Abstract
Numerous studies have proposed biomarkers based on magnetic resonance imaging (MRI) to detect and predict the risk of evolution toward Alzheimer’s disease (AD). Most of these methods have focused on the hippocampus, which is known to be one of the earliest structures impacted by the disease. To date, patch-based grading approaches provide among the best biomarkers based on the hippocampus. However, this structure is complex and is divided into different subfields, not equally impacted by AD. Former in-vivo imaging studies mainly investigated structural alterations of these subfields using volumetric measurements and microstructural modifications with mean diffusivity measurements. The aim of our work is to improve the current classification performances based on the hippocampus with a new multimodal patch-based framework combining structural and diffusivity MRI. The combination of these two MRI modalities enables the capture of subtle structural and microstructural alterations. Moreover, we propose to study the efficiency of this new framework applied to the hippocampal subfields. To this end, we compare the classification accuracy provided by the different hippocampal subfields using volume, mean diffusivity, and our novel multimodal patch-based grading framework combining structural and diffusion MRI. The experiments conducted in this work show that our new multimodal patch-based method applied to the whole hippocampus provides the most discriminating biomarker for advanced AD detection while our new framework applied into subiculum obtains the best results for AD prediction, improving by two percentage points the accuracy compared to the whole hippocampus.
Similar content being viewed by others
Introduction
Alzheimer’s disease (AD) is an irreversible neurodegenerative process leading to mental dysfunctions. Subjects presenting mild cognitive impairment (MCI) have a higher risk of developing AD1. To study the preclinical phase of the disease, the Alzheimer’s disease neuroimaging initiative (ADNI) has been set up based on two MCI definitions: early MCI (eMCI) and late MCI (lMCI). Subjects with eMCI have milder cognitive impairment than those with lMCI, both suffering from amnesic MCI2. Such clinical symptoms are caused by changes like synaptic and neuronal losses that lead to structural and microstructural alterations. Neuroimaging studies performed on AD subjects reveal that when an AD diagnosis is made, alterations of brain structure are already advanced, emphasizing the need to study the early stages of the disease.
The improvement of medical imaging techniques such as magnetic resonance imaging (MRI) has enabled the development of efficient biomarkers capable of detecting alterations caused by AD3. Over the past years, many methods have been proposed to perform automatic detection of alterations associated with AD. First, studies proposed methods based on specific regions of interest (ROI) capturing alterations at an anatomical scale. Among structures impacted by AD, previous investigations have been focused on the hippocampus4,5,6, entorhinal cortex (EC)7,8,9, parahippocampal gyrus, amygdala10, or parietal lobe11,12. Alterations of these structures are usually estimated using volume13,14, shape15,16, or cortical thickness17,18 measurements. Beside ROI-based methods, whole brain analysis performed on structural MRI (s-MRI) has been proposed to detect areas impacted by AD at a voxel scale. These methods are usually based on voxel-based morphometry (VBM) or tensor-based morphometry (TBM) frameworks19. It is interesting to note that both VBM and ROI-based studies confirmed that the medial temporal lobe is a key area in detecting the first signs of AD20,21,22,23,24,25. These studies also showed that the hippocampus is one of the earliest regions altered by AD in the medial temporal lobe26. Moreover, the hippocampus volume is one of the criteria that can be used to confirm the diagnosis of AD in clinical routines27. Recently, advanced methods were proposed to capture subtler structural alterations of the hippocampus9,28,29,30. Those techniques demonstrated an increase in detection and prediction performances at different AD stages when compared to volume-based methods30. Among them, patch-based grading (PBG) methods demonstrated competitive results to detect the earliest stages of AD before a clinical diagnosis can be made9,29,31. The main idea of this approach is to capture inter-subject pattern similarities via non-local comparisons between two groups of subjects. Such methods have shown their ability to predict AD more than seven years before the conversion to dementia32 and might enable a differential diagnosis33,34.
Thus, the hippocampus has been one of the most studied structures to diagnose AD. However, this structure is not homogeneous, so it is usually subdivided into different subfields. Initial efforts to define the hippocampus subfields were mainly based on cell size, shape, and connectivity35. The terminology differs across segmentation protocols36, but the most recognized definition37 divides hippocampus into the subiculum, the cornu ammonis (CA1/2/3/4), and the dentrate gyrus (DG). The CA1 subfield represents the biggest area in the hippocampus. It is composed of different layers called the stratum radiatum (SR), the stratum lacunosum (SL), the stratum molecular (SM), and the stratum pyramidale (SP). Interestingly, studies have shown that hippocampal subfields could have different functional specializations. It has been suggested that CA3 and DG might be responsible for encoding early retrieval38,39 while CA1 is responsible for consolidation, late retrieval and recognition40,41,42. Furthermore, hippocampal subfields are not equally impacted by AD43,44,45,46,47,48,49. Indeed, several MRI studies demonstrated that subfields are impacted differently according to AD stages. Postmortem and in vivo imaging studies showed that the CA1SR-L-M are the subfields impacted with the greatest atrophy in advanced AD45,46,48. Recently, it has been shown that the subiculum is the earliest affected hippocampal region49,50.
These studies indicate that a subfield analysis of hippocampus alterations at a finer scale with an analysis of the subiculum could provide better tools for AD detection and prediction. The subiculum lies between CA1 and the entorhinal cortex in the medial temporal lobe. It shows a columnar organization (parasubiculum, presubiculum, postsubiculum, prosubiculum) combined with a laminar organization and is the main output of the hippocampus. Aside from those from CA1, several other extrinsic afferents terminate within the subiculum from the temporal lobe cortex (entorhinal cortex, perirhinal cortex, parahippocampal cortex, and amygdala). The anterior thalamic nuclei also project densely upon the subicular complex. In terms of efferent pathways, the subiculum projects to more extrinsic sites than any other hippocampal area. Notably, the subiculum shows dense extrinsic projections toward the anterior thalamic nuclei, the mammillary bodies, and the retrospinal cortex. Regarding its function, the subiculum is implicated in working memory. Several rodent behavioral studies also have shown that subiculum lesions impair spatial memory tasks with spatial working memory having a higher sensitivity than reference memory51.
Although structural MRI is a valuable imaging technique for measuring global structural modifications, such modality is not able to capture microstructural degradation. However, the microstructural modifications caused by AD are believed to occur before the atrophy measured by structural MRI. Therefore, diffusion MRI (d-MRI) appears as a potential candidate in detecting the earliest sign of AD. Several diffusion tensor imaging (DTI) studies proposed automatic methods for detecting modifications of diffusion parameters into the whole white matter volume using machine learning techniques52,53,54. Others studies showed modifications of diffusion parameters for AD patients into specific white matter structures such as the corpus callosum55,56, the fornix57, the cingulum55, and also in gray matter tissues such as the hippocampus58. More advanced d-MRI studies using brain connectivity and fiber tracking have been proposed to extract features describing axonal fiber alterations57,59,60. Finally, it has been shown that the hippocampal mean diffusivity (MD) is correlated to pathology progression and thus could be used as an efficient biomarker of AD61. Moreover, it was demonstrated that MD increases with the development of AD in the gray matter62,63,64. Therefore, in previous work, we showed that patch-based features applied to DTI demonstrated competitive performances to classify the early stages of AD65. Although some studies showed the superiority of MD over volumetric measurement to detect early sign of AD, this difference remains unclear66,67. However, several methods showed the possibility of using volumetric and MD measurements to capture early alterations caused by AD68,69. Recently, a study combining volumetric measurements and mean diffusivity of hippocampus subfields demonstrated that the CA1 and subiculum are the most impacted subfields in late AD stage50.
