Abstract
The cephalopod fauna of the southwestern Atlantic is especially poorly-known because sampling is mostly limited to commercial net-fishing operations that are relatively inefficient at obtaining cephalopods associated with complex benthic substrates. Cephalopods have been identified in the diets of many large marine species but, as few hard structures survive digestion in most cases, the identification of ingested specimens to species level is often impossible. Samples can be identified by molecular techniques like barcoding and for cephalopods, mitochondrial 16S and COI genes have proven to be useful diagnostic markers for this purpose. The Amazon River estuary and continental shelf are known to encompass a range of different substrates with recent mapping highlighting the existence of an extensive reef system, a type of habitat known to support cephalopod diversity. The present study identified samples of the cephalopod fauna of this region obtained from the stomachs of red snappers, Lutjanus purpureus, a large, commercially-important fish harvested by fisheries using traps and hook-and-line gear that are capable of sampling habitats inaccessible to nets. A total of 98 samples were identified using molecular tools, revealing the presence of three squid species and eight MOTUs within the Octopodidae, representing five major clades. These include four known genera, Macrotritopus, Octopus, Scaeurgus and Amphioctopus, and one basal group distinct from all known octopodid genera described here as Lepidoctopus joaquini Haimovici and Sales, new genus and species. Molecular analysis of large predatory fish stomach contents was found to be an incredibly effective extended sampling method for biodiversity surveys where direct sampling is very difficult.
Similar content being viewed by others
Introduction
At least 800 species of cephalopods are known to exist worldwide1,2, and an increasing number of taxa are being described3,4,5,6,7, due largely to the application of new approaches, including genetic techniques, to identify cryptic species8,9,10. However, the difficulties of collecting cephalopod specimens are a major limitation for research. Most commercial fisheries are relatively ineffective at the capture of the majority cephalopod species, and in many cases, cephalopods are found in habitats that are difficult to fish11,12.
To circumvent these limitations, studies of the diets of the predators of cephalopods, including large fishes, marine mammals, seabirds, turtles, and even other cephalopods, can provide useful information on the occurrence of cephalopod species13,14,15,16. The identification of cephalopods found in stomach contents is hampered by the fact that their bodies are composed primarily of soft tissue, which is degraded rapidly by stomach acids. While the chitinous beaks are resistant to this acid, they typically permit identification only to genus or family level, making it impossible to differentiate sympatric congeneric species17,18. The identification of species can be achieved by using molecular techniques19,20,21, which have been used successfully when samples cannot be identified reliably based on their morphological characteristics.
In this context, DNA barcoding, which uses standardized protocols for the sequencing of a short fragment of the mitochondrial Cytochrome Oxidase I (COI) gene22, has proven to be very effective for the identification of species in many taxonomic groups including butterflies23, birds24, and plants25, as well as cephalopods26. In addition to the COI gene fragment, the large ribosomal subunit (16S rDNA) has also proven to be useful for the identification of cephalopod species at the intra-familial level9,27.
In northern Brazil, as in most of the tropical Atlantic, there are no fisheries specialized in harvesting cephalopods. Given this, samples are normally obtained as by-catch, usually from shrimp fisheries operating trawls over soft substrates, which can be an inefficient collection method for a variety of benthic cephalopod taxa. The present study focused on the poorly-known cephalopod fauna of the region of the Amazon Reef System (ARS), off the coast of northern Brazil28. This system was originally estimated to encompass an area of 9,500 km2, but it may be as large as 50.000 km2, and includes complex bottom habitats29, including biogenic reefs and basement outcrops, which are difficult to fish using nets. The basic consolidated substrates in the ARS are rhodolith beds, which might build extensive pavements. These pavements are often covered (and exposed) by sand wave migration, resulting in three-dimensional complexity. These features are intensively colonized by sponges, which results in additional complexity. Furthermore, fishes (e.g. Holocentridae Richardson, 1846) are excavating and reworking partially sand-recovered rhodolith beds, creating void spaces and building rhodolith mounds. Both voids and mounds are potential habitats for octopuses. On the other hand, the history of sea-level fall and rise over the Amazonian shelf has resulted in alternating deposition, formation and erosion of sedimentary deposits and rocks, where laterite outcrops seem to be particularly frequent in the southern sector of the ARS, offering abundant habitats for octopuses.
The study was based on the identification of the cephalopods found in the stomachs of red snapper, Lutjanus purpureus (Poey, 1875), a commercially important fish associated with these bottom habitats, which is harvested using traps and hook-and-line techniques. The red snapper is a large fish, which reaches 1 m in length and up to 10 kg in weight and is an active predator of benthic invertebrates. As it also tends to swallow its prey items whole, red snapper was selected as a likely candidate that could be used as an effective biological sampling tool for cephalopods in habitats that are difficult to survey30,31,32,33.
In this paper we describe the diversity of cephalopods found in the region, including a new genus and a new species of octopus that is both morphologically and genetically distinct from other genera and species in the Octopodidae.
Results
Genetic characterization
Squids
All squids sampled in this study (N = 14, 100%) were successfully amplified for the chosen fragment of the 16S rDNA gene but could not be amplified for the COI gene fragment. Analyses of squid data was therefore limited to 16S sequences. The GTR + I + G (−InL = 2738.3035) (AIC) and GTR + G (−InL = 2740.2735) (BIC) evolutionary models were selected by jModelTest2 for ML and BI analyses respectively. Comparisons with the GenBank sequences (selected using BLAST) associated the specimens to three species in the genera Doryteuthis Naef, 1912 and Abralia Gray, 1849 (Supplementary Data 1, Fig. 1). The majority (N = 12) of the individuals were genetically similar to Doryteuthis plei Blainville, 1823 (0.0–0.5% uncorrected p-distance), a species found on the Atlantic coast of Brazil9,34 (Supplementary Data 1). One specimen was genetically identical to samples identified as Doryteuthis pealeii Lesueur, 1821 (0.0% p-distance). A member of a probable species complex within D. pealeii is also found on the Atlantic coast of Brazil9. The last individual could only be assigned to the genus Abralia (grouping with other members in a clade with strong support: NJ/ML/BI 91%/88%/0.96) but was distinct from all the 16S sequences of Abralia species available in GenBank, with uncorrected p-distance ranging from 4.6% in relation to Abralia adamanica Goodrich, 1896 to 6.2% in the case of Abralia trigonura Berry, 1913 (Supplementary Data 1).
Phylogenetic tree based on Bayesian Inference of the mitochondrial 16S rDNA gene of the squid collected from the stomachs of the red snappers sampled in the present study. The values at the nodes show the bootstrap percentage for the Neighbor-Joining and Maximum Likelihood analyses, and a posteriori probabilities of Bayesian Inference analysis, respectively. Only values above 70%/0.7 are shown.
Octopods
Of the 130 sampled octopods, 84 were successfully sequenced for one or another target sequence, including 61 (~73%) for 16S rDNA and 61 (~73%) for COI. Unfortunately, successful amplification of both target sequences was only possible for 38 samples (Supplementary Data 2). For the 16S sequences, the evolutionary models selected were GTR + I + G (−InL = 2738.3035) (AIC) for ML analysis, and GTR + G (−InL = 2740.2735) (BIC) for BI analysis. In the case of the COI gene, the TIM2 + I + G (−InL = 3313.5079) model was selected (AIC and BIC) for both the ML and the BI analyses. Five principal monophyletic groups were recovered in these analyses, including the four genera, Macrotritopus Grimpe, 1922, Octopus Cuvier, 1797 (but note that Octopus itself is not monophyletic), Scaeurgus Troschel, 1857 and Amphioctopus Fischer, 1882, and a fifth group distinct from all octopodid genera that have sequences in GenBank (Fig. 2A).
