Abstract
The weed wall pellitory, Parietaria judaica, is one the most important pollen allergen sources in the Mediterranean area causing severe symptoms of hay fever and asthma in allergic patients. We report the expression of the major Parietaria allergens, Par j 1 and Par j 2 which belong to the family of lipid transfer proteins, in insect cells. According to circular dichroism analysis and gel filtration, the purified allergens represented folded and monomeric proteins. Insect cell-expressed, folded Par j 2 exhibited higher IgE binding capacity and more than 100-fold higher allergenic activity than unfolded Escherichia coli-expressed Par j 2 as demonstrated by IgE ELISA and basophil activation testing. IgE ELISA inhibition assays showed that Par j 1 and Par j 2, contain genuine and cross-reactive IgE epitopes. IgG antibodies induced by immunization with Par j 2 inhibited binding of allergic patients IgE to Par j 1 only partially. IgE inhibition experiments demonstrated that insect cell-expressed Par j 1 and Par j 2 together resembled the majority of allergenic epitopes of the Parietaria allergome and therefore both should be used for molecular diagnosis and the design of vaccines for allergen-specific immunotherapy of Parietaria allergy.
Similar content being viewed by others
Introduction
IgE-associated allergy is the most common hypersensitivity disease affecting more than 25% of the world´s population1,2. Mainly in the Mediterranean area but also at the southern coast of UK, France, Australia, California and Eastern Europe the weed Parietaria judaica, wall pellitory, is one the most common pollen allergen sources3,4,5,6,7. Parietaria judaica pollen is responsible for severe forms of respiratory allergy (i.e., severe allergic rhinitis, asthma) which may be due to the fact that the plant has an unusually long period of pollination ranging from February to November8,9,10. Already an early report from 1956 identified Parietaria as important allergen source responsible for allergic asthma, one of the most severe manifestations of allergy11. Approximately 40 years later the molecular nature of the two most important allergens from Parietaria, Par j 1 and Par j 2 was revealed by cDNA cloning techniques12,13. Par j 1 and Par j 2 are non-glycosylated allergens with a molecular weight of 15 kDa and 11.3 kDa respectively. They belong to the family of lipid transfer proteins (LTP) which typically consist of four alpha-helices that are linked and stabilized by four disulphide bonds14,15,16,17. In addition to Par j 1 and Par j 2, highly cross-reactive allergens belonging to the family of profilins and calcium-binding allergens are minor allergens in Parietaria which are recognized typically by less than 10% of the patients18,19. The expression and purification of recombinant Par j 1 and Par j 2 resembling the fold of the corresponding wild-type allergens is difficult and requires eukaryotic hosts such as yeast expression systems capable of forming correct disulphide bonds20. In fact, it was reported that yeast-expressed Par j 1 and Par j 2 showed a similar fold as the natural allergens. Yeast-expressed Par j 1 and Par j 2 were recognized by sera from Parietaria-allergic patients and exhibited allergenic activity as demonstrated by skin testing20. Accordingly the recombinant allergens were suggested to be useful for the diagnosis of allergy to Parietaria by component-resolved diagnosis. In fact, molecular allergy diagnosis offers important advantages over conventional allergen-extract-based diagnosis because it allows revealing the allergic patients molecular sensitization pattern and thus aids in the diagnostic resolution of difficult cases and in the refined prescription of allergen-specific immunotherapy21,22,23, the only causal and disease-modifying treatment for IgE-associated allergies24. In the meantime, molecular approaches for allergen-specific immunotherapy have been successfully evaluated in many clinical trials and hold promise to improve AIT in the future25,26. In the case of Parietaria allergy several approaches have been considered27,28,29 but the question remains what allergen molecules need to be included in the vaccine. Regarding Par j 1 and Par j 2 it is known that the allergens contain cross-reactive epitopes30 but the open question is if, due to cross-reactivity, one can replace the other allergen. Furthermore there are no detailed studies investigating the allergenic activity of the two allergens. Finally and importantly, it has not yet been determined what allergens can resemble the spectrum of allergenic epitopes of the Parietaria allergome to replace allergen extract-based vaccines.
In order to address these open questions we have expressed folded rPar j 1 and rPar j 2 in insect cells and studied the allergenic activity of both recombinant allergens by titrated basophil activation testing. Furthermore we studied the extent of IgE cross-reactivity of rPar j 1 and rPar j 2 by IgE inhibition studies and demonstrated that only both allergens together resemble the spectrum of IgE epitopes of natural Parietaria pollen extract.
Results
Physicochemical characterization of purified recombinant Par j 2 and Par j 1
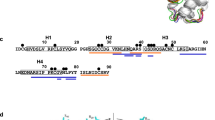
Figure 1a shows an alignment of the deduced amino acid sequences of Par j 2 (P55958) and Par j 1 (O04404). In the overlapping region the proteins show a 53% sequence identity. Par j 1 contains additional 37 amino acids at its C-terminal end. The hydrophobicity prediction performed with the Kyte and Doolittle algorithm shows that the proteins contain a highly hydrophobic N-terminus followed by a hydrophilic stretch and a region with intermediate hydrophobicity. The C-terminus of Par j 1 shows high hydrophilicity. Par j 1 and Par j 2 contain eight conserved cysteine residues (Fig. 1, boxed). Supplementary Fig. S1a displays the sequence identities of Par j 2 and Par j 1 with LTPs identified as allergens in other plants and in plant tissues other than pollen. Both, Par j 2 and Par j 1, showed a relatively low sequence identity of less than 35% with the other plant LTP allergens whereas sequence identities of up to >90% (e.g., Pru p 1 vs. Pru ar 3) between certain LTPs, mainly from somatic tissues (i.e., plant food) were found (Supplementary Fig. S1a, colors). Accordingly, the phylogenetical comparison analysis of the LTPs shows that Par j 1 and Par j 2 represent an independent branch highlighting the evolutionary differences with other proteins of this family as represented in Supplementary Fig. S1b. However, it should be noted that neither the degree of sequence identities nor the phylogenetic relationships of the different LTPs were associated with the expression in certain plant tissues (pollen versus somatic tissues) or with the botanical relationships of the corresponding plants (monocotyledonic versus dicotyledonic plants).
