Abstract
Prion diseases are caused by the conversion of physiological PrPC into the pathogenic misfolded protein PrPSc, conferring new properties to PrPSc that vary upon prion strains. In this work, we analyze the thermostability of three prion strains (BSE, RML and 22L) that were heated at 98 °C for 2 hours. PrPSc resistance to proteinase K (PrPres), residual infectivity by mouse bioassay and in vitro templating activity by protein misfolding cyclic amplification (PMCA) were studied. Heated strains showed a huge loss of PrPres and a radically different infectivity loss: RML was the most thermolabile strain (6 to 7 log10 infectivity loss), followed by 22L (5 log10) while BSE was the most thermostable strain with low or null infectivity reduction showing a clear dissociation between PrPres and infectivity. These results indicate that thermostability is a strain-specific feature, measurable by PMCA and mouse bioassay, and a great tool to distinguish prion strains.
Similar content being viewed by others
Introduction
Prion diseases are fatal neurodegenerative diseases that affect numerous mammal species and include kuru, Creutzfeldt-Jakob disease (CJD), fatal familial insomnia (FFI) and Gerstmann-Sträussler-Scheinker disease (GSS) in humans, scrapie in sheep and goats, bovine spongiform encephalopathy (BSE) in cattle and chronic wasting disease (CWD) in cervids1. According to the Protein-Only Hypothesis, prion diseases are caused by the conversion of the physiological cellular prion protein (PrPC) into a pathogenic β-sheets enriched isoform (PrPSc) that is able to self-propagate by recruiting and converting more PrPC 2.
PrPC conversion into PrPSc is a post-translational process where both isoforms share an identical amino acid sequence but differ in their conformation. This conformational change confers distinct physicochemical properties such as greater tendency to aggregate, greater insolubility in non-ionic detergents, partial resistance to protease digestion and high resistance to heat and chemical sterilization3,4,5,6,7. Notably, these properties vary upon prion agents which may result in distinctive prion disease phenotypes including incubation times, clinical signs, histopathological lesions and PrPSc deposition patterns in the brain. The existence of such prion strains has been proposed by many authors and are associated to the range of thermodynamically stable PrPSc conformers into which the PrPC can misfold during prion conversion8,9,10. Many prion features, apart from the ones directly related to disease phenotype and mentioned above are given by the particular prion strain. In this way, the “stability” of prions, understood as the ability to support certain physical-chemical conditions retaining its specific strain features, could be considered as other differential trait between strains. Related to this concept of prion stability, a direct correlation between the fragility of yeast prion fibrils and their rate of replication have been reported11,12,13 and extended to mammalian prions from synthetic mammalian prions14,15. By contrast, ovine and human prions seem not to follow this rule16,17. In addition, conformational stability assay of 30 different prion isolates revealed a linear relationship between the concentrations of guanidine hydrochloride (Gdn-HCl) required to denaturing 50% of PrPSc molecules and their incubation times15. Prion stability has also been associated with the differential ability of prions to invade the central nervous system (CNS)18. This suggests that highly neuroinvasive prion strains may be conformationally unstable in denaturing conditions and efficiently form diffuse, non-fibrillar PrP aggregates in the CNS, which produces a rapid progression to terminal disease in mice. On the other hand, weakly neuroinvasive strains may form dense, fibrillar plaques and mice progress to terminal disease more slowly18.
In this work, we analyze the stability of three prion strains from the point of view of stability against heat treatment (thermostability). For that purpose, three different mouse-adapted prion strains (BSE, RML and 22L) were heated and/or proteinase K (PK) digested and the effect of these treatments on their biological and biochemical properties as well as on their infectivity and templating activity was studied. Our results suggest that heat treatment substantially reduced PK resistant PrPSc (PrPres) decreases prion infectious titer without changing any prion strain feature and that prion resistance to heat treatment is strain dependent, BSE being the most thermostable strain with low or null infectivity and templating activity decrease. Differential thermostability between strains combined with PMCA appears as a suitable method for strain typing.
Results
Differential effect of the heat treatment on the biochemical and biological properties of prion strains
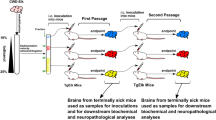
22L, RML and BSE inocula for this work were obtained by pooling brain homogenates of Tga20 transgenic mice (10% weight/volume in PBS) after serial passage. Tga20 transgenic mice were selected because its high overexpression of mouse PrP allow murine prion amplification with short mean survival times with a high sensitivity. These materials were subjected to heat treatment or heat treatment combined with PK digestion and then assayed for biochemical characteristics. First, Western blot (WB) analysis of the total PrP levels on the plain heated and non-heated samples were done (Fig. 1A). Heated samples showed a great reduction in their total PrP when compared to their non-heated counterparts. In the same line of results, WB analysis of the heated samples showed lower amounts of detectable PrPres than their respective non-heated counterparts (at least 64 fold reduction for 22L and BSE while between 16 to 64 fold reduction was quantified for RML) (Fig. 1B). In addition, samples subjected first to PK treatment and then heat pulse were included in the study. Since heat treatment caused such a high reduction on the PK resistant population, we applied the heat pulse to a sole PK resistant PrP population to test if any differences with the previous samples can be found. These samples also showed the same amounts of detectable PrPres than the plain heated ones (Fig. 1C). To obtain comparable signals, it was necessary to load a 2x or 4x fold amount of heated samples against a 1/20 dilution of the unheated samples. The reduction in the amount of PrPres in heated samples was independent of whether the PK treatment was done before (PK + 98 °C) or after heat treatment (98 °C + PK). Thus, suggesting that the PrP termoresistant population is also PK resistant but only a very limited part of the PrPres population is termoresistant.
