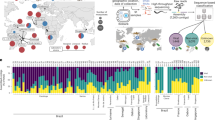
Abstract
Mosquitoes are colonized by a little-studied natural virome. Like the bacterial microbiome, the virome also probably influences the biology and immunity of mosquito vector populations, but tractable experimental models are lacking. We recently discovered two novel viruses in the virome of wild Anopheles and in colonies of the malaria vector Anopheles coluzzii: Anopheles C virus and Anopheles cypovirus. Here, we describe biological interactions between these two viruses and An. coluzzii mosquitoes. Viral abundance varies reproducibly during mosquito development. DNA forms of these viruses were not detected, and thus viral persistence is likely based on vertical transmission of RNA genomes. At least Anopheles C virus is vertically transmitted by an intraembryonic route. Relative abundance of the two viruses is inversely correlated in individual mosquitoes. One possible mechanism for this could be interactions with host immunity, and functional genomic analysis indicated differential influence of at least the Toll and JAK/STAT immune signaling pathways upon the viruses. The nonrandom distributions and interactions with host immunity suggest that these and other members of the natural virome may constitute a source of unrecognized heterogeneity in mosquito vector populations.
Similar content being viewed by others
Introduction
Anopheles species are the main vectors of human malaria. In addition to Plasmodium, Anopheles mosquitoes transmit filarial worms and arboviruses1,2. Anopheles mosquitoes also harbor a diverse natural virome of RNA viruses2,3,4,5. The Anopheles virome is composed mainly of insect specific viruses (ISVs) that multiply only in insects, but also includes relatives of arboviruses that can replicate in both insects and vertebrates. However, Anopheles-RNA virus interactions have been relatively unexamined, and it is unknown why Anopheles are efficient vectors of human malaria but do not transmit viruses as well as Aedes and Culex mosquitoes.
The only arbovirus known to be consistently transmitted by Anopheles mosquitoes is O’nyong nyong virus (ONNV, genus Alphavirus, family Togaviridae). ONNV is mainly transmitted by An. gambiae and An. funestus, two major African malaria vectors6. Millions of people were infected in known ONNV epidemics6,7,8. Anopheles species may also contribute to transmission of other arboviruses such as West Nile virus (WNV), Japanese encephalitis virus, and Wesselsbron virus2,5,9,10.
Anopheles antiviral immunity to ONNV has been studied11,12,13, but the mechanisms of competence and immunity of these mosquitoes to other RNA viruses are essentially unknown. RNA interference (RNAi), Toll, Imd and JAK/STAT are the main antiviral immune signaling pathways described in mosquitoes. The RNA interference (RNAi) pathway controls ONNV replication in An. coluzzii during the disseminated systemic infection11,12,14, but RNAi has no detectable protective function against the primary blood-induced infection of the midgut epithelium12. Conversely, the activity of the JAK/STAT and Imd pathways are required for protection against the primary midgut infection by ONNV, but play no protective role against the disseminated infection12. Thus, distinct antiviral mechanisms function against the primary midgut infection or the disseminated systemic infection.
However, besides ONNV, no studies have been reported on the functional interactions between Anopheles mosquitoes and other RNA viruses, to our knowledge. Studies of other RNA viruses in Anopheles would indicate the specific or general nature of the antiviral mechanisms observed for ONNV. Deep sequencing has recently facilitated discovery of replicating RNA viruses in insects, by identifying viral RNA (viRNA) products of siRNA cleavage of the double-stranded RNA intermediates of active viral replication15,16. By this approach, we recently identified two viruses in An. coluzzii: Anopheles C virus (AnCV) and Anopheles cypovirus (AnCPV)16. Wild Anopheles of diverse species collected in Senegal and Cambodia were positive for AnCPV, and AnCV was present in samples from Senegal. Therefore, both viruses belong to the natural virome of Anopheles.
A recent survey found published evidence of at least 51 viruses naturally associated with Anopheles5. Because Anopheles are persistently exposed in nature to members of the natural virome, it is likely that these ISVs have been the main evolutionary pressure shaping Anopheles antiviral immunity, rather than the less frequent exposure to arboviruses such as ONNV. Thus, here we characterize the biology and host in the interaction of Anopheles with two RNA viruses from the natural virome, AnCPV and AnCV. Understanding the biology underlying the apparent inefficiency of arbovirus transmission by Anopheles may reveal potential new tools to control virus transmission efficient arbovirus vectors such as Aedes and Culex.
Results
Relative composition of the virome varies during mosquito development
It was previously shown that Anopheles C virus (AnCV) and Anopheles cypovirus (AnCPV) are present in the natural virome of wild Anopheles, and also in Anopheles laboratory colonies, including all colonies of An. coluzzii tested to date16. Despite the presence of at least 51 viruses detected in Anopheles species worldwide5, AnCV and AnCPV were the only RNA viruses detected in the Ngousso colony of An. coluzzii16, and the biology of these two viruses is the topic of the current study. To determine the abundance of the two viruses during host development, we measured their prevalence by RT-PCR in different life stages of the An. coluzzii Ngousso strain (Fig. 1, Supplementary Table S1). Infection prevalence of AnCV decreases between pupae and newly emerged adults (p = 2.79e-07, chi-square = 41.08, df = 6, 3 biological replicates), suggesting an impact of metamorphosis on AnCV replication. In contrast, infection prevalence did not change between larvae and pupae, or between 1 week and 2-week-old adults for either virus. The infection prevalence of the two viruses displays a consistent reciprocal pattern in the aquatic larval and pupal stages, with high prevalence of AnCV, while AnCPV infection prevalence is low in larval stages and newly emerged adults, increasing in adults by 1 week after emergence.
Infection prevalence of viruses varies reproducibly during development of Anopheles coluzzii. Virus levels were measured by RT-PCR. Each line represents average infection prevalence for Anopheles C virus (AnCV, circles) and Anopheles cypovirus (AnCPV, squares). Error bars indicate standard deviation from three biological replicates (25–30 individuals per replicate) of each Anopheles stage: larvae (n = 90), pupae (n = 88), imagos (n = 89), adults 1 week (n = 90), adults 2 weeks (n = 88). Detailed descriptive statistics in Supplementary Table S1.