We expressed recombinant Par j 1 and Par j 2 proteins in baculovirus-infected insect cells and named them BvPar j 1 and BvPar j 2, respectively. Both proteins were compared with recombinant Par j 2 expressed in E. coli (EcPar j 2)12. Recombinant proteins were purified by Ni2+-affinity chromatography to homogeneity. Purified proteins were then analysed by SDS-PAGE under reducing and non-reducing conditions (Fig. 2a). BvPar j 2 migrated as single band of approximately 15 kDa under non-reducing conditions and as 14 kDa band under reducing conditions using ß-Mercaptoethanol. Under non-reducing conditions EcPar j 2 showed a main band of approximately 14 kDa and an additional band of 27 kDa and appeared as 13 kDa band when using ß-Mercaptoethanol. BvPar j 1 migrated as a monomeric band of approximately 18 kDa under non-reducing and as 17 kDa under reducing conditions (ß-Mercaptoethanol) (Fig. 2a).
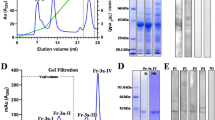
Biochemical and biophysical characterization of purified BvPar j 1, BvPar j 2 and EcPar j 2. Coomassie brilliant blue-stained SDS-PAGE containing purified BvPar j 1, BvPar j 2 and EcPar j 2 (a) separated under non-reducing conditions (panel NR), reducing conditions using β-Mercaptoethanol (panel Me) or Tris(2-carboxyethyl)phosphine (panel TCEP). Molecular weight markers are indicated on the left margins in kilo Dalton (kDa). Full-length gels are presented in Supplementary Fig. S1. Matrix-assisted laser desorption and ionization-time-of-flight (MALDI-Tof) analysis of (b) BvPar j 1, (c) BvPar j 2 (black) and EcPar j 2 (grey). The mass/charge ratios are indicated on the x-axes, and the intensities are shown as absorption units of the most intensive signals obtained in the mass range (y-axes). Analysis of (d) BvPar j 1, (e) BvPar j 2 (black) and EcPar j 2 (grey) by gel filtration. Cytochrome C (dotted peak) and carbonic anhydrase (indicated by an arrow at 29 kDa) served as molecular weight standards. Elution volumes are shown in milliliters - mL (x-axes) and the absorbances of the proteins at 280 nm are indicated in milli-absorbance units - mAU (y-axes).
In SDS-PAGE both insect cell-expressed proteins appeared to have greater masses than their calculated masses (BvPar j 2: 12,380 kDa; BvPar j 1: 16,020 kDa) whereas the mass of the bacterially expressed EcPar j 2 protein corresponded to the predicted mass (i.e., 12,744 kDa) under reducing and non-reducing conditions and a band of approximately 26 kDa likely to be a dimer. Taking into account that the structures of LTPs are stabilized by 4 disulphide bonds the discrepancy in masses might be explained by retained fold of the proteins even under denaturing conditions. We therefore performed SDS PAGE using Tris(2-carboxyethyl)phosphine (TCEP), a reagent which is more potent in reducing disulphide bonds than ß-Mercaptoethanol (Fig. 2a., right part). When TCEP was used EcPar j 2 appeared as a monomeric band of predicted mass and now BvPar j 1 and BvPar j 2 exhibited masses which corresponded to those which had been calculated (Fig. 2a).
Next, the purified proteins were analysed by MALDI mass spectrometry where BvPar j 1 showed a peak of 16,039 kDa (Fig. 2b) corresponding to the mass calculated from the amino acid sequence (16,020 kDa).
When BvPar j 2 and EcPar j 2 were checked by mass spectrometry, both showed high intensity peaks with masses of 12,376 kDa and 13,092 kDa, respectively (Fig. 2c). The measured mass of BvPar j 2 fitted the calculated mass of 12,380 kDa whereas the mass measured for EcPar j 2 was higher than the expected mass of 12,744 kDa which may be attributed to the presence of an N-terminal methionine. When comparing BvPar j 2 and EcPar j 2 in size exclusion experiments by gel filtration both insect cell- and E. coli-expressed proteins appeared at about the same elution volume as the standard Cytochrome C (EcPar j 2 > BvPar j 2) (12.4 kDa) (Fig. 2e). BvPar j 2 contained also a small dimer peak of approximately 25 kDa and EcPar j 2 showed the presence of dimers and trimers of approximately 26 kDa and 39 kDa, respectively (Fig. 2e). BvPar j 1 occurred as a strictly monomeric form confirming the data obtained by mass spectrometry (Fig. 2b).
We then compared the secondary structure of the proteins expressed in insect cells and E. coli by analysis of their far-ultraviolet circular dichroism (far-UV CD) spectra (Fig. 3). The spectra of BvPar j 1 (Fig. 3a) and BvPar j 2 (Fig. 3b) showed almost the same curves resembling α-helical structures with theoretical minima at 208 nm, a defined shoulder at 221 nm, and a maximum at 190 nm whereas EcPar j 2 showed an unordered conformation (Fig. 3b).