Total PrP and PrPres levels of non-heated (NH) and heated (98 °C) and PK digested (PK) samples. (A) The relative amount of total PrP in non-heated (NH) and heated (98 °C) samples was quantified by serial dilutions in WB using the Sha31 mAb. (B) PrPres quantification in non-heated (NH + PK) and heated (98 °C + PK) samples by serial dilutions in WB using the Sha31 mAb. Heated samples showed a general decrease on both total PrP and PrPres levels. (C) Comparison of PrPres glycoprofile between non-heated (NH + PK) and heated samples (98 °C + PK or PK + 98 °C) by loading equilibrated amounts of protein. To obtain comparable signals, it was necessary to load a 2x or 4x fold amount of heated samples against a 1/20 dilution of the unheated samples. The reduction in the amount of PrPres in heated samples was independent of whether the PK treatment was done before (PK + 98 °C) or after heat treatment (98 °C + PK). In all cases, samples were digested using PK at a 40 µg/ml concentration. Serial dilutions were done by diluting the original sample into loading buffer. Molecular weights in kD are shown.
These losses of both total PrP and PrPres signals were in the same order of magnitude for all the strains which suggests that this degradation may not be strain-dependent. In addition, the biochemical signature of the samples treated with the different conditions showed no apparent difference in the WB PrPres glycoprofile when compared to its non-heated controls (Fig. 1C).
Sha31 monoclonal antibody (mAb) was selected for the detection of PrP in WB because it binds the 144-WEDRYYRE-151 epitope of the mouse-PrP sequence19, which is in the core of the PrPres and allows detection of the different PrPres signatures with high sensitive. Other mAbs such as Saf84 (162-YRPVDQY-168 epitope of the mouse-PrP sequence)19 and 9A2 (98-WNK-100 epitope of the mouse-PrP sequence)20 were also tested producing very similar results (data not shown).
Finally, heated and non-heated samples were subjected to PK digestion using PK concentrations ranging from 2.5 to 40 µg/ml (Fig. 2). This will clarify if PrPres reduction detected on heated samples by WB was due to direct degradation by heating or due to an increased sensitivity to PK as a consequence of a conformational change during the heating process. Sha31 antibody was selected for the PK assay because it binds the 144-WEDRYYRE-151 epitope of the mouse-PrP sequence19, which is in the core of the PrPres and allows detection of several different PrPres signatures with high sensitivity. As expected, non-heated isolates of all the three strains remained stable after PK digestion. No PrPres signal was detected even when the heated samples were treated with low concentrations of PK.
Analysis on PrPres sensibility to PK treatment before and after heat treatment. PrPres sensibility to PK treatment was analyzed by WB using the Sha31 mAb. Different concentration of PK ranging from 40 to 2.5 µg/ml were applied to non-heated and heated samples. Non-heated samples showed a high and stable resistance to PK while heated samples showed no PrPres presence even at low concentrations of PK when only faint bands were detected. Molecular weights in kD are shown.
We next studied whether heat treatment altered the strains biological properties. For that purpose, plain heated and unheated BSE, RML and 22L samples were intracerebrally inoculated in Tga20 mice and mean survival time (ST) assessed. Mice inoculated with heated RML or heated 22L isolates showed 89% and 55% increased ST, respectively, compared to the animals inoculated with their correspondent original unheated samples (Table 1). By contrast, the heat treatment increased the ST of heated BSE inoculated mice by just a 7% compared to the animals inoculated with original unheated BSE, which is not statistically significant (Table 1).
Survival curves visually representing this data can be found in Fig. 3A. Brain PrPres of the inoculated animals proved to maintain the original electrophoretic signature of the prion strains (Fig. 3B). In the same line of results, for histopathological analysis no major differences of brain PrPSc distribution pattern was detected between animals infected with either heated or non-heated control samples (Fig. 3C,D). However increased intensity in the signal of the PET-blot pictures for animals inoculated with heated RML and 22L were detected, probably due to the longer survival time observed in these animals which could favor a greater accumulation of PrPres. Overall, this result indicates that no significant change in the neurological targeting of the three strains had occurred after heating.