All these elements indicate that a multimodal method based on hippocampal subfields using an advanced image analysis framework could improve AD detection and prediction. Consequently, in this paper, we propose the study of hippocampal subfield efficiencies using s-MRI and d-MRI modalities for AD detection and prediction. To that purpose, we have developed a novel multimodal patch-based grading fusion scheme to better capture such structural and microstructural alterations. First, we compare the performance of our novel method with volume and MD within the whole hippocampus. Secondly, we demonstrate state-of-the-art performances compared to more advanced d-MRI based methods. Finally, we study the efficiency of each hippocampal subfields in improving AD detection and prediction using volume, MD, and our multimodal patch-based grading method. Our results show that while PBG based on s-MRI obtains the best performance for AD diagnosis, d-MRI obtains the best performance for AD prognosis. Our novel multimodal patch-based grading method based on these two modalities obtains the best scores for both AD detection and prediction. These results highlight that our multimodal patch-based grading provides more robust features than PBG based on only a single modality. Moreover, we demonstrate that the study of the hippocampus at a finer scale improves AD prediction. The experiments conducted with our new multimodal patch-based grading show that the whole hippocampus provides better results for AD detection, but the subiculum is the best area for AD prediction.
Materials
Dataset
Data used in this work was obtained from the Alzheimer’s Disease Neuroimaging Initiative (ADNI) dataset (http://adni.loni.ucla.edu). ADNI is a North American campaign launched in 2003 with the aim of providing MRI, positron emission tomography scans, clinical neurological measures, and other biomarkers. This dataset includes AD patients, MCI, and control normal (CN) subjects. The group of MCI is composed of subjects who have abnormal memory dysfunctions. In this work, we used data from the ADNI-2 campaign that proposes eMCI and lMCI stages. The eMCI and lMCI subgroups were obtained with the Wechsler Scale-Revised Logistical Memory I and II tests in accordance with the education levels of each subject. ADNI-2 provides T1-weighted (T1w) MRI, DTI scans for 54 CN, 79 eMCI, 39 lMCI, and 47 AD subjects. Only patients who have T1w and DTI images were selected in our work. Hence, in this work, we used 52 CN, 99 MCI composed of 65 eMCI, 34 lMCI, and 38 AD instead of the whole initial ADNI-2 dataset. All MRI data and clinical status were collected at the baseline. The list of subjects involved in our experiments is available (http://bit.ly/scirep_mpbg_dataset). Table 1 shows the distribution of the data for each group. The s-MRI and d-MRI scans used for all considered subjects in this study were acquired with the same protocol (https://adni.loni.usc.edu/wp-content/uploads/2010/05/ADNI2_GE_3T_22.0_T2.pdf). T1w MRI acquisition protocol was done with the 3D accelerated sagittal IR-SPGR, according to the ADNI protocol70. The d-MRI is composed of 46 separate angles, 5 T2-weighted images with no diffusion sensitization (b0 images) and 41 directions (b = 1000 s/mm2). The d-MRI protocol was chosen to optimize the signal-to-noise ratio in a fixed scan time71. The native resolution of s-MRI and d-MRI was set to 1 mm3 and 2 mm3, respectively.
MRI processing
T1w images were processed using the volBrain system72 (http://volbrain.upv.es). This system is based on an advanced pipeline providing automatic segmentation of different brain structures from T1w MRI. The preprocessing is based on (a) a denoising step with an adaptive non-local mean filter73, (b) an affine registration in the MNI space74, (c) a correction of the image inhomogeneities75 and (d) an intensity normalization.
Afterward, segmentation of hippocampal subfields was performed with HIPS76 based on a combination of non-linear registration and patch-based label fusion77. This method uses a training library based on a dataset composed of high-resolution T1w images manually labeled according to the protocol proposed by Winterburn et al.37. To perform the segmentation, the images are up-sampled with a local adaptive super-resolution method to fit the training image resolution78. The method provides automatic segmentation of hippocampal subfields gathered into five labels: Subiculum, CA1SP, CA1SR-L-M, CA2-3, and CA4/DG (see Fig. 1). Then, the segmentation maps obtained from the up-sampled T1w images were down-sampled to fit the MNI space resolution. All the following experiments were carried out with images into the MNI space. Finally, an estimation of the total intra-cranial volume was performed79.
DTI processing
The preprocessing of the diffusion-weighted images is based on (a) a denoising step based on the LPCA filter80 and (b) a correction of the head motion using an affine registration. Afterward, we performed several steps to first obtain the mapping between the DWI native space and the MNI space and then to estimate the MD in the MNI space.
- (1)
Estimation of the mapping between DWI native space and MNI space: First, a diffusion tensor model81 estimated at each voxel using Dipy library82. The resulting MD is first linearly registered to the CSF map obtained from the T1w in the MNI space. Then, the MD (in the MNI space) is non-linearly registered to the CSF map (in the MNI space) to compensate for echo-planar imaging (EPI) distortions74. Afterward, the affine transformation and the non-linear deformations are concatenated into a single transformation to obtain the final mapping (including EPI distortion correction) from the DWI native space to the MNI space. It must be noted that the MD map estimated in the DWI native space is only used to estimate the mapping between both spaces.
- (2)
Estimation of the MD in the MNI space: The deformation field estimated at the previous step is used to register the b0 and each DWI direction from their native space into the MNI space using b-spline interpolations74. This is done to limit interpolation artifacts and to correct partial volume effect (PVE). It has been shown that up-sampling each DWI direction individually using interpolation before estimating DTI parameters enables the reduction of PVE present in DTI greatly83. Thus, the final diffusion tensor model is estimated in the MNI space using all the non-linearly registered DWI and b0.
To analyze microstructural modifications, the MD is estimated within each hippocampal subfield and the whole hippocampus structure with the segmentation described in the previous section. MD is defined as \(\frac{{\lambda }_{1}+{\lambda }_{2}+{\lambda }_{3}}{3}\) where λ1, λ2, λ3 are the three eigenvalues of the fitted tensor.
Finally, quality control is conducted to exclude data presenting segmentation errors or misregistration after MRI and DTI preprocessing step. Thus, 10 CN subjects, 18 eMCI, 5 lMCI, and 9 AD patients have been excluded from the initial considered ADNI2 dataset (see the dataset used in our experiments Table 1).
Methods
Patch-based grading
Patch-based grading was first proposed for s-MRI9. The main idea of this exemplar-based method is to use the capability of patch-based techniques in order to capture subtle signal modifications related to anatomical degradations caused by AD. To date, the PBG methods demonstrate state-of-the-art performances in the detection of the earliest stage of AD84. To determine the pathological status of the subject under study, the PBG methods estimate the state of cerebral tissues at each voxel by a similarity measurement. This measurement is performed between the anatomical pattern of the subject under study and those extracted from two training populations, one healthy and another one unhealthy.