(A) Phylogenetic tree based on Bayesian Inference of the mitochondrial 16S r DNA gene of the octopuses collected from the stomachs of the red snappers sampled in the present study. The values at the nodes show bootstrap percentage for the Neighbor-Joining and Maximum Likelihood analyses, and a posteriori probabilities of the Bayesian Inference analysis, respectively. Only values above 70%/0.7 are shown. (B) Phylogenetic tree based on Bayesian Inference for the analysis of the mitochondrial COI gene of the octopuses, showing the same parameters analyzed for the 16S gene. Green clade: Macrotritopus sp. and most similar sequence from GenBank. Grey clade: Octopus vulgaris species complex and most similar sequences from GenBank. Red clade: Lepidoctopus joaquini gen. et sp. nov. and most similar sequence from GenBank. Blue clade: Scaeurgus sp. and most similar sequences from GenBank. Yellow clade: Amphioctopus sp. and most similar sequences from GenBank.
The samples identified morphologically as Macrotritopus sp., grouped with sequences that have been given the name “Blandopus white V” in the phylogenies for both the 16S dataset (4.4–5.2% uncorrected p-distance) and the COI dataset, with 4.9–5.5% uncorrected p-distance (Supplementary Data 2A,B, Fig. 2). In the case of the samples identified as Octopus, the two lineages found in the 16S dataset were most closely associated with sequences published in GenBank as Octopus vulgaris Cuvier, 1797 from Brazil (KF843970/KF843972/KF843973/KF843974/KF843977) and from Venezuela (AJ252770/AJ390316). Although there was some overlap, within lineage uncorrected p-distance (0.0–0.9%, mean = 0.3%) was lower than between lineage uncorrected p-distance (0.5–1.6%, mean = 1.1%). For COI there was no overlap. Within lineage uncorrected p-distances were all 0.0%. A 0.5% distance exists between the first lineage containing only Brazilian samples and a second lineage containing the sample OvuPA184 (sequence KF844030). Both lineages were yet more divergent (2.1% uncorrected p-distance for the first, and 1.6% uncorrected p-distance for the second) from a third lineage including the Brazilian sample OvuPA173 (sequence KF844027) and other GenBank sequences (Fig. 2 Supplementary Data 3C,D). This indicates the presence of distinct haplogroups in the region that unfortunately cannot be associated to the described Types I or II35 due to the state of the voucher material. Furthermore, another sequence obtained in the present study was identical to the sequence of a sample previously identified as Octopus hummelincki Adam, 1936 from the northeastern Brazilian state of Ceará (0.0% uncorrected p-distance for both 16S and COI sequences).
The samples identified as Scaeurgus unicirrhus Delle Chiaje, 1839–1841 are represented by two lineages with low divergence (uncorrected p-distances of 0.0–1.2% for 16S and 0.0–1.0% for COI, Supplementary Data 2E,F). One of the lineages is more closely related to GenBank sequences for Scaeurgus unicirrhus based on 16S sequences, but both lineages were more closely related to each other and approximately equidistant from GenBank sequences of S. unicirrhus based on COI data, (Fig. 2B), indicating the likely existence of distinct stocks within the species.
For the genus Amphioctopus new sequences formed a strongly-supported monophyletic group along with available GenBank sequences (Fig. 2B, node with NJ/ML/BI support 99%/99%/0.99), and uncorrected p-distance values of between 0.5–2.74% for 16S and 0.2–4.2% for COI (Supplementary Data 2G,H) within the clade. These specimens were difficult to identify morphologically due to their degraded condition, but they could be identified based on their COI sequences. Two of the samples were genetically identical to AmspPA86 (GenBank access code KF844045)36 and unpublished sequences of specimens collected offshore from the state of Rio de Janeiro, in southeastern Brazil, which were identified morphologically as Amphioctopus burryi Voss, 1950 (M. Haimovici; unpublished data).
The fifth group of specimens could not be associated clearly with any known genus (herein described as Lepidoctopus gen. nov.). Some individuals were linked weakly with GenBank sequences named as Octopus rubescens Berry, 1953 based on the 16S data (BI = 0.96), but the uncorrected p-distance between these sequences was 12.0–12.2% (Fig. 2A, Supplementary Data 2G).
The complete phylogeny, based on the samples with data for both markers (concatenated dataset), provides a more reliable overview of the arrangement of the principal genera within the family Octopodidae (Fig. 3). This analysis revealed that Macrotritopus is most closely related to the other mimetic forms (Thaumoctopus Norman & Hochberg 2005, “Blandopus”, and Wunderpus Hochberg, Norman & Finn, 2006), with Abdopus aculeatus d’Orbigny, 1834 as a probable basal member of the clade (Fig. 3). Once again, the group of genetically distinct individuals that could not be assigned to any genus in the previous analyses (herein described as Lepidoctopus gen. nov.) formed a highly divergent lineage, with no well-supported relationship with any genus for which sequences are available. The phylogeny (Fig. 3) indicates a polytomy including this divergent lineage and a miscellaneous group including the clades ((Octopus rubescens + Muusoctopus longibrachus longibrachus Ibãnéz, Sepúlveda & Chong, 2006) + Eledone Leach, 1817), (Scaeurgus + Callistoctopus Taki, 1964), Octopus tehuelchus d’Orbigny, 1834 and (Amphioctopus + Hapalochlaena Robson, 1929).
Phylogenetic tree based on Bayesian Inference for the concatenated 16S + COI data set of the octopuses analyzed in the present study. The values at the nodes refer to the bootstrap percentage for Maximum Likelihood and a posteriori probabilities of the Bayesian Inference analysis, respectively. Only values above 70%/0.7 are shown. Green clade: Macrotritopus sp., represented by sample octo 97 in photo. Grey clade: Octopus spp., represented by sample octo 148 in photo. Blue clade: Scaeurgus sp., represented by samples octo 58, 59 and 60 in photo. Red clade: Lepidoctopus joaquini gen. et sp. nov., represented by sample octo 175 in photo. Yellow clade: Amphioctopus sp., represented by sample octo 114 in photo.
Morphological characterization
Squids
Squid could only be identified to family level because of the loss of most morphological diagnostic characteristics of the samples.
Octopods
Based on external morphological features alone, only three groups could be identified in the 130 octopuses examined. Some specimens, including those identified based on sequence identity as Scaeurgus unicirrhus and Amphioctopus burryi were too degraded to confirm morphological identification.
Macrotritopus sp
Small octopus, the largest sampled specimen was 237 mm total length and 31 mm mantle length. Mature females bore a large number of 1–2 mm long oocytes. The mantle is oval, mantle width is 32–41% of the mantle length. The mantle is grayish dorsally and cream ventrally, the eyes are large and protuberant, 11–17% the mantle length, the funnel is moderately long with more than half its length free. The gills have 9–10 lamellae in each of the two hemibranchs. The web is short, and the arms long and thin with two series of relatively small suckers. The ventral arms are the longest and reach five times the length of the mantle. A single male had both third arms and the right hectocotylized arm was 76% as long as the left arm. Short hectocotylus (2% of the arm length) with a small calamus, and a ligula with a longitudinal groove. These characteristics are broadly consistent with the description of Macrotritopus defilippi (Verany, 1851)37 and of Macrotritopus beatrixi Guerrero-Kommritz & Rodriguez-Bermudez38. The known distribution of Macrotritopus defilippi is in the Mediterranean and northeastern Atlantic Ocean, while some authors apply the same name to specimens with similar characteristics from the Caribbean and southwestern Atlantic Ocean39,40 and M. beatrixi is only described for the Caribbean Sea along Colombia38. To elucidate if our specimens pertain to either species or to an undescribed third one requires more reliable morphological data for a formal description.
Octopus vulgaris species complex
Most specimens of this type were in poor condition. One very well-preserved individual, which unfortunately we were not able to sequence, coincided with the description of the Octopus vulgaris species complex.
Specimens not associated with any known genus
Among the better-preserved specimens included a group of small octopuses with long arms and a moderately thick and short web. The mantle, head and base of the arms is covered with large rounded papillae, resembling dermal cushions, some of which have long conical cirri with multiple tips. In males, the third right hectocotylized arm has a very short calamus and a long, slender ligula with a deep longitudinal groove. This combination of characters was not recorded in any genus of the Family Octopodida35. We consider these specimens to be an undescribed species of a newly discovered genus, as follows:
Family Octopodidae d’Orbigny, 1840
Subfamily Octopodinae d’Orbigny, 1840
Lepidoctopus joaquini Haimovici and Sales, gen. et sp. nov.
