marker genes like AR, ACTA2, CNN1, StAR and PTGS1 (a). Inflammation-associated genes show significantly increased mRNA level of CCL2. PTX3, IL6, IL8 and PTGS2 are not changed (b). mRNA expression of growth factors, CXCL12 and GDNF (c). Graphs represent individual measurements and means ± SEM. Statistical analysis was executed with one-sample t-test, Asterisks show statistical significance, *p < 0.05, **p < 0.01, ***p < 0.001.
Discussion
In man, aging of many organs, including the testis, cannot be readily assessed. Information about cellular senescence of human testicular cells is likewise missing. HTPCs are the only human testicular cell type, which can be cultured and propagated until replicative senescence is witnessed. The present study combines ultrastructural 3D data from HTPCs during senescence with a proteomic analysis of cellular and secreted proteins and a qPCR analysis and provides a detailed picture of senescence-associated changes in these human testicular cells.
In general, replicative senescence of cells is characterized by cell cycle arrest29,30, changes in the cellular phenotype and telomere shortening31. As there is no universal marker, the combination of different hallmarks is typically used to characterize senescent cells32. All HTPCs analyzed in this study showed a combination of impaired proliferative competence, reduced telomere length, increased cell size, and increased β-galactosidase activity, i.e. the general hallmarks of senescent cells29,31,33,34. The study of these cells allowed us to identify HTPC-specific changes, which may be of relevance for the functions of the testis.
Detailed insights into cellular compartments of HTPCs, based on 3D reconstruction, identified three significant ultrastructural changes, associated with cellular senescence in HTPCs: i) a significant decrease of rough ER ii) a dramatic increase of lysosomes, iii) a decrease in the number of mitochondria. Ultrastructural reconstructions revealed that the ER is central in all these changes. It is connected to all lysosomes (Fig. 3a’), is in intimate contact to all mitochondria (Figs. 2g,g’: 3c), and involved in vacuole formation (Fig. 3a’). The ER was packed with ribosomes in a surprisingly high density in early and advanced passages, hence an efficient protein synthesis is visibly maintained in senescent cells. Both in vivo and in vitro (human) testicular peritubular cells secrete a plethora of proteins, mainly ECM components15. The proteomic data supported the general capacity for protein synthesis during all passages, however secreted ECM proteins are significantly decreased, concurring with the structural reduction of the ER from 31% to 4% of the cytoplasmic volume (Table 1). These results are in line with impaired protein homeostasis (proteostasis) in senescent HTPCs, which is associated with aging in many cells35.
The striking increase of lysosomes, which make up 60% of the cell volume in advanced passages, further argues for impaired proteostasis as a central event. The 3D reconstructions showed that in HTPCs lysosomes were connected to the ER in early and advanced passages (Fig. 3a’; Supplementary Fig. 1). Only in early passages, cellular polarity was observed with respect to a region located at one side of the nucleus, which was almost free of lysosomes and occupied by accumulation of parallel-arranged large sheets of rough ER (Fig. 2c,d). This cellular polarity was lost gradually in advanced passages. The massive accumulation of lysosomes reduced the space available for rough ER and implies steric hindrance of formation of rough ER. Similar data were recently published for large volume FIB/SEM reconstructions of HeLa cells: the dictyosomes, endosomes, lipid bodies and lysosomes form an interconnected system for Golgi degradation and reconstitution36. The massive increase of lysosomes, both in number and volume, may have different reasons. Thus, together with the proteomic data (Fig. 4b) and published physiological in vivo data37 the results indicate impaired proteostasis.
Small vacuoles are initially visible within rough ER sheets as lens shaped structures (Fig. 3a’) and subsequently, larger, spherical structures, still in luminal contact with ER, were found. They were considered to be nascent lysosomes and it seems likely that the formation of the vacuolar part of the mature lysosomes is a consequence of direct involvement of ER membranes and ER lumen. Similar autophagolysosomes/autophagosomes, degrading mitochondria, are described in podocytes of rats after acute ischemia38 and in hexa KO cells, shown in serial 3D reconstruction, and also indicateinvolvement of ER39.
The contact sites of ER with mitochondria are being discussed for Ca2+ exchange40 but also as a supply site of membrane components from the ER to the outer mitochondrial membrane41. Changes of the mitochondrial network and the reduction in surface area of mitochondria by a factor of four was qualitatively paralleled with a reduction of the rough ER (Table 1). The investigation of lysosomes revealed that the majority is composed of an electron dense matrix, which is, at least in part, formed by an aggregation of membranes. However, when looking at the mitochondria with large volume reconstruction, there are characteristic features: empty spaces, lacking cristae, within the mitochondrial matrix, similar in appearance to data from Szento et al.38 and contacts of mitochondria with lysosomes (Fig. 2g’; Supplementary Fig. 1) even at several segments of individual mitochondria, which may explain the reduction of mitochondrial volume by degradation (Fig. 3a; Supplementary Fig. 1). Compared to typical autophagy of cell organelles, as reviewed by Feng et al.42, where entire mitochondria or other cell organelles are engulfed by membranes to form an autophagosome43, the observed mechanism in HTPCs seems to be different. The mitochondrial vacuoles may enable fusion with ER-derived vacuoles and/or existing lysosomes. Mitochondria and rough ER are in direct contact to each other, as well as to lysosomes, thus forming a common, interconnected system shifting with progressing senescence to lysosome formation (Fig. 6).
