Abstract
Seeds are involved in the vertical transmission of microorganisms in plants and act as reservoirs for the plant microbiome. They could serve as carriers of pathogens, making the study of microbial interactions on seeds important in the emergence of plant diseases. We studied the influence of biological disturbances caused by seed transmission of two phytopathogenic agents, Alternaria brassicicola Abra43 (Abra43) and Xanthomonas campestris pv. campestris 8004 (Xcc8004), on the structure and function of radish seed microbial assemblages, as well as the nutritional overlap between Xcc8004 and the seed microbiome, to find seed microbial residents capable of outcompeting this pathogen. According to taxonomic and functional inference performed on metagenomics reads, no shift in structure and function of the seed microbiome was observed following Abra43 and Xcc8004 transmission. This lack of impact derives from a limited overlap in nutritional resources between Xcc8004 and the major bacterial populations of radish seeds. However, two native seed-associated bacterial strains belonging to Stenotrophomonas rhizophila displayed a high overlap with Xcc8004 regarding the use of resources; they might therefore limit its transmission. The strategy we used may serve as a foundation for the selection of seed indigenous bacterial strains that could limit seed transmission of pathogens.
Similar content being viewed by others
Introduction
Plant-associated microbial assemblages (a.k.a plant microbiome) can impact plant fitness through the modification of a number of traits including biomass production, flowering time or resistance to abiotic and biotic stresses1,2,3,4. Due to these important effects on plant growth and health, numerous studies have focused mainly on the microbial communities horizontally acquired from the environment and associated with the rhizosphere and phyllosphere5,6,7. Although the relative contribution of vertical transmission in the ultimate composition of the plant microbiome is probably limited8, vertically-transmitted microorganisms can also have an important impact on plant fitness9. For instance, vertically-transmitted microorganisms can promote germination of seeds through the production of cytokinins10,11 or increase nutrient availability in shoots and roots12. A well-documented example of vertical transmission is seed transmission of plant pathogenic agents. Seed transmission of plant pathogens, including viruses, bacteria and fungi, was initially identified in the late 1800s - early 1900s13. Seed-transmission of plant pathogenic agents serves as a major means of dispersal and can therefore be responsible for disease emergence and spread13,14. Very low degrees of seed contamination by bacterial pathogens can lead to efficient plant colonization15. Furthermore, seed transmission of plant pathogens is usually asymptomatic and can take place on non-host plants16,17, which can then serve as a reservoir of plant pathogens.
To control seed transmission of plant pathogens one should either eradicate them on seed-producing crops or perform seed treatments. Fungicide application during plant production or as seed treatment is an effective means of fungal pathogen management18. Since these chemical-base methods are unsatisfactory for bacterial plant pathogens and have a potentially harmful environmental impact19, alternative control methods have been proposed, including seed health testing, physical seed treatment (e.g thermotherapy) and biological methods such as seed coating of specific biocontrol agents14,18. Direct incorporation of biocontrol agents within the seed tissues through inoculation of seed-producing crops is of interest to restrict seed transmission of plant pathogenic agents. Recently, this approach (called EndoSeedTM) has been successfully performed by introducing the plant-growth promoting strain Paraburkholderia phytofirmans PsJN into seeds of multiple plant species through spray-inoculation of the parent plant at the flowering stage20. Therefore, inoculation of strain(s) that can compete with plant pathogens for resources and space may be of interest to limit seed transmission of phytopathogenic agents21.
Although inoculation of biocontrol microorganisms on or within the seeds may improve seedling health by protecting them against seed-borne or soil-borne pathogens, adequate formulation of these microorganisms for a successful seed colonization and subsequent persistence on seedlings is still challenging18. Further knowledge about the interactions between phytopathogenic agents and other members of the plant microbiome could provide clues on the selection of candidate strains that compete with the target plant pathogen. These microbial interactions can be estimated by analyzing the response of the seed microbiome following seed-transmission of plant pathogens. Indeed, this approach can potentially reveal co-occurence and more importantly exclusion between plant pathogens and resident members of the seed microbiota22. Recently, we initiated such an approach by studying the response of the radish seed microbiome to seed-transmission of two plant pathogens of brassicas, the bacterial strain Xanthomonas campestris pv. campestris 8004 (Xcc800423) and the fungal strain Alternaria brassicicola Abra43 (Abra4324,25). These plant pathogens are frequent seed colonizers of a range of Brassicaceae species including cabbage, cauliflower, turnip and radish26,27. Additionally, they differ in their seed transmission pathways: Xcc8004 being seed-transmitted through the systemic and floral pathways while Abra43 being transmitted through the external pathway28,29,30.
According to community profiling approaches, seed transmission of Abra43 impacted the taxonomic composition of the fungal fraction of the seed microbiome, while Xcc8004 did not change the structure of the seed microbiota24. The absence of changes in community structure following Xcc8004 seed transmission could be explained by differences in ecological niches between the plant pathogen and the members of the seed microbiome. In this work, we therefore investigated competition for resources (nutrient and space) between Xcc8004 and seed-borne bacterial strains (epiphytes and endophytes) of radish (Raphanus sativus var. Flamboyant5) through a combination of metagenomics, metabolic fingerprinting, bacterial confrontation assays, and fluorescence in situ hybridization approaches.
Results
Impact of Xcc8004 and Abra43 transmission on the structure of the seed microbiome
The relative abundances (RA) of Xcc8004 and Abra43 within the radish seed microbiome were assessed through mapping of the metagenomic reads against their reference genomic sequences. According to mapping results, Xcc8004 RA increased from 1% of paired-end reads in the control samples to 6% and 30% for X2013 and X2014, respectively (Additional file 1). Abra43 RA increased from 0.1% to 1.1% in 2013 and from 0.02% to 0.58% in 2014 (Additional file 1).
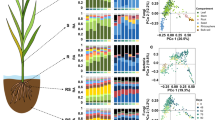
Metagenomic reads were used to predict the structure of the seed microbiome with a modified and parallelized version of Kraken (ParaKraken). The vast majority of the classified reads were derived from two bacterial families: Erwiniaceae and Pseudomonadaceae (Fig. 1a), while less than 0.06% of reads were classified as Dikarya (Additional file 2). At a genus level (Fig. 1g), the seed microbiome was mainly composed of Pantoea and Pseudomonas. In addition, we detected an increase in the RA of reads affiliated with Xanthomonas and to a lesser extent Alternaria in seed samples contaminated with Xcc8004 and Abra43, respectively (Fig. 1a,g).