First, a training library T composed of two datasets of images is built: one with images from CN subjects and the other one from AD patients. Next, for each voxel xi of the region of interest in the considered subject x, the PBG method produces a weak classifier denoted \({g}_{{x}_{i}}\). This weak classifier provides a surrogate of the pathological grading at the considered position. The weak classifier is computed using a measurement of the similarity between the patch \({P}_{{x}_{i}}\) surrounding the voxel xi belonging to the image under study and a set \({K}_{{x}_{i}}\) of the closest patches extracted from the library T. The most similar patches are found using an approximative nearest neighbor method85. The grading value \({g}_{{x}_{i}}\) at xi is defined as:
where \({P}_{{t}_{j}}\) is the patch surrounding the voxel j belonging to the training template \(t\in T\), and \(w({x}_{i},{t}_{j})\) is the weight assigned to the pathological status pt of the training image t. We estimate w such that:
where \(h=\,{\rm{\min }}\,\parallel {P}_{{x}_{i}}-{P}_{{t}_{j}}{\parallel }_{2}^{2}+\varepsilon \) and \(\varepsilon \to 0\). The pathological status pt is set to −1 for patches extracted from AD patient and to 1 for patches extracted from CN subject. Therefore, the PBG method provides a score representing an estimation of the alterations caused by AD at each voxel. Consequently, cerebral tissues strongly altered by AD have grading values close to −1 contrary to healthy one with scores close to 1.
Multimodal patch-based grading fusion
The patch-based method presented in the previous section was designed to capture structural alterations in T1w MRI. Recently, we proposed the extension this method to DTI modality in order to detect microstructural modifications65. We showed the efficiency of MD grading in improving the classification of the early stages of AD.
In this study, we propose a new framework to perform multimodal patch-based grading (MPBG). To this end, we developed an adaptive fusion of grading maps derived from different modalities (see the example of grading maps on Fig. 2). As shown in the following, this fusion provides more robust and accurate biomarkers compared to monomodal PBG biomarkers.
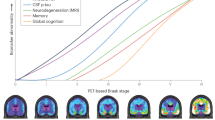
The presented results have been obtained using MRI from patients suffering from different severities of cognitive impairments. From top to bottom slices on the coronal plane of the segmentation maps, and the fusion of T1w and MD patch-based grading (i.e., MPBG: Multimodal Patch-Based Grading) with the proposed multimodal patch-based grading method. The blue and red colors represent healthy and altered tissues, respectively. To avoid bias due to overlap between training and testing datasets, the library has been constructed within a leave-one-out procedure.
As in the previous section, a training library of CN and AD subjects is built for each modality. Next, at each voxel within the ROI of the considered subject and for each modality, a set K of most similar patches is extracted. This step provides one set K of patches per modality \(m\in M\), where M corresponds to the set of the different modalities provided. Nevertheless, at each voxel, the quality of the grading estimation is not the same for all the modalities. Therefore, the degree of confidence is estimated with the function α defined as:
that reflects the confidence of the grading value \({g}_{{x}_{i}}\) for the modality m at the voxel xi. This confidence measure is derived from multi-feature fusion86. Thus, each modality provides a weak classifier at each voxel that is weighted with its degree of confidence \({\alpha }_{{x}_{i,m}}\). The multimodal grading denoted \({g}_{{x}_{i}}\), is given by:
In other words, the weights w and \({K}_{{x}_{i,m}}\) are estimated independently for each modality and combined afterward. Therefore, the proposed combination framework is spatially adaptive and takes advantage of the a local degree of confidence \({\alpha }_{{x}_{i,m}}\) for each modality m. When the matches found for a modality in the training library is composed of good candidates (i.e., patches very similar to the patch from the subject under study), our confidence \({\alpha }_{{x}_{i,m}}\) in the grading estimation for this modality is high. In the end, this modality will have a high weight in the mixing procedure described in (4).
Features estimation
Features were estimated in each hippocampal subfield and over the whole hippocampus as the union of all hippocampal subfields masks. To reduce the inter-individual variability, all volumes are normalized by the total intra-cranial volume87. Afterward, we aggregate weak local classifiers of the grading map into a single feature for each considered structure (i.e., hippocampal subfields, and whole hippocampus) by averaging them. Then, patch-based grading features are computed by an unweighted vote of the weak classifiers using the segmentation masks (see Fig. 3). Finally, to prevent the bias introduced as the structural alterations due to aging, all the features (i.e., volume, mean of MD and MPBG) are age corrected with a linear regression based on the CN group88.
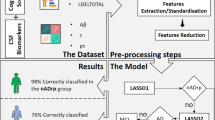
Proposed multimodal patch-based grading framework. At left, the input data: T1w images and MD maps into the MNI space. Data represented in this figure belongs to a CN subject. At the middle: a coronal view of hippocampal subfields segmentation on T1w, and the corresponding coronal view of a multimodal patch-based grading (MPBG) map estimated on T1w and MD. At right, the considered subfield biomarkers for all subjects under study. From top to bottom, the features are the volumes, the MPBG values, and the average of MD.
Implementation
We use the OPAL method to find the most similar patches in the training library89. OPAL is a fast approximate nearest neighbor patch search technique. This method processes each modality in about 4 seconds on a standard computer. A leave-one-out procedure was followed to construct the training library. Hence, for each test subject, a different training library is built. Consequently, the training library T is composed of 37 images from CN subjects and 37 images from AD subjects, for a total of 76 images. The number of patches extracted from both training libraries is K = 160 (i.e., 80 from CN subjects and 80 from AD patients) and the patch size is 5 × 5 × 5 voxels.
Furthermore, as done in our PBG DTI study65, we used zero normalized sum of squared differences for T1w to compute the L2 norm (see Eq. (2)). On the other hand, d-MRI is a quantitative imaging technique. Therefore, a straight sum of squared differences is used for MD in Eq. (2) in order to preserve the quantitative information.
Validation
To evaluate the efficiency of each considered biomarker in detection of AD alterations, the CN group is compared to the group of AD patients. In addition, to discriminate the impairment severity of MCI group, eMCI versus lMCI classification is conducted. The classification step is performed with linear discriminant analysis (LDA) within a repeated stratified 5-fold cross-validation with 200 iterations. Mean area under the curve (AUC) and mean accuracy (ACC) are computed to compare performance for each biomarker over the 200 iterations.
Statistical analyses
Statistical tests were conducted with an analysis of variances (ANOVA) procedure to determine the significance of biomarkers changes, related to the alterations caused by AD. The results of these tests have been corrected for multiple comparisons with Bonferroni’s method. Significant changes have been tested within six comparisons (i.e., CN-AD, CN-eMCI, CN-lMCI, eMCI-lMCI, eMCI-AD, and lMCI-AD). These comparisons have been achieved into each region of the hippocampus and with the three considered biomarkers (i.e., the volume, the average of MD, and our newly proposed MPBG). Finally, for each iteration of our stratified 5-fold cross-validation, we estimated the confidence interval of AUC using bootstrap iterated for 100 iterations90. Then an average of the minimum and maximum bounds are computed. The results presented in this paper show the average confidence interval based on these average bounds.
Results
In this section, the results are presented in three parts. In the first part, we compare the different approaches applied within the entire hippocampus structure to evaluate the performance of our new MPBG compared to usual biomarkers such as volume and average MD. In the second part, we compare the accuracy of each considered biomarker within hippocampal subfields in order to investigate the potential of hippocampal subfield analysis to improve the result of AD detection and prediction. Finally, we compare the results of our proposed multimodal biomarker with state-of-the-art methods based on d-MRI to show the competitive performance of our approach.