Schematic representation of the network of endoplasmic reticulum (ER), mitochondria (M), lysosomes (Ly), vacuoles (V) and their mutual relations based on electron microscopy data. All partners can be connected via ER. Sheets of rough ER enwrap mitochondria. Small lens shaped vacuoles form within the ER lumen. Mitochondria are elongated and range up to 15 µm. They have a minimal diameter of approx. 100 nm, without any cristae present. With progressing senescence, vacuoles form within the mitochondrial matrix, sometimes with degenerating cristae visible (asterisk). Both, mitochondria and rough ER are in direct contact to lysosomes forming a common, interconnected system.
Mass spectrometry revealed further striking changes of the cellular protein pattern, including the ubiquitous transcription factor NFκB1 that showed lower levels in senescent HTPCs. It was reported that a loss of NFκB1 may lead to early onset aging44. Furthermore, a reduced abundance of the RNA binding protein HNRNPA1 was found. It controls cellular senescence and the SASP via sirtuin1. Loss of HNRNPA1 induces a senescent phenotype in human diploid fibroblasts45 also due to its crucial role in telomere protection46. With increasing age, energy production is reduced as reviewed by Barzilai, et al.47. The GSEA results of the proteome suggest that the energy metabolism changes in senescent cells from an oxidative metabolism to anaerobic glycolysis, which correlates with decreased mtDNA, structural changes within the mitochondria as reviewed by Bratic et al.48 and increased LDHA abundance in advanced HTPC passages49. Senescent cells show changes in chromosome and chromatin architecture in general, as reviewed by Sun et al.50. GSEA showed that chromosome and chromatin organization are changed in advanced passages (Fig. 4a,b). This is paralleled by alterations of the shape of the nucleus, seen in many HTPCs (Fig. 3a’). In addition, the reduced DNA repair capacity in advanced passages, in combination with age-related increased levels of DNA damage, could reinforce cellular senescence51.
ROS production by HTPCs was described in previous studies20. The analysis of the secretome revealed that a number of factors, e.g. superoxide-dismutase 1 (SOD1), an antioxidative factor, were elevated in advanced passages. This may be a reaction to higher ROS levels, which can cause DNA damage, leading to cellular senescence in human fibroblast cell lines52. MIF, a pro-inflammatory cytokine, increased in senescent HTPCs, is expressed by immune cells and several other cell types, including smooth muscle cells53 (Fig. 4d). MIF is released into the extracellular space in response to various stimuli (e.g. mitogens and pro-inflammatory cytokines), and elevated levels indicate a pro-inflammatory milieu. MIF binds to cell surface receptors, either CXC chemokine receptors (CXCRs) or the CD74 receptor54. The G-protein–coupled chemokine receptors CXCR2, −4, and −7 are linked to chemotaxis of immune cells, in general. MIF could thereby contribute to an inflammatory environment within the human testis, also via suppression of the anti-inflammatory effect of glucocorticoids55. MIF actions may lead to enriched cytokine levels and can activate important pathways in senescence and aging, namely p53 and NFκB as reviewed by Salminen et al.56.
Within the testis, a special situation may exist, since MIF is a ligand for the CXCR4, expressed by SSCs, and is essential for postnatal maintenance of the SSCs in marmosets and mice57. If indeed applicable to the testis, higher levels of MIF may influence the SSC niche via CXCR4 binding. CXCR4-expressing cells in the testis may also be indirectly influenced by DPP4, which was recently described as a senescence marker for human fibroblasts58. DPP4 was significantly elevated in the secretome of advanced HTPC passages (Fig. 4d) and detected in vivo in human testicular samples in peritubular cells (Fig. 4e). DPP4 is able to cleave several neuropeptides but also chemokines, including CXCL12. This is of importance in hematopoiesis, angiogenesis, and stem cell homing. With respect to the testis, it was shown that at least in mice and marmosets CXCL12-CXCR4 signaling is required for postnatal maintenance of SSCs, SSC propagation and that it prevents SSC differentiation. As CXCL12 truncation by DPP4 can reduce CXCL12-CXCR4 signaling significantly59, DPP4 within the testis may have a negative impact on spermatogenesis and may antagonize other chemokine- and neuropeptide actions. The expression of DPP4 in the human testis, not shown before, raises the possibility that inhibitors for DPP4 may be potentially useful for therapy of diseases of the testis or age-related changes. In this respect, gliptins, i.e. established anti-diabetic drugs60 may be of interest, as they are inhibitors of DPP4 and were shown to affect stem cell niches61.
The levels of several ECM factors, including various laminins and collagens, are reduced in the secretome of advanced passages (Fig. 4d). They represent the major fractions of secreted proteins of HTPCs15, and reduced levels indicate an overall diminished secretory activity and/or impaired proteostasis of senescent HTPCs. Yet a reduced secretion is not in line with reported increased ECM-deposits in the tubular wall of elderly men6 or the changes seen in old rodent testes62. The human data is, however, being debated and questioned. It is possible that men with pre-existing infertility may have been included. As fibrosis of the peritubular wall is a hallmark of male infertility, such an inclusion may conceivably have biased the conclusion. If the results of the present study correlated with the in vivo situation, fibrosis of the peritubular wall is not be expected in testes of healthy elderly men without confounding issues8. Indeed, a recent light microscopic study by Pohl et al.19 did not detect differences in the tubular wall compartment from young and elderly men. Further, peritubular cells are smooth muscle-like cells, comparable with other smooth muscle cells of the body. Of note, senescent vascular smooth muscle cells, like senescent HTPCs, also showed reduced ECM protein secretion63.
