Abstract
Although the modulation of immune-related genes after viral infection has been widely described in vertebrates, the potential implications of non-coding RNAs (ncRNAs), especially long non-coding RNAs (lncRNAs), in immunity are still a nascent research field. The model species zebrafish could serve as a useful organism for studying the functionality of lncRNAs due to the numerous advantages of this teleost, including the existence of numerous mutant lines. In this work, we conducted a whole-transcriptome analysis of wild-type (WT) and heterozygous rag1 mutant (rag1+/−) zebrafish after infection with the pathogen spring viraemia of carp virus (SVCV). WT and rag1+/− zebrafish were infected with SVCV for 24 h. Kidney samples were sampled from infected and uninfected fish for transcriptome sequencing. From a total of 198,540 contigs, 12,165 putative lncRNAs were identified in zebrafish. Most of the putative lncRNAs were shared by the two zebrafish lines. However, by comparing the lncRNA profiles induced after SVCV infection in WT and rag1+/− fish, most of the lncRNAs that were significantly induced after viral challenge were exclusive to each line, reflecting a highly differential response to the virus. Analysis of the neighboring genes of lncRNAs that were exclusively modulated in WT revealed high representation of metabolism-related terms, whereas those from rag1+/− fish showed enrichment in terms related to the adaptive immune response, among others. On the other hand, genes involved in numerous antiviral processes surrounded commonly modulated lncRNAs, as expected. These results clearly indicate that after SVCV infection in zebrafish, the expression of an array of lncRNAs with functions in different aspects of immunity is induced.
Similar content being viewed by others
Introduction
The increasing genomic data available for model and non-model organisms has revealed that only a small percentage of the genome corresponds to protein-coding genes. A high proportion of the genome does not present coding potential but is transcribed into non-coding RNAs (ncRNAs). Among ncRNAs, the long non-coding RNAs (lncRNAs) present a length of over 200 nucleotides (nt) and are transcribed in the same way as mRNAs. LncRNAs regulate the expression of adjacent or distal genes through different mechanisms, such as transcription interference, initiation of chromatin remodeling, promoter inactivation, activation of accessory proteins, and activation of transcription factors1. Because most lncRNAs influence the expression of their neighbor genes by acting as local regulators, lncRNA expression is often positively or inversely correlated with the expression of their adjacent coding-genes1. Notably, lncRNAs play a role in different molecular processes and exhibit specific-transcription patterns in different cell types2, where these transcripts can promote or repress the gene expression3. Most likely due to the functional complexity of lncRNAs, studies concerning their activity remain scarce.
For understanding the role of lncRNAs, one of most useful species is zebrafish (Danio rerio). Zebrafish is a model fish species for genomic, developmental, biomedical and pharmacological studies, among other areas of inquiry. Studies on lncRNAs in zebrafish have demonstrated their roles in processes such as tissue and organ repair3, development and nervous system function4, cold acclimation5, the response to antibiotics6, and immune system function7.
In the case of the fish immunity, lncRNAs are key players that in turn can display complex mechanism to modulate the expression of Toll-like receptors, NF-kB activity, differentiation of the Th1/Th2 response, and other immune-relevant functions2. For instance, in mammals, the lncRNA Morrbid, HOTAIRM1 or lnc-DC mediates the development and differentiation of dendritic and myeloid cells8. Nevertheless, studies focused on the activity of lncRNAs in the teleost immune system are still scarce. In zebrafish, knockdown analysis demonstrated the role of the PU.1 gene in adaptive immunity and the negative regulation of this gene by its antisense long non-coding RNA (AS lncRNA)7. Furthermore, different expression patterns of lncRNAs have been described in response to a variety of pathogens in salmonids9,10,11,12, and also the modulation of the lncRNA patterns by HSP70 and Streptococcus agalactiae antigen stimulation in tilapia (Oreochromis niloticus) has been recently analyzed13.
Moreover, an excellent model for understanding the role of lncRNAs in the immune response and their modulation after infections is the zebrafish mutant line rag1. V(D)J recombination, carried out by the combined endonuclease activity of recombination activating gene 1 (RAG1) and RAG2, assembles the vast diversity of immunoglobulins and T cell receptor (TCR) genes14. A point mutation of the rag1 gene in zebrafish causes a premature stop codon in the rag1 catalytic domain; therefore, this mutation presumably abolishes the adaptive immune system15. Herein, SVCV is an enveloped, negative-sense, single-stranded RNA virus belonging to the Rhabdoviridae family, and this pathogen mainly affects cyprinids, including zebrafish16,17. Although numerous investigations concerning the immune system have been developed in zebrafish using SVCV18,19,20,21, to our knowledge, this is the first time that the impact of this virus on the lncRNA profile has been analyzed.
Comparison of the lncRNA expression pattern after SVCV challenge in wild-type (WT) and rag1-heterozygous mutant fish (rag1+/−) would allow us to elucidate the potential implications of the diversity of lncRNAs in different aspects of innate and adaptive immunity. Because heterozygous rag1 mutants are partially deficient in the generation of mature lymphocytes, the potential compensation mechanisms induced after SVCV challenge could reveal specific lncRNAs related to acquired immunity. This study gives new genomic knowledge of how lncRNAs are key molecular components of the immune system in teleost.
