Abstract
Differentiating between inherited renal hypouricemia and transient hypouricemic status is challenging. Here, we aimed to describe the genetic background of hypouricemia patients using whole-exome sequencing (WES) and assess the feasibility for genetic diagnosis using two founder variants in primary screening. We selected all cases (N = 31) with extreme hypouricemia (<1.3 mg/dl) from a Korean urban cohort of 179,381 subjects without underlying conditions. WES and corresponding downstream analyses were performed for the discovery of rare causal variants for hypouricemia. Two known recessive variants within SLC22A12 (p.Trp258*, pArg90His) were identified in 24 out of 31 subjects (77.4%). In an independent cohort, we identified 50 individuals with hypouricemia and genotyped the p.Trp258* and p.Arg90His variants; 47 of the 50 (94%) hypouricemia cases were explained by only two mutations. Four novel coding variants in SLC22A12, p.Asn136Lys, p.Thr225Lys, p.Arg284Gln, and p.Glu429Lys, were additionally identified. In silico studies predict these as pathogenic variants. This is the first study to show the value of genetic diagnostic screening for hypouricemia in the clinical setting. Screening of just two ethnic-specific variants (p.Trp258* and p.Arg90His) identified 87.7% (71/81) of Korean patients with monogenic hypouricemia. Early genetic identification of constitutive hypouricemia may prevent acute kidney injury by avoidance of dehydration and excessive exercise.
Similar content being viewed by others
Introduction
Uric acid (UA) is the final product of purine metabolism in humans1. After reuptake in the renal proximal tubule, only 10% of initially filtered UA is eliminated in the urine2. Serum UA level is determined by the balance between the rate of purine metabolism and clearance. Serum UA level converges to a normal distribution in general population3. At present, the heritability of serum urate has been estimated in several studies to account for 25% to 60% of the variance in serum UA level4. Common variants within SLC2A9 and ABCG2 were reported to be highly associated with serum UA levels with an additional 28 genetic loci affecting serum urate level in a genome-wide association study (GWAS) of more than 140,000 individuals of European ancestry5.
Hypouricemia, defined as extremely low serum UA level, is a rare condition which can be affected by malnutrition, and by genetic defects in critical pathways involving UA synthesis and reabsorption system. Deficiencies of xanthine dehydrogenase (XDH), Molybdenum Cofactor Sulfurase (MOCOS), purine nucleoside phosphorylase (PNP), and 5-phosphoribosyl-pyrophosphate (PRPP) are related to the defects in UA synthesis6. Renal hypouricemia (RHUC), with a prevalence of 0.19% to 0.53% in several studies, is diagnosed based on laboratory criteria as 1) hypouricemia (<2 mg/dL) and 2) increased fractional excretion of UA (>10%)7. RHUC is asymptomatic and rarely identified unless an individual presents with severe renal symptoms including exercise-induced acute kidney injury (EIAKI), renal failure and nephrolithiasis8. Despite these important clinical implications, differentiating between inherited and transient hypouricemia is challenging because a low level of UA may reflect malnutrition status, which can be resolved by genetic screening using a panel with well-established genetic variants9.
Two types of RHUC have been currently reported: type 1 (OMIM: 220150) caused by mutations in SLC22A12 and type 2 (OMIM: 612076) caused by mutations in SLC2A9. A Japanese study first identified the protein-truncating p.Trp258* mutation in the SLC22A12 gene, which encodes a drug transporter in the renal proximal tubule10. Recently, coding variants in SLC22A12 and SLC2A9 causal for RHUC has been reported in various ethnic groups including Israeli-Arab, Iraqi-Jewish, and Roma populations in the Czech Republic and Slovakia7,11,12,13,14,15.