Absence of DNA forms for both AnCV and AnCPV
DNA forms of non-retroviruses have been implicated in viral persistence in Drosophila and Aedes species17,18. We surveyed An. coluzzii to detect DNA forms of AnCV and AnCPV by PCR using three specific primer sets directed against each virus. The template was genomic DNA extracted from An. coluzzii pools, or cDNA generated from RNA of mosquito pools. Despite the high infection prevalence of these viruses, no specific amplification was detected from DNA template (Supplementary Fig. S1). As a positive control, PCR amplification was detected in all cDNA pools, and the larger size of the control ribosomal protein S7 (rpS7) product in the DNA reactions, due to the unspliced intron, confirmed the DNA source of the template. To further strengthen this observation, both virus sequences were used to query the An. gambiae PEST genome assembly using blastn, and no significant matches were detected. Taken together these results indicate that persistence of AnCV and AnCPV in An. coluzzii is probably not based on transmission of DNA forms of the virus genomes.
Transovarial intraembryonic route of AnCV persistence
It was previously observed that AnCV and AnCPV are vertically transmitted in An. coluzzii colonies16, but the transmission routes were not determined. Viruses can pass to progeny within the eggs by a transovarial intraembryonic route, or by egg surface contamination by a transovum route, for example in ovarian fluid. To distinguish between these routes, An. coluzzii eggs were treated with 0.025% sodium hypochlorite (NaOCl) to inactivate viruses on the egg surface, rinsed, and eggs were placed individually in a well of 24-well plates for hatching to prevent horizontal virus transmission between larvae. Controls were treated identically, but without exposure to NaOCl. Larvae (L3/L4 stages) were tested by RT-PCR. Only larvae could be tested because individual eggs reared to larvae in individual wells of a plate did not efficiently develop to the adult stage. Larvae from untreated eggs were infected with both viruses, and larvae from the NaOCl-treated group were infected with detectable levels of at least AnCV (Fig. 2A).
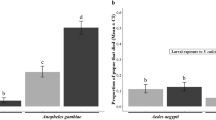
Vertical transmission of Anopheles C virus is intraembryonic. (A) Evaluation of transovarial and transovum routes of AnCV and AnCPV vertical transmission using an RT-PCR to detect both viral genomes in An. coluzzii larvae hatched from eggs in individual wells. Control larvae (black bars) were hatched from eggs mock-treated without NaOCl, and carry both viruses. Larvae grown from eggs treated with NaOCl (gray bars) carry AnCV, while AnCPV was not detected after treatment (N.D.). (B) Both viruses were present in oviposition fluid washed from the surface of An. coluzzii eggs (Ov. Fluid (A. col.)). Viruses were not detected in the pre-rinse water (H2O, N.D.). The virus-positive oviposition fluid was fed to uninfected An. stephensi in larval water, and the exposed larvae did not become detectably infected (Exposed larva (A. steph.), N.D.). Experiments were done in two replicates, error bars indicate standard deviation.
Liquid rinsed from the surface of An. coluzzii eggs containing oviposition fluid was tested using duplex Taqman real-time RT-PCR (Taqman RT-qPCR), to simultaneously measure abundance of both viruses. The oviposition fluid washed from eggs was orally administered to An. stephensi larvae by feeding in the water, to test for virus particles competent for horizontal transmission. The oviposition fluid from An. coluzzii was positive for RNA of AnCV and AnCPV, however the exposed An. stephensi larvae were not infected by either of the two viruses (Fig. 2B). As a positive control, we previously showed that exposure of An. stephensi larvae by the same methodology to filtered extract of homogenized An. coluzzii Ngousso larvae caused infection of 100% of exposed An. stephensi larvae16. Although we cannot rule out a dose effect of virus load difference between oviposition fluid and larval extract, this result may also suggest that viruses in oviposition fluid on the egg surface could be less infectious or non-infectious as virus particles. Taken together, these results suggest that vertical transmission is unlikely to be maintained by simple environmental contamination of oviposition sites with maternal oviposition fluid. These data indicate that an intraembryonic transovarial route is involved in the vertical transmission of at least AnCV, and consistent maintenance by an egg surface transovum route does not seem likely for either virus.
Abundances of AnCV and AnCPV are inversely correlated in mosquitoes
Host interaction with the virome establishes the basal physiological and immune environment encountered by arboviruses, which are probably not frequent components of the pre-existing virome. Thus, interactions among members of the virome might have implications for susceptibility of the host to arboviruses. For example, infection with Nhumirim virus, an insect-specific flavivirus found in the insect virome, reduced by 10,000-fold the viral load of West Nile virus in Ae. albopictus cells, and may serve as a barrier to WNV transmission by Culex19. Cases of homologous and heterologous viral interference were described with mosquito cells20. Knowledge of viral interactions is lacking in Anopheles.
An. coluzzii is persistently coinfected with both AnCV and AnCPV, and we examined the relationship between abundance of the two viruses in individual mosquitoes. Infection intensity and prevalence of both viruses was measured simultaneously in RNA of individual mosquitoes using duplex Taqman RT-qPCR. The abundance of the two viruses in individual mosquitoes is not independent, but rather is inversely correlated. Mosquitoes with high AnCV infection intensity displayed significantly lower infection intensity with AnCPV, and the reciprocal (Fig. 3A; combined p = 3e-05, chi-square = 26.81, df = 4; replicates shown in Supplementary Fig. S2). Co-infection occurs only when both viruses are present at low viral loads.