Methods
Zebrafish and virus
Six-month-old WT and rag1+/− zebrafish were obtained from the facilities of the Instituto de Investigaciones Marinas (Vigo, Spain), where zebrafish are maintained following established protocols22,23. Zebrafish were euthanized using a tricaine methanesulfonate (MS-222) overdose (500 mg/l). Fish care and challenge experiments were conducted according to the guidelines of the CSIC National Committee on Bioethics under approval number ES360570202001/16/FUN01/PAT.05/tipoE/BNG.
Spring viraemia of carp virus (SVCV) isolate 56/70 was propagated on epithelioma papulosum cyprini (EPC) carp cells (ATCC CRL-2872) containing MEM (Gibco) supplemented with 2% FBS (Gibco) and 100 µg/ml Primocin (InvivoGen) and was titrated in 96-well plates. The TCID50/ml was calculated according to the Reed and Muench method24.
Experimental design and samples
Twelve zebrafish from each line (WT and rag1+/−) were intraperitoneally (i.p.) injected with 20 μl of SVCV (3 × 102 TCID50/ml), and as control groups, the same number of fish were injected with an equal volume of MEM (Gibco, USA) supplemented with 2% FBS (Gibco) and 100 µg/ml Primocin (InvivoGen, USA). Kidney samples were collected at 24 h post-injection (hpi) and stored at −80 °C until RNA extraction.
In parallel, the effect of the virus on the survival of heterozygous mutants was evaluated. For this purpose, 20 individuals from each line (WT and rag1+/−) were also i.p. infected with the virus, and another 20 fish were inoculated with the medium. The 20 zebrafish from each group were divided into 2 batches of 10 fish/batch, which served as biological replicates for the survival analysis. Mortality was recorded over a period of 3 weeks. Kaplan-Meier survival curves were analyzed with a log-Rank (Mantel-Cox) test.
High-throughput transcriptome sequencing
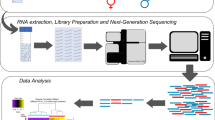
Kidney samples from infected and control WT and rag1+/− zebrafish were used for Illumina cDNA library preparations. Briefly, total RNA was extracted from each individual using the Maxwell® 16 LEV simplyRNA Tissue kit (Promega) with an automated Maxwell 16 Instrument following the manufacturer’s instructions. The quantity and purity of the total RNA was evaluated by using a Nanodrop ND-1000. RNA integrity was analyzed using the Bioanalyzer 2100 system (Agilent Technologies Inc., USA) according to the manufacturer’s instructions. Samples with a RIN over 8.0 were pooled and used for library preparation. Subsequently, using the RNA pools from the control and challenged groups (3 biological replicates; 4 fish/replicate), double-stranded cDNA libraries were constructed using the TruSeq RNA Sample Preparation Kit v2 (Illumina®, San Diego, CA, USA). Samples were sequenced by using the HiSeq (Illumina®) platform. The raw read sequences were deposited in the Sequence Read Archive (SRA) (https://www.ncbi.nlm.nih.gov/sra) under accession number PRJNA532380.
Sequence assembly and lncRNA identification
Sequence assembly was carried out using CLC Genomics Workbench v10.1 software (CLC Bio, Aarhus, Denmark). De novo assembly was performed using datasets from the zebrafish group. Assembly was conducted with an overlap criterion of 70% and a similarity of 0.9 to exclude paralogous sequence variants (PSVs)25. The settings were as follows: a mismatch cost of 2, deletion cost of 3, insert cost of 3, minimum contig length of 200 base pairs (bp) and trimming quality score of 0.05. After the assembly process, singletons were retained in the dataset as possible representatives of low-expression transcript fragments. However, the sequence redundancy of these fragments was removed by using the Duplicate Finder application incorporated in Geneious v8.0 software (Biomatters, Auckland, New Zealand). The assembled data were processed using CLC Genomics Workbench software following the previously described pipeline10,11. Briefly, following de novo assembly of the WT and rag1+/− zebrafish transcriptomes, several filters were applied to the consensus sequences of contigs. Sequences with an average coverage <50 were discarded. Then, BlastX analysis was performed for selected sequences, discarding all sequences with positive Blast hits (BlastX, e-value < 1e-05) against the proteins for all the bony fish species included in the NCBI GenBank and UniProt databases. Open reading frames (ORFs) were predicted from the remaining contigs, and all contigs with putative ORFs > 200 bp were discarded. Finally, the free Coding Potential Assessment Tool (CPAT) (http://lilab.research.bcm.edu/cpat/) was used to discard sequences with coding potential.
Genome mapping and pathway analysis
To gain a better understanding of the genomic context of lncRNAs in WT and rag+/− zebrafish, non-coding transcripts were mapped against the last version of the zebrafish genome (Assembly: GCA_000002035.4 GRCz11). Thus, lncRNAs were mapped using the CLC Genomics Workbench mapping algorithm and considering the following parameters: length fraction = 0.8, similarity fraction = 0.8 and mismatch, insertion and deletion costs of 2, 3 and 3, respectively. In addition, coding genes flanking up to 10,000 bases upstream and downstream from the annotated lncRNAs were identified and extracted for GO and KEGG analysis. Thus, GO enrichment analysis26 was conducted to identify the most represented biological process, cellular component and molecular function categories among the protein coding genes proximal to lncRNAs. The results were finally plotted using the REVIGO platform27, R and CYTOSCAPE software28.