Transmission of non-heated (NH) and heated (98 °C) samples into Tga20 mice. (A) Survival curves visually representing the data found in Table 1. (B) PrPres levels of original heated and non-heated samples in comparison to brain PrPres from Tga20 mice challenged with heated and non-heated samples were detected in western blot with Sha31 mAb. Molecular weights in kD are shown. (C) Histological characterization of Tga20 transgenic mice inoculated with RML, 22L and BSE heated and non-heated samples. Distribution of PrPres deposits as assayed by PET-blot with Sha31 mAb showed no change between animals inoculated with heated or non-heated samples. (D) Histological characterization of Tga20 transgenic mice inoculated with RML, 22L and BSE heated and non-heated samples. Hematoxylin and eosin showed no change between animals inoculated with heated or non-heated samples.
Differential effect of heat treatment in prion infectivity of different prion strains by mouse bioassay
The increases in ST observed in RML and 22L strains could have been the result of i) a decrease in prion infectivity, or ii) the appearance of new prion properties associated to PrPSc conformational alterations and therefore the emergence of a new different prion strain with longer ST. This last option was not much probable since previous results showed a concordance in both the WB profile and histological features. To fully discriminate between these two possibilities, heated and non-heated isolates were titrated by mouse bioassay. Ten-fold serial dilutions of these samples were intracranially inoculated in Tga20 mice and the infectivity of each dilution assessed as a function of their mean ST (Table 2).
As expected, no major differences in prion infectivity of heated and non-heated BSE samples were observed for any of the dilutions (Table 2), pointing to the high resistance to heat of this prion strain. By contrast, heat treatment decreased the infectious titer of RML and 22L prions by 6 and 5 log10, respectively.
Differential effect of the heat treatment plus PK digestion on the prion templating activity of prion strains in vitro
The key event causing prion diseases, the PrPC conversion into PrPSc, has been successfully reproduced in cell-free conditions by PMCA21. This technique is able to amplify different prion strains in healthy brain homogenates from different species, maintaining their strain-specific biological, biochemical and infectious properties22. The higher sensitivity of the technique compared to bioassay makes it possible to test prion templating activity, meaning the capacity of the prion agent to sustain effective PrPSc amplification in serial dilutions as other way to perform prion titration detecting subinfectious levels of prions23. We thus used it as an alternative and more rapid approach for assessing the differential thermostability of prion strains. For that purpose, serial 10-fold dilutions of heated, heated plus PK digested, PK digested plus heated and untreated isolates were subjected to PMCA amplification using non-infected Tga20 brain homogenate as substrate, as previously described23. The amplified material was then analyzed for PrPres content by immunobloting. Conversion capacity of each strain was assessed and the loss of templating activity after the different treatments was compared to that obtained by mouse bioassay (Table 2). Non-heated RML and 22L prions exhibited high conversion capacities, resulting in positive amplification with brain material diluted up to 10−9 and 10−8–fold, respectively (Table 3). After heat treatment, RML and 22L amplified up to 10−4 and 10−3 dilutions, which indicate a templating activity loss of 5 log10 for RML and 22L (Table 3, Fig. 4). Exactly the same results were obtained when heat treatment was combined with PK digestion, regardless if the PK digestion was done before or after the heat pulse. By contrast BSE strain showed identical conversion capacities up to 10−11 dilutions both without and with the different treatments (Table 3, Fig. 4). The templating activity losses assessed for each strain mostly overlapped those infectivity losses previously determined by mouse bioassay of the plain heated and unheated samples which supports that PMCA would be a suitable and fast procedure for assessing thermostability of prion strains, instead of using an in vivo approach which consume more time and requires a higher use of animals.
Templating activity analysis. Comparison of templating activity of non-heated (NH) with heated (98 °C) (A), heated and then PK digested (98 °C + PK) (B), PK digested and then heated (PK + 98 °C) (C) samples by PMCA on Tga20 mouse brains analyzed by WB using the Sha31 mAb. 22L and RML treated samples showed a lower templating activity when compared to its non-heated counterparts.
Discussion
Post-translational conversion of PrPC into PrPSc causes a deep change in both protein structure and physicochemical features. In this study, three well defined mouse-adapted prion strains (BSE, RML and 22L), were treated with different conditions that involved heating at 98 °C for two hours combined or not with PK digestion and their biological and biochemical properties, as well as residual infectivity and seeding activity were assessed in comparison to untreated control strains.