Inverse correlation between AnCV and AnCPV infection levels in Anopheles coluzzii. Spine plots depict the conditional probability of AnCPV infection load category given AnCV infection load category as measured by duplex Taqman RT-qPCR. The right y-axis indicates the proportion of mosquitoes with the given AnCPV load category (left y-axis) within each AnCV load category (x-axis). Mosquito viral loads were categorized as Non-infected (NI, green), Low infected (orange), and High infected (red) for each virus. Infection levels were defined as virus signal relative to An. coluzzii ribosomal protein rpS7 control, where Low and High levels were values smaller or greater than log10 = 0, respectively, and Non-infected displayed absence of specific virus PCR signal. Infection category of AnCV is indicated by the three histogram bars on the x-axis (AnCV load category), and AnCPV by the three bars on the left y-axis (AnCPV load category). Bar widths on the x-axis are proportional to the total number of mosquitoes in each AnCV load category. (A) Virus infection levels measured in untreated mosquitoes. (B) Virus infection levels measured in mosquitoes wounded by injection with control double stranded RNA, dsGFP. (N) indicates the number of biological replicates and (n) the total number of mosquitoes for all replicates. Statistical difference based on a null hypothesis of random distribution of the two viruses was first tested independently within replicates by the chi-square test using 100,000 permutations to assess the p-value. Individual replicates displayed a consistent direction of change (that is, inverse correlation of abundance of the two viruses, Supplementary Fig. S2), and consequently individual p-values were combined using the method of Fisher to generate a combined p-value, shown above each plot. Detailed statistics in Supplementary Table S4.
Physical injury of Anopheles triggers expression of a suite of wound-response genes that render mosquitoes more resistant to Plasmodium falciparum21. Therefore, we also measured virus abundance distributions in mosquitoes that were wounded by injection of buffer with an irrelevant double-stranded RNA directed against Green Fluorescent Protein. The wounded mosquitoes displayed the same significant inverse correlation of virus abundance as non-injected mosquitoes (Fig. 3B; combined p = 1e-12, chi-square = 76, df = 10; replicates shown in Supplementary Fig. S2). The p-value is more significant from the five combined replicates of wounded mosquitoes (Fig. 3B) as compared to the two combined non-injected replicates (Fig. 3A), however p-values of the individual replicates are comparable for untreated and wounded mosquitoes, indicating that wounding does not alter the conditions that influence distributions of the two viruses. These results indicate that infection efficiency of each virus is not independent of the other, but does not shed light on the mechanism. Potential mechanisms include interference by virus manipulation of the cellular environment and/or host immunity to exclude the other virus, or segregating host genetic differences that are favorable to one virus or the other. We next examined the interaction of the two viruses with host immunity.
The Toll signaling pathway influences AnCPV abundance in mosquitoes
Maintenance of ISVs in a non-pathogenic or commensal state probably requires active policing by basal host immunity, similar to the continual dialog between host immunity and the bacterial microbiome22. AnCV and AnCPV are widespread members of the natural Anopheles virome, and it is likely that the mosquito host deploys immunity to limit viral replication and potential pathogenesis. Here, we used RNAi-mediated gene silencing assays to query the influence of key immune signaling pathways on AnCV and AnCPV abundance.
The Toll pathway controls rodent malaria parasite infection in Anopheles23,24 and limits replication in Aedes of dengue (DENV, genus Flavivirus, family Flaviviridae) and Semliki forest viruses (SFV, genus Alphavirus, family Togaviridae)25,26. In Anopheles, the Toll pathway influence on viruses has only been studied for ONNV12. ONNV strongly inhibits activation of Toll in Anopheles, possibly as an adaptive mechanism because when ectopically activated, Toll significantly inhibits ONNV replication. DENV and SFV are also able to inhibit signaling of Toll in Aedes27,28.
We examined the role of the Toll pathway in the control of AnCV and AnCPV in An. coluzzii by silencing Cactus, a negative regulator of the Toll pathway, to activate Toll signaling (silencing validation shown in Supplementary Fig. S3). Mosquitoes were injected with double-stranded RNA targeting Cactus transcript (dsCactus), and virus abundance was measured by duplex Taqman RT-qPCR. Aggregate analysis of all mosquitoes after silencing Cactus displayed a non-significant tendency of lower AnCV infection intensity (Supplementary Fig. S3a) and inconsistent results for AnCPV infection intensity (Supplementary Fig. S3b).
However, considering the inverse correlation of abundance between the two viruses and potential interference effects (Fig. 3), we then analyzed the effect of Cactus silencing on AnCPV by focusing on the group with low or no AnCV load (equivalent to NI and Low infection categories in Fig. 3). This analysis revealed that Toll activation by dsCactus treatment significantly reduced AnCPV infection prevalence as compared with the dsGFP controls (Fig. 4; combined p = 0.0013, chi-square = 21.8, df = 6). The NI and Low AnCV groups were combined because NI mosquitoes alone were rare, and thus there was no power to compare AnCPV infection levels between these latter two groups. These results support the conclusion that AnCPV is sensitive to antiviral activity mediated by the Toll pathway, while AnCV is insensitive to the effects of the Toll pathway. High AnCV infection also appears to cause interference against AnCPV, which suggests the complexity of the trade-offs that structure the ecology of the virome.
Toll pathway activity limits levels of AnCPV in Anopheles coluzzii. Silencing the Toll negative regulator Cactus activates Toll signaling, and decreases the prevalence of AnCPV when analyzed in the AnCV low-infected group of mosquitoes to minimize the interference effect from AnCV. AnCV low infection level was defined as in Fig. 3 as the AnCV Non-infected and Low-infected groups combined. Plots show the prevalence of infection by AnCPV as a proportion, in three biological replicates ((A) replicate 1, (B) replicate 2, (C) replicate 3). dsCactus indicates mosquitoes injected with double-stranded RNA directed against Toll negative regulator Cactus transcript, and dsGFP indicates mosquitoes injected with irrelevant dsRNA. X-axis, dsRNA treatment, y-axis, AnCPV infection prevalence. (n) indicates the total number of mosquitoes for each of three biological replicates. Statistical differences were first tested independently within replicates by the Wilcoxon-Mann-Whitney test using 100,000 permutations to assess the p-value (given above each replicate plot). Individual replicates displayed a consistent direction of change, and consequently individual p-values were combined using the method of Fisher to generate a combined p-value (combined p = 0.0013, chi-square = 21.8, df = 6). Detailed statistics in Supplementary Table S4.