RNA-Seq analysis
The RNA-Seq settings included a minimum length fraction = 0.6 and a minimum similarity fraction (long reads) = 0.5. The expression values were set to the transcripts per million model (TPM). The distance metric was calculated using the Manhattan method, with the mean expression level in 5–6 rounds of k-means clustering subtracted. Finally, Kal’s statistical analysis test was used to compare gene expression levels in terms of the log2 fold-change (P = 0.0005; FDR corrected).
Quantitative PCR (qPCR) validation of lncRNA expression
cDNA synthesis from the samples was conducted with the NZY First-Strand cDNA Synthesis kit (NZYTech) using 0.5 μg of total RNA. A total of 12 lncRNAs were used to validate the RNA-Seq results. Specific qPCR primers for the selected lncRNAs were designed using Primer 3 software29, and their amplification efficiency was calculated with the threshold cycle (CT) slope method30. The primer sequences used for qPCR are listed in Supplementary Table S1. Individual qPCR assays were carried out in a 25 μl reaction volume containing 12.5 μl of SYBR GREEN PCR Master Mix (Applied Biosystems), 10.5 μl of ultrapure water, 0.5 μl of each specific primer (10 μM) and 1 μl of two-fold-diluted cDNA template in MicroAmp optical 96-well reaction plates (Applied Biosystems, USA). Reactions were conducted using technical triplicates in a 7300 Real-Time PCR System thermocycler (Applied Biosystems). The qPCR conditions were as follows: initial denaturation step (95 °C, 10 min), 40 cycles of a denaturation step (95 °C, 15 s) and a hybridization-elongation step (60 °C, 1 min). The relative expression level of the lncRNAs was normalized following the Pfaffl method30 and using the 18S ribosomal RNA (18 s) as a reference gene.
Results
Survival rate of WT and rag1+/− zebrafish after SVCV challenge
In contrast to what is observed in homozygous rag1 mutant zebrafish, which are more resistant to infection with SVCV compared to WT fish31,32, no significant differences in survival were found between the WT and rag1-heterozygous fish after SVCV challenge (Supplementary Fig. S1).
Sequence assembly and lncRNA identification
A total of 543,596,316 and 651,786,032 reads were obtained for WT and rag1+/− zebrafish, respectively. From the total assembly, 198,540 contigs were obtained from WT and rag1+/− zebrafish samples, among which 58,805 contigs represented coding sequences, and 12,165 were putative lncRNA sequences, with average lengths of 832 nt and 545 nt, respectively. The lncRNAs identified in zebrafish presented a length distribution ranging from 200 to 2,000 nt, whereas for the coding sequences, this range was 200 to 5,500 nt (Fig. 1). Moreover, 11,774 and 11,196 lncRNAs were identified from each dataset for WT and rag1+/− zebrafish, respectively, with the two strains sharing 10,859 lncRNAs (Fig. 2A) with similar expression profiles (Fig. 2B). Additionally, the locations of the lncRNAs identified for WT and rag1+/− zebrafish in the zebrafish genome were determined. Although few differences in lncRNA localization were observed for the two zebrafish lines, rag1+/− zebrafish did present some lncRNAs in different positions than the WT fish (Fig. 3).
LncRNA abundance and chromosome localization in WT and rag1+/− zebrafish. LncRNAs were mapped in the zebrafish genome to identify potential differences in the chromosome location between WT and rag1+/− zebrafish. In general terms, lncRNAs from both lines showed a similar chromosome distribution and abundance per chromosome. Nevertheless, some subtle differences were observed, revealing the specific position of WT and rag1+/− exclusive lncRNAs.
The neighboring genes of the exclusive lncRNAs of WT and rag1+/− zebrafish were annotated by GO analysis, showing different processes potentially modulated by these exclusive lncRNAs (Fig. 4). The biological processes that were putatively regulated by exclusive lncRNAs of WT zebrafish were mainly associated with different metabolic processes; for the cellular components, the greatest number of lncRNA-neighboring genes were associated with cytoplasm; and catalytic activity was the molecular function with the most 7-annotated genes (Fig. 4). In contrast, the exclusive lncRNAs of rag1+/− zebrafish were located near genes annotated to biological processes related to the cellular response to stimulus, cell communication, transport and immune response categories, among other processes. Among cellular components, intracellular and organelle components were the most abundant components annotated, and in the molecular function category, the most abundant neighboring genes were annotated as being associated with substrate-specific transporter activity and transmembrane transporter activity (Fig. 4).
GO analysis of exclusive lncRNA-neighboring genes identified in WT and rag1+/− zebrafish. Those lncRNAs exclusively detected in WT or rag1+/− fish were positioned on the genome and the protein-coding genes surrounding the lncRNAs were assigned to biological process, cellular component and molecular function categories.
Neighboring lncRNAs to rag1 and rag2 genes chromosomal localization and modulation
Five lncRNAs identified in WT and rag1+/− zebrafish showed a physical linkage with rag1 and rag2 genes on chromosome 25 (Fig. 5A). Expression analysis of these lncRNAs using the TPM values of the samples revealed that two of them were differentially expressed between WT and rag1+/− zebrafish (Fig. 5B). The Lnc_Contig0016722 and Lnc_Contig0057120 were up-regulated in rag1+/− compared to the WT zebrafish. The results suggest a putative regulatory role of Lnc_Contig0016722 and Lnc_Contig0057120 in the expression of rag1 and rag2 genes in rag1+/− zebrafish.