In this study, we investigated unrelated subjects with extremely low levels of UA using whole-exome sequencing (WES) to identify monogenic coding variants responsible for RHUC, which could be used for genetic screening of RHUC in Asians. After the discovery of candidate variants, we performed direct genotyping of the most frequent mutations (p.Trp258* and p.Arg90His) in SLC22A12 to replicate and quantify their contribution to RHUC in an independent Korean cohort, and to assess diagnostic feasibility of cost-effective genetic screening using these small subset of variants in hypouricemic patients.
Results
Hypouricemia prevalence and demographic information of 81 selected hypouricemic subjects
The prevalence of extreme hypouricemia (serum UA < 1.3 mg/dL) is 0.083% for the KoGES urban cohort (148/179,318). A total of 81 individuals (31 subjects in the KoGES urban cohort and 50 additional subjects from KCPS-II cohorts) were genetically tested for RHUC diagnostic assessment. Their baseline characteristics are summarized in Table 1. The 81 participants with RHUC (UA 0.74 ± 0.24 mg/dL; age, 47 ± 10 years; BMI, 23.3 ± 2.3 kg/m2; total cholesterol level, 189.2 ± 26.5 mg/dL) were healthy without chronic kidney disease, hypertension, diabetes mellitus or other metabolic diseases and without any history of smoking or malnutrition.
Identification of coding variants in SLC22A12 by whole-exome sequencing
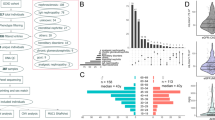
WES analysis was performed in 31 individuals with hypouricemia of the KoGES cohort (Fig. 1). The average depth coverage for these individuals was 85-fold. We performed variant calling and downstream filtering analyses assuming an autosomal recessive inheritance model. Coding variants in SLC22A12 were observed in 87.1% (27/31) of the individuals (Table 2). One subject was a compound heterozygote for SLC2A9 variants. In the remaining three individuals, variants within other genes were identified that appeared to have disease-causing potential and will be further investigated. 76% (24/31) individuals had variants previously reported in the Human Gene Mutation Database (HGMD) and the remaining 9.7% (3/31) exhibited novel missense mutations that had not been previously reported (Supplementary Table 3). 32.3% (10/31) of hypouricemia individuals were homozygous for the SLC22A12 p.Trp258* resulting in a premature stop codon, the most reported disease-causing variant to date. We also identified two homozygous individuals for the SLC22A12 p.Arg90His variants16. We found 12 compound heterozygous individuals for previously reported variants in the HGMD. In another individual, the p.Glu429Lys mutation was compound heterozygous with p.Trp258* in SLC22A12 (NIH17A8865148). Two novel SLC22A12 missense variants, p.Thr225Lys and p.Arg284Gln, were identified as compound heterozygotes (NIH17A8798528). Finally, we identified the novel p.Asn136Lys variant in the compound heterozygous state with the previously reported p.Leu418Arg variant (NIH17K4930892) (Supplementary Table 3).
The overall distribution of allele frequencies of the SLC22A12 variants within our study is shown in Fig. 2. The novel SLC22A12 variants were confirmed in the participant DNA samples by direct Sanger sequencing (Supplementary Fig. 1). Detailed properties of the four novel mutations in SLC22A12 are shown in Table 3 and Supplementary Table 3. This information was collected by querying several methods for functional prediction (Mutation Taster, Polyphen-2, SIFT, Condel). All four tools predicted the two SLC22A12 variants (p.Thr225Lys and p.Arg284Gln) reported in the NIH17A8798528 individual as deleterious. Amino acid sequence conservation was compared with R. macaque, M. musculus, C. lupus familiaris, and L. africana (Table 3). SLC22A12 p.Glu429Lys is not conserved in M. musculus and p.Asn136Lysd is not conserved in M. musculus, C. lupus familiaris, and L. africana.