Insect cell-expressed Par j 2 shows significantly higher IgE reactivity and allergenic activity than E. coli-expressed Par j 2: Importance of conformational IgE epitopes
IgE reactivity of BvPar j 2 and EcPar j 2 was compared by testing sera from 27 allergic patients (Supplementary Table S1) (Fig. 4a). Approximately one third of the sera showed higher IgE reactivity to BvPar j 2 as compared to EcPar j 2 whereas the other sera displayed similar IgE reactivity to the two proteins. Serum from a non-allergic subject showed no IgE reactivity. Correct coating of the proteins to the ELISA plates was confirmed with rabbit-anti-Par j 2 antibodies (data not shown). The comparison of the IgE levels specific for BvPar j 2 and EcPar j 2 demonstrated that BvPar j 2-specific IgE levels were significantly higher than EcPar j 2-specific IgE levels (Fig. 4b).
IgE reactivity of BvPar j 2 (black) and EcPar j 2 (grey) demonstrated by ELISA. IgE reactivity (y-axis: mean OD values +/− SD) of sera from 27 Parietaria allergic patients (x-axis) to BvPar j 2 and EcPar j 2. The buffer control was subtracted from the data and cut-off is represented by dashed line. (a) Box plot representation (whiskers = minimum and maximum; boxes = 25th to 75th percentiles, means: horizontal bars) of the IgE reactivity of the above sera to BvPar j 2 and EcPar j 2. (b) Statistically significant differences are indicated (*P < 0.05).
Next we compared the allergenic activity of BvPar j 2, BvPar j 1 and EcPar j 2 in basophil activation experiments. Rat basophil leukemia cells transfected with the human FcεRI receptor were loaded with serum IgE from nine Par j 1 and Par j 2 allergic patients and incubated with 5 different concentrations of each protein (0.01, 0.1, 1, 10, 100 ng/ml) or buffer only. BvPar j 2 was 100–1000-fold more potent in inducing mediator release as compared to EcPar j 2 when comparing the concentrations which induced a comparable release of mediators (Fig. 5). BvPar j 1 was less effective in inducing basophil activation as compared to BvPar j 2 in eight of the nine patients (Fig. 5). Stronger basophil activation with BvPar j 1 compared to BvPar j 2 was noted for two patients (i.e., patients, 11 and 13).
Comparison of the allergenic activity of BvPar j 1, BvPar j 2 and EcPar j 2. RBL cells were loaded with sera from nine Parietaria allergic patients (1, 2, 4, 9, 11, 13, 17, 22, 26) and stimulated with different allergen concentrations (x-axes). β-hexosaminidase releases are expressed as percentages of total mediator contents +/− SD (y-axes).
Par j 2 and Par j 1 show IgE cross-reactivity but represent distinct allergens
IgE cross-reactivity between Par j 1 and Par j 2 was investigated by IgE inhibition ELISA experiments (Fig. 6). Sera from Parietaria allergic patients were pre-adsorbed either with Par j 1 (grey) or Par j 2 (black) and then tested for remaining IgE reactivity to the same and to the other allergen (Fig. 6). Figure 6 shows the results of the IgE inhibitions as percentages of auto-inhibition obtained with the same allergen. Par j 2 inhibited IgE binding to Par j 1 better than Par j 1 to Par j 2 in seven of the nine tested patients (i.e., #5, 8, 10, 12, 15, 18). A similar cross-inhibition of IgE binding was found for two of the nine patients (i.e., #3, 20) (Fig. 6). When we calculated the mean percentages of IgE inhibition we found out that Par j 2 inhibited IgE binding to Par j 1 stronger (average 87%) than Par j 1 to Par j 2 (average 68%).
IgE cross-reactivity of Par j 1 and Par j 2 studied by IgE inhibition ELISA. Shown are the percentages of inhibition of IgE binding to BvPar j 1 and BvPar j 2 achieved by pre-incubating sera from patients (x-axis) with the other allergen (Inhibitors: BvPar j 1 grey column; BvPar j 2 black column). Results are expressed as percentages of auto-inhibition +/− SD (y-axis).
Next we tested rabbit IgG antibodies obtained by immunization with BvPar j 2 for their ability to inhibit allergic patients’ IgE binding to Par j 2 and Par j 1 (Table 1). We found that rabbit anti-BvPar j 2 antibodies inhibited strongly allergic patients IgE binding to Par j 2 (77.7–93.7%; 86.7% mean inhibition) whereas IgE binding to Par j 1 was inhibited to a much lower extent (41.2–73.3%; 56.8% mean inhibition) (Table 1), even for sera which had shown strong cross-reactivity (e.g., patients #3, #20, Fig. 6, Table 1).
Recombinant Par j 1 and Par j 2 resemble the majority of allergenic IgE epitopes in Parietaria judaica pollen
The comparison of IgE levels specific for BvPar j 1, BvPar j 2 and Parietaria judaica extract indicates that BvPar j 2-specific IgE levels represent a higher proportion of IgE-specific for natural Parietaria allergens than BvPar j 1 (Fig. 7a). Therefore, we were interested to study in detail the contribution of BvPar j 1 and BvPar j 2-specific IgE to IgE specific for natural Parietaria allergens. For this purpose, sera from Parietaria allergic patients were pre-incubated with BvPar j 1, BvPar j 2, a mix of BvPar j 1 and BvPar j 2 and for control purposes with natural Parietaria allergens and remaining IgE binding to natural Parietaria allergens was measured to calculate the percentage of inhibition (Fig. 7b). The inhibition of IgE binding to natural Parietaria allergens after pre-adsorption with BvPar j 1 ranged from 28.8% to 65.5% (mean 45.2%) and after pre-incubation with BvPar j 2 it ranged from 61.9% to 84.8% (mean 73.1%). The mix of BvPar j 1 and BvPar j 2 inhibited IgE binding to natural Parietaria allergens (i.e., 71.9–94.1%; mean 85.1%) almost as good as natural Parietaria allergens (i.e., 76.2–97.1%; mean 91%) (Fig. 7b).