LncRNA modulation during SVCV infection
RNA-Seq analysis of coding and non-coding transcripts in the zebrafish samples revealed two differentiated clusters of samples, one for control and SVCV-infected WT zebrafish and another cluster for both conditions in rag1+/− zebrafish, reflecting greater relevance of the zebrafish line than the infection (Fig. 6A,B). The heat map representation of protein-coding transcripts exhibited four clusters of contigs with a similar expression profile (Fig. 6A). Cluster 1 was highly expressed in both control and infected WT samples; cluster 2 showed greater representation in control and infected rag1+/− zebrafish and infected WT; cluster 3 was mainly expressed in control and infected rag1+/− zebrafish; and cluster 4 was mainly expressed in control WT fish. The protein-coding contigs included in each cluster are represented in Supplementary Table S2. Moreover, four main lncRNA clusters were identified from the heat map representation with similar expression profiles to those observed for the coding transcripts. The first (cluster 1) included those lncRNAs that were downregulated in rag1+/− control zebrafish (Fig. 6B), suggesting a role of these lncRNAs in the response to SVCV infection in rag1+/− zebrafish. On the other hand, cluster 2 was mainly up-regulated in kidney samples from control and infected WT zebrafish (Fig. 6B). Cluster 3 was highly regulated in control rag1+/− zebrafish and infected WT individuals. Finally, cluster 4 was more highly expressed in control and infected rag1+/− zebrafish (Fig. 6B). The lncRNAs included in the different clusters are represented in Supplementary Table S2.
Differentially expressed transcripts in response to SVCV infection. (A,B) Hierarchical clustering of expressed transcripts (TPM values) in kidney samples from different experimental conditions. (A) Coding transcripts. (B) LncRNAs. (C,D) Venn diagrams representing the significantly expressed, (C) coding transcripts and (D) lncRNAs (fold-change 4, p-value 0.01).
In addition, the differential expression analysis (absolute fold-change of 4 and p-value of 0.001) between SVCV-challenged and control WT and rag1+/− zebrafish showed a greater number of protein-coding and lncRNA transcripts that were modulated during infection in WT zebrafish (6,860 and 709, respectively) than in rag1+/− zebrafish (5,052 and 366). Interestingly, Venn diagrams showed that the majority of these transcripts were exclusively modulated after infection in each line, with 5,461 coding RNAs and 593 lncRNAs being exclusively modulated in WT and 3,653 coding RNAs and 250 lncRNAs being exclusively modulated in rag1+/− zebrafish (Fig. 6C,D). Moreover, a total of 116 common coding-genes and 1,399 common lncRNAs were identified between WT and rag1+/− zebrafish. Supplementary Table S3 shows the most modulated common and exclusive lncRNAs in WT and rag1+/− after infection, reflecting that some commonly modulated lncRNAs were regulated in opposite ways.
Enrichment of the neighboring genes of lncRNAs expressed during infection
Exclusive lncRNAs expressed after infection in WT and rag1+/− zebrafish were located in the genome to identify neighboring protein-coding genes located within 10 kb upstream and downstream from the lncRNAs (Supplementary Table S4). For the WT zebrafish-exclusive lncRNAs, GO enrichment of neighboring genes revealed that the most represented biological processes were different metabolic processes, growth, and oxidation-reduction process, among others (Fig. 7A). Interestingly, for the rag1+/− zebrafish-exclusive lncRNA-neighboring genes, several biological processes related to immunity, especially to adaptive immunity, were observed (immune system process, T cell-mediated immunity, regulation of adaptive immune response/built from immunoglobulin, regulation of adaptive immune response, cytokine production involved in immune response) (Fig. 7B). Similar results were observed for the GO enrichment analysis of exclusive coding RNAs expressed differentially during infection. A greater number of rag1+/− exclusive coding RNAs were associated with the immune response process (Supplementary Fig. S2).
GO enrichment analysis for exclusive lncRNA-neighboring genes annotated in WT and rag1+/− zebrafish that were highly regulated during viral infection. Although most of the lncRNAs modulated after an SVCV challenge were common to both zebrafish lines, some lncRNAs were exclusively regulated in WT and rag1+/− fish. The protein-coding genes located within 10 kb upstream and downstream from these lncRNAs were assigned to GO terms and a GO enrichment analysis was conducted.
Furthermore, the neighboring genes of lncRNAs that were expressed in both WT and rag1+/− zebrafish were annotated by GO enrichment and KEGG analysis (Fig. 8A,B). In this case, the GO terms were mainly associated with metabolic processes and RNA processing, although, as expected, several terms directly related to the immune system were also observed, such as positive regulation of TOR signaling, extrinsic apoptotic signaling pathway via death domain receptors, blood coagulation-fibrin clot formation, superoxide metabolic process and cellular response to oxidative stress (Fig. 8A). Numerous terms associated with cellular shape and cytoskeleton organization were also potentially regulated by lncRNAs induced after SVCV infection, regardless of the zebrafish line (Fig. 8A). The KEGG analysis showed a large number of processes linked to viral infection, such as endocytosis, the MAPK signaling pathway, herpes simplex infection, the Toll-like receptor signaling pathway, the RIG-I-like receptor signaling pathway, and the NOD–like receptor signaling pathway (Fig. 8B).