Molecular dynamic prediction of SLC22A12 and novel variant location
The amino acid substitutions in SLC22A12 (10 variants) were considered for a molecular dynamic prediction analysis. The predicted functional impact of the amino acid change is illustrated in Supplementary Table 4. Our overall organization of the SLC22A12 protein was similar to the molecular dynamics approach described by Clemencon et al.17. Steered dynamic simulations of urate transport were performed with mutations in SLC22A12 and are presented in Fig. 3. Assessing the extent of the effect of the variants in the S set is difficult in a qualitative analysis due to the large changes observed during the molecular dynamics trajectory. p.Arg90His, p.Thr217Met, p.Thr225Lys, p.Trp258*, and p.Leu418Arg for SLC22A12 were predicted to alter protein structure defect. p.Arg284Gln and p.Arg477His were predicted to affect transport of uric acid. p.Asn136Lys and p.Gln382Leu for SLC22A12 were predicted to affect binding of urate. SLC22A12 p.Arg477His was predicted to both lower binding of urate and block the transportation pathway.
Utility of screening with two genetic SLC22A12 variants: c.774G > A (p.Trp258*) and c.269G > A (p.Arg90His)
Among 50 hypouricemia individuals from the KCPS-II replication cohort, 47 individuals carried at least one of these two genetic variants (Supplementary Table 2): 10 individuals carried the c.774G > A (p.Trp258*) homozygous stop codon; one individual carried a c.269G > A (p.Arg90His) homozygous mutation; 22 individuals carried c.269G > A (p.Arg90His) and c.774G > A (p.Trp258*) in the compound heterozygous state; and 14 individuals carried either c.269G > A (p.Arg90His) or c.774G > A (p.Trp258*) heterozygous variants.
Discussion
In this study, we comprehensively evaluated the contribution of SLC22A12 to severe hypouricemia through WES of 31 RHUC cases and replication of two implicated SNVs in 50 RHUC cases for a total of 81 unrelated Korean subjects. This is the first study to evaluate causal genetic variants for their diagnostic potential for RHUC. Overall, our study confirmed the importance of two mutations (p.Trp258* and p.Arg90His) in SLC22A12 for RHUC diagnosis found in 71/81(87.7%) of hypouricemia subjects.
Among the individuals exhibiting SLC22A12 mutations, we described four novel variants that had not been previously reported in the HGMD: p.Asn136Lys, p.Thr225Lys, p.Arg284Gln, and p.Glu429Lys. p.Asn136Lys (exon2) was located at the end of an intracellular loop, p.Thr225Lys (exon4) was present at the beginning of an extracellular loop, p.Arg284Gln (exon5) was localized in the largest extracellular loop, and p.Glu429Lys, in which the distal end of exon 7 and the first part of exon 8 are connected via splicing, was found to be within the membrane before an intracellular loop (Fig. 2B.)16. p.Asn136Lys occurred together with p.Leu418Arg in the case of NIH17K4930892; however, we could not determine cis or trans configuration. p.Thr225Lys: p.Arg284Gln and p.Glu429Lys:p.Trp258* were found in the compound heterozygous state, respectively in in NIH17A8798528 and NIH17A8865148. None of these variants were not found in Japanese (OMIM #220150, RHUC type 1)16,18,19,20. Further studies are needed to elucidate the pathogenicity of rare variants of unknown significance located within novel genes in six unexplained cases. Family-based WES studies for cases not explained by the two founder variants in SLC22A12 might identify additional monogenic genes that cause extremely low serum UA levels.
Hypouricemia is often regarded as an unrecognized or neglected disorder from a public health aspect21. The prevalence of renal stone due to excess of UA excretion is 6–7 times higher in patients with RHUC than in individuals with normal uric acid levels16. Evidence of oxidative stress has accumulated not only in EIAKI and renal stone but also in neurodegenerative disease (e.g., Parkinson’s disease) in persons with RHUC, reflecting the ability of UA to act as a powerful scavenger of approximately 60% of peroxide radicals in the plasma22,23,24,25,26. The anti-oxidative stress hypothesis is also supported by the results of Facheris et al., which show that the SLC2A9 mutation, associated with lower serum UA, increases the risk for early onset of neurodegenerative diseases27. Early identification and intervention of hypouricemia (avoidance of hard exercise, adequate hydration, and pre-emptively taking XO inhibitors) may prevent adverse events, especially among military personnel and athletics. XO inhibitor use (allopurinol or febuxostat) may be beneficial by lowering filtered UA. Screening of just two SLC22A12 variants (p.Trp258*/rs121907892 and p.Arg90His/rs121907896) for soldiers or athletics will provide early diagnosis of inherited RHUC and increase awareness among primary care physicians and medical care professionals (e.g. military, sport physicians, urologists) of the potential adverse health outcomes in at-risk individuals.
