Abstract
Escherichia coli is a regular inhabitant of the gut microbiota throughout life. However, its role in gut health is controversial. Here, we investigated the relationship between the commensal E. coli strain CEC15 (CEC), which we previously isolated, and the intestine in homeostatic and disease-prone settings. The impact of CEC was compared to that of the probiotic E. coli Nissle 1917 (Nissle) strain. The expression of ileal and colonic genes that play a key role in intestinal homeostasis was higher in CEC- and Nissle-mono-associated wild-type mice than in germfree mice. This included genes involved in the turnover of reactive oxygen species, antimicrobial peptide synthesis, and immune responses. The impact of CEC and Nissle on such gene expression was stronger in a disease-prone setting, i.e. in gnotobiotic IL10-deficient mice. In a chronic colitis model, CEC more strongly decreased signs of colitis severity (myeloperoxidase activity and CD3+ immune-cell infiltration) than Nissle. Thus, our study shows that CEC and Nissle contribute to increased expression of genes involved in the maintenance of gut homeostasis in homeostatic and inflammatory settings. We show that these E. coli strains, in particular CEC, can have a beneficial effect in a chronic colitis mouse model.
Similar content being viewed by others
Introduction
Escherichia coli, a member of the Proteobacteria, is a Gram-negative facultative anaerobe with one of the most diverse lifestyles of all microbes and includes commensal, probiotic, and highly pathogenic strains1. This high phenotypic diversity is mirrored by the high genomic plasticity of E. coli, enriched by the acquisition of numerous mobile genetic elements.
The primary habitat of E. coli is the intestine, where it resides as a common and widespread inhabitant of the gut microbiota2. Upon birth, E. coli massively colonizes the gut after fetal exposure to maternal fecal matter and is predominant in the newborn gut microbiota3. Exposition to E. coli may even start in utero, as it is one of the most abundant microbes found in the maternal placental microbiota4. These early colonizers can change the structure and function of the intestinal epithelial in ways that may be critical for healthy microbiota development5 or provide a strong stimulus for B cell-maturation6, therefore contributing to intestinal homeostasis and immune maturation. In adulthood, E. coli remains a major member of the aero-tolerant fraction of the gut microbiota, although outnumbered by anaerobic bacteria, and stabilizes at approximately 107–108 CFU/g of fecal content7. Commensal E. coli resides on the mucus layer, for which its metabolism has adapted to use as a source of essential nutrients8, near the intestinal mucosa, due to the radial oxygen gradient9. Thus, this bacterial species, with its own nutritional requirements and metabolic capacity distinct from that of anaerobic bacteria, may play a specific role in the crosstalk between commensal bacteria and the host.
Several studies have assigned indigenous E. coli as pathobiont, a term that describes commensal microbes that induce disease only in certain genetic or environmental contexts10. Indeed, the disruption of the microbiota composition (or dysbiosis) that accompanies several human diseases is characterized by the expansion of Enterobacteriaceae, including E. coli. This has been shown for Crohn’s disease (CD)11, with a high prevalence of pro-inflammatory adherent-invasive E. coli (AIEC)12, colorectal cancer13, and, to a lesser extent, ulcerative colitis (UC)14. Similarly, the abundance of intestinal Escherichia also increases in several mouse models of inflammatory bowel disease (IBD)15,16. A more direct effect of certain indigenous strains of E. coli on intestinal inflammation has been demonstrated in gnotobiotic mouse models with a predisposition to inflammation. Thus, mono-association of mice genetically prone to inflammation with E. coli strains, originally isolated from mice gut microbiota, results in intestinal inflammation17,18,19.
However, the role of indigenous E. coli toward gut health is far from clear and requires further investigation. Indeed, another commensal E. coli isolate failed to induce disease in antibiotic-pretreated IBD-susceptible mice, despite robust colonization20. The inflammation-prone HLA B27 transgenic rat model responds only very moderately to E. coli strains isolated from CD patients, whereas other bacteria induce severe colitis21. Studies in a mono-associated IL2−/− mice model report divergent effects of indigenous E. coli strains in the induction of colitis19. Furthermore, E. coli strains originally isolated from human gut microbiota are the basis of at least two commercially available probiotic products, commercialized under the names Mutaflor and Symbioflor2. The E. coli Nissle 1917 (Nissle) strain, the active component of Mutaflor, is one of the most thoroughly investigated and documented probiotics22. Clinical trials have shown a beneficial effect of Nissle for the maintenance of remission in UC, similar to that of mesalazine23.
We previously isolated a primo-colonizing E. coli strain called CEC15 (CEC) from freshly pooled fecal samples of 15-day-old suckling rodents as a major representative of this environment. We reported that this E. coli strain elicits sequential remodelling of the colonic epithelium in gnotobiotic rodent models, affecting different arms of the intestinal epithelial defences required to achieve a microbiota-accommodating homeostasis5.
Here, we aimed (1) to gain further insight into the relationship between CEC and host intestinal health and (2) to compare the intestinal host response to CEC to that of the probiotic E. coli Nissle. We explored the ileal and colonic host response to these two strains under three different conditions: (i) disease-free mono-associated wild-type (WT) mice, (ii) mono-associated IL10 deficient (IL10−/−) mice, predisposed to intestinal inflammation; (iii) conventional IL10−/− mice exposed to inflammatory challenge as a chronic colitis model.
In summary, our data show that the host response to commensal strains of E. coli mobilized key genes that are involved in sustaining the symbiotic relationship with the gut microbiota. We further demonstrate that the host can over-mobilize its defence mechanisms when mono-associated with commensal E. coli in a setting in which the animals are predisposed to inflammation. Finally, we found that the impact of CEC is beneficial to the host in a chronic colitis IL10−/− mouse model.
Methods
Ethics approval
All procedures involving animal experimentation were carried out according to the European guidelines for the care and use of laboratory animals under the authority of a license issued by the French Veterinary Services (authorization number 78–122 specific to CC) and were approved by the French “Ministère de l’Enseignement Supérieur et de la Recherche” (authorization number APAFIS#3441-2016010614307552). Experiments involving gnotobiotic WT or IL10−/− mice were performed at the Anaxem facility of the MICALIS Institute (INRA, Jouy-en-Josas, France), which is accredited by the French “Direction Départementale de la Protection des Populations (DDPP78)”, accreditation number A78-322-6. Experiments involving conventional (CV) WT or IL10−/− mice were performed at the IERP facility (INRA, Jouy-en-Josas, France), which is accredited by the French “Direction Départementale de la Protection des Populations (DDPP78)”, accreditation number DDPP-VET-13-0124. GF WT mice and GF IL10−/− mice (generated from the CV IL10−/− mice (B6.129P2-Il10tm1Cgn/J; see below) using the standard procedure of cesarean delivery), were purchased from the GF rodent breeding facilities of the CNRS-TAAM (transgenesis, archiving and animal models) center (Orléans, France). They were delivered to Anaxem under sterile conditions and immediately transferred to the experimental isolator. After reception, GF mice were left undisturbed for eight days before starting the experiment. Conventional WT and IL10−/− (B6.129P2-Il10tm1Cgn/J)24 mice were born and bred at the IERP under standard conditions. The absence of IL10 gene expression was verified in IL10−/− GF and IL10−/− CV mice. All WT GF and CV mice used in experiments were C57BL/6. The genetic background of IL10−/− mice is a mix of C57BL/6 and 129/Ola. All animals used in this study were males.