Whole hippocampus
Results of the comparisons over the whole hippocampus are presented in Table 2. In this experiment, we compared the results of volume, mean of MD and PBG applied with both modality and MPBG over the whole hippocampus.
First, the hippocampus volume and its average of MD were compared. For CN versus AD classification, the volume obtains 86.6% of AUC, and the average of MD obtains 80.6%. For eMCI versus lMCI classification, the volume and the average of MD obtain 59.4% and 55.6% of AUC, respectively. The experiments demonstrate that the volume of the hippocampus results in better classification performances than the average of MD for all comparison, especially for CN versus AD. Second, PBG biomarkers applied with T1w and MD were compared. The results showed that T1w PBG provides better results than MD PBG with 92.6% of AUC for CN versus AD classification. However, for eMCI versus lMCI classification MD grading provides the best results with 69.5% of AUC. MPBG methods combining both modalities performed similarly to the best results for CN versus AD and eMCI versus lMCI with 92.1% and 69.5% of AUC, respectively. Finally, the proposed MPBG biomarker provides results similar to the best modalities for all considered comparisons. MPBG improves CN versus AD comparison result by 5.5% of AUC and by over 10% of AUC for eMCI versus lMCI comparison. Thus, MBPG biomarker has a good capability to capture modifications caused by AD at different stages of severity (see Fig. 2).
Hippocampal subfields
Figure 4 shows the distribution of volumes (A), the average of MD (B), and the MPBG (C) for each hippocampal subfield at different AD stages. For each comparison, a p-value was estimated with a multi-comparison test91. We can note that for all hippocampal subfields, alterations caused by the disease are related to volume and MPBG decrease with MD increase. The subiculum subfield presents the most significant differences for CN versus lMCI using volume and MD, for AD versus lMCI using MD, and for eMCI versus lMCI using MPBG. Indeed, it is the only subfield providing a p-value inferior to 0.05 for the comparison of CN versus eMCI using volume, a p-value inferior to 0.01 for lMCI versus AD using MD and a p-value inferior to 0.001 to eMCI versus lMCI using MPBG, which are the most challenging comparisons. The distribution of MPBG shows better discrimination between each group for all hippocampal subfields. Indeed, MPBG applied within CA1SP, and CA1SR-L-M provides p-values inferior to 0.01 for eMCI versus lMCI. Moreover, MPBG applied within the subiculum provides p-value inferior to 0.001 for the same comparison. Thus, MPBG enables AD detection using each subfield with an advantage for subiculum for the comparison of eMCI versus lMCI.
Distribution of the volume (A), MD (B), and MPBG (C) for the different considered groups. The normalized volumes are provided in mm3 in the MNI space for each subfield, MD is the mean of MD values into each subfield in mm2.s−1, and MPBG is the mean patch-based grading values into each subfield. Blue, cyan, orange, and red colors represent CN, eMCI, lMCI, and AD subjects, respectively. Statistical tests have been performed with ANOVA procedure and corrected for multiple comparisons with the Bonferroni’s method. The p-values inferior to 0.05, 0.01, and 0.001 are represented with *, **, and ***, respectively.
To estimate the efficiency of the considered biomarkers for AD detection, we also performed a classification experiment. Figure 5 shows the results of two comparisons, CN versus AD (part noted A in the figure) and eMCI versus lMCI (part noted B). First, for AD diagnosis (i.e., CN versus AD classification), the subfield providing the most discriminant volume is the CA1S-R-L-M with an AUC of 86.0%. Moreover, the most discriminating MD biomarker is given by the subiculum with an AUC of 88.1%. For this comparison, the MD of subiculum is the only biomarker performing better results than the whole hippocampus. The CA1SP provides the best results using MPBG feature with an AUC of 92.1%, followed by the CA1S-R-L-M and the subiculum.
AUC computed for CN versus AD (A), eMCI versus lMCI (B) comparisons with the different considered biomarkers in each hippocampal area. Results of subfields are grouped by features (i.e., the volume, the average of MD and the MPBG). Upper bounds of the confidence interval are represented with vertical bars. Whole hippocampus volume biomarker provides the best results with a mean AUC of 86.6% for CN versus AD comparison, followed by the CA1S-R-L-M volume that obtains a mean AUC of 86%. Subiculum volume provides the best results for eMCI versus lMCI with a mean AUC of 66.1%. The average of MD for subiculum obtains the best results for CN versus AD and eMCI versus lMCI with a mean AUC of 88.1% and 62.4%, respectively. Whole hippocampus MPBG obtains the best results for CN versus AD with a mean AUC of 92.1%. Subiculum MPBG obtains the best results for eMCI versus lMCI comparison with a mean AUC of 71.8%. This comparison shows that subiculum is the only biomarker providing better results than the whole hippocampus. This figure presents mean AUC and the mean confidence intervals that have been computed for each iteration of the stratified 5-fold cross-validation procedure carried out in our experiments.
Second, for eMCI versus lMCI classification, the subiculum provides the best results for each considered feature. Indeed, the subiculum obtained an AUC of 66.1% for the volume, 62.4% for the average of MD, and 71.8% for MPBG. Moreover, the subiculum also provided better results than the whole hippocampus for each considered method. Thus, the experiments conducted with three different biomarkers showed that the use of hippocampal subfields, especially the subiculum, results in better AD prediction than the whole hippocampal analysis.
Comparison with state-of-the-art methods
Direct comparison with other monomodal methods applied on ADNI1 is difficult since group definition (stable MCI and progressive MCI) are different. However, as recently shown, T1w PBG provides state-of-the-art performance on ADNI1 dataset, even compared to deep learning methods92. Consequently, the results presented in this paper with T1w PBG on ADNI2 can reasonably be considered competitive and can be used as a reference.
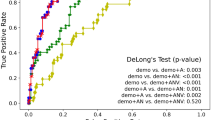
Consequently, to evaluate the performance of the proposed MPBG, we compared it with state-of-the-art multimodal methods using d-MRI. To this end, we used the ACC values published by the authors. Table 3 shows the comparison of our proposed biomarkers within the hippocampal area providing the best results (i.e. the whole hippocampus and the subiculum) with the state-of-the-art methods using similar dataset based on ADNI-2. We compared these biomarkers with a method using features based on tractography93, two different methods based on connectivity networks of the different brain structures60,94,95, and a voxel-based method that analyzes alterations of white matter96. The results of the comparison show that MPBG over the whole hippocampus obtains the best score for AD versus CN with 88.1% of accuracy while the best result is achieved by a voxel-based method with a feature selection96 that obtained 87.0% on similar ADNI2 dataset. For the best of our knowledge, the two works providing eMCI versus lMCI comparison60,94 using s-MRI and d-MRI from a similar ADNI2 dataset are based on a connectivity network and obtained 63.4% and 65.0%, respectively. These comparisons demonstrate the relevance of MPBG biomarkers for AD detection and prediction. Indeed, our method provides similar results than the best methods with similar dataset for CN versus AD classification and provides the best results for eMCI versus lMCI classification. Moreover, the proposed MPBG method based on the subiculum improves the performance for eMCI versus lMCI classification with an accuracy of 70.8%, that increases by 2% the accuracy based the whole hippocampus and over 6% compared to a connectivity network-based method.