qPCR validation of differentially expressed lncRNAs
Twelve putative lncRNAs were evaluated by qPCR. Among these lncRNAs, four were exclusively modulated in WT after SVCV infection, five were exclusively modulated in rag1+/−, and 3 were affected by the infection in both zebrafish lines. From the lncRNA qPCR results, similar expression profiles were observed between the fold-changes obtained via in silico analysis and the fold-changes determined by qPCR (Fig. 9A). Furthermore, a positive correlation was observed between the RNA-Seq and qPCR fold-change values evaluated using Pearson’s correlation coefficient (r = 0.94) (Fig. 9B).
qPCR validation of the lncRNA expression. (A) qPCR and in silico fold-change values of twelve lncRNAs modulated after SVCV challenge in WT and/or rag1+/− zebrafish transcriptomes, (B) Correlation between the qPCR and RNA-Seq fold-change values evaluated using Pearson’s correlation coefficient (r = 0.94).
Discussion
Members of the Rhabdoviridae family are important pathogens of valuable fish species in world aquaculture33,34,35, with SVCV being the main pathogenic rhabdovirus of cyprinids, such as zebrafish17. Evidence of the participation of lncRNAs in the antiviral immune response in teleosts is scarce9,11, and there is a total lack of such evidence related to fish rhabdoviruses. In the last few years, lncRNAs that are differentially expressed in virus-infected cells have been linked to increases or decreases in the expression of typical antiviral genes. Furthermore, the described modulatory functions of these lncRNAs have mainly been related to their impact on the expression of interferon-stimulated genes (ISGs)36,37,38,39. Additionally, viruses can alter the expression of certain host lncRNAs to favour their own replication40,41.
Recently, modulation of lncRNAs was described after rhabdovirus infection in mice infected with the rabies virus (RABV)42. The authors identified a total of 140 lncRNAs that were differentially expressed at 8 days post-infection. Moreover, by using GO and KEGG pathway analyses, the authors identified genes associated with immune-related processes (e.g., apoptosis, Toll-like receptor, TNF, MAPK, B cell receptor and T cell receptor signaling pathway genes) as potential targets of these lncRNAs, suggesting putative participation of these lncRNAs in the immune response42. In this regard, putative roles of lncRNAs have been described in Atlantic salmon in response to different pathogens (infectious salmon anaemia –ISA– virus, the intracellular bacterium Piscirickettsia salmonis and the ectoparasite copepod Caligus rogercresseyi)11. Moreover, in Atlantic salmon, tissue-specific lncRNA expression has been reported in response to ISAV, and a correlation between the expression of specific lncRNAs and antiviral transcript expression has been suggested9.
LncRNAs participate in the modulation of innate and adaptive immune responses2. Since the majority of lncRNAs are expressed in specific cell types2, the use of mutant animals lacking a particular immune cell type could help to elucidate the specific roles of the different lncRNA sets. This is the case for the rag1−/− zebrafish mutant, which is a T and B cell-deficient model, and as a consequence, these animals rely exclusively on their innate immunity43. It was previously reported that, under naïve conditions, adult homozygous rag1−/− fish possess enhanced innate immunity to compensate for the absence of adaptive immunity, and these fish have been shown to be more resistant to SVCV compared to WT fish31,32,44. In this study, when we analyzed the survival rates of WT and heterozygous rag1+/− mutants after challenge with SVCV, we did not observe significant differences in mortality. Therefore, because homozygous fish possess increased innate immunity, the use of heterozygous rag1+/− mutants could improve the identification of lncRNAs associated with adaptive immunity due to the presence of B and T lymphocytes in these animals. We assume that after SVCV challenge, the expression of those genes involved in the production of immunoglobulins and TCR recombination should be increased in rag1+/− fish compared to WT. In this work, we wanted to take advantage of this model to study lncRNA profiles after early infection with SVCV in both WT and rag1+/− zebrafish.
A total of 12,165 putative lncRNA sequences were identified in zebrafish kidneys, 10,859 of which were common to the two lines, while 915 were excusive to WT, and 337 were only found in rag1+/−. Interestingly, when the functionality of the neighboring protein-coding genes was analyzed, the biological process category immune system process appeared to be represented in rag1+/− but not in WT zebrafish. This difference could indicate that these exclusively expressed lncRNAs allow a compensatory response to the partial deficiency of functional Rag1 in rag1+/− fish, increasing the expression of genes associated with acquired immunity. Indeed, from the lncRNA genome location, five lncRNAs were located near rag1 and rag2 genes, and two of them were up-regulated in rag1+/− compared to WT fish. However, future functional studies are necessary to understand the role of these lncRNA. The other biological processes observed among the rag1+/– exclusive lncRNAs were associated with the cellular response, cell signaling and communication terms. WT-exclusive lncRNAs were located close to genes that are mainly involved in metabolic processes. If we take into consideration the energy cost of the immune response45, the compensatory and/or deficient mechanisms present in the mutant fish could explain the differences in metabolism-related GO terms between the two lines, although we cannot discard other possible collateral effects of the mutation.
The enrichment analysis performed for the neighboring genes of lncRNAs that were exclusively modulated in rag1+/− after SVCV challenge showed several immune-related GO terms, including T cell-mediated immunity, regulation of adaptive immune response, and regulation of adaptive immune response/built from immunoglobulin, among others. These processes are mediated by the adaptive immune system, which is nonexistent in rag1−/− fish but presumably partially present in rag1+/−. This difference is probably due to a compensatory mechanism present in rag1+/− zebrafish to effectively oppose SVCV infection with an efficient B and T lymphocyte response. Therefore, we can presuppose the involvement of these specific lncRNAs in the modulation of adaptive responses.
