Abstract
LGMD2L is a subtype of limb-girdle muscular dystrophy (LGMD), caused by recessive mutations in ANO5, encoding anoctamin-5 (ANO5). We present the analysis of five patients with skeletal muscle weakness for whom heterozygous mutations within ANO5 were identified by whole exome sequencing (WES). Patients varied in the age of the disease onset (from 22 to 38 years) and severity of the morphological and clinical phenotypes. Out of the nine detected mutations one was novel (missense p.Lys132Met, accompanied by p.His841Asp) and one was not yet characterized in the literature (nonsense, p.Trp401Ter, accompanied by p.Asp81Gly). The p.Asp81Gly mutation was also identified in another patient carrying a p.Arg758Cys mutation as well. Also, a c.191dupA frameshift (p.Asn64LysfsTer15), the first described and common mutation was identified. Mutations were predicted by in silico tools to have damaging effects and are likely pathogenic according to criteria of the American College of Medical Genetics and Genomics (ACMG). Indeed, molecular modeling of mutations revealed substantial changes in ANO5 conformation that could affect the protein structure and function. In addition, variants in other genes associated with muscle pathology were identified, possibly affecting the disease progress. The presented data indicate that the identified ANO5 mutations contribute to the observed muscle pathology and broaden the genetic spectrum of LGMD myopathies.
Similar content being viewed by others
Introduction
Limb-girdle muscular dystrophies (LGMD) form a group of inherited myopathies showing progressive limb-girdle weakness with sparing of the facial and distal muscles. According to the inheritance mode, LGMD is classified into two general categories: autosomal dominant (LGMD1) and autosomal recessive (LGMD2) forms. So far, 33 genes have been associated with the disease, nine dominant and 26 recessive1,2,3. Molecular pathophysiology and therefore clinical phenotypes of the LGMDs are highly heterogeneous as the mechanisms of disease include defects in numerous proteins1,4. The observation that mutations within the same gene could be associated with variable phenotypes was made for numerous LGMD associated genes, including desmin and dysferlin1,4,5,6,7,8. It is noteworthy that variable symptoms are often observed even within a family with the same causative mutation, indicating that a primary pathogenic variant combined with potentially modifying variants may exacerbate or mitigate the phenotype (for example9).
The LGMD2L subtype is associated with mutations in the anoctamin-5 gene (ANO5). ANO5 is localized to chr11p14.3, spans 90,192 bp and contains 22 exons. The gene encodes anoctamin-5 (ANO5), a ~100-kDa protein composed of 913 amino acids, containing eight transmembrane domains10,11. The protein belongs to a family of at least ten proteins (ANO1-10) involved in a variety of cellular functions such as ion transport, phospholipid scrambling, and regulation of other membrane proteins12,13. While ANO1, ANO2, ANO6, ANO8, and ANO9 were found to code for calcium-activated chloride channels (CaCC), an exact function of ANO5 still remains elusive. Currently ANO5 is believed to be engaged in processes related to muscle membrane repair and ion transport12,13. ANO5 localizes to the endoplasmic reticulum (ER) where it is postulated to function as an intracellular channel14. In addition, ANO5 is highly expressed in cardiac and skeletal muscle as well as in chondrocytes and osteoblasts, suggesting its important role(s) in the musculoskeletal system15.
The still-expanding spectrum of ANO5-related clinical phenotypes is highly variable. Although the majority of known mutations are recessive and cause muscle disease, rare autosomal dominant ANO5 mutations cause a totally different clinical entity, gnathodiaphyseal dysplasia (GDD)14,15,16. Moreover, the recent study linked dental tumor (giant cementoma) to an autosomal dominant mutation in ANO5 associated with gnathodiaphyseal dysplasia17. The functional differences between dominant and recessive ANO5 mutations are not yet fully understood although there is a report indicating that homodimer formation or interaction with other anoctamin isoforms could be considered17. The muscle phenotype ranges from asymptomatic hyperCKemia and exercise-induced myalgia to proximal and/or distal weakness. The most typical presentation is LGMD2L, diagnosed world-wide and is one of the most frequent in Northern Europe encompassing 10–20% of LGMD cases6,9,18. Miyoshi-like disease or Miyoshi muscular dystrophy 3 (MMD3), also caused by recessive mutations in ANO5, is less common19.
LGMD2L is characterized generally by asymmetric muscle involvement with prevalent quadriceps, biceps brachii atrophy, and pain following exercise. The disease onset is in adulthood and the patients are usually ambulant6. There are numerous mutations identified within ANO5 and many of them are localized to the exons coding for the cytoplasmically-localized domains, predominantly at the 299 amino acid-long N-terminal segment9,20. The first described and most common mutation described so far is c.191dupA (p.Asn64LysfsTer15) in exon 5, most probably due to a founder effect9,17,21,22.
In the study presented herein, we address the mechanisms of clinical variability of ANO5-associated muscular dystrophies. Thorough analyses of the clinical data of five patients with heterozygous mutations within ANO5 and variable phenotypes were performed. Variants identified in two of these patients were already reported in our previous study23 without addressing the consequences of mutations on the protein structure and clinical phenotype. Out of nine identified mutations, three were not reported in databases, p.Lys132Met and p.Trp401Ter, and one, p.Tyr23Ter, was only listed in the ExAC database. None of the them was described in literature. Clinical evaluation of the patients (including MRI imaging) and morphological examination of biopsied muscles were aided by the molecular dynamics-based structural analysis of the effects of missense mutations on the structure of ANO5 dimer, the model of which was created for the purpose of this study. Our results indicate that the location of the mutation within the polypeptide chain affects the protein structure not only locally, in the vicinity of the mutated residue, but also globally, influencing the dimer interface and dynamics, and seemingly the function.
Results
Comprehensive analyses of five patients admitted to the Department of Neurology of the Medical University of Warsaw with symptoms of LGMD were carried out; mutations in ANO5 were subsequently identified for each of them. The analyses included clinical and electrophysiological evaluation, morphological examination of biopsied muscles, MRI of muscles, and genetic analyses including whole exome sequencing (WES). In addition, extensive molecular modeling analyses were performed in order to evaluate the potential effects of the identified mutations on the ANO5 dimer structure and hence function.
Clinical evaluation
A clinical summary of the examined patients (numbered consecutively 1–5) is presented in Table 1. Pedigrees of the probands’ families as well as detailed case descriptions are presented in Supplementary Material 1 (parts I and II, respectively). Except for patient 1, family members of other patients were not affected.
Clinical characteristics of all examined patients fulfilled the criteria of limb-girdle muscular dystrophy. All patients presented with mild, slowly progressive muscles weakness and/or atrophy, more prominent in the proximal muscles of the lower legs. High creatine kinase (CK) levels (up to 50 times the upper limit), asymmetric weakness and myalgia were observed in all cases. The age of onset was in the third or fourth decade, however, the first symptoms such as gait disturbances, myalgia, muscle cramps, exercise intolerance or asymmetric weakness (of one leg) were variable.
Patients 2 and 3 (males) presented with mild generalized muscle hypertrophy. In two cases (patients 1 and 5), the posterior compartment of the lower leg muscles was substantially involved. On EMG, myopathic changes were detected, however, in two patients (3 and 4) EMG was normal.
Magnetic Resonance Imaging (MRI)
MRI examinations were performed for patients 2–5 and are presented in Fig. 1 and Supplementary Material 2. Patient 1 is deceased and thus could not be examined (see Table 1).
Magnetic Resonance Imaging (MRI). (A–D) Selected axial T1W images of muscles of patients 2–5. (A) Images of patient 2 thigh muscles - arrow points to fatty atrophy in right adductor longus muscle; asterisk marks biopsied left vastus lateralis muscle with a mild fatty degeneration (arrowhead). (B) Images of patient 3 thigh muscles - arrow points to severe fatty atrophy in left adductor longus muscle; asterisk marks biopsied left vastus lateralis muscle. (C) Images of patient 4 muscles; upper panel - thigh muscles with biopsied left vastus lateralis muscle (asterisk); lower panel - arrows point to fatty atrophy in medial head gastrocnemius muscles. (D) Images of patient 5 muscles; upper panel - arrows point to fatty infiltrations in biceps brachii muscles, asterisk marks biopsied left biceps brachii muscle; lower panel - arrows point to severe bilateral fatty infiltration of the soleus and gastrocnemius muscles.
The analysis revealed that patient 2 presented with very severe, symmetric atrophy of the medial head of gastrocnemius muscle as opposed to other patients who showed muscle fatty replacement with little volume loss or even relative hypertrophy (for details see Supplementary Material 2). Asymmetric changes and fat infiltration were visible in the adductor longus muscle at the thigh level (Fig. 1A, arrow), similar to patient 3 (see Fig. 1B). However, there was no decrease in muscle bulk. The biopsied left vastus lateralis muscle showed only mild fatty degeneration (Fig. 1A, asterisk and arrowhead; and Supplementary Material 2).