Changes in seed microbiome structure following seed transmission of Abra43 and Xcc8004. ParaKraken prediction of the taxonomic composition of the bacterial fraction of the radish seed microbiome (a). Measure of microbial alpha diversity through observed richness (number of predicted species), estimated richness (Chao1 index) and diversity (Shannon index) (b–d). Similarity in community membership (Jaccard index) (e) and community composition (Bray-Curtis index) (f) between seed samples. Changes in relative abundance of the 10 most abundant microbial genera following seed transmission of phytopathogenic agents (g).
To assess the impact of Xcc8004 and Abra43 transmission on the seed microbiome structure, we first removed the reads affiliated with the species Xanthomonas campestris and Alternaria brassicicola in the species table generated with ParaKraken. On average, 3,850 species (SD + 385) were detected in seed-associated microbial assemblages collected from control plots (Fig. 1b). This estimation is one order of magnitude higher than the richness predicted with 16S rRNA gene or gyrB24. According to Kruskal-Wallis non parametric analysis of variance followed by Dunn’s post-hoc test, the observed species richness, the predicted species richness and the species diversity were not significantly affected (P > 0.01) by seed transmission of both phytopathogenic agents (Fig. 1b). Differences in community membership and community composition between seed samples were assessed with Jaccard and Bray-Curtis indexes, respectively (Fig. 1e,f). While seed transmission of Xcc8004 and Abra43 did not alter community membership and composition (P > 0.01), the harvesting year explained 54.9% (P = 0.001) and 62.8% (P = 0.001) of variation in community membership and composition, respectively.
Although the overall structure of the radish seed microbiome was not impacted by Xcc8004 and Abra43 seed transmission, we assessed changes in specific taxa RA through fcros31 and DUO32,33 analyses. High abundance species (>0.1% RA) across all three conditions (i.e., Xcc8004 seed transmission (X), Abra43 seed transmission (A) and control (C)) were compared to identify differentially abundant species (Additional file 3). Of the 35 differentially abundant species across all conditions, 22 (63%) were Pseudomonas. DUO of the non-pathogen reads identified two networks with positive associations and no network with negative association (Additional file 4). The two networks were largely split by genus with one network having 67% of interacting species as Pantoea and the other with 67% of Pseudomonas. Both genera were predominant in the seed microbiome. None of the abundant species had a DUO score of >0.75, also suggesting that these pathogens had no major impact on the structure of the microbiome.
Functional composition of the seed microbiome
Taxonomic profiling of the seed microbiomes has shown no significant differences in the community structure between samples harvested from control plots and plots inoculated with Xcc8004 and Abra43. As genomes from different isolates of the same bacterial species can display considerable genetic variation34, seed-transmission of these phytopathogenic microorganisms could still influence the functional composition. We therefore evaluated the impact of seed transmission of plant pathogens on the functional profile of the seed microbiome.
A total of 682,374 coding sequences (CDSs) shared in 17,589 orthologous groups (OGs) were predicted across the different metagenomic samples. On average, each metagenomic sample was composed of 8,828 OGs (SD + 494, Fig. 2a). Seed transmission of Abra43 and Xcc8004 did not significantly alter (P > 0.01) the observed functional richness, the estimated functional richness and the functional diversity of the seed microbiome (Fig. 2a–c). While harvesting year significantly impacted functional membership (P < 0.0001, 44.4%) and functional composition (P < 0.0001, 52.6%) of the seed microbiome, no significant differences (P > 0.001) in functional membership or composition (Fig. 2d,e) were observed after seed transmission of Abra43 and Xcc8004. In addition, we did not detect any significant difference in RA of broad functional categories following pathogen transmission (Fig. 2f). Altogether these results highlight that seed transmission of Xcc8004 and Abra43 do not modify either the predicted structure or function of the seed microbiome.
Changes in the functional composition of the seed microbiome following seed transmission of Abra43 and Xcc8004. Measure of observed functional richness (number of OG), estimated functional richness (Chao1 index) and functional diversity (Shannon index) (a–c). Similarity in functional membership (d) and composition (e) as assessed with Jaccard and Bray-Curtis indexes. Changes in relative abundance of broad functional categories (f).
While broad functional changes were not observed with seed transmission, OG RAs clustered together based upon relative shifts within the microbiome. Markov cluster algorithm of DUO networks resulted in 16 clusters (>2 OGs) with positive associations and 6 clusters with negative associations (Additional file 5). One such positive cluster is suggestive of invasion/pathogenesis of eukaryotic cells and eukaryotic cell response (Additional file 5), denoting important host and pathogen domains in response to infection.
Visualization of Xanthomonas spp. within seed tissues
The absence of shift in structure and function of the seed microbiome following the plant-pathogen transmission suggests that the targeted plant pathogens and the other members of the seed microbiome do not compete for nutrient and space. We first investigated the spatial localization of Xcc8004 within seeds using fluorescence in situ hybridization. Using surface sterilized seeds, Xanthomonas spp. were detected inside inoculated seeds (Fig. 3). They were particularly present in embryonic tissues, especially at the surface of the radicle among other bacteria (Fig. 3a). They were also detected, however, in other embryonic zones of the seeds. Xanthomonas spp. were identified in the same tissues in control samples, but with lower abundance than Xcc8004-seeds (Fig. 3b). Using a photon detection method or analog integration on confocal microscope, similar results were obtained but with brighter signals from the specific probe using analog integration (Fig. 3c,d). Interestingly, cells were single or as packs of several bacteria (Fig. 3a–d). In control and Xcc8004-inoculated seeds hybridized with negative probe, no signal was identified (Fig. 3e,f). On the internal side of the seed coat, Xanthomonas spp. were further identified in Xcc8004 inoculated seeds, showing different niches of colonization (Fig. 3g,h).
Visualization of Xanthomonas spp. inside seeds. Photon counting detection of Xanthomonas spp. on the radicle of the embryo following seed transmission of Xcc8004 (a) or control seeds (b). Analog integration detection of Xanthomonas spp. on radicle of seeds contaminated with Xcc8004 (c) or not contaminated (d). Use of NONEUB probes on samples subjected to Xcc8004 (e) or control seeds (f). Detection of Xanthomonas spp. under the seed coat (g,h). Bacteria corresponding to Xanthomonas spp. appear as yellow/orange fluorescents after blue, green and red channels were merged or as red if channels are separated while other bacteria appear as green fluorescents.