Gnotobiotic mice
Six groups of gnotobiotic mice were studied (Supplementary Fig. 1A,B): either GF WT (GFWT), or IL10−/− (GFIL10); two groups of mono-colonized WT mice inoculated with either E. coli CEC or E. coli Nissle (CECWT and NissleWT groups, respectively) and two groups of mono-associated IL10−/− mice inoculated with either E. coli CEC or E. coli Nissle (CECIL10 and NissleIL10 groups, respectively). The absence of microbes was verified in GF mice by microscopic observation of fresh feces and culturing of fecal material on various bacterial culture media. All mice were maintained in Trexler type isolators and received the same standard diet (ad libitum, R03-40 SAFE sterilized by gamma irradiation at 45 kGy). CEC and Nissle inocula were prepared from fresh overnight cultures. Bacterial pellets were obtained by centrifugation (4 °C; 20 min; 4,700 × g), re-suspended in sterile PBS, immediately introduced into the isolator and 100 µL (containing 108 bacteria) used to inoculate GF mice. All mice were 8 to 10 weeks old at the time of inoculation and were sacrificed 21 days post-inoculation. Each experiment was carried out independently at least three times.
Experimental colitis in CV IL10−/− mice
Experimental chronic colitis was induced in six-to-eight-week-old CV IL10−/− mice by rectal injection with dinitrobenzene sulfonic acid (DNBS) (Supplementary Fig. 1C). The protocol was carried out according to25, except for the dose of DNBS, which was lower in our study. Prior to DNBS administration, mice were anesthetized with Xelamine/Ketamine and a 10 cm long piece of PE-90 tubing was attached to a syringe and inserted 3.5 cm into the colon. On the first day of the protocol, mice received one rectal dose of 150 mg/kg DNBS (ICN Biomedical Inc., Santa Ana, CA) in 30% ethanol. All mice received a subcutaneous injection of 1 ml saline solution (0.9% NaCl) for three days to prevent dehydration. Mice were allowed to recover for 21 days and then received a second DNBS injection at day 21, reactivating inflammation. Ten days before sacrifice, DNBS treated CVIL10−/− mice were given a dose of 1 × 109 CFU of CEC (DNBSIL10 + CEC) or Nissle (DNBSIL10 + Nissle) daily by oral gavage, or the same volume of PBS as a positive control for the disease (DNBSIL10) (Supplementary Fig. 1C). Mice were sacrificed 24 days after the first DNBS injection, i.e. three days after the second. Run in parallel, CV IL10/− mice received PBS instead of DNBS and were supplemented daily with PBS (Control), as a negative control group.
Sample collection
The day of sample collection, at 9:00 AM, mice were anesthetized with isoflurane, killed by cervical dislocation, and the ileum and colon promptly removed. An unflushed section of ileum and colon was kept and fixed in CARNOY (four weeks at 4 °C) for mucus layer thickness measurements as in5. The remainder of the intestinal tissues was quickly washed with PBS and immediately used either for ex vivo permeability measurements in an Ussing chamber, or fixed in 4% paraformaldehyde (PFA; 6 h at room temperature) for further histological analyses, or frozen at −80 °C for RNA extraction. Cecal contents were recovered and frozen at −80 °C before DNA extraction for 16S rRNA gene sequencing.
Histology and Immunostaining
CARNOY- and PFA-fixed ileum and colon were dehydrated and embedded in paraffin according to a standard protocol26. Staining was performed with Dako reagents according to the recommendations of the manufacturer. For PFA fixed samples, antigen retrieval was performed by boiling the slides for 40 min in 10 mM tri-sodium citrate pH 6.0. The primary antibodies used and corresponding dilutions were: anti-cadherin1 (Cdh-1, Invitrogen; 1/50), anti-CD3 (Abcam, 1/500), anti-mucin-2 (Muc2, Santa Cruz; 1/500), anti-mucin-13 (Muc13, Santa Cruz; 1/500), and anti-Ki67 (Dako; 1/50). Negative controls were performed by omitting the primary antibody from the reaction. For each section, Ki67+ cells were counted for 10 crypts and the results expressed as the percentage of total crypt cells. CD3+ cells were counted by microscopic field (x200). For mucus thickness analyses, CARNOY-fixed cuts were stained with the Muc-2 antibody. Tissues were observed with a 3DHistech Panoramic Scan and signal quantification was performed using Panoramic Viewer® or ImageJ software.
RNA extraction and cDNA preparation for real-time RT-PCR experiments
RNA isolation was carried out using the Ambion mirVana Kit (Life Technologies) according to the manufacturer’s recommendations. The RNA quality of all samples was checked using an Agilent 2100 Bioanalyzer (Agilent Technologies, Santa Clara, CA, USA) with an average RNA integrity number superior to 9 on a scale of 0 to 10. Complementary DNA (cDNA) was prepared using the RT RevertAid H Minus first Strand cDNA Synthesis Kit (Life Technologies).
Gene expression profiling using the TaqMan OpenArray System
For studies in gnotobiotic WT or IL10−/− mice, gene expression analysis was performed with a customized TaqMan OpenArray Real-time PCR System (Life Technologies). We designed two TaqMan OpenArrays, one dedicated to the ileum, the other to the colon, following a thorough review of the literature and searching the NCBI public repository Gene Expression Omnibus. For each TaqMan OpenArray, 220 genes were selected, plus four endogenous control genes (actb, gapdh, ubc, and tpb). The candidate genes selected for this study survey various functions of ileal and colonic cells, mainly involved in mucosal defence: immunological responses, intestinal barrier, oxidative stress, anti-microbial peptides, cellular signalling, regulation of cell proliferation and differentiation, detoxification, DNA-damage detection, growth factors, inflammasome, inflammation pathway, lipid synthesis and metabolism, pattern recognition, and solute transport (Supplementary Table 1–2, for the list of genes included in the ileum and colon array cards, respectively). For studies in DNBS treated Il10−/− mice, gene expression profiling was performed using the TaqMan OpenArray Mouse Inflammation Panel plate (Life Technologies) designed by the manufacturer, that consists of 632 gene targets selected for their involvement in inflammatory responses. The cDNA (10 µl) was mixed with the TaqMan OpenArray Real-Time PCR Master Mix and loaded onto the cards using the AccuFill™ System. The cards were cycled in an OpenArray NT Cycler System (Applied Biosystems) at the integrative microgenomic platform (@BRIDGe, INRA, Jouy-en-Josas) following the manufacturer’s protocol. The same sample was systematically loaded in each TaqMan OpenArray and used to check the reproducibility between the plates. Data were extracted using OpenArray Real-Time qPCR Analysis software (Applied Biosystems). The fold-change in gene expression (Rq or relative quantification) was calculated using the comparative 2−ΔΔCq method with global normalization of all gene expression data using GenEx software (Multid Analyses)27. The GF group was used as a calibrator for the gnotobiotic experiments and the Control group for gene expression profiling of the DNBS treated CV IL10−/− mice.
Principal component analysis, heatmap representation, and statistical analysis of the TaqMan OpenArray data
The R statistics environment was used for data analyses. Data obtained from TaqMan OpenArray experiments are represented either by Principal Component Analysis (PCA) plots or heatmaps. Rq values were used to build the PCA plots and heatmaps. Prior to PCA, we filtered the data using a one-way permutation test (oneway_test, R package “coin”) to remove genes from the datasets for which the variation in expression was less informative (threshold set to p < 0.01). PCA was then carried out on the filtered dataset using the R packages FactoExtra and FactoMineR. The number of genes included in the PCA analysis is specified in the figure legend for each comparison. Heatmaps were generated for genes showing a significant difference between the mono-associated mice and the GF group (p < 0.05; Man-Whitney test). Clustering on the gene expression profile was applied to the genes in the heatmap.
Single real-time quantitative PCR analyses of gene expression
Single real-time quantitative PCR assays were used to confirm the results obtained on the TaqMan Open Arrays system using the corresponding TaqMan assay. All gene expression results are expressed with the 2−∆∆Ct method (Rq), using gapdh as the housekeeping gene and values from the GF (gnotobiotic experiments) or Control (Chronic colitis experiments) mice as calibrators27.