JAK/STAT signaling activity promotes AnCPV infection in Anopheles coluzzii
The Janus kinase/signal transducer and activator of transcription (JAK/STAT) pathway is implicated in antiviral protection in Anopheles against ONNV12, and in Aedes against DENV and SFV25,26, as well as in mice and Drosophila29,30. We silenced STAT-A, a positive regulator of the JAK/STAT pathway, in An. coluzzii mosquitoes in order to inhibit activation of the pathway (silencing validation shown in Supplementary Fig. S4), and measured virus infection levels by duplex Taqman RT-qPCR. Inhibition of JAK/STAT decreased AnCPV infection prevalence of mosquitoes (Fig. 5, combined p = 0.02, chi-square = 11.4, df = 4, for 2 replicates). There was no effect on AnCPV infection intensity (replicate 1 p = 0.5, replicate 2 p = 0.29), and no effect on AnCV infection (replicate 1 p = 0.37, replicate 2 p = 0.62). This result indicates that the JAK/STAT immune pathway plays a significant role in promoting infection by this cypovirus member of the virome. Interestingly, unlike for the Toll pathway, the effect of STAT-A silencing was detected by the aggregate analysis of all mosquitoes after silencing STAT-A, and did not require analysis of only the low-AnCV fraction of mosquitoes for power to detect an effect. This suggests that, unlike for Toll, AnCV does not appear to influence activation of the JAK/STAT pathway, and/or the influence of the pathway on AnCPV replication.
JAK/STAT pathway activity promotes AnCPV infection in Anopheles coluzzii. Effect of STAT-A silencing on AnCPV prevalence is indicated as a proportion in two biological replicates ((A) replicate 1, (B) replicate 2). dsSTAT-A indicates mosquitoes injected with double-stranded RNA directed against STAT-A transcript, and dsGFP indicates mosquitoes injected with irrelevant dsRNA. X-axis, dsRNA treatment, y-axis, AnCPV infection prevalence. (n) indicates the total number of mosquitoes for each of the 2 replicates. Statistical differences were first tested independently within replicates by the Wilcoxon-Mann-Whitney test using 100,000 permutations to assess the p-value (given above each plot). Individual replicates displayed a consistent direction of change, and consequently individual p-values were combined using the method of Fisher to generate a combined p-value (combined p = 0.02, chi-square = 11.4, df = 4). Detailed statistics in Supplementary Table S4.
RNA interference, Imd, and Pastrel do not significantly influence AnCV and AnCPV levels in An. coluzzii
The RNA interference (RNAi) pathway is a major antiviral mechanism in Drosophila and mosquitoes31,32,33, including in Anopheles against ONNV11,12,14. Argonaute-2 (Ago2) is a key factor of the RNAi pathway, and an antagonist of ONNV in Anopheles. To assess the influence of the RNAi pathway for the control of AnCV and AnCPV in An. coluzzii, Ago2 was depleted by silencing, and virus levels were measured by duplex Taqman RT-qPCR (silencing efficiency shown in Supplementary Fig. S5). There was no consistent effect of inhibition of RNAi activity by Ago2 silencing upon the abundance of AnCV and AnCPV, and none of the replicates was individually significant (Supplementary Fig. S5a). Moreover, AnCPV prevalence in the group with low or no AnCV load (equivalent to NI and Low infection categories in Fig. 3) was also not significantly affected in two independent replicates (p = 1, combined Fisher test).
The immune deficiency (Imd) pathway plays an antiviral role against ONNV during the primary midgut infection of An. coluzzii12, but has no effect during the secondary disseminated systemic infection14. There are two isoforms of the Imd positive activator, Rel2: the longer Rel2-F and shorter Rel2-S. Silencing of both isoforms by targeting a shared region, and specifically silencing only Rel2-F, both caused increased ONNV infection in the midgut, and thus Imd-mediated protection against ONNV in the midgut requires at least the long Rel2-F isoform. Here, we targeted both isoforms by injecting a double-stranded RNA (dsRel2) directed against the common region (silencing efficiency by RT-qPCR shown in Supplementary Fig. S6). Depletion of Rel2 led to a consistent but non-significant tendency in three replicates towards lower infection prevalence for AnCPV (Supplementary Fig. S6a). AnCPV prevalence in the group with low or no AnCV load was not significantly affected in three independent replicates (p = 0.98, combined Fisher test).
Pastrel (pst) has been reported in Drosophila as an antiviral factor against picorna-like viruses and some dicistroviruses, such as Drosophila C-virus, a dicistrovirus related to AnCV34,35,36. There is currently no pst ortholog annotated in the An. gambiae PEST genome assembly. By searching for reciprocal best hits, we identified An. gambiae AGAP011771 as a candidate for Anopheles pst. We designed a double-stranded RNA construct to silence the gene, and measured viral loads by the duplex Taqman RT-qPCR assay (silencing efficiency shown in Supplementary Fig. S7). A non-significant increase of AnCPV-infected mosquitoes was observed after depletion of pst candidate AGAP011771 in four biological replicates (Supplementary Fig. S7a).
Discussion
The Anopheles virome is comprised of a number of viruses, among them insect specific viruses (ISVs)5. These ISVs are maintained by vertical transmission but their biology in Anopheles has not been examined. The goal of this study was to characterize the biological interaction of two ISVs, Anopheles C virus (AnCV) and Anopheles cypovirus (AnCPV), with An. coluzzii, the mosquito host in which they were discovered by deep sequencing of RNA and de novo assembly16. Both viruses have also both been identified in wild-caught Anopheles of various species.
Here, we studied the interaction and immune response to AnCV and AnCPV in co-infected An. coluzzii. In order to quantify the two viruses simultaneously, we developed a duplex Taqman real-time RT-PCR assay that is specific, sensitive, and rapid. We detected a consistent pattern of virus abundance corresponding to stages of host development. We showed that AnCV uses a transovarial, intraembryonic route of transmission. The mechanism of AnCPV persistence is less clear because we did not detect transovarial transmission. However, AnCPV is present at lower levels in the larval stage than AnCPV, and therefore the transmission experiment may have had less sensitivity to detect AnCPV. It was not possible to test adults emerging from the single-egg hatching experiment because larvae grown from eggs hatched in individual wells did not efficiently develop to adults. Both viruses are present in oviposition fluid and NaOCl-untreated eggs of An. coluzzii, but the virus-positive oviposition fluid did not infect An. stephensi larvae by feeding in larval water. Under the same conditions, we previously demonstrated that filtered infected total larval extract infected An. stephensi larvae. The difference could be explained if the viral particles in the oviposition fluid have low environmental stability and/or abundance. Taken together, transovarial transmission was demonstrated for AnCV, and this also appears to be the most likely explanation for AnCPV persistence, but the latter will require further work to detect. Transovarial transmission was demonstrated for DENV and Zika virus in Aedes mosquitoes37,38, but data are lacking on transmission modes of ONNV in Anopheles.