Diagnosis of the genera and species: small sized benthic octopod, largest examined specimen 40 mm mantle length (ML); mantle, head and base of arms covered by large rounded papillae-like dermal cushions more densely packed and larger on dorsal mantle and smaller on head and web; some papillae on dorsal mantle bear cirri branched in multiple tips; no lateral ridge observed; eyes moderate in size, slightly protruding and with single supraocular papillae with large branched cirrus, funnel half of ML; arms long, first and second typically around 4.5 times ML, third and fourth under 4.0 times ML; web typically half of ML; normal arms with up to 170 suckers, first 4 to 6 proximal suckers in single series, followed biserial to tips of arms; enlarged suckers in fourth to sixth biserial rows of males; third left arm of males hectocotylized, ~77% of the opposite third arm with 68 to 72 suckers, Short hectocotylus, ~22% of hectocotylized arm, with short conical calamus, 2% of ligula length, slender ligula with deep longitudinal groove ending in blunt tip.
Material examined
Eleven specimens, eight males and three females among the best-preserved specimens collected in stomach contents of the snapper Lutjanus purpureus commercially fished with a fish trap or lines off northern Brazil. All deposited in the mollusk collection of the Oceanography Museum of the University of Rio Grande (MORG) in Rio Grande, Rio Grande do Sul, Brazil.
Holotype
(Sample PV192) Mature male 28 mm ML, coastal waters off Cabo Norte, Amapá State, northern Brazil, 03°34′27″N, 50°03′49″W, 50–60 m. in 14 Dec. 2008. MORG coll. num. 51455. Molecular sequence: GenBank Accession No. MG010599 (COI)
Paratypes
Sample PV135 One mature male 36 mm ML, coastal waters off Pará State, northern Brazil, 00°05′53″N 47°14′08″W, 20–30 m in 17 Dec. 2007. MORG coll. num. 51448.
Sample PV137 One mature male 40 mm ML, coastal waters off Pará State, northern Brazil, 00°05′53″N 47°14′08″W, 20–30 m in 17 Dec. 2007. MORG coll. num. 51449. Molecular sequence: GenBank Accession No. MG010602 (COI).
Sample PV139 One mature male 33 mm ML, coastal waters off Pará State, northern Brazil, 00°05′53″N 47°14′08″W, 20–30 m in 17 Dec. 2007. MORG coll. num. 51450. Molecular sequence: GenBank Accession No. MG010603 (COI).
Sample PV142 One mature male 33 mm ML, coastal waters off Pará State, northern Brazil, 00°05′53″N 47°14′08″W, 20–30 m in 17 Dec. 2007. MORG coll. num. 51451. Molecular sequence: GenBank Accession No. MG010600 (COI).
Sample PV162 One mature male 29 mm ML, coastal waters off Pará State, northern Brazil, 00°05′53″N 47°14′08″W, 20–30 m in 12 Nov. 2008. MORG coll. num. 51452.
Sample PV182 One mature male 29 mm ML, coastal waters off Pará State, northern Brazil, 00°05′53″N 47°14′08″W, 20–30 m in 12 Nov. 2008. MORG coll. num. 51453. Molecular sequence: GenBank Accession No. MG010594 (COI).
Sample PV191 One mature male 31 mm ML, coastal waters off Pará State, northern Brazil, 00°05′53″N 47°14′08″W, 20–30 m in 14 Dec. 2008. MORG coll. num. 51454. Molecular sequence: GenBank Accession No. MG010598. (COI).
Sample PV175 One mature female 30 mm ML, coastal waters off Pará State, northern Brazil, 00°05′53″N 47°14′08″W, 20–30 m in 12 Nov. 2008. MORG coll. num. 51456. Molecular sequence: GenBank Accession No. MG010542 (16S) and MG010592 (COI).
Sample PV177 One mature female 33 mm ML, coastal waters off Pará State, northern Brazil, 00°05′53″N 47°14′08″W, 20–30 m in 12 Nov. 2008. MORG coll. num. 51457. Molecular sequence: GenBank Accession No. MG010543 (16S) and MG010602 (COI).
Sample PV188 One mature female 34 mm ML, coastal waters off Pará State, northern Brazil, 00°05′53″N 47°14′08″W, 20–30 m in 12 Nov. 2008. MORG coll. num. 51458.
Description
The description is based on eight males and three females in initial stages of digestion. Most had the skin damaged and either the arm tips or distal suckers were missing from some arms including all non-hectocotylized arms. However, a sufficient number of arms and web depth could be measured to calculate approximately the arm and web depth formulas. The largest male is a mature specimen, 40 mm ML, 274 mm TL, 36.2 g TW, with fully developed spermatophores in the spermatophoric sac. The largest female in fair condition presented a large ovary occupying the posterior half of the mantle cavity and was full of developing eggs. This female measured 34 mm ML, 218 TL and weighed 19.9 g TW. Morphometric measurements are presented in Table 1 and morphometric indices in Table 2. Values in the text are reported as mean value and range.
Morphology
Mantle oval and elongate, width 61% (50–68%) of ML; head is narrower than mantle, 45% (38–55%) of ML (Fig. 4a); Mantle, head, and arms in preserved specimens reddish brown dorsally, lighter cream ventrally (Supplementary Data 3 and 4); Skin of mantle, head and base of arms sculptured with large rounded, densely packed papillae-like dermal cushions with chromatophores on borders (Supplementary Data 5); Papillae on dorsal mantle larger, some modified with long conical cirri branched in multiple tips; Papillae on ventral mantle smaller with fewer chromatophores (Fig. 4b); No lateral ridge observed.
Lepidoctopus joaquini gen. et sp. nov.: (a) 40 mm mantle length male with reconstructed arms and sucker, (b) lateral view of the mantel and head with papillae and cirrus pattern. (c) Stylet. (d) Lateral view of lower (left) and upper (right) beaks. (e) Ventral view of lower beak (left) and dorsal view of upper beak (right), (f) radula. Each drawing has a specific scale bar. All imagens produced by contracted biological artist Leticia Cavole.
Eyes large, 14% (10–18%) of ML, slightly protruding; Supraocular papillae present with large branched cirrus (Fig. 4b).
Stylets present in mantle muscle at base of gills; Stylets relatively long, 35% of ML, with angle at 1/3 of its length, shorter posterior limb thicker and with distinct deep groove in which growth lines observed (Fig. 4c); Funnel relatively long, 48% (44–53%) of ML, free for 56% (41–60%) of its length, reaching anterior margin of eyes; Funnel organ not observed; Arms long and rather stout; First, second and third typically around 4.5 times ML fourth around 3.5 times ML (arm formula: 1 = 2 = 3 > 4); Arm of moderate width, 14.3% (11.4–17.2%) of ML; No difference in width observed between arms within individuals or between sexes; Webs moderately short, 76% of ML; Web formula non-consistent between specimens; First 4 to 6 proximal suckers in single row followed by biserial suckers to tips of arms; Suckers moderately sized, larger suckers did not differ between arms in females, 7.2% (5.7–7.9%) of ML; In males, 4 to 6 enlarged suckers present on all arms 11.6% (9.0–14.3%) of ML; Suckers on distal half of arms much smaller than those in proximal half (Fig. 4a); Third left arm of males hectocotylized and shorter (Supplementary Data 6), 77% (70–81%) of third right arm; Hectocotylized arm bears 68 to 82 suckers; Counts of number of suckers of all other arms not accurate due to degradation; Three largest counts ranged from 164 to 170; Ligula long, 16% (13–18%) of hectocotylized arm, and slender with deep longitudinal groove, ending in blunt tip; Calamus conical, very small, 5.5% of ligula length (Fig. 5a); Gills relatively long, 32% to 43% of ML, 7 lamellae on both inner and outer demibranchs in all examined specimens (gill lamellae number formula: L = 7/7; R = 7/7); Upper beak robust, with hooked rostrum and moderately long hood, 34% of total beak length, lower beak with rounded rostrum and hood measuring 62% of crest length (Fig. 4d,e); Radula with seven teeth and two marginal plates in each transverse row (Fig. 4f); Rhachidian tooth with one to two lateral cusps, typically two, on each side of large medial cone; lateral cusps in symmetrical seriation, migrating from lateral to medial position over approximately five to six transverse rows.