Here, we have shown that two Asian founder variants can provide a precision molecular diagnosis for 90% of inherited hypouricemia in the homogeneous Korean population. Recently, large scale WES have identified novel variants in SLC22A12 and SLC2A9 in individuals with European ancestry28. Like other genetic traits and conditions, RHUC shows genetic allelic and locus heterogeneity. Given that genetic architecture and causal variants, particularly rare variants, differ among ethnic and racial groups, collaborative genomic research may identify novel, population-specific variants associated with RHUC. Considering all of the population-specific rare variants observed in hypouricemia patients in Japanese, Roma, and African populations, a cosmopolitan screening panel may yield high diagnostic power even among heterogeneous populations that present with complex genetic admixture.
In summary, this study indicates the cost-effectiveness of screening for just two variants to diagnosis monogenic renal hypouricemia, and its potential utility in at-risk groups.
Materials and Methods
Study participants
This study was approved by the institutional review board of the Kangbuk Samsung Hospital (IRB# KBSMC 2016-12-016). We screened the subjects in the Korean genome and epidemiology study (KoGES) – KoGES health examinee study (urban cohort) and KoGES twin and family study. Out of 179,318 individuals, we selected 31 (M:11, F:20) individuals of hypouricemia (<1.3 mg/dL) who exhibited no other syndromic features or secondary causes (chronic kidney disease, hypertension, diabetes mellitus or any other metabolic diseases) and without any history of smoking. We also excluded people who have poor nutrition status. We obtained genomic DNA samples from the National Biobank of Korea29. In addition, 50 additional hypouricemic subjects without secondary causes were selected from the Korean Cancer Prevention Study (KCPS-II) cohort from the Severance Hospital, Seoul, Korea (IRB#4-2011-0277)30. Whole-exome sequencing (WES) was done in first 31 individuals, whereas SNaPshot genotyping of two variants (p.Trp258* and p.Arg90His) within SLC22A12 was performed to assess its screening purpose for second 50 subjects.
A total of 81 hypouricemic patients were therefore recruited for this study. All patients had given informed consent before they were enrolled in the study, which was conducted according to the Declaration of Helsinki. The overall flowchart for this study is presented in Fig. 1.
DNA preparation and whole-exome sequencing
Genomic DNA was obtained from peripheral blood leukocytes. We checked the quality of the DNA with an OD260/280 ratio of 1.8–2.0 by 1% agarose gel electrophoresis and PicoGreen® dsDNA Assay (Invitrogen, Waltham, MA, USA). SureSelect sequencing libraries were prepared (Agilent SureSelect All Exon kit 50 Mb, Santa Clara, CA, USA) and the enriched library was then sequenced using the HiSeq 2500 sequencing system (Illumina, San Diego, CA, USA). Image analysis and base calling were performed with the pipeline software using default parameters. Mapping was done using the human reference genome assembly (GRCh37/hg19), and all variants were called and annotated using CLC Genomic Workbench (version 9.0.1) software (QIAGEN bioinformatics, Redwood city, CA, USA).