Reconstruction of genomic sequences from the main bacterial population of the seed microbiome
Competition for nutritional resources between phytopathogenic agents and other members of the seed microbiome was subsequently estimated with genomic sequences. Given the limited number of fungal reads in our metagenomics datasets, we restricted this comparative genomic analysis to bacterial populations and Xcc8004. Bacterial genomic sequences were either recovered through metagenomic read sets or via genome sequencing of representative bacterial isolates (see materials and methods). Overall, 11 metagenome-assembled genomes (MAGs; >50% completeness, <10% contamination) and 21 genomes sequences were obtained (Table 1). These genomic sequences represented the major bacterial species detected with the ParaKraken taxonomic classification approach, and, based on their gyrB sequences, represented the main gyrB amplicon sequences variants (ASVs) of the radish seed microbiomes (Table 1 and Additional file 624,35). According to the average nucleotide identity based on blast (ANIb) values, these 32 genomic sequences were divided into 22 bacterial species (Additional file 7). The relative abundance of each genome sequence within the seed microbiome was estimated by mapping the metagenomics reads to these sequences and expressed as average coverage. Overall, genomic sequences related to Pantoea agglomerans and Pseudomonas viridiflava were highly abundant in all radish seed samples, with an average coverage of approximately 200X and 80X, respectively (Table 1 and Additional file 8).
Assessment of resources overlap among bacterial members of the seed microbiome
Overlap in nutritional resource use between Xcc8004 and bacterial populations from the seed microbiome was assessed by profiling nutrient consumption patterns of Xcc8004 and seed bacterial isolates with GEN III MicroPlate. Overall, the bacterial consumption pattern was largely grouped by the phylogenetic relationships between strains (Additional file 9). Overlap in nutritional resources between Xcc8004 and seed-associated bacterial strains ranged from 0.23 to 0.50, with the highest overlap being observed with strains CFBP13503 and CFBP13529 of Stenotrophomonas rhizophila (Fig. 4).
Resources overlap between Xcc8004 and bacterial populations of the seed microbiome. Resource consumption pattern was either assessed with Biolog GEN III MicroPlate or with orthologous groups (OGs) predicted from bin/genomic sequences. Resources overlap was defined as the number of shared resources between Xcc8004 and the tested bacterial strain, divided by the total number of resources used by Xcc8004 and the tested strain.
Since the compounds reduced in GEN III MicroPlates by the bacterial strains are neither representative of all seed exudates compounds nor a reflection of their actual concentrations36, we further assessed similarities/differences in nutritional resources consumption between Xcc8004 and the other bacterial members of the seed microbiome by using MAGs and draft genome sequences and genome sequences. We compared the number of predicted OGs associated with amino acid [E], carbohydrate [G], lipid [I] and inorganic ion [P] transport and metabolism (Fig. 4). The highest median overlap was observed with the functional categories E (0.61) and I (0.66), while the median overlap for G and P was below 0.5. In all cases, the most pronounced resource overlap was associated with genome sequences of Xanthomonadales (Fig. 4).
To determine if the observed resource overlap was associated with a decrease of Xcc8004 population during competition with these bacterial strains, co-inoculation of Xcc8004 was performed with each seed bacterial isolate on radish seed exudates media (see Materials and Methods). We observed a significant decrease (P < 0.01) in Xcc8004 CFU at 2 3, 4 and 5 dpi during co-inoculation with strains CFBP13503 and CFBP13529 (Fig. 5). This effect seems to be dependent on competition for nutritional resources, since no direct antagonism was observed between Xcc8004 and S. rhizophila strains during overlay assay (data not shown). To gain insight into potential competition for nutrients between Xcc8004 and S. rhizophila strains, we compared the set of OGs that were exclusively shared between these strains (Additional file 10). A total of 251 CDSs divided into 219 OGs were specifically shared between Xcc8004 and S. rhizophila. While, most of these OGs have no predicted function, nine CDSs were related to carbohydrate utilization (pectacte lyase, mannosidase and glucosidase). In addition, multiple protein-coding genes involved in rapid utilization of the limiting resource(s), such as TonB-dependent transporters (TBDTs) were also shared between the Xanthomonadales strains (Additional file 10).
Competition for resources between Xcc8004 and bacterial populations of the seed microbiome. Competition between Xcc8004 and seed bacterial isolates was performed at a 1:1 ratio on radish exudates media. Bacterial spots were removed 1, 2, 3, 4 and 5 days post-inoculation and the number of colony forming units of Xcc8004 (y-axis) was recorded on TSA10% supplemented with rifampicin and X-Gluc. Three independent biological replicates, each consisting of 3 technical replicates, were performed. Differences in number of CFUs per treatment were assessed by one-way ANOVA with post-hoc Tukey’s HSD test.
Dynamics of seed-associated microbial taxa during plant germination and emergence
Microbial populations associated with seeds could be either transient colonizers (i.e. seed-borne) or transmitted during germination and emergence to the seedling (seed-transmitted). To investigate which bacterial populations associated with radish seed samples contaminated with Xcc8004 were seed-transmitted, we performed community profiling with gyrB on germinating seeds and seedlings. According to the number of gyrB sequences detected during these early stages of the plant life’s cycle, ASVs belonging to the bacterial species P. agglomerans, P. viridiflava and species of the P. fluorescens group were efficiently transmitted to seedlings (Additional file 11). Moreover, ASVs belonging to Xcc8004 and S. rhizophila were also detected in germinating seeds and radish seedlings (Additional file 11). The competition for resources between Xcc8004 and S. rhizophila strains observed on exudates media (Fig. 5) and the co-occurrence of these two species within germinating seeds and seedlings either suggest that resources are not a limiting factor within these habitats or that both species are located in different tissues.
Discussion
Microbial interactions occurring on and around the seed are important for plant fitness, since seed-borne microorganisms are the primary source of inoculum for the plant. Depending on the outcome of the interactions, these microbial assemblages could either promote or reduce plant growth9. Understanding the processes involved in the assembly and resilience of the seed microbiota is especially relevant to comprehend the emergence of plant diseases. In the current work we assessed whether niche preemption occurred during seed-transmission of two phytopathogenic agents, Xcc8004 and Abra43, which differ in their transmission modes.