Lepidoctopus joaquini gen. et sp. nov.: (a) Copulatory organ. (b) Male reproductive tract. (c) Spermatophore. (d) Female reproductive tract. (e) Schematic view of maturing ovary. (f) Maturing oocyte. Each drawing has a specific scale bar. All imagens produced by contracted biological artist Leticia Cavole.
Digestive tract
Anterior salivary glands typically discoid, relatively small, 1/3 length of buccal mass; Posterior salivary glands large, each approximately as big as buccal mass; Elongated crop with distinct diverticulum; Stomach and caecum of same size; Digestive gland ovoid to lobed; Ink sac present, embedded in digestive gland (Fig. 4g); Distal half of intestine of specimens not recovered, presence of anal flaps undetermined.
Male reproductive system
Testicles in mature specimens oval, vas deferens very long thin and coiled; First spermatophore gland long and thin and second spermatophore gland short; Spermatophore sac of mature males containing (30 to 50) fully formed spermatophores; Terminal organ in mature males small, curved and with moderately large diverticulum (Fig. 5b); Spermatophores short, 10.5 to 12.3 mm, around 1/3 ML and thin (2–3 mm, mean 2.75 mm); sperm reservoir around half of spermatophore length (50–55% ML), with sperm cord coiled in (48–56 mm, mean 54.5 mm); Cement and ejaculatory apparatus approximately 16% and 27% of spermatophore length respectively (Fig. 5c).
Female reproductive system
Maturing females with ovary occupying posterior half of mantle cavity, short proximal oviducts, oviducal glands with distinct radial chambers and longer distal oviducts (Fig. 5d); Eggs small, striated, almost mature ovarian eggs of ovaries 1.5–2.1 mm long, with median length 1.7 mm (Fig. 5d,e); Fecundity of 35 mm ML female estimated by gravimetric method at around 4000 eggs.
Etymology
The genus name is a combination of lepido from scale in Greek, referring to the scaly appearance given by the large almost overlapping papillae (dermal cushions) on its skin giving it a peculiar appearance as if it is covered with scales, and octopus. The name joaquini refers to the young son of the first author of this paper (JBL Sales).
Discussion
The detailed morphological and genetic analyses presented here indicate that the red snapper, Lutjanus purpureus, can provide informative samples of the benthic and neritic benthopelagic cephalopods of the families Octopodidae and Loliginidae found in the southwestern Atlantic. In fact, the results of the present study indicate a much greater diversity of cephalopods in this region, in comparison with previous analyses. Barroso41 identified the genera Octopus and Loligo Lamarck, 1798 (now Doryteuthis) in the diet of L. purpureus (under the synonym Lutjanus aya Bloch, 1795) in specimens collected off northeastern Brazil. In the same region, Furtado-Ogawa & Menezes30 recorded a diet in this species (L. purpureus) dominated by fish, supplemented primarily by crustaceans and mollusks, including both loliginids and octopodids. In the diet of Lutjanus analis Cuvier, 1828 from Colombia, the only cephalopods identified by Duarte & Garcia33 were loliginid squid and octopuses of the genus Octopus. Fonseca42 found that the cephalopod prey of Lutjanus synagris Linnaeus, 1758 and Ocyurus chrysurus Bloch, 1791 collected in the Abrolhos Bank in the Brazilian state of Bahia included both octopodids and loliginids.
Squids
The fishermen who provided the snapper specimens reported that squids were rarely found in the stomachs of these fish, and this is confirmed by the relatively small number of squid samples collected in the present study. The identification of both Doryteuthis plei and Doryteuthis pealeii in the samples analyzed was expected, given the known occurrence of these species in the study region9, and their use of demersal habitats43.
The genus Abralia Gray, 1849 has 19 species, and the specimen collected in the present study is likely to belong to one of the five species known to occur in the Atlantic Ocean, and specifically, to one of the two species for which no 16S sequences are available. While the present specimen was associated with the three species (Abralia veranyi Ruppel, 1844, Abralia trigonura, and A. andamanica) for which 16S sequences are available, and could thus be assigned to this genus, the uncorrected p-distance values were well above the levels used to identify cephalopods species26,36. This indicates that the specimen represents one of the other Atlantic species, either Abralia redfieldi Voss, 1955 or Abralia grimpei Voss, 195944, for which no 16S sequences are available. Abralia are mesopelagic, upper slope squid that undertake diel vertical migration, remaining in deep water during the day (700–800 m in A. veranyi), and coming to the surface (40–60 m in A. veranyi) at night44,45. It is probably preyed on only occasionally by neritic lutjanids, and is likely to be captured only very rarely by any fishing technique.
Octopods
Macrotritopus sp
No sequences (COI or 16S) are available for Macrotritopus in GenBank. The closest taxon with which the specimens collected in the present study were associated was “Blandopus” White V, designated “White V Octopus”, Octopus sp.18 by Norman37, and subsequently as “Blandopus” by Hanlon et al.46. Both Macrotritopus cf. defilippi and “Blandopus” White V is long-armed species known to mimic other organisms in a manner similar to Thaumoctopus mimicus Norman & Hochberg, 2005. In particular, Macrotritopus cf. defilippi (probably now the species more recently described as M. beatrixi) has been observed mimicking flatfish when moving over sandy substrates in the Caribbean40. The complete phylogeny based on the concatenated data (Fig. 3) clearly groups together all the mimetic octopuses, Thaumoctopus mimicus47, Wunderpus photogenicus Hochberg, Norman & Finn48, “Blandopus” white V46, and Abdopus aculeatos49 with high support values in general, with slightly weaker support for the association Abdopus aculeatus at the base of the clade.
Octopus vulgaris species complex
These samples were identified genetically as Octopus vulgaris from the southwestern Atlantic, and most closely related to the O. vulgaris types I and II of Jereb et al.50. While type II has only been reported previously from the southwestern Atlantic, Fig. 3b indicates the existence of well-supported clades in O. vulgaris that include sequences of samples from Brazil (including one from the same region as the present study - see Sales et al.36), France, the mid-Atlantic Island of São Paulo, the southeastern Indian Ocean, Turkey and Venezuela. The genus Octopus has been the subject of taxonomic revisions5,6, but the cryptic species complex of the western Atlantic Octopus vulgaris has only recently begun to be unraveled51, showing that Octopus vulgaris is composed of multiple O. vulgaris-like species that are currently being incorrectly treated under a single species name. Our concatenated data indicate the presence of more than one lineage of the Octopus vulgaris species complex along the Brazilian Coast. Including more individuals (not from stomach samples, and from other regions of Brazil) increases support values of all clades (Europe, South Africa, and two lineages from Brazil) in phylogenetic analyses (unpublished data). More individuals from along the South American coast (including from stomach contents) will help shed new light in this puzzle.
Scaeurgus unicirrhus
Species of the genus Scaeurgus are deep-water, benthic octopuses, found on different substrates of the continental shelf and on slopes at depths of between 100 m and 500 m52,53. Scaeurgus species are found in all tropical and temperate seas. Until recently, the genus was divided into only two species, distributed in the Atlantic (S. unicirrhus) and the Pacific (Scaeurgus patagiatus Berry, 1913) oceans. However, Kubodera & Lu54 found evidence of a potential new species off the coast of Taiwan, and Norman et al.53 described three new species - S. tuber Norman, Hochberg & Boucher-Rodoni, 2005, S. jumeau Norman, Hochberg & Boucher-Rodoni, 2005 and S. nesisi Norman, Hochberg & Boucher-Rodoni, 2005 - from New Caledonia and sea-mounts in the nearby Coral Sea.
The samples analyzed in the present study produced contrasting topologies. While the individual 16S and COI datasets indicated the existence of two Scaeurgus clades (Fig. 2), the concatenated data pointed to a single group. Given the discrete levels of p-distance divergence found between clades (0.0–1.2% in the 16S and 1.0% in COI), the results of these analyses are consistent with the presence of a single species with population structuring.