Contribution of Par j 1 and Par j 2-specific IgE to IgE specific for natural Parietaria allergens. OD values corresponding to IgE levels specific for BvPar j 1, BvPar j 2 and natural Parietaria judaica pollen extract are shown as (a) box plot representation (whiskers = minimum and maximum; boxes = 25th to 75th percentiles, means: horizontal bars) (y-axis: OD values). (b) Inhibition of IgE binding of sera to Parietaria judaica extract by pre-incubation of sera with BvPar j 1, BvPar j 2, with a mix of the two proteins or with extract. Scatter plots showing the percentages of inhibition of IgE binding, horizontal lines indicate means ± SD.
Discussion
The weed wall pellitory, Parietaria judaica, is a major allergen source responsible for severe forms of respiratory allergy in the Mediterranean area in Europe as well as in certain parts of USA and Australia. Allergen-specific immunotherapy the only causative and disease-modifying treatment for allergy is based on the vaccination of allergic patients with the disease-causing allergens24,31. Allergy vaccines based on natural allergen extracts are difficult to produce in a form which meets the standards of modern vaccines in terms of quality and reproducible production according to good manufacturing practice (GMP) conditions32. Accordingly, new types of allergy vaccines based on recombinant allergens, recombinant hypoallergens or allergen-derived peptides are currently being developed and show promising results in clinical trials26,33,34. A first crucial step towards the development of a recombinant allergen-based vaccine approach for a given allergen source is the identification of the clinically most important allergen molecules which resemble the epitope spectrum of the allergen source35. Our study addresses several important open questions which need to be answered for the preparation of a molecular vaccine approach for the treatment of allergy to Parietaria judaica. The first important question is whether a Parietaria vaccine should contain only Par j 2 or Par j 1 or both proteins. In fact it has been shown that the major Parietaria allergens, Par j 1 and Par j 2 contain cross-reactive IgE epitopes but it has not been studied in detail if IgE cross-reactivity is extensive enough so that one of the two proteins can replace the other30. To address this question we have expressed folded versions of Par j 1 and Par j 2 in insect cells. The insect cell-expressed Par j 1 and Par j 2 expressed by us seemed to resemble natural Par j 1 and Par j 2 better than the previously reported recombinant Par j 1 and Par j 2 expressed in yeast because the yeast-expressed proteins were glycosylated whereas the natural proteins were not20. Of note, the yeast-expressed Par j 1 displayed even a mixture of two bands of 18.6 and 16.7 kDa. By contrast, our insect cell expressed proteins displayed single bands and resembled the exact molecular weight predicted according to their amino acid sequence without glycosylation. This is important, because it is known that hyperglycosylation as it occurs upon yeast expression can mask IgE epitopes36.
We found that recombinant folded Par j 2 had higher IgE binding capacity and allergenic activity than unfolded Par j 2 demonstrating the importance of conformational epitopes in addition to sequential epitopes for IgE reactivity to Par j 2. IgE inhibition experiments performed with Par j 1 and Par j 2 indicated that the allergens contain cross-reactive IgE epitopes but none of the two proteins could completely inhibit IgE binding to the other allergen. These results were also corroborated by the finding that IgG antibodies induced by immunization of rabbits with Par j 2 inhibited allergic patients IgE binding to Par j 2 almost completely but inhibited allergic patients IgE binding to Par j 1 only partially. We therefore concluded that both, Par j 1 and Par j 2 need to be part of a vaccine for allergy to Parietaria which is in contrast to a previously published studies concluding that Par j 2 is sufficient for diagnosis and allergen-specific immunotherapy (AIT)16,20,37.
Our conclusion, that both, Par j 1 and Par j 2 are needed for AIT of Parietaria allergy, was confirmed by two other observations: The comparison of the allergenic activity of Par j 1 and Par j 2 in basophil activation tests showed that both proteins are highly allergenic and induced strong basophil degranulation in vitro which is a good surrogate marker for the in vivo allergenic activity of an allergen38,39.
Importantly, we found different types of Parietaria allergic patients, one type, i.e., the majority of patients who were more sensitive to Par j 2 but also another type of patients who reacted stronger to Par j 1.
Finally, yet another important question remained to be answered: Do Par j 1 and Par j 2 resemble the majority of IgE epitopes of a natural Parietaria allergen extract or are other allergens needed? In fact, profilins and calcium-binding allergens have been described as cross-reactive allergens in Parietaria which are recognized by approximately 10% of the patients18,19. To address the latter question we performed IgE inhibition studies which demonstrated that a mix of Par j 1 and Par j 2 almost completely inhibited IgE binding to Parietaria extract and thus resembled the IgE epitopes of a complete natural Parietaria allergen extract. Of note, the mixture of the insect cell-expressed rPar j 1 and rPar j 2 reported by us seemed to inhibit IgE binding to Parietaria pollen extract better (mean inhibition 85.1% which was almost as good as natural Parietaria allergens mean inhibition 91%) than the previously reported rPar j 1 and rPar j 2 expressed in yeast20.