WES variant filtering analysis
We performed variant-filtering analysis assuming an autosomal recessive or X-linked recessive pattern according to the predominantly observed inheritance mode in hereditary RHUC31. First, we systematically excluded variants with minor allele frequency (MAF) > 1%, which has been the conventional threshold for a rare variant, using dbSNP database (version 150), 1000 Genomes Projects phase 3 data (2,504 individuals), Exome Aggregation Consortium (ExAC, http://exac.broadinstitute.org), and Genome Aggregation Database (gnomAD, http://gnomad.broadinstitute.org/)29. Second, variants present in the homozygous or hemizygous state in in-house database consisting of 46 healthy Koreans without hypouricemia were excluded. Third, non-synonymous variants, small insertion/deletion (indel) or splice-site variants were selected. In the further analysis, we excluded single heterozygous variants so that only bi-allelic variants (homozygous, compound heterozygous, hemizygous for male) finally remained
Direct Sanger sequencing
Confirmation of called variants was conducted via direct Sanger sequencing. The DNA sequences spanning the variants were amplified using specific primers (Supplementary Table 1) and sequenced using an Applied Biosystems 3500xl genetic analyzer3500XL (Applied Biosystems, Foster City, CA, USA).
SNaPshot method
The SNaPshot assay of rs121907896 (p.Arg90His) and rs121907892 (p.Trp258*) was performed according to the manufacturer’s instructions (ABI PRISM SNaPshot Multiplex kit, Foster City, CA, USA). The analysis was carried out using GeneMapper software (version 4.0; Applied Biosystems). The primer sets for the SNaPshot assay are described in Supplementary Table 1.
In silico analysis of novel missense variants
Prior to the analysis, known pathogenic variants of SLC22A12 were screened in the Human Gene Mutation Database (HGMD®) as a public reference. For the newly discovered missense SLC22A12 variants, we checked if the mutated amino acid resides are highly conserved across the vertebrate orthologs using the UCSC Genome Browser (https://genome.ucsc.edu/). Given the role of the nitrogen excretion function in the evolutionary process, we identified amino acid sequences in several mammals (Rhesus macaque, Mus musculus, Canis lupus familiaris, and Loxodonta africana) that share the urea cycle rather than direct UA excretion. Third, the prediction of the functional effect of missense variants was performed using the latest version of PolyPhen-2, SIFT, Condel, and Mutation Taster algorithms32,33,34,35.
In silico prediction of molecular dynamics
We initially predicted the structure of SLC22A12 using a homology modeling program, SWISS-MODEL (https://swissmodel.expasy.org/). The quality of predicted 3D structures was estimated on the basis of the geometrical analysis of the single model, global model quality estimation (GMQE) score and qualitative model energy analysis (QMEAN)36. The GenBank accession number used for each amino acid sequence was NP_653186 for SLC22A12. After homology modeling was completed, we selected a suitable SLC2A3 X-ray structure for SLC22A12 (PDB ID: 4ZW9, SLC2A3)37,38. For the more stable molecular dynamics simulations, we used I-Tasser generated models39. All models were generated and made publicly available and can be recovered together with the statistics from the server site (https://zhanglab.ccmb.med.umich.edu/I-TASSER/about.html). All graphical representations were made using the initial I-Tasser generated models to aid reproducibility. A qualitative evaluation of the mutation effect was conducted based on four simple criteria. Binding urate (U) indicates the effect of the mutation on binding or urate because of the exposure of the mutated residue to the vestibular region or the urate binding motif cavity and/or involves a polar/nonpolar mutation affecting the interaction with urate. The structural effect (S) was evaluated as an increase in the root mean square displacement (RMSD) deviation computed during 25 ns of molecular dynamics (after 25 ns of equilibration) measured against the conformations obtained during a 25 ns trajectory for the initial sequence using either a solvated model or a Feedback Restrained Molecular Dynamics model (FRMD). FRMD affords a simple protocol to maximally retain structural features during a molecular dynamics trajectory while minimizing distortions imposed by an external restrain40. The transport effect (T) indicates that the mutation intrudes into the vestibular area blocking the possible passage of urate and is assigned based on a reduction of the internal cavity volume. We used all the models to identify geometries compatible with the mutation extending the initial molecular dynamic trajectory for SLC22A12 (10 mutations) to 125 ns. All molecular dynamics calculations were performed using NAMD241 and the ff99SB force field in the NVT ensemble with typical settings (T = 298 K, 2fs integration time, 12A cutoffs) obtained using QwikMD with default parameters to prepare the input files.