In patient 3, muscle fatty degeneration changes in lumbar paraspinal muscles and lower limbs muscles with little or no change in the muscle bulk were seen (for details see Supplementary Material 2). At the pelvic level, the gluteus minimus muscle was bilaterally severely affected, more than the gluteus medius and maximus muscles. Asymmetric muscle changes and fatty degeneration were visible at the thigh level in the adductor longus muscle and all heads of the vastus muscle (Fig. 1B, arrow). Mild to moderate symmetric changes were observed in the adductor magnus muscle and hamstrings. In the lower legs, diffuse and severe changes were present in the medial head of gastrocnemius muscle and less severe in the lateral head of the gastrocnemius muscle with relative sparing of other muscles (Supplementary Material 2). The biopsied left vastus lateralis muscle (Fig. 1B, asteriks) showed only very mild fatty changes.
Patient 4 presented with mild fatty degeneration changes in the following muscles of the lower extremities: the gluteus muscles, the adductor magnus muscle and hamstrings (see Supplementary Material 2). Moderate fatty degeneration were visible in the medial head of the gastrocnemius muscle (Fig. 1C, lower panel, arrows). Also, MRI showed little decrease in the volume of the affected muscles. The biopsied left vastus lateralis muscle showed no fatty degeneration (Fig. 1C, upper panel, asterisk).
Patient 5 was the only patient in this study who showed abnormalities in the upper limbs with severe fatty degeneration and atrophy of the biopsied left biceps brachii muscle (Fig. 1D, upper panel, asterisk and arrow, and Supplementary Material 2). Changes in paraspinal muscles were more severe than those in patient 3. At the thigh level, the gluteus minimus and medius, adductor magnus and hamstrings were bilaterally severely affected with decreased muscle bulk (Supplementary Material 2). Also, some fatty degeneration changes were seen in the vastus muscles, similarly to those in patient 3 (Supplementary Material 2). In the lower legs, the patient presented with severe or complete fatty infiltration and relative hypertrophy of the medial head of the gastrocnemius muscle (Fig. 1D, lower panel, arrows), the soleus muscle, and the flexor hallucis longus muscle (Supplementary Material 2).
Moreover, in all the patients edema-like changes of the affected muscles (being a hallmark of ongoing disease activity in the muscles24), manifesting as increased signal in STIR (Short Tau Inversion Recovery) sequences,were seen (Supplementary Material 2).
Morphological analysis of biopsied muscles
Hematoxylin/eosin (HE) staining of muscle transverse sections of the patients’ biopsies revealed variable primary myopathic changes (Fig. 2 and Table 2). The changes were either mild as in the case of patient 2 or very severe as in the case of patient 1 (Fig. 2). Variability in the fiber size was observed in all the biopsied muscles and centrally positioned myonuclei were visible as well. Also, variable changes in stainings for the alkaline ATPase and NADH activities were visible, indicating a different state of muscle degeneration of biopsied muscle. The summary of the described morphological changes is presented in Table 2.
Genetic analyses
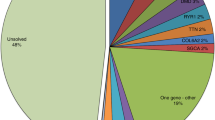
WES identified on average 66,873 genetic variants per patient, of which 45,156 off-target (defined as intergenic or intronic, but not affecting splice sites) were removed from further consideration. In total variants in 1,153 genes were further analyzed. In every filtering/prioritization approach mutations in ANO5 were ranked first on the results lists. The identified changes are rare, with a minor allele frequency below 0.1% (see Table 3). According to our ANO5 model that has been created for the purpose of this study, Tyr23, Asn64, p.Asp81 and Lys132 residues are situated within the cytoplasmic domains, Arg758 is facing the extracellular matrix, while the other residues (Trp401, Ser555, Tyr 671 and His841) are located in the central part of the molecule, mostly within the transmembrane α-helices (Figs 3 and 4). The variants were deemed pathogenic by at least one prediction program: Mutation Taster, Polyphen2 PROVEAN or SIFT, and were absent in our in-house database of 110 Polish exomes/genomes (see Table 3)25,26,27,28. In addition, His841Asp was predicted by dbscSNV as potentially altering splicing with both its adaptive boosting (ada_score) and random forest (rf_score) scores larger than 0.629. All these variants were individually assessed by a board of geneticists and clinicians according to the guidelines of the American College of Medical Genetics and Genomics30. Also, several other accompanying mutations in muscle genes have been identified. Unfortunately, segregation analysis could not be performed because of the unavailability of the DNA samples of the relatives. A detailed list of accompanying mutations for patients 1 and 2 was presented in23, and for patients 3–5 is presented in Supplementary Material 3.
Local effects of mutations on ANO5 structure. Most occupied hydrogen bonds and hydrophobic interactions (green and light purple dashed lines, respectively) for the wild-type (left panels) and mutant sites (right panels) at positions: p.Asp81Gly (A,B), p.Lys132Met (C,D), p.Ser555Ile (E,F), p.Tyr671Cys (G,H), p.Arg758Cys (I,J) and p.His841Asp (K,L). The mutated residues at positions 81, 132, 555, 671, 758 and 841 are shown in magenta, while all residues interacting with them are shown in blue. All remaining parts of the protein are shown as line ribbons and colored according to secondary structure assignment (α-helices in brown and β structures in green). Hydrogen bonds occurring in the trajectory but not in the average structures lack the dashed lines. Their appearance in the figure consists solely in the presence of both residue 81, 132, 555, 671, 758 or 841 (magenta) and its interacting partner (blue).
Global effects of mutations on ANO5 structure. (A,B) Models of ANO5 monomer and dimer, respectively. Residues undergoing missense and nonsense mutations are shown as yellow and purple sticks, respectively. (A) Flat ribbon representation of the ANO5 monomer. For space limitation, the residues are presented in the single-letter code. C- and N-termini are shown as olive CPK spheres, respectively. The α-helices A1 (residues 609–639) and A2 (residues 692–715) are shown in red. Cytoplasmic and extracellular domains are facing the top and bottom of the figure, respectively. (B) Model of ANO5 dimer in the tube representation with location of the mutated residues. Subunits A and B are colored in blue and orange, respectively, and residues forming dimerization interface are shown as spheres and color-coded as above. For space limitation, the residues are presented in the single-letter code. (C–I) Effects of mutations on the dimer interface. Left panels, close view of the central α-helices at the dimerization interface of the wild-type (C) and mutants (D–I) ANO5 structures. (D) p.Asp81Gly, (E) p.Lys132Met, (F) p.Ser555Ile, (G) p.Tyr671Cys, (H) p.Arg758C, (I) p.His841Asp. Helix A1, blue; helix A2, cyan, helix B1, red; and helix B2, orange. Subunits A and B are colored in green and straw, respectively. Hydrophobic interactions and hydrogen bonds between helices from subunits A and B are shown as light violet and green dashed lines, respectively. Right panels, concerted motions associated with the first eigenvectors (PC1) in PCA-2 for the four central α-helices: A1 (blue), A2 (cyan), B1 (red) and B2 (orange). The actual frames represent the average structures from the first mode of motion. Residues forming the helices are shown as C-α van der Waals spheres. Motions are represented by green arrows indicating the direction of motion and its amplitude.
Molecular modeling
To predict the impact of identified mutations on the ANO5 structure and hence function, we created a de novo model of ANO5 and its dimer (see Materials and Methods section, and Fig. 4A,B). The models were subjected to molecular dynamics (MD) analyses that enabled visualization of both local and global effects of the identified mutations (Figs 3–5 and Supplementary Materials 4–7, and Table 4). The analyses were not performed for the nonsense mutations (p.Trp401Ter, p. Tyr23Ter) and a frameshift p.Asn64LysfsTer15 mutation, which de facto also introduced the stop codon.
Eigenvector analysis. Plot of eigenvalues corresponding to the eigenvector index for the first 10 modes of motion in the PCA-1 (in (A)) or PCA-2 (in (B)) analyses carried out for the wild-type and mutant ANO5 dimer structures. (C), time development of projections of 2 features (distance-dependent functions) obtained from PCA-3 analysis of the central α-helices at the dimerization interface in the wild-type and mutant ANO5 structures. Color legend in (C) ANO5, black; p.Asp81Gly, red; p.Lys132Met, green; p.Ser555Ile, blue; p.Tyr671Cys, cyan; p.Arg758Cys, magenta; and p.His841Asp, orange.