Several distinct factors of importance for HTPCs (AR and GDNF), were not readily detectable by LC-MS/MS, but could be analyzed by qPCR (Fig. 5). AR was not significantly altered in advanced passages. AR-activation enhances the smooth muscle-phenotype and thereby may regulate contractility of HTPCs16. In full accordance with the proteomic data, the smooth muscle factors ACTA2 and CNN1 also remained stable, thus HTPCs keep their smooth muscle-like phenotype. GDNF contributes to spermatogenesis in mice14,17 and was also unchanged in advanced passages, like protein (LC-MS/MS) and mRNA levels of PTGS1. This enzyme is involved in the generation of prostaglandins and thereby in the regulation of GDNF64. CXCL12 also contributes to spermatogenesis and the mRNA and protein levels (Supplementary Fig. 4) were likewise not significantly altered. For all these transcript levels, considerable variations between individual human samples were, however, observed.
Several pro-inflammatory factors were also examined (IL6, IL8, CCL2, PTGS2 and PTX3), only CCL2 expression was slightly, but statistically significantly (about 2-fold) elevated in senescent HTPCs. This cytokine was identified in HTPCs21,22 and is involved in attracting monocytes and thus is considered a pro-inflammatory factor65. A related smooth muscle cell type in blood vessels showed higher levels of CCL2 with age, causing an inflammatory environment66. However, ELISA-measurements showed that not necessarily CCL2 content was changed in the cell culture media of HTPCs (Supplementary Fig. 5).
In summary, the results describe details of cellular senescence in human testicular cells. Senescence in HTPCs is not associated with a decreased expression of crucial HTPC genes, including contractility markers, AR and the growth factors GDNF and CXCL12. However, striking morphological changes in senescent HTPCs are accompanied by altered cellular protein levels, indicating impaired proteostasis. The secretome analysis revealed that specifically ECM-components are reduced. In contrast, inflammatory factors are increased. The HTPC-specific SASP includes MIF and DPP4, which both, directly or indirectly could influence the SSCs. In particular, DPP4 may be involved in degrading CXCL12. Thereby it could negatively influence the SSC niche. If so, it may serve as a potential drug target.
Reliable data on testicular aging and associated functional and structural changes in man are missing to the very day. Hence, the full in vivo-relevance of the changes in the proteome and secretome of HTPCs, in combination with ultrastructural large volume 3D-data, for the understanding of testicular aging in man remains to be shown.
Methods
HTPCs were isolated from testicular tissue fragments as described earlier13,21. The study was approved by the local ethical committee (Ethikkommission, Technische Universität München, Fakultät für Medizin, project number 309/14). For the scientific use of the tissue samples, the donors had granted written declaration of informed consent. HTPCs were cultured and propagated in DMEM High Glucose (Gibco, Paisly, UK) added with 10% fetal bovine serum (Capricorn Scientific, Ebsdorfergrund, Germany), 1% penicillin/streptomycin (Biochrom, Berlin, Germany) at 37 °C and 5% (v/v) CO2. Purity of the cell cultures was shown previously10. The cells studied derived from donors, with obstructive azoospermia and normal spermatogenesis. The age of the donors ranged from 39 to 55 years. HTPCs underwent serial passaging until the typical signs of senescence appeared (growth arrest, increase in size, β-galactosidase expression, telomere shorting). All these samples were compared with the respective early passage from the same donor. The number of individual patient-derived cells for the different experiments is provided below. All experimental methods were implemented in accordance with relevant guidelines and regulations (including all biosafety and laboratory regulations).
β-galactosidase staining
Cultured HTPCs were seeded onto coverslips. Senescence-associated β-galactosidase staining was performed using a commercial kit (Senescence β-Galactosidase Staining Kit, Cell Signaling Technology #9860, Danvers, MA, USA), according to manufacturer’s instructions. Staining was examined with a Zeiss Axiovert microscope (Zeiss GmbH, Oberkochen, Germany). HTPCs in early (P5 - P7) and advanced (P12 - P15) passages from 5 different donors were used for this experiment.
Cell size measurement
Cell size was determined using the CASY® Cell Counter on basis of variances in electrical resistance (Schärfe Systems, Reutlingen, Germany). HTPCs (from n = 7 individual donors, early (P3 – P7) and advanced (P11 – P14) passages) were trypsinized, centrifuged, resuspended in PBS and the measurement was implemented as described before67. For statistical analysis paired t-test (two-tailed) was applied.
DNA extraction
Total DNA extraction, from HTPCs in early (P3 – P8) and advanced (P12 – P20) passages (n = 6 different donors) was carried out with Wizard® SV Genomic DNA Purification System (Promega, Fitchburg, WI, USA) according to the manufacturer’s instructions.
Relative telomere length quantification
Total DNA from HTPCs (n = 6 donors, early (P3 – P8) and advanced (P12 – P20)) was analyzed with the Relative Telomere Length Quantification qPCR Assay Kit, (Science Cell, Carlsbad, CA, USA) according to the manufacturer’s instructions. The kit contains 2 primer sets, one for the recognition and amplification of the telomere sequence and a reference primer set for data normalization. qPCR was carried out with QuantiFast® SYBR Green PCR Kit (Qiagen, Hilden, Germany). A total amount of 5 ng DNA from cultured HTPCs was used in duplicates in a LightCycler® 96 System (Roche Diagnostics GmbH, Penzberg, Germany) with following conditions: Initial denaturation (95 °C, 10 min) and 32 cycles of denaturation/annealing/extension (95 °C, 20 s/52 °C 20 s/72 °C 45 s). Quantification was implemented with comparative ΔΔCq method. Statistical analysis was done via one-sample t-test.