Data Availability
Before the official release, the data are available on reasonable request. The datasets generated during and/or analysed during the current study are available from the corresponding author on reasonable request. The data will be available at CODA (Clinical & Omics Data Archive, http://coda.nih.go.kr).
References
Wu, X. W., Muzny, D. M., Lee, C. C. & Caskey, C. T. Two independent mutational events in the loss of urate oxidase during hominoid evolution. J Mol Evol 34, 78–84 (1992).
Anzai, N., Kanai, Y. & Endou, H. New insights into renal transport of urate. Current Opinion in Rheumatology 19, 151–157 (2007).
Riches, P. L., Wright, A. F. & Ralston, S. H. Recent insights into the pathogenesis of hyperuricaemia and gout. Hum Mol Genet 18, R177–184, https://doi.org/10.1093/hmg/ddp369 (2009).
Major, T. J., Topless, R. K., Dalbeth, N. & Merriman, T. R. Evaluation of the diet wide contribution to serum urate levels: meta-analysis of population based cohorts. BMJ 363, k3951, https://doi.org/10.1136/bmj.k3951 (2018).
Kottgen, A. et al. Genome-wide association analyses identify 18 new loci associated with serum urate concentrations. Nat Genet 45, 145–154, https://doi.org/10.1038/ng.2500 (2013).
Iwahana, H. & Itakura, M. [Inherited disorders of uric acid metabolism–classification, enzymatic- and DNA-diagnosis]. Nihon rinsho. Japanese journal of clinical medicine 54, 3303–3308 (1996).
Sebesta, I. et al. Diagnostic tests for primary renal hypouricemia. Nucleosides Nucleotides Nucleic Acids 30, 1112–1116 (2011).
Cheong, H. I. et al. Mutational analysis of idiopathic renal hypouricemia in Korea. Pediatr. Nephrol. 20, 886–890, https://doi.org/10.1007/s00467-005-1863-3 (2005).
Tseng, C. K. et al. In addition to malnutrition and renal function impairment, anemia is associated with hyponatremia in the elderly. Arch. Gerontol. Geriatr. 55, 77–81, https://doi.org/10.1016/j.archger.2011.06.019 (2012).
Enomoto, A. et al. Molecular identification of a renal urate anion exchanger that regulates blood urate levels. Nature 417, 447–452 (2002).
Stiburkova, B., Taylor, J., Marinaki, A. M. & Sebesta, I. Acute kidney injury in two children caused by renal hypouricaemia type 2. Pediatr. Nephrol. 27, 1411–1415 (2012).
Dinour, D. et al. URAT1 mutations cause renal hypouricemia type 1 in Iraqi Jews. Nephrol. Dial. Transplant. 26, 2175–2181 (2011).
Tasic, V. et al. Clinical and functional characterization of URAT1 variants. PLoS One 6, e28641–e28641 (2011).
Stiburkova, B. et al. Novel allelic variants and evidence for a prevalent mutation in URAT1 causing renal hypouricemia: biochemical, genetics and functional analysis. Eur. J. Hum. Genet. 21, 1067–1073 (2013).
Bhasin, B. et al. Hereditary renal hypouricemia: a new role for allopurinol? Am. J. Med. 127, e3–e4 (2014).
Ichida, K. et al. Clinical and molecular analysis of patients with renal hypouricemia in Japan-influence of URAT1 gene on urinary urate excretion. Journal of the American Society of Nephrology 15, 164–173 (2004).
Clémençon, B. et al. Expression, Purification, and Structural Insights for the Human Uric Acid Transporter, GLUT9, Using the Xenopus laevis Oocytes System. Plos One 9, e108852, https://doi.org/10.1371/journal.pone.0108852 (2014).