Ex vivo intestinal permeability measurements
After removal, segments of the ileum and colon were immediately mounted in Ussing chambers, as previously described5. Para-cellular permeability was further assessed by measuring mucosal-to-serosal flux of 4 kDa non metabolizable fluorescein isothiocyanate-labeled dextran (FD4) or 0.4 kDa FITC-sulfonic acid (FSA) for 90 min. Molecules were added to the mucosal side of the chamber at a final concentration of 0.4 mg/mL and the fluorescence intensity determined at the serosal side. Trans-epithelial conductance was measured by clamping the voltage and recording the change in the short-circuit current (Isc). At the end of the experiment, tissues were challenged with the cholinergic analog carbachol (CCh) on the serosal side (100 mM) and the ΔIsc was recorded to check the viability of the tissue.
Myeloperoxidase activity
Myeloperoxidase (MPO) activity was assayed according to25. MPO activity is expressed as the units per milligram of total protein. Lowry’s method was used for protein quantification.
16S rRNA gene sequencing and analysis
Cecal DNA of CV IL10−/− mice was extracted as described in28 and the V3-V4 hyper-variable region of the 16S rRNA gene amplified with the primers F343 (CTTTCCCTACACGACGCTCTTCCGATCTTACGGRAGGCAGCAG) and R784 (GGAGTTCAGACGTGTGCTCTTCCGATCTTACCAGGGTATCTAATCCT). The PCR amplicons were sent to the GeT-PlaGe platform (INRA, Toulouse) for sequencing using Illumina Miseq technology. Single multiplexing was performed and the multiplexes purified and loaded onto the Illumina MiSeq cartridge according to the manufacturer’s instructions. Raw sequences were analyzed using the FROGS pipeline to obtain the OTU (operational taxonomic units or phylotypes) abundance table, phylogenetic tree, and taxonomic table using the default parameters29. Statistical analyses of the 16S rRNA sequences were performed from the FROGS-generated outputs using R and environment version 3.2.3. Calculations of within-community diversity (α-diversity), between-community diversity (β-diversity), and relative abundance taxonomic summaries were performed using the add-on package “Phyloseq”.
Statistical analyses
R 3.3.1 and GraphPad 5 software were used to produce graphs and for statistical analyses. Data are expressed as the mean ± standard deviation in scatter plots. Differentially expressed genes from OpenArray data were obtained with non-parametric multiple comparison tests using false discovery rate (FDR) corrected p-values (the threshold for significance of differential expression was set to p < 0.05). Comparisons of other quantitative variables were performed using the non-parametric Mann-Whitney test (p < 0.05 was considered significant).
Results
Ileal and colonic gene expression profiles of CEC or Nissle mono-associated mice differ from those of GF mice
We carried out gene expression profiling by high-throughput open-array qPCR of the ileum and colon of CECWT, NissleWT, and GFWT mice (Fig. 1) using two customized gene-expression array plates. One was designed for the ileum and the other the colon, with a selection of candidate genes mainly involved in mucosal defence of the intestine (Supplementary Tables 1, 2). PCA revealed a clear separation of the gene expression profile between mono-associated and GFWT mice, both in the ileum (Fig. 1A) and colon (Fig. 1B). Accordingly, in the ileum, PC1 separated the mono-associated CECWT and NissleWT groups from the GFWT group, with both mono-associated groups overlapping. In the colon, PC1 also separated the gene expression profiles of the GFWT and mono-associated groups. The mono-associated CEC and Nissle groups tended to slightly diverge along PC2, revealing a stronger effect of the CEC strain on the colonic gene expression profile. We also included CVWT mice for comparison with the CECWT, NissleWT, and GFWT mice (Supplementary Fig. 2). As expected, PCA first separated the CVWT from all gnotobiotic groups along PC1. In the ileum and colon, the CECWT and NissleWT groups clustered together and were distinct from the GFWT and CVWT groups (Supplementary Fig. 2).
Intestinal gene expression profiles differ between GF and E. coli mono-associated WT mice. Germ-free wild-type mice (GFWT) were inoculated with the commensal CEC (CECWT) or probiotic Nissle 1917 (NissleWT) E. coli strain. Mono-colonized mice, CECWT and NissleWT, were sacrificed 21 days post-inoculation and the ileal and colonic gene expression profiles analyzed by high-throughput qPCR using TaqMan OpenArrays. (A) and (B) principal component analysis (PCA) plots of gene expression data in the ileum and colon, respectively. After pre-filtering the dataset, 64 genes were included for the PCA, both for the ileum and colon; n = 9–10 mice per group. (C) and (D) Gene-expression heatmaps for the ileum and colon, respectively. Genes that are shown are significantly differentially expressed relative to GFWT.
We next selected a subset of genes from high-throughput qPCR data that were differentially expressed between the GFWT and mono-associated groups and exhibited a Rq value > 1.5. Gene expression variation were plotted in a heatmap (Fig. 1C for the ileum and Fig. 1D for the colon). Again, there was a clear impact of the CEC and Nissle strains on the gene expression profile at both sites of the intestine (Fig. 1C,D). Differences in gene expression were confirmed in single TaqMan assays (Examples given in Supplementary Fig. 3 for the ileum and Supplementary Fig. 4 for the colon). Genes involved in several key intestinal functions in the ileum and colon of mice mono-associated with CEC or Nissle were upregulated. Indeed, genes were upregulated for: i) enzymes involved in ROS/RNS turnover, such as duox2 (dual oxidase 2), duoxa2 (dual oxidase activator 2), and nos2 (nitric oxide synthase 2) for both the ileum and colon; gpx2 (glutathione peroxidase 2) for the ileum only; and nox1 (NADPH oxidase 1) for the colon only; ii) antimicrobial peptide production and barrier functions: reg3-γ, reg3-β (regenerating islet-derived-γ and -β) and pla2g2a (secretory phospholipase A group IIA) for both the ileum and colon, ang4 (angiogenin-4) for the ileum only, and fut2 (α(1,2)-fucosyltransferase) for the colon only; and iii) factors involved in the immune response, such as IL18 (interleukin-18), cxcl10 (C-X-C motif chemokine ligand 10), tnfα (tumor necrosis factor α) and tap1 (transporter associated with antigen processing) for both the ileum and colon; IL2rb, IL4ra and IL2rg for the ileum only; and IL1β, IL6, IL17d, thpo (thrombopoietin), and ifnγ (interferon-γ) for the colon only (Fig. 1C,D) and (Supplementary Figs 3 and 4). Furthermore, CEC or Nissle modulated genes involved in the transport function of the intestine, such as the fructose transporter slc2a5 (solute carrier family 2 member 5; previously called glut5) and members of the aquaporin water channel family aqp3 and aqp7, which were dramatically upregulated in the ileum of CECWT and NissleWT mice, and the gene clca4 (Ca 2+-activated chloride channel 4), which was upregulated in the colon (Fig. 1C,D) and (Supplementary Figs 3 and 4).
CEC and Nissle maintain intestinal barrier function and epithelial integrity in gnotobiotic WT mice
We further investigated the impact of the CEC and Nissle strains on the barrier formed by the network of intestinal mucins, as it is a key player in physico-chemical protection. We examined the thickness of the mucus layer formed by the highly glycosylated secreted mucin2 (Muc2), and the abundance of membrane-associated mucins, Muc13 and Muc4. Muc2 staining showed the ileal mucus layer to be thicker in mono-colonized CECWT and NissleWT mice than GFWT mice, tending to become similar to that of CVWT mice (Fig. 2A,B). Similarly, Muc13 staining in the ileum of both CECWT and NissleWT mice was stronger than that of control GFWT mice (Fig. 2C), but not in the colon (Supplementary Fig. 5A). There were also no modifications in Muc4 staining for any of the groups (Supplementary Fig. 5B). We also investigated the effect of the two strains on expression of the epithelial proliferation marker Ki67 (Fig. 2D). In the NissleWT group, the percentage of Ki67+ cells was higher than that in the GFWT group in both the ileum and colon (Fig. 2D), whereas there was a significant increase in the percentage of proliferative epithelial cells in only the colon in the CECWT group (Fig. 2D).