We identified a significant dependence of the abundance distributions of the two viruses in individual mosquitoes. The inversely correlated abundance profiles could be a consequence of cellular interference, manipulation of host immunity, or segregating genetic variation in the host. In order to survey potential host immune effects, we queried the most important immune signaling pathways for influence on the two virome members. We identified a role of the Toll pathway in limiting AnCPV abundance. The effect was only detectable in the background of low infection with AnCV. One hypothesis could be that AnCV inhibits Toll pathway activation, which was previously shown for ONNV in Anopheles12. However, in the case of ONNV, adventitious stimulation of Toll displayed a deleterious effect on ONNV replication, while in the current study stimulation of Toll by depletion of Cactus did not influence AnCV levels. It thus appears more likely that AnCV may express an inhibitor of an unknown Toll-dependent antiviral effector, and not an inhibitor of Toll pathway activation. Hypothetically, AnCV could inhibit expression of the antiviral factor gene, or could directly inhibit the activity of the antiviral effector. Other dicistroviruses such as Drosophila C virus and Cricket paralysis virus express inhibitors of different targets in the RNAi pathway39. Further work could include assaying expression of Toll-response genes in the presence and absence of AnCV infection, in order to test the first hypothesis that AnCV inhibits antiviral gene expression.
We also identified a role of the JAK/STAT pathway in promoting infection of An. coluzzii by AnCPV. Silencing of the JAK/STAT positive regulator STAT-A to inhibit pathway activity reduces AnCPV infection prevalence. JAK/STAT is involved in maintaining homeostasis of the bacterial microbiome, and this result might suggest an implication of the bacterial microbiome in regulating virome composition. In contrast to the positive effect of JAK/STAT activity on AnCPV levels, the same pathway is antagonistic to ONNV during the primary midgut infection in An. coluzzii12, and has no effect on the disseminated systemic ONNV infection after intrathoracic injection14. In Drosophila, the loss of hop, which regulates the single STAT factor, STAT92E increases the viral loads of Drosophila C virus and elevates host mortality29. In contrast, depletion of STAT-A in An. gambiae reduces the number of Plasmodium oocysts in the midgut but increases their proportional survival40. These results observed for different infections indicate a diversity of effects of the JAK/STAT pathway in Anopheles depending on pathogen type and other factors.
Finally, we identified and functionally assayed a candidate for Anopheles pastrel. Mosquitoes depleted of transcript for this candidate displayed a non-significant tendency to increased infection by AnCPV. In order to further characterize the pastrel candidate, as well as a weak potential effect of the Imd pathway, it may be necessary to eliminate or control for the effects of the coinfection of AnCV and AnCPV, which are clearly not independent.
Work still remaining includes examining the influence of the virome upon malaria and ONNV susceptibility and transmission. Our initial attempts to test the influence of the two viruses upon P. falciparum infection were inconclusive, because the persistent infection of An. coluzzii colonies with negatively correlated loads of the two viruses is a confounding factor that requires large sample sizes for statistical power of correlation. An. coluzzii lines carrying only one of AnCV and/or AnCPV would permit controlled reinfection studies. There are interesting examples from other mosquitoes of virome interaction with transmissible pathogens. The insect specific flavivirus of Aedes, Cell fusing agent virus, enhances DENV replication and the reciprocal in Aedes cells41. Palm Creek virus, another insect specific flavivirus, causes reduced replication of the West Nile virus and Murray Valley encephalitis arboviruses in Aedes cells42. However, these studies were done in cultured cells, and studies on interactions between ISVs and transmissible pathogens such as arboviruses and Plasmodium in vivo in mosquitoes, closer to the natural biological conditions, remain to be done.
In the current work, we showed that An. coluzzii, one of main human malaria vectors in Africa, uses classical signaling pathways of immune response to control at least one ISV of the natural virome, AnCPV. The other ISV carried by this mosquito colony, AnCV, appears to be more difficult for the host to control, because its replication was not limited by the pathways or effectors tested. We speculate that AnCV may inhibit an effector of the Toll pathway, an antiviral effector that in turn reduces AnCPV replication when induced. However, Anopheles antiviral mechanisms against arbovirus infection may be quite efficient, based on the relative lack of arbovirus transmission by Anopheles despite highly anthropophilic feeding behavior, including on viremic hosts. Nevertheless, ONNV transmission is the exception that indicates arbovirus transmission by Anopheles is possible, so it is a biological puzzle that transmission is apparently restricted to just one virus.
Identifying the community of natural viruses inhabiting the Anopheles host niche will help clarify the biology underlying the apparent inefficiency of arbovirus transmission by Anopheles, and may suggest new tools to raise the barrier to arbovirus transmission by the more efficient Aedes and Culex vectors. The current study shows that Anopheles ISVs, because of their non-pathogenicity to humans, can be used as simple and tractable models to dissect the diversity of interactions between Anopheles mosquitoes and RNA viruses.
Methods
Mosquitoes
The Anopheles coluzzii Ngousso strain was initiated in Cameroon in 200643 and was obtained by the Institut Pasteur CEPIA facility in 2008. The Anopheles stephensi strain SDA500 was initiated in Pakistan in 198244 and was obtained by the Institut Pasteur CEPIA facility between 2000–2004. Larvae were grown in distilled water supplemented with 0.01% of mineral salt, and fed on Friskies cat food. Adults were reared at 28 °C ±1, at 80% ±5 humidity on a 12 h light–dark cycle. Adults were fed on sterile filtered and autoclaved 10% sucrose solution and females were blood-fed on anaesthetized rabbits for colony maintenance.
Molecular detection and quantification of virus RNA
Template RNA was extracted from individual or pooled mosquitoes using Direct-zol RNA MiniPrep reagents with DNase I treatment (Zymo Research). cDNA was synthesized from the RNA using Moloney Murine Leukemia Virus Reverse Transcriptase (Invitrogen) and random hexamer primers (Roche).