According to our metagenomic datasets, the radish seed microbiome was mostly composed of eubacteria. Whether this estimation reflects the actual structure of the radish seed microbiome or represents a bias due to DNA extraction/classification methods is currently unknown. Bacterial communities of radish seeds were dominated by two bacterial families, the Erwiniaceae and Pseudomonadaceae. More precisely, bacterial populations affiliated with Pantoea agglomerans, Erwinia persicina, Pseudomonas viridiflava, P. fluorescens and P. koreensis groups were associated with all radish seed samples. These species have been frequently reported in seeds of Brassicaceae24,35,37,38,39 and in other plant families such as Fabaceae37, Amaranthaceae38 or Poaceae39,40. The high prevalence of these taxa in various seed samples may be explained by specific determinants that could help these microorganisms to thrive in the seed habitat. For instance, seed-transmitted microorganisms should deal with the high level of basal defense response related to reproductive organs41,42,43 and tolerate desiccation occurring during seed maturation44. In addition, the high abundance and occurrence of these bacterial populations within seed samples suggested that these taxa could influence the community structure of the plant microbiome during its early developmental stages. This assumption remained to be validated experimentally through synthetic ecology approaches as recently performed in the maize rhizosphere45.
In our experimental design, the impact of Xcc8004 and Abra43 transmission on the structure and function of the seed microbiome was assessed during two consecutive years. The changes in taxonomic and functional profiles were mostly driven by the harvesting year, thus confirming previous results obtained through community profiling approaches35,37,46. This is perhaps not surprising, as abiotic factors, such as soil or field management practices, have been already observed to have a strong influence in the seed microbita37. In contrast, the seed transmission of the two plant pathogens, Xcc8004 and Abra43, did not impact the overall composition of the seed microbiome.
Previous reports have shown that Xcc8004 is seed-transmitted through the xylem and the stigma, while Abra43 is seed-transmitted via fruits30,31,32. Hence Xcc8004 is probably one of the primary colonists of the seed microbiota, while Abra43 is a late-arriving species. Although both phytopathogenic agents were efficiently transmitted to radish seeds, they impacted neither the structure nor the function of the seed microbiome, implying that they are not competing with the other members of the microbiome for the same ecological niches in the seed habitat. According to in situ hybridization, Xanthomonas sp. co-occurred with other bacterial taxa within the seed coat and at the surface of the seed embryo. Since no spatial separation was observed between Xcc8004 and other seed-borne bacterial populations, then differences in resources consumption can probably explain this co-existence. Based on our genomics prediction of resource overlap and competition assays performed on radish exudate media, most of the bacterial populations of the seed microbiome are not competing with Xcc8004 for nutritional resources. Although the competition assay performed does not necessary reflect the actual composition and concentration of nutrients that are available for microbial growth during seed development36,47, it can serve as a proxy for assessing competition between seed-borne bacterial species.
Under our experimental conditions, the only bacterial population that competes for nutritional resources with Xcc8004 belongs to the species S. rhizophila. Competition for a limiting resource can be divided into two categories: exploitative competition that is related to the rapid use of the limiting resource and contest competition that involves antagonistic interactions with production of antimicrobial compounds48. As we did not observe an antagonistic relationship with the overlay assay used in this work, the observed competition between S. rhizophila strains and Xcc8004 is likely due to resources use49. This contest competition is frequently related to efficient uptake of nutrients by the competing species48. Interestingly numerous OGs that are specifically shared between Xcc8004 and S. rhizophila strains CFBP13503 and CFBP13529 are related to TBDTs, which form a specific carbohydrate utilization system50.
According to the community profiling approach performed on more than 300 radish seed samples24,35, it appears that S. rhizophila strains and Xcc8004 co-exist within the seed habitat. Moreover, both bacterial species co-occurred on germinating seeds and seedlings during germination and emergence, suggesting that nutritional resources are not limited enough within these habitats for observing a strong niche preemption. Alternatively, we cannot rule out the hypothesis of both species being not spatially related. In contrast to Xcc8004, S. rhizophila is mainly located at the seed surface of radish, since no strains were recovered after seed surface sterilization (unpublished observations). Seed surface localization is in accordance with the epiphytic localization of S. rhizophila DSM 14405 on leaves and roots tissues of cotton, sweet pepper and tomato51. Owing to this predicted localization, it is tempting to postulate that S. rhizophila is a late colonist of the seed microbiome, being transmitted throug the external pathway.
Nowadays, sanitary quality of seeds is achieved through chemical methods and prophylaxis measures. Since reducing pesticide usage is an important objective for sustainable agriculture, the search for alternative seed treatments is essential. One of these alternative treatments consists in coating the seeds with microorganisms possessing biocontrol activities. However, biocontrol-based strategies require a better understanding of the colonization abilities of the microbial consortia within the seed habitat and its persistence within the spermosphere21. Our data showed that S. rhizophila and Xcc8004 have a high resource overlap and can compete in vitro for resources. An interesting approach to restrict seed transmission of Xcc8004 could be the introduction of S. rhizophila into the seed tissues with EndoSeedTM technology20. Inoculation would encourage the co-existence of both taxa within the same ecological niche and maybe limit the size of Xcc8004 population within seed tissues. In addition to this augmentative biological control strategy, the limitation of Xcc8004 transmission could be potentially obtained by sowing radish seeds in plots containing a high-level of S. rhizophila. Indeed, the composition of the spermosphere-associated microbial assemblages is highly influenced by local horizontal transmission52. All things considered, S. rhizophila seems to be a promising candidate to reduce seed transmission of the pathogen Xcc8004, although further seed transmission experiments are needed to corroborate the fate of these taxa in planta. Collectively, these data may serve as a foundation for further research towards the design of novel biocontrol-based strategies on seed samples.
Conclusions
Seed-transmission of phytopathogenic agents is significant for disease emergence and spread. In the present study, we have shown that the presence of Abra43 and Xcc8004 within radish seed samples did not modify the structure and function of the seed microbiome. The absence of community shift is explained in part by the low overlap in the use of resources between Xcc8004 and the main seed-borne bacterial populations. However, we have identified potential competitors of Xcc8004 belonging to S. rhizophila species. Understanding the pathways and determinants of seed transmission of Xcc8004 and S. rhizophila can result in the subsequent design of efficient management methods for restricting prevalence of phytopathogenic agents within seed samples through niche preemption21. More generally, the metagenomic sequences obtained in this work provide a starting point to investigate the genomic features involved in seed transmission through comparative genomics with bacterial isolates recovered from other plant habitats such as the rhizosphere or the phyllosphere53. Our approach of coupling community profiling approaches, bacterial confrontational assays and fluorescent in situ hybridization methods opens the way for further target research on seed inoculations with microorganisms possessing plant-beneficial properties.