Intestinal barrier function is preserved in E. coli mono-associated WT mice. Analyses of intestinal barrier markers were carried out on germ-free wild-type mice (GFWT), on mice mono-colonized with the CEC (CECWT) or Nissle (NissleWT) strains, and on conventional WT mice (CVWT). (A) Representative images of Muc2 (green) immunostaining of CARNOY-fixed tissues. (B) Thickness measurement of the ileal mucus layer (green layer at the top of the epithelium), n = 7–15 mice per group. (C) Muc13 (red) immunostaining of PFA-fixed tissues. Nuclei are stained with DAPI (blue). (D) Percentage of Ki67+ cells per total cells per crypt for the ileum and colon, n = 7–9 mice per group. (E) and (F) Analysis of ex vivo para-cellular permeability using the Ussing chamber system with (E) FITC-dextran (4 KDa; FD4) and (F) FITC-sulfonic acid (400 Da; FSA) or for the ileum and colon, n = 5–17 mice per group. (G) Immunostaining of the adhesion protein cadherin-1 (Cdh-1) in the ileum of PFA-fixed tissues. (H) Mean intensity of Cdh-1 staining, n = 5–7 mice/group. All values are presented as the means ± SEM. Mean values with letter designations are significantly different (non-parametric Mann-Whitney test).
We then assessed the effect of E. coli strains on intestinal permeability in the ileum and colon. We monitored the para-cellular passage from the mucosal to serosal side of high- and low-molecular weight molecules, FD4 (Fig. 2E) and FSA (Fig. 2F), respectively, in an Ussing chamber. There were no differences in permeability between the groups. Among the panel of tight junction or cell adhesion proteins we tested [claudins-4,-5,-7,-8,-12,-15, -17, ZO-1 and F11r (data not shown), Cld2 and Cld3 (Supplementary Fig. 5C,D)], we only observed greater immunostaining of cadherin-1 (Cdh1) in the CECWT than the GFWT mice (Fig. 2G,H). These data show that intestinal barrier function and epithelial integrity are preserved in gnotobiotic WT mice.
Key genes involved in mucosal defense and immune response are strongly mobilized in IL10−/− mono-associated mice
Gnotobiotic IL10−/− mice have been previously used as a model to investigate the colitis-inducing potential of individual bacterial strains, including indigenous strains of E. coli17,18. The colitis that IL10−/− mice develop requires microbial exposure, as GF mice are protected from disease17,18. We investigated the effect of CEC and Nissle in mono-associated IL10−/− mice, three weeks post-colonization, a time interval previously used to observe the effect of colitogenic indigenous E. coli strains in 129S6/SvEv IL10−/− mice18. There was no weight loss nor weight differences between GFIL10, CECIL10, and NissleIL10 mice (data not shown). There was also no histological mucosal damage to the intestines of CECI1L0 or NissleIL10 mice (Supplementary Fig. 6).
We further investigated the effect of CEC and Nissle on the ileal and colonic gene expression profile in gnotobiotic IL10−/− mice. We first randomly sampled four of the 10 individuals belonging to the GFIL10 and CECIL10 groups to assess the ileal gene expression profile using the customized gene-expression array plates. The data obtained were then compared to those previously obtained for GFWT and CECWT mice. PCA showed the CECIL10 group to strongly diverge from the others along PC1 (Fig. 3A). There was a striking difference of the gene expression profile from that of the CECWT group, and the difference between the CECIL10 and GFIL10 groups was higher than that between the CECWT and GFWT groups. This initial screening suggests that the intestinal responses to E. coli of IL10−/− mice are different from those of WT mice. Indeed, CEC and Nissle may have a stronger effect on the expression of ileal genes in IL10−/− than WT mice, such as shown for ileal reg3β and nox1 by qPCR single assays (Fig. 3B).
Key genes involved in mucosal defense and immune response are mobilized in IL10/- mono-associated mice. Gene expression profiling of the ileal epithelium of germ-free wild-type or IL10−/− mice (GFWT, GFIL10), or wild-type or IL10−/− mice mono-associated with the CEC (CECWT; CECIL10) or Nissle (NissleWT; NissleIL10) strain for 21 days. (A) Principal component analyses (PCA) plot of gene expression data in the ileum of GFWT, CECWT, GFIL10, and CECIL10; 61 genes were included in the PCA; n = 4–9 mice per group. (B) and (C) Relative gene expression (Rq), measured with single TaqMan Assays, of the ileum (reg3β and nox1) and colon (duox2, nox1, nos2, reg3γ, pl2g16, fut2, IL1β, IL22, stat1 and nlrc5) of GFIL10, CECWT, CECIL10, NissleWT, NissleIL10, and CVWT relative to GFWT, n = 8–11 mice/group. All values are presented as the means ± SEM. Mean values with letter designations are significantly different (non-parametric Mann-Whitney test).
We investigated whether this difference in the effect of CEC and Nissle between IL10−/− and WT mice also occurred in the colon by targeting key genes involved in mucosal defense using single qPCR assays (Fig. 3C). Thus, we investigated colonic expression of genes involved in the ROS/RNS turnover (duox2, nox1, nos2), antimicrobial peptide production (reg3γ and pl2g16), the mucosal intestinal barrier (fut2), immune response (IL1β,IL22), as well as stat1, which orchestrates antimicrobial responses and nod-like receptors 5 (nlrc5), an intracellular protein involved in the detection of microbes. These genes were more highly expressed in IL10−/− than WT mice, when they are mono-associated with CEC or Nissle (Fig. 3C).
The differential effect of the strains on IL10−/− and WT mice was not due to colonization levels, as both strains similarly colonized the WT and IL10−/− mice at a level of 1010 bacteria/g of stool (Supplementary Fig. 7). Overall, these results show that genes involved in intestinal defense mechanisms are strongly mobilized in the presence of CEC and Nissle in IL10−/− mice.
Intestinal barrier function is not altered in Il10−/− mice mono-colonized with CEC and Nissle
We assessed different markers to investigate whether intestinal barrier function, at the ileal and colonic level, is modified in Il10−/− mice mono-colonized with CEC and Nissle: Ki67 (Fig. 4A,B), cadherin1 (cdh1), a protein involved in cell adhesion (Fig. 4C,D) and the thickness of the mucus layer (Fig. 4E,F). The number of proliferative epithelial cells in the ileum and the colon of IL10 deficient mono-colonized mice was higher than that of the GFIL10 group (Fig. 4A,B for the ileum and colon respectively). Ileal Cdh1 staining was greater in mono-colonized IL10 deficient mice than in GFI1L0 mice (Fig. 4C,D). Furthermore, the mucus layer in the ileum was thicker in the 2 mono-colonized groups than in GFIL10 mice, similar to our observations for WT mice (Fig. 4E,F). But in contrast to WT mice, this increase also occurred in the colon of IL10−/− mice (Fig. 4E,F). Intestinal permeability was assessed in vitro at the level of the ileum and the colon. No difference in FSA passage among the groups was observed (Fig. 4G). In contrast, the ileal transepithelial conductance of CECIL10 and NissleIL10 mice was lower that than of the GFIL10 mice (Fig. 4H), suggesting a tendency to a lower intestinal permeability. Taken as a whole, this data reveals that CEC and Nissle do not alter intestinal permeability in a model of predisposition to inflammation. These strains can even impact positively on some markers of epithelial barrier function.