Virus detection of AnCV and AnCPV by simple RT-PCR followed by agarose gel analysis was carried out as described16. A duplex Taqman real-time RT-PCR assay (Taqman RT-qPCR) was developed (Supplementary Table S2) to detect and quantify the two viruses simultaneously. Taqman primers and probes were designed using Primer Express Software v2.0 (Applied Biosystems). Probe and primer sequences are listed in Supplementary Table S3. All have a minor groove binder/non-fluorescent quencher (MGB-NFQ) at the 3′ end and the 6-carboxyfluorescein (6-FAM), VIC and NED fluorophores at the 5′ end for AnCV, AnCPV and The An. coluzzii rpS7 gene respectively. Taqman RT-qPCR was performed with Taqman Universal Master Mix II, with UNG on a QuantStudio 12 K Flex instrument (Applied Biosystems). Primer concentration was 0.9 μM and probe concentration 0.25 μM. The cycling protocol was a hold step 50 °C for 2 min and 95 °C for 10 min, followed by the PCR stage of 40 cycles: 95 °C for 15 secs and 60 °C for 1 min. Negative controls were run with each test. A pool of 10 mosquitoes harboring both viruses was used to make the standard curve. Ten-fold serial dilutions were made between the five dilutions (dil1 to dil5) of the standard curve to test efficiency, specificity and sensitivity. All reactions were performed in triplicate.
No amplification was observed in negative controls with no template. Sensitivity was approved with amplification in the five dilutions of the standard sample (3000 ng for dil1 to 0.03 ng for dil5), and Ct values were inversely proportional to the expected amount of target nucleic acid in each dilution. Efficiency was over 90% in all assays. The An. coluzzii rpS7 gene was amplified in the same plate as the endogenous control gene for relative quantification.
Egg treatment to determine virus transmission route
Eggs were treated with sodium hypochlorite (NaOCl) to eliminate viruses on the egg surface. In a previous study, eggs of the beet armyworm (Spodoptera exigua) were decontaminated of nucleopolyhedroviruses (genus Alphabaculovirus, family Baculoviridae) by treatment with 0.25ppm (0.000025%) NaOCl45. We determined that Anopheles eggs were more resistant to NaOCl, possibly because of the impermeable egg chorion, and that eggs treated with 0.025% were still viable. Consequently, 0.025% NaOCl was used in the current study.
A batch of freshly laid An. coluzzii Ngousso strain eggs was divided into 3 batches. The experimental batch was treated with 0.025% NaOCl for 10 min, rinsed with distilled water, and placed as individual eggs in a well of a 24-well plastic culture plate for hatching and growth until L3/L4 larval stages. The control batch was mock-treated by only rinsing with distilled water (with no NaOCl treatment) and placed as individual eggs in a well of a 24-well plastic culture plate for hatching and growth until L3/L4 larval stages. L3/L4 larval stages from each batch were collected and tested using an RT-PCR assay for presence and abundance of AnCV and AnCPV virus genomes. The experiment was carried out in two replicates.
An additional egg batch was washed with 300 µl of distilled water, which was then collected and filtered with the 5 µm and 0.2 µm filters to remove bacteria. The filtered solution containing oviposition fluid was tested using the duplex Taqman RT-qPCR assay for presence and abundance of AnCV and AnCPV virus genomes. The liquid rinsed from the surface of eggs containing oviposition fluid was fed in the larval water to L2 larvae of An. stephensi. The experiment was carried out in two replicates.
Gene silencing assays
Gene-specific fragments of Ago2, Cactus, the pastrel candidate AGAP011771, Rel2, STAT-A, and GFP control were generated by PCR using primers tagged at their 5′ end with T7 promoter sequences. All T7 primer sequences are listed in Supplementary Table S3. The PCR products were used as template for in vitro dsRNAs synthesis using the MEGAscript RNAi Kit (Ambion). The targeted gene was silenced by injecting 500 ng of dsRNA into the thorax of ice-anesthetized 1–2 days old post-emergence of An. coluzzii females using a nanoinjector (Nanoject II; Drummond Scientific) and glass capillary needle as previously described46. Four days after the dsRNA treatment, silencing of the target gene was verified using total RNA from a pool of five mosquitoes by SYBR Green RT-qPCR for Ago2, Rel2 and pastrel, while RT-PCR followed by detection on agarose gels was used for Cactus and STAT-A. RNA was extracted using Direct-zol RNA MiniPrep reagents with DNase I treatment (Zymo Research). cDNA synthesis was done using Moloney Murine Leukemia Virus Reverse Transcriptase (Invitrogen) and random hexamer primer (Roche). The RT-qPCR primers used for gene expression are given in the Supplementary Table S3.
Total RNA extracted from individual mosquitoes 4 days post-dsRNA injection was used to assess virus infection intensity and prevalence by duplex Taqman real-time RT-PCR assay using relative quantification. The rpS7 gene was used as a housekeeping calibrator for normalization of nucleic acid quantity. Analysis of the expression of transcript relative to rpS7 was performed according to the 2−ΔΔCt method47. Intensity is the log10 of relative expression of virus to rpS7.
Statistical analysis
All statistical details are presented in Supplementary Table S4. For comparisons of virus infection prevalence of different mosquito developmental stages, the chi-square test was applied, and p-values with a null distribution were estimated by the Monte-Carlo method with 10,000 permutations. Multiple testing correction was done by the Bonferroni method. Box and bar plots were made using the beeswarm package in R48. For viral load (infection intensity), statistical significance was determined using the Wilcoxon–Mann–Whitney rank-sum test, and p-values were assessed with a null distribution of the statistical test approximated using Monte-Carlo resampling with 1,000,000 permutations. The p-values from independent tests of significance were combined using the meta-analytical approach of Fisher49.
Data Availability
All data are available in the article.
References
Muturi, E. J. et al. Relationship between malaria and filariasis transmission indices in an endemic area along the Kenyan Coast. J Vector Borne Dis 43, 77–83 (2006).
Villinger, J. et al. Arbovirus and insect-specific virus discovery in Kenya by novel six genera multiplex high-resolution melting analysis. Mol Ecol Resour 17, 466–480, https://doi.org/10.1111/1755-0998.12584 (2017).