Materials and Methods
Seed collection, DNA extraction and shotgun metagenomic libraries preparation
The seed samples used in this work were collected in 2013 and 2014 during experiments thoroughly described elsewhere24. Briefly, radish seeds (Raphanus sativus var. Flamboyant5) were harvested at the FNAMS experimental station (47°28′12.42″N, 0°23′44.30″W, Brain-sur-l′Authion, France). These seed samples were collected from plants inoculated with Alternaria brassicicola strain Abra43; plants inoculated with Xanthomonas campestris pv. campestris strain 8004 and control plants without pathogen inoculation. To assess variation in seed community composition, every seed sample (A2013, A2014, C2013, C2014, X2013 and X2014) was divided into 3 subsamples (pseudo-replicates) of 10,000 seeds, resulting in a total of 18 samples (Additional file 1). In addition, a fourth subsample of 10,000 seeds was collected from the X2013 sample for PacBio sequencing.
DNA extraction was performed on seed samples according to the procedure described earlier46 (see Text S1 in the Supplementary material for protocol details). DNA samples were sequenced at the Genome-Transcriptome core facilities (Genotoul GeT-PlaGe; France). Illumina sequencing libraries were prepared according to Illumina’s protocols using the Illumina TruSeq Nano DNA LT Library Prep Kit. DNA-sequencing experiments were performed on an Illumina HiSeq3000 using a paired-end read length of 2 × 150 bases with the Illumina HiSeq3000 Reagent Kit. Each library was sequenced on three lanes of HiSeq3000. PacBio sequencing was performed on a RSII system (details in Text S1). PacBio reads were used to improve the contigs length of the meta-assembly but were not used to assess differences in taxonomic/functional compositions. 0.25 nM of libraries were loaded per SMRTcell on the PacBio RSII System and sequencing was performed on 6 SMRTcells using the OneCellPerWell Protocol (OCPW) on C4P6 chemistry and 360 minutes movies.
Metagenomic read processing
Sequencing of the 18 metagenomic seed samples resulted in a mean of 15.3 million paired-end reads per sample (SD + 3.3 million). These paired-end reads were processed with cutadapt version 1.854 to remove Illumina’s adapters, bases with Qscore < 25, reads possessing more than one ambiguity and paired-end reads with a length < 100 bases. Host-related reads were removed with Bowtie2 version 2.2.455. Quality filtering of raw reads resulted in a mean of 12.5 million paired-end reads (SD + 2.7 million) per metagenomics read set, which corresponds to approximately 2.5 Gb per metagenome (Text S1 and Additional file 1).
Taxonomic inference of metagenomics reads
The relative abundances of Xcc8004 and Abra43 within seed samples were estimated by mapping metagenomic reads to their respective genome sequences23,56. Taxonomic profiling of each metagenome was estimated with a modified parallel version of Kraken57, named ParaKraken33, creating a count table of species and a relative abundance for each sample. Reads affiliated with the species Xanthomonas campestris and Alternaria brassicicola were removed in the count table for richness and diversity analyses (see Text S1 for details).
Richness and diversity were calculated with the R package phyloseq 1.22.358 on species count table. Differences in observed richness, estimated richness and diversity between samples were assessed through Kruskal-Wallis non parametric analysis of variance followed by Dunn’s post-hoc test. Differences in community membership and composition were estimated with the Jaccard and Bray-Curtis index. Principal coordinate analysis (PCoA) was used for ordination of Jaccard and Bray-Curtis distances. A permutational multivariate analysis of variance (PERMANOVA) was carried out to assess the impact of pathogen transmission on seed-associated microbial community profiles. To quantify the contribution of the seed transmission of phytopathogenic microorganisms on microbial community profiles, a canonical analysis of principal coordinates (CAP) was performed with the function capscale of the R package vegan 2.4.2, followed by PERMANOVA with the function adonis.
To investigate changes in the relative abundance between the different seed samples, rare species (those occurring in less than 75% of the samples) were first removed from the species count table. Fcros differential abundance analysis31 was then performed between each sample pair on all species and just the species with the highest abundance (>0.1% RA). A p-value cutoff of 0.05 and an f-score of 0.9 were used to determine differential abundance. The species scaled data was also run through DUO (https://github.com/climers/duo.git) with a threshold of >0.75, a program that identifies positive and negative relationships between species. Briefly, DUO first identifies abundance thresholds of 75th and 25th percentiles. The percentiles are then used to identify relationships between taxa by generating four categories (high-high, high-low, low-high, and low-low) based upon the CCC metric32.
Functional profiling of each metagenome
An assembly of Illumina and PacBio metagenomic reads was performed with IDBA_UD59 and HGAP360, resulting in a non-redundant assembly of circa 1 Gb containing 317,206 contigs (Additional file 12). A gene catalogue was created through gene prediction with Prokka 1.1261. Orthology inference of the 930,134 predicted CDSs was performed with DIAMOND62 against the EggNOG v4.5 database63. Approximately 73% of the predicted CDSs (682,374 CDSs) were affiliated with one orthologous group (OG).
To avoid an overrepresentation of orthologous groups (OGs) related to the inoculated phytopathogenic agents, reads that were mapped to Xcc8004 and Abra43 genomes were discarded. The remaining metagenomic reads were mapped with BowTie2 against all predicted CDSs and converted to bam files with Samtools v1.2.164. Bam files were used to count the number of reads occurrence within each predicted CDS.
Similar to the species counts data, the OG counts were run through DUO with a threshold of >0.75 to identify positive (up-up and down-down) and negative (down-up and up-down) relationships between OGs to hypothesize which functional elements may interact. OG DUO networks were split into a network of positive interactions and negative interactions and then clustered using MCL65 to identify groups of highly interconnected OGs.
Reconstruction of metagenome-assembled genomes
Initial binning was performed with Metabat v. 0.32.466. Metagenome-assembled genomes completeness, contamination and strain heterogeneity were evaluated with CheckM v 1.0.467. Taxonomic affiliation of each bin was performed by Average Nucleotide Identity based on BLAST (ANIb) with Jspecies68. Closest ANIb values were selected for reference genomes and used for Circle Packing representation with the D3.js JavaScript library representation (see details in Text S1).
Reconstruction of genomic sequences of seed-associated bacterial isolates
Approximately 200 bacterial strains from the seed samples used for the metagenomics analysis were isolated on 1/10 strength Tryptic Soy Agar (Text S1). A total of 21 representative isolates of the main bacterial taxa were subjected to Illumina HiSeq. 4000 PE150 sequencing (Table 1 and Additional File 6). These isolates were chosen by comparing their gyrB haplotypes with gyrB sequences previously obtained on radish seed samples24. All genome sequences were assembled with SOAPdenovo 2.0469 and VELVET 1.2.1070, annotated with prokka61 and EggNOG 4.563.