Local mutation effects
Local effects of the identified mutations were analyzed by investigating the conformational rearrangements around the site of mutation (Fig. 3A–L). To this end, hydrogen bonding and hydrophobic contacts throughout the simulations were accounted. The results indicate that two mutations to cysteine, p.Tyr671Cys and p.Arg758Cys, thoroughly change a role of mutated residues in local interaction networks (Fig. 3G–J). In particular, p.Arg758Cys mutation, the site of which is located on a distant-from-the-interface surface loop 753–822 (Figs 3I,J and 4B), caused the relatively short residue of Cys758 to lose almost entirely several hydrogen contacts with residues from the same loop (especially with Glu816, His821 and Asn822) and with Met823 from a short helical segment 823–825. All of them were maintained by the long and strongly polar residue of arginine 758. Instead, Cys758 maintains a strong H-bond contact with Gln824 and a weak contact with Asn817. In addition, positions of Arg758 and Cys758 are stabilized on a similar level (although more persistently in the case of Arg758) due to hydrophobic contacts with Ile756 and Met823 (to Arg 758) and Val760 and Tyr761 (to Cys758) (Fig. 3I,J, Supplementary Material 4: Tables 8 and 9). Furthermore, there is a weakened stabilization of the mutated residue of Cys671 compared to the wild-type residue of Tyr671 (Fig. 3G,H). This site is located in α-helix 669–684 in the core of each subunit ~15 Å from the dimerization interface (Figs 3G and 4A,B). In the wild-type structure, Tyr671 maintains medium persistent H-bonds with Glu675 from the same α-helix and with Gly668 from a short loop 664–668 as well as strong H-bond with Arg234 from a surrounding loop 222–237 (Fig. 3G). In the p.Tyr671Cys mutant structure, Cys671 loses the H-bonds with Glu675 and Arg234, and only preserves the H-bond with Gly668 (Fig. 3H). In addition, Tyr671 interacts hydrophobically with Met560 from a long α-helix 549–568 (strong contact) and Lys233 from the loop 222–237 (periodic contact), compared with Cys671 loosing entirely the former and almost entirely the latter contacts (Fig. 3G,H; Supplementary material 4: Tables 6 and 7). Moreover, the replacement of tyrosine with cysteine within the 669–684 α-helix perturbs the symmetry of the N-terminal end of this α-helix (residues 669–674), resulting in the disruption of periodic hydrogen bonding and subsequently weakened, less persistent local α-helical structure (see secondary structure assignment for this α-helix in the wild type and Tyr671Cys mutant in Supplementary Material 5).
However a different picture can be seen in the case of the p.His841Asp mutation, which is located in the surface α-helix 834–849, nearby the site of the p.Arg758Cys mutation (Fig. 4A,B). In the mutant structure, Asp841 is coordinated with hydrogen bonds more persistently than His841 in the wild-type structure(Fig. 3K,L). However, the imidazole side chain of His841 is engaged in several hydrophobic contacts, including with Phe844 and Leu845 from the same α-helix and with Arg364 from a surrounding short surface loop 232–234 (Fig. 3K). Such a stabilization cannot be reached upon mutation to a polar residue of aspartate and hence the p.His841Asp mutant loses all the mentioned hydrophobic contacts, rendering histidine (His841 in the wild-type structure) as an overall better stabilized residue (Fig. 3K,L, Supplementary Material 4: Tables 10 and 11).
The other two mutants, p.Asp81Gly (Fig. 3A,B) and p.Ser555Ile (Fig. 3E,F), show less distinct rearrangements, accounted for as differences in local interaction networks with respect to the wild-type structure. Asp81, located at the dimerization interface in the loop 71–83 (Fig. 4B), is strongly saturated with H-bonds to Arg78; these interactions appear to be crucial in stabilizing the position of Asp81 in the wild-type structure. Nevertheless, other H-bond contacts of Asp81 are very limited and even less persistent than the hydrogen bonds maintained in the mutant structure by Gly81 (Fig. 3A,B, Supplementary Material 4: Table 1). Ser555, located in the core α-helix 549–568 (Fig. 4A), maintains a very strong H-bond with neighboring Glu544 and two much less persistent H-bonds with Glu551 and Lys559 (Fig. 3E), compared to the lack of the Glu544 contact and more persistent contacts with Glu551 and Lys559, maintained in the p.Ser555Ile mutant structure by Ile555 (Fig. 3F). In addition, the aliphatic side chain of Ile555 is mildly engaged in hydrophobic interaction with the pyrrolidine side chain of Pro546, whereas Ser555 lacks the hydrophobic stabilization at all (Fig. 3E,F, Supplementary Material 4: Tables 4 and 5).
Contrary to the other analyzed mutations, the local effect of the p.Lys132Met mutation, located close to the dimerization interface at the C-terminal end of the β-strand 126–132 (Fig. 4B), is neither unfavorable nor at most neutral as it seems to stabilize the local protein structure. This can be observed in the larger occupations of both hydrogen bonds and especially hydrophobic interactions maintained by Met132 in comparison to Lys132 (Fig. 3C,D). Met132 forms, both as a donor and acceptor, persistent H-bonds with Leu115 from a surrounding loop 106–115. Moreover, there is a strong hydrophobic contact between side chains of Met132 and Leu115 (Fig. 3C,D). Conversely, Leu132 interacts with Leu115 less strongly, forming with it a less persistent H-bond (only as a donor) and maintaining a much less persistent hydrophobic contact (Fig. 3C). Besides, unlike Leu132, Met132 maintains yet another stabilizing interaction, i.e. a very strong hydrophobic contact with Ala135 from a surrounding short loop 133–136 (Fig. 3C,D, Supplementary Material 4: Tables 2 and 3).
Global mutation effects
Stability of the dimerization interface in view of the effects on central α-helices: Parameters from the PDBePISA evaluation of dimerization interfaces in the average structures from the stable parts of trajectories of the wild-type ANO5 and its six single mutants are shown in Table 4. There is a diversity of the calculated interface surfaces and the estimated solvation free energy gains upon formation of the interface. The ΔG P-values, indicating the P-value of the observed solvation free energy gain, are either below or above 0.5. This allows the distinction between hydrophobic, interaction-specific interfaces, observed for wild type ANO5 and its mutants: p.Tyr671Cys, p.His841Asp and p.Lys132Met (ΔG P-values below 0.5), from interfaces that are more “random”, i.e. less hydrophobic than they optimally could be, observed for ANO5 mutants p.Asp81Gly, p.Ser555Ile and p.Arg758Cys (ΔG P-values significantly above 0.5). Therefore, the ΔG P-value is an important overall indication of the differences in hydrophobic character of the dimerization interfaces in the modeled sample.
The dimerization interface in the ANO5 structure and its mutants is composed of four central α-helices. Each subunit contributes two α-helices [609–639 (A1) and 692–715 (A2) in subunit A and, symmetrically, 1523–1552 (B1) and 1605–1628 (B2) in subunit B] together with surrounding loops and a few patches from other helices (see Fig. 4A,B). As expected, based on their buried location in the protein, these central α-helices are of mostly hydrophobic character. The conservation of this hydrophobic core during MD simulations can be evaluated by counting up the hydrophobic contacts present in the trajectories and in the average structures between hydrophobic residues from each of the central α-helices (Supplementary Material 4: Table 12, and Supplementary Material 5). These results are in accordance with the PDBePISA ΔG P-values. The highest number of contacts between subunits (20) was predicted for the wild type protein with 12 contacts between subunits A and B of the dimer (Supplementary Material 4: Table 12; Fig. 4C, left panel). The low and thus disadvantageous number of contacts is calculated for the p.Asp81Gly and p.Arg758Cys mutants with five and four contacts, respectively, and only with three contacts between residues in subunits A and B of both mutants (Supplementary Material 4: Table 12 and Fig. 4D,H, left panels). The higher number of contacts, 15, is predicted for the p.Ser555Ile mutant with nine intersubunit contacts (Supplementary Material 4: Table 12 and Fig. 4F, left panel). Similar values were calculated for other mutants with 17(6) for p.Lys132Met, 14(9) for p.His841Asp and 14(7) for p.Tyr671Cys (numbers in parenthesis reflect the intersubunit contacts) (Supplementary Material 4: Table 12; Fig. 4E,H,I, left panels). The percentage of trajectory occupancies of all hydrophobic contacts between residues of the four central α-helices of the analyzed structures are listed in Supplementary Material 4: Table 12. Moreover, only the wild-type protein shows the contacts from the A1 α-helix distributed equally to the B1 and B2 α-helices, whereas in all the mutant structures the contacts from the A1 α-helix reach exclusively the B1 α-helix, putting the B2 α-helix totally aside. It means the larger segment of interface stabilization, embracing the A1, B1 and B2 central α-helices, to be present only in the wild-type structure, in contrast to all mutant structures contributing to the stabilization pool only the A1-B1 α-helices’ interactions. Furthermore, in the trajectory of the wild-type structure there are two strong H-bonds linking the A1 α-helix with the B1 α-helices (Tyr632-Ile1540, occupied at 71.9% of the analyzed trajectory) and B2 (Ile627-Trp1623, occupied at 57.4%), which seem to consolidate even further the dimerization interface. On the contrary, in their own trajectories, the mutant structures lack similarly persistent H-bonds at the central helices. Besides, all H-bonds occupied above 10% of the mutants’ trajectories (between Asn626-Asn1550, occupied at 45.2% in the p.Asp81Gly mutant; between Trp638-Thr1532, occupied at 31.7% in the p. His841Asp mutant; and between Trp638-Thr1532, occupied 27.5% in p.Lys132Met) are formed solely between the A1 and B1 α-helices. Unlike in the wild-type protein, the B2 α-helix is not engaged in all those H-bonds (Fig. 4C–I and Supplementary Material 4: Table 13).