Fauver, J. R. et al. West African Anopheles gambiae mosquitoes harbor a taxonomically diverse virome including new insect-specific flaviviruses, mononegaviruses, and totiviruses. Virology 498, 288–299, https://doi.org/10.1016/j.virol.2016.07.031 (2016).
Colmant, A. M. G. et al. A New Clade of Insect-Specific Flaviviruses from Australian Anopheles Mosquitoes Displays Species-Specific Host Restriction. mSphere 2, https://doi.org/10.1128/mSphere.00262-17 (2017).
Nanfack Minkeu, F. & Vernick, K. D. A Systematic Review of the Natural Virome of Anopheles Mosquitoes. Viruses 10, https://doi.org/10.3390/v10050222 (2018).
Williams, M. C., Woodall, J. P., Corbet, P. S. & Gillett, J. D. O’nyong-Nyong Fever: An Epidemic Virus Disease in East Africa. 8. Virus Isolations from Anopheles Mosquitoes. Trans R Soc Trop Med Hyg 59, 300–306 (1965).
Sanders, E. J. et al. O’nyong-nyong fever in south-central Uganda, 1996-1997: description of the epidemic and results of a household-based seroprevalence survey. J Infect Dis 180, 1436–1443, https://doi.org/10.1086/315073 (1999).
Rwaguma, E. B. et al. Emergence of epidemic O’nyong-nyong fever in southwestern Uganda, after an absence of 35 years. Emerg Infect Dis 3, 77, https://doi.org/10.3201/eid0301.970112 (1997).
Su, C. L. et al. Molecular epidemiology of Japanese encephalitis virus in mosquitoes in Taiwan during 2005-2012. PLoS Negl Trop Dis 8, e3122, https://doi.org/10.1371/journal.pntd.0003122 (2014).
Mancini, G. et al. Mosquito species involved in the circulation of West Nile and Usutu viruses in Italy. Vet Ital 53, 97–110, https://doi.org/10.12834/VetIt.114.933.4764.2 (2017).
Keene, K. M. et al. RNA interference acts as a natural antiviral response to O’nyong-nyong virus (Alphavirus; Togaviridae) infection of Anopheles gambiae. Proc Natl Acad Sci USA 101, 17240–17245, https://doi.org/10.1073/pnas.0406983101 (2004).
Carissimo, G. et al. Antiviral immunity of Anopheles gambiae is highly compartmentalized, with distinct roles for RNA interference and gut microbiota. Proc Natl Acad Sci USA 112, E176–185, https://doi.org/10.1073/pnas.1412984112 (2015).
Carissimo, G., Pain, A., Belda, E. & Vernick, K. D. Highly focused transcriptional response of Anopheles coluzzii to O’nyong nyong arbovirus during the primary midgut infection. BMC Genomics 19, 526, https://doi.org/10.1186/s12864-018-4918-0 (2018).
Waldock, J., Olson, K. E. & Christophides, G. K. Anopheles gambiae antiviral immune response to systemic O’nyong-nyong infection. PLoS Negl Trop Dis 6, e1565, https://doi.org/10.1371/journal.pntd.0001565 (2012).
Webster, C. L., Longdon, B., Lewis, S. H. & Obbard, D. J. Twenty-Five New Viruses Associated with the Drosophilidae (Diptera). Evol Bioinform Online 12, 13–25, https://doi.org/10.4137/EBO.S39454 (2016).
Carissimo, G. et al. Identification and Characterization of Two Novel RNA Viruses from Anopheles gambiae Species Complex Mosquitoes. PLoS One 11, e0153881, https://doi.org/10.1371/journal.pone.0153881 (2016).
Goic, B. et al. RNA-mediated interference and reverse transcription control the persistence of RNA viruses in the insect model Drosophila. Nat Immunol 14, 396–403, https://doi.org/10.1038/ni.2542 (2013).
Goic, B. et al. Virus-derived DNA drives mosquito vector tolerance to arboviral infection. Nat Commun 7, 12410, https://doi.org/10.1038/ncomms12410 (2016).
Goenaga, S. et al. Potential for Co-Infection of a Mosquito-Specific Flavivirus, Nhumirim Virus, to Block West Nile Virus Transmission in Mosquitoes. Viruses 7, 5801–5812, https://doi.org/10.3390/v7112911 (2015).
Condreay, L. D. & Brown, D. T. Exclusion of superinfecting homologous virus by Sindbis virus-infected Aedes albopictus (mosquito) cells. J Virol 58, 81–86 (1986).
Nsango, S. E. et al. AP-1/Fos-TGase2 axis mediates wounding-induced Plasmodium falciparum killing in Anopheles gambiae. J Biol Chem 288, 16145–16154, https://doi.org/10.1074/jbc.M112.443267 (2013).
Buchon, N., Broderick, N. A. & Lemaitre, B. Gut homeostasis in a microbial world: insights from Drosophila melanogaster. Nat Rev Microbiol 11, 615–626, https://doi.org/10.1038/nrmicro3074 (2013).
Frolet, C., Thoma, M., Blandin, S., Hoffmann, J. A. & Levashina, E. A. Boosting NF-kappaB-dependent basal immunity of Anopheles gambiae aborts development of Plasmodium berghei. Immunity 25, 677–685, https://doi.org/10.1016/j.immuni.2006.08.019 (2006).
Mitri, C. et al. Fine pathogen discrimination within the APL1 gene family protects Anopheles gambiae against human and rodent malaria species. PLoS Pathog 5, e1000576, https://doi.org/10.1371/journal.ppat.1000576 (2009).
Souza-Neto, J. A., Sim, S. & Dimopoulos, G. An evolutionary conserved function of the JAK-STAT pathway in anti-dengue defense. Proc Natl Acad Sci USA 106, 17841–17846, https://doi.org/10.1073/pnas.0905006106 (2009).
Xi, Z., Ramirez, J. L. & Dimopoulos, G. The Aedes aegypti toll pathway controls dengue virus infection. PLoS Pathog 4, e1000098, https://doi.org/10.1371/journal.ppat.1000098 (2008).
Fragkoudis, R. et al. Semliki Forest virus strongly reduces mosquito host defence signaling. Insect Mol Biol 17, 647–656, https://doi.org/10.1111/j.1365-2583.2008.00834.x (2008).