Characterization of resource overlap between seed-associated bacterial isolates
Nutritional resource consumption patterns of bacterial strains were assessed with Biolog GEN III MicroPlateTM (Biolog Inc) (see details in Text S1 in Supplementary data). One Biolog GEN III MicroPlateTM was used per isolate. The nutritional resources overlap was also predicted by comparing the numbers of OGs that are classified in the following functional categories: [E] amino acid transport and metabolism, [G] carbohydrate transport and metabolism, [I] lipid transport and metabolism and [P] inorganic ion transport and metabolism. Competition for resources and direct antagonisms between Xcc8004 and the selected bacterial strains were assessed on radish exudates and TSA 10% media (details in Text S1).
Community profiling of germinating seeds and radish seedlings
Seed samples harvested in plot X2014 were used to determine the dynamics of bacterial population during germination and emergence of radish. DNA extraction and subsequent gyrB amplicon library preparation were performed on germinating seed (n = 6) and seedling (n = 8) samples according to the procedure described earlier46 (see details in Text S1). Libraries were sequenced with a MiSeq reagent kit v2 (500 cycles). Fastq files were processed with DADA2 1.671 using the parameters described in the workflow for “Big Data: Paired-end”. The only modification made regarding this protocol was a change in the truncLen argument according to the quality of the sequencing run. Species abundance was assessed on germinating seed and seedling samples with the R package phyloseq58 (ASVs taxonomic affiliation details in Text S1).
DOPE-FISH and CLSM microscopy of Xcc8004 infected seeds
Seeds were surface-sterilized, cut in half and fixed in a paraformaldehyde solution. DOPE-FISH was performed with probes from Eurofins (Austria) labeled at both the 5′ and 3′ end positions according to Glassner et al.72 using an EUBmix targeting all bacteria (EUB338, EUB338II, EUB338III) coupled with the fluorochrome Cy373,74, and a Xanthomonas spp. targeting probe (5′-TCATTCAATCGCGCGAAGCCCG-3′) coupled with Cy575. A NONEUB probe76 coupled with Cy3 or Cy5 was also used independently as a negative control (further protocol details in Text S1). Samples were observed under a confocal microscope (Olympus Fluoview FV1000 with multiline laser FV5-LAMAR-2 HeNe(G) and laser FV10-LAHEG230-2) (see Text S1 for image processing details). Pictures were cropped, and whole pictures were sharpened to better observe the image details. All experiments were repeated on eight seeds for each condition. Images presented in this publication are the average of colonization.
Data Availability
The datasets supporting the conclusions of this article are available in the SRA database under the accession number PRJNA454584. Whole Genome Shotgun projects have been deposited at GenBank under the accessions QFZV00000000-QGAM00000000. All bacterial strains have been deposited at the CIRM-CFBP (https://www6.inra.fr/cirm_eng/CFBP-Plant-Associated-Bacteria; https://doi.org/10.15454/1.5103266699001077E12).
References
Panke-Buisse, K., Poole, A. C., Goodrich, J. K., Ley, R. E. & Kao-Kniffin, J. Selection on soil microbiomes reveals reproducible impacts on plant function. The ISME journal 9, 980–9 (2015).
Lau, J. A. & Lennon, J. T. Rapid responses of soil microorganisms improve plant fitness in novel environments. Proc. Natl. Acad. Sci. USA 109, 14058–62 (2012).
Mendes, R. et al. Deciphering the rhizosphere microbiome for disease-suppressive bacteria. Science 332, 1097–100 (2011).
Sugiyama, A., Bakker, M. G., Badri, D. V., Manter, D. K. & Vivanco, J. M. Relationships between Arabidopsis genotype-specific biomass accumulation and associated soil microbial communities. Botany 91, 123–126 (2013).
Bulgarelli, D. et al. Revealing structure and assembly cues for Arabidopsis root-inhabiting bacterial microbiota. Nature 488, 91–95 (2012).
Maignien, L., DeForce, E. A., Chafee, M. E., Eren, A. M. & Simmons, S. L. Ecological Succession and Stochastic Variation in the Assembly of Arabidopsis thaliana Phyllosphere Communities. mBio 5, (2014).
Edwards, J. et al. Structure, variation, and assembly of the root-associated microbiomes of rice. Proc. Natl. Acad. Sci. USA 112, E911–920 (2015).
Leff, J. W., Lynch, R. C., Kane, N. C. & Fierer, N. Plant domestication and the assembly of bacterial and fungal communities associated with strains of the common sunflower, Helianthus annuus. New Phytol. 214, 412–423 (2017).
Shade, A., Jacques, M.-A. & Barret, M. Ecological patterns of seed microbiome diversity, transmission, and assembly. Curr. Opin. Microbiol. 37, 15–22 (2017).
Rodrigues Pereira, A. S., Houwen, P. J. W., Deurenberg-Vos, H. W. J. & Pey, E. B. F. Cytokinins and the bacterial symbiosis of Ardisia species. Zeitschrift für Pflanzenphysiologie 68, 170–177 (1972).
Goggin, D. E., Emery, R. J., Kurepin, L. V. & Powles, S. B. A potential role for endogenous microflora in dormancy release, cytokinin metabolism and the response to fluridone in Lolium rigidum seeds. Annals of botany 115, 293–301 (2015).
Vázquez-de-Aldana, B. R., García-Ciudad, A., García-Criado, B., Vicente-Tavera, S. & Zabalgogeazcoa, I. Fungal Endophyte (Epichloë festucae) Alters the Nutrient Content of Festuca rubra Regardless of Water Availability. PLOS ONE 8, e84539 (2013).
Baker, K. F. & Smith, S. H. Dynamics of Seed Transmission of Plant Pathogens. Annual Review of Phytopathology 4, 311–332 (1966).
Gitaitis, R. & Walcott, R. The epidemiology and management of seedborne bacterial diseases. Annual Review of Phytopathology 45, 371–97 (2007).
Darrasse, A., Bureau, C., Samson, R., Morris, C. & Jacques, M.-A. Contamination of bean seeds by Xanthomonas axonopodis pv. phaseoli associated with low bacterial densities in the phyllosphere under field and greenhouse conditions. European Journal of Plant Pathology 119, 203–215 (2007).
Darrasse, A. et al. Transmission of Plant-Pathogenic Bacteria by Nonhost Seeds without Induction of an Associated Defense Reaction at Emergence. Applied and Environmental Microbiology 76, 6787–6796 (2010).