The secondary structure content throughout the simulations (see Supplementary Material 5) shows that the p.Asp81Gly and, to a lesser extent, p.His841Asp mutations perturb the central helices. This results in their accumulated average α-helical assignments of 70.4% and 74.1%, respectively, compared to 78.7% for the wild-type structure. The values for the p.Tyr671Cys, p.Ser555Ile and p.Lys132Met mutants were similar to that of the wild type and were 76.7%, 77.8% and 77.9%, respectively. Interestingly, the p.Arg758Cys mutation even strengthened the α-helical assignment of the four central helices, bringing it to an accumulated average of 85.2% (Supplementary Material 5).
Essential Dynamics: Essential dynamics was performed to enable identification of concerted motions that are often fundamental for protein functions31,32,33. In the present work, three Principal Component Analyses (PCA) were performed: first based on the decomposition of the global conformational space (PCA-1), second on the conformational space explored by four central helices (PCA-2), and the third on the distances between the central helices (PCA-3) (Figs 4C–I, right panels, and 5, as well as Supplementary Materials 6 and 7).
The eigenvalues against the corresponding eigenvector index for the first 10 modes of motion are plotted for the PCA-1 and PCA-2 analyses (Fig. 5A,B, respectively). It demonstrates that only few modes of motion are required to explain the majority of variance in the motions. Indeed, the first eigenvectors (PC1) alone explain 29.5–37.6% or 13.4–40.6% of the total variance in the motions observed in PCA-1 or PCA-2, respectively. Together with the second (PC2) as well as the second (PC2) and third eigenvector (PC3) the explanatory rate increases to 41.0–50.3% and 30.5–62.6%, for PCA-1 and PCA-2, respectively. The correlation coefficients (r2), computed from the time evolution of trajectories projected on the first eigenvector and plotted for each pair composed of the wild-type protein and mutant, show the motions associated with PC1 being either analogous (ANO5 versus p.Ser555Ile and ANO5 versus p.His841Asp) or strongly analogous between the wild-type protein and the mutant systems in PCA-1. On the other hand, r2 are very poorly correlated between ANO5 and its mutants in PCA-2 (Supplementary Materials 6 and 7). It demonstrates that while the motions are similar on the global scale, the local motions pertaining to the central helices are not similar at all, i.e. they do not repeat the same pattern.
Investigating further the eigenvalues against eigenvector index plots in Fig. 5B, a very shallow PCA-2 profile can be observed for ANO5, indicating the concerted motions associated with different eigenvectors in ANO5 to be of similar explanatory relevance. In addition, the PC1 motions in mutant structures have slightly (p.Ser555Ile) through moderately to largely (p.His841Asp and p.Lys132Met) higher amplitudes compared to the PC1 motion in ANO5. It indicates that the central helices in mutated structures undergo larger PC1 motions compared to the wild-type protein (see Fig. 4C–I, right panels, Supplementary Materials 6 and 7). It points to supposedly destabilizing effects of the mutations, as can be concluded also from the comparison of the plots depicting the PCA-2 motions associated with PC1 (Fig. 5 and Supplementary Material 7). The consolidated position and moderate motions of the central helices in the average structure from the PC1 motion mode of ANO5, compared with less ordered position and more “random” motions of those helices in all mutant structures are noticed there. Moreover, the “subtle” PCA-2 motions associated with PC2 and PC3 in ANO5 (see Supplementary Materials 6 and 7), worth of considering due to similar explanatory relevance of the first three modes of motion of ANO5, do not overrule the picture emerged from the comparison of the motions associated with PC1.
An incisive dependence pattern of the amplitudes (eigenvalues) associated with eigenvectors, observed in PCA-2, does not occur in PCA-1, where the amplitudes of the PC1 motions are less mutually different and the eigenvalue associated with PC1 in ANO5 is not the smallest one in the series (Fig. 5A,B, and Supplementary Materials 6 and 7). Nevertheless, the concerted motions associated with PC1 in ANO5 appear to be the most ordered in the series, with few surface segments and the central loop 1409–1424 from subunit B moving while the rest of the structure remaining virtually still (Supplementary Material 7). Overall, all mutants appear to be more “restless”, with concerted motions involving large parts of their structures, which is especially evident in the p.Tyr671Cys, p.Lys132Met, p.R758C and p.His841Asp mutants (see Supplementary Material 7). But given that, the results of PCA-1 do not seem to provide a clear message on the differences potentially destabilizing the mutant structures compared to the wild-type protein.
The PCA-3 analysis can be considered as complementary to PCA-2. The two obtained features were able to explain ~90% of variance in the dataset. Results plotted on Fig. 5C show the time development of projections of the functions characterizing distances between the central α-helices along PC1 vs PC2. Figure 5C presents a separate localization and concentrated appearance of the wild-type protein (black) and different localizations and more dispersed appearances of all mutant structures, though less so of the p.Ser555Ile mutant (blue). It suggests (and possibly confirms) the stable behavior of the central α-helices in the wild-type structure in contrast to less stable/more perturbed behaviors of those helices and, in turn, of the dimerization interface, in mutant structures. In particular, the p.Asp81Gly and p.His841Lys mutants (red and orange, respectively) appear to be significantly different from all other mutants and from the wild-type protein, while mutants p.Lys132Met (green), p.Arg758Cys (magenta), p.Tyr671Cys (cyan) and p.Ser555Ile (blue) also differ from the wild-type protein but exhibit mutual similarity when described with two PCA-derived features. p.Asp81Gly and p.His841Asp mutants are the ones in the series with weakened α-helical secondary structure assignments within the central α-helices. Besides, A1 and A2 α-helices in the p.His841Asp mutant are found wandering quite a lot during the first mode of motion (see Supplementary Material 7), the one explaining as much as 46% of the total motion variance in PCA-2. This wandering perturbs their between-helices-distances, which appears to be reflected in a very strongly dispersed appearance of His841Asp on the plot, with A1 and A2 helices either in contraction or separated (approximately two parts of the His841Asp appearance on the plot in Fig. 5C).
Discussion
We presented herein clinical and diagnostic data of five Polish patients with ANO5 heterozygous mutations and clinical diagnosis of LGMD2L. Analyses of both clinical phenotypes and muscle morphology (based on histochemistry and MRI) revealed a high degree of variability. The MRI imaging also showed a high degree of variability of changes in different muscles in the same patient, making the LGMD2L picture even more complicated.
Genetic analysis identified nine mutations, among them three: p.Tyr23Ter, p.Lys132Met and p.Trp401Ter were not yet characterized and the latter two not even registered in the gnomAD database. The other six mutations: p.Asn64LysfsTer15, p.Asp81Gly, p.Ser555I, p.Tyr671C, p.Arg758C and p.His841Asp have been already described in the European population9,21,22 as damaging, implying their contribution to the observed pathology.
The two newly characterized nonsense mutations introduce a premature stop codon and most probably lead to degradation of a truncated protein through proteasomal pathways23,31,32. The lack of the full length ANO5 protein was recently shown for the homozygous p.Asn64LysfsTer15 (c.191dupA) mutation that in fact introduce a premature stop codon (see34). In many genes (e.g. DMD, RYR1) the presence of frameshift or nonsense mutations is often associated with a severe disease course35.
To characterize the effects of missense mutations on the ANO5 structure, and possibly function, we employed molecular dynamics analyses. For this purpose, models of the monomer and dimer structure were created. The models were generally analogous to the models presented by Andreeva et al.17. However, the number of transmembrane helices depicted in our model surpasses the number declared by those authors. Unfortunately, more detailed comparison of both models is impossible due to the lack of access to the aforementioned models.