CEC15 and Nissle 1917 strengthen parameters of the intestinal barrier function in gnotobiotic Il10−/− mice. (A,B) Percentage of Ki67+ cells per total cells per crypt of the ileum (A) and colon (B) of germ-free IL10−/− mice (GFIL10), or IL10−/− mice mono- colonized with the CEC (CECIL10) or Nissle 1917 (NissleIL10) strain, n = 8–9 mice/group. (C) immunostaining of the tight junction protein cadherin-1 (Cdh-1) of the ileum of PFA-fixed tissues. (D) Mean intensity of Cdh-1, n = 5–7 mice/group. (E) Muc2 immunostaining of the ileum and colon of GFIL10, CECIL10 and NissleIL10 mice. Tissues were fixed in CARNOY and nuclei were stained with DAPI (blue). (F) Quantification of mucus layer thickness (red layer on top of the epithelium) in the ileum and colon, n = 7–11 mice/group. (G) Analysis of ex vivo para-cellular permeability with FITC-sulfonic acid (400 Da; FSA) of the ileum and colon. (H) Conductance of the ileum, n = 10–13 mice/group. (G,H) were measured using the Ussing chamber system, n = 5–19 mice/group. All values are presented as the means ± SEM. Mean values with letter designations are significantly different (non-parametric Mann-Whitney test).
CEC and Nissle partially reduce the severity of inflammation of DNBS-treated CV IL10−/− mice
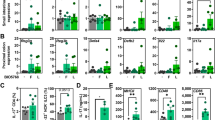
We next investigated the effect of CEC on inflammation using DNBS-treated CV IL10−/− mice as a model of a chronic intestinal colitis. Although the Ameho score was unexpectedly close to zero (Supplementary Fig. 8), DNBS treated CV IL10−/− mice exhibited several signs of inflammation that were apparent in both the ileum and colon: the mice showed a decrease in body weight (Fig. 5A), shortening of the small intestine and colon (Fig. 5B), increased myeloperoxidase (MPO) activity, a marker of neutrophil infiltration (Fig. 5C), infiltration of immune cells (CD3+ T lymphocytes) (Fig. 5D,E), a pro-inflammatory gene expression profile (Supplementary Fig. 9A,B).
CEC15 decreases inflammation markers in DNBS-treated CV IL10−/− mice. Conventional IL10−/− mice treated intra-rectally with dinitrobenzene sulfonic acid (DNBS) were supplemented with CEC15 (DNBSIL10 + CEC) or Nissle 1917 (DNBSIL10 + Nissle) or PBS (DNBSIL10); a group of mice, that received intra-rectally PBS instead of DNBS, was used as a negative control for the disease (Control). (A) Mean weight evolution during the 24 days of the experiment. DNBS was injected on days 1 and 21, n = 19–26 mice/group. (B) Length of the small intestine and colon, n = 10–14 mice/group. (C) Levels of myeloperoxidase (MPO), a neutrophil infiltration marker, in the ileum and colon, n = 9–14 mice/group. (D) Number of CD3+ cell in the colonic mucosa, n = 9–12 mice/group. (E) Representative picture of CD3+ (white arrows) staining. All values are presented as the means ± SEM. Mean values with letter designations are significantly different (non-parametric Mann-Whitney test).
Although CEC and Nissle were unable to induce recovery of the body weight in the model (Fig. 5A), both strains rescued the inflammation-associated reduction of ileal and colon length and MPO activity to control values (Fig. 5B,C). CEC had an additional positive effect on the number of ileal and colonic CD3+ T lymphocytes (Fig. 5D,E). Moreover, the permeability of the intestinal barrier did not increase after CEC treatment (Supplementary Fig. 10).
The gene expression profile of E. coli DNBS-treated groups was skewed towards an anti-inflammatory profile, close to that of the control group
We then analysed the inflammatory profile in Control (negative control for the disease), DNBSIL10, and DNBSIL10 + CEC/Nissle mice by focusing on a panel of genes involved in inflammation. Genes for which the resulting Rq values showed them to be significantly modified by DNBS treatment and rescued by E. coli strains, were selected and the corresponding Rq values plotted in a heatmap for the ileum and colon (Figs 6A and 7A).
Ileal gene expression reverts towards the control profile when DNBS-treated IL10−/− mice receive CEC15 or Nissle1917. Conventional IL10−/− mice were treated intra-rectally with dinitrobenzene sulfonic acid (DNBS) and supplemented with CEC15 (DNBSIL10 + CEC), Nissle 1917 (DNBSIL10 + Nissle) or PBS (DNBSIL10), as a positive control for the disease; a group of mice, that received intra-rectally PBS instead of DNBS, was used as a negative control for the disease (Control). Gene expression profiling of the ileal mucosa was performed using the TaqMan Open Array system Mouse Inflammatory Panel. (A) Heatmap of the relative gene expression (Rq) to that of the Control group of the main modified genes of the ileum (B) Relative expression (Rq), measured with single TaqMan Assays of IL6, IL17α and ltf (lactotransferrin) in the ileum;n = 8–10 mice/group. All values are presented as the means ± SEM. Mean values with letter designations are significantly different (non-parametric Mann-Whitney test).
Supplementation of DNBS-treated IL10−/− mice with CEC15 shifts colonic gene expression towards an anti-Inflammatory profile. Conventional IL10−/− mice were treated intra-rectally with dinitrobenzene sulfonic acid (DNBS) and supplemented with CEC15 (DNBSIL10 + CEC), Nissle 1917 (DNBSIL10 + Nissle) or PBS (DNBSIL10), as a positive control for the disease; a group of mice, that received intra-rectally PBS instead of DNBS, was used as a negative control for the disease (Control). Gene expression profiling of the ileal mucosa was performed using the TaqMan Open Array system Mouse Inflammatory Panel. (A) Heatmap of the relative gene expression (Rq) to that of the Control group of the main modified genes of the colon. (B) Relative expression (Rq), measured with single TaqMan Assays, of a2m (alpha-2-macroglobulin), IL6, IL17α, orl1 (oxidized low density lipoprotein receptor 1), clu (clusterin) and cfi (complement component factor I) in the colon, n = 8–10 mice/group. All values are presented as the means ± SEM. Mean values with letter designations are significantly different (non-parametric Mann-Whitney test).
The gene expression profile in the ileum of the control group was partially restored when DNBSIL10 mice were treated with the commensal E. coli strains CEC or Nissle (Fig. 6A). Thus, single TaqMan assay experiments confirmed that increased expression of interleukins such as IL6, IL17α and ltf (lactotransferrin) observed for DNBSIL10 relative to those of the control group was counteracted when mice received CEC or Nissle (Fig. 6B).
Similarly, the gene expression profile observed in the colon of DNBSIL10 + CEC was close to that of the control group (Fig. 7A). The effect of CEC was stronger than that of Nissle (Fig. 7A), with the recovery of Control expression levels for genes, such as the AMPs (camp, pla2g2e, reg3α, and reg3β) and several cytokines (tnfα, cxcl11, IL22 and IL6), following E. coli treatment. Single TaqMan assays confirmed that the increased expression of the a2m (alpha-2-macroglobulin), IL6, IL17α, orl1 (oxidized low density lipoprotein receptor 1), clu (clusterin) and cfi (complement component factor I) genes following DNBS treatment decreased to Control values when the DNBS treated mice received CEC (Fig. 7B).
The effect of CEC on gene expression is independent of an effect on gut microbiota composition
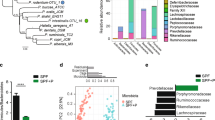
We investigated whether the effect of the E. coli strains involved modifications of the gut microbiota by analyzing the intestinal bacterial composition of the different experimental groups. We found no differences in the α-diversity, which measures the taxonomic richness of the cecal microbiota communities, between groups (Fig. 8A). In addition, β-diversity analysis, which measures the degree of similarity between the gut microbial communities, revealed no clustering of the mice according to the DNBS or E. coli treatments (Fig. 8B). At the phylum level, there was a decrease in the relative abundance of Firmicutes and an increase in that of Proteobacteria in the DNBSIL10 mice relative to that of the Control group, revealing dysbiosis in our model (Fig. 8C). Neither CEC nor Nissle treatment corrected the changes in microbiota composition induced by DNBS treatment.