Relationship with cognitive scores
To investigate relationships between cognitive scores and MPBG values, we performed a generalized linear analysis with the following model: MPBG = β0 + β1.ages + β2.sex + β3.MMSE + β4.RAVLT + β5.FAQ + β6.CDRSB + β7.ADAS11 + β8.ADAS13. We found significant relationship of hippocampal MPBG with sex (p < 0.01), MMSE (p < 0.05) and ADAS 13 (p < 0.01). This correlation with MMSE and ADAS scores is valid for all subfields of the hippocampus. We found no specific model for a given subfield, all presented a similar pattern. These results are in line with relationships obtained between hippocampus subfields volumes and MMSE and ADAS97.
Discussion
In this work, multimodal analysis of the hippocampal subfields alterations caused by AD is proposed. First, the structural and microstructural alterations were captured from two MRI modalities with different methods. Then, the use of volume, MD, and the proposed MPBG methods were investigated to achieve this analysis. In this section, the efficiency of these different methods applied to the whole hippocampus, and each hippocampal subfield are discussed.
Whole hippocampus biomarkers
We first compared the performance of different methods applied to the whole hippocampus (see Table 2). The experiments showed that volume and average of MD of the hippocampus do not provide the most discriminating biomarkers to detect early stages of AD. Indeed, the proposed MPBG method obtains better results compared to the volume and the average of MD. However, for CN vs. AD, our MPBG method obtained lower results than T1w PBG when applied to the hippocampus. Therefore, the substantial structural differences between these two populations seem to be better captured using T1w modality. This probably comes from the better native resolution of this modality. On the other hand, for eMCI vs. lMCI, MPBG and MD PBG obtained the best result. Therefore, the subtle alterations between both populations seem to be better captured using DTI modality. This may come from the capability of this modality to measure microstructural modifications. Finally, when applied on the whole hippocampus, our MPBG demonstrates state-of-the-art performances for AD detection and prediction hippocampus compared to recent methods (see Table 3).
These results emphasize the relevance of using more accurate biomarker, such as MPBG, to study the effectiveness of hippocampal subfields for AD detection and prediction.
Hippocampal subfield biomarkers
The main contribution of this study is the multimodal analysis of hippocampal subfields. Indeed, most of the proposed biomarkers based on the hippocampus focus only on the whole structure or study alterations of hippocampal subfields with methods that do not provide sensitive biomarkers to detect early modification caused by AD. The lack of work studying alterations of hippocampal subfields with advanced biomarkers could be explained by the fact that automatic segmentation of the hippocampal subfields is a complex task due to subtle borders dividing each area.
In this work, we compared the efficiency of diffusion MRI and multimodal patch-based biomarkers for AD detection and prediction over the hippocampal subfields. Comparisons based on MD, volume and multimodal patch-based biomarkers showed that the subiculum is the most discriminating structure in the earliest stage of AD providing the best results for AD prediction (see Figs 4 and 5). However, whole hippocampus structure, followed by CA1SR-L-M, obtains best results for AD detection.
These results are in accordance with literature studies based on animal model and in vivo imaging combining volume and MD demonstrating that the subiculum is the earliest hippocampal region affected by AD49,50. Moreover, postmortem studies showed that hippocampal degeneration in the early stages of AD is not uniform. After the apparition of alterations in the EC, the pathology spreads to the subiculum, CA1, CA2-3 and finally the CA4 and DG subfields43,44,49,98. It is interesting to note that the results of our experiments using volume-based biomarkers are also coherent with the previous in-vivo imaging studies that analyzed the atrophy of each hippocampal subfield at the advanced stage of AD. These studies showed that CA1 is the subfield impacted with the most severe atrophy45,46,99,100. Furthermore, studies using the ultra-high field at 7T, enabling CA1 layers discrimination showed that CA1SR-L-M are the subfields showing the greatest atrophy at advanced stages of AD47,48.
Comparison with state-of-the-art methods
In the past years, a large number of studies dedicated to automatic detection of Alzheimer’s disease have been proposed53,69,93,101. For a fair comparison, we consider only methods based on similar modalities and validated on the same ADNI2 dataset. Direct comparison with other monomodal methods applied on ADNI1 is difficult because group definition and pathological status definition are different. However, we can observe that the results obtained by the proposed method are in line with recently published results for AD vs. CN102.
Strengths and limitations
The major strength of our work comes from studying the effectiveness of using multimodal hippocampal subfields alterations for AD classification with a novel multi-modal patch-based grading framework. Nonetheless, we acknowledge that our multi-modal framework is not without potential limitations. The main limitation is the large voxel size of DWI in native space that is prone to PVE by merging signal from CSF with the signal from brain tissues. This results in an increase of MD coefficients, especially for structures with severe atrophies. However, to limit this aspect, we corrected the PVE83. Indeed, it has been shown that the use of up-sampling methods over individual DWI direction enables reduction of the PVE effect. Nevertheless, this study does not aim to provide an interpretation of DTI parameters modification, but to study the effectiveness of the use of hippocampal subfields for AD classification with multimodal patch-based grading method. Finally, although our method extracts patches independently from both s-MRI and d-MRI modalities to estimate grading maps from both modalities, the fusion of the two grading maps requires accurate alignment of images from each modality. Consequently, the correction of EPI distortions is crucial in ensuring that each voxel corresponds to the location.
Conclusion
In this paper, we analyzed hippocampal subfield alterations with a multimodal framework based on structural and diffusion MRI. In addition, to study tenuous modifications occurring in each hippocampal subfield, we developed a new multimodal patch-based framework using T1w and DTI. Our novel MPBG method was compared to the volume and the average of MD over the whole hippocampus. This comparison demonstrated that our MPBG method improves performances for AD detection and prediction. Also, a comparison with state-of-the-art diffusion-based methods showed the competitive performance of MPBG biomarkers. Finally, volume, average MD and MBPG methods were used to analyze hippocampal subfields. Although CA1 is the subfields with the greater atrophy in the late stage of AD, the experiments demonstrated that the whole hippocampus provides the best biomarker for AD detection while the subiculum provides the best biomarker for AD prediction.
Data Availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on reasonable request.
Change history
30 June 2020
An amendment to this paper has been published and can be accessed via a link at the top of the paper.
References
Petersen, R. C. et al. Current concepts in mild cognitive impairment. Archives of neurology 58, 1985–1992 (2001).
Aisen, P. S. et al. Clinical core of the Alzheimer’s Disease Neuroimaging Initiative: progress and plans. Alzheimer’s and Dementia 6, 239–246 (2010).
Bron, E. E. et al. Standardized evaluation of algorithms for computer-aided diagnosis of dementia based on structural MRI: The CADDementia challenge. NeuroImage 111, 562–579 (2015).
Hyman, B. T., Van Hoesen, G. W., Damasio, A. R. & Barnes, C. L. Alzheimer’s disease: cell-specific pathology isolates the hippocampal formation. Science 225, 1168–1170 (1984).
West, M. J., Coleman, P. D., Flood, D. G. & Troncoso, J. C. Differences in the pattern of hippocampal neuronal loss in normal ageing and Alzheimer’s disease. The Lancet 344, 769–772 (1994).
Braak, H. & Braak, E. Staging of Alzheimer’s disease-related neurofibrillary changes. Neurobiology of aging 16, 271–278 (1995).