Analysis of the local effects indicated that the most visible changes were observed for p.Tyr671C and p.Arg758C mutants, both associated with the introduction of a cysteine residue, much smaller than the original arginine and tyrosine residues but also prone to form new interactions. On the other hand, p.Asp81Gly (introducing a small glycine residue) and p.Ser555Ile (introducing a hydrophobic isoleucine residue) mutations have much less impact on local conformational rearrangements. Contrary to that, the novel p.Lys132Met (introducing a similar in molecular mass but non-polar and sulphur containing methionine residue) mutation seems to rather stabilize (stiffen) the local protein structure. Interestingly, Asp81 and Tyr671 as well as Lys132 residues are situated either at the dimerization interface (Asp81) or in its vicinity (Tyr671 and Lys132). Thus the local effects of the mutations on the protein structure are variable and do not depend on the localization of the mutated residues within the molecule. However, they could influence the protein dimerization and so we examined the effects of the mutations on the dimerization interface.
We predict that all the analyzed mutations affect dimer formation. In particular, the Asp81Gly and Arg758Cys mutations appear to have a strong detrimental effect on the stability of the dimerization interface as shown by the high ΔG P-values, low overall numbers and lack of balanced pattern of hydrophobic contacts, lower SSC (for p.Asp81Gly) as well as lack of (for p.Arg758Cys) or weakened (for p.Asp81Gly) hydrogen bonds. More elaborate although ultimately disadvantageous seem to be the effects of the p.Ser555Ile (the high ΔG P-value, lack of balanced hydrophobic interactions pattern and lack of strong hydrogen bonds) and p.His841Asp (lower SSC, lack of balanced hydrophobic interactions pattern, weaker hydrogen bonds compared with the wild-type system) mutations. The other two mutations, p.Lys132Met and p.Tyr671Cys, exhibit both a less balanced hydrophobic interactions pattern, and the lack of (for p.Tyr671Cys) or weakened (for p.Lys132Met) hydrogen bonds, with both these features expectedly lowering the stability of the dimerization interface.
Moreover, essential dynamics analysis revealed that all analyzed mutations affect the concerted motions of the ANO5 dimer and have supposedly destabilizing effects as central α-helices of mutant structures undergo larger motions with respect to the wild type protein. All the mutants appear to be more motile (restless) with larger parts of the protein engaged in the concerted motions. In particular, this was most evident in p.Lys132Met, p.Tyr671C, p.Arg758C and in particular in p.His841Asp. Thus the results of essential dynamics highlight the differences between the wild-type protein and the mutant structures on the ground of less stable central α-helices at the dimerization interface, that in mutant proteins undergo larger and seemingly more random motions. These dynamic structural differences could therefore result in damaging/deleterious effects on the protein function.
The question arises whether and how particular mutations affect the protein conformation and influence the patients’ phenotype. Patient 1, with most severe dystrophic changes, bore a p.Trp401Ter truncation mutation and p.Asp81Gly missense mutation. The nonsense mutation introduces premature stop codon located in the proximity of the final exon-junction thus suggesting elimination of the mRNA through the nonsense mediated decay. The latter mutation was also found in patient 2 where it co-existed with a missense p.Arg758Cys mutation, which also had negative impact on the dimerization interface. Despite carrying mutations that negatively affect dimerization, patient 2 presented mild myopathic changes (in clinical and MRI examinations) but with early disease onset. Interestingly, the same pair of mutations has been already identified in a Finnish male patient with the similar disease onset and clinical phenotype20. Of note, the p.Arg758Cys mutation is considered as a founder mutation in the Finnish population36. Patient 3, bearing the novel p.Tyr23Ter truncation and known missense p.Ser555Ile mutation (with disadvantageous effect on dimer formation), also presented mild muscle involvement but with disease onset in third decade of life. Patient 4 bears a novel p.Lys132Met mutation accompanied by a known p.His841Asp mutation that was first described in a British patient as homozygous and was predicted to be pathogenic22. Our analyses showed that both mutations affect the dimerization interface with p.His841Asp being more disadvantageous. This patient, with onset in his fourth decade presented a moderate phenotype. Patient 5 carries the most common p.Asn64LysfsTer15 mutation predicted to have deleterious effect on the protein function, and p.Tyr671Cys missense mutation, which as we showed strongly affected ANO5 conformation. This patient had disease onset in her third decade and presented advanced dystrophic changes.
The complexity to the picture is aided by the fact that numerous WES analyses have revealed mutation burden in genes other than ANO5 that are also associated with LGMD phenotypes (for example9,20,23). These were depicted by prediction softwares either as benign/moderate or as deleterious/damaging and hence possibly pathogenic. In our study, such accompanying variants were also found and differed among patients. Here, we will focus on those depicted by the prediction programs as potentially most pathogenic. Patient 1 had potentially phenotype-modifying missense variants found in BAG3, coding for BAG3 (BCL2 associated athanogene 3), a protein involved in the chaperone-assisted selective autophagy (CASA) pathway37 and in FLNC, coding for filamin C (FLNC), a protein involved in actin cytoskeleton organization. Interestingly, a missense FLNC variant was also identified in patient 4. Both FLNC variants, predicted as damaging, are located within Ig-like domains, mutations in which are also often associated with cardiomyopathy phenotypes38. It is noteworthy that patient 1 had cardiac failure but so far patient 4 has not reported any heart problems. The reported herein p.Ser185Ile BAG3 variant could affect chaperone function of the protein as even a mild impairment of the function could cause abnormalities in the sarcomere structure or dysfunctional protein folding quality control mechanisms39,40. Therefore it cannot be excluded that for patient 1 the observed muscle phenotype of ANO5 could be also exacerbated by BAG3 and FLNC variants. Patient 2 carries a variant in NEB, encoding a large protein regulating the length of thin filament in the contractile apparatus. NEB mutations are associated with a wide pattern of myopathic symptoms, and are often linked with nemaline myopathies41,42. We did not find nemaline rods in patient’s 2 biopsy indicating that the effect of this NEB mutation on the clinical phenotype seems to be rather minor. Patient 3 carries a known variant in RYR1, encoding skeletal muscle-type ryanodine receptor 1 (RYR1). Over 500 mutations have been mapped to RYRs and the majority of them are implicated in human diseases such as malignant hyperthermia as well as RYR1 myopathies, previously classified as central core disease (CCD) and multiminicore disease43,44. Identified RYR1 mutation, affecting a repeat within the cytoplasmic part of the protein, is reported in the ClinVar database with a note that there are conflicting interpretations of pathogenicity. Therefore we cannot conclude whether or not this variant could contribute to the observed phenotype of patient 3. Patient 4, besides the FLNC mutation mentioned above, carries mutations in COL12A1 and LAMA2, encoding isoforms of collagen 12 and extracellular matrix glycoprotein laminin A, respectively. Mutations within both genes could have deleterious effects on the skeletal muscle organization and function as they are associated with several myopathies, including LGMD, collagenoses and merosin deficient congenital muscular dystrophy45,46. The COL12A1 variant is listed in the ClinVar database as likely benign while the one in LAMA2 has contradictory predictions from benign to deleterious. We saw an increase in endomysial connective tissue infiltrations that could result from changes in collagen structure, however, they are much smaller than the ones observed in patient 1, who did not have variants in the gene encoding this collagen isoform. Moreover, it is also possible that minimal endomysial infiltrations could be associated with effects of mutation with LAMA2. However, mutations in both genes do not seem to greatly contribute to the observed phenotype. Interestingly, variants in several other genes involved in cell adhesion, namely AGRN, encoding agrin, a protein crucial for the development of the neuromuscular junction (NMJ), ITGA7, encoding integrin subunit α7, and LAMB2, encoding laminin B2 isoform, were identified. Both AGRN and LAMB2 are associated with congenital myasthenic syndromes, associated with basal-lamina dysfunction47 and ITGA7 mutations are associated with congenital muscular dystrophy48. Identified variants in these three genes could possibly affect sarcolemma and synaptic activity at the NMJ. Thus they could to some extent explain the observed phenotype, especially the changes in muscle morphology (as examined by MRI and histochemistry).
There is also another possibility that is beyond a scope of this study that changes evoked by ANO5 mutations (and/or by mutations in other muscle-related genes) could have effect on functions of unmodified genes. It could be speculated that impairment of ANO5 might affect other anoctamin isoforms and/or proteins involved in the same processes, in which ANO5 is engaged.
In conclusion, we have described five Polish LGMD2L patients with recessive mutations in ANO5 that according to the criteria proposed by Antonarakis et al.49 and MacArthur et al.50 are highly likely to be disease-causing according to ACMG criteria (see Table 3). Our data, in frame with other recent results, indicate that recessive ANO5 mutations are a frequent cause of muscular dystrophy and are population-specific51,52,53,54. We showed that ANO5 mutations (possibly in concerto with other variants in numerous genes linked with myopathies) are associated with the observed phenotypes. The severity of the clinical and morphological muscle impairment seems to be associated with the location of the mutation, and therefore its effect on ANO5 structure and function. However, there is not a direct correlation between the effects of mutations and clinical picture, probably due to mutation burden in other muscle-related genes. Finally, our data seem to support the ongoing hypothesis that LGMD and other dystrophies/myopathies could be in fact oligogenic disorders, with few mutated genes contributing unevenly to the development of the disease23,55.