Mitochondrial DNA copy number quantification
The mtDNA copy number was quantified by qPCR (n = 6 different donors, early (P3 – P8) and advanced (P12 – P20)). A mitochondrial and a nuclear locus were compared as described elsewhere68. qPCR was executed with QuantiFast® SYBR Green PCR Kit (Qiagen) using 5 ng DNA and two different primer sets (for nuclear receptor coactivator three (NCOA3) and mtDNA) (Supplementary Table 7). qPCR conditions: 5 min, 95 °C preincubation, 40 cycles of amplification including denaturation at 95 °C for 10 s, annealing temperature 60 °C for 30 s and a melting step by heating from 65 °C to 95 °C, followed by a cool down to 37 °C for 30 s in a LightCycler® 96 System (Roche). Comparative ΔΔCq method was used for mtDNA quantification and statistically analyzed with one-sample t-test.
FIB/SEM
Cells were seeded on laser marked slides69. For light microscopic investigations cells were fixed in cacodylate buffer and stained with DAPI, as described by Luckner and Wanner69. From each passage 10 representative interphase cells were selected in phase contrast light microscopy according to their cell size and shape and documented with a CCD camera with the corresponding epifluorescence DAPI image. Cells were post-fixed/stained with either osmium tetroxide or reduced osmium-TCH-osmium (=rOTO) and ultra-thin embedded with epoxy resin as described36,69. After polymerization, the cells were documented again by bright field light microscopy for visualization and control of the heavy metal staining again. For high resolution SEM and FIB milling, cells were processed as described in detail by Luckner and Wanner69. Images were recorded with 3072 × 2048 pixel. The resulting data sets were aligned using AmiraTM (Thermo Fisher Scientific, Waltham, MA, USA), first automatically with the module “align slices” and corrected with the “shear” function. The quality of the alignment had to be verified manually by fine correction. Image stacks were segmented and reconstructed in AmiraTM (Thermo Fisher Scientific) and/or processed with a volume-rendering algorithm (volren) for direct visualization. 3D reconstructions/correlations were performed with AmiraTM. Dependent on the scientific demand for 3D reconstruction pixel sizes from 7.5 µm in x/y down to 2 nm isovoxel were chosen.
Nano LC-MS/MS
Cultured HTPCs in early (P3 – P7) and advanced (P11 – P14) passages (n = 6 different donors) were washed five times in serum-free DMEM to remove FBS and cell debris, and incubated for additional 24 h in serum-free DMEM. Cell pellets were harvested, suspended and homogenized in 8 M urea and 50 mM ammonium bicarbonate, as described previously70. The conditioned media (CM) were collected and centrifuged at 1,000 g for 3 min. The supernatant was transferred to Amicon 3 kDa centrifugal filter devices (Millipore) to desalt and concentrate the secreted proteins. The remaining concentrate solute was dried in a vacuum centrifuge. Protein concentration was determined using the Pierce 660 nm Protein Assay (Thermo Fisher Scientific)71. Protein samples from cells (20 µg) and their CM (10 µg) were digested in consecutive incubation steps with Lys-C (enzyme/substrate: 1:100; Wako) for 4 h at 37 °C and trypsin (enzyme/substrate: 1:50; Promega) overnight at 37 °C, as described earlier70. Peptide samples were analyzed by nano-LC-MS/MS on an UltiMateTM 3000 RSLCnano system (Thermo Scientific) coupled to a TripleTOF® 5600 + mass spectrometer (Sciex). 2.5 µg of peptides were separated at a flow rate of 200 nl/min with an analytical column (AcclaimTM PepMapTM RSLC C18, 75 μm × 50 cm, 2 μm, Thermo Fisher Scientific) in consecutive linear gradients: 5–25% solvent B (0.1% formic acid in acetonitrile) in 255 min and 25–50% B in 60 min. MS data were acquired in scan cycles of one survey scan (m/z 400–1250) followed by 70 data dependent CID fragmentation scans. MS raw data were processed using MaxQuant (v. 1.6.1.0). Database search parameters were set to SCIEX TOF instruments and protein identification was performed using the human Swiss-Prot subset (release 2018–10) and the MaxQuant common contaminants database at a false discovery rate of 1%. Label-free quantification (LFQ) was used as quantification strategy with a LFQ min. ratio count of 1 and the match between runs feature enabled. Data analysis and statistics was done in Perseus72 and R73. Gene Ontology (GO) and KEGG pathway annotations were retrieved from UniProt. Protein identifications in the conditioned media data were filtered for extracellular locations to define the secretome protein dataset. The presence of a signal peptide position and the keyword “secreted” in UniProt were used as indication for classical secretion. Paired t-tests were used to identify significantly differentially abundant proteins with an s0 value of 0.174. The FDR was controlled to be < 0.05. Gene set enrichment analysis (GSEA) was done using the Reactome and KEGG databases, as well as GO categories (molecular function, biological process, cellular component). Permutation type was set to gene type, enrichment statistic was weighted and as metric a t-test was used, while the rest of the settings were kept at their defaults. Enrichment maps were generated in Cytoscape75 using the Enrichment map76 and clusterMaker277 apps. The following settings were used for the enrichment maps: FDR q-value was 0.05 and as metric for edge generation an overlap index of 0.5 was used. REVIGO78 was used to summarize significant gene sets by clustering similar ontology terms.