Gómez-Isla, T. et al. Profound loss of layer ii entorhinal cortex neurons occurs in very mild Alzheimer’s disease. Journal of Neuroscience 16, 4491–4500 (1996).
Du, A. et al. Magnetic resonance imaging of the entorhinal cortex and hippocampus in mild cognitive impairment and Alzheimer’s disease. Journal of Neurology, Neurosurgery & Psychiatry 71, 441–447 (2001).
Coupé, P. et al. Scoring by nonlocal image patch estimator for early detection of Alzheimer’s disease. NeuroImage: clinical 1, 141–152 (2012).
Jack, C. R. et al. Medial temporal atrophy on MRI in normal aging and very mild Alzheimer’s disease. Neurology 49, 786–794 (1997).
Ross, S. et al. Progressive biparietal atrophy: an atypical presentation of Alzheimer’s disease. Journal of Neurology, Neurosurgery & Psychiatry 61, 388–395 (1996).
Kaida, K.-I., Takeda, K., Nagata, N. & Kamakura, K. Alzheimer’s disease with asymmetricx parietal lobe atrophy: a case report. Journal of the neurological sciences 160, 96–99 (1998).
Jack, C. R., Petersen, R. C., O’brien, P. C. & Tangalos, E. G. Mr-based hippocampal volumetry in the diagnosis of Alzheimer’s disease. Neurology 42, 183–183 (1992).
Jack, C. R. et al. Hypothetical model of dynamic biomarkers of the Alzheimer’s pathological cascade. The Lancet Neurology 9, 119–128 (2010).
Scher, A. et al. Hippocampal shape analysis in Alzheimer’s disease: a population-based study. Neuroimage 36, 8–18 (2007).
Achterberg, H. C. et al. Hippocampal shape is predictive for the development of dementia in a normal, elderly population. Human brain mapping 35, 2359–2371 (2014).
Fischl, B. & Dale, A. M. Measuring the thickness of the human cerebral cortex from magnetic resonance images. Proceedings of the National Academy of Sciences 97, 11050–11055 (2000).
Eskildsen, S. F. et al. Prediction of Alzheimer’s disease in subjects with mild cognitive impairment from the ADNI cohort using patterns of cortical thinning. Neuroimage 65, 511–521 (2013).
Ashburner, J. & Friston, K. J. Voxel-based morphometry—the methods. Neuroimage 11, 805–821 (2000).
Good, C. D. et al. Automatic differentiation of anatomical patterns in the human brain: validation with studies of degenerative dementias. Neuroimage 17, 29–46 (2002).
Karas, G. et al. Global and local gray matter loss in mild cognitive impairment and Alzheimer’s disease. Neuroimage 23, 708–716 (2004).
Hirata, Y. et al. Voxel-based morphometry to discriminate early Alzheimer’s disease from controls. Neuroscience letters 382, 269–274 (2005).
Klöppel, S. et al. Automatic classification of MR scans in Alzheimer’s disease. Brain 131, 681–689 (2008).
Ferreira, L. K., Diniz, B. S., Forlenza, O. V., Busatto, G. F. & Zanetti, M. V. Neurostructural predictors of Alzheimer’s disease: a meta-analysis of VBM studies. Neurobiology of aging 32, 1733–1741 (2011).
Wolz, R. et al. Multi-method analysis of MRI images in early diagnostics of Alzheimer’s disease. PloS one 6, e25446 (2011).
Frisoni, G. B., Fox, N. C., Jack, C. R., Scheltens, P. & Thompson, P. M. The clinical use of structural MRI in Alzheimer disease. Nature Reviews Neurology 6, 67–77 (2010).
Hill, D. L. et al. Coalition against major diseases/european medicines agency biomarker qualification of hippocampal volume for enrichment of clinical trials in predementia stages of Alzheimer’s disease. Alzheimer’s & Dementia 10, 421–429 (2014).
Gerardin, E. et al. Multidimensional classification of hippocampal shape features discriminates Alzheimer’s disease and mild cognitive impairment from normal aging. Neuroimage 47, 1476–1486 (2009).
Tong, T. et al. Multiple instance learning for classification of dementia in brain MRI. Medical image analysis 18, 808–818 (2014).
Sørensen, L. et al. Differential diagnosis of mild cognitive impairment and Alzheimer’s disease using structural MRI cortical thickness, hippocampal shape, hippocampal texture, and volumetry. NeuroImage: Clinical (2016).
Liu, M., Zhang, D., Shen, D. & Alzheimer’s Disease Neuroimaging Initiative. Ensemble sparse classification of Alzheimer’s disease. NeuroImage 60, 1106–1116 (2012).
Coupé, P. et al. Detection of Alzheimer’s disease signature in MR images seven years before conversion to dementia: Toward an early individual prognosis. Human brain mapping 36, 4758–4770 (2015).
Koikkalainen, J. et al. Differential diagnosis of neurodegenerative diseases using structural MRI data. NeuroImage: Clinical 11, 435–449 (2016).
Tong, T. et al. Five-class differential diagnostics of neurodegenerative diseases using random undersampling boosting. NeuroImage: Clinical 15, 613–624 (2017).
Lorente de Nó, R. Studies on the structure of the cerebral cortex. ii. continuation of the study of the ammonic system. Journal für Psychologie und Neurologie (1934).
Yushkevich, P. A. et al. Quantitative comparison of 21 protocols for labeling hippocampal subfields and parahippocampal subregions in in vivo MRI: towards a harmonized segmentation protocol. Neuroimage 111, 526–541 (2015).
Winterburn, J. L. et al. A novel in vivo atlas of human hippocampal subfields using high-resolution 3T magnetic resonance imaging. Neuroimage 74, 254–265 (2013).
Hasselmo, M. E. The role of hippocampal regions CA3 and CA1 in matching entorhinal input with retrieval of associations between objects and context: theoretical comment on Lee et al. (2005). Behavioral Neuroscience 119, 342–345 (2005).
Acsády, L. & Káli, S. Models, structure, function: the transformation of cortical signals in the dentate gyrus. Progress in brain research 163, 577–599 (2007).
Wan, H., Aggleton, J. P. & Brown, M. W. Different contributions of the hippocampus and perirhinal cortex to recognition memory. Journal of Neuroscience 19, 1142–1148 (1999).
Nakazawa, K., McHugh, T. J., Wilson, M. A. & Tonegawa, S. Nmda receptors, place cells and hippocampal spatial memory. Nature Reviews Neuroscience 5, 361 (2004).
Hunsaker, M. R. & Kesner, R. P. Evaluating the differential roles of the dorsal dentate gyrus, dorsal ca3, and dorsal ca1 during a temporal ordering for spatial locations task. Hippocampus 18, 955–964 (2008).
Braak, E. & Braak, H. Alzheimer’s disease: transiently developing dendritic changes in pyramidal cells of sector CA1 of the ammon’s horn. Acta neuropathologica 93, 323–325 (1997).
Braak, H., Alafuzoff, I., Arzberger, T., Kretzschmar, H. & Del Tredici, K. Staging of Alzheimer disease-associated neurofibrillary pathology using paraffin sections and immunocytochemistry. Acta neuropathologica 112, 389–404 (2006).