Our study thus identifies Par j 1 and Par j 2 as the two allergen molecules which need to be included in a molecular vaccine for the treatment of Parietaria allergy. In fact, the two recombinant folded versions of Par j 1 and Par j 2 could be formulated as allergy vaccine containing wildtype-like recombinant allergens as has been shown for a birch pollen allergy vaccine based on recombinant wildtype-like major birch pollen allergen Bet v 1 which was effective in clinical trials40,41. Likewise it was shown that a mix of the major recombinant grass pollen allergens, Phl p 1, Phl p 2, Phl p 5 and Phl p 6 which were part of a recombinant allergen-based grass pollen vaccine was effective to treat grass pollen allergy42. Accordingly one may consider formulating a recombinant wild-type allergen vaccine based on Par j 1 and Par j 2 for the treatment of Parietaria allergy. However, the recombinant wild-type-like folded Par j 1 and Par j 2 molecules may be used also as fully allergenic benchmarks for the construction of vaccines based on recombinant hypoallergenic derivatives of Par j 1 and Par j 2 as described for the recently developed hypoallergenic grass pollen allergy vaccine, BM3243,44,45.
Materials and Methods
Paritaria judaica pollen extract
An allergen extract was prepared from Parietaria judaica pollen which was obtained from ALK-Abelló (Madrid, Spain). An aqueous extract was prepared by stirring 1 g of pollen in physiologic buffer (PBS, pH = 7.4) during 4 hours at RT followed by two rounds of centrifugation at 15.000xg for 15 min to remove insoluble particles. The supernatant was dialyzed against dH2O overnight at 4 °C and stored in aliquots at −20 °C. The protein concentration of the dialysed samples was determined using a Bradford protein assay kit (Bio-Rad Laboratories, Hercules, CA).
Recombinant production of Par j 2 and Par j 1 in baculovirus-infected insect cells (BvPar j 2 and BvPar j 1)
Cloning and generating of high molecular weight recombinant bacmid DNA
Synthetic genes, codon-optimized for expression in insect cells coding for Par j 2.0101 (340 base pairs; accession number P55958) and for Par j 1.0102 (451 base pairs; accession number O04404) both including an additional 3′ sequence coding for a C-terminal hexahisitidine tag (Table 1), were subcloned into plasmid pTM1 via the BamHI/SmaI sites (ATG: biosynthetics, Merzhausen, Germany). The plasmids were transformed into DH10Bactm E. coli cells to generate high molecular weight recombinant bacmid DNA which was then used to transfect Sf9 insect cells to obtain virus stocks46.
Expression and purification of recombinant BvPar j 2, BvPar j 1 and EcPar j 2
For expression of the recombinant BvPar j 2 and BvPar j 1 proteins 150 μl/ml of viral stock was added to 20 ml of 1 × 106 cells/ml of SF9 cells in culture medium (Sf-900 II SFM, Gibco, Life technologies, Carlsbad, CA) supplemented only with gentamicin (10 μg/ml, Life technologies) but without FBS. The cells were then incubated under continuous shaking (120 rpm) for 48 hours at 27 °C. Cells and supernatant were then separated by centrifugation (1000 rpm, 10 min, 4 °C) and the supernatant was dialysed against buffer A used for purification by Ni2+-affinity chromatography (50 mM NaH2PO4, 300 mM NaCl, 10 mM Imidazole, pH 8.0) at 4 °C. Affinity purification of recombinant proteins was achieved by using Ni-agarose (Qiagen, Hilden, Germany)46.
Bacterially-expressed recombinant Par j 2 was expressed and purified as described and received from Prof. Paolo Colombo12.
The purity of the recombinant proteins was assessed by SDS-PAGE under non-reducing and reducing conditions with either ß-Mercaptoethanol or Tris(2-carboxyethyl)phosphine (TCEP) (see Supplementary Fig. S2). Aliquots of 2 μg of BvPar j 1, BvPar j 2 and EcPar j 2 were loaded on a 14% analytical gel47. Pre-stained protein ladder (PageRulerTMPlus, Thermo Scientific) was used as a molecular weight marker to determine the apparent masses of the proteins after staining of the gels with Coomassie blue.
Biochemical and biophysical characterization of purified BvPar j 1, BvPar j 2 and EcPar j 2
The determination of the mass of the purified proteins was performed by means of MALDI-ToF MS as previously described48. The fold of the proteins was analyzed by circular dichroism (CD) measurements as described49. Aggregation behaviour of purified proteins was analysed by gel filtration on an UltiMate™ 3000 HPLC (Dionex-Thermo Fisher Scientific, Vienna, Austria) using a BioSep-SEC-s3000 column (Phenomenex, Torrance, CA). Gel filtration of recombinant proteins was done in PBS pH 7.5. Cytochrome C (12.4 kDa) (Sigma-Aldrich, St. Louis, Mi) and carbonic anhydrase (29 kDa) (Sigma-Aldrich) were used for calibration under identical conditions and the molecular masses (MMs) of the elution peaks were calculated based on the elution time of the standard proteins49.
Sera from Paritaria judaica allergic patients
Sera from patients (n = 27; age 7 to 62 years; 19 females and 8 males) suffering from symptoms of pollinosis during the flowering period of Parietaria were obtained in the Associated Centers for Molecular Allergology, Rome, Italy. Clinical symptoms were recorded, IgE sensitization to Parietaria was confirmed by the measurement of Par j 2-specific IgE Abs (ImmunoCAP ISAC, Phadia, Uppsala, Sweden) and IgE sensitizations to other allergen sources were determined by ImmunoCAP ISAC measurements of IgE specific for micro-arrayed allergens (see Supplementary Table S1).
Serum samples were analyzed in an anonymized manner with approval of the ethics committee of the Medical University of Vienna, Austria (EK1641/2014). All experiments were performed in accordance with relevant guidelines and regulations.