Iwai, N. et al. A high prevalence of renal hypouricemia caused by inactive SLC22A12 in Japanese. Kidney Int. 66, 935–944 (2004).
Ichida, K. et al. Age and origin of the G774A mutation in SLC22A12 causing renal hypouricemia in Japanese. Clin. Genet. 74, 243–251 (2008).
Taniguchi, A. et al. A common mutation in an organic anion transporter gene, SLC22A12, is a suppressing factor for the development of gout. Arthritis & rheumatism 52, 2576–2577 (2005).
Kuwabara, M. et al. Prevalence and complications of hypouricemia in a general population: A large-scale cross-sectional study in Japan. Plos One 12, e0176055, https://doi.org/10.1371/journal.pone.0176055 (2017).
Kawachi, M. et al. Decreased renal clearance of xanthine and hypoxanthine in a patient with renal hypouricemia: a new defect in renal handling of purines. Nephron 61, 428–431, https://doi.org/10.1159/000186961 (1992).
Windpessl, M., Ritelli, M., Wallner, M. & Colombi, M. A Novel Homozygous SLC2A9 Mutation Associated with Renal-Induced Hypouricemia. Am. J. Nephrol. 43, 245–250, https://doi.org/10.1159/000445845 (2016).
Okabayashi, Y. et al. Rare case of nephrocalcinosis in the distal tubules caused by hereditary renal hypouricaemia 3 months after kidney transplantation. Nephrology (Carlton) 21(Suppl 1), 67–71, https://doi.org/10.1111/nep.12774 (2016).
Sugihara, S. et al. Depletion of Uric Acid Due to SLC22A12 (URAT1) Loss-of-Function Mutation Causes Endothelial Dysfunction in Hypouricemia. Circ. J. 79, 1125–1132, https://doi.org/10.1253/circj.CJ-14-1267 (2015).
Mou, L. J., Jiang, L. P. & Hu, Y. A novel homozygous GLUT9 mutation cause recurrent exercise-induced acute renal failure and posterior reversible encephalopathy syndrome. J. Nephrol. 28, 387–392, https://doi.org/10.1007/s40620-014-0073-0 (2015).
Facheris, M. F. et al. Variation in the uric acid transporter gene SLC2A9 and its association with AAO of Parkinson’s disease. J. Mol. Neurosci. 43, 246–250, https://doi.org/10.1007/s12031-010-9409-y (2011).
Tin, A. et al. Large-scale whole-exome sequencing association studies identify rare functional variants influencing serum urate levels. Nature Communications 9, 4228, https://doi.org/10.1038/s41467-018-06620-4 (2018).
Kim, Y. & Han, B. G. Cohort Profile: The Korean Genome and Epidemiology Study (KoGES) Consortium. Int. J. Epidemiol. 46, 1350, https://doi.org/10.1093/ije/dyx105 (2017).
Cho, S. K., Kim, S., Chung, J. & Jee, S. H. Discovery of URAT1 SNPs and association between serum uric acid levels and URAT1. BMJ open 5, e009360–e009360 (2015).
Sperling, O. Hereditary renal hypouricemia. Mol. Genet. Metab. 89, 14–18, https://doi.org/10.1016/j.ymgme.2006.03.015 (2006).
Adzhubei, I. A. et al. A method and server for predicting damaging missense mutations. Nat Methods 7, 248–249, https://doi.org/10.1038/nmeth0410-248 (2010).
Kumar, P., Henikoff, S. & Ng, P. C. Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat Protoc 4, 1073–1081, https://doi.org/10.1038/nprot.2009.86 (2009).
Gonzalez-Perez, A. & Lopez-Bigas, N. Improving the assessment of the outcome of nonsynonymous SNVs with a consensus deleteriousness score, Condel. Am J Hum Genet 88, 440–449, https://doi.org/10.1016/j.ajhg.2011.03.004 (2011).