Amphioctopus
All Amphioctopus samples suffered a high degree of degradation from digestive processes and could only be identified by molecular techniques. Amphioctopus was recently revalidated in reviews of the genus Octopus (Octopus aegina group), and includes species targeted by commercial fisheries, especially in Southeast Asia5,48,55. Amphioctopus burryi (Voss, 1950) is the only species documented from the southwestern Atlantic Ocean. A single individual, identified only as Amphioctopus sp. (AmspPA86), was described from this region by Sales et al.36. The samples identified in the present study (151 and 152) were identical to AmspPA86. Together with well-preserved material obtained recently from Mexico and the Brazilian state of Rio de Janeiro, all known specimens from the region can be identified as A. burryi (Figs 2 and 3). Further analyses will nevertheless be necessary to confirm the phylogenetic position of this taxon in relation to the genera Amphioctopus, Hapalochlaena and Callistoctopus considering the polyphyletic characteristics of Amphioctopus, as determined by Dai et al.26.
Lepidoctopus joaquini Haimovici and Sales, gen. et sp. nov
This clade appears to be part of a weakly supported polytomic clade (0.84 posterior probability) including the genera Eledone, Muusoctopus, Scaeurgus, Callistoctopus (sensu Strugnell et al.56) and the species O. rubescens and O. tehuelchus, (Fig. 3). Octopus tehuelchus has been considered more closely related to Callistoctopus than other Octopus species57. Based on the form of the hectocotylus of the better-preserved specimens, the new taxon appears to be more similar morphologically to Callistoctopus, which is represented by a single species in the eastern Atlantic (Callistoctopus macropus). However, the uncorrected p-distances of 12.6% from Callistoctopus luteus and 11.9% from Callistoctopus minor, indicate that the specimens are only distantly related to Callistoctopus, and, when compared with divergence values between described cephalopod genera, supports the hypothesis that these samples represent a different genus. Additionally, none of the other 21 genera within the Octopodidade35 share the morphological characters of dense dermal cushions and a long ligula with a longitudinal groove, supporting the generic status of this new taxon.
Sampling cephalopod diversity through the analysis of stomach contents
The cephalopod diversity of South and Central America is still relatively poorly known, and the findings of the present study have further highlighted the ongoing trend of species descriptions over the past 60 years8,58,59,60, and the increasing evidence of genetic divergence along the Atlantic coast of the Americas34.
While many previous studies have used molecular tools to identify species to understand the diet of a particular predator species or increase sampling efficiency of specific known prey taxa61,62 none have used molecular studies of stomach contents as an approach for surveying the general biodiversity of a taxonomic group in relatively inaccessible habitats. The selection of a suitable predator that feeds in or around the target environment is fundamental to the success of this approach. In the present study, cephalopod samples were obtained from substrates that are not normally targeted by fisheries using gear suitable for the capture of these organisms. The sampling of stomach contents also circumvents the problem of vertical migrations of the target cephalopods, given that the predators may “sample” their prey at any time of day. While, this aggregative effect may imply imprecision of the sampling location, the ability to adequately identify morphological characters and produce DNA sequence data implies a degree of sample freshness and at least a moderate precision of the location. Although the digestive degradation of the specimens does limit formal taxonomic descriptions in some cases, repeated sampling may well provide more intact specimens suitable for morphological description and identification.
The use of a metagenomic approach to screen stomach contents rapidly for a range of prey organisms (e.g. Berry et al.63) should increase considerably the occurrence data for the target species. This approach should be complemented with classic morphological analyses, when possible, and individual molecular diagnoses, to provide a systematic record of the principal items in the diet of the predator, in addition to reference material for the description of new taxa.
This study demonstrated the potential of a combined morphological and molecular approach for the identification of the cephalopod fauna of areas that are difficult to sample directly. While many samples had been degraded by the digestive process, it was possible to obtain sequences from 100% of squid samples and approximately 63% of the octopod samples collected (14/14 squid and 82/130 octopods). This reinforces the potential of molecular tools for the identification of the species in aquatic food chains, as well as the understanding of the diversity of poorly-studied regions and hard-to-sample environments, such as the continental shelf and reef systems found off the mouth of the Amazon River.
Methods
Ethics statement
All methods were carried out in accordance with relevant guidelines and regulations. This included full license for the collection of biological material from already deceased animals (Under Brazilian biodiversity collection authorization license: SISBIO Permanent License 5857–1). As all material sampled in this project obtained from commercial fishermen was already dead, there was no requirement for ethical approval of sampling protocols as it did not include live organisms.
Sampling and sample identification
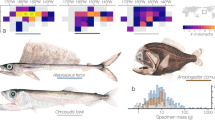
Partially digested and near-intact (which generally lacked the suckers or the tips of the tentacles) cephalopods were collected from the stomachs of recently fished red snapper, Lutjanus purpureus, between 2006 and 2008 at two fishing grounds in the continental shelf off the Amazon coast of the Brazilian states of Amapá and Pará (Fig. 6A). One area is located in the northern sector of the Amazon Reef System (00°05′53″N, 50°03′49″W) where the plume of the Amazon River flows primarily to the northwest, and the Photosynthetically Active Radiation (PAR) almost never reaches the bottom, even though the euphotic layer increases with increasing depth. The second area is located in the central sector of the reef system (03°34′27″N, 47°14′08″W), where the PAR often reaches the bottom, but with substantial spatial and temporal variation related to the dynamics of the Amazon plume, resulting in a shallow, mesophotic environment28.
(A) Area in which the red snappers analyzed in the present study were captured. The red dots indicate the points at which the traps were set. (B) Cephalopods found in the stomachs of snappers recently captured within the study area. Amapá Coast sampling point: 00°05′53″N, 50°03′49″W. Pará State sampling point: 03°34′27″N, 47°14′08″W.
The red snappers were caught by a small-scale artisanal fishery fleet based in the town of Bragança, Pará. Crew members confirmed fishing in areas at depths of up to 80 m over rocky and muddy substrates where nets cannot be used, and traps are frequently lost due to both to the characteristics of the substrate and the extremely turbulent currents. The cephalopod samples were washed immediately after collection to remove external contaminants, and then rinsed with diluted bleach to degrade any remaining red snapper DNA, washed again with water and then preserved directly in absolute ethanol for transportation to the laboratory in the town of Bragança (Fig. 6B). In total, 130 octopod samples and 14 squid samples were obtained during the study period. A small tissue sample was removed for molecular analyses, and the remaining specimen was fixed in a 10% formalin solution before being shipped to the Laboratory for Demersal Resources and Cephalopods at the Federal University of Rio Grande, Rio Grande do Sul state, Brazil for examination and identification based on external morphology. The complete list of specimens sampled are found in Supplementary Data 7.
As many of the squid specimens lost their skin, suckers, and part of their arms, they could only be identified morphologically to family (Loliginidae Lesueur, 1821). Some octopuses were better preserved, and it was possible to measure the length and width of the mantle, total length, the length and width of some arms, and the characteristics of the hectocotylized arm in males. In some cases, the diameter and the number of oocytes, the number and length of the spermatophores and the number of gill lamellae could also be determined, and a more detailed morphological description was possible.
Extraction, amplification and sequencing of the DNA
The DNA was extracted using three different protocols. The phenol-chloroform protocol64 was applied to samples that were relatively well-preserved. The DNA of the more degraded samples was extracted using either the Qiagen DNeasy kit (Valencia, CA) or the Invitrogen Pure Link Genomic DNA Mini kit (Carlsbad, CA), using the mouse tail protocol in both cases. The fragments of the mitochondrial 16S rRNA and Cytochrome Oxidase I (COI) genes were amplified by PCR using the procedure described by Sales et al.36 and the primers L1987/H2609 proposed by Palumbi et al.65 and LCO1490/HC02198 Folmer et al.66.
Molecular analyses
Sequences were aligned automatically using ClustalW67, run in BioEdit v.7.0.368, and then visually inspected for errors. In cases of doubt, bases were called conservatively. Sequences were deposited in GenBank under the following accession numbers: For 16S squid: MG010606 - MG010619, for 16S octopus: MG010487–MG010544 and for COI Octopus: MG010545–MG010605. Only the 16S rRNA sequence (495 bps) was sequenced successfully in the case of squid samples, providing a single data set. For octopuses, both markers were amplified in only a subset of specimens, creating three distinct data sets: (i) 16S rRNA (541 bps), (ii) COI (659 bps), and (iii) the concatenated 16S and COI sequences. For the individual markers, GenBank sequences were only included in the analyses when similarity was at least 95%. For the concatenated data set, additional sequences representing all the genera of the family Octopodidae d’Orbigny, 1840 were obtained from GenBank to determine the most likely phylogenetic position of the samples that could not be identified morphologically. As all the BOLD sequences were also available in GenBank, only the GenBank sequences were included in the analyses. For all analyses, Argonauta nodosa Lightfoot, 1786 was used as outgroup.