Furthermore, the GO enrichment analysis of the genes surrounding lncRNAs that were induced after viral infection in both WT and rag1+/− showed high representation of terms related to RNA processing and metabolism. The high representation of RNA processing mechanisms (RNA processing, translational initiation, mRNA cleavage, RNA splicing, tRNA splicing, maturation of 5.8S rRNA, ribosomal small subunit assembly, and cytoplasmic mRNA processing body assembly) highlight the relevance of SVCV-induced lncRNAs in the transcriptional and translational machinery of the host cells. An increasing number of lncRNAs have been linked to the modulation of these processes, especially splicing regulation46. Among the identified metabolic processes, we found terms consistent with the activation of immune cells (tricarboxylic acid cycle, gluconeogenesis, superoxide metabolic process, and phospholipid biosynthetic process). Increased lipid biosynthesis and decreased fatty acid oxidation are also key mechanisms involved in such activation47. Indeed, the enriched term positive regulation of TOR signaling is also closely related to this process. The mammalian target of rapamycin (mTOR) is directly involved in the induction of aerobic glycolysis and promotes the synthesis of fatty acids48. mTOR is also an inhibitor of autophagy49, a controversial mechanism that has been described as both antiviral and pro-viral machinery50. The extrinsic apoptotic signaling pathway via death domain receptors, which was enriched in this analysis as well, is especially relevant in the elimination of virus-infected cells51. On the other hand, lncRNAs regulating blood coagulation-fibrin clot formation are also expected due to the highly haemorrhagic nature of SVCV.
Other interesting groups of enriched GO terms identified after infection were related to cell morphogenesis (actin filament depolymerization, cytoskeleton organization, positive regulation of pseudopodium assembly, regulation of cell morphogenesis). Virion binding to the cell surface triggers signaling mechanisms that alter the cell surface and activate endocytosis52. One of the most enriched KEGG pathways among the potential targets of the SVCV-induced lncRNAs in both lines was endocytosis. LncRNAs that are potentially involved in virion internalization remain unexplored, but some lncRNAs participating in cytoskeletal remodeling have been identified in tumors53,54.
The KEGG pathway analysis of SVCV-induced lncRNAs indicated that these lncRNAs were mainly neighbors of genes involved in metabolic pathways and, particularly, that the insulin pathway showed strong representation. It has been well established, especially for the hepatitis C virus, that infection induces insulin resistance55,56. Numerous lncRNAs have been linked to insulin resistance and diabetes, and these lncRNAs therefore represent a very interesting target for novel therapeutic strategies57. Nevertheless, this effect could be related to changes in glucose metabolism suffered by the immune cells upon activation. As expected, numerous immune-related pathways were also identified (MAPK signaling pathway, herpes simplex infection, Toll-like receptor signaling pathway, RIG-I-like receptor signaling pathway and NOD-like receptor signaling pathway), including some common pathways similar to those observed in mice after RABV infection42.
Therefore, we can conclude that SVCV infection in zebrafish induces strong modulation of lncRNAs with a potential role in the immune response, and the heterozygous rag1 mutant could be a useful tool for the identification of lncRNAs linked to adaptive immunity. Furthermore, the lncRNAs that were modulated in both lines after viral infection represent an excellent source of information for further functional studies focused on the identification of their specific roles under infection. Nevertheless, we are far from understanding all of these coding gene-lncRNA interactions in detail. Future functional investigations could clarify the specific roles of the various lncRNAs modulated in response to the virus.
Data Availability
The raw read sequences were deposited in the Sequence Read Archive (SRA) (https://www.ncbi.nlm.nih.gov/sra) under accession number PRJNA532380.
References
Ponting, C. P., Oliver, P. L. & Reik, W. Evolution and functions of long noncoding RNAs. Cell 136, 629–641 (2009).
Aune, T. M. & Spurlock, C. F. Long non-coding RNAs in innate and adaptive immunity. Virus Res. 212, 146–160 (2016).
Kaushik, K. et al. Dynamic expression of long non-coding RNAs (lncRNAs) in adult zebrafish. PloS One 8, e83616 (2013).
Chen, W. et al. Comprehensive analysis of coding-lncRNA gene co-expression network uncovers conserved functional lncRNAs in zebrafish. BMC Genomics 19, 112 (2018).
Jiang, P. et al. Characterization of lncRNAs involved in cold acclimation of zebrafish ZF4 cells. PloS One 13, e0195468 (2018).
Wang, X. et al. Screening and functional identification of lncRNAs under β-diketone antibiotic exposure to zebrafish (Danio rerio) using high-throughput sequencing. Aquat. Toxicol. 182, 214–225 (2017).
Wei, N. et al. Knockdown of PU. 1 mRNA and AS lncRNA regulates expression of immune-related genes in zebrafish Danio rerio. Dev. Comp. Immunol. 44, 315–319 (2014).
Atianand, M. K., Caffrey, D. R. & Fitzgerald, K. A. Immunobiology of long noncoding RNAs. Annu. Rev. Immunol. 35, 177–198 (2017).
Boltaña, S., Valenzuela-Miranda, D., Aguilar, A., Mackenzie, S. & Gallardo-Escárate, C. Long noncoding RNAs (lncRNAs) dynamics evidence immunomodulation during ISAV-Infected Atlantic salmon (Salmo salar). Sci. Rep. 6 (2016).