Heat treatment caused an increase in the ST of inoculated mice when compared to mice challenged with unheated control material. Such increase was dependent on the prion strain being BSE the most thermostable strain (just a 7% increase in the mean ST) whereas RML and 22L were much more thermolabile (89% and 55% increase in the mean ST respectively). Increase in ST could be attributed to a shift of the prion strain as a consequence of the heat treatment. In a previous report studying prion thermostability, scrapie prion strain ME7 retained its general strain features while showing incubation periods longer than the controls at equivalent doses once autoclaved at 126 °C24. This result was interpreted as differences in the structure of the causal agent. However, none of the strain features like PrPres glycoprofile, PrPres deposition pattern and lesion profiles in the brains of inoculated mice were altered when comparing heated and non-heated isolates in our study. Therefore, it seems that the heat treatment just caused a reduction in the infectivity titer for our tested prion strains. Our end-point titration studies by both mouse bioassay and PMCA clearly proved that hypothesis finally discarding the possibility of a strain change. Indeed, PMCA also proved that combination of PK digestion with the heat treatment (either after or before the heat pulse) resulted in no change in the templating activity of the samples compared to plain heated samples. This work proves that PMCA can be used as a tool to discriminate between more or less thermostable strains faster than mouse bioassay.
It is important to note that heated isolates tested on this study showed differential reduction on infectivity (as proved by both mouse bioassay and PMCA) while showing high reductions in PrPC and PrPres by WB. The most extreme case is the one of BSE in which near total disappearance of PrPres resulted in no reduction of the infectivity. Indeed, there are some kind of inverse relationship between the amount of PrPres retained after heat treatment and maintenance of infectivity for the strains used on this study: 22L and RML retained some PrPres but their infectivity was strongly affected whereas BSE infectivity was intact although showing no detectable PrPres. Partial and total dissociation between prion infectivity and PrPres presence have been detected recurrently in the prion field. In the very beginning, such fact was used to claim against the Prion Protein-Only Hypothesis25. Later, attempts to clarify where exactly prion infectivity resides have been done although giving contradictory results. Regarding the size of the prion fibrils, for certain strains, infectivity was mapped to small and soluble PrPSc oligomers rather than larger fibrils26,27. Other studies, demonstrate the existence of protease-sensitive oligomers named sPrPSc (for “sensitive” PrPSc) which may contribute to prion pathogenesis28,29,30,31 or not depending on the prion strain32. In this sense, our PMCA analysis using PK digested and heated material showed that the particle retaining the infectivity in the case of BSE is both resistant to heat and PK treatment. For 22L and RML, other PrP subpopulations affected for the different treatments subjected on this work were also responsible of part of the infectivity. At a subcellular level, strong correlation between infectivity and presence of lipid rafts have been traditionally done33. In fact, addition of certain lipids and polyanions to PMCA reactions tends to increase the efficiency of purified recombinant and cellular PrP conversion34,35,36. In contrast, other reports also point to presence of infectivity related to mitochondria and endoplasmic reticulum enriched fractions37,38,39. Thus, ours results provide another observation about dissociation of PrPres and infectivity and more research is needed to clarify the puzzle conformed by PrPres, sPrPSc and other co-factors important for the maintenance of the infectivity. However, the possibility that each strain refold differently or require different refolding kinetic after the heat treatment could not be ruled out. The retained electrophoretic mobility and glycoform pattern argue against changes in the secondary and tertiary structure but quaternary structure could have been modified after the heat treatment, explaining the differences between strains.
Previous reports that studied differential resistance to heat in prion strains proposed the existence of a heteromeric structure involving two types of macromolecules in which one macromolecule, independent of the host, differs in its covalent structure between TSE strains24,40, being compatible with the “virino hypothesis” which proposes a host-independent informational molecule protected by the host protein PrP41. Later, the existence of prion strains as a result of different tertiary and quaternary structures while sharing the same amino acid sequence were proposed42. Therefore, both previous reports24,37,43 and our current results indicate that thermostability is a strain-specific physicochemical feature like the PrPres glycoprofile, lesion profile or other well-known prion strain characteristics. Thus, it would be related to some extent to the PrPSc tertiary and/or quaternary structure. Regarding how prion molecules manage to survive to heat treatment, other authors postulated that dehydration of the prion agents by means of the removal of the water of solvation could stabilize the interactions between prion protein and some TSE agent-specific ligands43. In the same line, it was recently proposed that dehydration may also protect prion against inactivation via freezing and thawing processes44.
Two of the strains employed on this work, RML and 22L, are mouse scrapie isolates which proved to be very thermolabile. On the other hand, BSE was very much resistant to heat treatment. The ability to reach the brain in a faster way was linked to poor resistance to heat18. However, BSE is reported here as very stable being also well known for its high neuroinvasion capacity45. Indeed, BSE has already been described as an extremely stable prion strain and extremely resistant to environmental conditions46,47,48,49. As a matter of fact, BSE thermostability was related to its own origin, since spreading through meat-and-bone meal (MBM) implied heating of cow tissues for MBM production. Specifically, BSE emergence in the UK in the late 1980s is related to a replacement of batch dry-rendering plants by continuous-rendering systems which supposed that both the cooking temperature and the drying time were reduced50. Therefore, special efforts must be done to avoid residual prion contamination especially in the case of highly thermostable strains like BSE.