IgE reactivity and allergenic activity of purified BvPar j 1, BvPar j 2 and EcPar j 2
The reactivity of recombinant allergens with patients IgE was tested by ELISA. Nickel pre-coated ELISA 96-well plates (Thermo Fisher Scientific, Pierce, Rockford, IL) with specific binding capacity for His-tagged proteins were coated with 2 μg/mL of EcPar j 2, 2 μg/mL of BvPar j 2 and, for control purposes, with 2 μg/mL BSA in carbonate buffer for 1 hour RT. In pilot experiments we tested different dilutions of sera to assure conditions of allergen excess. The plates were then washed 3 times with PBST (PBS + 0.05% Tween 20) and incubated overnight with sera (diluted 1:10 PBST + BSA) from allergic patients, serum from 3 non-allergic subjects. Bound IgE antibodies were detected with horseradish peroxidase–coupled goat anti-human IgE antibodies (KPL, Gaithersburg, MD) diluted 1:2500. The colour reaction was started by adding 1.7 mM 2.2′-azino-bis (3-ethylbenzothiazoline-6-sulfonic acid, Sigma-Aldrich), in 60 mM citric acid, 77 mM Na2HPO4.2H2O, and 3 mM H2O2. Optical density values (OD) were measured on ELISA reader (Spectra Max PLUS 384, Molecular Devices, Biberach, Germany) at different time points. All determinations were conducted as triplicates and results are displayed as means +/−SD. The buffer control was subtracted from the data and cut-off values of IgE reactivity were defined as mean OD plus three standard deviations of three sera from non-Parietaria-sensitized individuals.
To analyze the allergenic activity of the recombinant allergens rat basophil leukemia (RBL) cell-release assays were performed. Rat basophil leukemia (RBL) cells (clone RBL-703/21)50 transfected with human FcεRI (triplicates) were loaded with serum samples of nine Parietaria-allergic patients diluted 1:10 and incubated overnight at 37 °C. Degranulation of the cells was induced by adding BvPar j 1, BvPar j 2, EcPar j 2 (100, 10, 1, 0.1, 0.01 ng/mL) or buffer as a control. The release of β -hexosaminidase was measured and the results are shown as the percentage of total β-hexosaminidase release51.
Testing for IgE cross-reactivity by ELISA inhibition experiments
For IgE ELISA competition experiments, 96-well plates (Nunc Maxisorb, Roskilde, Denmark) were coated with BvPar j 1 or BvPar j 2 at a concentration 1 μg/mL or, in certain experiments with Parietaria judaica extract in a concentration of 5 μg/ml, and blocked as described for the IgE ELISA. Serum samples from Parietaria-allergic patients and from a non-allergic subject (1:20 dilution) were pre-incubated with purified recombinant allergens (BvPar j 2 or BvPar j 1) at a concentration of 2 μg/mL, with Parietaria judaica extract (10 μg/ml) or, for control purposes, with BSA (2 μg/ml). Plates were then incubated with pre-adsorbed sera overnight at 4 °C. and bound IgE was detected with horseradish peroxidase–coupled goat anti-human IgE antibodies (KPL, Gaithersburg, MD) diluted 1:2000 as described for the IgE ELISA. All determinations were conducted as duplicates with a variation of less than 5%. The percentages of inhibition obtained by pre-incubating sera with different allergens and allergen extract were calculated as follows: Percentage inhibition = 100 − (mean OD inhibited with allergen/mean OD non-inhibited) × 100. OD non-inhibited corresponds to pre-incubation with BSA as control protein. For certain experiments the percentage of inhibition obtained with another allergen is expressed as percentage of inhibition obtain with the very same allergen (i.e., auto-inhibition).
Inhibition of patients IgE binding with specific IgG antibodies obtained by immunizing rabbits with BvPar j 2
Par j 2-specific rabbit antibodies were obtained by immunizing a New Zealand white rabbit three times with the purified protein (200 μg, using once Freund’s complete and twice Freund’s incomplete adjuvant) by Charles River (Chatillon, sur Chalaronne, France). The ability of Par j 2-specific rabbit IgG to inhibit allergic patients IgE binding to Par j 1 and Par j 2 was determined by using an IgE competition ELISA52. The concentrations of BvPar j 1 and BvPar j 2 used for coating were 1 μg/mL in these experiments. The rabbit anti-Par j 2 antiserum was diluted 1:50 and sera from allergic patients were diluted 1:20 for IgE detection. All determinations were carried out in duplicates with a variation of less than 5%, results are expressed as means. Percentages of inhibition of IgE binding were calculated as follows: Percentage inhibition = 100 – (OD rabbit immune serum/OD rabbit pre-immune serum) × 100.
Statistical analysis and prediction tools
Statistical analysis was performed using GraphPad Prism 6 Software. Mann-Whitney U-test was used for comparison of differences between the groups. All calculated probability (p) values were two-tailed, and differences were considered as statistically significant if the p-value was lower than 0.05. For sequence alignment Clustal Omega program was used (https://www.ebi.ac.uk/Tools/msa/clustalo/). Sequences of LTPs with homology to Par j 2 were identified by comparing the amino acid sequences of Par j 2 and Par j 1 with the sequences deposited in the UniProt data base (www.uniprot.org/blast/) using the BLAST P program.
A hydrophobicity plot of Par j 1 and Par j 2 was generated using the ProtScale bioinformatics tool from the ExPASY server53 (http://web.expasy.org/protscale/) using the algorithm of Kyte and Doolittle54.
Phylogenetic comparisons of the amino acid sequence of Par j 2 with LTPs from different species were performed using the molecular evolutionary genetics analysis program (MEGA) version 7.0.2655,56. We used Neighbor-Joining algorithm with 100 replicate bootstraps57.
Data availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
References
Anto, J. M. et al. Mechanisms of the Development of Allergy (MeDALL): Introducing novel concepts in allergy phenotypes. J Allergy Clin Immunol. 139, 388–399 (2017).
Valenta, R. et al. Molecular Aspects of Allergens and Allergy. Adv Immunol. 138, 195–256 (2018).