Immunohistochemistry
Immunohistochemical staining was performed as published previously67. Sections from patients with normal spermatogenesis (n = 6, age 48–50 years) were studied (Ethikkommission, Technische Universität München, Fakultät für Medizin, project number 309/14). Primary polyclonal goat anti-human DPP4 antibody (1:40, R&D Systems, Minneapolis, MN, USA) was used. For negative control purposes, the primary antiserum was omitted and replaced by non-immune serum. Hematoxilin was used to counterstain the sections. Examination of the sections was done with a Zeiss Axiovert light microscope (Zeiss GmbH).
Reverse transcription and qPCR
RNeasy Plus Micro Kit (Qiagen) was used for total RNA isolation from HTPCs in early (P3 – P8) and advanced (P12 – P16) passages. 200 ng RNA were reverse transcribed, random 15mer primer and SuperScriptTM II (Invitrogen, Darmstadt, Germany). A LightCycler® 96 System (Roche) and QuantiFast® SYBR Green PCR Kit (Qiagen) were used. Primer are listed in Supplementary Table 7 Results were analyzed according to 2−ΔΔCq method, mRNA expression was normalized to HPRT and RPL19, which served as endogenous references. Results were depicted as means ± SEM. Statistical analysis was done with one-sample t-test of ΔΔCq values using GraphPad Prism 6.0 Software (GraphPad Software, San Diego, CA, USA). Negative controls consisted of non-reverse transcription and non-template reactions.
References
Wu, F. C. et al. Hypothalamic-pituitary-testicular axis disruptions in older men are differentially linked to age and modifiable risk factors: the European Male Aging Study. J Clin Endocrinol Metab 93, 2737–2745, https://doi.org/10.1210/jc.2007-1972 (2008).
Kuhnert, B. & Nieschlag, E. Reproductive functions of the ageing male. Human reproduction update 10, 327–339, https://doi.org/10.1093/humupd/dmh030 (2004).
Honore, L. H. Ageing changes in the human testis: a light-microscopic study. Gerontology 24, 58–65, https://doi.org/10.1159/000212237 (1978).
Meacham, R. B. & Murray, M. J. Reproductive function in the aging male. The Urologic clinics of North America 21, 549–556 (1994).
Paniagua, R., Nistal, M., Amat, P., Rodriguez, M. C. & Martin, A. Seminiferous tubule involution in elderly men. Biology of reproduction 36, 939–947 (1987).
Paniagua, R., Nistal, M., Saez, F. J. & Fraile, B. Ultrastructure of the aging human testis. Journal of electron microscopy technique 19, 241–260, https://doi.org/10.1002/jemt.1060190209 (1991).
Harris, I. D., Fronczak, C., Roth, L. & Meacham, R. B. Fertility and the aging male. Reviews in urology 13, e184–190 (2011).
Nieschlag, E., Lammers, U., Freischem, C. W., Langer, K. & Wickings, E. J. Reproductive functions in young fathers and grandfathers. J Clin Endocrinol Metab 55, 676–681, https://doi.org/10.1210/jcem-55-4-676 (1982).
Welter, H. et al. Partial loss of contractile marker proteins in human testicular peritubular cells in infertility patients. Andrology 1, 318–324, https://doi.org/10.1111/j.2047-2927.2012.00030.x (2013).
Walenta, L. et al. ATP-mediated Events in Peritubular Cells Contribute to Sterile Testicular Inflammation. Sci Rep 8, 1431, https://doi.org/10.1038/s41598-018-19624-3 (2018).
Meineke, V., Frungieri, M. B., Jessberger, B., Vogt, H. & Mayerhofer, A. Human testicular mast cells contain tryptase: increased mast cell number and altered distribution in the testes of infertile men. Fertility and sterility 74, 239–244 (2000).
Welter, H., Kohn, F. M. & Mayerhofer, A. Mast cells in human testicular biopsies from patients with mixed atrophy: increased numbers, heterogeneity, and expression of cyclooxygenase 2 and prostaglandin D2 synthase. Fertility and sterility 96, 309–313, https://doi.org/10.1016/j.fertnstert.2011.05.035 (2011).
Albrecht, M., Ramsch, R., Kohn, F. M., Schwarzer, J. U. & Mayerhofer, A. Isolation and cultivation of human testicular peritubular cells: a new model for the investigation of fibrotic processes in the human testis and male infertility. J Clin Endocrinol Metab 91, 1956–1960, https://doi.org/10.1210/jc.2005-2169 (2006).
Spinnler, K., Kohn, F. M., Schwarzer, U. & Mayerhofer, A. Glial cell line-derived neurotrophic factor is constitutively produced by human testicular peritubular cells and may contribute to the spermatogonial stem cell niche in man. Human reproduction (Oxford, England) 25(2181–2187), 170, https://doi.org/10.1093/humrep/deq. (2010).
Flenkenthaler, F. et al. Secretome analysis of testicular peritubular cells: a window into the human testicular microenvironment and the spermatogonial stem cell niche in man. J Proteome Res 13, 1259–1269, https://doi.org/10.1021/pr400769z (2014).
Mayer, C. et al. Insights into the role of androgen receptor in human testicular peritubular cells. Andrology 6, 756–765, https://doi.org/10.1111/andr.12509 (2018).