Materials and Methods
Ethics statement
Animal experiments were performed in strict accordance with the recommendations of the Code for Methods and Welfare Considerations in Behavioral Research with Animals (Directive 86/609EC and 2010/63/EU). Experiments in mice were carried out in CISA-INIA (Madrid) and authorized by the Committee on the Ethics of Animal Experiments (CEEA) of the Spanish Instituto Nacional de Investigación y Tecnología Agraria y Alimentaria (INIA); Permit Numbers: PROEX263/15 and CEEA2009/004.
Prion samples
Three mouse adapted strains, BSE, RML and 22L, were used on this study. Such prion isolates have been obtained through serial transmission into mouse-PrP Tga20 mice which overexpress the mouse wild-type PrPC 51. Prion isolates consisted on pooled brain homogenates (10% weight/volume in PBS).
Heat treatment and PK digestion of prion strains
Several combinations of heat plus PK treatment were studied on this work: Plain heat treatment, heat treatment plus PK digestion and finally PK digestion plus heat treatment.
Heat treatment (98 °C): Brain homogenates of Tga20 mice infected with BSE, RML or 22L strains were aliquoted (500 µl), placed in safe-lock tubes (Eppendorf) and heated at 98 °C for 2 hours in a thermocycler (Primus 96 Plus Thermal Cycler, MWG AG Biotech). Samples were removed, allowed to cool gradually to room temperature and harvested at −20 °C.
Heat treatment plus PK digestion (98 °C + PK): The heating process was done as stated above. After the gradually cooling of the samples, 250 µl were taken and aliquoted in 50 µl. Individual aliquots were mixed with 150 µl of healthy sheep brain and subjected to PK digestion with 40 µg/ml of PK using the reagents of the TeSeE (Bio-Rad) enzyme-linked immunosorbent assay at 37 °C for 15 min. The reaction was stopped by adding PMSF to a final concentration of 2 mM. Finally, samples were subjected to centrifugation (15,000 g, 7 min, 20 °C) and pellets were resuspended in 50 µl of PBS.
PK digestion plus heat treatment (PK + 98 °C): PK digestion was done as described above using as starting material 50 µl of unheated brain homogenates. After centrifugation, pellets were resuspended in 50 µl of PBS and subjected to heat treatment as described above.
Total PrP and PrPres detection by Western blot in the treated samples (inocula)
For total PrP detection, 15 µl of the different samples were analyzed by WB. Samples were prepared as previously detailed52. Serial dilutions were done by diluting the original sample into loading buffer just before electrophoresis. For immunoblotting, membranes were incubated with Sha31 mAb19 that recognizes 144-WEDRYYRE-151 epitope of the mouse-PrP sequence. To determine the presence of PrPres, 20 μl of the different samples were analyzed by WB as previously described53. For immunoblotting, membranes were incubated with Sha31 mAb. Immunocomplexes were detected with horseradish peroxidase-conjugated anti-mouse IgG (Amersham Pharmacia Biotech) after incubating the membranes for 1 hour. Immunoreactivity was visualized by chemiluminescence with ECL Select (GE Healthcare Amersham Biosciences). When necessary, WB images were quantified using ImageLab software v5.2.1(BioRad).
Measurement of prion infectivity by mouse bioassay
The relative infectivity in the unheated and heated BSE, RML or 22L samples (10% brain homogenates) was calculated as a function of the survival times (ST) observed after their inoculation in Tga20 mice. Briefly, groups of 6–9 individually identified (six to seven weeks old) Tga20 mice were anesthetized with isoflurane and intracerebrally inoculated with 20 µl of the inocula (ten-fold serial dilution of each unheated and heated sample) in the right parietal lobe using a 25-gauge disposable hypodermic needle. Mice were examined biweekly for the development of neurological signs of the prion disease and humanely euthanized by cervical dislocation when progression of the disease was evident or at the end of the titration study: 2-fold the mean ST of the corresponding non-diluted sample for each strain. A mouse was considered positive for neurological disease when it showed two or three out of the 10 signs of neurological dysfunction previously described54,55. Once euthanized, brains were removed and analyzed for the presence of PrPres by WB as described above. ST and attack rate were calculated for each inoculum. ST was expressed as the mean of the survival dpi of all the mice scored positive for brain PrPres, with its correspondent standard deviation (ST). Attack rate was determined as the proportion of mice scored positive for brain PrPres from all the inoculated mice. Infectious dose (ID) was subsequently assessed by the Reed-Muench method56 and expressed in ID50 per gram (ID50 g−1) of tissue (Table 2) as previously described57.
Statistical information
Mann-Whitney’s t test was applied to the ST data in order to find statistically significant differences between heated and un-heated inoculated animals. Significance level was adjusted to 0.05 and the test had two tails.