Materials and Methods
Magnetic Resonance Imaging (MRI)
The whole body MRI (WB MRI) was performed on 1.5 T scanner (Siemens Avanto) with a multi-sequence imaging protocol. T1-weighted, T2-weighted and STIR (Short Tau Inversion Recovery) sequences were obtained in coronal planes with 5 mm slice thickness while T1-weighted and STIR sequences were obtained in axial planes with 8 mm slice thickness. Muscle bulk was assessed on T1-weighted images. Muscles were scored on axial T1-weighted sequences with the semi-quantitative56 visual scale modified by Fischer et al.57 to estimate the degree of muscle degeneration. STIR sequences were used to assess the presence and symmetry of edema-like changes.
Muscle biopsy
The open muscle biopsies of the probands biceps brachii or vastus lateralis were performed, and the muscle specimens were routinely processed for further analyses.
Light microscopy
Biopsied muscles were frozen in isopentane cooled in liquid nitrogen, cut on a cryomicrotome with a slice thickness of 8 µm and stained with the routine battery of histological and histochemical methods.
Genetic analyses
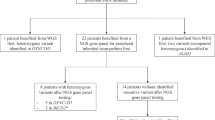
DNA was extracted from peripheral blood of the proband using standard methods58. Whole exome sequencing (WES) was performed in two institutions: samples of patients 1 and 2 were sequenced commercially at Oxford Gene Technology (Oxfordshire, United Kingdom), and samples of patients 3, 4 and 5 were sequenced at the Broad Institute’s Genomics Platform as part of the MYO-SEQ project coordinated by the John Walton Muscular Dystrophy Research Centre, Newcastle University.
Samples 1 and 2 were sequenced using SureSelect Human All Exon v4 enrichment kit and 100 bp paired-end sequencing on the Illumina HiSeq2000 platform. Fastq read files were generated from the sequencing platform via the Illumina software. The paired-end reads were aligned to the human genome build 37 (hg19) reference using Burrows-Wheeler Alignment (BWA) package59. Duplicate reads were removed with Picard and base quality Phred scores were recalibrated using Genome Analysis Toolkit (GATK) covariance recalibration60. Alignments were viewed with Integrative Genomics Viewer (IGV)61. Sequencing resulted in >80% of targets covered at least 20x and a mean target coverage was 55x. Single Nucleotide Polymorphism (SNPs) and insertions/deletions (indels) were called using the GATK Unified Genotyper Annovar for initial variant annotation62 and further annotation, filtering and analysis performed on the Galaxy platform (on PL-Grid Infrastructure).
WES for samples 3, 4 and 5 was performed at the Broad Institute’s Genomics Platform, using Illumina exome capture and 76 bp paired-end reads on the Illumina platform. Fastq read files were generated from the sequencing platform via the Illumina software. The paired-end reads were aligned to the human genome build 37 (hg19) reference using Burrows-Wheeler Alignment (BWA) package59. Duplicate reads were removed with Picard and base quality Phred scores were recalibrated using Genome Analysis Toolkit (GATK) covariance recalibration (McKenna et al.60). Alignments were viewed with IGV. Sequencing resulted in >80% of targets covered at least 20x and a mean target coverage was above 80x. SNPs and indels were called using the GATK HaplotypeCaller Default filters were applied to SNP and indel calls using the GATK Variant Quality Score Recalibration (VQSR) approach. Lastly, the variants were annotated using Variant Effect Predictor (VEP).
Filtering was based on the allele frequency in ExAC database (<3% for variants in genes already associated with LGMD, and <1% for variants in other genes), and predicted pathogenicity (predicted pathogenic by at least one out of Mutation Taster, PolyPhen2, and SIFT software). Prioritization was based on inheritance mode, on the predicted effect with truncating and elongating variants being evaluated more carefully, on predicted pathogenicity, and on known association with myopathic phenotypes reported in the Human Phenotype Ontology. Standardized phenotypic information was uploaded to PhenoTips63. Analyses were made with Exomiser2, PhenIX and Exome Walker with prioritization of variants based on possible association with Human Phenotype Ontology term “limb-girdle muscular dystrophy” and based on random-walk analysis of protein interaction networks64. ANO5 mutations and selected mutations in other genes (BAG3, FLNC, RYR1) were confirmed using direct, fluorescent sequencing (ABI 3130 Gemetic Analyzer, Applied Biosystems, USA).
Models of ANO5 and its mutants
A model of dimeric wild-type ANO5 protein was prepared in two steps: (i) the monomeric structure of ANO5 was generated using the intensive mode of Phyre2 algorithm65 based on the human ANO5 wild-type sequence (Q75V66 in the UnitProt Knowledgebase), and (ii) the monomer into dimer conversion was carried out using the M-ZDOCK server66. Modeling of both steps was performed with default settings implemented in the Phyre2 intensive mode and M-ZDOCK algorithms. 602 out of 913 residues in the ANO5 sequence (66%) were modeled by homology and rated by the Phyre2 algorithm as highly confident (above 90% of accuracy), while the rest of the structure was modeled ab initio due to the lack of respective homology motifs.
The following single mutations were incorporated in the ANO5 dimer wild-type model in order to investigate their effect on the 3D structural level: Asp81Gly, Lys132Met, Ser555Ile, Tyr671Cys, Arg758Cys and His841Asp (Fig. 4A,B). Both the wild-type structure and all mutant structures were prepared to MD simulations according to the following protocol. Hydrogen atoms were added in tleap from AmberTools1667 according to standard protonation states of the amino acids. Resulting structures were immersed in truncated octahedral boxes of water molecules (TIP3P model68) extended up to 9 Å away from the protein surfaces. Monovalent sodium ions were placed in calculated positions to keep the neutrality of the simulation boxes.
Molecular dynamics (MD)
The simulation setup was as follows. Initially, the models were subjected to mixed steepest descent and conjugate gradient energy minimizations carried out in two steps involving (i) only solvent molecules and (ii) the whole systems. Then, gradual heating to 300 K was performed in three 10 ps intervals (0 → 100 K, 100 K → 200 K and 200 K → 300 K), followed by equilibrations at 300 K lasting for 10 ns. Finally, productive MD simulations in the NpT ensemble (conserved number of atoms (N), pressure (p) at 1 atm and temperature (T) at 300 K) were performed and lasted for 130, 180 or 230 ns, depending on the simulation. Pressure and temperature were monitored by the isotropic position scaling and the weak-coupling algorithms, respectively, with both coupling times of 1 ps. Periodic boundary conditions were applied throughout simulations. Long-range electrostatic interactions were handled using the particle mesh Ewald method69 with a charge grid spacing of approx. 0.9 Å. All bonds involving hydrogen atoms were constrained using the SHAKE algorithm70 with a relative tolerance of 0.00001 Å, allowing a time step of 2 fs. Trajectories were saved at 2 ps intervals. All minimizations and MD simulations except the productive part were carried out with sander from AmberTools16 using the ff14sb force field71. Productive MD simulations were performed with pmemd using the same force field.
Trajectory analysis
Root mean square deviations (RMSD) of the backbone atoms from the collected trajectories have been calculated with cpptraj72 from AmberTools16 to assess the conformational stability of the wild-type and mutant models. Those stable parts of each trajectory which spanned at least 40 ns were used for further analysis with cpptraj, including computing average structures, hydrogen bond (HB) and hydrophobic interaction (HI) occupancies and secondary structure content. The average structures have been evaluated with the Protein interfaces, surfaces and assemblies service PISA at the European Bioinformatics Institute (http://www.ebi.ac.uk/pdbe/prot_int/pistart.html)73,74. The HB occupancies were calculated based on the following limits: 3.2 Å for the donor-acceptor distance and 120° for the donor-hydrogen-acceptor bond angle. The HI occupancies were calculated for the hydrophobic contacts between two residues in the range of 3.5–5.5 Å.
Essential dynamics (ED)
In general, the method is based on the construction of the covariance matrix of coordinate fluctuations, which is diagonalized in order to obtain the eigenvectors and eigenvalues that represent the concerted motion in the system. While the eigenvectors indicate the directions of motion, the corresponding eigenvalues reflect the amplitudes of motion along those directions. The trajectories have been sampled uniformly either to obtain 2,500 frames for each individual analysis (PCA-1 and PCA-2) or every 50th frame to obtain 4,406 frames for combined analysis (PCA-3). In the latter analysis, all possible distances between four central α-helices were measured. The objective was to reduce the number of features describing those distances to 2. The helices were defined as follows: 609–639 (A1) and 692–715 (A2) from subunit A and 1523–1552 (B1) and 1605–1628 (B2) from subunit B. All analyses were carried out with either pyPCAZIP, a python-based reimplementation of the PCAZIP package (PCA-1 and PCA-2)75 or scikit-learn Python package (PCA-3)76. The plots of concerted motions were prepared with the help of cpptraj and the Normal Mode Wizard VMD plugin77,78 and rendered with POV-Ray 3.6 (PCA-1 and PCA-2)79.