Chen, L. Y., Willis, W. D. & Eddy, E. M. Targeting the Gdnf Gene in peritubular myoid cells disrupts undifferentiated spermatogonial cell development. Proceedings of the National Academy of Sciences of the United States of America 113, 1829–1834, https://doi.org/10.1073/pnas.1517994113 (2016).
Paul, C. & Robaire, B. Ageing of the male germ line. Nature reviews. Urology 10, 227–234, https://doi.org/10.1038/nrurol.2013.18 (2013).
Pohl, E. et al. Ageing in men with normal spermatogenesis alters spermatogonial dynamics and nuclear morphology in Sertoli cells. Andrology, https://doi.org/10.1111/andr.12665 (2019).
Kampfer, C. et al. Evidence for an adaptation in ROS scavenging systems in human testicular peritubular cells from infertility patients. International journal of andrology 35, 793–801, https://doi.org/10.1111/j.1365-2605.2012.01281.x (2012).
Schell, C. et al. Exploring human testicular peritubular cells: identification of secretory products and regulation by tumor necrosis factor-alpha. Endocrinology 149, 1678–1686, https://doi.org/10.1210/en.2007-1064 (2008).
Mayer, C. et al. Sterile inflammation as a factor in human male infertility: Involvement of Toll like receptor 2, biglycan and peritubular cells. Sci Rep 6, 37128, https://doi.org/10.1038/srep37128 (2016).
Goto, M. Inflammaging (inflammation + aging): A driving force for human aging based on an evolutionarily antagonistic pleiotropy theory? Bioscience trends 2, 218–230 (2008).
Campisi, J. From cells to organisms: can we learn about aging from cells in culture? Experimental gerontology 36, 607–618 (2001).
Gil, J. Cellular senescence causes ageing. Nature reviews. Molecular cell biology 20, 388, https://doi.org/10.1038/s41580-019-0128-0 (2019).
McHugh, D. & Gil, J. Senescence and aging: Causes, consequences, and therapeutic avenues. The Journal of cell biology 217, 65–77, https://doi.org/10.1083/jcb.201708092 (2018).
Rodier, F. et al. Persistent DNA damage signalling triggers senescence-associated inflammatory cytokine secretion. Nature cell biology 11, 973–979, https://doi.org/10.1038/ncb1909 (2009).
Petkova, D. S., Viret, C. & Faure, M. IRGM in autophagy and viral infections. Frontiers in immunology 3, 426, https://doi.org/10.3389/fimmu.2012.00426 (2012).
Hayflick, L. & Moorhead, P. S. The serial cultivation of human diploid cell strains. Experimental cell research 25, 585–621 (1961).
Lopez-Otin, C., Blasco, M. A., Partridge, L., Serrano, M. & Kroemer, G. The hallmarks of aging. Cell 153, 1194–1217, https://doi.org/10.1016/j.cell.2013.05.039 (2013).
Harley, C. B., Futcher, A. B. & Greider, C. W. Telomeres shorten during ageing of human fibroblasts. Nature 345, 458–460, https://doi.org/10.1038/345458a0 (1990).
Hernandez-Segura, A., Nehme, J. & Demaria, M. Hallmarks of Cellular Senescence. Trends in cell biology, https://doi.org/10.1016/j.tcb.2018.02.001 (2018).
Kurz, D. J., Decary, S., Hong, Y. & Erusalimsky, J. D. Senescence-associated (beta)-galactosidase reflects an increase in lysosomal mass during replicative ageing of human endothelial cells. Journal of cell science 113(Pt 20), 3613–3622 (2000).
Dimri, G. P. et al. A biomarker that identifies senescent human cells in culture and in aging skin in vivo. Proceedings of the National Academy of Sciences of the United States of America 92, 9363–9367 (1995).
Kaushik, S. & Cuervo, A. M. Proteostasis and aging. Nature medicine 21, 1406–1415, https://doi.org/10.1038/nm.4001 (2015).
Luckner, M. & Wanner, G. Precise and economic FIB/SEM for CLEM: with 2 nm voxels through mitosis. Histochemistry and cell biology 150, 149–170, https://doi.org/10.1007/s00418-018-1681-x (2018).
Carmona-Gutierrez, D., Hughes, A. L., Madeo, F. & Ruckenstuhl, C. The crucial impact of lysosomes in aging and longevity. Ageing research reviews 32, 2–12, https://doi.org/10.1016/j.arr.2016.04.009 (2016).
Szeto, H. H. et al. Mitochondria Protection after Acute Ischemia Prevents Prolonged Upregulation of IL-1beta and IL-18 and Arrests CKD. Journal of the American Society of Nephrology: JASN 28, 1437–1449, https://doi.org/10.1681/asn.2016070761 (2017).
Nguyen, T. N. et al. Atg8 family LC3/GABARAP proteins are crucial for autophagosome-lysosome fusion but not autophagosome formation during PINK1/Parkin mitophagy and starvation. The Journal of cell biology 215, 857–874, https://doi.org/10.1083/jcb.201607039 (2016).
Rizzuto, R. et al. Close contacts with the endoplasmic reticulum as determinants of mitochondrial Ca2+ responses. Science (New York, N.Y.) 280, 1763–1766 (1998).
English, A. R. & Voeltz, G. K. Endoplasmic reticulum structure and interconnections with other organelles. Cold Spring Harbor perspectives in biology 5, a013227, https://doi.org/10.1101/cshperspect.a013227 (2013).