PrPres detection by Western blot in the brain of transgenic mice after mouse bioassay
A total of 175 mg of whole brain tissue was homogenized in 5% glucose in distilled water in grinding tubes (Bio-Rad) and adjusted to 10% (w/v) by using a TeSeETM Precess 48TM homogenizer (Bio-Rad) following the manufacturer’s instructions. Homogenates were pressed through 0.4 mm needles of a calibration syringe and immediately frozen at −20 °C. To determine the presence of PrPres, 20 μl of 10% brain homogenates were analyzed by WB as described above. For immunoblotting, membranes were incubated with Sha31 mAb19.
Measurement of prion templating activity by PMCA
Unheated and heated BSE, RML or 22L samples (10% brain homogenates) were titrated for their templating activity by miniaturized-bead PMCA (mb-PMCA)23. Mouse brain lysates from healthy Tga20 mice were prepared and used as substrate. PMCA was performed in a final volume of 36 µl of lysate per well, in 96-well PCR microplate (Axygen, Union City, CA, USA). Each well was first filled with teflon beads (2.381 mm in diameter; Marteau et Lemarié, Pantin, France). A 4-µl aliquot of each original prediluted BSE, RML or 22L brain homogenates was suspended in 36 µl of healthy Tga20 brain lysate to obtain the first 10−3 dilution. Then, a series of 10-fold dilutions were made. Microplates were placed on a Plexiglas rack designated for the cup horn of the Q700 sonicator (Misonix, Farmingdale, NY, USA, or Delta Labo, Colombelles, France) and subjected to 96 cycles of 30 s of sonication at 200–220 W power (~30% amplitude of the Q700 sonicator) followed by 29 min 30 s of incubation at 37 °C. The cup horn was filled with 300 ml of water circulating with rubber tubing in a water bath maintained at a temperature of 37 °C. At the end of the PMCA, the microplates were removed and aliquots from each sample were taken to be analyzed for their PrPres content by dot- and WB analysis as previously described23. Negative controls were run in parallel. They were composed of unseeded substrate or seeded with uninfected brain homogenate.
Data Availability
The authors declare that all other data supporting the findings of this study are available within the paper.
References
Aguilar-Calvo, P., García, C., Espinosa, J. C., Andreoletti, O. & Torres, J. M. Prion and prion-like diseases in animals. Virus Res. 207, 82–93 (2015).
Ma, J. & Wang, F. Prion disease and the ‘protein-only hypothesis’. Essays Biochem. 56, 181–91 (2014).
Caughey, B. & Chesebro, B. Transmissible spongiform encephalopathies and prion protein interconversions. Adv Virus Res. 56, 277–311 (2001).
Collinge, J. Prion diseases of humans and animals: their causes and molecular basis. Annu Rev Neurosci. 24, 519–550 (2001).
McKinley, M. P., Masiarz, F. R., Isaacs, S. T., Hearst, J. E. & Prusiner, S. B. Resistance of the scrapie agent to inactivation by psoralens. Photochem Photobiol. 37, 539–545 (1983).
Collinge, J. & Clark, A. R. A General Model of Prion Strains and Their Pathogenicity. Science. 318, 930–936 (2007).
Riesner, D. Biochemistry and structure of PrP(C) and PrP(Sc). Br Med Bull. 66, 21–33 (2003).
Prusiner, S. B. P. Proc Natl Acad Sci USA 95, 13363–13383 (1998).
Wiltzius, J. J. et al. Molecular mechanisms for protein-encoded inheritance. Nat Struct Mol Biol. 16, 973–978 (2009).
Angers, R. C. et al. Prion strain mutation determined by prion protein conformational compatibility and primary structure. Science. 328, 1154–1158 (2010).
Immel, F. et al. In vitro analysis of SpUre2p, a prion-related protein, exemplifies the relationship between amyloid and prion. J Biol Chem. 282, 7912–7920 (2007).
Tanaka, M., Collins, S. R., Toyama, B. H. & Weissman, J. S. The physical basis of how prion conformations determine strain phenotypes. Nature. 442, 585–589 (2006).
Tanaka, M., Chien, P., Naber, N., Cooke, R. & Weissman, J. S. Conformational variations in an infectious protein determine prion strain differences. Nature. 428, 323–8 (2004).
Sun, Y. et al. Conformational stability of PrP amyloid fibrils controls their smallest possible fragment size. J Mol Biol. 376, 1155–1167 (2008).
Legname, G. et al. Continuum of prion protein structures enciphers a multitude of prion isolate-specified phenotypes. Proc Natl Acad Sci USA 103, 19105–19110 (2006).
Laferrière, F. et al. Quaternary structure of pathological prion protein as a determining factor of strain-specific prion replication dynamics. PLoS Pathog. 9, e1003702 (2013).
Cescatti, M. et al. Analysis of Conformational Stability of Abnormal Prion Protein Aggregates across the Spectrum of Creutzfeldt-Jakob Disease Prions. J Virol. 90, 6244–6254 (2016).
Bett, C. et al. Biochemical properties of highly neuroinvasive prion strains. PLoS Pathog. 8, e1002522 (2012).