Ethics Statement
Written informed consent was obtained from all patients for muscle biopsy analyses as well as for genetic testing, according to the Declaration of Helsinki. The study was approved by the Ethics Committee of the Warsaw Medical University (Warsaw, Poland) in compliance with national legislation and with the Code of Ethical Principles for Medical Research Involving Human Subjects of the World Medical Association.
Data Availability
The datasets generated during and/or analyzed during the current study are available from the corresponding author on a reasonable request.
References
Vissing, J. Limb girdle muscular dystrophies: classification, clinical spectrum and emerging therapies. Curr. Opin. Neurol. 29, 635–641 (2016).
Servián-Morilla, E. et al. A POGLUT1 mutation causes a muscular dystrophy with reduced Notch signaling and satellite cell loss. EMBO Mol. Med. 8, 1289–1309 (2016).
Straub, V. et al. 229th ENMC international workshop: Limb girdle muscular dystrophies - Nomenclature and reformed classification Naarden, the Netherlands, 17-19 March 2017. LGMD workshop study group. Neuromuscul. Disord. 28, 702–710 (2018).
Mitsuhashi, S. & Kang, P. B. Update on the genetics of limb girdle muscular dystrophy. Sem. Pediatric Neurol. 19, 211–218 (2012).
Azzimato, V., Gennebäck, N., Tabish, A. M., Buyandelger, B. & Knöll, R. Desmin, desminopathy and the complexity of genetics. J. Mol. Cell. Cardiol. 92, 93–95 (2016).
Nigro, V. & Savarese, M. Genetic basis of limb-girdle muscular dystrophies: the 2014 update. Acta Myol. 33, 1–12 (2014).
Amato, A. A. & Brown, R. H. Jr. Dysferlinopathies. Handb. Clin. Neurol. 101, 111–118 (2011).
Fanin, M. & Angelini, C. Progress and challenges in diagnosis of dysferlinopathy. Muscle Nerve 54, 821–835 (2016).
Savarese, M. et al. Next generation sequencing on patients with LGMD and nonspecific myopathies: Findings associated with ANO5 mutations. Neuromuscul. Disord. 25, 533–541 (2015).
Katoh, M. & Katoh, M. Identification and characterization of TMEM16E and TMEM16F genes in silico. Int. J. Oncol. 2, 1345–1349 (2004).
Hicks, D. et al. A founder mutation in Anoctamin 5 is a major cause of limb-girdle muscular dystrophy. Brain 134, 171–182 (2011).
Pedemonte, N. & Galietta, L. J. Structure and function of TMEM16 proteins (anoctamins). Physiol. Rev. 9, 419–459 (2014).
Tian, Y., Schreiber, R. & Kunzelmann, K. Anoctamins are a family of Ca2+-activated Cl2 channels. J. Cell Sci. 125, 4991–4998 (2012).
Tsutsumi, S. et al. The novel gene encoding a putative transmembrane protein is mutated in gnathodiaphyseal dysplasia (GDD). Am. J. Hum. Genet. 74, 1255–1261 (2004).
Mizuta, K. et al. Molecular characterization of GDD1/TMEM16E, the gene product responsible for autosomal dominant gnathodiaphyseal dysplasia. Biochem. Biophys. Res. Commun. 357, 126–132 (2007).
Jin, L. et al. Three novel ANO5 missense mutations in Caucasian and Chinese families and sporadic cases with gnathodiaphyseal dysplasia. Sci. Rep. 7, 40935, https://doi.org/10.1038/srep40935 (2017).
Andreeva, V. et al. Whole exome sequencing links dental tumor to an autosomal-dominant mutation in ANO5 gene associated with gnathodiaphyseal dysplasia and muscle dystrophies. Sci Rep. 6, 26440, https://doi.org/10.1038/srep26440 (2016).
Witting, N. et al. Anoctamin 5 muscular dystrophy in Denmark: prevalence, genotypes, phenotypes, cardiac findings, and muscle protein expression. J. Neurol. 260, 2084–2093 (2013).
Bolduc, V. et al. Recessive mutations in the putative calcium-activated chloride channel Anoctamin 5 cause proximal LGMD2L and distal MMD3 muscular dystrophies. Am. J. Hum. Genet. 86, 213–221 (2010).
Penttilä, S. et al. Eight new mutations and the expanding phenotype variability in muscular dystrophy caused by ANO5. Neurology 78, 897–903 (2012).
Papadopoulos, C. et al. Hyperckemia and myalgia are common presentations of anoctamin-5-related myopathy in French patients. Muscle Nerve 56, 1096–1100 (2017).
Sarkozy, A. et al. ANO5 gene analysis in a large cohort of patients with anoctaminopathy: confirmation of male prevalence and high occurrence of the common exon 5 gene mutation. Hum. Mutat. 34, 1111–1118 (2013).
Fichna, J. P. et al. Whole-exome sequencing identifies novel pathogenic mutations and putative phenotype-influencing variants in Polish limb-girdle muscular dystrophy patients. Hum Genomics 12, 34, https://doi.org/10.1186/s40246-018-0167-1 (2018).
Wattjes, M. P., Kley, R. A. & Fischer, D. Neuromuscular imaging in inherited muscle diseases. Eur. Radiol. 20, 447–2460 (2010).
Schwarz, J. M., Cooper, D. N., Schuelke, M. & Seelow, D. MutationTaster2: mutation prediction for the deep-sequencing age. Nat. Methods 11, 361–362 (2014).
Adzhubei, I. A. et al. A method and server for predicting damaging missense mutations. Nat. Methods 7, 248–249 (2010).
Choi, Y. & Chan, A. P. PROVEAN web server: a tool to predict the functional effect of amino acid substitutions and indels. Bioinformatics 31, 2745–2747 (2015).
Sim, N. L. et al. SIFT web server: predicting effects of amino acid substitutions on proteins. Nucleic Acids Res. 40, W452–457 (2012).
Liu, X., Jian, X. & Boerwinkle, E. dbNSFP: a lightweight database of human nonsynonymous SNPs and their functional predictions. Hum. Mutat. 32, 894–899 (2011).
Richards, S. et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet. Med. 17, 405–424 (2015).
Amadei, A., Linssen, A. B. M. & Berendsen, H. J. C. Essential dynamics of proteins. Proteins 17, 412–425 (1993).
Amadei, A., Linssen, A. B. M., de Groot, B. L., van Aalten, D. M. & Berendsen, H. J. C. An efficient method for sampling the essential subspace of proteins. J. Biomol. Struct. Dyn. 13, 615–625 (1996).
Daidone, I. & Amadei, A. Essential dynamics: foundation and applications. WIREs Comput. Mol. Sci. 2, 762–770 (2012).
Vihola, A. et al. Diagnostic anoctamin-5 protein defect in patients with ANO5-mutated muscular dystrophy. Neuropathol. Appl. Neurobiol., https://doi.org/10.1111/nan.12410 (2017)
Amburgey, K. et al. Genotype-phenotype correlations in recessive RYR1-related myopathies. Orphanet J. Rare Dis. 8, 117, https://doi.org/10.1186/1750-1172-8-117 (2013).
Ylikallio, E. et al. Decreased aerobic capacity in ANO5-muscular dystrophy. J. Neuromuscul. Dis. 3, 475–485 (2016).
Arndt, V. et al. Chaperone-assisted selective autophagy is essential for muscle maintenance. Curr. Biol. 20, 143–148 (2010).
Kiselev, A. et al. De novo mutations in FLNC leading to early-onset restrictive cardiomyopathy and congenital myopathy. Hum. Mutat. 39, 1161–1172 (2018).
Kostera-Pruszczyk, A. et al. BAG3-related myopathy, polyneuropathy and cardiomyopathy with long QT syndrome. J. Muscle Res. Cell Motil. 36, 423–432 (2015).
Selcen, D. et al. Mutation in BAG3 causes severe dominant childhood muscular dystrophy. Ann. Neurol. 65, 83–89 (2009).
Piga, D. et al. New Mutations in NEB Gene Discovered by Targeted Next-Generation Sequencing in Nemaline Myopathy Italian Patients. J. Mol. Neurosci. 59, 351–359 (2016).