MEGA 6.069 was used to calculate uncorrected p-distances for all datasets to allow direct comparison across datasets and with published data. For production of phylogenetic trees, distances were based on the best evolutionary model selected for jModelTest270 for each database. Neighbor-Joining (NJ) trees71 were produced, with support for the nodes based on 1000 bootstrap generations72. Maximum Likelihood (ML) and Bayesian Inference (BI) approaches were used to verify the clusters produced by the NJ trees and to analyze the concatenated dataset including all the species. The COI data set was partitioned (codon positions: 1 vs. 2 vs. 3; 1 + 2 vs. 3; and 1 + 2 + 3) based on Akaike Information Criterion (AIC), and ML trees produced using PhyML 3.073 with support for nodes determined by 1000 bootstrap generations74. BI trees were produced in Mr. Bayes 3.275, based on MCMC (Markov Chain Monte Carlo) sampling, with four simultaneous runs of 10 million generations, each consisting of four chains (one cold and three hot). Bayesian posterior probabilities were defined using a 70% consensus rule, random seeds, and sampling every 100 generations. To guarantee reliability, the first 25% of trees sampled in each MCMC run were discarded as burn-in. Log-likelihood scores were then plotted in Tracer v. 1.476 to confirm the validity of the burn-in criterion. The post burn-in samples were used to construct a strict consensus tree.
Morphological analysis
All cephalopods collected from stomach contents were partly digested and lost parts of the arms and suckers. However, some were sufficiently preserved to allow some measurements and counts. Octopuses had their total weight (TW), total length (TL) dorsal mantle length (DML, measured from the posterior end to the midpoint between the eyes), mantle width (MW), funnel length (FL) free funnel length (FFL), the length (AL) and width (AW) of some arms and in males the hectocotylus (Hc), ligula length (LL) and calamus length (CL) were measured. In some cases, the diameter and number of intraovaric oocytes, the number and length of spermatophores and the number of gill lamellae could be measured or counted. All measurements were carried out following Roper and Voss77 and indices were expressed as quotients relative to the mantle length except for the free funnel index (FFI = FFL/FL) and the calamus index (CL/LL). Males with a fully developed hectocotylus and/or fully developed spermatophores in their spermatophoric sac and females with developed oviducal glands and/or developing oocytes in their ovaries were considered as mature for description purposes.
References
Roper, C. E. F, Sweeney, M. J. & Nauen, C. E. Cephalopods of the World: An Annotated Illustrated Catalogue of Species of Interest to Fisheries. FAO Fish Synopsis, 125 (1984).
Lindgren, A. R., Giribet, G. & Nishiguchi, M. K. A combined approach of the phylogeny of Cephalopoda (Mollusca). Cladistics 20, 454–486, https://doi.org/10.1111/j.1096-0031.2004.00032.x (2004).
Allcock, A. L., Hochberg, F. G., Rodhouse, P. G. K. & Thorpe, J. P. Adelieledone, a new genus of octopodid from Southern Ocean. Antartic Science 15, 415–424, https://doi.org/10.1017/S0954102003001512 (2003).
Norman, M. D., Boucher-Rodoni, R. & Hocheberg, F. G. The sharkclub octopus, Galeoctopus lateralis, a new genus and species of deep-water octopus from the Western Pacific Ocean (Cephalopoda: Octopodidae). Journal of Molluscan Studies 70, 247–256, https://doi.org/10.1093/mollus/70.3.247 (2004).
Norman, M. D. & Hochberg, F. G. The current state of octopus taxonomy. Proceedings of the International Workshop and Symposium of Cephalopod International Advisory Council, Phuket, 2003. Phuket Marine Biological Center Special Publication 66, 127–154 (2005a).
Norman, M. D., Boucher-Rodoni, R. & Hocheberg, F. G. A new genus and two new species of mesobenthic octopuses from Australia and New Caledonia. Journal of Molluscan Studies 75, 323–336, https://doi.org/10.1093/mollus/eyp027 (2009).
Jorgensen, E. M., Strugnell, J. M. & Allcock, A. L. Description and Phylogenetic relationships of a new genus of Octopus, Sasakiopus (Cephalopoda: Octopodidae), from the Bering Sea, with a redescription of Sasakiopus salebrosus (Sasaki, 1920). Journal of Molluscan Studies 76, 57–66, https://doi.org/10.1093/mollus/eyp046 (2010).
Leite, T. S., Haimovici, M., Molina, W. & Warnke, K. Morphological and genetic description of Octopus insularis, a new species in the Octopus vulgaris complex (Cephalopoda: Octopodidae) from the tropical southwestern Atlantic. Journal of the Molluscan Studies 74, 63–74, https://doi.org/10.1093/mollus/eym050 (2008).
Sales, J. B. L. et al. New molecular phylogeny of the squids of the family Loliginidae with emphasis on the genus Doryteuthis Naef, 1912: Mitochondrial and nuclear sequences indicate the presence of cryptic species in the southern Atlantic Ocean. Molecular Phylogenetics and Evolution 68, 293–299, https://doi.org/10.1016/j.ympev.2013.03.027 (2013a).
Gleadall, I. G. Octopus sinensis d’Orbigny, 1841 (Cephalopoda: Octopodidae): Valid Species Name for the commercially Valuable East Asian Common Octopus. Species Diversity 21, 31–42, https://doi.org/10.12782/sd.21.1.031 (2016).
Clarke, M. R. The role of cephalopods in the world’s oceans. Philosophical Transactions of the Royal Society B 351, 977–1112, https://doi.org/10.1098/rstb.1996.0088 (1996).
Xavier, J. C. et al. Future challenges in cephalopod research. Journal of the Marine Biological Association of the United Kingdom 95(5), 999–1015, https://doi.org/10.1017/S0025315414000782 (2015).
Santos, R. A. & Haimovici, M. Cephalopods in the trophic relations off southern Brazil. Bulletin of Marine Science 71(2), 753–770 (2002).
Veske, T. J., Vooren, C. M. & Lessa, R. P. Feeding strategy of Yellowfin tuna (Thunnus albacares), and Wahoo (Acanthocybium solandri) in the Saint Peter and Saint Paul Archipelago, Brazil. B. Inst. Pesca 29(2), 173–181 (2003).
Markaida, U. & Sosa-Nishizaki, O. Food and feeding habits of the blue shark Prionace glauca caught off Ensenada, Baja California, Mexico, with a review on its feeding. J. Mar. Biol. Associ. 90, 1–18, https://doi.org/10.1017/S0025315409991597 (2010).
Galván-Magaña, F. et al. Shark predation on cephalopods in the Mexican and Ecuadorian Pacific Ocean. Deep-Sea. Research II 95, 52–62, https://doi.org/10.1016/j.dsr2.2013.04.002 (2013).
Cherel, Y. & Hobson, A. Stable isotopes, beaks and predators: a new tool to study the trophic ecology of cephalopods, including giant and colossal squids. Proc. R. Soc. B 272, 1601–1607 (2005).
Chen, X. et al. Species identification of Ommastrephes bartramii, Dosidicus gigas, Stenoteuthis oualaniensis and Illex argentinus (Ommastrephidae) using beak morphological variables. Scientia Marina 76(3), 473–481, https://doi.org/10.3989/scimar.03408.05B (2012).
Deagle, B. E., Jarman, S. N., Pemberton, D. & Gales, N. J. Genetic Screening for Prey in the Gut Contents from a Giant Squid (Architeuthis sp.). Journal of Heredity 96(4), 417–423, https://doi.org/10.1093/jhered/esi036 (2005).