Feraudet, C. et al. Screening of 145 Anti-PrP Monoclonal Antibodies for Their Capacity to Inhibit PrPSc Replication in Infected Cells. J. Biol. Chem. 280, 11247–11258 (2005).
Langeveld, J. P. M. et al. Rapid and discriminatory diagnosis of scrapie and BSE in retro-pharyngeal lymph nodes of sheep. BMC Vet Res. 2, 19 (2006).
Saborio, G. P., Permanne, B. & Soto, C. Sensitive detection of pathological prion protein by cyclic amplification of protein misfolding. Nature. 411, 810–813 (2001).
Castilla, J. et al. Cell-free propagation of prion strains. EMBO J. 27, 2557–66 (2008).
Moudjou, M. et al. Highly infectious prions generated by a single round of microplate-based protein misfolding cyclic amplification. MBio. 5, e00829–13 (2014).
Taylor, D. M., Fernie, K., Steele, P. J., McConnell, I. & Somerville, R. A. Thermostability of mouse-passaged BSE and scrapie is independent of host PrP genotype: implications for the nature of the causal agents. J Gen Virol. 83, 3199–3204 (2002).
Rubenstein, R. et al. Concentration and distribution of infectivity and PrPSc following partial denaturation of a mouse-adapted and a hamster-adapted scrapie strain. Arch Virol. 139, 301–311 (1994).
Silveira, J. R. et al. The most infectious prion protein particles. Nature. 437, 257–261 (2005).
Tixador, P. et al. The physical relationship between infectivity and prion protein aggregates is strain dependent. PLoS Pathog. 6, e1000859 (2010).
Kim, C. et al. Protease-sensitive conformers in broad spectrum of distinct PrP Sc structures in sporadic Creutzfeldt-Jakob disease are indicator of progression rate. PLoS Pathog. 7, e1002242 (2011).
Colby, D. W. et al. Protease-sensitive synthetic prions. PLoS Pathog. 6, e1000736 (2010).
Sajnani, G. et al. PK-sensitive PrP is infectious and shares basic structural features with PK-resistant PrP. PLoS Pathog. 8, e1002547 (2012).
Zou, W. Q. et al. Variably protease-sensitive prionopathy: a new sporadic disease of the prion protein. Ann Neurol. 68, 162–172 (2010).
Cronier, S. et al. Detection and characterization of proteinase K-sensitive disease-related prion protein with thermolysin. Biochem J. 416, 297–305 (2008).
Lewis, V. & Hooper, N. M. The role of lipid rafts in prion protein biology. Front Biosci. 16, 151–168 (2011).
Wang, F., Wang, X., Yuan, C. G. & Ma, J. Generating a prion with bacterially expressed recombinant prion protein. Science. 327, 1132–1135 (2010).
Nishina, K., Deleault, N. R., Lucassen, R. W. & Supattapone, S. (2004) In vitro prion protein conversion in detergent-solubilized membranes. Biochemistry. 43, 2613–2621 (2004).
Deleault, N. R., Harris, B. T., Rees, J. R. & Supattapone, S. Formation of native prions from minimal components in vitro. Proc Natl Acad Sci USA 104, 9741–9746 (2007).
Millson, G. C., Hunter, G. D. & Kimberlin, R. H. An experimental examination of the scrapie agent in cell membrane mixtures. II. The association of scrapie activity with membrane fractions. Comp Pathol. 81, 255–265 (1971).
Aiken, J. M., Williamson, J. L. & Marsh, R. F. Evidence of mitochondrial involvement in scrapie infection. J Virol. 63, 1686–1694 (1989).
Lewis, V. et al. Prion subcellular fractionation reveals infectivity spectrum, with a high titre-low PrPres level disparity. Mol Neurodegener. 26, 7–18 (2012).
Somerville, R. A. et al. Characterization of thermodynamic diversity between transmissible spongiform encephalopathy agent strains and its theoretical implications. J Biol Chem. 277, 11084–11089 (2002).
Farquhar, C. F., Somerville, R. A. & Bruce, M. E. Straining the prion hypothesis. Nature. 391, 345–346 (1998).
Scott, M. et al. Transgenetic investigations of the species barrier and prion strains in Prion Biology and Diseases (ed. Cold Spring Harbor, N. Y.) 435–482 (Cold Spring Harbor Laboratory Press, 2004).
Fernie, K., Steele, P. J., Taylor, D. M. & Somerville, R. A. Comparative studies on the thermostability of five strains of transmissible-spongiform-encephalopathy agent. Biotechnol Appl Biochem. 47, 175–183 (2007).
Yuan, Q., Telling, G., Bartelt-Hunt, S. L. & Bartz, J. C. Dehydration of Prions on Environmentally Relevant Surfaces Protects Them from Inactivation by Freezing and Thawing. J Virol. 92, e02191–17 (2018).