Liccardi, G. et al. Allergy in adolescent population (14-18 years) living in Campania region (Southern Italy). A multicenter study. Eur Ann Allergy Clin Immunol. 51 2019.
Terzioğlu, E. et al. Sensitivity to Parietaria pollen in Izmir, Turkey as determined by skin prick and serum specific IgE values. J Investig Allergol Clin Immunol. 8, 180–3 (1998).
Cvitanović, S., Marusić, M., Zekan, L. & Köhler-Kubelka, N. Allergy induced by Parietaria officinalis pollen in southern Croatia. Allergy. 41, 543–5 (1986).
Holgate, S. T., Jackson, L., Watson, H. K. & Ganderton, M. A. Sensitivity to Parietaria pollen in the Southampton area as determined by skin-prick and RAST tests. Clin Allergy. 18, 549–56 (1988).
Kaufman, H. S. Parietaria: an unrecognized cause of respiratory allergy in the United States. Ann Allergy. 64, 293–6 (1990).
Ariano, R. et al. Parietaria pollination duration: myth or fact? Eur Ann Allergy Clin Immunol. 49, 6–10 (2017).
Gelardi, M. et al. Nasal inflammation in Parietaria-allergic patients is associated with pollen exposure. J Investig Allergol Clin Immunol. 24, 352–3 (2014).
Sala-Cunill, A. et al. Prevalence of asthma and severity of allergic rhinitis comparing 2 perennial allergens: house dust mites and Parietaria judaica pollen. J Investig Allergol Clin Immunol. (3), 145–5 (2013).
Panzani, R. Pollen asthma from Parietaria in France. Presse Med. 64, 908–10 (1956).
Duro, G. et al. cDNA cloning, sequence analysis and allergological characterization of Par j 2.0101, a new major allergen of the Parietaria judaica pollen. FEBS Lett 399, 295–8 (1996).
Duro, G. et al. Isolation and characterization of two cDNA clones coding for isoforms of the Parietaria judaica major allergen Par j 1.0101. Int Arch Allergy Immunol. 112, 348–55 (1997).
Colombo, P. et al. The allergens of Parietaria. Int Arch Allergy Immunol 130, 173–179 (2003).
González-Rioja, R., Asturias, J. A., Martínez, A., Goñi, F. M. & Viguera, A. R. Par j 1 and Par j 2, the two major allergens in Parietaria judaica, bind preferentially to monoacylated negative lipids. FEBS J. 276, 1762–75 (2009).
Stumvoll, S. et al. Identification of cross-reactive and genuine Parietaria judaica pollen allergens. J Allergy Clin Immunol 111, 974–79 (2003).
Amoresano, A. et al. Assignment of disulphide bridges in Par j 2.0101, a major allergen of Parietaria judaica pollen. Biol. Chem. 384, 1165–1172 (2003).
Bonura, A. et al. Isolation, expression and immunological characterization of a calcium-binding protein from Parietaria pollen. Mol Immunol. 57, 220–5 (2014).
Bonura, A. et al. Cloning, expression in E. coli and immunological characterization of Par j 3.0201, a Parietaria pollen profilin variant. Mol Immunol. 45, 2465–73 (2008).
Gonzalez-Rioja, R. et al. Diagnosis of Parietaria judaica pollen allergy using natural and recombinant Par j 1 and Par j 2 allergens. Clin Exp Allergy. 37, 243–250 (2007).
Valenta, R. et al. The recombinant allergen-based concept of component-resolved diagnostics and immunotherapy (CRD and CRIT). Clin Exp Allergy. 29, 896–904 (1999).
Valenta, R., Twaroch, T. & Swoboda, I. Component-resolved diagnosis to optimize allergen-specific immunotherapy in the mediterranean area. Investig Allergol Clin Immunol. 17, 88–92 (2007).
Matricardi, P. M. et al. EAACI Molecular Allergology User’s Guide. Pediatr Allergy Immunol. 27(Suppl 23), 1–250 (2016).
Larché, M., Akdis, C. A. & Valenta, R. Immunological mechanisms of allergen-specific immunotherapy. Nat Rev Immunol. 6, 761–71 (2006).
Valenta, R. et al. From allergen genes to allergy vaccines. Annu Rev Immunol 28, 211–41 (2010).
Curin, M. et al. Next-Generation of Allergen-Specific Immunotherapies: Molecular Approaches. Curr Allergy Asthma Rep. 18, 39 (2018).
Bonura, A. et al. Characterization of a Par j 1/Par j 2 mutant hybrid with reduced allergenicity for immunotherapy of Parietaria allergy. Clin Exp Allergy. 42, 471–80 (2012).
González-Rioja, R. et al. Genetically engineered hybrid proteins from Parietaria judaica pollen for allergen-specific immunotherapy. J Allergy Clin Immunol. 120, 602–9 (2007).
Bonura, A. et al. Modulating allergic response by engineering the major Parietaria allergens. J Allergy Clin Immunol. 141, 1142–1144 (2018).
Asturias, J. A., Gómez-Bayón, N., Eseverri, J. L. & Martínez, A. Par j 1 and Par j 2, the major allergens from Parietaria judaica pollen, have similar immunoglobulin E epitopes. Clin Exp Allergy. 33, 518–24 (2003).
Shamji, M. H. & Durham, S. R. Mechanisms of allergen immunotherapy for inhaled allergens and predictive biomarkers. J Allergy Clin Immunol. 140, 1485–1498 (2017).
Valenta, R. et al. Allergen Extracts for In Vivo Diagnosis and Treatment of Allergy: Is There a Future? Allergy Clin Immunol Pract. 6, 1845–1855 (2018).