Darsonval, A. et al. The Type III secretion system of Xanthomonas fuscans subsp. fuscans is involved in the phyllosphere colonization process and in transmission to seeds of susceptible beans. Applied and Environmental Microbiology 74, 2669–78 (2008).
Valeria, M. & Gianfranco, R. Seed treatments to control seedborne fungal pathogens of vegetable crops. Pest Management Science 70, 860–868 (2013).
Wightwick, A., Walters, R., Allinson, G., Reichman, S. & Menzies, N. Environmental Risks of Fungicides Used in Horticultural Production Systems. Fungicides, doi:10.5772/13032 (2010).
Mitter, B. et al. A New Approach to Modify Plant Microbiomes and Traits by Introducing Beneficial Bacteria at Flowering into Progeny Seeds. Front. Microbiol. 8, (2017).
Barret, M., Guimbaud, J.-F., Darrasse, A. & Jacques, M.-A. Plant microbiota affects seed transmission of phytopathogenic microorganisms. Molecular Plant Pathology 17, 791–795 (2016).
Jakuschkin, B. et al. Deciphering the Pathobiome: Intra- and Interkingdom Interactions Involving the Pathogen Erysiphe alphitoides. Microb. Ecol. 72, 870–880 (2016).
Qian, W. et al. Comparative and functional genomic analyses of the pathogenicity of phytopathogen Xanthomonas campestris pv. campestris. Genome research 15, 757–67 (2005).
Rezki, S. et al. Differences in stability of seed-associated microbial assemblages in response to invasion by phytopathogenic microorganisms. PeerJ 4, e1923 (2016).
Avenot, H. et al. Isolation of 12 polymorphic microsatellite loci in the phytopathogenic fungus Alternaria brassicicola. Molecular Ecology Notes 5, 948–950 (2005).
Vicente Joana, G. & Holub Eric, B. Xanthomonas campestris pv. campestris (cause of black rot of crucifers) in the genomic era is still a worldwide threat to brassica crops. Molecular Plant Pathology 14, 2–18 (2012).
Verma, P. R., Saharan, G. S. & Canada. Agriculture and Agri-Food Canada. Research Branch. Monograph on Alternaria diseases of crucifers. (Ottawa: Research Branch, Agriculture and Agri-Food Canada, 1994).
Maude, R. B. Seedborne diseases and their control: principles and practice. (1996).
Pochon, S. et al. The Arabidopsis thaliana-alternaria brassicicola pathosystem: a model interaction for investigating seed transmission of necrotrophic fungi. Plant Methods 8, (9 May 2012)-(9 May 2012) (2012).
Wolf, J. M., van der, Zouwen, P. Svander & Heijden, L. van der. Flower infection of Brassica oleracea with Xanthomonas campestris pv. campestris results in high levels of seed infection. Eur J Plant Pathol 136, 103–111 (2013).
Dembélé, D. & Kastner, P. Fold change rank ordering statistics: a new method for detecting differentially expressed genes. BMC Bioinformatics 15, 14 (2014).
Climer, S. et al. A Custom Correlation Coefficient (CCC) Approach for Fast Identification of Multi‐SNP Association Patterns in Genome‐Wide SNPs Data. Genetic epidemiology 38, (610–621 (2014).
Garcia, B. J. et al. Phytobiome and transcriptional adaptation of Populus deltoides to acute progressive drought and cyclic drought. Phytobiomes. https://doi.org/10.1094/PBIOMES-04-18-0021-R (2018)
Lukjancenko, O., Wassenaar, T. M. & Ussery, D. W. Comparison of 61 sequenced Escherichia coli genomes. Microb. Ecol. 60, 708–720 (2010).
Rezki, S. et al. Assembly of seed-associated microbial communities within and across successive plant generations. Plant Soil 422, 67–79 (2018).
Torres-Cortés, G. et al. Functional microbial features driving community assembly during seed germination and emergence. Front Plant Sci 9, 902 (2018).
Klaedtke, S. et al. Terroir is a key driver of seed-associated microbial assemblages. Environ Microbiol 18, 1792–1804 (2016).
Lopez-Velasco, G., Carder, P. A., Welbaum, G. E. & Ponder, M. A. Diversity of the spinach (Spinacia oleracea) spermosphere and phyllosphere bacterial communities. Fems Microbiology Letters 346, 146–154 (2013).
Links, M. G. et al. Simultaneous profiling of seed-associated bacteria and fungi reveals antagonistic interactions between microorganisms within a shared epiphytic microbiome on Triticum and Brassica seeds. The New phytologist 202, 542–53 (2014).
Yang, L. et al. Dominant groups of potentially active bacteria shared by barley seeds become less abundant in root associated microbiome. Front. Plant Sci. 8 (2017).
Terrasson, E. et al. Identification of a molecular dialogue between developing seeds of Medicago truncatula and seedborne xanthomonads. Journal of Experimental Botany, doi:10.1093/jxb/erv167 (2015).
Terras, F. R. G. et al. Small cysteine-rich antifungal proteins from radish - their rôle in host-defense. Plant Cell 7, 573–588 (1995).
Meldau, S., Erb, M. & Baldwin, I. T. Defence on demand: mechanisms behind optimal defence patterns. Ann. Bot. 110, 1503–1514 (2012).
Leprince, O., Pellizzaro, A., Berriri, S. & Buitink, J. Late seed maturation: drying without dying. J Exp Bot 68, 827–841 (2017).
Niu, B., Paulson, J. N., Zheng, X. & Kolter, R. Simplified and representative bacterial community of maize roots. Proc. Natl. Acad. Sci. USA 114, E2450–E2459 (2017).
Barret, M. et al. Emergence Shapes the Structure of the Seed Microbiota. Applied and Environmental Microbiology 81, 1257–1266 (2015).
Nelson, E. B. Microbial dynamics and interactions in the spermosphere. Annual Review of Phytopathology 42, 271–309 (2004).
Hibbing, M. E., Fuqua, C., Parsek, M. R. & Peterson, S. B. Bacterial competition: surviving and thriving in the microbial jungle. Nat. Rev. Microbiol. 8, 15–25 (2010).
Fukami, T. Historical Contingency in Community Assembly: Integrating Niches, Species Pools, and Priority Effects. Annual Review of Ecology, Evolution, and Systematics 46, 1–23 (2015).
Jacques, M.-A. et al. Using Ecology, Physiology, and Genomics to Understand Host Specificity in Xanthomonas. Annu Rev Phytopathol 54, 163–187 (2016).