Deagle, B. E. et al. Studying Seabird Diet through Genetic Analysis of Feaces: A Case Study on Macaroni Penguins (Eudyptes chrysolophus). PLoS ONE 2(9), e831, https://doi.org/10.1371/journal.pone.0000831 (2007).
Dunn, M. R., Szabo, A., McVeagh, M. S. & Smith, P. J. The diet of deep-water sharks and the benefits of using DNA identification of prey. Deep-Sea Research I 57, 923–930, https://doi.org/10.1016/j.dsr.2010.02.006 (2010).
Hebert, P. D. N., Cywinska, A., Ball, S. L. & de Waard, J. R. Biological identifications through DNA barcodes. Proceedings of the Royal Society B: Biological Sciences 270, 313–322, https://doi.org/10.1098/rspb.2002.2218 (2003).
Lukhtanov, V. A., Sourakov, A., Zakharov, E. V. & Hebert, P. D. N. DNA barcoding Central Asian butterflies: increasing geographical dimension does not significantly reduce the success of species identification. Molecular Ecology Resources 9(5), 1302–1310, https://doi.org/10.1111/j.1755-0998.2009.02577.x (2009).
Kerr, K. C. R. et al. Comprehensive DNA barcode coverage of North American birds. Molecular Ecology Notes 7, 535–543, https://doi.org/10.1111/j.1471-8286.2007.01670.x (2007).
Kress, W. L., Wurdack, K. J., Zimmer, E. A., Weigt, L. A. & Janzen, D. H. Use of DNA barcodes to identify flowering plants. Proceedings of the National Academy of Sciences of the United States of America 102, 8369–8374, https://doi.org/10.1073/pnas.0503123102 (2005).
Dai, L., Zheng, X., Kong, L. & Li, Q. DNA barcoding analysis of Coleoidea (Mollusca: Cephalopoda) in Chinese waters. Molecular Ecology Resources 12, 437–477, https://doi.org/10.1111/j.1755-0998.2012.03118.x (2012).
Bonnaud, L., Boucher-Rodoni, R. & Monnerot, M. Phylogeny of decapod cephalopods based on partial 16S rDNA nucleotide sequences. Comptes Rendus de I’Académie des Sciences – Series III 317, 581–588 (1994).
Moura, R. L. et al. An extensive reef system at the Amazon River mouth. Science Advances 2, e1501252, https://doi.org/10.1126/sciadv.1501252 (2016).
Francini-Filho, R. B. et al. Perspectives on the Great Amazon Reef: extension, biodiversity and threats. Frontiers in Marine Science 5, 142, https://doi.org/10.3389/fmars.2018.00142 (2018).
Furtado-Ogawa, E. & Menezes, M. F. Alimentação do pargo Lutjanus purpureus Poey, no Nordeste Brasileiro. Arquivos de Ciências do Mar 12(2), 105–108 (1972).
Allen, G. R. Snappers of the world. Vol. 6.FAO Fish.Synop. 125(6), 208 (1985).
Cervigón, F. R., et al. Fichas FAO de identificación de especies para los fines de la pesca. Guía de campo de las especies comerciales marinas y de aquas salobres de la costa septentrional de Sur América. FAO, Rome. Preparado con el financiamento de la Comisión de Comunidades Europeas y de NORAD.513 p. (1992).
Duarte, L. O. & Garcia, C. B. Diet of the Mutton snapper Lutjanus analis (Cuvier) from the Gulf of Salamanca, Colombia, Caribbean Sea. Bulletin of Science 65(2), 453–465 (1999).
Sales, J. B. L. et al. Divergence of cryptic species of Doryteuthis plei Blainville, 1823 (Loliginidae, Cephalopoda) in the Western Atlantic Ocean is associated with the formation of the Caribbean Sea. Molecular Phylogenetics and Evolution 106, 44–54, https://doi.org/10.1016/j.ympev.2016.09.014. (2017).
Norman, M. D., Finn J. K. & Hochberg F. G. Family Octopodidae d´Órgigny, 1840 p: 36–201. In Jereb, P.; Roper, C. F. E., Norman, M. D. & Finn, J. K. (Eds) Cephalopods of the world. An annotated and illustrated catalogue of cephalopod species known to date. Volume 3. Octopods and Vampire Squids. FAO Species Catalogue for Fishery Purposes. No. 4, Vol. 3. Rome, FAO. 370 p. 11 colour plates.(2013).
Sales, J. B. L. et al. Phylogeographical features of Octopus vulgaris and Octopus insularis in the Southeastern Atlantic Based on the analysis of Mitochondrial Markers. Journal of Shellfish Research 32(2), 325–339, https://doi.org/10.2983/035.032.0211 (2013).
Norman, M. D. Cephalopods of the World: a world guide. Hakenhein, Germany. ConchBook. 320pp (2003).
Guerrero-Kommritz J. & Rodríguez-Bermúdez A. Soft-bottom octopods (Cephalopoda: Octopodidae) of the Southern Caribbean with the description of a new species of Macrotritopus. Journal of Marine Biodiversity. 1867–1624 (Online), https://doi.org/10.1007/s12526-018-0903-8 (2018).
Voss, G. L. A note on some Cephalopods from Brazil with a description of a new species of octopod, Eledone massyae. Bulletin of Marine Science of the Gulf and Caribbean 14(3), 511–516 (1964).
Hanlon, R. T., Watson, A. C. & Barbosa, A. A “Mimic Octopus” in the Atlantic: Flatfish Mimicry and Camouflage by Macrotritopus cf. defilippi. Biol. Bull. 218, 15–24, https://doi.org/10.1086/BBLv218n1p15 (2010).
Barroso, L. M. Regime alimentar do pargo (Lutjanus aya Bloch, 1795) no Nordeste Brasileiro. Boletim do Est. Pesca 5(3), 7–16 (1965).
Fonseca, J. F. Estudo da dieta do Lutjanus synagris (Linnaeus, 1758) e Ocyurus chrysurus (Bloch, 1791), Teleostei: Perciformes: Lutjanidae, no banco dos Abrolhos, Bahia, Brasil e pesca das principais espécies de Lutjanideos e Serranideos na Região. Master Thesis. Universidade Estadual Paulista “Julio de Mesquita Filho”, Campus Rio Claro. 159p. (2009).
Boyle, P. & Rodhouse, P. G. Cephalopods: ecology and fisheries. UK, Blackwell Science. 452 p. (2005).
Jereb, J. P. & Roper, C. F. E. Myopsid and Oegopsid squids. In Cephalopods of the world. An annotated and illustrated catalogue of cephalopod species know to date. Volume 2.FAO species Catalogue for Fishery Purposes. No. 4, Vol. 2, 605p (2010).
Young, R. E. & Tsuchiya, K. Abralia Gray 1849.Version 26 July 2009 (under construction), http://tolweb.org/Abralia/19642/2009.07.26. in The Tree of Life Web Project, http://tolweb.org/ (2009).
Hanlon, R. T., Conroy, L. A. & Forsythe, J. W. Mimicry and foraging behavior of two tropical and-flat octopus species off North Sulawesi, Indonesia. Biological Journal of the Linnean Society 93, 23–28, https://doi.org/10.1111/j.1095-8312.2007.00948.x (2008).
Norman, M. D. & Hochberg, F. G. The “mimic Octopus” (Thaumoctopus mimicus n. gen. et sp.), a new octopus from the tropical Indo-West Pacific (Cephalopoda: Octopodidae). Molluscan Research 25, 57–70 (2005b).
Hochberg, F. G., Norman, M. D. & Finn, J. Wunderpus fotogenicus n. gen. and sp., a new octopus from the shallow waters of the Indo-Malayan Archipelago (Cephalopoda: Octopodidae). Molluscan Research 26, 128–140 (2006).
Huffard, C. L. Locomotion by Abdopus aculeatus (Cephalopoda: Octopodidae): Walking the line between primary and secondary defenses. J. Exp. Biol. 209, 3697–3707, https://doi.org/10.1242/jeb.02435 (2006).
Jereb, J. P., Roper, C. F. E., Norman, M. D. & Finn, J. K. Octopods and Vampire Squids. Cephalopods of the world. An annotated and illustrated catalogue of cephalopod species know to date. Vol. 3, No. 4, FAO. 382pp (2014).