Apostolova, L. G. et al. Conversion of mild cognitive impairment to Alzheimer disease predicted by hippocampal atrophy maps. Archives of neurology 63, 693–699 (2006).
La Joie, R. et al. Hippocampal subfield volumetry in mild cognitive impairment, Alzheimer’s disease and semantic dementia. NeuroImage: Clinical 3, 155–162 (2013).
Kerchner, G. et al. Hippocampal CA1 apical neuropil atrophy in mild Alzheimer disease visualized with 7-T MRI. Neurology 75, 1381–1387 (2010).
Kerchner, G. A. et al. Hippocampal CA1 apical neuropil atrophy and memory performance in Alzheimer’s disease. Neuroimage 63, 194–202 (2012).
Trujillo-Estrada, L. et al. Early neuronal loss and axonal/presynaptic damage is associated with accelerated amyloid-β accumulation in aβpp/ps1 Alzheimer’s disease mice subiculum. Journal of Alzheimer’s Disease 42, 521–541 (2014).
Li, Y.-D., Dong, H.-B., Xie, G.-M. & Zhang, L.-J. Discriminative analysis of mild Alzheimer’s disease and normal aging using volume of hippocampal subfields and hippocampal mean diffusivity: an in vivo magnetic resonance imaging study. American Journal of Alzheimer’s Disease & Other Dementias 28, 627–633 (2013).
Aggleton, J. P. & Christiansen, K. The subiculum: the heart of the extended hippocampal system. In Progress in brain research, vol. 219, 65–82 (Elsevier, 2015).
O’Dwyer, L. et al. Using support vector machines with multiple indices of diffusion for automated classification of mild cognitive impairment. PloS one 7, e32441 (2012).
Dyrba, M. et al. Robust automated detection of microstructural white matter degeneration in Alzheimer’s disease using machine learning classification of multicenter DTI data. PloS one 8, e64925 (2013).
Dyrba, M. et al. Predicting prodromal Alzheimer’s disease in subjects with mild cognitive impairment using machine learning classification of multimodal multicenter diffusion-tensor and magnetic resonance imaging data. Journal of Neuroimaging 25, 738–747 (2015).
Nir, T. M. et al. Effectiveness of regional DTI measures in distinguishing Alzheimer’s disease, MCI, and normal aging. NeuroImage: clinical 3, 180–195 (2013).
Wang, Z. et al. Interhemispheric functional and structural disconnection in Alzheimer’s disease: a combined resting-state fMRI and DTI study. PLoS One 10, e0126310 (2015).
Liu, Y. et al. Diffusion tensor imaging and tract-based spatial statistics in Alzheimer’s disease and mild cognitive impairment. Neurobiology of aging 32, 1558–1571 (2011).
Rose, S. E., Andrew, L. & Chalk, J. B. Gray and white matter changes in Alzheimer’s disease: a diffusion tensor imaging study. Journal of Magnetic Resonance Imaging 27, 20–26 (2008).
Wee, C.-Y. et al. Identification of MCI individuals using structural and functional connectivity networks. Neuroimage 59, 2045–2056 (2012).
Prasad, G. et al. Brain connectivity and novel network measures for Alzheimer’s disease classification. Neurobiology of aging 36, S121–S131 (2015).
Fellgiebel, A. & Yakushev, I. Diffusion tensor imaging of the hippocampus in MCI and early Alzheimer’s disease. Journal of Alzheimer’s Disease 26, 257–262 (2011).
Kantarci, K. et al. DWI predicts future progression to Alzheimer disease in amnestic mild cognitive impairment. Neurology 64, 902–904 (2005).
Müller, M. J. et al. Functional implications of hippocampal volume and diffusivity in mild cognitive impairment. Neuroimage 28, 1033–1042 (2005).
Fellgiebel, A. et al. Predicting conversion to dementia in mild cognitive impairment by volumetric and diffusivity measurements of the hippocampus. Psychiatry Research: Neuroimaging 146, 283–287 (2006).
Hett, K. et al. Patch-based DTI grading: Application to Alzheimer’s disease classification. In International Workshop on Patch-based Techniques in Medical Imaging, 76–83 (Springer, 2016).
Mak, E. et al. Multi-modal MRI investigation of volumetric and microstructural changes in the hippocampus and its subfields in mild cognitive impairment, Alzheimer’s disease, and dementia with Lewy bodies. International psychogeriatrics 29, 545–555 (2017).
Clerx, L., Visser, P. J., Verhey, F. & Aalten, P. New MRI markers for alzheimer’s disease: a meta-analysis of diffusion tensor imaging and a comparison with medial temporal lobe measurements. Journal of Alzheimer’s Disease 29, 405–429 (2012).
Cui, Y. et al. Automated detection of amnestic mild cognitive impairment in community-dwelling elderly adults: a combined spatial atrophy and white matter alteration approach. Neuroimage 59, 1209–1217 (2012).
Li, M., Qin, Y., Gao, F., Zhu, W. & He, X. Discriminative analysis of multivariate features from structural mri and diffusion tensor images. Magnetic resonance imaging 32, 1043–1051 (2014).
Jack, C. R. et al. The Alzheimer’s disease neuroimaging initiative (ADNI): MRI methods. Journal of magnetic resonance imaging 27, 685–691 (2008).
Jahanshad, N. et al. Diffusion tensor imaging in seven minutes: determining trade-offs between spatial and directional resolution. In Biomedical Imaging: From Nano to Macro, 2010 IEEE International Symposium on, 1161–1164 (IEEE, 2010).
Manjón, J. V. & Coupé, P. volbrain: An online MRI brain volumetry system. Frontiers in neuroinformatics 10 (2016).
Manjón, J. V., Coupé, P., Mart-Bonmat, L., Collins, D. L. & Robles, M. Adaptive non-local means denoising of MR images with spatially varying noise levels. Journal of Magnetic Resonance Imaging 31, 192–203 (2010).
Avants, B. B. et al. A reproducible evaluation of ANTs similarity metric performance in brain image registration. Neuroimage 54, 2033–2044 (2011).
Tustison, N. J. et al. N4ITK: improved N3 bias correction. IEEE transactions on medical imaging 29, 1310–1320 (2010).
Romero, J. E., Coupe, P. & Manjon, J. V. Hips: A new hippocampus subfield segmentation method. NeuroImage 163, 286–295 (2017).
Romero, J. E., Coupé, P. & Manjón, J. V. High resolution hippocampus subfield segmentation using multispectral multiatlas patch-based label fusion. In International Workshop on Patch-based Techniques in Medical Imaging, 117–124 (Springer, 2016).
Coupé, P., Manjón, J. V., Chamberland, M., Descoteaux, M. & Hiba, B. Collaborative patch-based super-resolution for diffusion-weighted images. NeuroImage 83, 245–261 (2013).
Manjón, J. et al. Nice: non-local intracranial cavity extraction. International Journal of Biomedical Imaging (2014).
Manjón, J. V. et al. Diffusion weighted image denoising using overcomplete local pca. PloS one 8, e73021 (2013).
Basser, P. J., Mattiello, J. & LeBihan, D. Mr diffusion tensor spectroscopy and imaging. Biophysical journal 66, 259–267 (1994).
Garyfallidis, E. et al. Dipy, a library for the analysis of diffusion MRI data. Frontiers in neuroinformatics 8, 8 (2014).