Schmidt, C. S., Alavi, M., Cardinale, M., Müller, H. & Berg, G. Stenotrophomonas rhizophila DSM14405T promotes plant growth probably by altering fungal communities in the rhizosphere. Biol Fertil Soils 48, 947–960 (2012).
Nelson, E. B. The seed microbiome: Origins, interactions, and impacts. Plant Soil 422, 7–34 (2018).
Bai, Y. et al. Functional overlap of the Arabidopsis leaf and root microbiota. Nature 528, 364–369 (2015).
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal 17, 10–12 (2011).
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nature methods 9, 357–9 (2012).
Belmas, E. et al. Genome Sequence of the Necrotrophic Plant Pathogen Alternaria brassicicola Abra43. Genome Announc. 6, e01559–17 (2018).
Wood, D. E. & Salzberg, S. L. Kraken: ultrafast metagenomic sequence classification using exact alignments. Genome Biology 15, R46 (2014).
McMurdie, P. J. & Holmes, S. phyloseq: An R Package for Reproducible Interactive Analysis and Graphics of Microbiome Census Data. PLOS ONE 8, e61217 (2013).
Peng, Y., Leung, H. C. M., Yiu, S. M. & Chin, F. Y. L. IDBA-UD: a de novo assembler for single-cell and metagenomic sequencing data with highly uneven depth. Bioinformatics 28, 1420–1428 (2012).
Chin, C.-S. et al. Nonhybrid, finished microbial genome assemblies from long-read SMRT sequencing data. Nat Meth 10, 563–569 (2013).
Seemann, T. Prokka: rapid prokaryotic genome annotation. Bioinformatics 30, 2068–2069 (2014).
Buchfink, B., Xie, C. & Huson, D. H. Fast and sensitive protein alignment using DIAMOND. Nat Meth 12, 59–60 (2015).
Huerta-Cepas, J. et al. eggNOG 4.5: a hierarchical orthology framework with improved functional annotations for eukaryotic, prokaryotic and viral sequences. Nucleic Acids Res 44, D286–D293 (2016).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
van Dongen, S., Abreu-Goodger, C. & Using, M. C. L. to extract clusters from networks. Methods Mol. Biol. 804, 281–295 (2012).
Kang, D. D., Froula, J., Egan, R. & Wang, Z. MetaBAT, an efficient tool for accurately reconstructing single genomes from complex microbial communities. PeerJ 3, e1165 (2015).
Parks, D. H., Imelfort, M., Skennerton, C. T., Hugenholtz, P. & Tyson, G. W. CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res. 25, 1043–1055 (2015).
Richter, M. & Rosselló-Móra, R. Shifting the genomic gold standard for the prokaryotic species definition. Proc. Natl. Acad. Sci. USA 106, 19126–19131 (2009).
Luo, R. et al. SOAPdenovo2: an empirically improved memory-efficient short-read de novo assembler. Gigascience 1, 18 (2012).
Zerbino, D. R. & Birney, E. Velvet: Algorithms for de novo short read assembly using de Bruijn graphs. Genome Res. 18, 821–829 (2008).
Callahan, B. J. et al. DADA2: High-resolution sample inference from Illumina amplicon data. Nature Methods 13, 581–583 (2016).
Glassner, H. et al. Bacterial niches inside seeds of Cucumis melo L. Plant Soil 422, 101–113 (2018).
Amann, R. I. et al. Combination of 16S rRNA-targeted oligonucleotide probes with flow cytometry for analyzing mixed microbial populations. Appl. Environ. Microbiol. 56, 1919–1925 (1990).
Daims, H., Brühl, A., Amann, R., Schleifer, K.-H. & Wagner, M. The Domain-specific Probe EUB338 is Insufficient for the Detection of all Bacteria: Development and Evaluation of a more Comprehensive Probe Set. Systematic and Applied Microbiology 22, 434–444 (1999).
Darrasse, A., Barret, M., Cesbron, S., Compant, S. & Jacques, M.-A. Niches and routes of transmission of Xanthomonas citri pv. fuscans to bean seeds. Plant Soil 422, 115–128 (2018).
Günter, W., Rudolf, A. & Wolfgang, B. Optimizing fluorescent in situ hybridization with rRNA‐targeted oligonucleotide probes for flow cytometric identification of microorganisms. Cytometry 14, 136–143 (1993).
Acknowledgements
The authors wish to thank Guillaume Chesneau for his help with bacterial competition assays, Laurent Noel for providing the strain pTac::GUS-GFP, the FNAMS for running field experiments, the ANAN platform (SFR QuaSav) for amplicon sequencing and BGI for sequencing the bacterial genomes. We would also like to thank BBRIC network for all the bioinformatic support. This research was supported by the grant awarded by the Region des Pays de la Loire (metaSEED, 2013 10080); RFI Objectif Végétal (DynaSeedBiome) and the AgreenSkills + fellowship programme, which has received funding from the EU’s Seventh Framework Programme under grant agreement n° FP7-609398. This research used resources from the Oak Ridge Leadership Computing Facility at the Oak Ridge National Laboratory, which is supported by the Office of Science of the U.S. Department of Energy under Contract No. DE-AC05-00OR22725. This research was also supported by the Plant-Microbe Interfaces Scientific Focus Area in the Genomic Science Program, the Office of Biological and Environmental Research (BER) in the U.S. Department of Energy Office of Science, and by the Department of Energy, Laboratory Directed Research and Development funding (ProjectID 8321).
Author information
Authors and Affiliations
Contributions
G.T.C., B.G., S.C., D.J. and M.B. designed the study and made substantial contributions to the analysis and interpretation of the results. G.T.C., B.G., J.P. and M.Br. carried out bioinformatic analyses and participated actively in the interpretation of the results. S.C. performed F.I.S.H. analyses. S.R. and A.P. performed all the microbiological analyses and Biolog profiles. A.R. and O.B. carried out the HiSeq and PacBio sequencing of the environmental DNA. G.T.C., B.G. and M.B. wrote the manuscript with input from the other authors. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Torres-Cortés, G., Garcia, B.J., Compant, S. et al. Differences in resource use lead to coexistence of seed-transmitted microbial populations. Sci Rep 9, 6648 (2019). https://doi.org/10.1038/s41598-019-42865-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-42865-9
This article is cited by
-
Characterization of Pseudomonas viridiflava isolates associated with a new leaf spot disease in Cichorium species
Journal of Plant Pathology (2022)
-
The microbial community associated with pea seeds (Pisum sativum) of different geographical origins
Plant and Soil (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.