CEC15 supplementation does not affect the composition of the gut microbiota. Conventional IL10−/− mice were treated intra-rectally with dinitrobenzene sulfonic acid (DNBS) and supplemented with CEC15 or Nissle 1917; PBS supplementation was used as a control. Analysis of the microbiota composition was performed by 16S rRNA gene sequencing of the cecal content. (A) α-diversity of the cecal microbiota measured with the Shannon index (a calculation method to measure the richness of the community (α-diversity) (B) β-diversity (measure of between-community diversity) of the cecal microbiota communities measured with the unweighted unifrac index. (C) Phylum abundance of Firmicutes, Bacteroidetes, Actinobacteria, and Proteobacteria. For all analyses n = 9–12 mice/group. All values are presented as the means ± SEM. Mean values with letter designations are significantly different (non-parametric Mann-Whitney test).
Discussion
Most studies demonstrating that several human diseases are associated with an expansion in the gut of Enterobacteriaceae, including E. coli, are based on sequencing data. However, this approach has insufficient resolution to detect genetically similar organisms of differing ecophysiology and impact on health. Considering phenotypic diversity at the strain level would aid the assessment of how resident E. coli may affect gut health and disease outcomes, as demonstrated by a recent study30.
Here, we extend our knowledge on a commensal E. coli strain that we previously isolated from suckling rodents, that we named CEC15, showing dynamic transcriptional responses of the ileum and the colon to CEC15. The expression of genes that play a key role in mucosal defence and the maintenance of mutualistic host-microbiota interactions was elevated in CEC mono-associated mice, similar to observations made with the probiotic Nissle 1917. Indeed, we show a core-response to both CEC and Nissle strains, in both the ileum and colon. This includes a set of genes involved in ROS turnover: duox2 and nox1, both generating extracellular H2O2; the expression of duoxa2, required for the maturation of functional duox2, also increased, consistent with increased duox2 expression. Expression of the AMPs, reg3β and reg3ϒ, ang4, and pla2g2a also increased, as well as that of fut2. We further demonstrated that components of the mucosal immune system recognize and react to the presence of commensal E. coli by increasing their expression, consistent with previous results that showed that GF mice colonized with members of the gut microbiota undergo immune system activation and development31.
Previous studies have shown that the intestinal expression of duox232, members of the reg3 family33, and fut234 are upregulated by the gut microbiota in mice and that certain single commensal or probiotic bacterial strains can contribute to this effect33,34,35. These are key components of the innate immune system that help to fight against pathogens36,37,38 while preserving the symbiotic nature of the relationship between the gut microbiota and the host. In mice, duox2-generated H2O2 and members of the reg3 family play a key role by spatially segregating indigenous bacteria, thereby dampening microbiota-induced mucosal immune responses35,36. Fut2 is involved in the fucosylation of glycol-conjugates expressed on epithelial cells, providing a source of host-derived complex carbohydrates for the gut microbiota. The induction of fut2 plays a role in colonization resistance against pathogens by restoring commensal diversity38.
Genes modulated in mono-associated mice include those that encode Ca2+-activated chloride channel 4, (clca4), the fructose transporter Slc2a5 (previously called GLUT5), and members of the water and glycerol channel aquaglyceroporin family (AQP -3 and -7), suggesting that the primary functions of the intestine may also be modulated in mono-associated mice39. CLCA4 expression is downregulated in UC, suggesting that the control of electrolyte balance may be part of the defence mechanism against luminal microbes40. As recently shown, AQP3 transports H2O2 generated at the cell surface by NOX1 and DUOX2 to mediate signal transduction in colonic epithelia41. This further argues for a central role of E. coli strains on ROS turnover and signalling.
We tested the effect of the CEC strain in IL10−/− mice, a well-recognized model for immune-mediated colitis, given its relevance to human IBD42. We speculate that the impact of the genetic background was low in the models we used. Indeed, the genetic background of wild type GF mice was C57BL/6 and that of GF IL10−/− mice was a mix of C57BL/6 and 129/Ola. In our study, we found that, in the absence of IL10, the host response to commensal CEC and Nissle was stronger 21 days post-inoculation than in WT mice, as observed for duox2, duox2a, nox1, nos2, reg3β, reg3ϒ, and fut2 expression. We also observed strong activation of immune responses, as observed for cxcl10, tap1, il1β, and IL22. In particular, neither the expression of stat1 nor that of nod-like receptor 5 (nlrc5), were modified in WT mice by the presence of the two bacterial strains, whereas they were four to eight fold higher in mono-associated IL10−/− than GFIL10 mice. A previous study suggested that IL10 may be involved in the control of the homeostatic relationship between indigenous strains of E. coli and the host43. In our study, intensification of the host response may have compensated for the absence of IL10.
We found that nos2 gene expression can be altered by the bacterial status of mice as seen in the difference between the CVWT and GFWT models. Both CEC and Nissle also upregulated nos2 expression in the WT mice but to a lesser extent than in the CVWT group. This E. coli related increase of nos2 is intensified in Il10−/− mono-colonized mice. The role of intestinal nos2 in E. coli growth has been demonstrated in an inflammation setting. Previous studies have shown an elevated intestinal expression of nos2 during the inflammation process and that the host-nos2-derived by-products of reactive nitrogen species contribute to the proliferation of E. coli44. However, the physiological role of nos2 in this context has not been fully investigated and needs further studies.
We observed no evidences for intestinal inflammation in the mono-associated IL10−/− mice, neither in the ileum or colon, despite a high level of colonization. Furthermore, the onset of inflammation was previously shown to be preceded by increased ileal and colonic permeability in the IL10−/− mouse model45. In contrast, we found that some parameters of ileal and colonic permeability and integrity were improved in the CEC and Nissle mono-associated IL10−/− groups, although we did not find any major differences in the passage of molecules. Thus, we found decreased electrical conductance associated with stronger staining of cadherin-1 in the ileum of IL10−/− mice mono-colonised with E. coli. In addition, we found the epithelial integrity to be preserved, based on the positive effect of the strains on an epithelial proliferative marker. In addition, the ileal expression of Muc13 increased. Finally, we showed that CEC and Nissle can induce an increase in the thickness of the mucus layer in the ileum, which extended to the colon in the IL10−/− mice.
We further investigated the effect of strain CEC in a disease setting using a model of conventional Il10−/− mice exposed to DNBS. As previously mentioned, the IL-10 deficient mice model shares similarities with human IBD patients42. As the onset of colitis can vary among mice according to several factors46, a DNBS administration protocol, previously validated in our lab25, was used to synchronize colitis in this model. Our data show a beneficial role of CEC in promoting gut homeostasis upon mucosal injury in IL10−/− mice. Although CEC was unable to reverse DNBS-associated weight loss, it attenuated several types of DNBS-induced damage, and reversed the gene expression profile towards that of the control group, both in the ileum and colon. The beneficial effect of CEC appeared to be stronger than that of the Nissle strain in the colon, based on the gene expression profile.
Several studies have reported that Nissle has an inhibitory effect on other E. coli species47,48,49. Regarding our 16S data, we did not observe any impact of CEC or Nissle on the gut microbiota composition. However, as 16S data analysis has insufficient resolution to distinguish genetically close organisms, we cannot exclude a remodelling of E. coli population by the administration of CEC15 or Nissle.
Our results that show a beneficial effect of CEC compared to those of other studies, strongly suggest that different E. coli strains of the gut microbiota may differ dramatically in their colitogenic or probiotic potential. For example, indigenous E. coli strains have a colitis-inducing potential in a genetically predisposed model for inflammation, whereas others have no negative impact19. The genetic determinants underlying such divergent behaviour are currently far from being understood. Preliminary data show that the genome of the CEC strain does not contain the genomic pks island (data not shown). This is in contrast to the Nissle strain, for which the pks island was shown. This gene cluster encodes non-ribosomal peptide synthetases (NRPS) and polyketide synthetases (PKS) and produces colibactin (a peptide-polyketide genotoxin) that can induce DNA damage by inducing double-strand breaks50. However, the role of pks in gut health is a matter of debate. Indeed, the probiotic properties of Nissle are linked to the pks island, more precisely to a lipopeptide encoded in this this region51. Sequencing of the CEC15 genome and genomic comparison of other commensal E. coli strains with a known phenotype will help to identify the molecular signatures that can distinguish E. coli strain impact. Preliminary analysis, based on a PCR method enabling an E. coli isolate to be assigned to one of the E. coli phylogroups52 reveals that the CEC15 belongs to the phylogroup C, a group of strains closely related to the phylogroup B1. So, interestingly, CEC15, which had a stronger anti-inflammatory effect than Nissle, belongs to a different phylogroup (Phylogroup B2 for Nissle53).