Feng, Y., He, D., Yao, Z. & Klionsky, D. J. The machinery of macroautophagy. Cell research 24, 24–41, https://doi.org/10.1038/cr.2013.168 (2014).
Lamb, C. A., Yoshimori, T. & Tooze, S. A. The autophagosome: origins unknown, biogenesis complex. Nature reviews. Molecular cell biology 14, 759–774, https://doi.org/10.1038/nrm3696 (2013).
Bernal, G. M. et al. Loss of Nfkb1 leads to early onset aging. Aging 6, 931–943, https://doi.org/10.18632/aging.100702 (2014).
Shimada, N., Rios, I., Moran, H., Sayers, B. & Hubbard, K. p38 MAP kinase-dependent regulation of the expression level and subcellular distribution of heterogeneous nuclear ribonucleoprotein A1 and its involvement in cellular senescence in normal human fibroblasts. RNA biology 6, 293–304 (2009).
LaBranche, H. et al. Telomere elongation by hnRNP A1 and a derivative that interacts with telomeric repeats and telomerase. Nature genetics 19, 199–202, https://doi.org/10.1038/575 (1998).
Barzilai, N., Huffman, D. M., Muzumdar, R. H. & Bartke, A. The critical role of metabolic pathways in aging. Diabetes 61, 1315–1322, https://doi.org/10.2337/db11-1300 (2012).
Bratic, I. & Trifunovic, A. Mitochondrial energy metabolism and ageing. Biochimica et biophysica acta 1797, 961–967, https://doi.org/10.1016/j.bbabio.2010.01.004 (2010).
Ross, J. M. et al. High brain lactate is a hallmark of aging and caused by a shift in the lactate dehydrogenase A/B ratio. Proceedings of the National Academy of Sciences of the United States of America 107, 20087–20092, https://doi.org/10.1073/pnas.1008189107 (2010).
Sun, L., Yu, R. & Dang, W. Chromatin Architectural Changes during Cellular Senescence and Aging. Genes 9, https://doi.org/10.3390/genes9040211 (2018).
Collin, G., Huna, A., Warnier, M., Flaman, J. M. & Bernard, D. Transcriptional repression of DNA repair genes is a hallmark and a cause of cellular senescence. Cell death & disease 9, 259, https://doi.org/10.1038/s41419-018-0300-z (2018).
d’Adda di Fagagna, F. et al. A DNA damage checkpoint response in telomere-initiated senescence. Nature 426, 194–198, https://doi.org/10.1038/nature02118 (2003).
Jankauskas, S. S., Wong, D. W. L., Bucala, R., Djudjaj, S. & Boor, P. Evolving complexity of MIF signaling. Cellular signalling 57, 76–88, https://doi.org/10.1016/j.cellsig.2019.01.006 (2019).
Subbannayya, T. et al. An integrated signal transduction network of macrophage migration inhibitory factor. Journal of cell communication and signaling 10, 165–170, https://doi.org/10.1007/s12079-016-0326-x (2016).
Calandra, T. & Bucala, R. Macrophage Migration Inhibitory Factor (MIF): A Glucocorticoid Counter-Regulator within the Immune System. Critical reviews in immunology 37, 359–370, https://doi.org/10.1615/CritRevImmunol.v37.i2-6.90 (2017).
Salminen, A. & Kaarniranta, K. Control of p53 and NF-kappaB signaling by WIP1 and MIF: role in cellular senescence and organismal aging. Cellular signalling 23, 747–752, https://doi.org/10.1016/j.cellsig.2010.10.012 (2011).
Heckmann, L., Pock, T., Trondle, I. & Neuhaus, N. The C-X-C signalling system in the rodent vs primate testis: impact on germ cell niche interaction. Reproduction (Cambridge, England) 155, R211–r219, https://doi.org/10.1530/rep-17-0617 (2018).
Kim, K. M. et al. Identification of senescent cell surface targetable protein DPP4. Genes &. development 31, 1529–1534, https://doi.org/10.1101/gad.302570.117 (2017).
Janssens, R. et al. Truncation of CXCL12 by CD26 reduces its CXC chemokine receptor 4- and atypical chemokine receptor 3-dependent activity on endothelial cells and lymphocytes. Biochemical pharmacology 132, 92–101, https://doi.org/10.1016/j.bcp.2017.03.009 (2017).
Campbell, R. K. Rationale for dipeptidyl peptidase 4 inhibitors: a new class of oral agents for the treatment of type 2 diabetes mellitus. The Annals of pharmacotherapy 41, 51–60, https://doi.org/10.1345/aph.1H459 (2007).
Christopherson, K. W. II, Hangoc, G., Mantel, C. R. & Broxmeyer, H. E. Modulation of hematopoietic stem cell homing and engraftment by CD26. Science (New York, N.Y.) 305, 1000–1003, https://doi.org/10.1126/science.1097071 (2004).
Matzkin, M. E. et al. Aging in the Syrian hamster testis: Inflammatory-oxidative status and the impact of photoperiod. Experimental gerontology 124, 110649, https://doi.org/10.1016/j.exger.2019.110649 (2019).
Rong, N. et al. Restoring extracellular matrix synthesis in senescent stem cells. FASEB journal: official publication of the Federation of American Societies for Experimental Biology, fj201900377R, https://doi.org/10.1096/fj.201900377R (2019).