Amor, M. et al. Morphological assessment of the Octopus vulgaris species complex evaluated in light of molecular-based phylogenetic inferences. Zoologica Scripta 46(3), 275–288, https://doi.org/10.1111/zsc.12207 (2016).
Nesis, K. N. Cephalopods of the World. English Translation from Russian. Levitov, B. S. (transl.), Burgess, L, A. (Ed.). Neptune City: T.F.H. Publications, Inc. 351 pp. (1987).
Norman, M. D., Hochberg, F. G. & Boucher-Rodoni, R. A revision of the deep-water octopus genus Scaeurgus (Cephalopoda: Octopodidae) with description of three new species from the Southwest Pacific Ocean. Journal of Molluscan Studies 71, 319–337, https://doi.org/10.1093/mollus/eyi033 (2005c).
Kubodera, T. & Lu, C. C. A review of cephalopod fauna in Chinese-Japanese subtropical region. Proceedings of the 3rd and 4th Symposia on Collection Building and Natural History Studies in Asia and the Pacific Rim. National Science Museum Monographs 22, 159–171 (2002).
Guzik, M. T., Norman, M. D. & Crozier, R. H. Molecular phylogeny of benthic shallow-water octopuses (Cephalopoda: Octopodidae). Molecular Phylogenetics and Evolution 37, 235–248, https://doi.org/10.1016/j.ympev.2005.05.009 (2005).
Strugnell, J. M., Norman, M. D., Vecchione, M., Guzik, M. & Allcock, A. L. The ink sac clouds octopod evolutionary history. Hydrobiologia 725(1), 215–235, https://doi.org/10.1007/s10750-013-1517-6 (2014).
Acosta-Jofré, M. S., Sahade, R., Laudien, J. & Chiappero, M. B. A contribution to the understanding of phylogenetic relationships among species of the genus Octopus (Octopodidae: Cephalopoda). Scientia Marina 76, 311–318, https://doi.org/10.3989/scimar.03365.03B (2012).
Voss, G. L. & Ramirez, M. S. Octopus maya: a new species from the Bay of Campeche, Mexico. Bulletin of Marine Science 16, 615–625 (1966).
Levy, J. A., Haimovici, M. & Conceição, M. Genetic Evidence for two species to genus Eledone (Cephalopoda: Octopodidae) in South Brazil. Comparative Biochemistry and Physiology 90B, 275–277 (1988).
Guerra, A., Cortez, T. & Rocha, F. Redescripción del pulpo de los changos (Octopus mimus Gould, 1852) del litoral chileno-peruano (Mollusca, Cephalopoda). Iberus 17, 37–57 (1999).
Maroneze, D. M. et al. Fish as ecological tool to complement biodiversity inventories of benthic macro invertebrates. Hydrobiologia 673, 29–40, https://doi.org/10.1007/s10750-011-0747-8 (2011).
Deagle, B. E., Kirkwood, R. & Jarman, S. N. Analysis of Australian fur seal diet by pyrosequencing prey DNA in feaces. Molecular Ecology 18(9), 2022–2038, https://doi.org/10.1111/j.1365-294X.2009.04158.x (2009).
Berry, O. et al. Comparison of morphological and DNA metabarcoding analyses of diets in exploited marine fishes. Marine Ecology Progress Series 540, 167–181 (2015).
Sambrook, J. & Russel, D. Molecular Cloning: A laboratory manual. Cold Spring Harbor Laboratory Press, NY. 2344p (2001).
Palumbi, S. R. et al. The Simple Fool’s Guide t PCR. (Department of Zoological Special Publication, University of Honolulu, 1991).
Folmer, O., Black, M., Hoech, W., Lutz, R. & Vrienhock, R. DNA primers for amplification of mitochondrial Cytochrome C subunit I from diverse metazoan invertebrates. Molecular Marine Biology and Biotechnology 3, 294–299 (1994).
Thompson, J. D., Gibson, T. J., Plewniak, F., Jeanmougin, F. & Higgins, D. G. The CLUSTALX windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tool. Nucleic Acids Research 24, 4876–4882 (1997).
Hall, T. A. BioEdit: A user-friendly biological sequence alignment edit and analysis program for Windows 95/98/NT. Nucleic Acids Symposium Series 41, 95–98 (1999).
Tamura, K., Stecher, G., Peterson, D., Filipski, A. & Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis Version 6.0. Molecular Biology and Evolution 30(12), 2725–2729, https://doi.org/10.1093/molbev/mst197 (2013).
Darriba, D., Taboada, G. L., Doallo, R. & Posada, D. jModelTest 2: more models, new heuristics and parallel computing. Nature methods 9, 772, https://doi.org/10.1038/nmeth.2109 (2012).
Saitou, N. & Nei, M. The neighbor-joining method: A new method for reconstruction phylogenetic trees. Molecular Biology and Evolution 4, 406–425 (1987).
Kimura, M. A simple method for estimating evolutionary rate of base substitutions through comparative studies of nucleotide sequences. Journal of Molecular Evolution 16, 111–120 (1980).
Felsenstein, J. Confidence limits on phylogenies: an approach using the bootstrap. Evolution 39, 783–791 (1985).
Guidon, S. et al. New Algorithms and Methods to Estimate Maximum-Likelihood Phylogenies: Assessing the Performance of PhyML 3.0. Systematic Biology 59(3), 307, https://doi.org/10.1093/sysbio/syq010 (2010).
Ronquist, F. et al. MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Systematc Biology 22, 539–542, https://doi.org/10.1093/sysbio/sys029 (2012).
Rambaut, A. & Drummond, A. J. Tracer v.1.5. Available from http://beast.bio.ed.ac.uk/LogCombiner (accessed in 03 January 2018) (2007).
Roper, C. F. E. & Voss, G. L. Guidelines for taxonomic description of cephalopod species. In: Roper, C. F. E., Lu, C. C. & Hochberg, F. G. Memoirs of the National Museum of Victoria. Proceedings of the workshop on the biology and resource potential of cephalopods, Melbourne, Australia. March, 1981 no 44. 48–64 (1983).
Acknowledgements
Funding for this research was provided by CNPq (Grants 306233/2 009-6 to IS, 306233/2009-6 to HS and 303561/2015-7 to MH), FAPESPA (Grants PET0035/20 10 and APP064/20 11 to IS) and FAPESPA (PRONEX 2007 to HS). We would like the fishermen from Bragança for supplying specimens. The authors also thank Marcio de Araujo Freire and Davi de Vasconcellos Machado for their technical support. The remaining authors dedicate this work to coauthor Horacio Schneider who passed away during the final review of this manuscript and was a reference for genetic research in Brazil.
Author information
Authors and Affiliations
Contributions
J.B.d.L.S. and R.F.C.S. participated on sample collection. J.B.L.S., Y.F. and J.d.C.S.P. performed molecular analysis. M.H. performed the morphometric analysis. H.S. and I.S. provided the infrastructure necessary for producing data. J.B.d.L.S., J.S.R., M.H., N.E.A. and L.F.C.C. participated in data analysis. J.B.d.L.S., J.S.R., M.H. and N.E.A. participated in sample design and manuscript writing.
Corresponding author
Ethics declarations
Competing Interests
The authors declare no competing interests.
Additional information
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
de Luna Sales, J.B., Haimovici, M., Ready, J.S. et al. Surveying cephalopod diversity of the Amazon reef system using samples from red snapper stomachs and description of a new genus and species of octopus. Sci Rep 9, 5956 (2019). https://doi.org/10.1038/s41598-019-42464-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-019-42464-8
This article is cited by
-
Progress in the analysis of English Channel loliginid squid diets using DNA-metabarcoding techniques
Marine Biology (2024)
-
The zoogeographic regionalization for cephalopoda linked to the canary current upwelling system
Marine Biology (2024)
-
Biodiversity of octopuses in the Americas
Marine Biology (2024)
-
A new species of pygmy Paroctopus Naef, 1923 (Cephalopoda: Octopodidae): the smallest southwestern Atlantic octopod, found in sea debris
Marine Biodiversity (2021)
-
Novel insights into the evolution of genome size and AT content in mollusks
Marine Biology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.