Recently, a study identified a respiratory pathway that is only operational during episodes of inflammation and that can drive Enterobacteriaceae expansion during colitis54. Selective inhibition of this pathway prevents dysbiotic overgrowth of Enterobacteriaceae, including E. coli, in murine models of colitis55. The implementation of this strategy, based on a better knowledge of the effect of commensal E. coli, has the potential to control the population of E. coli, making it possible to discard potential harmful microbes while preserving the potentially beneficial ones.
Data Availability
Full data will be available from the corresponding author on reasonable request.
References
Tenaillon, O., Skurnik, D., Picard, B. & Denamur, E. The population genetics of commensal Escherichia coli. Nat Rev Microbiol 8, 207–217, https://doi.org/10.1038/nrmicro2298 (2010).
Secher, T., Brehin, C. & Oswald, E. Early settlers: which E. coli strains do you not want at birth? Am J Physiol Gastrointest Liver Physiol 311, G123–129, https://doi.org/10.1152/ajpgi.00091.2016 (2016).
Adlerberth, I. & Wold, A. E. Establishment of the gut microbiota in Western infants. Acta Paediatr 98, 229–238, https://doi.org/10.1111/j.1651-2227.2008.01060.x (2009).
Aagaard, K. et al. The placenta harbors a unique microbiome. Sci Transl Med 6, 237ra265, https://doi.org/10.1126/scitranslmed.3008599 (2014).
Tomas, J. et al. Early colonizing Escherichia coli elicits remodeling of rat colonic epithelium shifting toward a new homeostatic state. ISME J 9, 46–58, https://doi.org/10.1038/ismej.2014.111 (2015).
Lundell, A. C. et al. Infant B cell memory differentiation and early gut bacterial colonization. J Immunol 188, 4315–4322, https://doi.org/10.4049/jimmunol.1103223 (2012).
Smati, M. et al. Quantitative analysis of commensal Escherichia coli populations reveals host-specific enterotypes at the intra-species level. Microbiologyopen 4, 604–615, https://doi.org/10.1002/mbo3.266 (2015).
Chang, D. E. et al. Carbon nutrition of Escherichia coli in the mouse intestine. Proc Natl Acad Sci USA 101, 7427–7432, https://doi.org/10.1073/pnas.0307888101 (2004).
Albenberg, L. et al. Correlation between intraluminal oxygen gradient and radial partitioning of intestinal microbiota. Gastroenterology 147, 1055–1063 e1058, https://doi.org/10.1053/j.gastro.2014.07.020 (2014).
Chow, J. & Mazmanian, S. K. A pathobiont of the microbiota balances host colonization and intestinal inflammation. Cell Host Microbe 7, 265–276, https://doi.org/10.1016/j.chom.2010.03.004 (2010).
Pascal, V. et al. A microbial signature for Crohn’s disease. Gut 66, 813–822, https://doi.org/10.1136/gutjnl-2016-313235 (2017).
Palmela, C. et al. Adherent-invasive Escherichia coli in inflammatory bowel disease. Gut 67, 574–587, https://doi.org/10.1136/gutjnl-2017-314903 (2018).
Feng, Q. et al. Gut microbiome development along the colorectal adenoma-carcinoma sequence. Nat Commun 6, 6528, https://doi.org/10.1038/ncomms7528 (2015).
Kotlowski, R., Bernstein, C. N., Sepehri, S. & Krause, D. O. High prevalence of Escherichia coli belonging to the B2+D phylogenetic group in inflammatory bowel disease. Gut 56, 669–675, https://doi.org/10.1136/gut.2006.099796 (2007).
Carvalho, F. A. et al. Transient inability to manage proteobacteria promotes chronic gut inflammation in TLR5-deficient mice. Cell Host Microbe 12, 139–152, https://doi.org/10.1016/j.chom.2012.07.004 (2012).
Wohlgemuth, S., Haller, D., Blaut, M. & Loh, G. Reduced microbial diversity and high numbers of one single Escherichia coli strain in the intestine of colitic mice. Environ Microbiol 11, 1562–1571, https://doi.org/10.1111/j.1462-2920.2009.01883.x (2009).
Kim, S. C. et al. Variable phenotypes of enterocolitis in interleukin 10-deficient mice monoassociated with two different commensal bacteria. Gastroenterology 128, 891–906 (2005).
Kim, S. C., Tonkonogy, S. L., Karrasch, T., Jobin, C. & Sartor, R. B. Dual-association of gnotobiotic IL-10−/− mice with 2 nonpathogenic commensal bacteria induces aggressive pancolitis. Inflamm Bowel Dis 13, 1457–1466, https://doi.org/10.1002/ibd.20246 (2007).
Waidmann, M. et al. Bacteroides vulgatus protects against Escherichia coli-induced colitis in gnotobiotic interleukin-2-deficient mice. Gastroenterology 125, 162–177 (2003).
Bloom, S. M. et al. Commensal Bacteroides species induce colitis in host-genotype-specific fashion in a mouse model of inflammatory bowel disease. Cell Host Microbe 9, 390–403, https://doi.org/10.1016/j.chom.2011.04.009 (2011).
Rath, H. C., Wilson, K. H. & Sartor, R. B. Differential induction of colitis and gastritis in HLA-B27 transgenic rats selectively colonized with Bacteroides vulgatus or Escherichia coli. Infect Immun 67, 2969–2974 (1999).
Wassenaar, T. M. Insights from 100 Years of Research with Probiotic E. Coli. Eur J Microbiol Immunol (Bp) 6, 147–161, https://doi.org/10.1556/1886.2016.00029 (2016).
Henker, J., Muller, S., Laass, M. W., Schreiner, A. & Schulze, J. Probiotic Escherichia coli Nissle 1917 (EcN) for successful remission maintenance of ulcerative colitis in children and adolescents: an open-label pilot study. Z Gastroenterol 46, 874–875, https://doi.org/10.1055/s-2008-1027463 (2008).
Kuhn, R., Lohler, J., Rennick, D., Rajewsky, K. & Muller, W. Interleukin-10-deficient mice develop chronic enterocolitis. Cell 75, 263–274 (1993).
Barone, M. et al. A Versatile New Model of Chemically Induced Chronic Colitis Using an Outbred Murine Strain. Front Microbiol 9, 565, https://doi.org/10.3389/fmicb.2018.00565 (2018).
Cherbuy, C. et al. Microbiota matures colonic epithelium through a coordinated induction of cell cycle-related proteins in gnotobiotic rat. Am J Physiol Gastrointest Liver Physiol 299, G348–357, https://doi.org/10.1152/ajpgi.00384.2009 (2010).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25, 402–408, https://doi.org/10.1006/meth.2001.1262 (2001).
Tomas, J. et al. Primocolonization is associated with colonic epithelial maturation during conventionalization. FASEB J 27, 645–655, https://doi.org/10.1096/fj.12-216861 (2013).
Escudie, F. et al. FROGS: Find, Rapidly, OTUs with Galaxy Solution. Bioinformatics 34, 1287–1294, https://doi.org/10.1093/bioinformatics/btx791 (2018).
Matamouros, S. et al. Adaptation of commensal proliferating Escherichia coli to the intestinal tract of young children with cystic fibrosis. Proc Natl Acad Sci USA 115, 1605–1610, https://doi.org/10.1073/pnas.1714373115 (2018).