Rey-Ares, V. et al. Prostaglandin E2 (PGE2) is a testicular peritubular cell-derived factor involved in human testicular homeostasis. Molecular and cellular endocrinology 473, 217–224, https://doi.org/10.1016/j.mce.2018.01.022 (2018).
Deshmane, S. L., Kremlev, S., Amini, S. & Sawaya, B. E. Monocyte chemoattractant protein-1 (MCP-1): an overview. Journal of interferon & cytokine research: the official journal of the International Society for Interferon and Cytokine Research 29, 313–326, https://doi.org/10.1089/jir.2008.0027 (2009).
Song, Y. et al. Aging enhances the basal production of IL-6 and CCL2 in vascular smooth muscle cells. Arteriosclerosis, thrombosis, and vascular biology 32, 103–109, https://doi.org/10.1161/atvbaha.111.236349 (2012).
Schell, C. et al. 15-Deoxy-delta 12-14-prostaglandin-J2 induces hypertrophy and loss of contractility in human testicular peritubular cells: implications for human male fertility. Endocrinology 151, 1257–1268, https://doi.org/10.1210/en.2009-1325 (2010).
Jackson, C. B., Gallati, S. & Schaller, A. qPCR-based mitochondrial DNA quantification: influence of template DNA fragmentation on accuracy. Biochem Biophys Res Commun 423, 441–447, https://doi.org/10.1016/j.bbrc.2012.05.121 (2012).
Luckner, M. & Wanner, G. From Light Microscopy to Analytical Scanning Electron Microscopy (SEM) and Focused Ion Beam (FIB)/SEM in Biology: Fixed Coordinates, Flat Embedding, Absolute References. Microscopy and microanalysis: the official journal of Microscopy Society of America, Microbeam Analysis. Society, Microscopical Society of Canada 24, 526–544, https://doi.org/10.1017/s1431927618015015 (2018).
Schmid, N. et al. Characterization of a nonhuman primate model for the study of testicular peritubular cells - comparison with human testicular cells. Mol. Hum. Reprod., https://doi.org/10.1093/molehr/gay025 (2018).
Antharavally, B. S., Mallia, K. A., Rangaraj, P., Haney, P. & Bell, P. A. Quantitation of proteins using a dye-metal-based colorimetric protein assay. Anal. Biochem. 385, 342–345, https://doi.org/10.1016/j.ab.2008.11.024 (2009).
Tyanova, S. et al. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat. Methods 13, 731–740, https://doi.org/10.1038/nmeth.3901 (2016).
R: A Language and Environment for Statistical Computing (R Foundation for Statistical Computing, Vienna, Austria, 2018).
Tusher, V. G., Tibshirani, R. & Chu, G. Significance analysis of microarrays applied to the ionizing radiation response. Proceedings of the National Academy of Sciences of the United States of America 98, 5116–5121, https://doi.org/10.1073/pnas.091062498 (2001).
Shannon, P. et al. Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504, https://doi.org/10.1101/gr.1239303 (2003).
Merico, D., Isserlin, R., Stueker, O., Emili, A. & Bader, G. D. Enrichment map: a network-based method for gene-set enrichment visualization and interpretation. PLoS One 5, e13984, https://doi.org/10.1371/journal.pone.0013984 (2010).
Morris, J. H. et al. clusterMaker: a multi-algorithm clustering plugin for Cytoscape. BMC Bioinformatics 12, 436, https://doi.org/10.1186/1471-2105-12-436 (2011).
Supek, F., Bosnjak, M., Skunca, N. & Smuc, T. REVIGO summarizes and visualizes long lists of gene ontology terms. PLoS One 6, e21800, https://doi.org/10.1371/journal.pone.0021800 (2011).
Acknowledgements
The authors thank Astrid Tiefenbacher for skillful technical assistance. The work was supported by grants from the Deutsche Forschungsgemeinschaft (DFG, MA 1080/27-1; AR 362/9-1).
Author information
Authors and Affiliations
Contributions
N.S. performed the majority of the cellular experiments and evaluated results. K.G.D. was involved in the culture and characterization of cells and cellular experiments. M.L. performed rOTO and ultra-thin embedding. N.S. and G.W. performed FIB/SEM-tomography and 3D-reconstruction. F.F., J.B.S., T.F. and G.A. performed the proteomic studies and evaluated the results. J.U.S., F.M.K. provided testicular tissues and provided together with L.K. conceptual input. A.M. conceived of the study, supervised the experiments. All authors provided helpful comments. N.S., G.W. and A.M. drafted the manuscript, and all authors contributed to the final version.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Schmid, N., Flenkenthaler, F., Stöckl, J.B. et al. Insights into replicative senescence of human testicular peritubular cells. Sci Rep 9, 15052 (2019). https://doi.org/10.1038/s41598-019-51380-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-51380-w
This article is cited by
-
Integrative Proteomics-Metabolomics of In Vitro Degeneration of Cardiovascular Cell Lines
Applied Biochemistry and Biotechnology (2024)
-
Autophagy, a critical element in the aging male reproductive disorders and prostate cancer: a therapeutic point of view
Reproductive Biology and Endocrinology (2023)
-
Functional histology of human scrotal wall layers and their overlooked relation with infertility: a narrative review
International Journal of Impotence Research (2023)
-
Inflammation and aging: signaling pathways and intervention therapies
Signal Transduction and Targeted Therapy (2023)
-
The landscape of human tissue and cell type specific expression and co-regulation of senescence genes
Molecular Neurodegeneration (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.