Schwarz, J. M., Cooper, D. N., Schuelke, M. & Seelow, D. MutationTaster2: mutation prediction for the deep-sequencing age. Nat Methods 11, 361–362, https://doi.org/10.1038/nmeth.2890 (2014).
Benkert, P., Biasini, M. & Schwede, T. Toward the estimation of the absolute quality of individual protein structure models. Bioinformatics 27, 343–350, https://doi.org/10.1093/bioinformatics/btq662 (2011).
Deng, D. et al. Molecular basis of ligand recognition and transport by glucose transporters. Nature 526, 391–396 (2015).
Nomura, N. et al. Structure and mechanism of the mammalian fructose transporter GLUT5. Nature 526, 397–401 (2015).
Yang, J. et al. The I-TASSER Suite: protein structure and function prediction. Nat. Methods 12, 7–8 (2015).
Cachau, R. E., Erickson, J. W. & Villar, H. O. Novel procedure for structure refinement in homology modeling and its application to the human class Mu glutathione S-transferases. Protein Eng. 7, 831–839 (1994).
Phillips, J. C. et al. Scalable molecular dynamics with NAMD. J Comput Chem 26, 1781–1802 (2005).
Acknowledgements
The bioresources for this study were provided by the National Biobank of Korea, Centers for Disease Control and Prevention, Republic of Korea. This research was supported by the Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education (NRF-2016R1A6A3A11933380 to S.K.C and 2015R1D1A1A01056685 to H.Y.G) and the Korea Health Technology R&D Project through the Korea Health Industry Development Institute (KHIDI), funded by the Ministry of Health & Welfare, Republic of Korea (Grant Number: HI17C2372 to S.K.C). We thank the Research Division, NEXBiO Co. Ltd for their contribution to our sequencing. We are grateful to Young Sup Cho MD, PhD and Hyekyung Son MD, PhD for their academic advice during this project. This Research was supported partly by the Intramural Research Program of the NIH, National Cancer Institute, Center for Cancer Research. This project has been funded in part with federal funds from the National Cancer Institute, National Institutes of Health, under contract HHSN26120080001E. The content of this publication does not necessarily reflect the views or policies of the Department of Health and Human Services, nor does the mention of trade names, commercial products, or organizations imply endorsement by the U.S. Government.
Author information
Authors and Affiliations
Contributions
Sung Kweon Cho, Do Hyeon Cha, Heon Yung Gee, Jong Mun Choi and Sun Ha Jee planned and designed the study and directed its implementation, including quality assurance and control. Sung Kweon Cho, Do Hyeon Cha, Jong Mun Choi, Daeui Park and Raul Cachau analyzed the data and designed the study’s analytic strategy. Sung Kweon Cho, Cheryl A Winkler, Kyeong Kyu Kim and Seungho Ryu helped supervise field activities. Sung Kweon Cho, Seungho Ryu, Cheryl A Winkler, Kyeong Kyu Kim, Hong-Hee Won, Sophie Limou and Woojae Myung helped conduct the literature review and prepare the Materials and Methods and the Discussion sections of the text. Sung Kweon Cho and Do Hyeon Cha interpreted the results. Sung Kweon Cho and Do Hyeon Cha drafted the manuscript. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cha, D.H., Gee, H.Y., Cachau, R. et al. Contribution of SLC22A12 on hypouricemia and its clinical significance for screening purposes. Sci Rep 9, 14360 (2019). https://doi.org/10.1038/s41598-019-50798-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-50798-6
This article is cited by
-
Genotype and Phenotype of Renal Hypouricemia: A Single-Center Study from China
Molecular Diagnosis & Therapy (2024)
-
Causal and putative pathogenic mutations identified in 39% of children with primary steroid-resistant nephrotic syndrome in South Africa
European Journal of Pediatrics (2022)
-
Rational selection of a biomarker panel targeting unmet clinical needs in kidney injury
Clinical Proteomics (2021)
-
Polygenic analysis of the effect of common and low-frequency genetic variants on serum uric acid levels in Korean individuals
Scientific Reports (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.