Sim, S. & Dimopoulos, G. Dengue virus inhibits immune responses in Aedes aegypti cells. PLoS One 5, e10678, https://doi.org/10.1371/journal.pone.0010678 (2010).
Dostert, C. et al. The Jak-STAT signaling pathway is required but not sufficient for the antiviral response of drosophila. Nat Immunol 6, 946–953, https://doi.org/10.1038/ni1237 (2005).
Karst, S. M., Wobus, C. E., Lay, M., Davidson, J. & Virgin, H. W. T. STAT1-dependent innate immunity to a Norwalk-like virus. Science 299, 1575–1578, https://doi.org/10.1126/science.1077905 (2003).
Myles, K. M., Morazzani, E. M. & Adelman, Z. N. Origins of alphavirus-derived small RNAs in mosquitoes. RNA Biol 6, 387–391 (2009).
Myles, K. M., Wiley, M. R., Morazzani, E. M. & Adelman, Z. N. Alphavirus-derived small RNAs modulate pathogenesis in disease vector mosquitoes. Proc Natl Acad Sci USA 105, 19938–19943, https://doi.org/10.1073/pnas.0803408105 (2008).
Sabin, L. R., Hanna, S. L. & Cherry, S. Innate antiviral immunity in Drosophila. Curr Opin Immunol 22, 4–9, https://doi.org/10.1016/j.coi.2010.01.007 (2010).
Magwire, M. M. et al. Genome-wide association studies reveal a simple genetic basis of resistance to naturally coevolving viruses in Drosophila melanogaster. PLoS Genet 8, e1003057, https://doi.org/10.1371/journal.pgen.1003057 (2012).
Martins, N. E. et al. Host adaptation to viruses relies on few genes with different cross-resistance properties. Proc Natl Acad Sci USA 111, 5938–5943, https://doi.org/10.1073/pnas.1400378111 (2014).
Cao, C., Cogni, R., Barbier, V. & Jiggins, F. M. Complex Coding and Regulatory Polymorphisms in a Restriction Factor Determine the Susceptibility of Drosophila to Viral Infection. Genetics 206, 2159–2173, https://doi.org/10.1534/genetics.117.201970 (2017).
Li, C. X. et al. Vector competence and transovarial transmission of two Aedes aegypti strains to Zika virus. Emerg Microbes Infect 6, e23, https://doi.org/10.1038/emi.2017.8 (2017).
Joshi, V., Mourya, D. T. & Sharma, R. C. Persistence of dengue-3 virus through transovarial transmission passage in successive generations of Aedes aegypti mosquitoes. Am J Trop Med Hyg 67, 158–161 (2002).
Nayak, A. et al. Cricket paralysis virus antagonizes Argonaute 2 to modulate antiviral defense in Drosophila. Nat Struct Mol Biol 17, 547–554, https://doi.org/10.1038/nsmb.1810 (2010).
Gupta, L. et al. The STAT pathway mediates late-phase immunity against Plasmodium in the mosquito Anopheles gambiae. Cell Host Microbe 5, 498–507, https://doi.org/10.1016/j.chom.2009.04.003 (2009).
Zhang, G., Asad, S., Khromykh, A. A. & Asgari, S. Cell fusing agent virus and dengue virus mutually interact in Aedes aegypti cell lines. Sci Rep 7, 6935, https://doi.org/10.1038/s41598-017-07279-5 (2017).
Hobson-Peters, J. et al. A new insect-specific flavivirus from northern Australia suppresses replication of West Nile virus and Murray Valley encephalitis virus in co-infected mosquito cells. PLoS One 8, e56534, https://doi.org/10.1371/journal.pone.0056534 (2013).
Harris, C. et al. Polymorphisms in Anopheles gambiae immune genes associated with natural resistance to Plasmodium falciparum. PLoS Pathog 6, e1001112, https://doi.org/10.1371/journal.ppat.1001112 (2010).
Feldmann, A. M. & Ponnudurai, T. Selection of Anopheles stephensi for refractoriness and susceptibility to Plasmodium falciparum. Med Vet Entomol 3, 41–52 (1989).
Virto, C. et al. Gender-mediated differences in vertical transmission of a nucleopolyhedrovirus. PLoS One 8, e70932, https://doi.org/10.1371/journal.pone.0070932 (2013).
Mitri, C. et al. An Evolution-Based Screen for Genetic Differentiation between Anopheles Sister Taxa Enriches for Detection of Functional Immune Factors. PLoS Pathog 11, e1005306, https://doi.org/10.1371/journal.ppat.1005306 (2015).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(−Delta Delta C(T)) Method. Methods 25, 402–408, https://doi.org/10.1006/meth.2001.1262 (2001).
R Core Team. R: A language and environment for statistical computing, http://www.R-project.org/ (2012).
Fisher, R. A. Statistical Methods for Research Workers. (Oliver & Boyd, 1925).
Acknowledgements
We thank the Center for Production and Infection of Anopheles platform (CEPIA) at the Institut Pasteur, Paris for rearing mosquitoes. This work received financial support to K.D.V. from the European Commission, Horizon 2020 Infrastructures #731060 Infravec2; European Research Council, Support for frontier research, Advanced Grant #323173 AnoPath; and French Laboratoire d’Excellence “Integrative Biology of Emerging Infectious Diseases” #ANR-10-LABX-62-IBEID. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: F.N.M., K.D.V., E.B. (Belda) Performed the experiments: F.N.M., C.M., I.C. Analysed the data: F.N.M., C.M., E.B. (Bischoff), K.D.V. Wrote the manuscript: F.N.M., C.M., E.B. (Bischoff), K.D.V. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Nanfack-Minkeu, F., Mitri, C., Bischoff, E. et al. Interaction of RNA viruses of the natural virome with the African malaria vector, Anopheles coluzzii. Sci Rep 9, 6319 (2019). https://doi.org/10.1038/s41598-019-42825-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-42825-3
This article is cited by
-
Factors enhancing the transmission of mosquito-borne arboviruses in Africa
VirusDisease (2022)
-
Characterization of the virome associated with Haemagogus mosquitoes in Trinidad, West Indies
Scientific Reports (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.