Van Keulen, L., Bossers, A. & Van Zijdderveld, F. TSE pathogenesis in cattle and sheep. Vet. Res. 39, 24, https://doi.org/10.1051/vetres:2007061 (2008).
Maluquer de Motes, C., Cano, M. J., Torres, J. M., Pumarola, M. & Girones, R. Detection and survival of prion agents in aquatic environments. Water Res. 42, 2465–2472 (2008).
Maluquer de Motes, C. et al. Persistence of the bovine spongiform encephalopathy infectious agent in sewage. Environ Res. 117, 1–7 (2012).
Marín-Moreno, A. et al. An assessment of the long-term persistence of prion infectivity in aquatic environments. Environ Res. 151, 587–594 (2016).
Böhnlein, C., Groschup, M. H., Maertlbauer, E., Pichner, R. & Gareis, M. Stability of bovine spongiform encephalopathy prions: absence of prion protein degradation by bovine gut microbiota. Zoonoses Public Health. 59, 251–255 (2012).
Skovgaard, N. Bovine spongiform encephalitis: the case of an unexpected and rapidly spreading epidemic in cattle with serious consequences. Case Studies in Food Safety and Authenticity. Lessons from Real-Life Situations in Woodhead Publishing Series in Food Science, Technology and Nutrition (ed. Woodhead Publishing) 206–214 (Woodhead Publishing, 2012).
Fischer, M. et al. Prion protein (PrP) with amino-proximal deletions restoring susceptibility of PrP knockout mice to scrapie. EMBO J. 15, 1255–12564 (1996).
Aguilar-Calvo, P. et al. Role of the goat K222-PrP(C) polymorphic variant in prion infection resistance. J Virol. 88, 2670–2676 (2014).
Padilla, D. et al. Sheep and Goat BSE Propagate More Efficiently than Cattle BSE in Human PrP Transgenic Mice. PLoS Pathog. 7, e1001319 (2011).
Scott, M. et al. Transgenic mice expressing hamster prion protein produce species- specific scrapie infectivity and amyloid plaques. Cell. 59, 847–857 (1989).
Scott, M. et al. Propagation of prions with artificial properties in transgenic mice expressing chimeric PrP genes. Cell. 73, 979–988 (1993).
Reed, L. J. & Muench, H. A simple method of estimating fifty per cent endpoint. Am. J. Hyg. 27, 493–497 (1938).
Aguilar-Calvo, P. et al. Effect of Q211 and K222 PRNP Polymorphic Variants in the Susceptibility of Goats to Oral Infection With Goat Bovine Spongiform Encephalopathy. J Infect Dis. 212, 664–672 (2015).
Acknowledgements
This work was supported by grants from the Spanish Ministerio de Economía y Competitividad (AGL2016-78054-R (AEI/FEDER, UE), the French Fondation pour la Recherche Médicale (FRM) (DEQ. 20150331689) and European Community (FP7 222887 “Priority”). A.M.M. was supported by a fellowship from the INIA (FPI-SGIT-2015-02) and P.A.C. was supported by a fellowship from the Spanish Ministerio de Economía y Competitividad (BES-2010-040922). The funders had no role in study design, data collection and interpretation, or the decision to submit the work for publication. We thank Juan Píquer, Irene Prieto, Patricia Lorenzo and Ana Villa for their technical assistance; and the staff of the Biosafety Level 3 animal facility and the Biosafety Office at the CISA-INIA (Valdeolmos-Madrid) for their excellent animal care and work.
Author information
Authors and Affiliations
Contributions
J.M.T., J.C.E. and V.B. contributed in work conception and design. J.M.T., J.C.E., V.B., M.M., P.A.C. and A.M.M. contributed in data acquisition, analysis and interpretation. A.M.M. and J.M.T. drafted the work and all the other authors contributed to revision. J.M.T., J.C.E., V.B., M.M., P.A.C. and A.M.M. approved the submitted version and agreed both to be personally accountable for the author’s own contributions and to ensure that questions related to the accuracy or integrity of any part of the work, even ones in which the author was not personally involved, are appropriately investigated, resolved, and the resolution documented in the literature.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Marín-Moreno, A., Aguilar-Calvo, P., Moudjou, M. et al. Thermostability as a highly dependent prion strain feature. Sci Rep 9, 11396 (2019). https://doi.org/10.1038/s41598-019-47781-6
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-47781-6
This article is cited by
-
Classical BSE dismissed as the cause of CWD in Norwegian red deer despite strain similarities between both prion agents
Veterinary Research (2024)
-
Experimental transmission of ovine atypical scrapie to cattle
Veterinary Research (2023)
-
Cell biology of prion strains in vivo and in vitro
Cell and Tissue Research (2023)
-
Polymorphic Alpha-Synuclein Oligomers: Characterization and Differential Detection with Novel Corresponding Antibodies
Molecular Neurobiology (2023)
-
Stability of BSE infectivity towards heat treatment even after proteolytic removal of prion protein
Veterinary Research (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.