Valenzuela-Miranda, D. & Gallardo-Escárate, C. Novel insights into the response of Atlantic salmon (Salmo salar) to Piscirickettsia salmonis: Interplay of coding genes and lncRNAs during bacterial infection. Fish Shellfish Immunol. 59, 427–438 (2016).
Tarifeño-Saldivia, E., Valenzuela-Miranda, D. & Gallardo-Escárate, C. In the shadow: The emerging role of long non-coding RNAs in the immune response of Atlantic salmon. Dev. Comp. Immunol. 73, 193–205 (2017).
Valenzuela-Muñoz, V., Valenzuela-Miranda, D. & Gallardo-Escárate, C. Comparative analysis of long non-coding RNAs in Atlantic and Coho salmon reveals divergent transcriptome responses associated with immunity and tissue repair during sea lice infestation. Dev. Comp. Immunol. 87, 36–50 (2018).
Luo, H. et al. LncRNA and mRNA profiling during activation of tilapia macrophages by HSP70 and Streptococcus agalactiae antigen. Oncotarget 8, 98455–98470 (2017).
Bassing, C. H., Swat, W. & Alt, F. W. The mechanism and regulation of chromosomal V(D)J recombination. Cell 109, S45–S55 (2002).
Wienholds, E., Schulte-Merker, S., Walderich, B. & Plasterk, R. H. A. Target-selected inactivation of the zebrafish rag1 gene. Science 297, 99–102 (2002).
Ahne, W. et al. Spring viremia of carp (SVC). Dis. Aquat. Organ. 52, 261–272 (2002).
Sanders, G. E., Batts, W. N. & Winton, J. R. Susceptibility of zebrafish (Danio rerio) to a model pathogen, spring viremia of carp virus. Comp. Med. 53, 514–521 (2003).
López-Muñoz, A., Roca, F. J., Sepulcre, M. P., Meseguer, J. & Mulero, V. Zebrafish larvae are unable to mount a protective antiviral response against waterborne infection by spring viremia of carp virus. Dev. Comp. Immunol. 34, 546–552 (2010).
Encinas, P. et al. Identification of multipath genes differentially expressed in pathway-targeted microarrays in zebrafish infected and surviving Spring Viremia Carp Virus (SVCV) suggest preventive drug candidates. PloS One 8, e73553 (2013).
Varela, M. et al. Cellular visualization of macrophage pyroptosis and onterleukin-1β release in a viral hemorrhagic infection in zebrafish larvae. J. Virol. 88, 12026–12040 (2014).
Pereiro, P. et al. Zebrafish Nk-lysins: First insights about their cellular and functional diversification. Dev. Comp. Immunol. 51, 148–159 (2015).
Nusslein-Volhard, C. & Dahm, R. Zebrafish: A Practical Approach (Oxford University Press, 2002).
Westerfield, M. The Zebrafish Book: A Guide For The Laboratory Use Of Zebrafish (University of Oregon Press, 2000).
Reed, L. J. & Muench, H. A simple method of estimating fifty percent endpoints. Am. J. Epidemiol. 27, 493–497 (1938).
Renaut, S., Nolte, A. W. & Bernatchez, L. Mining transcriptome sequences towards identifying adaptive single nucleotide polymorphisms in lake whitefish species pairs (Coregonus spp. Salmonidae). Mol. Ecol. 19, 115–131 (2010).
Huang, D. W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 4, 44–57 (2009).
Supek, F., Bosnjak, M., Skunca, N. & Smuc, T. REVIGO summarizes and visualizes long lists of Gene Ontology terms. PloS One 6, e21800 (2011).
Shannon, P. et al. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Rozen, S. & Skaletsky, H. Primer3 on the WWW for general users and for biologist programmers. Methods Mol. Biol. 132, 365–368 (2000).
Pfaffl, M. W. A new mathematical model for relative quantification in real-time RT-PCR. Nucleic Acids Res. 29, 2002–2007 (2001).
García-Valtanen, P. et al. Zebra fish lacking adaptive immunity acquire an antiviral alert state characterized by upregulated gene expression of apoptosis, multigene families, and interferon-related genes. Front. Immunol. 8, 121 (2017).
Novoa, B. et al. Rag1 immunodeficiency-induced early aging and senescence in zebrafish are dependent on chronic inflammation and oxidative stress. Aging Cell 18, e13020 (2019).
Pereiro, P., Figueras, A. & Novoa, B. Turbot (Scophthalmus maximus) vs. VHSV (viral hemorrhagic septicemia virus): a review. Front. Physiol. 7, 192 (2016).
Dixon, P., Paley, R., Alegria-Moran, R. & Oidtmann, B. Epidemiological characteristics of infectious hematopoietic necrosis virus (IHNV): a review. Vet.Res. 47, 63 (2016).
Purcell, M. K., Laing, K. J. & Winton, J. R. Immunity to fish rhabdoviruses. Viruses 4, 140–166 (2012).
Kambara, H. et al. Negative regulation of the interferon response by an interferon-induced long non-coding RNA. Nucleic Acids Res. 42, 10668–10680 (2014).
Ouyang, J. et al. NRAV, a long noncoding RNA, modulates antiviral responses through suppression of interferon-stimulated gene transcription. Cell Host Microbe 16, 616–626 (2014).