Valenta, R., Campana, R. & Niederberger, V. Recombinant allergy vaccines based on allergen-derived B cell epitopes. Immunol Lett. 189, 19–2 (2017).
Valenta, R., Campana, R., Focke-Tejkl, M. & Niederberger, V. Vaccine development for allergen-specific immunotherapy based on recombinant allergens and synthetic allergen peptides: Lessons from the past and novel mechanisms of action for the future. J Allergy Clin Immunol. 137, 351–7 (2016).
Valenta, R. et al. Recombinant allergens: what does the future hold? J Allergy Clin Immunol. 127, 860–4 (2011).
Zafred, D., Nandy, A., Pump, L., Kahlert, H. & Keller, W. Crystal structure and immunologic characterization of the major grass pollen allergen Phl p 4. J Allergy Clin Immunol. 132, 696–703 (2013).
Ciprandi, G., Comite, P., Ferrero, F. & Mussap, M. A real life comparison between allergenic extracts and allergenic molecules. Iran J Allergy Asthma Immunol. 16, 39–44 (2017).
Purohit, A. et al. Poor association between allergen-specific serum immunoglobulin E levels, skin sensitivity and basophil degranulation: a study with recombinant birch pollen allergen Bet v 1 and an immunoglobulin E detection system measuring immunoglobulin E capable of binding to Fc epsilon RI. Clin Exp Allergy. 35, 186–92 (2005).
Shamji, M. H. et al. Biomarkers for monitoring clinical efficacy of allergen immunotherapy for allergic rhinoconjunctivitis and allergic asthma: an EAACI Position Paper. Allergy. 72, 1156–1173 (2017).
Pauli, G. et al. Efficacy of recombinant birch pollen vaccine for the treatment of birch-allergic rhinoconjunctivitis. J Allergy Clin Immunol. 122, 951–60 (2008).
Nony, E. et al. Development and evaluation of a sublingual tablet based on recombinant Bet v 1 in birch pollen-allergic patients. Allergy. 70, 795–804 (2015).
Jutel, M. et al. Allergen-specific immunotherapy with recombinant grass pollen allergens. J Allergy Clin Immunol. 116, 608–13 (2005).
Focke-Tejkl, M. et al. Development and characterization of a recombinant, hypoallergenic, peptide-based vaccine for grass pollen allergy. J Allergy Clin Immunol. 135, 1207–7 (2015).
Zieglmayer, P. et al. Mechanisms, safety and efficacy of a B cell epitope-based vaccine for immunotherapy of grass pollen allergy. EBioMedicine. 11, 43–57 (2016).
Niederberger, V. et al. Safety and efficacy of immunotherapy with the recombinant B-cell epitope-based grass pollen vaccine BM32. J Allergy Clin Immunol. 142, 497–509 (2018).
Pahr, S. et al. Biochemical, biophysical and IgE-epitope characterization of the wheat food allergen, Tri a 37. PLoS One. 4(9), 11 (2014).
Fling, S. P. & Gregerson, D. S. Peptide and protein molecular weight determination by electrophoresis using a high-molarity tris buffer system without urea. Anal Biochem. 155, 83–8 (1986).
Resch, Y. et al. Molecular characterization of Der p 10: a diagnostic marker for broad sensitization in house dust mite allergy. Clin Exp Allergy. 41, 1468–1477 (2011).
Campana, R. et al. Altered IgE epitope presentation: A model for hypoallergenic activity revealed for Bet v 1 trimer. Mol Immunol. 48, 431–41 (2011).
Nakamura, R. et al. A convenient and sensitive allergy test: IgE crosslinking‐induced luciferase expression in cultured mast cells. Allergy. 65, 1266–1273 (2010).
Gieras, A. et al. Molecular determinants of allergen‐induced effector cell degranulation. J Allergy Clin Immunol. 119, 384–390 (2007).
Focke, M. et al. Non-anaphylactic synthetic peptides derived from B cell epitopes of the major grass pollen allergen, Phl p 1, for allergy vaccination. FASEB J. 15, 2042–4 (2001).
Gasteiger, E. et al. Protein identification and analysis tools on the ExPASy server. Totowa, N.J. In The proteomics protocols handbook (ed. Walker, J. M.) 571–607 (Humana Press; 2005).
Kyte, J. & Doolittle, R. F. A simple method for displaying the hydropathic character of a protein. J Mol Biol. 157, 105–132 (1982).
Hall, B. G. Building phylogenetic trees from molecular data with MEGA. Mol Biol Evol. 30, 1229–35 (2013).
Kumar, S., Tamura, K. & Nei, M. MEGA: Molecular Evolutionary Genetics Analysis software for microcomputers. Comput Appl Biosci. 10, (189–91 (1994).
Saitou, N. & Nei, M. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol Biol Evol. 4, 406–425 (1987).
Acknowledgements
This work was funded by the Austrian Science Fund (FWF), projects DK W1248-B30, F4605, P29991 and the Medical University of Vienna. Rudolf Valenta is a recipient of a Megagrant of the Government of the Russian Federation, grant number 14.W03.31.0024. We thank Prof Ryosuke Nakamura for kindly providing the huRBL cell line.
Author information
Authors and Affiliations
Contributions
M.F. and R.V. designed, supervised and coordinated the study, interpreted the results and wrote the manuscript. Y.D. conducted experiments, analysed the data, prepared the figures, wrote the manuscript. P.C., M.B., A.M. and R.K. provided with experimental materials. All authors reviewed and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Dorofeeva, Y., Colombo, P., Blanca, M. et al. Expression and characterization of recombinant Par j 1 and Par j 2 resembling the allergenic epitopes of Parietaria judaica pollen. Sci Rep 9, 15043 (2019). https://doi.org/10.1038/s41598-019-50854-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-50854-1
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.