Dyrby, T. B. et al. Interpolation of diffusion weighted imaging datasets. NeuroImage 103, 202–213 (2014).
Tong, T. et al. A novel grading biomarker for the prediction of conversion from mild cognitive impairment to Alzheimer’s disease. IEEE Transactions on Biomedical Engineering 64, 155–165 (2017).
Barnes, C., Shechtman, E., Finkelstein, A. & Goldman, D. Patchmatch: A randomized correspondence algorithm for structural image editing. ACM Transactions on Graphics-TOG 28, 24 (2009).
Sutour, C., Deledalle, C.-A. & Aujol, J.-F. Adaptive regularization of the NL-means: Application to image and video denoising. IEEE Transactions on image processing 23, 3506–3521 (2014).
Whitwell, J. L., Crum, W. R., Watt, H. C. & Fox, N. C. Normalization of cerebral volumes by use of intracranial volume: implications for longitudinal quantitative MR imaging. American Journal of Neuroradiology 22, 1483–1489 (2001).
Dukart, J., Schroeter, M. L. & Mueller, K., Alzheimer’s Disease Neuroimaging Initiative. Age correction in dementia–matching to a healthy brain. PloS one 6, e22193 (2011).
Giraud, R. et al. An optimized patchmatch for multi-scale and multi-feature label fusion. NeuroImage 124, 770–782 (2016).
Zweig, M. H. & Campbell, G. Receiver-operating characteristic (roc) plots: a fundamental evaluation tool in clinical medicine. Clinical chemistry 39, 561–577 (1993).
Hochberg, Y. & Tamhane, A. Multiple comparison procedures (John Wiley, 1987).
Hett, K. et al. Adaptive fusion of texture-based grading for alzheimer’s disease classification. Computerized Medical Imaging and Graphics 70, 8–16 (2018).
Nir, T. M. et al. Diffusion weighted imaging-based maximum density path analysis and classification of alzheimer’s disease. Neurobiology of aging 36, S132–S140 (2015).
Zhan, L., Liu, Y., Zhou, J., Ye, J. & Thompson, P. M. Boosting classification accuracy of diffusion MRI derived brain networks for the subtypes of mild cognitive impairment using higher order singular value decomposition. In Biomedical Imaging (ISBI), 2015 IEEE 12th International Symposium on Biomedical Imaging, 131–135 (IEEE, 2015).
La Rocca, M., Amoroso, N., Monaco, A., Bellotti, R. & Tangaro, S. A novel approach to brain connectivity reveals early structural changes in alzheimer’s disease. Physiological Measurement (2018).
Maggipinto, T. et al. Dti measurements for alzheimer’s classification. Physics in Medicine and Biology 62, 2361 (2017).
Khan, W. et al. Automated hippocampal subfield measures as predictors of conversion from mild cognitive impairment to alzheimer’s disease in two independent cohorts. Brain topography 28, 746–759 (2015).
Thal, D. R. et al. Alzheimer-related τ-pathology in the perforant path target zone and in the hippocampal stratum oriens and radiatum correlates with onset and degree of dementia. Experimental neurology 163, 98–110 (2000).
Mueller, S. et al. Measurement of hippocampal subfields and age-related changes with high resolution MRI at 4T. Neurobiology of aging 28, 719–726 (2007).
Carlesimo, G. A. et al. Atrophy of presubiculum and subiculum is the earliest hippocampal anatomical marker of Alzheimer’s disease. Alzheimer’s & Dementia: Diagnosis, Assessment & Disease Monitoring 1, 24–32 (2015).
Oishi, K. et al. Multi-modal MRI analysis with disease-specific spatial filtering: initial testing to predict mild cognitive impairment patients who convert to alzheimer’s disease. Frontiers in neurology 2, 54 (2011).
Arbabshirani, M. R., Plis, S., Sui, J. & Calhoun, V. D. Single subject prediction of brain disorders in neuroimaging: Promises and pitfalls. NeuroImage 145, 137–165 (2017).
Acknowledgements
This study has been carried out with financial support from the French State, managed by the French National Research Agency (ANR) thanks to the funding of the project DeepvolBrain (ANR-18-CE45-0013) and in the frame of the Investments for the future Program IdEx Bordeaux, Cluster of excellence CPU and labex TRAIL (BigDataBrain ANR-10-LABX-57). The study presented in this work is a part of the thesis entitled “Multi-scale and multimodal imaging biomarkers for the early detection of Alzheimer’s disease” defended by the same author. Data collection and sharing for this project was funded by the Alzheimer’s Disease Neuroimaging Initiative (ADNI) (National Institutes of Health Grant U01 AG024904) and by the National Institute on Aging, the National Institute of Biomedical Imaging and Bioengineering, and through generous contributions from the following: AbbVie, Alzheimer’s Biogen; Bristol-Myes Squibb Company; CereSpir, Inc.; Cogstate; Eisai Inc.; Elan Pharmaceuticals, Inc.; Eli Lilly and Company; EuroImmun; F. Hoffman-La Roche Ltd. and its affiliated company Genentech, Inc.; Fujirebio; GE Healthcare; IXICO Ltd.; Janssen Pharmaceutical Research & Development LLC.; NeuroRx Research; Neurotrack Technologies; Novartis Pharmaceuticals Corporation; Pfizer Inc.; Piramal Imaging; Servier; Takeda Pharmaceutical providing funds to support ADNI clinical sites in Canada. Private sector contributions are facilitated by the Foundation for the National Institutes of Health (www.fnih.org). The grantee organization is the Northern California Institute of Research and Education, and the study is coordinated by the Alzheimer’s Therapeutic Research Institute at the University of Southern California. ADNI data are disseminated by the Laboratory for Neuro Imaging at the University of Southern California.
Author information
Authors and Affiliations
Consortia
Contributions
K.H., J.V.M., V.-T.T. and P.C. carried out the experiment and wrote the manuscript with support from T.T. and G.C. All authors reviewed the manuscript. The data used in this manuscript is obtained from Alzheimer’s Disease Neuroimaging Initiative (ADNI) database (http://adni.loni.usc.edu). As such, the investigators within the ADNI contributed to the design and implementation of ADNI and/or provided data but did not participate in analysis or writing of this report. A complete listing of ADNI investigators can be found at: http://adni.loni.usc.edu/wpcontent/uploads/how_to_apply/ADNI_Acknowledgement_List.pdf.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
A comprehensive list of consortium members appears at the end of the paper
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Hett, K., Ta, VT., Catheline, G. et al. Multimodal Hippocampal Subfield Grading For Alzheimer’s Disease Classification. Sci Rep 9, 13845 (2019). https://doi.org/10.1038/s41598-019-49970-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-49970-9
This article is cited by
-
Combining Nonlinear Features of EEG and MRI to Diagnose Alzheimer’s Disease
Annals of Data Science (2024)
-
Altered hippocampal subfield volumes in major depressive disorder with and without anhedonia
BMC Psychiatry (2023)
-
Structural connectivity mapping in human hippocampal-subfields using super-resolution hybrid diffusion imaging: a feasibility study
Neuroradiology (2022)
-
Volumetric changes within hippocampal subfields in Alzheimer’s disease continuum
Neurological Sciences (2022)
-
Author Correction: Classification and Visualization of Alzheimer’s Disease using Volumetric Convolutional Neural Network and Transfer Learning
Scientific Reports (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.