Qian, X., Xu, C., Zhao, P. & Qi, Z. Long non-coding RNA GAS5 inhibited hepatitis C virus replication by binding viral NS3 protein. Virology 492, 155–165 (2016).
Nishitsuji, H. et al. Long noncoding RNA #32 contributes to antiviral responses by controlling interferon-stimulated gene expression. Proc. Natl. Acad. Sci. USA 113, 10388–10393 (2016).
Imam, H., Bano, A. S., Patel, P., Holla, P. & Jameel, S. The lncRNA NRON modulates HIV-1 replication in a NFAT-dependent manner and is differentially regulated by early and late viral proteins. Sci. Rep. 5, 8639 (2015).
Wang, J. et al. Host long noncoding RNA lncRNA-PAAN regulates the replication of influenza A virus. Viruses 10, 330 (2018).
Zhao, P. et al. Analysis of expression profiles of long noncoding RNAs and mRNAs in brains of mice infected by rabies virus by RNA sequencing. Sci. Rep. 8, 11858 (2018).
Jima, D. D. et al. Enhanced transcription of complement and coagulation genes in the absence of adaptive immunity. Mol. Immunol. 46, 1505–1516 (2009).
Petrie-Hanson, L., Hohn, C. & Hanson, L. Characterization of rag1 mutant zebrafish leukocytes. BMC Immunol. 10, 8 (2009).
Georgiev, A. V., Kuzawa, C. W. & McDade, T. W. Trade-offs between acquired and innate immune defenses in humans. Evol. Med. Public Health 2016, 1–16 (2016).
Benhamed, M., Romero-Barrios, N., Crespi, M., Legascue, M. F. & Ariel, F. Splicing regulation by long noncoding RNAs. Nucleic Acids Res. 46, 2169–2184 (2018).
Hubler, M. J. & Kennedy, A. J. Role of lipids in the metabolism and activation of immune cells. J. Nutr. Biochem. 34, 1–7 (2016).
Jones, R. G. & Pearce, E. J. MenTORing immunity: mTOR signaling in the development and function of tissue-resident immune cells. Immunity 46, 730–742 (2017).
Jung, C. H., Ro, S.-H., Cao, J., Otto, N. M. & Kim, D.-H. mTOR regulation of autophagy. FEBS Letters 584, 1287–1295 (2010).
Shoji-Kawata, S. & Levine, B. Autophagy, antiviral immunity, and viral countermeasures. Biochim. Biophys. Acta 1793, 1478–1484 (2009).
Zhou, X., Jiang, W., Liu, Z., Liu, S. & Liang, X. Virus infection and death receptor-mediated apoptosis. Viruses 9, 316 (2017).
Taylor, M. P., Koyuncu, O. O. & Enquist, L. W. Subversion of the actin cytoskeleton during viral infection. Nat. Rev. Microbiol. 9, 427 (2011).
Ma, C. et al. Long intergenic noncoding RNA 00673 promotes non-small-cell lung cancer metastasis by binding with EZH2 and causing epigenetic silencing of HOXA5. Oncotarget 8, 32696–32705 (2017).
Tang, Y. et al. LncRNAs regulate the cytoskeleton and related Rho/ROCK signaling in cancer metastasis. Mol. Cancer 17, 77 (2018).
Parvaiz, F. et al. Hepatitis C virus infection: molecular pathways to insulin resistance. Virol. J. 8, 474 (2011).
Shintani, Y. et al. Hepatitis C virus infection and diabetes: direct involvement of the virus in the development of insulin resistance. Gastroenterology 126, 840–848 (2004).
Leti, F. & DiStefano, K. J. Long noncoding RNAs as diagnostic and therapeutic targets in type 2 diabetes and related complications. Genes 8, E207 (2017).
Acknowledgements
This work was funded by projects BIO2017-82851-C3-1-R of the Spanish Ministerio de Economía y Competitividad, IN607B 2016/12 from Consellería de Economía, Emprego e Industria (GAIN, Xunta de Galicia), FONDAP # 15110027 and FONDECYT #1180867 from CONICYT-Chile. Patricia Pereiro wishes to thank the Axencia Galega de Innovación (GAIN, Xunta de Galicia) for her postdoctoral contract (IN606B-2018/010), and Margarita Álvarez-Rodríguez was the recipient of an FPU fellowship from the Spanish Ministerio de Educación (FPU014/05517).
Author information
Authors and Affiliations
Contributions
V.V.-M., C.G.-E., A.F. and B.N. conceived and designed the project. P.P. and M.A.-R. conducted the experimental infections, sampling and RNA isolation. V.V.-M., C.G.-E. and A.F. performed the reads trimming, assembly, RNA-Seq and statistical analyses. V.V.-M., P.P. and C.G.-E. analyzed the generated data. P.P. conducted the qPCR validation. V.V.-M., P.P., C.G.-E. and B.N. wrote the manuscript. All listed authors revised, edited, read and approved the manuscript.
Corresponding authors
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Valenzuela-Muñoz, V., Pereiro, P., Álvarez-Rodríguez, M. et al. Comparative modulation of lncRNAs in wild-type and rag1-heterozygous mutant zebrafish exposed to immune challenge with spring viraemia of carp virus (SVCV). Sci Rep 9, 14174 (2019). https://doi.org/10.1038/s41598-019-50766-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-50766-0
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.