Feingold-Zadok, M. et al. Mutations in the NEB gene cause fetal akinesia/arthrogryposis multiplex congenita. Prenat. Diagn. 37, 144–150 (2017).
Santulli, G., Lewis, D. R. & Marks, A. R. Physiology and pathophysiology of excitation-contraction coupling: the functional role of ryanodine receptor. J. Muscle Res. Cell Motil. 38, 37–45 (2017).
Hwang, J. H., Zorzato, F., Clarke, N. F. & Treves, S. Mapping domains and mutations on the skeletal muscle ryanodine receptor channel. Trends Mol. Med. 18, 644–657 (2012).
Chiquet, M., Birk, D. E., Bönnemann, C. G. & Koch, M. Collagen XII: Protecting bone and muscle integrity by organizing collagen fibrils. Int. J. Biochem. Cell Biol. 53, 51–54 (2014).
Quijano-Roy, S., Sparks, S.E. & Rutkowski, A. LAMA2-Related Muscular Dystrophy. (In: Adam, M. P. et al. editors. GeneReviews® [Internet]. Seattle (WA): University of Washington, Seattle) 1993–2018 (2012).
Beeson, D. Congenital myasthenic syndromes: recent advances. Curr. Opin. Neurol. 29, 565–571 (2016).
Hayashi, Y. K. et al. Mutations in the integrin alpha7 gene cause congenital myopathy. Nat. Genet. 19, 94–97 (1998).
Antonarakis, S.E. & Cooper, D.N. Human Gene Mutation: Mechanisms and Consequences. (In: Speicher, M., Antonarakis, S. E. & Motulsky, A. G. Vogel and Motulsky’s Human Genetics. 4th ed. Berlin Heidelberg: Springer) 319–364 (2010).
MacArthur, D. G. et al. Guidelines for investigating causality of sequence variants in human disease. Nature 508, 469–476 (2014).
Pathak, P. et al. Limb girdle muscular dystrophy type 2A in India: a study based on semi-quantitative protein analysis, with clinical and histopathological correlation. Neurol India 58, 549–554 (2010).
Fanin, M. et al. Frequency of LGMD mutations in Italian patients with distinct clinical phenotypes. Neurology 72, 1432–1435 (2009).
van der Kooi, A. J. et al. Limb-girdle muscular dystrophy in the Netherlands. Gene defect identified in half of the families. Neurology 68, 2125–2128 (2007).
Sveen, M. L., Schwartz, M. & Vissing, J. High prevalence and phenotype-genotype correlations of limb girdle muscular dystrophy type 2I in Denmark. Ann. Neurol. 59, 808–815 (2006).
Di Fruscio, G., Garofalo, A., Mutarelli, M., Savarese, M. & Nigro, V. Are all the previously reported genetic variants in limb girdle muscular dystrophy genes pathogenic? Eur. J. Hum. Genet. 24, 73–77 (2016).
Mercuri, E. et al. Clinical and imaging findings in six cases of congenital muscular dystrophy with rigid spine syndrome linked to chromosome 1p (RSMD1). Neuromuscul. Disord. 12, 631–638 (2002).
Fischer, D. et al. Distinct muscle imaging patterns in myofibrillar myopathies. Neurology 71, 758–765 (2008).
Miller, S. A., Dykes, D. D. & Polesky, H. F. A simple salting out procedure for extracting DNA from human nucleated cells. Nucleic Acids Res. 16, 1215 (1988).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
Robinson, J. T. et al. Integrative genomics viewer. Nat Biotechnol 29, 24–6 (2011).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164, https://doi.org/10.1093/nar/gkq603 (2010).
Girdea, M. et al. PhenoTips: patient phenotyping software for clinical and research use. Hum. Mutat. 34, 1057–1065 (2010).
Smedley, D. & Robinson, P. N. Phenotype-driven strategies for exome prioritization of human Mendelian disease genes. Genome Med. 7, 81, https://doi.org/10.1186/s13073-015-0199-2 (2015).
Kelley, L. A., Mezulis, S., Yates, C. M., Wass, M. N. & Sternberg, M. J. E. The Phyre2 web portal for protein modeling, prediction and analysis. Nat. Protoc. 10, 845–858 (2015).
Pierce, B. G. et al. ZDOCK Server: Interactive docking prediction of protein-protein complexes and symmetric multimers. Bioinformatics 30, 1771–1773 (2014).
Case, D. A. et al. AMBER 2016. (University of California, San Francisco, 2016).
Jorgensen, W. L., Chandrasekhar, J., Madura, D., Impey, R. W. & Klein, M. L. Comparison of simple potential functions for simulating liquid water. J. Chem. Phys. 79, 926–935 (1983).
Essmann, U. et al. A smooth particle mesh Ewald method. J. Chem. Phys. 103, 8577–8593 (1995).
Ryckaert, J. P., Ciccotti, G. & Berendsen, H. J. C. Numerical integration of the Cartesian equations of motion of a system with constraints: molecular dynamics of n-Alkanes. J. Comput. Phys. 23, 327–341 (1977).
Maier, J. A. et al. ff14SB: Improving the Accuracy of Protein Side Chain and Backbone Parameters from ff99SB. J. Chem. Theory Comput. 11, 3696–3713 (2015).
Roe, D. R. & Cheatham, T. E. III. PTRAJ and CPPTRAJ software for processing and analysis of molecular dynamics trajectory data. J. Chem. Theory Comput. 9, 3084–3095 (2013).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Krissinel, E. Crystal contacts as nature’s docking solutions. J. Comput. Chem. 31, 133–143 (2010).
Shkurti, A. et al. pyPCAZIP: A PCA-based toolkit for compression and analysis of molecular simulation data. SoftwareX 5, 44–50 (2016).
Pedregosa, F. et al. Scikit-learn: Machine Learning in Python. J. Mach. Learn. Res. 12, 2825–2830 (2011).
Bakan, A., Meireles, L. M. & Bahar, I. ProDy: Protein dynamics inferred from theory and experiments. Bioinformatics 27, 1575–1577 (2011).
Bakan, A. et al. Evol and ProDy for bridging protein sequence evolution and structural dynamics. Bioinformatics 30, 2681–2683 (2014).
Persistence of Vision Pty. Ltd. Persistence of Vision Raytracer (Version 3.6) [Computer Software], http://www.povray.org/download/ (2004).
Acknowledgements
This work was supported by grants from Polish National Science Center (NCN 2013/09/B/NZ4/03258 and NCN 2012/05/D/NZ4/02978) as well as the statutory funds for the Nencki Institute, for the Department of Neurology of Medical University of Warsaw and for Mossakowski Medical Research Centre from the Ministry of Science and Higher Education. Also, it was supported by the KNOW-MARC project financed by the Ministry of Science and Higher Education (to J.P.F.). The bioinformatic part of the research (J.P.F.) was supported by PL-Grid Infrastructure financed by the Polish Ministry of Science and Higher Education and co-funded by the European Regional Development Fund as part of the Innovative Economy program. The MYO-SEQ project was supported by Sanofi Genzyme, Ultragenyx, LGMD2I Research Fund, Samantha J Brazzo Foundation, LGMD2D Foundation, Kurt + Peter Foundation, Muscular Dystrophy UK and Coalition to Cure Calpain 3. The funding sources had no involvement in the experiment design, analysis, data interpretation and manuscript preparation. Computations were carried out with the support of the Interdisciplinary Centre for Mathematical and Computational Modeling (ICM) at University of Warsaw (grant GB67-19 to AJ). We acknowledge use of CePT infrastructure financed by the European Union: the European Regional Development Fund within the Operational Programme ‘Innovative Economy’ for 2007–2013.
Author information
Authors and Affiliations
Contributions
All the authors were involved in planning the experiments, data analysis and critical reading of the manuscript draft. M.J.R. and A.M.K. conceived the manuscript; A.L., A.M.b and A.M.K. were involved in clinical evaluation; J.P.F., A.M.c, K.J., A.T. and V.S. performed genetic analyses; E.R. performed MRI examination; A.M.K. and M.T. were involved in muscle morphology analysis; A.J., K.S. and S.D.-H. performed molecular dynamics analyses; M.J.R., A.M.K., J.P.F. and A.J. wrote the manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Jarmula, A., Łusakowska, A., Fichna, J.P. et al. ANO5 mutations in the Polish limb girdle muscular dystrophy patients: Effects on the protein structure. Sci Rep 9, 11533 (2019). https://doi.org/10.1038/s41598-019-47849-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-47849-3
This article is cited by
-
A Journey with LGMD: From Protein Abnormalities to Patient Impact
The Protein Journal (2021)
-
Diagnostic muscle biopsies in the era of genetics: the added value of myopathology in a selection of limb-girdle muscular dystrophy patients
Acta Neurologica Belgica (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.