Mathieu, E. et al. Paradigms of Lung Microbiota Functions in Health and Disease, Particularly, in Asthma. Front Physiol 9, 1168, https://doi.org/10.3389/fphys.2018.01168 (2018).
Sommer, F. & Backhed, F. The gut microbiota engages different signaling pathways to induce Duox2 expression in the ileum and colon epithelium. Mucosal Immunol 8, 372–379, https://doi.org/10.1038/mi.2014.74 (2015).
Natividad, J. M. et al. Differential induction of antimicrobial REGIII by the intestinal microbiota and Bifidobacterium breve NCC2950. Appl Environ Microbiol 79, 7745–7754, https://doi.org/10.1128/AEM.02470-13 (2013).
Bry, L., Falk, P. G., Midtvedt, T. & Gordon, J. I. A model of host-microbial interactions in an open mammalian ecosystem. Science 273, 1380–1383 (1996).
Grasberger, H. et al. Increased Expression of DUOX2 Is an Epithelial Response to Mucosal Dysbiosis Required for Immune Homeostasis in Mouse Intestine. Gastroenterology 149, 1849–1859, https://doi.org/10.1053/j.gastro.2015.07.062 (2015).
Brandl, K., Plitas, G., Schnabl, B., DeMatteo, R. P. & Pamer, E. G. MyD88-mediated signals induce the bactericidal lectin RegIII gamma and protect mice against intestinal Listeria monocytogenes infection. J Exp Med 204, 1891–1900, https://doi.org/10.1084/jem.20070563 (2007).
Flores, M. V. et al. Dual oxidase in the intestinal epithelium of zebrafish larvae has anti-bacterial properties. Biochem Biophys Res Commun 400, 164–168, https://doi.org/10.1016/j.bbrc.2010.08.037 (2010).
Pham, T. A. et al. Epithelial IL-22RA1-mediated fucosylation promotes intestinal colonization resistance to an opportunistic pathogen. Cell Host Microbe 16, 504–516, https://doi.org/10.1016/j.chom.2014.08.017 (2014).
Deschemin, J. C. et al. The microbiota shifts the iron sensing of intestinal cells. FASEB J 30, 252–261, https://doi.org/10.1096/fj.15-276840 (2016).
Schniers, A. et al. The Proteome of Ulcerative Colitis in Colon Biopsies from Adults - Optimized Sample Preparation and Comparison with Healthy Controls. Proteomics Clin Appl 11, https://doi.org/10.1002/prca.201700053 (2017).
Thiagarajah, J. R., Chang, J., Goettel, J. A., Verkman, A. S. & Lencer, W. I. Aquaporin-3 mediates hydrogen peroxide-dependent responses to environmental stress in colonic epithelia. Proc Natl Acad Sci USA 114, 568–573, https://doi.org/10.1073/pnas.1612921114 (2017).
Keubler, L. M., Buettner, M., Hager, C. & Bleich, A. A Multihit Model: Colitis Lessons from the Interleukin-10-deficient Mouse. Inflamm Bowel Dis 21, 1967–1975, https://doi.org/10.1097/MIB.0000000000000468 (2015).
Wu, C., Sartor, R. B., Huang, K. & Tonkonogy, S. L. Transient activation of mucosal effector immune responses by resident intestinal bacteria in normal hosts is regulated by interleukin-10 signalling. Immunology 148, 304–314, https://doi.org/10.1111/imm.12612 (2016).
Spees, A. M. et al. Streptomycin-induced inflammation enhances Escherichia coli gut colonization through nitrate respiration. MBio 4, https://doi.org/10.1128/mBio.00430-13 (2013).
Madsen, K. L. et al. Interleukin-10 gene-deficient mice develop a primary intestinal permeability defect in response to enteric microflora. Inflamm Bowel Dis 5, 262–270 (1999).
Martin, R. et al. Using murine colitis models to analyze probiotics-host interactions. FEMS Microbiol Rev 41, S49–S70, https://doi.org/10.1093/femsre/fux035 (2017).
Fang, K., Jin, X. & Hong, S. H. Probiotic Escherichia coli inhibits biofilm formation of pathogenic E. coli via extracellular activity of DegP. Sci Rep 8, 4939, https://doi.org/10.1038/s41598-018-23180-1 (2018).
Hancock, V., Dahl, M. & Klemm, P. Probiotic Escherichia coli strain Nissle 1917 outcompetes intestinal pathogens during biofilm formation. J Med Microbiol 59, 392–399, https://doi.org/10.1099/jmm.0.008672-0 (2010).
Mohsin, M., Guenther, S., Schierack, P., Tedin, K. & Wieler, L. H. Probiotic Escherichia coli Nissle 1917 reduces growth, Shiga toxin expression, release and thus cytotoxicity of enterohemorrhagic Escherichia coli. Int J Med Microbiol 305, 20–26, https://doi.org/10.1016/j.ijmm.2014.10.003 (2015).
Nougayrede, J. P. et al. Escherichia coli induces DNA double-strand breaks in eukaryotic cells. Science 313, 848–851, https://doi.org/10.1126/science.1127059 (2006).
Perez-Berezo, T. et al. Identification of an analgesic lipopeptide produced by the probiotic Escherichia coli strain Nissle 1917. Nat Commun 8, 1314, https://doi.org/10.1038/s41467-017-01403-9 (2017).
Clermont, O., Christenson, J. K., Denamur, E. & Gordon, D. M. The Clermont Escherichia coli phylo-typing method revisited: improvement of specificity and detection of new phylo-groups. Environ Microbiol Rep 5, 58–65, https://doi.org/10.1111/1758-2229.12019 (2013).
Olier, M. et al. Genotoxicity of Escherichia coli Nissle 1917 strain cannot be dissociated from its probiotic activity. Gut Microbes 3, 501–509, https://doi.org/10.4161/gmic.21737 (2012).
Hughes, E. R. et al. Microbial Respiration and Formate Oxidation as Metabolic Signatures of Inflammation-Associated Dysbiosis. Cell Host Microbe 21, 208–219, https://doi.org/10.1016/j.chom.2017.01.005 (2017).
Zhu, W. et al. Precision editing of the gut microbiota ameliorates colitis. Nature 553, 208–211, https://doi.org/10.1038/nature25172 (2018).
Acknowledgements
We thank the @BRIDGe platform, UMR 1313 GABI, 78350, Jouy-en-Josas, France, for technical support and advice, in particular C. Bevilacqua and M.N. Rossignol for the high-throughput gene expression experiments and the histology facility team, including J. Riviere, A. Boukadiri and M. Violette. We also thank the IERP and ANAXEM Animal Facilities of the INRA of Jouy-en-Josas for the animal experiments and technical support.
Author information
Authors and Affiliations
Contributions
U.E. and C.C. designed the study, collected, analysed, and interpreted the data. S.V. helped in collecting and analysing the samples. R.M. and F.C. helped for the design of the study. U.E., C.C., P.L. and M.T. wrote the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Escribano-Vazquez, U., Verstraeten, S., Martin, R. et al. The commensal Escherichia coli CEC15 reinforces intestinal defences in gnotobiotic mice and is protective in a chronic colitis mouse model. Sci Rep 9, 11431 (2019). https://doi.org/10.1038/s41598-019-47611-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-47611-9
This article is cited by
-
Diets Supplemented with Probiotics Improve the Performance of Broilers Exposed to Heat Stress from 15 Days of Age
Probiotics and Antimicrobial Proteins (2023)
-
Comprehensive probiogenomics analysis of the commensal Escherichia coli CEC15 as a potential probiotic strain
BMC Microbiology (2023)
-
Effects of GLP-1 receptor agonist on changes in the gut bacterium and the underlying mechanisms
Scientific Reports (2021)
-
Efficient protection of microorganisms for delivery to the intestinal tract by cellulose sulphate encapsulation
Microbial Cell Factories (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.